Abstract
This study investigated plant growth-promoting (PGP) mechanisms in Priestia aryabhattai VMYP6 and Paenibacillus sp. VMY10, isolated from tomato roots. Their genomes were initially assessed in silico through various approaches, and these observations were then compared with results obtained in vitro and in vivo. Both possess genes associated with the production of siderophores, indole acetic acid (IAA) and cytokinins (CKs), all of which have been shown to promote plant growth. The two strains were able to produce these compounds in vitro. Although both genomes harbor genes for phosphorus solubilization, only VMY10 demonstrated this ability in vitro. Genes linked to flagellar assembly and chemotaxis were identified in the two cases. Both strains were able to colonize plant roots, even though VMYP6 lacked motility and no flagella were observed microscopically. In the greenhouse, tomato plants inoculated with the strains showed increased biomass, leaf area, and root length. These findings underscore the importance of integrating in vitro assays, genomic analyses, and plant trials to gain a comprehensive insight into the PGP mechanisms of rhizobacteria like VMYP6 and VMY10. Such insight may contribute to improving the selection of strains used as biofertilizers in tomato, a major crop worldwide.
Similar content being viewed by others
Introduction
Plant growth-promoting rhizobacteria (PGPR) are a diverse group of bacteria that adhere to and colonize the rhizosphere. They are known to improve soil quality, suppress plant disease, stimulate plant growth, and increase plant yield1,2. When used as inoculants, their performance depends on their survival and colonization rates, both of which are influenced by environmental factors. Nevertheless, these bacteria are particularly well-equipped to face adverse conditions, thanks to their ability to produce endospores, their motility mechanisms, and their root-associated lifestyle3,4. PGPR include members of the genera Bacillus, Pseudomonas, Burkholderia, and Azospirillum, among others5,6,7.
In 2020, several Bacillus species were reclassified into the newly proposed genus Priestia. Some of these species have shown PGP activity. For instance, Priestia aryabhattai promotes soybean growth by increasing the levels of phytohormones such as abscisic acid (ABA) and gibberellins (GA3), while also exhibiting tolerance to oxidative stress8. Similarly, Priestia megaterium enhances growth in mustard and pepper through phosphate and zinc solubilization, respectively9.
On the other hand, Paenibacillus spp. were originally included within the genus Bacillus but were reclassified as a separate genus in 1993. They are also effective PGPR with agricultural and horticultural applications as soil inoculants10. The genus encompasses more than 150 species. Paenibacillus polymyxa is the most studied but many remain unexplored, while others are yet to be classified11,12.
Sequencing technology has made it possible to analyze the whole genome of isolated strains to reveal potential microbial functions. However, many PGP mechanisms associated with the genera Priestia and Paenibacillus have not been elucidated yet13,14.
Tomato (Solanum lycopersicum L.) is the most in-demand horticultural crop after potato in many places around the world15, including Argentina16,17. Its cultivation takes place mainly in greenhouses and requires particularly high levels of fertilizers18. PGPR alternatives can thus be particularly useful in this case.
This study explored the genomes of two bacterial strains from the rhizosphere of tomato which can promote plant growth. It also sought to understand the mechanisms involved in this promotion. The two strains were taken from a collection gathered in an earlier study, in which they demonstrated positive results in vitro for phosphate solubilization, siderophore synthesis, and phytohormone production. All these capabilities are associated with plant growth promotion. Here, whole genome sequencing and annotation were performed to identify the strains at the species level and locate genes potentially involved in PGP activity. The genotypic data were then correlated with phenotypes observed in vitro and effects recorded in tomato seedlings in vivo.
The overall goal was to enhance our understanding of bacterial PGP mechanisms through a comprehensive and comparative approach that comprises phenotypic, genotypic, and in planta analyses. The information thus gathered can contribute to improving the selection of strains with potential applications as biofertilizers in tomato.
Results
Selection and identification of bacteria with PGP traits
In a previous study, 106 rhizobacteria were isolated from healthy tomato plants19. In the present study, those isolates were screened for the production of indole acetic acid (IAA) and siderophores (Fig. S1a), as well as for phosphate solubilization (Fig. S1b). VMYP6 was selected because of its capacity to produce IAA (5.42 ± 0.14 µg mL-1) and siderophores (4.06 ± 2.52 mm). VMY10 was selected based on its ability to produce IAA (5.89 ± 0.15 µg mL-1) and siderophores (3.84 ± 1.08 mm), and also to solubilize phosphate (6.43 ± 0.75 mm). These traits made the two rhizobacterial isolates potential plant growth promoters. Additionally, both were identified as Gram-positive rods and spore-forming bacilli (Fig. S1c). Analysis of their 16S rDNA gene sequence revealed that VMYP6 belongs to the genus Priestia (GenBank accession number: PQ362315) and VMY10 to Paenibacillus (GenBank accession number: PQ362316).
Whole genome sequencing and phylogenomic analysis
Identification of VMYP6 and VMY10 at the species level was carried out through whole genome sequencing. The sequences were obtained on the Illumina HiSeq 2500 platform. After being assembled, their final sizes were 6,073,033 bp for VMYP6 and 6,702,291 bp for VMY10. They respectively contained 46 and 22 contigs, and 37.3% and 46.91% GC. Assembly completeness and contamination were confirmed to meet the 95% and 5% thresholds (Table S1), as established by previous studies8. Then, the values for average nucleotide identity (ANI) and DNA-DNA hybridization (dDDH) were calculated through comparison with type strains of Paenibacillus and Priestia genera. VMYP6 exhibited over 96% ANI and 70% dDDH with type strain P. aryabhattai JCM 13839. It was thus classified as a member of that species (Table S2a-b). On the other hand, although VMY10 showed the highest ANI and dDDH with type strain Paenibacillus illinoisensis NBRC 15959 (Table S2c-d), the values did not surpass species delineation thresholds.
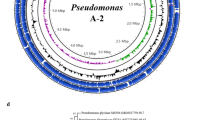
In a further attempt at identification, a core-based phylogenetic tree was built for VMY10 and other Paenibacillus spp. strains with available genome sequences. In this tree, VMY10 clustered with strains identified as P. illinoisensis, including type strain NBRC 15959 (Fig. 1b). VMY10 and the strain 7523-1 were grouped together (Fig. 1b), and shared ANI and dDDH values of 98.5% and 87.8%, respectively. However, neither VMY10 nor 7523-1 shared the required ANI or dDDH values with NBRC 15959 to be classified as P. illinoisensis. This suggests that both strains should be considered a new species, from here on referred to as Paenibacillus sp.
