Abstract
After allogeneic HSCT (allo-HSCT), the diversity of the intestinal microbiota significantly decreases. The changes can be rapid and are thought to be caused by chemotherapy, antibiotics, or intestinal inflammation. Most patients are exposed to prophylactic and therapeutic antibiotics during neutropenia and several patients are colonized by ESBL bacteria. We investigated the changes in gut microbiota composition in allo-HSCT, aiming at investigating if the acquisition of ESBL colonization may affect gut microbiome diversity during allo-HSCT. This was a single-center prospective pilot study. All patients consecutively admitted to the Haematological Unit of the City of Health and Science, Molinette Hospital in Turin, Italy, and undergoing allo-HSCT between August 2017 to August 2020 were enrolled in the study. Microbiome analysis on fecal samples were collected every 7 days from hospital admission to discharge and until 1 year after HSCT. 48 patients were enrolled in the study. At baseline 14 patients (29.16%) were colonized by MDR bacteria, mostly extended-spectrum beta-lactamase (ESBL)-producing gram negatives (N = 11; 78.57%). During allo-HSCT, one patient had a positive rectal swab for a carbapenemase-producing Klebsiella pneumoniae and eight patients lost the colonization during the hospital stay. Microbiota composition was compared between patients colonized by ESBL at baseline and non-colonized patients. Patients colonized by ESBL had a greater abundances of Bifidobacterium, Blautia, Clostridium, Coprococcus, L-Ruminococcus Mogibacteriaceae, Peptostreptococceae and Oscillospira, while non-colonized ESBL patients had a greater abundance of Actinomycetales, Staphylococcus and Sutterella. Moreover, microbiota composition of colonized by ESBL that retained colonization after HSCT showed an increased in abundances of Akkermansia, Dialister, Erysipelotrichaceae and Methanobrevibacter when compared with patients that become negative at rectal swabs. From a clinical perspective, the evolution of this prospective pilot study will be to investigate markers of gut barrier functions, SCFA productions and to correlate the predictivity of these parameters with risk of invasive infections and clinical outcomes in allo-HSCT population.
Similar content being viewed by others
Introduction
A substantial body of evidence has demonstrated that the use of antibiotics has long-term effects on the ecology of gut microbiota, with reductions in taxonomic diversity which can persist in most individuals1,2,3,4,5,6.
During prolonged antibiotic treatments, collateral damage to the gut microbiota can predispose to colonization with deleterious microbes and become a significant “reservoir” of multi-drug resistant (MDR) microorganisms with “nosocomial profiles” of antibiotic resistance6,7, including ESBL. Intestinal colonization mediated by MDR bacteria or fungi like Candida, has been demonstrated as a significant risk factor for the development of subsequent invasive infection, such as carbapenemase-producing Klebsiella pneumoniae (CP-Kp), ESBL Enterobacteriaceae, vancomycin resistant Enterococci (VRE) or by Candida spp.8,9,10.
Martin et al.11 and Gorrie et al.12 reported that gastrointestinal carriage of CP-Kp in hospitalized patients was associated with a greater risk of developing an invasive infection due to the same colonizing microorganism. Additionally, gut colonization by MDR bacteria has been described as an essential risk factor for invasive infections, especially in solid organ and hematopoietic stem cell transplantation (HSCT), in which the mortality rate due to invasive infections by MDR bacteria is high8,13,14.
Associations between certain bacteria and allo-HSCT outcomes have been found repeatedly: several studies showed that the commensal microorganisms residing in the gut microbiota play an essential role in the development of several complications after allo-HSCT, including bloodstream infections15,16. In the setting of allo-HSCT, ESBL represent one of the major epidemiological issues and being colonized by ESBL may lead to higher risk of translocation and invasive infections by ESBL bacteria. Moreover the colonization status may impact empiric antibiotic therapies in febrile neutropenia17,18,19.
Considering the above-presented data, we investigated the changes in gut microbiota composition after allo-HSCT focusing on the effect of ESBL colonization on gut microbiota diversity.
Material and methods
Patient recruitment and sample collection
This was a single-center prospective pilot study. All patients consecutively admitted to the Haematological Unit of the City of Health and Science, Molinette Hospital in Turin, Italy, and undergoing allo-HSCT between August 2017 to August 2020 were enrolled in the study.
Patients did not receive any antibiotic prophylaxis for neutropenia. Antiviral prophylaxis was administered in all patients with acyclovir. In patients at risk for Cytomegalovirus disseminated infection Letermovir was administered and in patients, at risk for HBV-reactivation, prophylaxis with lamivudine or entecavir or tenofovir disoproxil fumarate was administered. Primary or secondary antifungal prophylaxis was administered with azoles or micafungin according to haematological protocols for HSCT. TMP-SMX has been administered to all patients after engraftment.
