Abstract
Clostridium perfringens is an important foodborne pathogen that produces diverse toxins and is often associated with foodborne gastroenteritis. In this sense, novel biopreservatives with anti-C. perfringens activity are of interest. Among them, bacteriocins produced by lactic acid bacteria stand out as potential candidates. This study describes leucocyclicin C, a novel variant of the bacteriocin leucocyclicin Q, capable of inhibiting C. perfringens. The bacteriocin comprises 61 amino acids, has a molecular mass of 6,081.44 Da, and is produced by the strain Leuconostoc lactis APC 3969. Like many circular bacteriocins, leucocyclicin C has a broad spectrum of activity, is protease resistant, and has high stability against thermic and pH stresses. The leucocyclicin C genetic cluster comprises ten genes instead of the five genes previously described for leucocyclicin Q. Also, this genetic cluster seems to be part of a putative composite transposon. Leucocyclicin C has a minimum inhibitory concentration (MIC) of 3.288 µM against C. perfringens, comparable with other antimicrobial peptides. These results suggest that leucocyclicin C has the potential as a biopreservative for controlling C. perfringens in food.
Similar content being viewed by others
Introduction
Clostridium perfringens is an anaerobic, gram-positive, rod-shaped, spore-forming bacterium that is commonly associated with foodborne gastroenteritis1. This pathogen is ubiquitous and can be found in different sources such as raw materials in feed production, soil, mammalian gut, and plants, among others. However, C. perfringens can cause serious foodborne illness since its spores survive food processing and germinate in the gastrointestinal (GI) tract of humans. Indeed, the short germination time of the spores, along with their high resilience combined with inadequate hygiene conditions, all facilitate the survival of C. perfringens2. On entering the GI tract, C. perfringens can sporulate and produce enterotoxins, leading to illness symptoms, such as abdominal pain, diarrhoea, and gas production3. There are seven types of C. perfringens (A to G), which are categorized based on their patterns of toxin production [Alpha, Beta, Epsilon, Iota, enterotoxin CPE and necrotic enteritis B-like toxin (NetB)]. Among them, type A isolates produce only alpha toxins and are commonly reported in food-poisoning outbreaks; type B isolates (Alpha, Beta and Epsilon toxins) are usually associated with haemorrhagic enteritidis in sheep, foals and calves; type C isolates (Alpha, Beta toxins and may present CPE enterotoxin) may cause serious clostridial necrotizing enteritis; type D isolates (Alpha, Epsilon toxins and may present CPE enterotoxin) are associated with enterotoxaemia in goats, sheep and cattle; type E isolates (Alpha, Iota toxins and may present CPE enterotoxin) are commonly associated with enterotoxaemia in lambs and calves; type F isolates (Alpha toxin and CPE enterotoxin) may cause human food-poisoning and non-foodborne diarrhoea; and type G isolates (Alpha toxin and NetB toxin) are suggested to be the aetiological agents of avian necrotic enteritis4,5. To mitigate such risks, different approaches have been investigated and developed to inactivate vegetative cells and spores of C. perfringens, including the use of strict processing conditions2, physical treatments (thermal and pressure), chemical additives (preservatives, organic acids) and biological compounds (plant extracts, animal lysozyme and microbial metabolites)6. Recently, studies have shown promising results using bacteriophages, bacteriocinogenic bacteria and purified bacteriocins with anti-C. perfringens activities7,8,9.
Microbial secondary metabolites with antimicrobial properties, especially bacteriocins, have attracted attention as promising alternatives to traditional antibiotics and chemical food preservatives. Bacteriocins are small ribosomally-synthesized peptides that can demonstrate narrow- or broad-spectrum antimicrobial activity. They may be classified into two major classes: Class I, post-translationally modified peptides, such as head-to-tail circularization, and Class II, peptides without any modification10. One example of Class I bacteriocins are lantibiotics, small peptides with characteristic amino acids containing thioether bridges (lanthionine and β-methyllanthionine). Nisin, the most well-studied bacteriocin, is the principal representative of this class11. Class I also includes sactibiotics, linaridins, thiopeptides, lasso peptides and glycocins, circular bacteriocins12, among others10. The Class II bacteriocins can be divided into five subgroups: (i) pediocin-like bacteriocins with a consensus YGNGVXC N-terminal sequence; (ii) two-component peptides where both are required for activity; (iii) linear, non-pediocin-like, non-two component bacteriocins; (iv) defensin-like bacteriocins13; and (v) leaderless bacteriocins10,14. The circular bacteriocins are an interesting subgroup due to their circular structure associated with a globular conformation, which limits the accessibility of proteases. In addition, one study indicated that the circular structure is not essential for bioactivity but reinforces the overall structural stability and inhibitory activity. It is important to highlight that the linear form of the bacteriocin enterocin AS-48 presented lower inhibitory activity when compared to its circular form15. Such structural advantages render these bacteriocins ideal scaffolds for the design of novel drugs and preservatives for food applications16. The widely studied circular bacteriocin, AS-48, exerts antibacterial activity through interactions of its charged and hydrophobic residues with phospholipids of the target bacterial membrane, thus integrating into and permeabilizing the cell membrane17. In addition, circular bacteriocins, such as circularin A and plantacyclin B21AG, exhibit high antibacterial activity on gram-positive bacteria, such as C. perfringens9,18.
