Abstract
White lupin (Lupinus albus L.) is an ancient grain legume that is still undergoing improvement of domestication traits, including vernalization-responsiveness, providing frost tolerance and preventing winter flowering in autumn-sowing agriculture, and vernalization-independence, conferring drought escape by rapid flowering in spring-sowing. A recent genome-wide association study highlighted several loci significantly associated with the most contrasting phenotypes, including deletions in the promoter of the FLOWERING LOCUS T homolog, LalbFTc1, and some DArT-seq/silicoDArT loci. The present study aimed to develop and validate a versatile PCR marker array enabling molecular selection of spring- and winter-type white lupin ecotypes. Candidate DArT-seq and silicoDArT loci were transformed into cleaved amplified polymorphic sequence (CAPS) or derived CAPS markers. Developed markers, together with those previously published for LalbFTc1 INDELs and quantitative trait loci from linkage maps, were implemented for screening of white lupin germplasm panel subjected to 2-year phenotyping of phenology traits. Three DArT-seq, two silicoDArT and seven LalbFTc1 INDEL markers were positively validated, constituting a convenient PCR-based marker assay for rapid and accurate reselection of white lupin germplasm towards early flowering and thermoneutrality or late flowering and vernalization-responsiveness, as well as for tracking high genetic and phenotypic diversity within white lupin landraces, revealed in the present study.
Similar content being viewed by others
Introduction
White lupin (Lupinus albus L.) is a grain legume crop cultivated in Greece and Egypt since ancient times1,2,3. During the last centuries, this species has been domesticated and distributed into numerous countries including, among others, Italy, France, Spain, Germany, Switzerland, Poland, Ethiopia and Australia4,5,6,7,8,9. White lupin has recently been recognized as a valuable nutritional replacement for soybean protein source in livestock feeding (mostly pigs and broilers) and in aquaculture (salmonids, cyprinids and perciform fish)10,11,12,13,14,15. High protein content, moderate oil content with a desirable ratio of omega-6 to omega-3 acids as well as other specific components, such as oligosaccharides, antioxidants and non-starch carbohydrates, make white lupin also an attractive component in human nutrition16. White lupin requires less mineral fertilization than many other crops due to symbiotic nitrogen fixation and physiological adaptations to phosphorus deficiency involving proteoid root development17,18. Moreover, in rotation farming, residual nitrogen can be partially recovered by succeeding crops such as wheat (Triticum aestivum L.) and oilseed rape (Brassica napus L.)19.
Despite the relatively long history of cultivation, white lupin requires significant improvement to become a competitive crop for temperate climate agroecosystems. The major breeding efforts have been focusing lately on anthracnose resistance and low alkaloid content with significant progress achieved in deciphering the molecular components underlying these traits7,8,9,20,21. Moreover, sequence-specific markers were provided to assist molecular selection for low alkaloid content and anthracnose resistance in breeding programs7,8,20,22.
Numerous accessions of white lupin, as many other temperate plant species, are responsive to vernalization during vegetative growth phase23,24. During domestication of the species, two ecotypes were selected by breeders: a winter-type, which requires at least 2 weeks of vernalization with temperatures about 1–6 °C to induce flowering, and a spring-type, with much lower vernalization requirements, including even thermoneutral accessions23,25,26,27,28,29. As vernalization responsiveness is positively correlated with frost tolerance during winter, it is the fundamental trait underlying white lupin adaptation to autumn sowing28,30,31.
Thus, the domestication process of the species in the Mediterranean temperate climate selected winter ecotypes and autumn sowing since antiquity, as these were adapted to the winter frost events and vernalization, utilizing better the late winter and early spring humidity and precipitation rates of the particular temperate climate. Breeders have recognized these traits and driven their selection for further improvement of white lupin in ‘60 s-70s32,33. Autumn sowing is carried out in warm cropping areas such as the Mediterranean basin, Western Europe and Australia, where white lupin was imported in the late 60s34. In contrast, in colder regions of temperate climate, which include central and eastern Europe, Russia, the Northern United States and southeastern Canada, spring sowing is preferred4,5,35,36,37. Vernalization responsiveness is disadvantageous in spring types because it unnecessarily delays crop flowering in the absence of low temperature after sowing and interferes with a drought escape strategy based on early phenology38.
Recent years have witnessed unprecedented advances in molecular resources for numerous plant species, including white lupin. Such resources developed for this species include, among others, a high-density linkage map carrying sequence-based markers originating from a recombinant inbred line (RIL) mapping population descending from a cross between the early flowering vernalization-independent Ukrainian cultivar Kiev Mutant and the late flowering vernalization-responsive Ethiopian landrace P27174, two reference genome sequences, and a pangenome assembly for 40 accessions representing cultivars and landraces including Kiev Mutant and P271747,39,40,41. Quantitative trait loci (QTL) mapping in the Kiev Mutant × P27174 RIL population revealed the presence of several QTLs for flowering date related to vernalization responsiveness7. PCR-based markers anchored in those QTLs were developed and implemented for correlation analysis in a germplasm panel carrying both wild and domesticated accessions, highlighting significant marker-trait associations42. Nevertheless, for the most contrasting phenotypes (i.e. very early or very late) no single marker applicable for molecular selection was identified, probably due to different genetic backgrounds in those accessions and parental lines of the mapping population. To address this issue, a white lupin diversity panel43 carrying genotypes originating from three main climate zones (tropical, subtropical and temperate)44,45 was subjected to genome-wide association study (GWAS) of flowering time in controlled conditions (greenhouse without vernalization) and in three environments differing by intensity of vernalization, i.e. Mediterranean and subcontinental climate regions in Italy (autumn sowing) and suboceanic climate in France (spring sowing). GWAS revealed several loci significantly associated with flowering time, including newly identified deletions in the promoter region of the FLOWERING LOCUS T homolog, LalbFTc1 gene (Lalb_Chr14g0364281), and DArT-seq/silicoDArT markers matching previously published QTL regions and designating a few novel loci on other chromosomes46. That study provided also several candidate DArT-seq/silicoDArT loci associated with the most contrasting phenotypes, awaiting implementation into a PCR array.
Therefore, the present study aimed to the development a versatile PCR array for marker-assisted selection of white lupin towards spring- and winter ecotypes, including rare alleles correlated with very early or very late flowering. Single nucleotide polymorphisms (SNPs, DArT-seq markers) from recent GWAS study46 were directly transformed into PCR markers using cleaved amplified polymorphic sequence (CAPS)47 or derived CAPS (dCAPS)48 approaches, whereas presence/absence variants (silicoDArT markers) were sequenced using flanking primers and eventually transformed into allele-specific PCR or CAPS/dCAPS markers. Newly developed PCR markers together with those previously published for white lupin flowering time QTLs42,49 and LalbFTc1 gene promoter INDELs46 were validated in the set of 300 white lupin accessions and confronted with observations of plant phenology (days from sowing to floral bud emergence, start of flowering and end of flowering) from a 2-year controlled environment study.
Results
Controlled environment phenotyping highlights large variability of white lupin phenology traits
300 white lupin lines originating from several geographical localizations representative of different climatic conditions (Supplementary Table S1) were phenotyped in a greenhouse under ambient long-day photoperiod for plant phenology traits, i.e. the number of days from sowing to floral bud emergence, the start of flowering and end of flowering. To highlight differences in plant phenology between winter and spring ecotypes, neither pre-sowing nor post-sowing vernalization was applied. The mean number of days from sowing to floral bud emergence reached 57.9 ± 15.3 in 2020 year and 54.0 ± 16.1 in 2021, ranging from 36.0 ± 0.0 to 111.7 ± 0.9 days and from 33.0 ± 0.0 to 114.7 ± 3.1 days, respectively. The beginning of flowering (i.e. development of the first fully colored petal) was observed about two weeks after floral bud emergence, namely at 72.0 ± 16.5 days from sowing in the first year of the study and 69.1 ± 16.3 in the second year. The range of observed values spanned from 41.3 ± 0.5 to 127.0 ± 1.4 in 2020 year and from 40.7 ± 0.5 to 121.0 ± 0.0 in 2021. The end of flowering on the main inflorescence occurred about 10 days after flowering initiation, thus at 82.9 ± 14.1 days from sowing in 2020 year and 79.0 ± 15.3 in 2021, with the range of observed values from 62.0 ± 0.0 to 129 ± 0.0 and from 54.0 ± 1.4 to 128.0 ± 0.0, respectively. As the experiment was performed in a controlled environment, very high correlation coefficients between years were observed, reaching 0.97 for days from sowing to floral bud emergence, 0.96 for start of flowering and 0.95 for end of flowering (P-values 6.5 × 10–177, 2.4 × 10–62, and 4.3 × 10–156, respectively). A slightly decreasing value of the correlation coefficient (albeit still very high) during the progress of phenology phases may reflect the increasing influence of the environment during plant development. Observation data for studied traits and replicates are provided in Supplementary Table S2 whereas calculated mean values and standard deviation values are in Supplementary Table S3.
Heritability
A consistent genetic influence on the expression of white lupin phenology traits was observed in both greenhouse experiments, conducted in 2020 and 2021. Namely, for the year 2020, the heritability values for days from sowing to the floral bud formation, the start of flowering, and the end of flowering were estimated at 0.41, indicating a moderate genetic contribution to the phenotypic variance in these traits. Similarly, in 2021, the heritability values for these traits were uniformly estimated at 0.40. These findings suggest a stable genetic effect on flowering traits across years, which emphasizes the potential for breeding programs to enhance these characteristics in white lupin by artificial selection towards desired phenology.