Febrile neutropenia was defined as an oral temperature > 38.3 °C or two consecutive readings > 38.0 °C for 2 h and an absolute neutrophil count < 0.5 × 109/L or a count expected to fall below 0.5 × 109/L20. Febrile neutropenia was empirically treated with cefepime plus amikacin or piperacillin/tazobactam plus amikacin, except for patients colonized at baseline with CP-Kp, in which case empiric therapy with meropenem or piperacillin/tazobactam plus tigecycline was administered. GVHD prophylaxis was administered to all patients according to standard guidelines21. Acute GVHD (aGVHD) was defined as clinical symptoms typically (maculopapular erythematous rash, gastrointestinal symptoms, or cholestatic hepatitis) occurring within 100 days after HSCT or donor leukocyte infusion22.
Only grade 2–4 of aGVHD was collected. Bloodstream infections (BSI) were defined as positive blood culture, either central or peripheral, with the same organism(s) isolated from two or more blood cultures drawn on separate occasions within a 48-h period in a patient with at least one of the following signs and symptoms: fever (> 38 °C), chills, rigors, hypotension. Pneumonia was defined as clinical and radiological respiratory findings with or without positive culture from bronchoalveolar lavage or sputum.
Study procedures
For each patient enrolled in the study stool sample and rectal swab were collected every seven days (± 1) from hospital admission (T0) until 28 days (T4) after allo-HSCT. T0 was collected before conditioning regimen. Patients were followed up until 12 months after transplant and stool samples and rectal swabs were prospectively collected on days 100 (T5), 180 (T6) and 1 (T7) year after allo-HSCT.
Patients who died within 7 days from hospital admission or patients with insufficient samples (defined as less than 3 samples collected at different time points) or patients lost at follow-up (defined as patients who missed more than 1 sample collection after discharge) were excluded from the analysis.
In patients previously colonized by multi-drug resistant organism (i.e., ESBL, carbapenem resistant Enterobacteriacae -CRE- and vancomycin-resistant enterococci -VRE-), decolonization was assessed for each subsequent sample, using the criteria of at least two consecutive negative samples performed weekly. Patients were counted as decolonized according to this criterion. Decolonized patients were still monitored weekly with stool samples to detect colonization until discharge.
Microbiological samples
A proactive surveillance program to detect intestinal colonization by carbapenem resistant Enterobacteriacae (CRE), vancomycin-resistant enterococci (VRE) and ESBL have been adopted in onco-haematology wards at the time of admission on a weekly basis.
Rectal swabs were collected using the Fecal Swab™ system (Copan, Brescia, Italy) with a liquid transport medium, containing 2 ml of Modified Amies Medium that was suitable for culture. Specimens were processed by automated direct plating using Wasp® instrument (Copan, Brescia, Italy). Ten μL of the FecalSwab™ liquid medium were inoculated on Brilliance CRE medium (Oxoid Ltd, Hampshire, United Kingdom), BBL Chromagar ESBL (Becton Dickinson GmbH, Heidelberg, Germany), and VRE Brilliance agar (Oxoid), for carbapenemase resistant Gram-negative (GN), extended-spectrum beta-lactamases producing GN, and vancomycin-resistant Enterococcus isolation, respectively. Inoculated plates were incubated for 18–24 h at 35 ± 2 °C. The grown colonies were identified by MALDI-TOF analysis (Bruker DALTONIK GmbH, Bremen, Germany) and their antimicrobial susceptibility was determined by a commercial microdilution method (Microscan Walkaway 96 Plus- Beckman Coulter)23.
Antimicrobial susceptibility testing for GN and Enterococcus spp. isolates were performed using Panel 83 (Microscan) which included ESBL confirmation test and Panel 33 which included vancomycin and teicoplanin testing. Carbapenemase activity was evidenced in Enterobacterales isolates using combination disc diffusion test “Mastdiscs® combi Carba plus” (Mast Group Ltd, UK) and confirmed by genotypic assay “Xpert Carba-R” on GeneXpert platform (Cepheid, Sunnyvale, CA) The Cepheid Xpert Carba-R assay (Cepheid, Sunnyvale, CA, USA) is an automated in vitro diagnostic test for the qualitative detection of the blaKPC, blaNDM, blaVIM, blaOXA-48, and blaIMP genes. The assay is performed on the GeneXpert instrument systems and can be performed on rectal swab specimens (a validated U.S. in vitro diagnostics [US-IVD] and European Union in vitro diagnostics [EU-IVD] method) or on pure cultures of carbapenem-nonsusceptible bacterial isolates (a validated US-IVD method).
DNA extraction and amplicon sequencing
All stool samples were transported to laboratory immediately after collection and stored at − 80 °C for DNA extraction. Total DNA from faecal sample collected during the trial was extracted following the standard operating procedures (SOP’s) developed by the International Human Microbiome Standard Consortium. Total DNA was extracted by using the The QIAamp Power Fecal Pro DNA Kit (Qiagen, Milan, Italy) according to manufacturing instructions. After extraction, DNA was quantified by using the QUBIT dsHS kit, and standardized at 5 ng/uL. The V3-V4 region (16SF-5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG-3′, 16SR-5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC-3′) of the 16S rRNA gene was amplified according to Klindworth et al.24
Amplicons were then purified, tagged, and pooled according to the Illumina metagenomic guidelines [https://support.illumina.com/documents/documentation/chemistry_documentation/16s/16s-metagenomic-library-prep-guide-15044223-b.pdf]. Sequencing was performed with a MiSeq Illumina instrument (Illumina) with V2 chemistry and generated 250 bp paired-end reads according to the manufacturer’s instructions.