Phylogenetic trees for VMYP6 and VMY10 with their related strains. A total of 133 and 136 common ancestral genes were identified across all studied strains using BLASTN searches with an E-value of 10–30. Genes of P. megaterium ATCC 14581 (a) and P. illinoisensis NBRC 15959 (b) were respectively used as queries. The identified genes were aligned and trimmed individually. For the Priestia genus (a), genes with at least 80% query coverage and 80% identity were selected. For the Paenibacillus genus (b), the selection criteria were set at 90% query coverage and 90% identity. The trees were built with the RAxML algorithm77, and visualized and annotated using iTOL78. Tree reliability was assessed through bootstrapping with 200 and 300 replicates, respectively, using the MRE-based Bootstopping criterion77. Type strains are indicated with a blue star.
The phylogenetic tree made for VMYP6 showed it in a cluster with type strain P. aryabhattai JCM 13839 (Fig. 1a), which confirms the taxonomic classification made on the basis of whole genome sequence analysis.
PGP traits of VMYP6 and VMY10
To investigate the presence of genes that encode proteins involved in promoting plant growth, a BLASTP search was conducted and agnostic annotation was performed on RAST. In VMYP6, 364 proteins with functions reported in PGPR strains were identified through BLASTP. Notably, 30% of those proteins are involved in motility, chemotaxis, and adhesion to plant surfaces, while 13% are associated with phytohormone production and modulation (Fig. 2a). For the same genome, RAST annotated 2077 features in 334 subsystems (Table S3).
PGP genes identified in Priestia aryabhattai VMYP6 and Paenibacillus sp. VMY10. A total of 504 and 390 annotated genes are respectively shown for VMYP6 (a) and VMY10 (b). BLASTP was implemented first, then RAST, and finally PSI-BLAST. Only non-redundant genes identified by each method are shown.
Only 13 protein-encoding genes were detected in VMY10 with BLASTP (Fig. 2b). This may be due to the strain’s lower phylogenetic relatedness to those bacteria used as queries, whose PGP proteins are well-documented. RAST annotated 1724 features in 334 subsystems for the VMY10 genome (Table S3), which resulted in the identification of 159 additional proteins linked to PGP. Twenty-four percent of these are associated with motility, chemotaxis, and attachment to plant surfaces, 19% are involved in sulfur metabolism; 16% in phosphorus metabolism, and 15% in iron transport and siderophore production (Fig. 2b). Identifying more divergent proteins, beyond those that have already been characterized, is challenging20. To address this, query proteins were used to define position-specific scoring matrices (PSSMs), and to identify homologs among the predicted proteins encoded in the VMY10 genome using psi-BLAST. Another 218 PGP-related proteins were identified in this way. A significant proportion (32%) is implicated in the production of antagonist traits, while others (15%) are connected with phytohormone production and modulation (Fig. 2b).
The genome of VMYP6 was explored with psi-BLAST as well. This approach led to the identification of 89 additional proteins (Fig. S2), 74% of which are associated with the production of antagonist traits (Fig. 2a).
As described above, the in silico analyses not only found a plethora of genes that encode PGP proteins, but also others that signal the presence of antagonist traits. Since such traits are clearly related to the synthesis of secondary metabolites, we looked into the possible existence of gene clusters for this synthesis21. VMYP6 has clusters for three terpenes (two that contain RRE and one which is RIPP-like) and clusters for T3PKS, Ni-siderophore, and phosphonate production (Table S4). VMY10 features clusters for two terpenes, one lasso peptide, non-ribosomal peptide synthetase, T3PKS, opine-like metallophore, proteusin, and phosphonate production. In all cases, the cluster components were identified by BLAST, RAST, or PSI-BLAST, except for non-ribosomal peptide synthetase in VMY10 (Table S4).
Table 1 shows the genes involved in the synthesis of phytohormones such as IAA and cytokinins (CKs), and those related to iron uptake, phosphate solubilization and mineralization, motility, chemotaxis, and adhesion to plant surfaces. A complete description of the genes found in VMYP6 and VMY10 is given in Table S5a, b.
Phytohormone production
The in silico analyses revealed numerous genes related to the pathways for auxin and cytokinin biosynthesis in the genomes of VMYP6 and VMY10 (Table 1). These data were supported by phenotypic observations made in vitro: both strains were able to produce IAA, which was assessed in the initial screening, and several other phytohormones. These included different CKs, namely kinetin (KIN), zeatin (ZEA), and 6-benzylaminopurine (BAP), as well as salicylic acid (SA), GA3, and ABA. However, only VMY10 produced indole butyric acid (IBA) and jasmonic acid (JA) (Table 2).
Motility and root colonization
VMYP6 and VMY10 possess genes involved in chemotaxis and the assembly and regulation of flagellum-dependent movement (Table 1).
As shown by transmission electron microscopy (TEM), VMYP6 lacks flagellar filaments while VMY10 has multi-flagellar filaments of considerable length (Fig. 3a). Flagellar functionality was assessed with a 0.3% agar swimming assay. After 15 h of incubation at 28 °C, VMY10 showed swimming motility but VMYP6 did not (Fig. 3b). Motility was likewise evident in VMY10 and absent in VMYP6 when the strains were observed through light microscopy (videos S1 and S2, respectively).
Transmission electron micrographs of VMYP6 and VMY10. The arrows indicate the flagellar basal bodies (a). Swimming motility of VMYP6 and VMY10 on a soft agar plate. Bacteria were stabbed in the center of the agar plate and incubated at 28 °C for 15 h. The ‘ ↔ ’ indicates positive motility by diffuse growth around the inoculation point.
Scanning electron microscopy (SEM) showed that the two strains adhered to the root surface of 10-day-old tomato seedlings in vitro, though colonization density differed from one to the other (Fig. 4). No bacterial cells were found on the root surface of the control treatment.
Scanning electron micrographs of the root surface of 10-day-old tomato plants: uncolonized control treatment; colonization by VMYP6; colonization by VMY10.
Effects of inoculation with VMYP6 and VMY10 on tomato growth under greenhouse conditions
Inoculation with either VMYP6 or VMY10 had a positive effect on the growth of tomato plants compared to the uninoculated control (Fig. 5). All the inoculated plants showed a significant increase (P < 0.05) in stem and leaf dry weight, leaf area, and root length with respect to the control.
Effect of treatment with VMYP6 and VMY10 on plant growth attributes in tomato plants 75 days after sowing: (a) leaf dry weight; (b) stem dry weight; (c) leaf area; and (d) root length. Bars represent mean values ± standard deviation (SD) of three independent replicates. FW: Fresh weight. For each variable, different lowercase letters between columns indicate significant differences at P < 0.05 according to Fisher’s LSD test.
