Abstract
Atopic dermatitis (AD) is a multifactorial, chronic relapsing disease. Staphylococcus aureus is the key microbial factor in AD, linked to disease activity. However, there is limited knowledge of genomic prevalence characteristics and phenotypic features of S. aureus in AD patients in China. We investigated 108 S. aureus of AD in China and globally publicly available genome sequences of 579 S. aureus of AD. Sequence type (ST) 7, ST15 and ST188 were the major lineages in China. Genes esaC, esxB, and sea were only detected in ST7, potentially contributing to its prevalence in AD. ST188 exhibited high virulence and adhesion, possibly due to the cna gene. Phylogenetic and population structure analysis revealed that 579 strains of global AD were classified into 15 sequence clusters (SCs), with SC5, SC2, and SC7 dominating. S. aureus of Chinese AD patients was mainly distributed in SC2, SC7, and SC12. Comparative genomic highlighted genes linked to AD, including enterotoxins (seh, selk, selq, entH), adhesion genes (fnbA, fnbB, sdrD, map, fib, narH). From China and global perspectives, we analyzed S. aureus’s genomic epidemic traits, phylogeny, and population structure in AD skin. These findings contribute to understanding S. aureus-host interactions and genomic diversity in AD.
Similar content being viewed by others
Introduction
Atopic dermatitis (AD) is one of the most prevalent inflammatory skin diseases. Typically, it develops in childhood and may last until adulthood. AD is characterized by recurrent, pruritic, localized eczema, frequently with seasonal fluctuations1. AD has a complicated and multifaceted pathogenesis. It encompasses hereditary conditions, abnormalities of the epidermal barrier, alterations in immunological responses, and disturbances in the skin’s microbial equilibrium2. S. aureus was found to colonize the skin of AD patients at a higher density and abundance than that of healthy patients by culture and amplicon sequencing techniques, and to increase with increasing severity3,4. The more severe the patients’ skin lesions are, the lower the microbial diversity of the skin surface and the higher the number of S. aureus. In some cases, it may even be that only S. aureus is present. The occurrence and development of AD can be mediated by S. aureus in a variety of ways, including forming biofilms by adhering to tissue cells, secreted virulence factors, and activating the inflammatory immune response of the host5,6. The prevalence and incidence of AD have increased over the past several decades. According to the Global Burden of Disease study, there is a prevalence of 15% to 20% among children and up to 10% among adults7. Different aspects of AD, such as chronic pruritus, psychosocial distress, and inflammation, can lead to anxiety, depression, or suicidality8. A retrospective study showed that the severity of AD was linked to higher health care consumption and annual treatment expenses for patients with the condition compared to matched controls without it9.
Whole-genome sequencing (WGS) is a high-resolution genotyping tool. Through the use of whole genome sequencing for genetic description, it is feasible to reconstruct the evolutionary events that produced bacterial populations and analyze how they adapt and differentiate within their host throughout colonization. Additionally, WGS can predict potential antimicrobial resistance and virulence patterns based on complete gene banks of clinical isolates10. Current typing methods, such as multi-locus sequence typing (MLST), have been used to study the population structure of S. aureus strains with respect to their specificity for human or animal infection. The molecular characteristics of S. aureus, including MLST types and spa types, show variation depending on the time and geographic region of isolation. This variability highlights the importance of considering temporal and spatial factors in epidemiological studies of S. aureus. Whole genome sequencing and metagenomic research have demonstrated that S. aureus isolated from AD is more suited to the AD environment. Though reports of this have come from Singapore, the United States, Japan, Denmark and the United Kingdom11,12,13,14,15,16,17,18. Zhongjie Wang et al. found significant changes in gene content and functions by using WGS to compare S. aureus strains from the AD and healthy control (HC) individuals from global and local scales19. However, there is currently a lack of systematic studies on the epidemiological characteristics and phylogeny of S. aureus isolated from Chinese AD patients.
In this study, we prospectively isolated and cultured S. aureus from the skin of Chinese AD patients, and performed whole-genome sequencing and phenotypic experiments to characterize the molecular epidemiological and phenotypic characteristics. Subsequently, we included globally publicly available genomes of S. aureus isolated from AD patients to analyze the phylogeny and population structure. Through comparative genomic analyses and pan-genome-wide association study (PGWAS), we discovered several S. aureus sequence types and genes that are associated with AD. Therapeutic therapies may target the virulence genes discovered by comparative genome methods and PGWAS.
Results
Clinical characteristics and antibiotic susceptibility results of the S. aures isolates
A total of 59 skin swabs from HC were collected for isolation and culture, but only two strains of S. aureus were isolated, so they were not included in this study for follow-up. In total, 108 nonduplicate clinical S. aureus isolates were collected from patients with atopic dermatitis in a tertiary hospital in Guangzhou city between October 2019 and May 2023. Relative information is listed in Table 1. Of these, 38 strains (35.2%) came from outpatients, 70 strains came from hospitalized patients (64.8%). 84.3% (59/70) of hospitalized patients had an increase in IgE, and 80.0% (56/70) of hospitalized patients had an increase in eosinophil count (EOS). There were roughly equal numbers of strains isolated from males (53.7%, 58/108) and females (46.3%, 50/108). The median age of AD patients was 17.00 years (interquartile range: 9.25–28.75), and more than half of the S. aureus strains (53.7%, 58/108) were isolated from patients aged 18 years and younger. Fifty-five patients had a combination of other allergic dermatoses such as food allergy, asthma, and allergic rhinitis. Additionally, 22 patients had co-occurring conditions such as folliculitis, erythroderma, and skin infections (Table S1).
Most strains were susceptible to a wide range of antibiotics. In particular, none of strains was resistant to teicoplanin, vancomycin, daptomycin and ceflorin (Fig. S1). Penicillin G resistance was the most common among all isolates; 82.4% (89/108) were resistant to this beta-lactam antibiotic. In addition, there was a high rate of resistance to erythromycin (37.0%, 40/108) and clindamycin (19.4%, 21/108). Thirteen strains (12.04%, 13/108) of S. aureus were resistant to cefoxitin and/or oxacillin and were categorized as MRSA. In total, 25.0% (27/108) of the S. aureus isolated were multidrug-resistant (MDR). The prevalence of MDR in MRSA strains (100%, 13/13) was significantly higher than that in MSSA strains (14.74%, 14/95). There were significant differences in MDR between MRSA and MSSA isolates (Table 1).