Bioinformatics analysis
After sequencing, paired-end reads were imported in QIIME225. Sequences were filtered for quality and for chimeric sequences using the QIIME DADA2 denoise-paired script26. Amplicon-sequence variants (ASVs) obtained were then used for taxonomic assignment against the Greengenes database v13. ASV table displays the lowest taxonomy resolution that was reached; when the taxonomy assignment was not able to reach the genus, the family name was displayed. Taxonomic assignment was checked by BLAST27.
Statistical analysis
The variables were compared in ESBL colonized and non-colonized groups using the Epi Info 7.2.5 program (CDC-Atlanta- USA). Chi-square test in StatCalc was used for univariate analysis with a significance threshold of p < 0.05.
QIIME2-diversity script was then used to calculate alpha diversity of the study population at and beta-diversity for each patient at each timepoint. Not-normally distributed variables were presented as median (interquartile range). Differences in gut microbiota were calculated by Wilcoxon matched paired test. Box plots represented the interquartile range between the first and the third quartile, with the error bars showing the lowest and the highest value.
P values were adjusted for multiple testing using the Benjamini- Hochberg procedure, which assesses the falsediscovery rate (FDR).
Ethics approval and consent to participate
Each participant gave their written informed consent to participate in the study. The study protocol was approved by the Ethical Committee of the “AOU Città Della Salute e Della Scienza” Hospital of Turin, Italy (Prot. N.0038691; August 2017).
Results
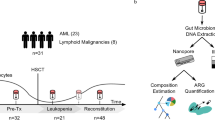
Fifty-eight patients were enrolled in the study between August 2017 and August 2020. Ten patients were subsequently excluded for early death (N = 3), insufficient number of specimens (N = 5) or loss to follow-up (N = 2). One hundred ninety-five faecal samples from 48 patients were analyzed (Table 1). The most frequent indication for allo-HSCT was acute myelogenous leukemia (AML; 29; 60%). The most common comorbidities were cardiovascular diseases (N = 10; 20.8%), COPD, diabetes mellitus and obesity (grade I or II) (Table 1). Thiotepa plus busulfan plus fludarabine was the most widely used (N = 16;33%) conditioning regimen and most of the patients (83.33%) received bone marrow from a matched unrelated donor (MUD) and received antiviral and antifungal prophylaxis as shown in Table 1. Furthermore, the most GVHD prophylactic regimen was cyclosporine plus anti-thymocyte globulin and methotrexate (47.91%, N = 23). At baseline 14 patients (29.16%) were colonized by MDR bacteria, mostly ESBL-producing gram negatives (N = 11; 78.57%; Table 1). During allo-HSCT, one patient had a positive rectal swab for a carbapenemase-producing Klebsiella pneumoniae (N = 1, KPC) and eight patients lost the colonization within 21 days after allo-HSCT (N = 7, ESBL and N = 1 VRE) (Table 1).
Around 93.75% of patients developed febrile neutropenia (N = 45) and were treated with combination antibiotic therapy, usually regimens including ceftazidime, cefepime or piperacillin/tazobactam plus amikacin, except for colonized patients who were treated according to local guidelines as described in the methods section. The most frequent antibiotics administered for febrile neutropenia are described in Table 1. About 42% of patients received carbapenems as empiric or targeted therapy during the hospital stay and thirty-three patients (68.75%) received at least one anti-anaerobe agent.
Episodes of infections during allo-HSCT
Fourteen infection episodes were observed during the hospital stay. In most cases infections were blood-stream infections (BSI; 11; 78.57%) or Clostridioides difficile infections (CDI; 4; 28.58%) (Table 1). The most frequent isolates were gram-positive bacteria (i.e., methicillin-resistant CoNS) causing BSI. Among BSI, we described two ESBL-positive strains of bacteremia and one CP-Kp BSI in one patient. Moreover, one ESBL BSI and the CP-Kp BSI occurred in patients previously colonized by MDR organisms (Table 1).
aGVHD and survival
aGVHD occurred in thirteen patients (N = 13; 27,08%) mostly skin related (N = 8) and mostly grade I-II as detailed in Table 1. Most patients (97.91%; N = 47) survived during the period of study with a follow-up of 12 months after HSCT.