No differences were observed in the number of leaves or flowers between plants treated with the bacteria and the control. In addition, no symptoms of disease were visible in roots or shoots (Fig. S3), which suggests the strains are not pathogenic in tomato plants.
Total chlorophyll was significantly higher in plants inoculated with VMYP6 than in the control, but the carotenoid content remained unchanged. In contrast, total chlorophyll did not differ between plants treated with VMY10 and the control, but the carotenoid content was lower than in the control (Fig. 5).
Discussion
This work adopted a holistic approach to study two native bacterial strains isolated from the tomato rhizosphere, VMYP6 and VMY10. It compared genomic data with the results of in vitro and in planta assays to explore PGP traits in these bacteria that could benefit tomato growth and productivity.
The strains were taxonomically classified as a P. aryabhattai (VMYP6) sp. and a Paenibacillus sp. (VMY10). Members of these genera are used as PGP inoculants in many crops22,23. In addition to engaging in diverse growth-promoting mechanisms, they are capable of forming spores. This makes them more resilient to a wide range of environmental stresses and ensures their long-term survival in the soil4.
Gene-encoded functions in a given bacterium determine its phenotype and behavior in the environment24. To find bacterial species with plant-beneficial traits, an effective strategy consists in searching their genomes for encoded proteins that have already been confirmed to exist in the genome of other PGPR reference strains20. In this study, three complementary tools were used to explore the genomes of VMYP6 and VMY10, in search of homologs of proteins known to be involved in direct or indirect PGP activity: (i) BLASTP, for close homolog assignation; (ii) de novo annotation on RAST, which depends on reliable subsystem-based assertions for genome annotation; and (iii) PSI-BLAST with custom PSSMs, to identify more distantly related evolutionary homologs.
As expected, protein identification via BLASTP was more successful when homolog proteins were more phylogenetically similar to query proteins. When there was a lower degree of relatedness, RAST was crucial to uncover the broader genomic potential of VMYP6 and VMY10. Given that the latter belongs to a species which is more distantly related to the query-harboring strains, the use of PSI-BLAST was also fundamental. This tool, moreover, enabled the identification of antagonist traits in VMYP6, likely because these pathways often comprise more heterogeneous members25.
An important characteristic in PGP bacteria is the production of siderophores, which make it possible to scavenge iron from the environment. Through the action of siderophores, PGPRs may not only make iron more readily available for plant growth but also limit iron availability for pathogens in the same niche. In turn, this may protect plants from pathogenic attack and facilitate bacterial colonization of plant surfaces26,27. According to our results, VMYP6 and VMY10 have gene clusters associated with siderophore production (rhb, yfm)28. Additionally, VMYP6 harbors the efe and feo gene cluster, implicated in iron transport29. The two strains produced siderophores in vitro. Interestingly, a cluster in the genome of VMY10 (identified with antiSMASH) showed 100% similarity with the complete cluster dhbACEBF, associated with a siderophore called bacillibactin30. This secondary metabolite has been studied in B. amyloliquefaciens as a biocontrol agent31.
Another significant PGP trait is the ability to dissolve inorganic phosphorus (P), an essential macronutrient. Eighty percent of soluble P in the soil is immobile and unavailable for plant uptake32, but rhizobacteria can increase its availability by solubilizing it33. We found that VMY10 dissolved inorganic phosphorus in vitro and harbors genes gcd, phoARP, pstABCS and pqqEFG, which play a key role in P solubilization34,35.
Although VMYP6 features the same genes, it did not solubilize P in vitro. Other authors reported that certain microorganisms with the necessary orthologous genes for P solubilization (e.g. pqq and gcd) do not solubilize the element in vitro without an external pqq coenzyme36. Likewise, VMYP6 might only solubilize P if other inhabitants in its microbiome provide it with a necessary coenzyme. This shows the importance of considering soil microbiota as a whole when studying a given microorganism.
Successful colonization of the rhizosphere is essential for PGP4. Colonization involves reaching the root surface through passive movement or active flagellar propulsion37,38. Both VMYP6 and VMY10 were able to attach to and colonize tomato roots in this study, and both contain genes associated with flagellar synthesis and its regulation, including flh, fli, flg39 and motAB40. They also possess genes linked with chemotaxis, such as cheABDRVWY41. However, transmission electron microscopy (TEM) and a swimming assay on agar did not produce similar results for the two strains. In the case of VMY10, TEM revealed peritrichous flagella, which is consistent with the characteristics reported for other Paenibacillus spp.42. Moreover, this strain was able to swim on agar. In contrast, VMYP6 exhibited no swimming motility and TEM failed to detect any flagella in it. Flagellum assembly and functioning require the coordinated expression of over 30 proteins43. Consequently, flagellar presence or absence depend on the comprehensive regulation of gene expression at the transcriptional, translational, and transport levels44. In another study, the root exudates of potato plants were found to have a profound impact on gene expression levels in B. mycoides, including the motility gene set45. Similarly, the expression of the flagellum-related genes identified in silico in VMYP6 may rely on tomato root exudates which were absent in vitro. This is a possible explanation for the lack of flagella ascertained through TEM in this strain.
In the greenhouse, tomato plants inoculated with VMYP6 or VMY10 had increased stem and leaf dry weight, leaf area, and root length with respect to the control. Similar results were obtained when tomato plants were inoculated with other strains of the genera Paenibacillus or Priestia42,46. These improvements are likely due in part to the production of phytohormones by the bacteria. Phytohormones regulate a range of physiological processes in plants, including growth, development, and nutrient uptake40,47. In this study, VMYP6 and VMY10 produced auxins in vitro, such as IAA, gibberellins, and CKs.
Bacterial production of IAA has been reported to have PGP effects48,49,50. Our genomic analyses found that VMYP6 and VMY10 respectively contain trpABC and trpABCDEF cluster genes, which are related to tryptophan pathways that intervene in IAA synthesis. Furthermore, genes yclC, yclB, ysnE, ipdC, and dhaS in VMY10 encode enzymes responsible for the pathways of indole-3-pyruvic acid (IPyA) and tryptamine (TAM)-dependent IAA synthesis in bacteria51,52,53.
VMYP6 and VMY10 also contain several genes associated with CK biosynthesis, including miaA, miaB, yvdD, and log. CKs perform a specific role in delaying plant senescence by regulating protein levels and promoting chlorophyll synthesis in leaves, thereby mitigating leaf yellowing54. Tomato plants treated with the two strains used in this study showed a significant increase in total chlorophyll content, a phenotype to which the presence of CKs may contribute.