Transformation of DArT-seq and silicoDArT loci into PCR markers enables precise selection of white lupin winter ecotypes
The set of 11 loci from the recent genome-wide association study46 was subjected to transformation into PCR-based markers. This set included five DArT-seq (Chr06_14434379, Chr08_12044717, Chr11_14834409, Chr13_1469866 and Chr16_572706) and six silicoDArT (Chr02_2625564_D, Chr07_16560064_D, Chr08_3090075_D, Chr11_5890565_D, Chr13_12561729_D, Chr25_4002891_D) loci (Supplementary Table S4). All DArT-seq loci were successfully transformed into CAPS47 markers using flanking PCR primers and commercially available enzymes recognizing target SNP sites. Observed agarose gel patterns of enzyme cleavage products matched those calculated in silico for both allelic phases50,51.
The procedure was more complex for silicoDArT loci, which required resequencing and search for candidate loci that could eventually underlie observed polymorphism. Such an approach yielded one allele-specific PCR (Chr13_12561729_D_PCR), one CAPS (Chr07_16560064_D_CAPS) and three derived CAPS (dCAPS)48 markers (Chr02_2625564_D_dCAPS, Chr08_3090075_D_dCAPS and Chr11_5890565_D_dCAPS). Due to the lack of DNA sequence polymorphism in the resequenced region, one silicoDArT locus, Chr25_4002891_D, remains unsolved. Besides expected products, two markers, Chr11_5890565_D_dCAPS and Chr11_14834409_CAPS, yielded additional rare absence alleles. As CAPS and dCAPS markers show codominant appearance, we were able to mark off heterozygotes. Nevertheless, heterozygosity level observed in this white lupin germplasm panel was rather low and ranged from 0–1.0% (markers Chr02_2625564_D_dCAPS, Chr08_12044717_CAPS, Chr11_5890565_D_dCAPS, Chr11_14834409_CAPS and Chr13_12561729_D_PCR) through 2.3–5.7% (markers Chr06_14434379_CAPS, Chr13_1469866_CAPS, Chr07_16560064_D_CAPS and Chr08_3090075_D_dCAPS) to 9.3% (marker Chr16_572706_CAPS). Heterozygotes usually had an intermediate phenotype as compared to lines carrying the opposite alleles in a homozygous state. The list of positively validated markers including selected restriction enzymes, the lengths of expected restriction products and allele count observed in the white lupin germplasm panel is provided in Table 1. Results of white lupin germplasm genotyping with DArT-seq and silicoDArT PCR-based markers are provided in Supplementary Table S5. Marker sequences were deposited in the public repository Zenodo under DOI https://doi.org/10.5281/zenodo.10689061.
As the set of white lupin lines used in this study majorly overlapped with those used for GWAS study46 we were able to compare segregation patterns between original DArT-seq or silicoDArT loci and their corresponding PCR-based markers. Observed patterns were identical for Chr06_14434379_CAPS and Chr08_12044717_CAPS markers, as well as very similar (more than 96% identical scores) for Chr11_14834409_CAPS, Chr13_1469866_CAPS and Chr16_572706_CAPS markers. All of those markers were based on DArT-seq loci. SilicoDArT loci revealed much lower similarity between original and PCR-based scoring, revealing 87% identity for Chr11_5890565_D_dCAPS, 77% for Chr13_12561729_D_PCR, 67% for Chr07_16560064_D_CAPS, whereas below 50% for Chr02_2625564_D_dCAPS and Chr08_3090075_D_dCAPS markers.
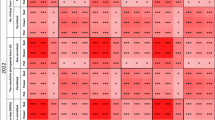
All PCR-based markers anchored in DArT-seq and silicoDArT loci except the Chr02_2625564_D_dCAPS and Chr11_5890565_D_dCAPS markers revealed statistically significant correlations between marker scores and plant phenotype for all analyzed traits in both years. The direction of the relationship between genotype and phenotype was always the same among years and traits for a particular marker. The strongest correlations were observed for Chr16_572706_CAPS (mean r value − 0.53, P-value 5.6 × 10–23), Chr07_16560064_D_CAPS (mean r value − 0.36, P-value 1.4 × 10–10), Chr13_1469866_CAPS (mean r value 0.33, P-value 3.2 × 10–9) and Chr13_12561729_D_PCR (mean r value 0.33, P-value 5.5 × 10–9) markers. The Chr02_2625564_D_dCAPS marker turned out to be selective only to two Ethiopian genotypes (minor allele frequency, MAF, 0.7%), whereas the Chr08_12044717_CAPS marker (mean r value 0.20, P-value 5.7 × 10–4) just to four very late flowering genotypes representing French winter-type cultivars Luxe and Aster (MAF 1.3%). Similarly, the Chr11_14834409_CAPS marker (mean r value 0.16, P-value 0.006) was also selective to French winter-type cultivars (Adam and Aster), however, with a 1.0% false-positive calls, including two heterozygotes. The Chr13_1469866_CAPS marker was revealed to be fairly selective for late flowering non-domesticated germplasm from Azores and Canaries, which may be beneficial in breeding programs involving those landraces. Visualization of correlation coefficients for particular marker-trait associations is provided in Fig. 1.
Correlation heatmap reporting Spearman rank correlation coefficients for each trait vs DArT-seq and silicoDArT PCR-based marker comparison. Reference alleles were coded as 0, heterozygotes as 1, whereas alternative alleles as 2. Observations were performed during 2020 and 2021 growing seasons in a greenhouse at the Institute of Plant Genetics, Polish Academy of Sciences, Poznań, Poland (52°26′ N 16°54′ E). The bar below the heatmap indicates the color legend of correlation coefficients. Asterisk (*) indicates significant correlations in the following scheme: ****, p < 0.00001; ***, 0.00001 ≤ p < 0.0001; **, 0.0001 ≤ p ≤ 0.001; *, 0.001 ≤ p ≤ 0.05.; –, non-significant.
White lupin spring ecotypes can be efficiently selected with PCR markers anchored in LalbFTc1 gene INDELs
PCR array of recently published 17 markers (Supplementary Table S4) tagging LalbFTc1 gene promoter INDELs46 was used for white lupin genotyping to supplement the set of markers developed in the present study for several DArT-seq and silicoDArT loci. MAF ranged from 3.3% to 29.7%, with five markers with MAF below 10% (PR_36a–3.3%, PR_35a–5%, PR_58b–6.3%, PR_71b–7.7% and PR_42a–8.2%). Due to the dominant type of most of the LalbFTc1 gene INDEL markers, heterozygotes were scored only for 5 markers. The level of heterozygosity ranged from 0.3% (PR_42a) to 7.0% (PR_58a). Comparing mean values, heterozygous lines for markers PR_35a and PR_71a conferred an intermediate phenotype as compared to lines carrying the opposite alleles in a homozygous state, whereas PR_58a marker heterozygotes were earlier than the corresponding homozygous lines. The list of LalbFTc1 gene promoter INDEL markers including the lengths of products for reference and alternative alleles and allele count observed in white lupin germplasm panel is provided in Table 2. Results of white lupin germplasm genotyping with LalbFTc1 gene promoter INDEL markers are provided in Supplementary Table S6.
All LalbFTc1 gene promoter INDEL markers except PR_30, PR_36a and PR_41 revealed a statistically significant correlation between marker genotype and plant phenology (Fig. 2). The direction of the relationship was the same for a particular marker in both years. The strongest correlations were observed for markers PR_71d (mean r value − 0.51, P-value 5.6 × 10–21), PR_58c (mean r value − 0.44, P-value 1.5 × 10–15), PR_36b (mean r value − 0.36, P-value 1.4 × 10–10), PR_71b (mean r value − 0.34, P-value 1.2 × 10–9), PR_42a (mean r value − 0.34, P-value 1.3 × 10–9), PR_71a (mean r value − 0.33, P-value 4.5 × 10–9) and PR_70 (mean r value − 0.30, P-value 9.6 × 10–8). As all those markers had the same sign of the correlation coefficient (negative) but differed by a proportion of reference and alternative alleles, they provide a possibility to select the set of about 20 earliest white lupin lines (markers PR_42a and PR_71b ) or direct towards late flowering germplasm carrying about 70-80 white lupin lines (PR_36b and PR_58c). Together with the set of PCR-based markers anchored in DArT-seq and silicoDArT loci (Chr16_572706_CAPS, Chr08_12044717_CAPS and Chr11_14834409_CAPS), they constitute a convenient molecular tool to select with lupin germplasm resources expressing the most contrasting phenotypes of plant phenology.
Correlation heatmap reporting Spearman rank correlation coefficients for each trait vs LalbFTc1 gene INDEL PCR marker comparison. Reference alleles were coded as 0, heterozygotes as 1, whereas alternative alleles as 2. Observations were performed during 2020 and 2021 growing seasons in a greenhouse at the Institute of Plant Genetics, Polish Academy of Sciences, Poznań, Poland (52°26′ N 16°54′ E). The bar below the heatmap indicates the color legend of correlation coefficients. Asterisk (*) indicates significant correlations in the following scheme: ****, p < 0.00001; ***, 0.00001 ≤ p < 0.0001; **, 0.0001 ≤ p ≤ 0.001; *, 0.001 ≤ p ≤ 0.05.; –, non-significant.
QTL markers from the white lupin linkage map provide lower selection efficiency than DArT-seq, silicoDArT and LalbFTc1 gene INDEL markers
White lupin linkage mapping studies reported hitherto several QTLs associated with flowering time in this species7,52, that were recently supplemented with PCR-based markers enabling their tracking in white lupin germplasm42,49. In the present study, we analyzed the polymorphism pattern of 15 QTL markers originating from those studies (Supplementary Table S4). Screening of the white lupin germplasm panel revealed that only three QTL markers had MAF values below 15%, namely QTL15 (0.3%), QTL12 (1.5%) and QTL09 (2.5%). As all those markers appeared as codominant, we were able to distinguish heterozygotes from homozygotes. The heterozygosity level for this set of markers was remarkably higher than in two other marker systems reported in the present study, reaching a mean value of 6.1%, with a range from 0.0% (QTL15) and 0.3% (QTL11) to 13.7% (QTL12) and 14.0% (QTL10). The list of QTL markers including the lengths of products for reference and alternative alleles and allele count observed in the white lupin germplasm panel is provided in Table 3. Results of white lupin germplasm genotyping with QTL markers are provided in Supplementary Table S7.