Shift in gut microbiota composition during allo-HSCT
A total of 9.134.587 reads were used for data analysis with an average of 50.924 reads per sample and a sample coverage > 99%. Alpha diversity calculation (Faith’s phylogenetic diversity, Shannon index) were analyzed as a function of the overall population during allo-HSCT, from admission to 1 year after transplant however phylogenetic diversity measures does not show significant difference according to sample time (data not shown). Beta diversity based on Bray–Curtis distance matrix display differences when sample at T0 were compared with samples at T3 (ANOSIM P = 0.038). Dendrogram analysis performed on microbiota on patients showed three main clusters (Fig. 1). Cluster 1 includes Corynebacterium, Staphylococcus, Gemellaceae and Granulicatella belonging to subject 24, 20, 52, 35, 12, 27 and 29; cluster 2 mainly characterized by Dorea, L-Ruminococcus, Oscillospira, Erysipelotrichaceae and Akkermansia belonging to subject 13, 17, 52, 38, 15 and 8; cluster 3 characterized by Parabacteroides, Christensenellaceae, Butyricimonas, Methanobrevibacter, Desulfovibrio, Catenibacterium, Sutterella and Phascolarctobacterium belonging to subject 13, 15, 20, 22, 50, 29, 37, 35 and 41.
Average-linkage clustering based on the Spearman distance of fecal samples of patients undergoing allo-HSC. Rows and columns are clustered by Ward linkage hierarchical clustering.
Comparing microbiota composition across time, at the lowest taxonomic resolution reached we observed an increase in relative abundance of the class Clostridia from T1 to T2; an increase in the relative abundances of the families Enterobacteriacae and Christensenellaceae from T3 to T4, and increases in Enterococcus and of Gemellacae from T0 to T4 (Fig. 2, FDR < 0.05).
Boxplots of differentially abundant ASVs in fecal samples across patients undergoing allo-HSCT. ASVs frequency is obtained from ASV table summarized at the higher taxonomy resolution and testing procedure based on Wilcoxon matched pairs test (FDR < 0.05).
ESBL baseline colonized vs baseline negative patients
Microbiota composition was compared between patients colonized by ESBL at baseline (T0; n = 11) and non-colonized patients. We observed greater abundances of Bifidobacterium, Blautia, Clostridium, Coprococcus, L-Ruminococcus Mogibacteriaceae, Peptostreptococceae and Oscillospira in patients colonized by ESBL at baseline, compared with non-colonized (Fig. 3, FDR < 0.05). Moreover, we found that ESBL-colonized patients, there was a greater abundance of Actinomycetales, Staphylococcus and Sutterella.
Boxplots of differentially abundant ASVs in fecal samples across ESBL carriers at baseline (T0; n = 11) and non-colonized patients. ASVs frequency is obtained from ASV table summarized at the higher taxonomy resolution and testing procedure based on Wilcoxon matched pairs test (FDR < 0.05).
ESBL baseline colonized that persisted after HSCT vs patients who lost colonization during follow-up rectal swabs
Microbiota composition of colonized by ESBL that retained colonization after HSCT (T0 to T6; n = 4) showed an increased in abundances of Akkermansia, Dialister, Erysipelotrichaceae and Methanobrevibacter when compared with patients who lost colonization at rectal swab (Fig. 4A and B, FDR < 0.05). Patients who lost ESBL colonization showed an increase in the relative abundance of Actinomycetales, Arcanobacterium, Corynebacterium, Streptococcus and Lactobacillus.
(A) Boxplots of differentially abundant ASVs in fecal samples across ESBL carriers that persisted after HSCT compared to patients who lost ESBL colonization at follow-up rectal swabs. ASVs frequency is obtained from ASV table summarized at the higher taxonomy resolution and testing procedure based on Wilcoxon matched pairs test (FDR < 0.05). (B) Boxplots of differentially abundant ASVs in fecal samples across ESBL carriers that persisted after HSCT compared to patients who lost ESBL colonization at follow-up rectal swabs the. ASVs frequency is obtained from ASV table summarized at the higher taxonomy resolution and testing procedure based on Wilcoxon matched pairs test (FDR < 0.05).
Despite the low number of observations, we found a relatively higher rate of C. difficile infection complications in the ESBL-colonised group than in non-colonised patients (3 vs. 1; Table 1). Comparing the microbiota by Wilcoxon test of ESBL positive CDI-positive patients, CDI-negative patients had significantly fewer members of Bacteroides, Clostridium cluster IV, Bifidobacterium spp., Faecalibacterium prausnitzii and Prevotella in their faecal microbiota if compared with ESBL positive CDI negative group (Figure supplementary 1). The abundances of Akkermansia muciniphila, Lactobacillus, Escherichia/Klebsiella were higher in the CDI group compared with the CDI negative group (P < 0.05).
Discussion
Several data described the impact of allo-HSCT on gut microbiome, but so far, no data are available about the relevance of ESBL colonization in gut microbiome composition. The results of this explorative prospective pilot study provide preliminary insights into whether there are differences in microbiota between patients colonized with ESBL compared to those who were not.
During allo-HSCT, the diversity of the intestinal microbiota significantly decreases as anaerobic commensal taxa are lost, and the microbiota becomes dominated by potentially pathogenic organisms19,28,29,30. It has been suggested that aerobic and facultative anaerobic bacteria consume oxygen in the distal intestine and maintain hypoxia in the lumen, leading to colonisation by strict anaerobes and the production of short-chain fatty acids (SCFAs)29. Different studies have been reported the potential impact of gut microbiome composition on several complications of allo-HSCT, including the risk of BSI after HSCT31,32,33,34.