Our research found that the presence of genes encoding PGP traits in bacterial genomes is sometimes but not necessarily correlated with phenotypic observations, in agreement with previous findings55,56. Nevertheless, it highlights the importance of genomic analysis to predict PGPR behavior, as previously reported55,57,58. Even greater insight may be gained by complementing functional annotation with in vitro and in vivo results, as was done here with two different PGPRs tested on tomato. This comprehensive approach provides a more in-depth understanding of the mechanisms underlying PGP activity in relevant genera. It also helps to narrow the gap between laboratory findings and real outcomes in the complex rhizosphere environment, and may be a valuable means of identifying good bacterial candidates to be used as biofertilizers.
Materials and methods
Isolation of strains and screening for PGP traits
The bacteria chosen for this study were taken from a collection gathered and described by19. The strains in this collection were isolated from five horticultural farms in Villa María and Villa Nueva, Córdoba, Argentina. To find those with PGP traits, Gram-positive isolates were separated from the rest and assessed for IAA and siderophore production, as well as for phosphorus solubilization. IAA production was detected through a long-standing method59, following the protocol described in another study60. Siderophore production was determined with chrome azurol S (CAS) and hexadecyltrimethylammonium bromide (HDTMA) as indicators61, with slight modifications62. Phosphate solubilization was tested through a plate assay using the National Botanic Research Institute’s phosphate medium (NBRIP), which contains insoluble tricalcium phosphate as the sole source of phosphorus63.
Bacterial cultures
A colony of each strain (VMYP6 and VMY10) was pre-cultured in Luria broth base (LB) in an orbital shaker at 28 °C and 120 rpm, until reaching a concentration of 108 colony forming units (CFU) mL−1. Each culture was harvested by centrifugation at 5000 rpm for 10 min, and the pellets were washed with sterile distilled water. For the inoculations in planta, the cultures were diluted in sterile distilled water at a concentration of 105 CFU mL−1.
The 16S rRNA gene was amplified by polymerase chain reaction (PCR) using universal primers 27F y 1462R. Genomic DNA was extracted with the Wizard DNA Purification kit (Promega) according to the manufacturer’s instructions, and sent to MACROGEN Inc. to be sequenced. The sequences were analyzed with the BLAST algorithm in GenBank (http://www.ncbi.nlm.nih.gov), and aligned with sequences retrieved from that database.
Sequencing, quality control, and de novo assembly of VMYP6 and VMY10
The genomes of VMYP6 and VMY1 were sequenced on the Illumina NovaSeq 6000 platform at Novogene Co. Briefly, genomic DNA was randomly sheared into short fragments, which were then end-repaired, A-tailed, and further ligated with Illumina adapters. The fragments with adapters were amplified by PCR, size selected, and purified. The library was checked using Qubit and real-time PCR for quantification, and Bioanalyzer for size distribution detection. Low-quality reads were filtered, and quality control (QC) reports were generated for each sequence on FastQC (Version 0.11.98)64. The genomes were assembled de novo on SPAdes 3.15.465. The quality of the resulting assemblies was checked on ABySS (Version 2.3.5)66,67. To this end, genome size, contig number, and N50 values were compared to mean values for these parameters. Such values corresponded to the database genomes of other members of the genera to which each strain belongs. Quality thresholds were set as follows: genome size was compared to the mean ± 2 standard deviations (SD), N50 to the mean—2 SD, and the number of contigs to the mean + 2 SD. The completeness and contamination of the assembled genomes were assessed using the BV-BRC implementation in CheckM (BV-BRC Version 3.32.13a). Data management and mathematical operations were performed with the R programming language (version 4.4.1) and the tidyverse package (version 2.0.0)68.
Annotation and gene mining
Genes were annotated and their functions were predicted with the NCBI Prokaryotic Genome Annotation Pipeline (PGAP) and the RAST de novo annotation platform (https://rast.nmpdr.org/ accessed 2023-12-01). Genes of interest were additionally mined with BLAST and PSI-BLAST (version 2.14.0)69. To construct the PSSMs, amino acid sequences of interest served as queries using PSI-BLAST and four iterations. The resulting matrices were then the input for a PSI-BLAST search within the assembled sequences of VMYP6 and VMY10. Only those hits with an E value of 10–5 or less were retained.
Gene clusters involved in secondary metabolite production were predicted with the antiSMASH annotation tool (version 7.1.0) using the default parameters.
Identification of isolates at the species level using genomic data
Genomic sequence datasets and predicted coding sequences for the strains used in the comparative analysis were retrieved from Refseq (ftp.ncbi.nlm.nih.gov/genomes/), using the Download Genomes tool (https://github.com/torresmanno/Download_Genomes). The datasets were composed of sequences submitted until 2023-02-09. Reference or type sequences were defined on the basis of the RefSeq or EZBioCloud databases70. The assembly accession numbers of the selected sequences are detailed in Supplementary Table S6a-b.
Average nucleotide identity (ANI) values were calculated on FastANI V1.3271 using the default parameters.
Maximum likelihood phylogenetic trees were constructed as previously reported20,72 with minor modifications. Only those strains with ANI > 75 were used. Briefly, the genes present in all the strains analyzed (common ancestral genes) were identified by BLASTN searches (Blast + 2.7.1)73 using an E-value of 10–30. To contra-select potential paralogs, coverage and identity percentage cut-offs were set at 90% for the genus Paenibacillus and 80% for the genus Priestia20. The resulting genes were aligned individually on MAFFT version 7.49074, trimmed on Gblock 0.91b75, and concatenated on AMAS76. To optimize the analysis, those sequences that were highly similar to each other were excluded, since they provided no new information. Finally, phylogenetic relationships were inferred through the GTR substitution model with Gamma distribution on RAxML 8.2.1277. Inferred tree reliability was evaluated by bootstrapping. The number of replications was determined by the MRE-based Bootstopping criterion (autoMRE parameter)77 and plotted with iTOL78.
Phytohormone production
To study phytohormone production by VMYP6 and VMY1, the strains were grown in flasks containing nutrient broth supplemented with 0.5 mg mL-1 of L-tryptophan. The flasks were incubated at 28 °C with shaking at 150 rpm for 72 h. Then, 10 mL of the supernatants were collected, centrifuged at 5000 rpm for 15 min, and filtered through a membrane filter (pore size 0.22 μm). These filtered supernatants were used to determine SA, GA3, KIN, 6-BAP, and ABA by liquid chromatography coupled to tandem mass spectrometry (LC-MSMS, Waters and Micromass respectively), as detailed by other authors79. Briefly, the deuterated internal standards for the hormones mentioned above were added, and the partitioning technique was performed with organic solvents. Next, the organic phases were collected, evaporated to dryness, resuspended with methanol (100%), and filtered through 0.45 µm syringe filters. Finally, they were diluted in 50 µL of pure methanol, and 10 µL were injected into the liquid chromatograph.