Taking into consideration correlation with plant phenology, ten QTL markers (Fig. 3) revealed significant correlation (i.e. with a P-value below 0.05), however, only one of these markers, QTL02, had a correlation coefficient value and P-value (mean r value 0.47, P-value 5.3 × 10–18) similar to those calculated for the most correlated DArT-seq, silicoDArT and LalbFTc1 gene INDEL markers. It should be clarified here, that the QTL02 marker is anchored in the LalbFTc1 gene promoter and amplifies the region carrying markers PR_70 and PR_71a-d. The two other QTL markers highly correlated with plant phenology were QTL08 (mean r value 0.28, P-value 1.2 × 10–6) and QTL10 (mean r value 0.27, P-value 1.8 × 10–6). Those markers are located in the major flowering time QTLs on chromosomes Lalb_Chr02 and Lalb_Chr13, respectively. Nevertheless, the usefulness of the QTL10 marker for marker-assisted selection may be seriously affected by excessive heterozygosity.
Correlation heatmap reporting Spearman rank correlation coefficients for each trait vs white lupin linkage map QTL PCR marker comparison. Reference alleles were coded as 0, heterozygotes as 1, whereas alternative alleles as 2. Observations were performed during 2020 and 2021 growing seasons in a greenhouse at the Institute of Plant Genetics, Polish Academy of Sciences, Poznań, Poland (52°26′ N 16°54′ E). The bar below the heatmap indicates the color legend of correlation coefficients. Asterisk (*) indicates significant correlations in the following scheme: ****, p < 0.00001; ***, 0.00001 ≤ p < 0.0001; **, 0.0001 ≤ p ≤ 0.001; *, 0.001 ≤ p ≤ 0.05.; –, non-significant.
To summarize, twelve markers, namely Chr16_572706_CAPS, PR_71d, QTL02, PR_58c, Chr07_16560064_D_CAPS, PR_36b, PR_71b, PR_42a, Chr13_1469866_CAPS, PR_71a and Chr13_12561729_D_PCR, PR_70 revealed strong correlations with plant phenology (P-value < 1 × 10–7). These markers also revealed significant correlations (P-value < 1 × 10–4) with cumulative growing degree days (GDDs) for all studied traits in both years (Supplementary Tables S8 and S9). As markers differ in allele frequencies (Tables 1, 2, 3) and direction of allelic effects (Fig. 4), they constitute a versatile tool for breeders, enabling the reselection of desired alleles in wide genetic background. In routine screening of the progeny, only a few markers from this panel need to be used, depending on the target phenotype (earliness or vernalization responsiveness) and allelic phases detected in parental lines.
Allelic effects on days from sowing to floral bud emergence (Bud), the start of flowering (Start) and the end of flowering (End) from a 2-year controlled environment study (2020 and 2021) for white lupin markers significantly correlated with plant phenology: Chr16_572706_CAPS (A), PR_71d (B), QTL02 (C), PR_58c (D), Chr07_16560064_D_CAPS (E), PR_36b (F), PR_71b (G), PR_42a (H), Chr13_1469866_CAPS (I), PR_71a (J), Chr13_12561729_D_PCR (K) and PR_70 (L). R stands for a reference allele, V is for a variant allele, whereas H is for a heterozygote. Observations were performed at the Institute of Plant Genetics, Polish Academy of Sciences, Poznań, Poland (52°26′ N 16°54′ E).
Since landraces are dynamic populations that are diverse in their genetic composition both within and between populations, we compared phenotypic variability within and between genotypes of a particular landrace with genetic diversity (Supplementary Table S10). Mean standard deviation values calculated for phenology traits ranged from 1.3 and 1.9 within genotypes and from 3.2 to 3.9 between genotypes (a, b, c, d) within landraces, whereas mean percentages of markers showing different scores between genotypes within landraces ranged from 0.0% to 42.9% for 42 PCR-based markers (mean value of 17.2%) and from 0.8% to 27.7% for 10 720 DArT-seq46 markers (mean value of 19.7). There were significant correlations of genetic diversity between PCR-based and DArT-seq markers (0.64, P-value 6.9 × 10–12), as well as between genetic diversity of both types of markers and phenotypic diversity (standard deviations) within landraces for all studied phenology traits in both years (from 0.27 to 0.37, P-values from 0.0003 to 0.0065). Therefore, a designed PCR array may also be useful to distinguish one genotype from the others within a particular landrace (Fig. 5).
Allelic composition heatmap of PCR markers that revealed the highest correlations with white lupin phenology. Reference alleles were coded as 0, heterozygotes as 1, whereas alternative alleles as 2. Observations were performed during the 2020 and 2021 growing seasons in a greenhouse at the Institute of Plant Genetics, Polish Academy of Sciences, Poznań, Poland (52°26′ N 16°54′ E). The bar below the heatmap indicates the color legend of phenological observations (days from sowing to bud emergence, start of flowering, and end of flowering) and PCR marker alleles. Only genotypes that revealed the most extreme phenology are represented on the heatmap.
Discussion
The current status of the PCR array for molecular selection of agronomic traits in white lupin
The present study highlighted moderate heritability of time to flowering in the absence of vernalization in white lupin (0.40–0.41). It highlighted quantitative inheritance of this trait in white lupin, contrary to single gene inheritance in the narrow-leafed lupin (L. angustifolius) with heritability value about 0.8153 and oligogenic inheritance in the yellow lupin (L. luteus) with reported heritability about 0.7154, whereas other white lupin studies revealed heritability about 0.60–0.6455,56. Similar heritability values were also reported in many other legume species, including, inter alia, Glycine max57,58, Medicago truncatula59,60, Vigna unguiculata61 and Cicer arietinum62. It should be noted that the occurrence of vernalization decreases heritability values for flowering time and this reduction is proportional to the length of vernalization59. Therefore, heritability values calculated for field observations with some effective vernalization days are usually lower than those obtained for a controlled environment without any vernalization. Moreover, genetic heterogeneity of landraces may result in underestimation of heritability values due to phenotypic diversity of particular genotypes within a landrace. Our study highlighted relatively high genetic diversity within landraces, resulting in 17–20% polymorphic marker scores between particular genotypes. This finding is coherent with the cross-pollination rate, estimated in white lupin as 10–30%63,64.
In theory, moderate and high heritability traits may be effectively scored by traditional phenotyping and marker-assisted selection does not provide large benefits in the means of efficiency65,66,67. Nevertheless, even in such cases, molecular markers may still be very beneficial for the selection of traits that are time-consuming in scoring the phenotypes and/or are environmentally sensitive, such as vernalization responsiveness or disease resistance68. Moreover, for the specific traits that are expressed in the adult plants (i.e. induction of flowering) molecular screening and selection of plants can be performed at the juvenile phase of growth, providing an opportunity to reduce the number of lines subjected to further crossings or seed multiplication69. The current PCR marker array applicable for white lupin MAS includes several transformed DArT-seq and silicoDArT markers for plant phenology developed and positively validated in this study, several candidate LalbFTc1 insertion-deletion polymorphism underlying vernalization independence46 also positively validated here, a few markers for anthracnose resistance elucidated from linkage mapping study7,22, albeit not confirmed in the independently controlled environment and field conditions survey9, and a PCR marker tagging a functional mutation (SNP) responsible for the low alkaloid pauper phenotype20,21. Moreover, there are several candidate SNP loci significantly associated with field-relevant anthracnose resistance awaiting transformation into PCR array8,70. Recently, several kompetitive allele-specific PCR (KASP) markers targeting the novel source of white lupin anthracnose resistance have been developed and preliminary validated71.
For a closely-related major lupin crop species, L. angustifolius, numerous loci linked to agronomic traits have been reported and mapped in the genome, including vernalization responsiveness (Ku/Julius), pod shattering (tardus and lentus), soft seededness (mollis), low alkaloid content (iucundus), white seed color (leucospermus), Phomopsis stem blight (Phr1, PhtjR) and anthracnose resistance (Lanr1, LanrBo, AnMan)72,73,74,75,76,77,78,79,80. Functional mutations were identified only for vernalization responsiveness (FLOWERING LOCUS T gene, LanFTc1) and low alkaloid content (APETALA2/ethylene response transcription factor, RAP2-7)81,82,83. Nevertheless, molecular markers for agronomic trait selection in narrow-leafed lupin breeding were developed for all major traits78,80,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99. Contrary to the statement of100 that “the vast majority of the favorable alleles at these identified QTL reside in journals on library shelves rather than in cultivars that have been improved through the introgression or selection of these favorable QTL alleles” all those loci were introduced into narrow-lupin breeding with the great aid of molecular selection101. However, it was quite feasible as all of those were single- or double-gene traits. In white lupin, the situation is different because many agronomic traits have quantitative inheritance indicating polygenic control7,8,9,23,38,42,43,102,103,104,105,106,107,108,109. For such traits, traditional selection with a set of PCR-based markers was usually ineffective110 and as such has been replaced by genomics-assisted breeding111,112.
Perspectives for genomic selection in white lupin breeding
As the cost of genotyping has significantly decreased, the use of high-density SNP arrays for genomic-enabled selection has become more feasible in modern breeding113. During the recent decade, commercially available SNP arrays have been developed for numerous plant species, including cereals, oilseeds, horticultural crops as well as major legumes such as groundnut, chickpea, pigeon pea, cowpea, common bean and soybean112,114. New genotyping methods have been consequently introduced into legume breeding and already resulted in the development of improved cultivars115,116. Lupin breeding, despite relatively advanced molecular research, was technologically backward as compared to other species, exploiting traditional PCR marker systems for selection, supplemented a few years ago with a Fluidigm nanofluidic array74,75,77,101. Just very recently, a multispecies low-cost DNA genotyping platform for chickpeas, field peas, lentils and lupins has been developed by Grains Research Development Corporation and Agriculture Victoria in Australia117. The vast majority of mentioned molecular resources are available only for the narrow-leafed lupin, with white lupin lagging behind modern breeding.