In this report, we found an overall significant increase in the abundance of Clostridia, especially from T1 to T2 – the very early period after HSCT (7–14 days after allo-HSCT)- and the highest period of immunosuppression. The abundance of Firmicutes and Clostridia has been associated with better survival in the HSCT population with less GVHD-related or relapse-related mortality and less need for GVHD treatment with systemic steroids35. Furthermore, Clostridial species, such as those in clusters IV and XIVa, are notable for generating high amounts of SCFAs (such as butyrate) and inducing T-regs may be essential for mitigating GVHD36.
In addition, we observed a significant increase in the abundance of Enterobacteriaceae and Christensenellaceae from T3 to T4 (day 21 to day 28 after allo-HSCT) and a higher abundance of Enterococcus and Gemellacae (the latter belonging to Firmicutes phylum) at T3-T4 respect to T0. This could be described because of the lower gut microbial diversity within the early post transplantation period (1–28 days), characterised by an abundance of presumptive pathogenic taxa, such as Enterobacteriaceae (i.e., Klebsiella, Escherichia) or Enterococcus37,38,39,40,41. Christensenellaceae are a recently described as a family belonging to Firmicutes and are usually significant depleted in many diseases, especially metabolic disorders, and gastrointestinal pathologies primarily inflammatory bowel disease38. However, a higher abundance of Christensenellaceae is not always associated with a healthy phenotype; there is evidence, though scarce, of a possible link between this family and a delayed gastrointestinal transit time and constipation38,42,43. Therefore, Christensenellaceae modulated colonic transit time and this may impact the colonic metabolism, leading to changes in metabolomic43.
We further explored the effect of ESBL colonization on gut microbiome diversity. Though there is some literature assessing the microbiota features of ESBL carriers, data are particularly lacking from patients with hematologic malignancy undergoing allo-HSCT. To the best of our knowledge, this is the first study to assess differences from ESBL carriers compared to non-ESBL carriers, as well as compare microbiota over time following HSCT between those with persistent colonization and those without.
In our study, baselines ESBL carriers, compared to non ESBL -colonised patients, showed a significatively higher relative abundance of several obligate anaerobes, such as Clostridium, Blautia, Ruminococcus, Coprococcus and Oscillospira. Moreover, we found that ESBL-colonized patients, had a greater abundance of the order Actinomycetales. A possible explanation for this baseline differences may be found in the previous exposure of patients to antibiotics, chemotherapy or previous hospitalization which may have modulate gut microbiota even before allo-HSCT.
Moreover, according to a number of recent investigations, the composition of the Blautia population in the gut, as well as changes in that population, are connected to a variety of parameters, including the age of the host, the location of the host, the food, the genotype, the health of the host, the disease state, and other physiological situations44. Additionally, it has been shown that this genus is involved in biotransformation and engaging in crosstalk with other bacteria that are found in the digestive tract. Despite the fact that Blautia has demonstrated a number of possible probiotic qualities, there is no thorough knowledge of this genus and this is most likely owing to the absence of a comprehensive review45,46.
In patients with persistent ESBL colonization we noticed an increased in abundances of Akkermansia, Dialister, Erysipelotrichaceae and Methanobrevibacter when compared with patients who lost the rectal ESBL colonization, in which there was increase in the relative abundance of Streptococcus and Lactobacillus. These data are in contrast with the recent matched-control study by Ducarmon et al., found no differences in diversity parameters or in relative abundance between ESBL-positive and ESBL-negative non -HSCT patients, based on bacterial species-level composition47. These differences may be related to the different design of the study and different type of population. The authors stated that asymptomatic gut carriage of ESBL-producing E. coli is not associated with an altered microbiome composition or function47. Another proof of the minor difference between EBSL-negative and ESBL-positive groups outside the haematological setting, comes from Davies et al. among a group of travellers, the diversity and community composition did not differ among travellers before or during travel in terms of ESBL-E acquisition48.
Ilett et al.49 found a lower abundance of Akkermansia muciniphila early post-aHSCT in those who developed aGVHD. It has been reported that Akkermansia has the ability to activate T-cells to support the stability of the intestinal microenvironment50. Given the above, Akkermansia has a close correlation with inflammation, but more details are needed regarding its cause51. In our study ESBL-colonized patients that retained gut colonization presented with a higher abundance of Akkermansia, theoretically increasing the stability of gut microbiome52,53. On the opposite site, Akkermansia seems to play a role in bioaccumulation of the drugs, and we can speculated that this could be one of the different factor modulating gut microbiota exposure to drugs and therefore promoting persistent colonization and increasing the risk of translocation of ESBL bacteria.