Electron microscopy and motility assay
Negative staining The methodology proposed by other authors80 was performed with some modifications. The bacterial cultures were grown in LB broth and statically cultured at 28 °C for 8 h. To obtain TEM images, Formvar-coated nickel grids were floated on 15 µl of bacterial suspension for 10 min. After that, the grids with absorbed bacteria were incubated with a 2% phosphotungstic acid solution for 1 min. They were subsequently rinsed with distilled water and the excess liquid was drained off with the edge of a filter paper. The preparations were air-dried for 10 min. Finally, the specimens were examined under a Hitachi HT7800 electron microscope (Hitachi, Tokyo, Japan) operated at 80 kv, and photographed with a NanoSprint15 AMT camera (Advanced Microscopy Techniques Corp., Woburn, MA, USA).
Swimming motility was studied as described previously81. Three µl of bacterial suspensions (OD600 = 0.3) were inoculated in the center of 90 mm plates containing LB medium with 0.3% agar. The plates were sealed with parafilm to prevent dehydration and incubated at 28 ± 2 °C for 15 h before swimming was measured.
Bacterial root colonization observed by SEM
Bacterial colonization of the rhizosphere and the rhizoplane was qualitatively demonstrated in vitro by SEM. Seeds were hydrated with tap water for 6 h and rinsed three times with sterile distilled water. Next, they were placed for 15 min in shakers containing a suspension of VMYP6 or VMY10, adjusted to an OD600 of 103 UFC.ml−1. Control seeds were placed in sterile distilled water without the bacterial inoculum. The seeds were germinated on 110 mm plates containing 50% MS medium82 with 0.8% agar. These plates were incubated for 10 days at 26 °C with a photoperiod of 16/8 h (l/d). At that time, root samples were excised and fixed in 2.5% glutaraldehyde (v/v) in 0.1 M phosphate buffer (pH 7.2) for 4 h. The samples were dehydrated through a graded ethanol series (30–100%, v/v), and dried in CO2 by the critical point method. To prevent damage to the tissues, pressure was released slowly and they were shadowed with gold (22 nm) before observation under a scanning electron microscope (Zeiss supra 55vp).
Greenhouse experiments
The in vivo study was carried out between February and May 2022 in a greenhouse located on an experimental field belonging to the National University of Villa Maria, Argentina (32° 22′ 56.5″ S 63° 15′ 38.7″ W). The experiment consisted in inoculating the roots of tomato plants (Solanum lycopersicum) with VMYP6 or VMY10. Control plants were not inoculated, and there were 10 plants per treatment.
Tomato seeds (hybrid Ichiban), purchased from Bayer CropScience, were washed with tap water for 6 h and sown in speedlings for germination at 28 °C, until cotyledons were visible. The plants were transferred to the greenhouse and kept under natural conditions, at an average temperature of 20.2 °C (± 7 °C). Thirty days after sowing (DAS), similarly sized seedlings were transplanted into single 1 L pots containing GrowMix MultiPro. The plants were inoculated seven and 15 days after being transplanted. The bacterial suspensions for the first and second inoculations were respectively 5 mL (105 CFU mL−1) and 10 mL (105 CFU mL−1). The plants were irrigated periodically throughout the study. The following growth parameters were determined 75 DAS: dry weight of leaves and stems (g), leaf area (cm2), and root length (cm). To find out the leaf area, leaves were scanned and measured in square cm2 on the software ImageJ. Chlorophyll content was also measured 75 DAS (see below).
Determination of photosynthetic pigment content
To determine the photosynthetic pigment content, leaf discs weighing 100 mg were taken from inoculated and control plants. Afterward, they were incubated in 5 mL of 95% ethanol in the dark, until they became colorless. The concentrations of chlorophyll a and b were measured spectrophotometrically at 664 nm and 648 nm, respectively. Carotenoid concentration was determined at 470 nm following another study83. The values were expressed as µg g (fresh weight or FW)−1.
Statistical analysis
All the experiments were repeated three times. An analysis of variance (ANOVA) followed by an a posteriori test for comparison of means (Fisher’s LSD) were conducted for multiple comparisons. A value of P < 0.05 was considered statistically significant. For ANOVA, parametric assumptions were tested using the Shapiro–Wilk test for normality, Levene’s test for homogeneity, and a test of sample independence of the scatter plots of residuals (y-axis) and cases (x-axis). All the data were statistically analyzed on Infostat version 202084, and expressed as mean ± SD.
Data availability
The datasets supporting the conclusions of this article are included within the article and its additional files. Sequencing and assembly data are available in the NCBI BioProject database under the accession PRJNA1041798.
References
Normander, B. O. & Prosser, J. I. Bacterial origin and community composition in the barley phytosphere as a function of habitat and presowing conditions. Appl. Environ. Microbiol. 66(10), 4372–4377. https://doi.org/10.1128/AEM.66.10.4372-4377.2000 (2000).
Ikiz, B., Dasgan, H. Y. & Gruda, N. S. Utilizing the power of plant growth promoting rhizobacteria on reducing mineral fertilizer, improved yield, and nutritional quality of Batavia lettuce in a floating culture. Sci. Rep. 14, 1616. https://doi.org/10.1038/s41598-024-51818-w (2024).
Ansari, F. A. & Ahmad, I. Fluorescent Pseudomonas -FAP2 and Bacillus licheniformis interact positively in biofilm mode enhancing plant growth and photosynthetic attributes. Sci. Rep. 9, 4547. https://doi.org/10.1038/s41598-019-40864-4 (2019).
Gupta, A. et al. Plant growth promoting rhizobacteria (PGPR): A sustainable agriculture to rescue the vegetation from the effect of biotic stress—A review. Lett. Appl. NanoBiosci. 10, 2459–2465. https://doi.org/10.33263/LIANBS103.24592465 (2021).
Pérez-Rodriguez, M. M. et al. Pseudomonas fluorescens and Azospirillum brasilense increase yield and fruit quality of tomato under field conditions. J. Soil Sci. Plant Nutr. 20(1614–1624), 2020. https://doi.org/10.1007/s42729-020-00233-xh (2020).
Cochard, B. et al. Endophytic PGPR from tomato roots: Isolation, in vitro characterization and in vivo evaluation of treated tomatoes (Solanum lycopersicum L.). Microorganisms 2022 10, 765. https://doi.org/10.3390/microorganisms10040765 (2022).