Molecular types of S. aureus isolates
108 S. aureus strains were typed according to MLST, spa-, agr- and SCCmec-typing (Table S2). The S. aureus was assigned to 33 distinct sequence types (STs), of which one was never reported before. ST7 (13.89%, 15/108), ST15 (13.89%, 15/108) ST188 (11.11%, 12/108) and ST2990 (6.48%, 7/108) were most prevalent. ST15 (17.14%, 12/70), ST7 (12.86%, 9/70), and ST188 (12.86%, 9/70) were prevalent among inpatients, while ST7 (15.80%, 6/38) and ST72 (13.16%, 5/38) were prevalent among outpatients. A total of 56 spa types were identified, with the top of two prevalent spa types of t91 (17.43%, 19/108) and t189 (10.09%, 11/108). The most prevalent ST188-t189 (9.26%, 10/108), ST7-t91 (8.33%, 9/108), and ST2990-t91 (6.48%, 7/108) combinations were found while combining ST type with spa typing. Notably, all ST2990 S. aureus isolates were t91 (100%, 7/7). In contrast, some STs included variable spa types. The S. aureus isolates of the major ST15 clonotype were composed of at least 8 spa types, including t84 (26.67%, 4/15), t85 (26.67%, 4/15), t335 (6.675%, 1/15), t360 (6.675%, 1/15), t385 (6.675%, 1/15), t5766 (6.675%, 1/15), t10647 (6.675%, 1/15), t14014(6.675%, 1/15), and nondetectable (6.675%, 1/15). ST72 S. aureus isolates included four spa types (t148, t3169, 9131 and nondetectable). ST88 S. aureus isolates included five spa types (t1376, t2526, t2855, t2894 and t4431). agr-I represented the most frequent agr type, which was found in 56.48% (61/108) of the strains. A lower prevalence was observed for the agr-II and agr-III, which composed 28.70% (31/108) and 13.89% (15/108) of the strains, respectively. While no agr-IV strains were isolated and one strain was nondetectable. A total of 13 MRSA strains were isolated in this study, and four SCCmec types were identified as type IVa (23.08%, 3/13), IVc (23.08%, 3/13), IVg (7.7%, 1/13), and V (7.7%, 1/13), while five strains were not identified.
Virulence gene and antimicrobial resistance gene profiles of the S. aureus isolates
Using VFDB, a total of 81 virulence factor genes (VFGs) caried by S. aureus were identified (Fig. 1). Most VFGs belonging to the categories of adhesion, enzymes, immune evasion, secretion system, and toxins. Most VFGs had more than 90% detection rates, but eta, isdD, lukS-PV, seb, sed, she, and selq genes had detection rates of less than 10%. Immune evasion cluster (IEC) genes are located on bacteriophages. Bacteriophages are mobile genetic elements that can be transferred between strains20. This gene cluster improves the capacity of S. aureus to evade the human immune response. IEC genes were found in 83.33% (90/108) of isolates. There were five types in IEC system: type A (sea-sak-chp-scn, n = 7), type B (sak-chp-scn, n = 29), type C (chp-scn, n = 18), type D (sea-sak-scn, n = 26), type E (sak-scn, n = 10) (Fig. S2).
Phylogenetic analysis and patterns of the presence of virulence factor genes (VFGs) and antimicrobial resistance genes (ARGs). Maximum likelihood (ML) tree showing the phylogenetic structure of 108 S. aureus isolates from atopic dermatitis (AD). The ML tree was constructed based on the core-genome alignment of 36,744 SNP sites. AMR, antimicrobial resistance.
To explore the prevalence differences among the major types of ST7, ST15 and ST188, we statistically analyzed their VFGs. S. aureus isolates of the predominant STs showed differential carriage of immune evasion gene chp, adherence gene cna, secretion system genes esxB and esaC, enzyme gene sak, and toxin gene sea (Fig. 2A). Notably, sak was detected in ST7 and ST188 with a 100% detection rate, whereas it was detected in ST15 with a 0% detection rate; sea was not detected in ST15 and ST188, with an 86.67% detection rate in ST7 (P < 0.05); the cna gene was not detected in both ST7 and ST15, but it was partially detected in ST188 (P < 0.05); esxB and esaC was only detected in ST7 with a 100% detection rate. The most common IEC type in ST7 (86.67%, 13/15) was Type D. In ST188, type B constituted the majority (75%, 9/12), in contrast to ST15, when type C made up 100% (15/15).
Antimicrobial resistance genes (ARGs) were consistent with the results of antibiotic susceptibility characteristic (Fig. 1). The strains under investigation exhibited a high level of \(\beta\)-lactam, macrolide and phosphomycin resistance genes. The \(\beta\)-lactam resistance genes consisted mainly of blaZ (71.30%, 77/108) and mecA (7.41%, 8/108). The macrolide resistance gene included erm(A) (4.63%, 5/108), erm(B) (5.56%, 6/108), erm(C) (21.30%, 23/108), erm(T) (0.93%, 1/108), lnu(A) (19.44%, 21/108), lnu(B) (2.78%, 3/108), lnu(G) (3.70%, 4/108), mph(C) (1.85%, 2/108) and msr(A) (1.85%,2/108). The presence of phosphomycin resistance gene fosD were 43.52% (47/108).
Comparison of adhesion capacity and virulence of major STs of S. aureus
The adherence of S. aureus to human skin is probably an important step in the pathogenesis of AD patients. We thus compared the adherence ability to HaCaT cells with major STs (ST7, ST15 and ST188) and Newman. The results indicated that ST188 isolates showed a significantly higher adherence ability than Newman, ST7 and ST15 (P < 0.05) (Fig. 2B).
To compare the differences in virulence among the major STs, we randomly selected four S. aureus strains from each of the ST7, ST15, and ST188 strains to infect the larvae of the G. mellonella. In G. mellonella larvae models, the survival of the G. mellonella infected with ST15 was significantly reduced (P = 0.006, 0.004) in comparison with the Newman and ST7 (Fig. 2C). The survival of the G. mellonella infected with ST188 was also significantly reduced (P = 0.003, P < 0.001) in comparison with the Newman and ST7 (Fig. 2C).