Development of reference genome assemblies for the white lupin cultivar Amiga40,41 as well as resequencing of a set of modern cultivars, landraces and wild germplasm resources39 opened possibilities for GWAS and genomic predictions. Such studies in white lupin have been already performed, targeting anthracnose resistance, drought tolerance, grain yield, yield-related morphological traits, plant height, winter survival and time to flowering8,9,46,102,103,104,118. Depending on the trait and population size reported predictive abilities in those studies ranged from about 0.28 to 0.86.
The cost-effectiveness of genomic selection in plant breeding depends on many factors, including potential genetic gain, the predictive accuracy of the selected model, cost of implementation, and, last but not least, the availability of cost-effective high-throughput SNP arrays or next-generation sequencing platforms119,120. Taking into consideration genotyping technology, two platforms were exploited in white lupin: genotyping-by-sequencing with genome complexity reduced with restriction enzyme ApeKI121 and Diversity Arrays Technology sequencing (DArT-seq)122. Both methods are commercially available and provide a large number of markers but are relatively expensive, with a cost per sample of roughly 40–50 USD, whereas newer technology based on liquid chips has reduced genotyping cost to just a few dollars per sample123,124,125. As white lupin is a minor crop species with a low global protein market when compared to other grain legumes, the genotyping cost may be a limiting factor for the implementation of genomic selection into breeding practice. At least in Poland, which in the year 2022 was the second lupin producer in the world, with 193 360 ha of harvested lupin acreage (FAOSTAT, 2024), two major breeding companies working on this species prefer PCR-based genotyping to genomic selection due to financial constraints (personal communication).
Ongoing climate change may also force reselection of germplasm towards lower vernalization responsiveness due to difficulties in fulfilling vernalization requirements. Central Europe has experienced an increase in air temperature by about 2–2.5 °C since the 1950s in the spring months March-May126. For instance, the April-June period in Poland was 2.03 °C warmer in the last decade than in the 1970s, revealing a relatively stable trend of 0.50 ± 0.13 °C per decade127. As a consequence of global warming, delayed flowering has already been observed in natural plant populations with high vernalization requirements128, as well as in winter wheat in temperate climate, resulting in yield sensitivity to vernalization variations129. German wheat breeders compensated for rising temperature in the past recent 50 years by the consecutive release of cultivars with the negative trend for the heading time (about 4 days of earliness improve per decade)130. A similar strategy can be adapted for white lupin, with the use of validated markers reported in this study, providing an opportunity for rapid, accurate and effective reselection of early flowering, thermoneutral germplasm in the progeny.
Materials and methods
Plant material
The plant material encompassed 300 genotypes: 52 accessions provided by Poznań Plant Breeding Ltd. (Wiatrowo, Poland) and 248 genotypes randomly selected from 120 accessions representing 11 landrace and 2 cultivar pools43 provided by Council for Agricultural Research and Economics (Lodi, Italy). Selection of genotypes was performed due to observed high variability of phenology within accessions during 2004–2005 growing season43. From two to four genotypes were retained per accession, including early, late and intermediate flowering time. Genotypes originate from 26 countries and differ by domestication status: 243 are landraces, 34 cultivars, 18 wild or primitive accessions and 5 breeding lines. Collection sites represent different climatic conditions such as tropical and subtropical highland (Ethiopia), cold semi-arid (Anatolia and Maghreb), dry-summer Mediterranean (sites around the Mediterranean Basin), warm-summer Mediterranean (Azores and Madeira), humid temperate (i.e. oceanic: French cultivars), temperate subcontinental (other cultivars and breeding lines). These regions diverged also by photoperiod during the juvenile phase of white lupin growth, ranging from about 9–10 h in winter sowing in northern regions of the Mediterranean Basin to 11–12 h in Ethiopia and 12–17 h in spring sowing in other regions of Europe. The list of accessions with countries/regions of origin, domestication status and germplasm donors is provided in Supplementary Table S1.
Phenotyping of plant phenology in white lupin germplasm panel
Phenotyping of plant phenology was performed in a controlled environment (greenhouse) without pre-sowing vernalization. Experiments were performed at the Institute of Plant Genetics, Polish Academy of Sciences, Poznań, Poland (52°26′ N 16°54′ E) in Poznań (western Poland) with spring sowing (19th March 2020 and 11th March 2021) and under ambient long-day photoperiod, increasing during plant cultivation from about 12 h in March to 16 h in June and July. Automatic heating was used to keep the minimum air temperature above 18 °C. The mean minimum temperature recorded in greenhouse was 18.1 °C in both years, whereas the mean maximum temperature reached 27.1 °C in 2020 and 26.7 °C in 2021. Watering was provided every second day to avoid drought stress. Potting soil was composed of TS1 medium basic rec. 085 substrate pH 6.0 (Klasmann-Deilmann GmbH, Geeste, Germany) mixed with sand in equal proportions. No artificial fertilization was provided during the experiments. Floral bud emergence was scored when the bud was visible after parting the leaves surrounding the apical meristem (inspected every second day). The start of flowering was recorded as the number of days from sowing to the observation of the first fully colored petal on the main stem, whereas the end of flowering was when half of the petals on the main inflorescence faded. Observations were made in at least three biological replicates (depending on the seed availability, from 3 to 10 plants per genotype were analyzed). Experimental design in '2020 and '2021 controlled environment phenotyping of white lupin germplasm panel is provided in Supplementary Table S11. Cumulative growing degree days (GDDs) were calculated using the formula:
where t and n are days from sowing and the total number of days from sowing to the observed phenology (floral bud emergence, start of flowering and end of flowering), Td is a daily mean temperature, whereas Tb corresponds to the base temperature of white lupin considered in this study as 3°C24,28. GDD values for fractional days were calculated on a linear scale.
Daily mean temperature was calculated using the formula:
where Tmax and Tmin are daily maximum and minimum temperatures.
DNA isolation from white lupin germplasm panel
Two young upper leaves (about 50–100 mg tissue), collected from 5-week-old plants cultivated in a greenhouse, were placed into 2 ml microcentrifuge tubes (Eppendorf, Hamburg, Germany), immediately frozen in liquid nitrogen and stored at − 80 °C. Frozen plant tissue was homogenized for 45 s at 30 rpm using TissueLyser II (Qiagen, Hilden, Germany) and two stainless steel beads (ø 5 mm, Qiagen). DNA isolation was performed with Maxwell® RSC PureFood GMO and Authentication Kit (Promega, Mannheim, Germany)131 and automated isolation station Maxwell® RSC 48 Instrument (Promega). No changes to the standard protocol were made. DNA concentration and quality were estimated using a NanoDrop 2000 (ThermoFisher Scientific, Warsaw, Poland). Automated DNA isolation protocol yielded about 95 µl of mixture with an average concentration of 1011 ± 323 ng/ µl DNA (min. 407 ng/µl, max 1998 ng/µl). Results of DNA isolation are provided in Supplementary Table S12. Three biological replicates were analyzed per genotype.
Development of PCR markers for novel loci associated with flowering time in white lupin germplasm panel
A recently published GWAS highlighted several new loci significantly associated with white lupin flowering time in a range of environments46. These loci included two types of Diversity Array Technology (DArT) sequencing data: presence/absence (dominant) markers (SilicoDArT) and standard single nucleotide polymorphism (SNP) markers (DArT-seq). SNP markers were directly transformed into PCR markers using cleaved amplified polymorphic sequence (CAPS)47 or derived CAPS (dCAPS)48 techniques. SilicoDArT markers required a more complex approach because they may represent several possible types of polymorphisms, including SNPs and small INDELs in restriction enzyme recognition sites, larger insertions/deletions in restriction fragments or methylation variation at restriction sites. Potential sequence polymorphism underlying SilicoDArT markers could be identified by Sanger sequencing, therefore, PCR primers flanking SilicoDArT loci by 200–250 bp in each direction were designed and used for PCR amplification on the DNA templates isolated from lines showing opposite allelic phases. Next, the obtained amplicons were sequenced. Based on recognized polymorphism, PCR presence/absence, CAPS or dCAPS markers were designed. Sanger sequencing was performed using BigDye® Terminator v3.1 Cycle Sequencing Kit (ThermoFisher Scientific) and 96-capillary 3730xl DNA Analyzer (Applied Biosystems, ThermoFisher Scientific) by Genomed (Warsaw, Poland).
Sequence alignments were performed using the progressive Mauve algorithm132 assuming genome collinearity. Primers were designed using Primer 3 Plus133 implemented in Geneious Prime134. Restriction enzymes for the CAPS approach were identified using SNP2CAPs50 and current (28.07.2023) REBASE update51 whereas restriction enzymes and mismatches in primers for the dCAPS approach were identified using a web-based version of dCAPS Finder 2.0 http://helix.wustl.edu/dcaps/135. The list of all designed markers with primer sequences and their coordinates in white lupin genome sequence is provided in Supplementary Table S3. To facilitate germplasm genotyping with new PCR markers, PCR with a gradient of primer annealing temperature in the range between 56 °C and 64 °C was performed first. If amplification efficiency within this range was too low to enable routine screening, a gradient between 50 °C and 58 °C was tested. Primer annealing temperature providing the strongest PCR bands without significant unspecific products or stutter bands was selected.