Regarding the abundances of Dialister in ESBL carriers, it is well known that Dialister, is able of generating both acetate and propionate, and the abundance of this bacterium is reduced in gut diseases such as Chron’s disease, as with other Firmicutes52. Recent studies have shown that a higher abundance of Dialister is associated with more inflammation in autoimmune articular diseases53, therefore we can consider this finding as another marker of inflammation in ESBL setting. Moreover, some reports described an increase in the abundance of Dialister in Sars-Cov2 critically ill patients colonized by MDR bacteria. Commonly, these bacteria have been recognized as opportunistic pathogens54, suggesting that the MDR colonization could derive from the presence and/or the negative effects of these microorganisms in virus-infected patients. These bacteria could be involved in the MDR colonization in these patients because they have been reported to have antibiotic resistance and have been implicated in inflammatory processes as well as SARS-CoV-2 infections.
In our series, patients that not-retained ESBL gut colonization found a higher abundance of Streptococcus spp. that in theory, from the previous studies, define a microbiome at higher risk to GVHD events in the HSCT population55,56. Increases in Streptococcus abundance was negatively associated with faecal butyrate and indole levels in Galloway-Pena study and have also been associated with acute GVHD or GVHD-related death, according to Enterococcus55. Similarly, Han et al. showed a lower abundance of Erysipelotrichaceae in the aGVHD group compared with the non-aGVHD group56.
This study had several limitations, the most important of which is that the population studied is relatively small and heterogenous, we did not investigate previous antibiotic exposure, hospitalization, chemotherapies before allo-HSCT, which may have a relevant role in the baseline composition of gut microbiota and baseline gut colonisation. Moreover, PCR on faecal samples was not performed to define loss of colonization. Despite the low numbers, we observed an increased episodes of CDI in ESBL patients, so further studies and a larger sample would be useful to deepen the analysis.
The evolution in the future of this prospective study will be to investigate markers of gut barrier functions, SCFA productions and to correlate the predictivity of these parameters with clinical outcomes and risk of infections in HSCT population. A better understanding of gut microbiota composition in the setting of MDR colonization will help clinicians to target microbiota and to personalized antibiotic treatment either in the setting of prophylaxis of febrile neutropenia and empiric therapy in special populations with high risk of mortality for severe infections.
Data availability
Sequencing data were deposited at the Sequence Read Archive of the National Center for Biotechnology Information under the BioProject accession number PRJNA1175421.
Abbreviations
- HSCT:
-
Hematopoietic stem cell transplant
- IQR:
-
Interquartile range
- BMI:
-
Body mass index
- AML:
-
Acute myeloid leukemia
- MDS:
-
Myelodisplasia
- MF:
-
Mycosis fungoides
- CML:
-
Chronic myeloid leukemia
- MM:
-
Multiple myeloma
- AA:
-
Aplastic anaemia
- CLL:
-
Chronic lymphocitic leukemia
- MUD:
-
Matched unrelated donor
- APLO:
-
Haploidentical
- TMP/SMX:
-
Trimethoprim/sulfametossaxole
- CSA:
-
Cyclosporine A
- ATG:
-
Thymoglobulin
- MMF:
-
Mycophenolate mofetil
- MTX:
-
Methotrexate
- COPD:
-
Chronic obstructive pulmonary disease
- IBD:
-
Inflammatory bowel disease
- CKD:
-
Chronic kidney disease
- ESBL:
-
Extended spectrum beta-lactamases
- VRE:
-
Vancomycin-resistant enterococcus
- KPC:
-
Klebsiella producers carbapenemases
- PTZ:
-
Piperacillin/Tazobactam
- CDI:
-
Clostridioides difficile infections
- BSI:
-
Blood-stream infection
- GVHD:
-
Graft versus host disease
- aGVHD:
-
Acute GVHD
- ESBL_colB_retain:
-
Extended-spectrum beta-lactamases colonization baseline retained
- Nocol_BL_NO_Ongoing:
-
No colonization baseline, no colonization ongoing
References
De Vos, W. M., Tilg, H., Van Hul, M. & Cani, P. D. Gut microbiome and health: Mechanistic insights. Gut 71, 1020–1032 (2022).
Gasaly, N., de Vos, P. & Hermoso, M. A. Impact of bacterial metabolites on gut barrier function and host immunity: A focus on bacterial metabolism and its relevance for intestinal inflammation. Front. Immunol. 12, 658354 (2021).
O’Toole, P. W. & Jeffery, I. B. Gut microbiota and aging. Science 350, 1214–1215 (2015).
Mohajeri, M. H. et al. The role of the microbiome for human health: From basic science to clinical applications. Eur. J. Nutr. 57, 1–14 (2018).
Weersma, R. K., Zhernakova, A. & Fu, J. Interaction between drugs and the gut microbiome. Gut 69, 1510–1519 (2020).
Iizumi, T., Battaglia, T., Ruiz, V. & Perez Perez, G. I. Gut microbiome and antibiotics. Arch. Med. Res. 48, 727–734 (2017).
Gargiullo, L., Del Chierico, F., D’Argenio, P. & Putignani, L. Gut Microbiota modulation for multidrug-resistant organism decolonization: Present and future perspectives. Front. Microbiol. 10, 1704 (2019).
Kaundal, S. et al. Impact of broad-spectrum antibiotic exposures and multidrug-resistant gram-negative bacteremia on hematopoietic cell transplantation outcomes. Transpl. Infect. Dis. 23, e13717 (2021).