Comparison of major STs of S. aureus isolates. (A) Differentiated VFGs in major STs. (B) Adhesion of S. aureus to human immortalized keratinocytes (HaCaTs). Colony counts of adhesive and internalized bacteria on/in HaCaT cells after infection. n = 3. The statistical significance was measured by the one-way analysis of variance (ANOVA). (C) Kaplan–Meier survival plots of G. mellonella larvae infected with major STs of S. aureus. Larvae were injected with 104 CFU of S. aureus. Survival was analysed by Kaplan–Meier survival plots and the log-rank test. Log-rank test results are as follows: Newman and ST7: P = 0.5157, Newman and ST15: P = 0.0055, Newman and ST188: P = 0.0003, ST7 and ST15: P = 0.0004, ST7 and ST188: P < 0.0001, ST15 and ST188: P = 0.2127. Data are shown from 3 independent experiments using 10 larvae per group for each S. aureus.
Phylogenetic analysis of S. aureus isolates of AD based on cgSNPs
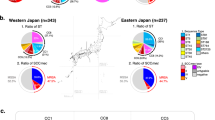
To explore the evolution of S. aureus of AD patients and characterize the population structure, we constructed a phylogenetic tree with our 108 isolates and 471 isolates of AD patients from NCBI SRA database (Fig. 3, Table S2). Using hierBAPS analysis, the maximum likelihood (ML) phylogenetic tree of 579 samples consisted of 15 sequence clusters (SCs). The prevalent SC was SC5 (14.68%, 85/579), which was mainly composed by ST188, and SC2 (11.92%, 69/579) mainly comprised ST6, ST12, ST72, ST88 and ST2990 and SC7 (11.57%, 67/579) consisted primarily of ST15 and ST582. Although phylogeny and population structure agreed well, it’s interesting to see that SC2 was divided into six portions in the phylogenetic tree. It contained strains from 6 countries and 19 different ST types. It is divided into 6 sections in the phylogenetic tree, SC2A mainly contains ST6 and ST12 and ST72, SC2B mainly contains ST88 and ST2990. S. aureus isolates from AD was diverse and sequence clusters were not regionally specific. Most Chinese S. aureus isolates were concentrated in SC2, SC7 and SC12 (Fig. 3).
Phylogenetic analysis and population structure of 579 S. aureus isolated from AD patients. Maximum likelihood phylogeny of 579 S. aureus isolated from AD patients is constructed on the whole-genome alignment of 92035 cgSNP sites. Pie plots represent the source distribution of isolates for each SC. Dominant multilocus sequence typing types for each SC are labelled. SC sequence cluster.
Comparative genomic analysis and PGWAS of S. aureus between AD patients and HC
The S. aureus was assigned to 61 distinct STs in 579 isolates from AD patients. ST188 (13.64%, 79/579), ST8 (9.33%, 54/579) and ST1 (8.81%, 51/579) were most prevalent. To further analyze the characteristics of S. aureus genome from AD and HC, we included 339 S. aureus genomes of HC. These isolates were assigned to 42 distinct STs. ST8 (12.68%, 43/339), ST188 (11.80%, 40/339) and ST15 (8.85%, 30/339) were prevalent. Comparing the STs in healthy and AD, we found that STs, including S. aureus ST1, ST7, and ST582 were significantly enriched in AD as compared with HC (P < 0.05) (Fig. 4A). And S. aureus ST30, ST6, ST508, ST630 and ST22 were significantly enriched in HC (P < 0.05). In addition, ST582 was detected only in AD skin. It is noteworthy that 9 of the 10 ST582 strains originated from Denmark, showing a clear geographical specificity. To illustrate the distribution of ST582 in AD patients and healthy controls from various countries, more strains must be included for analysis. A total of 71 STs and 243 spa types were identified in all 918 isolates. The top three prevalent STs were ST188 (12.96%, 119/918), ST8 (10.57%, 97/918) and ST15 (8.71%, 80/918). The top three prevalent spa types were t189 (11.00%, 101/918), t84 (3.92%, 36/918) and t127 (3.81%, 35/918). When combining ST type and spa typing, the most common combinations identified were ST188-t189 (10.46%, 96/918), ST1-t127 (3.38%, 31/918) and ST8-t8 (3.05%, 28/918). Overall agr typing was generally consistent with this study, with agr-I to IV accounting for 59.15%, 22.00%, 16.45% and 3.29%, respectively.
In addition, we compared the differences in VFGs and ARGs between S. aureus from AD and HC. We found significant differences in the carriage of few genes between strains from AD and HC. Strains from AD had considerably enriched cap8I, cap8J, cap8K, coa, ebp, essC, fnbA, fnbB, lukF-PV, sdrD, seh, selk, and selq, while cap8D, cap8E, cap8F, cap8G, tsst-1, and vWbp were significantly enriched in strains from HC (P < 0.05) (Fig. 4B). blaZ, fosD, erm(A), ant (9)-Ia, blaTEM-116, and aph (3’)-IIa were more frequently detected in strains from HC, whereas aac(6’)-aph(2”), fusC, aadD, mecA, tet(K), and lnu(A) were more frequently detected in strains from AD (P < 0.05) (Fig. 4C).
Comparative genomic analysis of S. aureus between AD patients and healthy controls. (A) Comparison of the proportion of different ST types in S. aureus. (B) Comparison of the proportion of VFGs in S. aureus. (C) Comparison of the proportion of ARGs in S. aureus. The statistical significance was measured by the chi-square test.
To systematically identify specific gene of S. aureus associated with AD, we performed a pan-genome-wide association study (PGWAS) of 918 isolates using Scoary and with 196 genes more prevalent in S. aureus from AD (Benjamini-Hochberg p-value < 0.05) (Fig. 5, Table S3). We identified genes involved in adhesion and biofilm formation including map, fib, and narH, highlighting the importance of keratinocyte attachment in pathogenesis. We also found genes shown to be necessary for virulence, immune evasion, and adaptation to toxic chemicals and environmental stressors-for example, aur, selX, sigA, ssl4, ssl7, isaB, setC, gltB and entH. Despite being enriched in AD, these genes had limited specificity and sensitivity, indicating that they could not be employed as markers to distinguish between S. aureus in HC and AD. The COG functional categories of differential genes belonged mainly to defense mechanisms (8.11%) and amino acid transport and metabolism (7.03%) and replication, recombination and repair (7.03%). A total of 79 hits (42.70%) had no known COG annotation or were classified as function unknown (category S).