Genotyping of white lupin germplasm panel with PCR markers associated with flowering time
The set of PCR markers used for white lupin germplasm genotyping (Supplementary Table S3) included those developed in this study as well as those recently published for QTL loci42 and insertion-deletion polymorphism in a promoter region of white lupin FLOWERING LOCUS T homolog, LalbFTc1 gene (Lalb_Chr14g0364281)46. All PCR reactions were performed using a DNA polymerase GoTaq® Flexi (Promega), a thermal cycler Labcycler Gradient (Sensoquest, Göttingen, Germany), 96-well PCR plates (4titude, Wotton, Surrey, UK) and standard pipet tips (Neptune Scientific, San Diego, USA). Thermo Fisher Scientific and New England Biolabs (Ipswich, USA) were restriction enzyme providers. PCR amplicons and restriction products were resolved by gel electrophoresis using standard (Wide Range, Serva, Heidelberg, Germany) or high-resolution agarose (3:1, Serva) with concentration (1–3%) adjusted according to the size of the expected digestion products. Electrophoresis buffer and gels were prepared using standard TAE: Tris base, acetic acid and EDTA (Serva). To transfer samples between PCR plates and gels, an electronic expandable multichannel pipette (Matrix, Thermo Fisher Scientific) was used. Results of electrophoresis separation were visualized by in-gel SYBRSafe (ThermoFisher Scientific) staining and FastGene FAS-DIGI PRO (Nippon Genetics Europe, Düren, Germany) gel documentation system.
Data analysis
In the exploration of genetic markers associated with flowering time in white lupin, a statistical framework was employed, utilizing the capabilities of R along with its essential libraries, such as tidyverse and ggplot. The relationship between markers and flowering traits was assessed through Spearman’s rank correlation, with the cor.test function providing a means to determine the significance of these correlations, yielding p-values for the correlation coefficients.
For the analysis of greenhouse data, linear mixed models were pivotal. The model, facilitated by the SpATS package136, was articulated as:
where \(y\) denotes the response variable (flowering traits), μ represents the overall mean, \({\alpha }_{i}\) and \({\beta }_{j}\) are the random effects for row and column, respectively, \({\gamma }_{ij}\) captures the spatial trend within the greenhouse, and \({g}_{k}\) denotes the genotype effect as a random factor, \(\epsilon\) symbolizes the residual error. The spatial trend component was estimated using the SAP function137 from the SpATS package enhancing the model’s capacity to accurately reflect spatial variability. Following this, the predict function was utilized to derive breeding values (BLUP) for the response variable, ensuring an accurate estimation of genetic potential across different lines and traits. This approach was methodically applied to each dataset, segmented by year, to ensure a nuanced analysis reflective of annual variations. Heritability was estimated using the generalized broad sense heritability formula:
where ki represents the eigenvalues derived from the genetic effects model, m denotes the total number of eigenvalues, and s indicates the number of eigenvalues that are zero due to model constraints. This formula is effectively capturing both additive and non-additive genetic variances138. This approach offered a comprehensive measure of genetic influence on flowering time in white lupin embodying the analytical rigor and depth of investigation, consistent with the standards of scientific inquiry in elucidating the genetic determinants. To complement the analysis, a violin plot illustrating the distribution of flowering time across lines with different alleles of the analyzed markers was created. This visualization was achieved using the ggplot2 package in R, enhanced by the functionalities of the grid and gridExtra packages.
All methods were carried out in accordance with relevant guidelines and regulations.
Data availability
All data generated during this study are included in this published article, its Supplementary Information files and in the public repositories as follows: sequences of white lupin PCR-based markers targeting DArT-seq and silicoDArT loci significantly associated with white lupin phenology were deposited in Zenodo under DOI https://doi.org/https://doi.org/10.5281/zenodo.10689061 whereas sequence variant data in the European Variation Archive (EVA) at EMBL-EBI (project PRJNA939025, accession number ERZ16297462). Full-length agarose gel electrophoregrams for cropped gel images presented in Supplementary Figure S1 are provided in Supplementary Figure S2.
References
van der Veen, M. Consumption, trade and innovation. Exploring the botanical remains from the Roman and Islamic Ports at Quseir a-Qadim, Egypt. 313 (Africa Magna Verlag, 2012).
Kurlovich, B. S. Lupins: Geography, classification, genetic resources and breeding. 468 (Intan, 2002).
Zohary, D., Hopf, M. & Weiss, E. Domestication of Plants in the Old World. 4th edn, 264 (Oxford University Press, 2012).
Gresta, F. et al. in Lupins in European cropping systems. Legumes in cropping systems (eds D. Murphy-Bokern, F. Stoddard, & C. Watson) 88–108 (CABI Publishing, 2017).
Adhikari, K. N., Thomas, G., Diepeveen, D. & Trethowan, R. Overcoming the barriers of combining early flowering and anthracnose resistance in white lupin (Lupinus albus L.) for the Northern Agricultural Region of Western Australia. Crop Pasture Sci. 64, 914–921. https://doi.org/10.1071/cp13249 (2013).
Yeheyis, L. et al. The search for commercial sweet white lupin (Lupinus albus L.) adaptive to Ethiopian growing condition seems not successful: what should be done?. Z. Naturforsch C J Biosci. 78, 317–325. https://doi.org/10.1515/znc-2023-0033 (2023).
Książkiewicz, M. et al. A high-density consensus linkage map of white lupin highlights synteny with narrow-leafed lupin and provides markers tagging key agronomic traits. Sci. Rep. 7, 15335. https://doi.org/10.1038/s41598-017-15625-w (2017).
Alkemade, J. A. et al. Genome-wide association study reveals white lupin candidate gene involved in anthracnose resistance. Theor. Appl. Genet. 135, 1011–1024. https://doi.org/10.1007/s00122-021-04014-7 (2022).
Alkemade, J. et al. A high-throughput phenotyping tool to identify field-relevant anthracnose resistance in white lupin. Plant Dis. 105, 1719–1727. https://doi.org/10.1094/pdis-07-20-1531-re (2021).
Batterham, E. S., Andersen, L. M., Lowe, R. F. & Darnell, R. E. Nutritional value of lupin (Lupinus albus)-seed meal for growing pigs: availability of lysine, effect of autoclaving and net energy content. Br. J. Nutr. 56, 645–659. https://doi.org/10.1079/BJN19860145 (2007).
Kasprowicz-Potocka, M. et al. The effect of Lupinus albus seeds on digestibility, performance and gastrointestinal tract indices in pigs. J. Anim. Physiol. Anim. Nutr. 101, e216–e224. https://doi.org/10.1111/jpn.12594 (2017).
Pisarikova, B., Zraly, Z., Bunka, F. & Trckova, M. Nutritional value of white lupine cultivar Butan in diets for fattening pigs. Veterinární medicína 53, 124–134. https://doi.org/10.17221/1943-VETMED (2008).
Nalle, C. L., Ravindran, V. & Ravindran, G. Nutritional value of white lupins (Lupinus albus) for broilers: apparent metabolisable energy, apparent ileal amino acid digestibility and production performance. Animal 6, 579–585. https://doi.org/10.1017/S1751731111001686 (2012).
Straková, E., Všetičková, L., Kutlvašr, M., Timová, I. & Suchý, P. Beneficial effects of substituting soybean meal for white lupin (Lupinus albus, cv. Zulika) meal on the biochemical blood parameters of laying hens. Ital. J. Anim. Sci. 20, 352–358. https://doi.org/10.1080/1828051X.2021.1884006 (2021).
Szczepański, A. et al. Lupin: A promising alternative protein source for aquaculture feeds?. Aquac. Rep. 26, 101281. https://doi.org/10.1016/j.aqrep.2022.101281 (2022).
Prusinski, J. White lupin (Lupinus albus L.)—Nutritional and health values in human nutrition—A review. Czech. J. Food Sci. 35, 95–105. https://doi.org/10.17221/114/2016-CJFS (2017).
Neumann, G., Massonneau, A., Martinoia, E. & Römheld, V. Physiological adaptations to phosphorus deficiency during proteoid root development in white lupin. Planta 208, 373–382. https://doi.org/10.1007/s004250050572 (1999).
Schulze, J., Temple, G., Temple, S. J., Beschow, H. & Vance, C. P. Nitrogen fixation by white lupin under phosphorus deficiency. Ann. Bot. 98, 731–740. https://doi.org/10.1093/aob/mcl154 (2006).
Mayer, J., Buegger, F., Jensen, E. S., Schloter, M. & Heß, J. Residual nitrogen contribution from grain legumes to succeeding wheat and rape and related microbial process. Plant Soil 255, 541–554. https://doi.org/10.1023/A:1026081015076 (2003).
Rychel, S. & Książkiewicz, M. Development of gene-based molecular markers tagging low alkaloid pauper locus in white lupin (Lupinus albus L.). J. Appl. Genet. 60, 269–281. https://doi.org/10.1007/s13353-019-00508-9 (2019).
Mancinotti, D. et al. The causal mutation leading to sweetness in modern white lupin cultivars. Sci. Adv. 9, eadg8866. https://doi.org/10.1126/sciadv.adg8866 (2023).
Rychel-Bielska, S. et al. Development of PCR-based markers and whole-genome selection model for anthracnose resistance in white lupin (Lupinus albus L.). J. Appl. Genet. 61, 531–545. https://doi.org/10.1007/s13353-020-00585-1 (2020).
Adhikari, K. N., Buirchell, B. J. & Sweetingham, M. W. Length of vernalization period affects flowering time in three lupin species. Plant Breed. 131, 631–636. https://doi.org/10.1111/j.1439-0523.2012.01996.x (2012).
Huyghe, C. Winter growth of autumn-sown white lupin (Lupinus albus L.) main apex growth model. Ann. Bot. 67, 429–434. https://doi.org/10.1093/oxfordjournals.aob.a088178 (1991).
Rahman, M. & Gladstones, J. Control of lupin flower initiation by vernalization, photoperiod and temperature under controlled environment. Aust. J. Exp. Agric. 12, 638–645. https://doi.org/10.1071/ea9720638 (1972).
Clapham, W. M. & Willcott, J. B. Thermosensitivity in spring white lupin. Ann. Bot. 76, 349–357. https://doi.org/10.1006/anbo.1995.1107 (1995).
Putnam, D. H., Simmons, S. R. & Hardman, L. L. Vernalization and seeding date effects on yield and yield components of white lupin. Crop Sci. 33, 1076–1083. https://doi.org/10.2135/cropsci1993.0011183X003300050040x (1993).