Nakahara, H., Matsuzoe, N., Taniguchi, T. & An, P. Effect of Burkholderia sp. and Pseudomonas spp. inoculation on growth, yield, and absorption of inorganic components in tomato ‘Micro-Tom’ under salinity conditions. J. Plant Interact. 17(1), 277–289. https://doi.org/10.1080/17429145.2022.2035439 (2022).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055. https://doi.org/10.1101/gr.186072.114 (2015).
Bhatt, K. & Maheshwari, D. K. Zinc solubilizing bacteria (Bacillus megaterium) with multifarious plant growth promoting activities alleviates growth in Capsicum annuum L. 3 Biotech 10(2), 36. https://doi.org/10.1007/s13205-019-2033-9 (2020).
Ash, C., Priest, F. G. & Collins, M. D. Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test: proposal for the creation of a new genus Paenibacillus. Antonie van leeuwenhoek 64, 253–260 (1993).
Soni, R., Rawal, K. & Keharia, H. Genomics assisted functional characterization of Paenibacillus polymyxa HK4 as a biocontrol and plant growth promoting bacterium. Microbiol. Res. 2021(248), 126734. https://doi.org/10.1016/j.micres.2021.126734 (2021).
Luo, Y. et al. Complete genome sequence of industrial biocontrol strain Paenibacillus polymyxa HY96-2 and further analysis of its biocontrol mechanism. Front. Microbiol. 9, 1520. https://doi.org/10.3389/fmicb.2018.01520 (2018).
Deng, C. et al. Molecular mechanisms of plant growth promotion for methylotrophic Bacillus aryabhattai LAD. Front. Microbiol. 13, 917382. https://doi.org/10.3389/fmicb.2022.917382 (2022).
Patowary, R. & Deka, H. Paenibacillus. In Beneficial Microbes in Agro-Ecology 339–361 (Academic Press, 2020). https://doi.org/10.1016/B978-0-12-823414-3.00017-4
Ali, M. Y. et al. Nutritional composition and bioactive compounds in tomatoes and their impact on human health and disease: A review. Foods https://doi.org/10.3390/foods10010045 (2021).
Ávila, G., Salusso, F. & Wester, O. Horticulture in the Córdoba Province. Analysis of some contextual aspects. Hortic. Argentina 39, 21–34 (2020).
Lozano, F. et al. Overview of the Argentinean horticulture sector: 1. Characterisation and priorities of national horticulture. Hortic. Argentina 39, 76–102 (2020).
Alconada Magliano, M. M. Salt-affected soils in protected production systems, irrigation water and production management. In H. Rimski-Korsakov y RS Lavado (Comp.), Actas VI Congreso de la Red Argentina de Salinidad 140–145 (2019).
Felipe, V., Bianco, M., Terrestre, M. & Yaryura, P. Biocontrol of tomato bacterial spot by novel Bacillus and Pseudomonas strains. Eur. J. Plant Pathol. 160, 935–948. https://doi.org/10.1007/s10658-021-02297-6 (2021).
Torres Manno, M. A. et al. GeM-Pro: A tool for genome functional mining and microbial profiling. Appl. Microbiol. Biotechnol. 103(7), 3123–3134. https://doi.org/10.1007/s00253-019-09648-8 (2019).
Dhanabalan, S. et al. Unleashing Bacillus species as versatile antagonists: Harnessing the biocontrol potentials of the plant growth-promoting rhizobacteria to combat Macrophomina phaseolina infection in Gloriosa superba. Microbiol. Res. https://doi.org/10.1016/j.micres.2024.127678 (2024).
Rios-Ruiz, W. F., Tuanama-Reátegui, C., Huamán-Córdova, G. & Valdez-Nuñez, R. A. Co-inoculation of endophytes Bacillus siamensis TUR07-02b and Priestia megaterium SMBH14-02 promotes growth in rice with low doses of nitrogen fertilizer. Plants 12(3), 524. https://doi.org/10.3390/plants12030524 (2023).
Hussain, A. et al. Plant-growth-promoting Bacillus and Paenibacillus species improve the nutritional status of Triticum aestivum L. Plos One 15(12), e0241130. https://doi.org/10.1371/journal.pone.0241130 (2020).
Karlsen, S. T., Rau, M. H., Sánchez, B. J., Jensen, K. & Zeidan, A. A. From genotype to phenotype: computational approaches for inferring microbial traits relevant to the food industry. FEMS Microbiol. Rev. 47(4), fuad030. https://doi.org/10.1093/femsre/fuad030 (2023).
Aleti, G., Sessitsch, A. & Brader, G. Genome mining: Prediction of lipopeptides and polyketides from Bacillus and related Firmicutes. Comput. Struct. Biotechnol. J. 13, 192–203 (2015).
Chincholkar, B. L., Chaudhari, B. L. & Rane, M. R. Microbial siderophore: A state of art. In Microbial Siderophores (eds Varma, A. & Chincholkar, S. B.) 233–242 (Springer-Verlag, 2007).
Soares, E. V. Perspectiva sobre la producción biotecnológica de sideróforos bacterianos y su uso. Appl. Microbiol. Biotechnol. 106, 3985–4004. https://doi.org/10.1007/s00253-022-11995-y (2022).
Timofeeva, A. M., Galyamova, M. R. & Sedykh, S. E. Plant growth-promoting soil bacteria: Nitrogen fixation, phosphate solubilization, siderophore production, and other biological activities. Plants 12, 4074. https://doi.org/10.3390/plants12244074 (2023).
Gómez-Garzón, C., Barrick, J. E. & Payne, S. M. Disentangling the evolutionary history of feo, the major ferrous iron transport system in bacteria. mBio 13, e03512-e3521. https://doi.org/10.1128/mbio.03512-21 (2022).
Yue, Z. et al. Bacillus sp. WR12 alleviates iron deficiency in wheat via enhancing siderophore- and phenol-mediated iron acquisition in roots. Plant Soil 471, 247–260. https://doi.org/10.1007/s11104-021-05218-y (2022).
Dimopoulou, A. et al. Direct antibiotic activity of bacillibactin broadens the biocontrol range of Bacillus amyloliquefaciens MBI600. mSphere 6, e00376-21. https://doi.org/10.1128/mSphere.00376-21 (2021).
Anzuay, M. S. et al. Growth promotion of peanut (Arachis hypogaea L.) and maize (Zea mays L.) plants by single and mixed cultures of efficient phosphate solubilizing bacteria that are tolerant to abiotic stress and pesticides. Microbiol. Res. 199, 98–109 (2017).
Pongrac, P., Fischer, S., Thompson, J. A., Wright, G. & White, P. J. Earl responses of Brassica oleracea roots to zinc supply under sufficient and sub-optimal phosphorus supply. Front. Plant Sci. 10, 1645. https://doi.org/10.3389/fpls.2019.01645 (2020).