Pan-genome-wide analysis results of accessory gene for S. aureus isolates from AD and HC. Volcano plot of different accessory genes between AD and HC. The y-axis represents the -log10 (Benjamini_H_p) values derived from the genome-wide association analysis. The x-axis represents the accessory gene prevalence in AD minus the accessory gene prevalence in HC. Blue dots indicate differential genes with higher prevalence in HC. Red dots indicate differential genes with higher prevalence in AD. P value was adjusted using the Benjamini-Hochberg method.
Discussion
Skin dysbiosis is an important factor in AD. Clinical studies have shown a link between S. aureus and AD3,4. However, the exact roles played by S. aureus in vivo are not completely understood. In this study, epidemiological investigation of 108 unduplicated S. aureus strains from AD patients was undertaken by comprehensive analyses incorporating genotypic and phenotypic data to examine population structure, distribution of VFGs and ARGs, adhesion to HaCaT cells and virulence to G. mellonella. These data provide in-depth insights into S. aureus prevalence in AD in China and will strengthen control strategies. Currently, the genomic epidemiology and phenotypic characterization of S. aureus in Chinese AD patients are unknown. As far as we know, this study is the first to investigate the genomic epidemiology and phenotypic characterization of S. aureus in AD patients in South China.
Phylogenetic analysis and the identification of specific ST lineages of S. aureus using WGS revealed different trends among these countries. ST7 and ST15 were the most common in China. While ST188 and ST7 were the most common in Japan and Italy, respectively12,16,17, ST1 was the most popular in Denmark, the UK, and Singapore11,13,14. Although S. aureus ST1 was abundant in Denmark, the UK, and Singapore, it was not present in the United States18. ST8 was the dominant type of AD patients in the US15,18. Despite the assumption that ST30 was previously only found in HC in Japan17, our research revealed that ST30 strains were also detected in AD patients from Denmark, the UK, Singapore, and China. Except for a few heterogeneous STs, the majority of STs are homogeneous with spa. The diversified spa types associated with ST15, ST72 and ST88 partially revealed the genetic heterogeneity in the three S. aureus clones. In contrast to Lu Liu et al.’s publication21, our investigation identified t2152 and t2612 types in ST188. The accessory gene regulator (agr) quorum sensing (QS) system is a major regulator of virulence phenotypes in S. aureus. There are four agr specificity groups each with a different autoinducer peptide sequence encoded by the agrD gene22. A study of the genomes of about 40,000 S. aureus showed that the distribution of the four agr groups was about 60% type I, about 22% type II, about 14% type III, and about 2.5% type IV23. Our results are consistent with this study, suggesting that agr groups were not associated with AD. Therefore, we need to focus on the role of agr virulence in the pathogenesis of AD in humans.
Important VFGs carried in S. aureus isolates from specific STs may be closely related to their prevalence and transmission. The proportions of VFGs and ARGs in the predominant STs of AD skin-derived isolates were investigated in this study. ST7, ST15 and ST188 mainly carried different types of IEC genes. The gene sak was not detected in the ST15 strains in this study. According to the literature sak is associated with host specificity24, suggesting that the ST15 strains may be of livestock-specific strains origin. Staphylokinase (sak) is a virulence factor acting as a thrombolytic agent, which leads to tissue damage and improves bacterial invasiveness. The SAK protein, encoded by the sak gene, could promote skin infections25. The virulence and adherence of ST7 in Guanghzou were weaker than that of ST188, and we hypothesized that the cause of its prevalence might be related to the esaC, esxB, and sea genes, which are enriched only in ST7. SEA is a staphylococcal enterotoxin that causes staphylococcal food poisoning globally. EsxB and EsaC are secreted by the ESAT-6 secretion system. S. aureus has two ESAT-6-like proteins (EsxA and EsxB), and EsxB plays an important role in intracellular internalization of S. aureus26. EsaC is a secretion substrate of the Ess pathway and implements its pathogenic function during infection27. In addition, cna was detected in some ST188 strains, while it was absent in both ST7 and ST15 strains. The Collagen (Cn)-binding protein Cna is one of the cell surface adhesion proteins that bind to host extracellular mesenchymal proteins. It plays a significant role in bacterial-host adhesion and immune evasion28. This suggests that the adhesion ability and virulence of ST188 is highly correlated with cna gene. However, its specific regulatory mechanism remains to be further investigated. Notably, spa gene was prevalent in S. aureus. spa detection rates varied between STs possibly due to limitations of the sequencing method. Both genotypic and phenotypic analysis were performed in our study in order to assess the antibiotic resistance profile of all the 108 S. aureus strains. The results of antibiotic susceptibility testing indicated that most strains were highly resistant to penicillin G, erythromycin, and clindamycin. In this study, 25.0% of the S. aureus isolated exhibited multidrug resistance. The resistance rate of the S. aureus isolates in this study was lower than that of S. aureus reported by the China Antimicrobial Resistance Surveillance System (https://www.carss.cn/Report/).
The 579 isolates from the NCBI database and China of AD patients were grouped into 15 SCs. Our results indicated that SC5, SC2 and SC7 were predominant SCs in AD. S. aureus isolates of AD were diverse and SCs were not regionally specific. S. aureus in Chinese AD patients was mainly concentrated in SC2, SC7 and SC12. The majority of the phylogeny and population structure studies agreed well with one other, however there were differences in SC2, which requires further investigation.