Huyghe, C. & Papineau, J. Winter development of autumn sown white lupin: Agronomic and breeding consequences. Agronomie 10, 709–716. https://doi.org/10.1051/agro:19900902 (1990).
Rahman, M. & Gladstones, J. Effects of temperature and photoperiod on flowering and yield components of lupin genotypes in the field. Aust. J. Exp. Agric. 14, 205–213. https://doi.org/10.1071/EA9740205 (1974).
Annicchiarico, P. & Iannucci, A. Winter survival of pea, faba bean and white lupin cultivars across contrasting Italian locations and sowing times, and implications for selection. J. Agric. Sci. 145, 611–622. https://doi.org/10.1017/s0021859607007289 (2007).
Shield, I. F., Scott, T., Stevenson, H. J., Leach, J. E. & Todd, A. D. The causes of over-winter plant losses of autumn-sown white lupins (Lupinus albus) in different regions of the UK over three seasons. J. Agric. Sci. 135, 173–183. https://doi.org/10.1017/S0021859699008047 (2000).
Abraham, E. M. et al. The use of lupin as a source of protein in animal feeding: genomic tools and breeding approaches. Int. J. Mol. Sci. 20, 851. https://doi.org/10.3390/ijms20040851 (2019).
Zafeiriou, I. et al. Mediterranean White Lupin Landraces as a valuable genetic reserve for breeding. Plants. https://doi.org/10.3390/plants10112403 (2021).
Gladstones, J. S. Lupins in Western Australia. 1. Species and varieties. J. Dept. Agric. Western Aus. Ser. 4(10), 318–324 (1969).
López-Bellido, L., Fuentes, M., Lhamby, J. C. B. & Castillo, J. E. Growth and yield of white lupin (Lupinus albus) under Mediterranean conditions: effect of sowing date. Field Crops Res. 36, 87–94. https://doi.org/10.1016/0378-4290(94)90057-4 (1994).
Yakovenko, G. L., Lukashevisn, M. I., Ageeva, P. A., Novik, N. V. & Zakharova, M. V. Status and prospects of breeding of cultivated species of Lupin in Russia. IOP Conf. Ser. Earth Environ. Sci. 663, 012014. https://doi.org/10.1088/1755-1315/663/1/012014 (2021).
Faluyi, M. A. et al. Seed quality of sweet white lupin (Lupinus albus) and management practice in eastern Canada. Eur. J. Agron. 13, 27–37. https://doi.org/10.1016/S1161-0301(00)00057-5 (2000).
Annicchiarico, P., Romani, M. & Pecetti, L. White lupin (Lupinus albus) variation for adaptation to severe drought stress. Plant Breed. 137, 782–789. https://doi.org/10.1111/pbr.12642 (2018).
Hufnagel, B. et al. Pangenome of white lupin provides insights into the diversity of the species. Plant Biotechnol. J. 19, 2532–2543. https://doi.org/10.1111/pbi.13678 (2021).
Xu, W. et al. The genome evolution and low-phosphorus adaptation in white lupin. Nat. Commun. 11, 1069. https://doi.org/10.1038/s41467-020-14891-z (2020).
Hufnagel, B. et al. High-quality genome sequence of white lupin provides insight into soil exploration and seed quality. Nat. Commun. 11, 492. https://doi.org/10.1038/s41467-019-14197-9 (2020).
Rychel-Bielska, S. et al. Quantitative control of early flowering in white lupin (Lupinus albus L.). Int. J. Mol. Sci 22, 3856. https://doi.org/10.3390/ijms22083856 (2021).
Annicchiarico, P., Harzic, N. & Carroni, A. M. Adaptation, diversity, and exploitation of global white lupin (Lupinus albus L.) landrace genetic resources. Field Crops Res 119, 114–124. https://doi.org/10.1016/j.fcr.2010.06.022 (2010).
Parisi, S. G., Cola, G., Gilioli, G. & Mariani, L. Modeling and improving Ethiopian pasture systems. Int. J. Biometeorol. 62, 883–895. https://doi.org/10.1007/s00484-017-1492-0 (2018).
Shimabukuro, R., Tomita, T. & Fukui, K.-I. Update of global maps of Alisov’s climate classification. Prog. Earth Planet Sci. 10, 19. https://doi.org/10.1186/s40645-023-00547-1 (2023).
Rychel-Bielska, S. et al. A GWAS study highlights significant associations between a series of indels in a FLOWERING LOCUS T gene promoter and flowering time in white lupin (Lupinus albus L.). BMC Plant Biol. 24, 722 (2024).
Konieczny, A. & Ausubel, F. M. A procedure for mapping Arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. Plant J. 4, 403–410. https://doi.org/10.1046/j.1365-313X.1993.04020403.x (1993).
Neff, M. M., Neff, J. D., Chory, J. & Pepper, A. E. dCAPS, a simple technique for the genetic analysis of single nucleotide polymorphisms: experimental applications in Arabidopsis thaliana genetics. Plant J. 14, 387–392. https://doi.org/10.1046/j.1365-313X.1998.00124.x (1998).
Rychel, S., Książkiewicz, M., Tomaszewska, M., Bielski, W. & Wolko, B. FLOWERING LOCUS T, GIGANTEA, SEPALLATA and FRIGIDA homologs are candidate genes involved in white lupin (Lupinus albus L.) early flowering. Mol. Breed. 39, 43. https://doi.org/10.1007/s11032-019-0952-0 (2019).
Thiel, T., Kota, R., Grosse, I., Stein, N. & Graner, A. SNP2CAPS: a SNP and INDEL analysis tool for CAPS marker development. Nucleic Acids Res. 32, e5–e5. https://doi.org/10.1093/nar/gnh006 (2004).
Roberts, R. J., Vincze, T., Posfai, J. & Macelis, D. REBASE: a database for DNA restriction and modification: Enzymes, genes and genomes. Nucleic Acids Res. 51, D629–D630. https://doi.org/10.1093/nar/gkac975 (2022).
Phan, H. T. T., Ellwood, S. R., Adhikari, K., Nelson, M. N. & Oliver, R. P. The first genetic and comparative map of white lupin (Lupinus albus L.): Identification of QTLs for anthracnose resistance and flowering time, and a locus for alkaloid content. DNA Res. 14, 59–70. https://doi.org/10.1093/dnares/dsm009 (2007).
Taylor, C. M. et al. A multiplex PCR marker distinguishes between a series of four LanFTc1 alleles regulating flowering time in narrow-leafed lupin (Lupinus angustifolius). Plant Breed. 140, 1090–1101. https://doi.org/10.1111/pbr.12982 (2021).
Iqbal, M. M. et al. The first genetic map for yellow lupin enables genetic dissection of adaptation traits in an orphan grain legume crop. BMC Genet. 20, 68–68. https://doi.org/10.1186/s12863-019-0767-3 (2019).
Croxford, A. E., Rogers, T., Caligari, P. D. S. & Wilkinson, M. J. High-resolution melt analysis to identify and map sequence-tagged site anchor points onto linkage maps: A white lupin (Lupinus albus) map as an exemplar. New Phytol. 180, 594–607. https://doi.org/10.1111/j.1469-8137.2008.02588.x (2008).
Beyene, C. Genetic variation among white lupin (Lupinus albus L.) landraces from Northwestern and Southern Ethiopia for agronomic traits and nutrient contents of grain. J. Plant Breed. Crop Sci. 12, 156–169. https://doi.org/10.5897/JPBCS2019.0869 (2020).
Zhang, J. et al. Genome-wide association study for flowering time, maturity dates and plant height in early maturing soybean (Glycine max) germplasm. BMC Genom. 16, 217. https://doi.org/10.1186/s12864-015-1441-4 (2015).
Mao, T. et al. Association mapping of loci controlling genetic and environmental interaction of soybean flowering time under various photo-thermal conditions. BMC Genom. 18, 415. https://doi.org/10.1186/s12864-017-3778-3 (2017).
Gay, L. et al. Evolution of flowering time in a selfing annual plant: Roles of adaptation and genetic drift. Ecol. Evol. 12, e8555. https://doi.org/10.1002/ece3.8555 (2022).
Pierre, J. B., Huguet, T., Barre, P., Huyghe, C. & Julier, B. Detection of QTLs for flowering date in three mapping populations of the model legume species Medicago truncatula. Theor. Appl. Genet. 117, 609–620. https://doi.org/10.1007/s00122-008-0805-4 (2008).
Paudel, D., Dareus, R., Rosenwald, J., Muñoz-Amatriaín, M. & Rios, E. F. Genome-wide association study reveals candidate genes for flowering time in cowpea (Vigna unguiculata [L.] Walp.). Front. Genet. https://doi.org/10.3389/fgene.2021.667038 (2021).
Nabati, J. et al. Identification of diverse agronomic traits in chickpea (Cicer arietinum L.) germplasm lines to use in crop improvement. Legume Sci. 5, e167. https://doi.org/10.1002/leg3.167 (2023).
Huyghe, C. White lupin (Lupinus albus L.). Field Crops Res. 53, 147–160. https://doi.org/10.1016/S0378-4290(97)00028-2 (1997).
Brebaum, S. & Boland, G. J. Sweet white lupin: A potential crop for Ontario. Can. J. Plant Sci. 75, 841–849. https://doi.org/10.4141/cjps95-140 (1995).
Hospital, F., Moreau, L., Lacoudre, F., Charcosset, A. & Gallais, A. More on the efficiency of marker-assisted selection. Theor. Appl. Genet. 95, 1181–1189. https://doi.org/10.1007/s001220050679 (1997).
Lande, R. & Thompson, R. Efficiency of marker-assisted selection in the improvement of quantitative traits. Genetics 124, 743–756. https://doi.org/10.1093/genetics/124.3.743 (1990).