Lu, J. L. et al. Remarkable effects of microbial factors on soil phosphorus bioavailability: A country-scale study. Global Change Biol. 28(14), 4459–4471. https://doi.org/10.1111/gcb.16213 (2022).
Dai, Z. et al. Long-term nutrient inputs shift soil microbial functional profiles of phosphorus cycling in diverse agroecosystems. ISME J. 14(3), 757–770. https://doi.org/10.1038/s41396-019-0567-9 (2020).
Leontidou, K. et al. Plant growth promoting rhizobacteria isolated from halophytes and drought-tolerant plants: Genomic characterisation and exploration of phyto-beneficial traits. Sci. Rep. 10(1), 14857. https://doi.org/10.1038/s41598-020-71652-0 (2020).
Harshey, R. M. Bacterial motility on a surface: Many ways to a common goal. Annu. Rev. Microbiol. 57, 249–273. https://doi.org/10.1146/annurev.micro.57.030502.091014 (2003).
Mahmoud, F. M. et al. Comparative genomic analysis of strain Priestia megaterium B1 reveals conserved potential for adaptation to endophytism and plant growth promotion. Microbiol. Spectr. 12, e00422-e424. https://doi.org/10.1128/spectrum.00422-24 (2024).
Yan, Y., Xu, W., Chen, W., Hu, Y. & Wang, Z. Complete genome sequence of Bacillus velezensis YYC, a bacterium isolated from the tomato rhizosphere. Arch. Microbiol. 204, 1–5. https://doi.org/10.1007/s00203-021-02709-5 (2022).
Dahmani, M. A. et al. Unearthing the plant growth-promoting traits of Bacillus megaterium RmBm31, an endophytic bacterium isolated from root nodules of Retama monosperma. Front. Plant Sci. 27(11), 124. https://doi.org/10.3389/fpls.2020.00124 (2020).
Dong, X. et al. The genome of Bacillus velezensis SC60 provides evidence for its plant probiotic effects. Microorganisms 10, 767. https://doi.org/10.3390/microorganisms10040767 (2022).
Lee, Y., Kwon, S., Balaraju, K. & Jeon, Y. Influence of phenotypic variation of Paenibacillus polymyxa E681 on growth promotion in cucumbers. Front. Microbiol. 15, 1427265. https://doi.org/10.3389/fmicb.2024.1427265 (2024).
Thormann, K. M. Dynamic hybrid flagellar motors—fuel switch and more. Front. Microbiol. 13, 863804. https://doi.org/10.3389/fmicb.2022.863804 (2022).
do Amaral, F. P. et al. Diverse bacterial genes modulate plant root association by beneficial bacteria. mBio 11, e03078-20. https://doi.org/10.1128/mBio.03078-20 (2020).
Yi, Y., de Jong, A., Frenzel, E. & Kuipers, O. P. Comparative transcriptomics of Bacillus mycoides strains in response to potato-root exudates reveals different genetic adaptation of endophytic and soil isolates. Front. Microbiol. 8, 1487. https://doi.org/10.3389/fmicb.2017.01487 (2017).
Katsenios, N. et al. Evaluation of plant growth promoting bacteria strains on growth, yield and quality of industrial tomato. Microorganisms 2021(9), 2099. https://doi.org/10.3390/microorganisms9102099 (2021).
Adedayo, A. A. et al. The application of plant growth-promoting rhizobacteria in Solanum lycopersicum production in the agricultural system: A review. PeerJ https://doi.org/10.7717/peerj.13405 (2022).
Wagi, S. & Ahmed, A. Bacillus spp.: Potent microfactories of bacterial IAA. PeerJ 7, e7258. https://doi.org/10.7717/peerj.7258 (2019).
Rehan, M., Al-Turki, A., Abdelmageed, A. H. A., Abdelhameid, N. M. & Omar, A. F. Performance of plant-growth-promoting rhizobacteria (PGPR) Isolated from sandy soil on growth of tomato (Solanum lycopersicum L). Plants 12, 1588. https://doi.org/10.3390/plants12081588 (2023).
Al-Kurtany, A. E. S., Ali, S. A. M. & Oleawy, M. F. Tomato seedling production using an inoculum prepared with plant growth-promoting rhizobacteria (PGPR) isolates. SABRAO J. Breed. Genet. 55(1), 230–236. https://doi.org/10.54910/sabrao2023.55.1.21 (2023).
Shao, J. et al. Analysis and cloning of the synthetic pathway of the phytohormone indole-3-acetic acid in the plant-beneficial Bacillus amyloliquefaciens SQR9. Microb. Cell Factories 14, 1–13. https://doi.org/10.1186/s12934-015-0323-4 (2015).
Shao, J. et al. Participating mechanism of a major contributing gene ysnE for auxin biosynthesis in Bacillus amyloliquefaciens SQR9. J. Basic Microbiol. 61(6), 569–575. https://doi.org/10.1002/jobm.202100098 (2021).
Sun, H. et al. Identification and combinatorial engineering of indole-3-acetic acid synthetic pathways in Paenibacillus polymyxa. Biotechnol. Biofuels 15, 81. https://doi.org/10.1186/s13068-022-02181-3 (2022).
Guo, Y. et al. Leaf senescence: progression, regulation, and application. Mol. Hortic. 1, 5. https://doi.org/10.1186/s43897-021-00006-9 (2021).
Guerrieri, M. C. et al. Integrated genomic and greenhouse assessment of a novel plant growth-promoting rhizobacterium for tomato plant. Front. Plant Sci. https://doi.org/10.3389/fpls.2021.660620 (2021).
Silva, U. C. et al. Genomic and phenotypic insights into the potential of rock phosphate solubilizing bacteria to promote millet growth in vivo. Front. Microbiol. 11, 574550. https://doi.org/10.3389/fmicb.2020.574550 (2021).
Zhang, L., Chen, W., Jiang, Q., Fei, Z. & Xiao, M. Genome analysis of plant growth-promoting rhizobacterium Pseudomonas chlororaphis subsp. aurantiaca JD37 and insights from comparasion of genomics with three Pseudomonas strains. Microbiol. Res. 237, 126483. https://doi.org/10.1016/j.micres.2020.126483 (2020).
Fadiji, A. E., Ayangbenro, A. S. & Babalola, O. O. Genomic assessment of Enterobacter mori AYS9: A potential plant growth-promoting drought-resistant rhizobacteria. Span. J. Soil Sci. 13, 11302. https://doi.org/10.3389/sjss.2023.1130 (2023).
Loper, J. E. & Schroth, M. N. Influence of bacterial sources of indole-3-acetic acid on root elongation of sugar beet. Phytopathology 76(4), 386–389 (1986).