Further incorporating the S. aureus genomes of HC for comparative genomics, we found that STs, including S. aureus ST1, ST7, and ST582 were the dominant profiles in AD. And S. aureus ST30, ST6, ST508, ST630 and ST22 were the dominant spectrum in HC. ST582 was detected exclusively in AD skin with a clear geographic specificity in these studies. In addition, our study demonstrates that S. aureus of AD may have greater adherence and virulence than S. aureus from healthy human. According to Aziz and colleagues, specific staphylococcal enterotoxins (SEs) may play a role in AD29. Consistent with this we found that the enterotoxin genes seh, selk, and selq were significantly enriched in AD. SEs predominantly engage with the variable regions of T-cell receptor \(\beta\) chains (TCR V\(\beta\)), triggering cytokine production and mitogenic responses in T cells, thereby driving inflammatory onset30,31. They also trigger the production of specific immunoglobulin E (IgE) in the host. IgE-associated superantigens stimulate mast cell and basophil degranulation, leading to the release of mediators like cytokines and chemokines. This promotes skin rashes and itching in dermatitis patients32. Patients with AD also showed a substantial enrichment of lukF-PV, whereas very little lukS-PV was detected in AD and HC. Production of the Panton-Valentine leukocidin (PVL) by S. aureus is mediated via the genes lukS-PV and lukF-PV. Anne Tristan et al. indicated that processed LukS-PV signal peptide was sufficient to significantly enhance the ability of S. aureus to attach to extracellular matrix (ECM) components33. This implies that S. aureus enrichment in AD patients’ skin might not be connected to lukF-PV. However, the exact nature of this mechanism needs to be further investigated. fnbA and fnbB and sdrD were also detected at higher rates in AD. fnbA and fnbB encode fibronectin-binding proteins A and B (FnBPA and FnBPB) which mediate the adhesion of S. aureus to fibrinogen, elastin and fibronectin34. Serine aspartate repeat containing protein D (SdrD) is involved in adhesion to human squamous cells isolated from the nose. Fatemeh Askarian et al. found that genetic deletion of sdrD in S. aureus NCTC8325-4 through allelic replacement resulted in decreased bacterial adherence to Dsg1-expressing HaCaT cells in vitro35. Through the incorporation of publicly available databases, PGWAS identified several VFGs involved in adhesion and biofilm formation and immune evasion that were associated with AD. Qi Peng et al. showed that there was a close relationship between GltB and SaeRS36. In particular, we found that the splA gene was enriched in AD. SplA induces IL-8 in keratinocytes37. Taken together, these differential genes may be one of the reasons why S. aureus frequently colonizes the skin of AD patients. Of course, further research is necessary to confirm the precise role that these genes play in the pathophysiology of AD. Even though these genes are enriched in AD, more research is necessary to determine how these genes contribute to the pathophysiology of AD.
A limitation of our work is that our isolates were all derived from Guangdong province. Multicenter studies are needed to comprehensively assess S. aureus infections in AD patients. Second, this study did not include the very small amount of S. aureus that was extracted from the skin of HC. Future research should consider expanding the skin sample region or gathering nose swabs for culture and isolation. The common colonization site of S. aureus is the anterior nasal cavity. Studies have indicated that nasal carriage of S. aureus considerably increases the risk of autoinoculation and endogenous infection38. Detection of VFGs with WGS also has several general limitations. There is disagreement over the appropriate gene coverage and similarity cutoffs for using WGS to identify virulence genes. We applied a threshold of 80% for both similarity and gene coverage, which has also been used previously39. Although the analyses of VFGs indicate that some VFGs were enriched in AD, this has not been experimentally confirmed in virulence models. Another limitation of this study is that we only performed detailed adhesion and virulence analyses of the three most common STs, while other STs were not covered. Despite these limitations, the study provides a valuable reference direction for subsequent more in-depth and extensive investigations.
Conclusion
In conclusion, our work offers the first thorough characterization of S. aureus isolates from AD patients in China. Chinese AD patients had a higher prevalence rate of S. aureus ST7, ST15, and ST188, according to our study. ST188 was highly lethal to G. mellonella larvae and showed strong adhesion to HaCaT cells. In addition, we analyzed the molecular prevalence characteristics and phylogeny of S. aureus strains globally. The significant difference in numbers of VFGs and ARGs and pan-genes may be associated with the pathogenesis and development of AD. These findings will further our knowledge of the connection between S. aureus and AD and help in the prevention and treatment of S. aureus infections in AD patients.
Materials and methods
Study design and ethics statement
This study evaluated patients with AD, who visited the Guangdong Hospital of Traditional Chinese Medicine between October 2019 and May 2023. The protocol was approved by the Guangdong Hospital of Traditional Chinese Medicine (approval no. BE2019-165-01 and BF2022-145-01) and was performed in accordance with the Declaration of Helsinki. Written informed consent was obtained from all participants. AD was diagnosed according to the diagnostic criteria previously by Williams40. Patients who had used antibiotics within the preceding week were excluded from the study. Total IgE was measured by chemiluminescence using Roche cobas e602, and eosinophils were counted by flow cytometry using mindray BC7500.
Sample collection and S. aureus isolation and identification
Samples from left elbow socket area of HC and lesion of AD within 4-cm2 frame were collected using a swab. And each area was rubbed for 120 s with a sterile swab moistened with saline (medicine). Twenty-four hours before all sampling time points, patients had to avoid washing and the use of topical emollients at the sampling skin site. All samples were refrigerated at \(4\,^{\circ }\)C after collection, and transported for laboratory processing within 2 h of collection.
Skin swabs were placed in tubes containing 3 mL of tryptic soy broth (TSB, Oxoid, UK), immediately capped, and shaken for 30 s using a vortex mixer to suspend the bacterial cells. A volume of 10 \(\upmu \hbox {l}\) of each swab suspension was plated onto Columbia blood and Chocolate agar medium and (Jiangmen Kailin Co., Ltd, China). After 18–24 h incubation at 37 °C, the bacterial colonies were identified by matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry41 and counted. From those samples where S. aureus was detected, a single colony was amplified and stored in a maintenance freeze medium (15% glycerol) at \({-80}\,^{\circ }\)C until use.
Accessing microbiome data for published AD studies
We queried the National Center for Biotechnology Information (NCBI, https://www.ncbi.nlm.nih.gov/) database in December 2023 for publicly available whole genomes using the search terms “Atopic dermatitis and Staphylococcus aureus”. Subsequently, we further reviewed the articles through BioProject to exclude articles with incomplete clinical information and multiple strains of the same individual. Genomic FASTQ files for all SRA accessions were obtained from the NCBI SRA database using prefetch and fastq-dump from the SRA toolkit. A total of 837 genomes of S. aureus strains from AD and HC were collected from public databases representing a broad global spectrum of sites where strains were originally isolated.
Whole-genome sequencing procedures and analyses
After growing the isolates for 18–24 h at 37 °C in Tryptone Soya Agar (TSA, Oxoid, UK), genomic DNA was extracted and purified using a SteadyPure Bacterial Genomic DNA Extraction Kit (Accurate Biology, Bactria, China). DNA was sequenced (Illumina novaseq 6000). Illumina contigs were assembled using SPAdes (version 3.15.3)42. CheckM2 (version 1.0.1)43 was used to calculate the completeness and contamination of the 108 isolates from China and 837 isolates from the global dataset. To minimize batch in the combined global whole genome databases, we only used isolates with completeness \(\ge\) 90%, contamination \(\le\) 5%, N50 > 10,000. Out of the 945 isolate genomes from AD and HC, 918 passed QC (including 579 S. aureus isolates from AD patients and 339 S. aureus isolates from HC).