Knapp, S. J. Marker-assisted selection as a strategy for increasing the probability of selecting superior genotypes. Crop Sci. 38, 1164–1174. https://doi.org/10.2135/cropsci1998.0011183X003800050009x (1998).
Hasan, N., Choudhary, S., Naaz, N., Sharma, N. & Laskar, R. A. Recent advancements in molecular marker-assisted selection and applications in plant breeding programmes. J. Genet. Eng. Biotechnol. 19, 128. https://doi.org/10.1186/s43141-021-00231-1 (2021).
Collard, B. C. & Mackill, D. J. Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos. Trans. R. Soc. B Biol. Sci. 363, 557–572. https://doi.org/10.1098/rstb.2007.2170 (2008).
Schwertfirm, G. et al. in Breeding research for sustainable cultivation of white lupin (Lupinus albus). XVI International Lupin Conference Breeding, Cultivation and Use of Lupins for a Sustainable Agriculture—Recent Developments 47 (Rostock, Germany, 2023).
Patyi, A. et al. in Development and validation of molecular markers tagging anthracnose resistance in white lupin (Lupinus albus L.). XVI International Lupin Conference Breeding, Cultivation and Use of Lupins for a Sustainable Agriculture—Recent Developments 48 (Rostock, Germany, 2023).
Boersma, J. G. et al. Construction of a genetic linkage map using MFLP and identification of molecular markers linked to domestication genes in narrow-leafed lupin (Lupinus angustifolius L.). Cell. Mol. Biol. Lett. 10, 331–344 (2005).
Nelson, M. N. et al. The first gene-based map of Lupinus angustifolius L.-location of domestication genes and conserved synteny with Medicago truncatula. Theor. Appl. Genet. 113, 225–238. https://doi.org/10.1007/s00122-006-0288-0 (2006).
Kamphuis, L. G. et al. Transcriptome sequencing of different narrow-leafed lupin tissue types provides a comprehensive uni-gene assembly and extensive gene-based molecular markers. Plant Biotechnol. J. 13, 14–25. https://doi.org/10.1111/pbi.12229 (2015).
Hane, J. K. et al. A comprehensive draft genome sequence for lupin (Lupinus angustifolius), an emerging health food: Insights into plant-microbe interactions and legume evolution. Plant Biotechnol. J. 15, 318–330. https://doi.org/10.1111/pbi.12615 (2017).
Yang, H. et al. Draft genome sequence, and a sequence-defined genetic linkage map of the legume crop species Lupinus angustifolius L.. PLoS ONE 8, e64799. https://doi.org/10.1371/journal.pone.0064799 (2013).
Zhou, G. et al. Construction of an ultra-high density consensus genetic map, and enhancement of the physical map from genome sequencing in Lupinus angustifolius. Theor. Appl. Genet. 131, 209–223. https://doi.org/10.1007/s00122-017-2997-y (2018).
Yang, H. et al. Development of molecular markers using MFLP linked to a gene conferring resistance to Diaporthe toxica in narrow-leafed lupin (Lupinus angustifolius L.). Theor. Appl. Genet. 105, 265–270. https://doi.org/10.1007/s00122-002-0925-1 (2002).
Fischer, K. et al. Characterization and mapping of LanrBo: a locus conferring anthracnose resistance in narrow-leafed lupin (Lupinus angustifolius L.). Theor. Appl. Genet. 128, 2121–2130. https://doi.org/10.1007/s00122-015-2572-3 (2015).
Yang, H., Renshaw, D., Thomas, G., Buirchell, B. & Sweetingham, M. A strategy to develop molecular markers applicable to a wide range of crosses for marker assisted selection in plant breeding: a case study on anthracnose disease resistance in lupin (Lupinus angustifolius L.). Mol. Breed. 21, 473–483. https://doi.org/10.1007/s11032-007-9146-2 (2008).
Nelson, M. N. et al. The loss of vernalization requirement in narrow-leafed lupin is associated with a deletion in the promoter and de-repressed expression of a Flowering Locus T (FT) homologue. New Phytol. 213, 220–232. https://doi.org/10.1111/nph.14094 (2017).
Taylor, C. M. et al. INDEL variation in the regulatory region of the major flowering time gene LanFTc1 is associated with vernalization response and flowering time in narrow-leafed lupin (Lupinus angustifolius L.). Plant Cell Environ. 42, 174–187. https://doi.org/10.1111/pce.13320 (2019).
Kroc, M. et al. Transcriptome-derived investigation of biosynthesis of quinolizidine alkaloids in narrow-leafed lupin (Lupinus angustifolius L.) highlights candidate genes linked to iucundus locus. Sci. Rep. 9, 2231. https://doi.org/10.1038/s41598-018-37701-5 (2019).
Boersma, J. G., Nelson, M. N., Sivasithamparam, K. & Yang, H. Development of sequence-specific PCR markers linked to the Tardus gene that reduces pod shattering in narrow-leafed lupin (Lupinus angustifolius L.). Mol. Breed. 23, 259–267. https://doi.org/10.1007/s11032-008-9230-2 (2009).
Boersma, J. G., Buirchell, B. J., Sivasithamparam, K. & Yang, H. Development of a PCR marker tightly linked to mollis, the gene that controls seed dormancy in Lupinus angustifolius L. Plant Breed. 126, 612–616. https://doi.org/10.1111/j.1439-0523.2007.01417.x (2007).
Boersma, J. G., Buirchell, B. J., Sivasithamparam, K. & Yang, H. Development of two sequence-specific PCR markers linked to the le gene that reduces pod shattering in narrow-leafed lupin (Lupinus angustifolius L.). Genet. Mol. Biol. 30, 623–629. https://doi.org/10.1590/s1415-47572007000400020 (2007).
Boersma, J. G., Buirchell, B. J., Sivasithamparam, K. & Yang, H. Development of a sequence-specific PCR marker linked to the Ku gene which removes the vernalization requirement in narrow-leafed lupin. Plant Breed. 126, 306–309. https://doi.org/10.1111/j.1439-0523.2007.01347.x (2007).
You, M. et al. A PCR-based molecular marker applicable for marker-assisted selection for anthracnose disease resistance in lupin breeding. Cell. Mol. Biol. Lett. 10, 123–134 (2005).
Yang, H., Boersma, J. G., You, M., Buirchell, B. J. & Sweetingham, M. W. Development and implementation of a sequence-specific PCR marker linked to a gene conferring resistance to anthracnose disease in narrow-leafed lupin (Lupinus angustifolius L.). Mol. Breed. 14, 145–151. https://doi.org/10.1023/b:molb.0000038003.49638.97 (2004).
Yang, H. et al. Application of whole genome re-sequencing data in the development of diagnostic DNA markers tightly linked to a disease-resistance locus for marker-assisted selection in lupin (Lupinus angustifolius). BMC Genom. 16, 660. https://doi.org/10.1186/s12864-015-1878-5 (2015).
Yang, H. et al. Rapid development of molecular markers by next-generation sequencing linked to a gene conferring phomopsis stem blight disease resistance for marker-assisted selection in lupin (Lupinus angustifolius L.) breeding. Theor. Appl. Genet. 126, 511–522. https://doi.org/10.1007/s00122-012-1997-1 (2013).
Yang, H. et al. Application of next-generation sequencing for rapid marker development in molecular plant breeding: a case study on anthracnose disease resistance in Lupinus angustifolius L.. BMC Genom. 13, 318. https://doi.org/10.1186/1471-2164-13-318 (2012).
Li, X., Yang, H. & Yan, G. Development of a co-dominant DNA marker linked to the gene lentus conferring reduced pod shattering for marker-assisted selection in narrow-leafed lupin (Lupinus angustifolius) breeding. Plant Breed. 131, 540–544. https://doi.org/10.1111/j.1439-0523.2012.01978.x (2012).
Li, X., Buirchell, B., Yan, G. & Yang, H. A molecular marker linked to the mollis gene conferring soft-seediness for marker-assisted selection applicable to a wide range of crosses in lupin (Lupinus angustifolius L.) breeding. Mol. Breed. 29, 361–370. https://doi.org/10.1007/s11032-011-9552-3 (2012).
Li, X., Yang, H., Buirchell, B. & Yan, G. Development of a DNA marker tightly linked to low-alkaloid gene iucundus in narrow-leafed lupin (Lupinus angustifolius L.) for marker-assisted selection. Crop Pasture Sci. 62, 218–224. https://doi.org/10.1071/cp10352 (2011).
Yang, H. et al. Development of sequence-specific PCR markers associated with a polygenic controlled trait for marker-assisted selection using a modified selective genotyping strategy: a case study on anthracnose disease resistance in white lupin (Lupinus albus L.). Mol. Breed. 25, 239–249. https://doi.org/10.1007/s11032-009-9325-4 (2010).
Li, X., Renshaw, D., Yang, H. & Yan, G. Development of a co-dominant DNA marker tightly linked to gene tardus conferring reduced pod shattering in narrow-leafed lupin (Lupinus angustifolius L.). Euphytica 176, 49–58. https://doi.org/10.1007/s10681-010-0212-1 (2010).
Yang, H. & Buirchell, B. in Strategies in developing molecular markers for marker assisted selection in lupin breeding in Australia. Lupins for health and wealth, Proceedings of the 12th International Lupin Conference. (eds J. A. Palta & J. B. Berger) 267–270 (International Lupin Association).
Sweetingham, M. W., Yang, H., Buirchell, B. J., Shea, G. & Shield, I. in Resistance to rust in narrow-leafed lupin and development of molecular markers. México, where old and new world lupins meet. 11th International Lupin Conference. (eds E. van Santen & G. D. Hill) 14–16 (International Lupin Association).
Bernardo, R. Molecular markers and selection for complex traits in plants: Learning from the last 20 years. Crop Sci. 48, 1649–1664. https://doi.org/10.2135/cropsci2008.03.0131 (2008).
Książkiewicz, M. & Yang, H. in Molecular marker resources supporting the Australian lupin breeding program. The Lupin Genome (eds Karam B. Singh, Lars G. Kamphuis, & Matthew N. Nelson) 73–86 (Springer International Publishing, 2020).