Ahmad, F., Ahmad, I. & Khan, M. S. Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiol. Res. 163(2), 173–181. https://doi.org/10.1016/j.micres.2006.04.001 (2008).
Schwyn, B. & Neilands, J. B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 160(1), 47–56. https://doi.org/10.1016/0003-2697(87)90612-9 (1987).
Pinter, I. F., Salomon, M. V., Berli, F., Bottini, R. & Piccoli, P. Characterization of the As (III) tolerance conferred by plant growth promoting rhizobacteria to in vitro-grown grapevine. Appl. Soil Ecol. 109, 60–68. https://doi.org/10.1016/j.apsoil.2016.10.003 (2017).
Nautiyal, C. S. An efficient microbiological growth medium for screening phosphate solubilizing microorganisms. FEMS Microbiol. Letters 170(1), 265–270 (1999).
Andrews, S. et al. FastQC: A quality control tool for high throughput sequence data. Babraham Bioinformatics (Babraham Institute, 2010).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19(5), 455–477. https://doi.org/10.1089/cmb.2012.0021 (2012).
Simpson, J. T. et al. ABySS: A parallel assembler for short read sequence data. Genome Res. 19(6), 1117–1123. https://doi.org/10.1101/gr.089532.108 (2009).
Jackman, S. D. et al. ABySS 20: Resource-efficient assembly of large genomes using a bloom filter. Genome Res. 27(5), 768–777. https://doi.org/10.1101/gr.214346.116 (2017).
Wickham, S. et al. Welcome to the Tidyverse. J. Open Source Softw. 4(43), 1686. https://doi.org/10.21105/joss.01686 (2019).
Camacho, C. & Madden, T. BLAST+ Release Notes. 2013 Mar 12 [Updated 2024 Jun 25]. In: BLAST® Help [Internet]
Yoon, S. H. et al. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 67, 1613–1617. https://doi.org/10.1099/ijsem.0.001755 (2017).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9(5114), 1–8. https://doi.org/10.1038/s41467-018-07641-9 (2018).
Espariz, M., Zuljan, F. A., Esteban, L. & Magni, C. Taxonomic identity resolution of highly phylogenetically related strains and selection of phylogenetic markers by using genome-scale methods: the Bacillus pumilus group case. PLoS One 11, e0163098. https://doi.org/10.1371/journal.pone.0163098 (2016).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res 36, W5–W9. https://doi.org/10.1093/nar/gkn201 (2008).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30(4), 772–780. https://doi.org/10.1093/molbev/mst010 (2013).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577. https://doi.org/10.1080/10635150701472164 (2007).
Borowiec, M. L. AMAS: A fast tool for alignment manipulation and computing of summary statistics. PeerJ 4, e1660. https://doi.org/10.7717/peerj.1660 (2016).
Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313. https://doi.org/10.1093/bioinformatics/btu033 (2014).
Letunic, I. & Bork, P. Interactive tree of life v2: Online annotation and display of phylogenetic trees made easy. Nucleic Acids Res. 39, W475–W478. https://doi.org/10.1093/nar/gkr201 (2011).
Masciarelli, O., Llanes, A. & Luna, V. A new PGPR co-inoculated with Bradyrhizobium japonicum enhances soybean nodulation. Microbiol. Res. 169(7–8), 609–615. https://doi.org/10.1016/j.micres.2013.10.001 (2014).
Li, Y. et al. A flagella hook coding gene flgE positively affects biofilm formation and cereulide production in emetic Bacillus cereus. Front. Microbiol. 13, 897836. https://doi.org/10.3389/fmicb.2022.897836 (2022).
Adler, J. Chemotaxis in bacteria: Motile Escherichia coli migrate in bands that are influenced by oxygen and organic nutrients. Science 153(3737), 708–716. https://doi.org/10.1126/science.153.3737.708 (1966).
Murashige, T. & Skoog, F. A Revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x (1962).
Lichtenthaler, H. K. & Buschmann, C. Extraction of photosynthetic tissues: Chlorophylls and carotenoids. Curr. Protocols Food Anal. Chem. 1(1), F4–F2 (2001).
Di Rienzo, J. et al. InfoStat Statistical Software (2003).
Acknowledgements
This work was funded by the Fondo para la Investigación Científica y Tecnológica (FONCyT) of the Agencia Nacional de Promoción de la Investigación, el Desarrollo Tecnológico y la Innovación (Argentina): PICT 2019 N° 3274. It was also partially financed by PID-2023-80020220700089UR from the UNR and DTT-2023-021 granted by the Agencia Santafesina de Ciencia Tecnología e Innovación (ASaCTeI). VA, MIB, ME, and PY are Career Investigators of the Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Argentina. AMR is a professor at the University of Buenos Aires, Argentina. CA, TDP and MAP are recipients of a doctoral fellowship from CONICET, Argentina. The authors thank the TEM service of the Instituto de Investigaciones en Ciencias de la Salud (INICSA)-CONICET, Electronic Microscopy Center, Facultad de Ciencias Médicas, Universidad Nacional de Córdoba, Argentina.
Author information
Authors and Affiliations
Contributions
C.A: Experimentation, Methodology, Research and Formal analysis, Writing—review & editing of the manuscript. T.D.P: Methodology, Research and Formal analysis, Writing—review & editing of the manuscript. M.A.P: Methodology, Writing—review & editing of the manuscript. A.M.R: review & editing of the manuscript. V.A: Methodology, review & editing of the manuscript. M.I.B: Writing—review & editing of the manuscript. M.E: Research supervision, Methodology and Formal analysis, Writing—review & editing of the manuscript. P.M.Y: Fund acquisition and Project management, Conceptualization, Research supervision and Formal analysis, Writing—review & editing of the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Video S1.
Supplementary Video S2.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Almirón, C., Petitti, T.D., Ponso, M.A. et al. Functional and genomic analyses of plant growth promoting traits in Priestia aryabhattai and Paenibacillus sp. isolates from tomato rhizosphere. Sci Rep 15, 3498 (2025). https://doi.org/10.1038/s41598-025-87390-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-87390-0
This article is cited by
-
Iterative enrichment cultivation and multiomic analysis reveal potential endophytic bacteria affecting the sinomenine synthesis in Sinomenium acutum
Microbial Cell Factories (2026)
-
Improvement of Growth and Selenium Absorption Characteristics of Tomato Plant by Lactic Acid Bacterial Biochar Fertilizer
Current Microbiology (2026)
-
Genomically-selected antifungal Bacillaceae strains improve wheat yield and baking quality
Applied Microbiology and Biotechnology (2025)