The assembled contigs of each isolate were used for molecular typing. STs were inferred using the PubMLST database44. The spa and SCCmec types were determined using spa Typer 1.0 (https://cge.food.dtu.dk/services/spaTyper-1.0/) and SCCmecFinder (https://cge.food.dtu.dk/services/SCCmecFinder-1.2/), respectively. Draft genomes were annotated using Prokka (version 1.13.3)45. Pangenome analysis was done using Roary (version 3.11.2)46, and core genome single-nucleotide polymorphisms (cgSNPs) extracted using SNP-sites47. cgSNPs generated were used to construct the approximate maximum-likelihood phylogenetic (ML) tree of the isolates by using RAxML48, and bootstrap supports for bipartitions were estimated using 1000 bootstrap replicates. The visualization and annotation were performed with the tvBOT tool49. Population structure was assessed using cgSNPs with hierBAPS50, which was run four times using maximum clustering sizes of 20, 40, 60 and 80. The ARGs and VFGs were identified by ABRicate (version 1.0.0) (https://github.com/tseemann/abricate) based on ResFinder51 and VFDB21 database, respectively. PGWAS was performed by Scoary \((version 1\cdot 6\cdot 16)\)52, which uses a gene presence/absence dataset. EggNOG-mapper53 was used to annotate the differential gene. Sequencing data are available through NCBI BioProject (accession number PRJNA907199).
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing of 15 antimicrobial agents including penicillin G, erythromycin, cefoxitin, gentamicin, vancomycin, tigecycline, ticoranin, linezolid, benzazolin, rifampin, daptomycin, levofloxacin, clindamycin, ceflorin and cotrimoxazole was determined according to the protocols provided by Clinical and Laboratory Standards Institute (CLSI, 2022). S. aureus ATCC 29213 and ATCC 25923 strains were used for quality control.
Adhesion of S. aureus of major STs to human immortalized epidermal cells (HaCaT)
HaCaT were cultured in Dulbecco’s Modified Eagle Medium (DMEM) with 10% fetal bovine serum (FBS) at \(37\,^{\circ }\)C and 5% CO2. S. aureus was grown to the exponential growth phase in TSB and washed three times with DMEM. Approximately 2.5 \(\times\) \(10^5\) cells were added into 24-well plates. Then the cells were infected with S. aureus at a multiplicity-of-infection (MOI) = 10 and co-incubated at \(37\,^{\circ }\)C and 5% CO2 for 2 h. Subsequently, the supernatant was discarded and the plates were washed three times with sterile PBS to remove loosely adherent bacteria. HaCaT cells were lysed by the addition of 0.05% Triton X-100. The bacteria colony-forming unit (CFU) was enumerated by serial dilution of the cell lysates and spreading of the diluted lysates on TSA.
Comparison of survival rates of Galleria mellonella larvae infected with S. aureus of major STs
S. aureus was grown to the exponential growth phase in TSB and harvested by centrifugation (8000 rpm, 1 min), washed trice with sterile PBS, then resuspended in PBS and adjusted to a concentration of 1 \(\times\) \(10^6\) CFU/ml. Groups of 10 larvae of G. mellonella weighing 250–300 mg (Tianjin huiyude Biotech, Tianjin, China) were used in all assays. G. mellonella was injected with 10 \(\upmu \hbox {l}\) of S. aureus suspension in the last left posterior leg using Hamilton syringe. Larvae with injected with 10 \(\upmu \hbox {l}\) PBS as control. Then the treated larvae were placed in sterile petri plates and kept at \(37\,^{\circ }\)C for observation. The experiment was repeated at least three times.
Statistical analysis
Statistical analyses and random selection were done using GraphPad Prism (version 9.3.0). Differences in carriage of VFGs and ARGs in S. aureus of major STs types were assessed using the chi-square test (n \(\ge\) 40) or Fisher’s exact test (n < 40). One way ANOVA was used for comparing the results of adhesion assay. G. mellonella survival profiles were determined using the Log-rank test and Kaplan-Meier survival plots. P values < 0.05 was reported as statistically significant. Error bars in the figures represented the standard deviation of a data set (mean ± standard). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Data availability
Data are deposited in the National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/, NCBI) with accession number PRJNA907199.
Change history
17 March 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41598-025-93289-7
References
Ständer, S. Atopic dermatitis. N. Engl. J. Med. 384(12), 1136–1143. https://doi.org/10.1056/NEJMra2023911 (2021).
Sroka-Tomaszewska, J. & Trzeciak, M. Molecular mechanisms of atopic dermatitis pathogenesis. Int. J. Mol. Sci. 22, 8. https://doi.org/10.3390/ijms22084130 (2021).
Geoghegan, J. A., Irvine, A. D. & Foster, T. J. Staphylococcus aureus and atopic dermatitis: A complex and evolving relationship. Trends Microbiol. 26(6), 484–497. https://doi.org/10.1016/j.tim.2017.11.008 (2018).
Paller, A. S. et al. The microbiome in patients with atopic dermatitis. J. Allergy Clin. Immunol. 143(1), 26–35. https://doi.org/10.1016/j.jaci.2018.11.015 (2019).
Patrick, G. J. et al. Epicutaneous Staphylococcus aureus induces il-36 to enhance ige production and ensuing allergic disease. J. Clin. Investig. 131(5), 334. https://doi.org/10.1172/jci143334 (2021).
Tauber, M. et al. Staphylococcus aureus density on lesional and nonlesional skin is strongly associated with disease severity in atopic dermatitis. J. Allergy Clin. Immunol. 137(4), 1272–12743. https://doi.org/10.1016/j.jaci.2015.07.052 (2016).
Nutten, S. Atopic dermatitis: Global epidemiology and risk factors. Ann. Nutr. Metab. 66, 8–16. https://doi.org/10.1159/000370220 (2015).
Silverberg, J. I. Comorbidities and the impact of atopic dermatitis. Ann. Allergy Asthma Immunol. 123(2), 144–151. https://doi.org/10.1016/j.anai.2019.04.020 (2019).
Laughter, M. R. et al. The global burden of atopic dermatitis: Lessons from the global burden of disease study 1990–2017. Br. J. Dermatol. 184(2), 304–309. https://doi.org/10.1111/bjd.19580 (2021).