Guastalegname, M., Russo, A., Falcone, M., Giuliano, S. & Venditti, M. Candidemia subsequent to severe infection due to Clostridium difficile: Is there a link?. Clin. Infect. Dis. 57, 772–774 (2013).
Corcione, S. et al. Epidemiology and risk factors for mortality in bloodstream infection by CP-Kp, ESBL-E, Candida and CDI: A single center retrospective study. Eur. J. Intern. Med. 48, 44–49 (2018).
Martin, R. M. et al. Molecular epidemiology of colonizing and infecting isolates of Klebsiella pneumoniae. mSphere 1, e00261-e316 (2016).
Gorrie, C. L. et al. Gastrointestinal carriage is a major reservoir of Klebsiella pneumoniae infection in intensive care patients. Clin. Infect. Dis. 65, 208–215 (2017).
Patriarca, F. et al. Risk factors and outcomes of infections by multidrug resistant gram-negative bacteria in patients undergoing hematopoietic stem cell transplantation. Biol. Blood Marrow Transplant. 23, 333–339 (2017).
Peric, Z. et al. Gut colonization by multidrug-resistant gram-negative bacteria is an independent risk factor for development of intestinal acute graft-versus-host disease. Biol. Blood Marrow Transplant. 23, 1221–1222 (2017).
Thomas, J., Bowe, C. & Dains, J. E. Antibiotic therapy and gastrointestinal graft-versus-host disease in the allogeneic stem cell transplantation population. J. Adv. Pract. Oncol. 13, 61–69 (2022).
Shono, Y. & van den Brink, M. R. M. Gut microbiota injury in allogeneic haematopoietic stem cell transplantation. Nat. Rev. Cancer 18, 283–295 (2018).
Devaux, C. A., Million, M. & Raoult, D. The butyrogenic and lactic bacteria of the gut microbiota determine the outcome of allogenic hematopoietic cell transplant. Front. Microbiol. 11, 1642 (2022).
Kusakabe, S. et al. Pre- and post-serial metagenomic analysis of gut microbiota as a prognostic factor in patients undergoing haematopoietic stem cell transplantation. Br. J. Haematol. 188, 438–449 (2020).
Smibert, O. C., Trubiano, J. A., Slavin, M. A. & Kwong, J. C. An infectious diseases perspective on the microbiome and allogeneic stem cell transplant. Curr. Opin. Infect. Dis. 33, 426–432 (2020).
Freifeld, A. G. et al. Clinical practice guideline for the use of antimicrobial agents in neutropenic patients with cancer: 2010 update by the Infectious Diseases Society of America. Clin. Infect. Dis. 52, 56–93 (2011).
Penack, O. et al. Prophylaxis and management of graft versus host disease after stem-cell transplantation for haematological malignancies: Updated consensus recommendations of the European Society for Blood and Marrow Transplantation. Lancet Haematol. 7, e157–e167 (2020).
Harris, A. C. et al. International, multicenter standardization of acute graft-versus-host disease clinical data collection: A report from the Mount Sinai Acute GVHD International Consortium. Biol. Blood Marrow 22, 4–10 (2016).
www.microbiome-standards.org (Last access, 1 February 2024)
Klindworth, A. et al. Evaluation of General 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1 (2013).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods. 7, 335–336 (2010).
Callahan, B. J. et al. DADA2: High resolution sample inference from amplicon data. Nat. Methods 13, 581–583 (2016).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
He, G. et al. Noninvasive measurement of anatomic structure and intraluminal oxygenation in the gastrointestinal tract of living mice with spatial and spectral EPR imaging. Proc. Natl. Acad. Sci. USA 96, 4586–4591 (1999).
Espey, M. G. Role of oxygen gradients in shaping redox relationships between the human intestine and its microbiota. Free Radic. Biol. Med. 55, 130–140 (2013).
Peled, J. U. et al. Microbiota as predictor of mortality in allogeneic hematopoietic-cell transplantation. N Engl. J. Med. 382, 822–834 (2020).
Peled, J. U. et al. Intestinal microbiota and relapse after hematopoietic-cell transplantation. J. Clin. Oncol. 35, 1650–1659 (2017).
Kumari, R., Palaniyandi, S. & Hildebrandt, G. C. Microbiome: An emerging new frontier in graft-versus-host disease. Dig. Dis. Sci. 64, 669–677 (2019).
Kelly, M. S. et al. Gut colonization preceding mucosal barrier injury bloodstream infection in pediatric hematopoietic stem cell transplantation recipients. Biol. Blood Marrow Transpl. 25, 2274–2280 (2019).
Montassier, E. et al. Pretreatment gut microbiome predicts chemotherapy-related bloodstream infection. Genome Med. 8, 49 (2016).
Schwabkey, Z. I. & Jenq, R. R. Microbiome anomalies in allogeneic hematopoietic cell transplantation. Ann. Rev. Med. 71, 137–148 (2020).
Khuat, L. T., Dave, M. & Murphy, W. J. The emerging roles of the gut microbiome in allogeneic hematopoietic stem cell transplantation. Gut Microbes 13, 1966262 (2021).