Pecetti, L. et al. White lupin drought tolerance: Genetic variation, trait genetic architecture, and genome-enabled prediction. Int. J. Mol. Sci. 24, 2351. https://doi.org/10.3390/ijms24032351 (2023).
Annicchiarico, P., Nazzicari, N. & Ferrari, B. in Genetic and genomic resources in white lupin and the application of genomic selection. The Lupin Genome (eds Karam B. Singh, Lars G. Kamphuis, & Matthew N. Nelson) 139–149 (Springer International Publishing, 2020).
Annicchiarico, P. et al. Genomic prediction of grain yield in contrasting environments for white lupin genetic resources. Mol. Breed. 39, 142. https://doi.org/10.1007/s11032-019-1048-6 (2019).
Kroc, M. et al. Quantitative and qualitative analysis of alkaloids composition in the seeds of a white lupin (Lupinus albus L.) collection. Genet. Resour. Crop. Evol. 64, 1853–1860. https://doi.org/10.1007/s10722-016-0473-1 (2017).
Annicchiarico, P. & Thami-Alami, I. White lupin improvement for countries of the Mediterranean basin. Legume Perspect. 10, 12–14 (2015).
Raman, R. et al. Localisation of loci involved in resistance to Diaporthe toxica and Pleiochaeta setosa in white lupin (Lupinus albus L.). Open J. Genet. https://doi.org/10.4236/ojgen.2014.43022 (2014).
Vipin, C. A. et al. Construction of integrated linkage map of a recombinant inbred line population of white lupin (Lupinus albus L.). Breed. Sci. 63, 292–300. https://doi.org/10.1270/jsbbs.63.292 (2013).
Annicchiarico, P., Harzic, N., Huyghe, C. & Carroni, A. M. Ecological classification of white lupin landrace genetic resources. Euphytica 180, 17–25. https://doi.org/10.1007/s10681-011-0364-7 (2011).
Bernardo, R. Bandwagons I, too, have known. Theor. Appl. Genet. 129, 2323–2332. https://doi.org/10.1007/s00122-016-2772-5 (2016).
Varshney, R. K. et al. Designing future crops: Genomics-assisted breeding comes of age. Trends Plant Sci. 26, 631–649. https://doi.org/10.1016/j.tplants.2021.03.010 (2021).
Budhlakoti, N. et al. Genomic selection: a tool for accelerating the efficiency of molecular breeding for development of climate-resilient crops. Front. Genet. https://doi.org/10.3389/fgene.2022.832153 (2022).
Crossa, J. et al. Genomic selection in plant breeding: Methods, models, and perspectives. Trends Plant Sci. 22, 961–975. https://doi.org/10.1016/j.tplants.2017.08.011 (2017).
Salgotra, R. K. & Stewart, C. N. Jr. Genetic Augmentation of Legume Crops Using Genomic Resources and Genotyping Platforms for Nutritional Food Security. Plants. https://doi.org/10.3390/plants11141866 (2022).
Raina, A. et al. Editorial: Legume breeding in transition: Innovation and outlook. Front. Genet. 14, 1221551. https://doi.org/10.3389/fgene.2023.1221551 (2023).
Varshney, R. K. et al. Toward the sequence-based breeding in legumes in the post-genome sequencing era. Theor. Appl. Genet. 132, 797–816. https://doi.org/10.1007/s00122-018-3252-x (2019).
Kaur, S., Keeble-Gagnere, G., Sudheesh, S., Wong, D. & Hayden, M. Multi-species DNA chip to boost chickpea improvement. GRDC Groundcover Suppl. 167, 4–5 (2023).
Annicchiarico, P. et al. White lupin adaptation to moderately calcareous soils: Phenotypic variation and genome-enabled prediction. Plants 12, 1139. https://doi.org/10.3390/plants12051139 (2023).
Krishnappa, G. et al. Integrated genomic selection for rapid improvement of crops. Genomics 113, 1070–1086. https://doi.org/10.1016/j.ygeno.2021.02.007 (2021).
Bhat, J. A. et al. Genomic selection in the era of next generation sequencing for complex traits in plant breeding. Front. Genet. 7, 221. https://doi.org/10.3389/fgene.2016.00221 (2016).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6, e19379. https://doi.org/10.1371/journal.pone.0019379 (2011).
Kilian, A. et al. Diversity arrays technology: A generic genome profiling technology on open platforms. Methods Mol. Biol. 888, 67–89. https://doi.org/10.1007/978-1-61779-870-2_5 (2012).
Liu, Y. et al. GenoBaits Soy40K: A highly flexible and low-cost SNP array for soybean studies. Sci. China Life Sci. 65, 1898–1901. https://doi.org/10.1007/s11427-022-2130-8 (2022).
Guo, Z. et al. Development of high-resolution multiple-SNP arrays for genetic analyses and molecular breeding through genotyping by target sequencing and liquid chip. Plant Commun. 2, 100230. https://doi.org/10.1016/j.xplc.2021.100230 (2021).
Lawrie, R. D. & Massey, S. E. Agrigenomic diversity unleashed: Current single nucleotide polymorphism genotyping methods for the agricultural sciences. Appl. Biosci. 2, 565–585 (2023).
Kempf, M. Enhanced trends in spectral greening and climate anomalies across Europe. Environ. Monit. Assess. 195, 260. https://doi.org/10.1007/s10661-022-10853-8 (2023).
Rychel-Bielska, S. et al. Molecular selection of soybean towards adaptation to Central European agroclimatic conditions. J. Appl. Genet. https://doi.org/10.1007/s13353-024-00889-6 (2024).
Cook, B. I., Wolkovich, E. M. & Parmesan, C. Divergent responses to spring and winter warming drive community level flowering trends. Proc. Natl. Acad. Sci. 109, 9000–9005. https://doi.org/10.1073/pnas.1118364109 (2012).
Wu, X., Liu, H., Li, X., Tian, Y. & Mahecha, M. D. Responses of winter wheat yields to warming-mediated vernalization variations across temperate Europe. Front. Ecol. Evol. https://doi.org/10.3389/fevo.2017.00126 (2017).
Rezaei, E. E., Siebert, S., Hüging, H. & Ewert, F. Climate change effect on wheat phenology depends on cultivar change. Sci. Rep. 8, 4891. https://doi.org/10.1038/s41598-018-23101-2 (2018).
McGaughey, K. D. et al. Comparative evaluation of a new magnetic bead-based DNA extraction method from fecal samples for downstream next-generation 16S rRNA gene sequencing. PLoS ONE 13, e0202858. https://doi.org/10.1371/journal.pone.0202858 (2018).
Darling, A. C., Mau, B., Blattner, F. R. & Perna, N. T. Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 14, 1394–1403. https://doi.org/10.1101/gr.2289704 (2004).
Untergasser, A. et al. Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res. 35, W71-74. https://doi.org/10.1093/nar/gkm306 (2007).
Kearse, M. et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649. https://doi.org/10.1093/bioinformatics/bts199 (2012).
Neff, M. M., Turk, E. & Kalishman, M. Web-based primer design for single nucleotide polymorphism analysis. Trends Genet. 18, 613–615. https://doi.org/10.1016/s0168-9525(02)02820-2 (2002).
Rodríguez-Álvarez, M. X., Boer, M. P., van Eeuwijk, F. A. & Eilers, P. H. C. Correcting for spatial heterogeneity in plant breeding experiments with P-splines. Spat. Stat. 23, 52–71. https://doi.org/10.1016/j.spasta.2017.10.003 (2018).
Rodríguez-Álvarez, M. X., Lee, D.-J., Kneib, T., Durbán, M. & Eilers, P. Fast smoothing parameter separation in multidimensional generalized P-splines: The SAP algorithm. Stat. Comput. 25, 941–957. https://doi.org/10.1007/s11222-014-9464-2 (2015).
Oakey, H., Verbyla, A., Pitchford, W., Cullis, B. & Kuchel, H. Joint modeling of additive and non-additive genetic line effects in single field trials. Theor. Appl. Genet. 113, 809–819. https://doi.org/10.1007/s00122-006-0333-z (2006).
Acknowledgements
The authors would like to acknowledge National Research Institute for Agriculture, Food and Environment (INRAE) and Paolo Annicchiarico from the Council for Agricultural Research and Economics, Research Centre for Animal Production and Aquaculture for providing the landrace accessions of white lupin subjected to controlled environment phenotyping in the study. We would like to thank Jolanta Belter from the Department of Gene Structure and Function, Institute of Plant Genetics, Polish Academy of Sciences for assistance in greenhouse experiments.
Funding
This research was funded by National Science Centre, Poland (SONATINA3, 2019/32/C/NZ9/00055 and SONATA17, 2021/43/D/NZ9/00293). An open access charge was co-financed by Wroclaw University of Environmental and Life Sciences as well as by National Science Centre, Poland. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Author information
Authors and Affiliations
Contributions
Author contributions (CRediT): conceptualization, M.K., S.R.-B.; methodology, M.K., A.S., S.R.-B.; software, B.K.; validation, A.S., M.K., S.R.-B.; formal analysis, M.K., B.K.; investigation, A.S., S.R.-B., W.B.; resources, A.S., S.R.-B; data curation, A.S., B.K., M.K.; writing—original draft preparation, M.K.; writing—review and editing, A.S., S.R.-B., W.B., B.K., R.G.; visualization, A.S., M.K., B.K.; supervision, M.K., S. R.-B.; project administration, S. R.-B.; funding acquisition, S. R.-B., R.G. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Surma, A., Książkiewicz, M., Bielski, W. et al. Development and validation of PCR marker array for molecular selection towards spring, vernalization-independent and winter, vernalization-responsive ecotypes of white lupin (Lupinus albus L.). Sci Rep 15, 2659 (2025). https://doi.org/10.1038/s41598-025-86482-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-86482-1