Bakour, S. et al. Identification of virulence factors and antibiotic resistance markers using bacterial genomics. Future Microbiol. 11(3), 455–466. https://doi.org/10.2217/fmb.15.149 (2016).
Chia, M. et al. Shared signatures and divergence in skin microbiomes of children with atopic dermatitis and their caregivers. J. Allergy Clin. Immunol. 150(4), 894–908. https://doi.org/10.1016/j.jaci.2022.01.031 (2022).
Conte, A. L. et al. Atopic dermatitis-derived Staphylococcus aureus strains: What makes them special in the interplay with the host. Front. Cell. Infect. Microbiol. 13, 1194254. https://doi.org/10.3389/fcimb.2023.1194254 (2023).
Edslev, S. M., Clausen, M. L., Agner, T., Stegger, M. & Andersen, P. S. Genomic analysis reveals different mechanisms of fusidic acid resistance in Staphylococcus aureus from danish atopic dermatitis patients. J. Antimicrob. Chemother. 73(4), 856–861. https://doi.org/10.1093/jac/dkx481 (2018).
Harkins, C. P. et al. The widespread use of topical antimicrobials enriches for resistance in Staphylococcus aureus isolated from patients with atopic dermatitis. Br. J. Dermatol. 179(4), 951–958. https://doi.org/10.1111/bjd.16722 (2018).
Key, F. M. et al. On-person adaptive evolution of Staphylococcus aureus during treatment for atopic dermatitis. Cell Host Microbe 31(4), 593–6037. https://doi.org/10.1016/j.chom.2023.03.009 (2023).
Nakamura, Y. et al. Staphylococcus agr virulence is critical for epidermal colonization and associates with atopic dermatitis development. Sci. Transl. Med. 12(551), 4068. https://doi.org/10.1126/scitranslmed.aay4068 (2020).
Obata, S. et al. Comprehensive genomic characterization of Staphylococcus aureus isolated from atopic dermatitis patients in Japan: Correlations with disease severity, eruption type, and anatomical site. Microbiol. Spectr. 11(4), 0523922. https://doi.org/10.1128/spectrum.05239-22 (2023).
Saheb Kashaf, S. et al. Staphylococcal diversity in atopic dermatitis from an individual to a global scale. Cell Host Microbe 31(4), 578–5926. https://doi.org/10.1016/j.chom.2023.03.010 (2023).
Wang, Z. et al. Genomic and functional divergence of Staphylococcus aureus strains from atopic dermatitis patients and healthy individuals: Insights from global and local scales. Microbiol. Spectr. 1, 0057124. https://doi.org/10.1128/spectrum.00571-24 (2024).
Bano, A. et al. Exploring the virulence potential of immune evasion cluster genes in methicillin-resistant Staphylococcus aureus from cancer patients. Saudi J. Biol. Sci. 30(11), 103835. https://doi.org/10.1016/j.sjbs.2023.103835 (2023).
Liu, B., Zheng, D., Zhou, S., Chen, L. & Yang, J. Vfdb 2022: A general classification scheme for bacterial virulence factors. Nucleic Acids Res. 50(D1), 912–917. https://doi.org/10.1093/nar/gkab1107 (2022).
Williams, P., Hill, P., Bonev, B. & Chan, W. C. Quorum-sensing, intra- and inter-species competition in the staphylococci. Microbiology (Reading) 169(8), 1381. https://doi.org/10.1099/mic.0.001381 (2023).
Raghuram, V. et al. Species-wide phylogenomics of the Staphylococcus aureus agr operon revealed convergent evolution of frameshift mutations. Microbiol. Spectr. 10(1), 0133421. https://doi.org/10.1128/spectrum.01334-21 (2022).
Sassi, M. et al. Forecasting Staphylococcus aureus infections using genome-wide association studies, machine learning, and transcriptomic approaches. mSystems 7(4), 0037822. https://doi.org/10.1128/msystems.00378-22 (2022).
Kwiecinski, J. et al. Staphylokinase promotes the establishment of Staphylococcus aureus skin infections while decreasing disease severity. J. Infect. Dis. 208(6), 990–999. https://doi.org/10.1093/infdis/jit288 (2013).
Jang, K. O., Lee, Y. W., Kim, H. & Chung, D. K. Complement inactivation strategy of Staphylococcus aureus using decay-accelerating factor and the response of infected hacat cells. Int. J. Mol. Sci. 22(8), 4015. https://doi.org/10.3390/ijms22084015 (2021).
Burts, M. L., DeDent, A. C. & Missiakas, D. M. Esac substrate for the esat-6 secretion pathway and its role in persistent infections of Staphylococcus aureus. Mol. Microbiol. 69(3), 736–746. https://doi.org/10.1111/j.1365-2958.2008.06324.x (2008).
Herman-Bausier, P. et al. Mechanical strength and inhibition of the Staphylococcus aureus collagen-binding protein cna. mBio 7(5), 16. https://doi.org/10.1128/mBio.01529-16 (2016).
Aziz, F. et al. Staphylococcus aureus isolated from skin from atopic-dermatitis patients produces staphylococcal enterotoxin y, which predominantly induces t-cell receptor v\(\alpha\)-specific expansion of t cells. Infect. Immun. 88(2), 19. https://doi.org/10.1128/iai.00360-19 (2020).
Fraser, J. D. & Proft, T. The bacterial superantigen and superantigen-like proteins. Immunol. Rev. 225, 226–243. https://doi.org/10.1111/j.1600-065X.2008.00681.x (2008).
Sundberg, E. J., Deng, L. & Mariuzza, R. A. Tcr recognition of peptide/mhc class ii complexes and superantigens. Semin. Immunol. 19(4), 262–271. https://doi.org/10.1016/j.smim.2007.04.006 (2007).
Brunner, P. M., Leung, D. Y. M. & Guttman-Yassky, E. Immunologic, microbial, and epithelial interactions in atopic dermatitis. Ann. Allergy Asthma Immunol. 120(1), 34–41. https://doi.org/10.1016/j.anai.2017.09.055 (2018).
Tristan, A. et al. The signal peptide of Staphylococcus aureus panton valentine leukocidin luks component mediates increased adhesion to heparan sulfates. PLoS ONE 4(4), 5042. https://doi.org/10.1371/journal.pone.0005042 (2009).