Henig, I., Yehudai-Ofir, D. & Zuckerman, T. The clinical role of the gut microbiome and fecal microbiota transplantation in allogeneic stem cell transplantation. Haematologica 106, 933–946 (2021).
Waters, J. L. & Ley, R. E. The human gut bacteria Christensenellaceae are widespread, heritable, and associated with health. BMC Biol. 17, 83 (2019).
Taur, Y. et al. The effects of intestinal tract bacterial diversity on mortality following allogeneic hematopoietic stem cell transplantation. Blood 124, 1174–1182 (2014).
Taur, Y. et al. Intestinal domination and the risk of bacteremia in patients undergoing allogeneic hematopoietic stem cell transplantation. Clin. Infect. Dis. 55, 905–914 (2012).
Stein-Thoeringer, C. K. et al. Lactose drives Enterococcus expansion to promote graft-versus-host disease. Science 366, 1143–1149 (2019).
Xu, R. et al. Dysbiosis of the intestinal microbiota in neurocritically ill patients and the risk for death. Crit. Care 23, 195 (2019).
Roager, H. et al. Colonic transit time is related to bacterial metabolism and mucosal turnover in the gut. Nat. Microbiol. 1, 16093 (2016).
Liu, X. et al. Blautia-a new functional genus with potential probiotic properties?. Gut Microbes 13(1), 1–21. https://doi.org/10.1080/19490976.2021.1875796 (2021).
Odamaki, T. et al. Age-related changes in gut microbiota composition from newborn to centenarian: A cross-sectional study. BMC Microbiol. 16(1), 1–12. https://doi.org/10.1186/s12866-016-0708-5 (2016).
Laverde Gomez, J. A. et al. Formate cross-feeding and cooperative metabolic interactions revealed by transcriptomics in co-cultures of acetogenic and amylolytic human colonic bacteria. Environ. Microbiol. 21(1), 259–271. https://doi.org/10.1111/1462-2920.14454 (2019).
Ducarmon, Q. R. et al. Gut colonisation by extended-spectrum β-lactamase-producing Escherichia coli and its association with the gut microbiome and metabolome in Dutch adults: A matched case-control study. Lancet Microbe 3, e443–e451 (2022).
Davies, M. et al. Enterobacteriaceae and Bacteroidaceae provide resistance to travel-associated intestinal colonization by multi-drug resistant Escherichia coli. Gut Microbes 14, 2060676 (2022).
Ilett, E. E. et al. Associations of the gut microbiome and clinical factors with acute GVHD in allogeneic HSCT recipients. Blood Adv. 4, 5797–5809 (2020).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. United States Am. 110, 9066–9071 (2013).
Zhu, S. et al. Combination of mesenchymal stem cell and endothelial progenitor cell infusion accelerates injured intestinal repair by regulating gut microbiota after hematopoietic cell transplantation. Transplant. Cell. Therapy. 27, 152-152.e9 (2021).
Shi, H. et al. The gut microbiome as mediator between diet and its impact on immune function. Sci. Rep. 12, 51 (2022).
Tito, R. Y. et al. Dialister as a microbial marker of disease activity in spondyloarthritis. Arthritis Rheumatol. 69, 114–121 (2017).
Murphy, E. C. & Frick, I. M. Gram-positive anaerobic cocci–commensals and opportunistic pathogens. FEMS Microbiol. Rev. 37, 520–553 (2013).
Galloway-Peña, J. R. et al. Gut microbiome signatures are predictive of infectious risk following induction therapy for acute myeloid leukemia. Clin. Infect. Dis. 71, 63–71 (2020).
Han, L. et al. A gut microbiota score predicting acute graft-versus-host disease following myeloablative allogeneic hematopoietic stem cell transplantation. Am. J. Trans. 20, 1014–1027 (2020).
Acknowledgements
We really thank Dr. D. Snydman and Dr. C. Thorpe, Tufts Medical Center, Boston, MA, USA for reviewing results and sharing comments and thoughts on this project. This project has been presented at ECCMID 2018 as oral presentation.
Funding
This study was funded by Ethical Committee funds for no-profit studies of the “A.O.U. Città Della Salute e Della Scienza” Hospital of Turin, Italy. This research was supported by EU funding within the MUR PNRR Extended Partnership initiative on Emerging Infectious Diseases (Project no. PE00000007, INF-ACT).
Author information
Authors and Affiliations
Contributions
SC, FDR, BB, AB conceived the study; GB, CDC, SC, CC, BB, IF, LB, SB, LG worked on the acquisition, analysis, and interpretation of data. TL, SC, IF, AB, LG, FDR drafted the paper. All authors substantively revised it and worked on the submitted version of the manuscrpit. All authors approved the submitted version and agreed both to be personally accountable for the author’s own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Corcione, S., Ferrocino, I., Lupia, T. et al. Influence of ESBL colonization status on gut microbiota composition during allogenic hematopoietic stem cell transplantation. Sci Rep 15, 1275 (2025). https://doi.org/10.1038/s41598-025-85128-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85128-6