Murai, M., Moriyama, H., Hata, E., Takeuchi, F. & Amemura-Maekawa, J. Variation and association of fibronectin-binding protein genes fnba and fnbb in Staphylococcus aureus Japanese isolates. Microbiol. Immunol. 60(5), 312–25. https://doi.org/10.1111/1348-0421.12377 (2016).
Askarian, F. et al. The interaction between Staphylococcus aureus sdrd and desmoglein 1 is important for adhesion to host cells. Sci. Rep. 6, 22134. https://doi.org/10.1038/srep22134 (2016).
Peng, Q. et al. Purn is involved in antibiotic tolerance and virulence in Staphylococcus aureus. Antibiotics (Basel) 11(12), 702. https://doi.org/10.3390/antibiotics11121702 (2022).
De Donato, D. P. et al. Staphylococcus aureus serine protease-like protein a (spla) induces il-8 by keratinocytes and synergizes with il-17a. Cytokine 180, 156634. https://doi.org/10.1016/j.cyto.2024.156634 (2024).
Masiuk, H., Wcisłek, A. & Jursa-Kulesza, J. Determination of nasal carriage and skin colonization, antimicrobial susceptibility and genetic relatedness of Staphylococcus aureus isolated from patients with atopic dermatitis in Szczecin, Poland. BMC Infect. Dis. 21(1), 701. https://doi.org/10.1186/s12879-021-06382-3 (2021).
Fröding, I. et al. Extended-spectrum-\(\beta\)-lactamase- and plasmid ampc-producing Escherichia coli causing community-onset bloodstream infection: Association of bacterial clones and virulence genes with septic shock, source of infection, and recurrence. Antimicrob. Agents Chemother. 64(8), 19. https://doi.org/10.1128/aac.02351-19 (2020).
Williams, H. C. et al. The UK working party’s diagnostic criteria for atopic dermatitis. I. Derivation of a minimum set of discriminators for atopic dermatitis. Br. J. Dermatol. 131(3), 383–396. https://doi.org/10.1111/j.1365-2133.1994.tb08530.x (1994).
Calderaro, A. & Chezzi, C. Maldi-tof ms: A reliable tool in the real life of the clinical microbiology laboratory. Microorganisms 12(2), 322. https://doi.org/10.3390/microorganisms12020322 (2024).
Bankevich, A. et al. Spades: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19(5), 455–477. https://doi.org/10.1089/cmb.2012.0021 (2012).
Chklovski, A., Parks, D. H., Woodcroft, B. J. & Tyson, G. W. Checkm2: A rapid, scalable and accurate tool for assessing microbial genome quality using machine learning. Nat. Methods 20(8), 1203–1212. https://doi.org/10.1038/s41592-023-01940-w (2023).
Jolley, K. A., Bray, J. E. & Maiden, M. C. J. Open-access bacterial population genomics: Bigsdb software, the pubmlst.org website and their applications. Wellcome Open Res. 3, 1 (2018).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 30(14), 2068–2069. https://doi.org/10.1093/bioinformatics/btu153 (2014).
Page, A. J. et al. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 31(22), 3691–3693. https://doi.org/10.1093/bioinformatics/btv421 (2015).
Page, A. J. et al. Snp-sites: Rapid efficient extraction of snps from multi-fasta alignments. Microb. Genom. 2(4), 000056. https://doi.org/10.1099/mgen.0.000056 (2016).
Togkousidis, A., Kozlov, O. M., Haag, J., Höhler, D. & Stamatakis, A. Adaptive raxml-ng: Accelerating phylogenetic inference under maximum likelihood using dataset difficulty. Mol. Biol. Evol. 40(10), 227. https://doi.org/10.1093/molbev/msad227 (2023).
Xie, J. et al. Tree visualization by one table (tvbot): A web application for visualizing, modifying and annotating phylogenetic trees. Nucleic Acids Res. 51(W1), 587–592. https://doi.org/10.1093/nar/gkad359 (2023).
Tonkin-Hill, G., Lees, J. A., Bentley, S. D. & Frost, S. D. W. Rhierbaps: An r implementation of the population clustering algorithm hierbaps. Wellcome Open Res. 3, 93. https://doi.org/10.12688/wellcomeopenres.14694.1 (2018).
Florensa, A. F., Kaas, R. S., Clausen, P., Aytan-Aktug, D. & Aarestrup, F. M. Resfinder—An open online resource for identification of antimicrobial resistance genes in next-generation sequencing data and prediction of phenotypes from genotypes. Microb. Genom. 8(1), 748. https://doi.org/10.1099/mgen.0.000748 (2022).
Brynildsrud, O., Bohlin, J., Scheffer, L. & Eldholm, V. Rapid scoring of genes in microbial pan-genome-wide association studies with scoary. Genome Biol. 17(1), 238. https://doi.org/10.1186/s13059-016-1108-8 (2016).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggnog-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38(12), 5825–5829. https://doi.org/10.1093/molbev/msab293 (2021).
Acknowledgements
We are very grateful to all the participants in this study and all team members for their support of this study.
Funding
This research was funded by Fundamental Research Project of Guangzhou Municipal School (College) Joint Funding (Grant No. 2024A03J0026) and Guangdong Basic and Applied Basic Research Fundation (Grant No. 2021A1515110857).
Author information
Authors and Affiliations
Contributions
Conceptualization, C.C. and P.Q. and W.Y.; methodology, C.Y. and M.L.; software, X.C.; investigation, H.L. and S.L.; resources, D.H. and J.F.; data curation, M.Z.; writing-original draft preparation, C.Y.; writing-review and editing, P.Q. and S.L.; visualization, C.Y. and X.C.; supervision, C.C.; project administration, S.L.; funding acquisition, S.L. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The protocol was approved by the Guangdong Hospital of Traditional Chinese Medicine (approval no. BE2019-165-01 and BF2022-145-01).
Consent to participate
Informed consent was obtained from each participant included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained error an error in the legend of Figure 5, where the legend was duplicated from Figure 4. Full information regarding the corrections made can be found in the correction for this Article.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, C., Chen, X., Li, M. et al. Genomic epidemiology and phenotypic characterization of Staphylococcus aureus isolated from atopic dermatitis patients in South China. Sci Rep 15, 4773 (2025). https://doi.org/10.1038/s41598-025-87317-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-87317-9
Keywords
This article is cited by
-
An antimicrobial daptide from human skin commensal Staphylococcus hominis protects against skin pathogens
Nature Communications (2025)