Bacteriocins produced by lactic acid bacteria (LAB) are particularly appealing to the food industry due to the GRAS (generally recognized as safe) status of LAB, which means that the bacteriocins can be applied directly, e.g. nisin, as fermentates, e.g. Microcin® or as safety cultures19,20. Despite this, very few bacteriocins are commercially available as food biopreservatives10. Yet, given the consumer demand for natural alternatives to chemical preservatives in food, there is a need for more bacteriocins with potent activity against foodborne pathogens and that exhibit stability during food processing, Leuconostoc species are members of the LAB group and are prevalent in various foods, such as plants, raw milk, wine, and fermented products. Due to their ability to coexist with Lactobacillus sp., their high tolerance to extreme conditions and their capacity to produce flavour compounds, they are commonly used as dairy starter cultures21. Like other LAB strains, the bacteriocin-producing capacity of Leuconostoc spp. provides the microbes with competitive advantages in different environments. Studies have shown that this genus can produce multiple bacteriocins with different antibacterial spectra. For example, Leuconostoc mesenteroides TA33a produces three leucocins (A-TA33a, B-TA33a and C-TA33a)22. Leuconostoc pseudomesenteroides QU 15 can produce leucocin A-QU 15, leucocin Q and leucocin N23. Previously, one study identified a cyclic bacteriocin, leucocyclicin Q, produced by L. mesenteroides TK41401. Leucocyclicin Q showed a broad inhibition spectrum against various gram-positive bacteria but weakly affected gram-negative strains24.
In this study, we identified and characterized a new variant of leucocyclicin Q, termed leucocyclicin C, produced by Leuconostoc lactis APC 3969 isolated from raw bovine milk collected in bulk. We focused on its inhibitory spectrum against important foodborne and clinical pathogens including C. perfringens, its stability in temperature and pH extremes, and sensitivity to proteases, and we investigated the genes involved in its production. This new variant possesses a broad inhibition spectrum and high resistance to heat and pH adjustments similar to other circular bacteriocins. However, unlike leucocyclicin Q which is reported to be encoded by five genes25, we showed that the biosynthetic gene cluster of leucocyclicin C is composed of 10 genes and is part of a putative composite transposon while the active peptides differ by a single amino acid substitution. This is the first report of a circular bacteriocin produced by L. lactis. Given its stability and potent activity against important foodborne and clinical pathogens, particularly C. perfringens, the characterisation of leucocylicin C in this study will contribute to its development as a novel food biopreservative in the future.
Results
The inhibitory spectrum of Leuconostoc lactis APC 3969
The L. lactis APC 3969 strain was obtained from a screening of antimicrobial-producing bacteria from raw bovine milk collected in bulk samples. Two techniques were used to assess the antimicrobial activity of the strain: the spot-on-lawn assay and the well diffusion assay (WDA). From the 26 indicator strains tested (see Table 1), 16 strains (61.54%) were inhibited as indicated by the presence of clear halos; 5 strains (19.23%) displayed growth reduction with the absence of clear halos; and five strains (19.23%) were not inhibited (Table 1). Both Clostridium strains (Clostridium perfringens EM124 and Clostridium tyrobutyricum DSM 663) were inhibited in the WDA using cell-free supernatant (CFSN) of the producer strain. L. lactis APC 3969 showed a broad inhibitory spectrum capable of inhibiting bacteria from different gram-positive genera, including Staphylococcus, Enterococcus, Streptococcus, Lactococcus, Listeria, and Clostridium. It also inhibited gram-negative microorganisms Escherichia coli and Pseudomonas aeruginosa.
Sensitivity of L. lactis APC 3969 antimicrobial activity against different treatments
The sensitivity of the strain’s antimicrobial activity to different treatments was evaluated using the agar-spot assay (proteases and 0.2 M NaOH) or well diffusion assay (temperatures and pHs), with Lactococcus lactis HP as the indicator strain. The results revealed that the antimicrobial activity produced by the strain is sensitive to proteinase K, which confirmed the proteinaceous nature of the antimicrobial activity. However, the activity resisted treatment with 0.2M NaOH, trypsin and carboxypeptidase A. The cell-free supernatant stability of the strain was stable over a wide range of temperatures (60° to 121 °C) and pH (4.0 to 9.0; Table 2; Fig. S1).
Colony mass spectrometry analysis of L. lactis APC 3969
The colony mass spectrum of the strain L. lactis APC 3969 revealed a high-intensity peak of 6,081.45 Da (Fig. 1), suggesting that the antimicrobial activity may be associated with a compound with this mass.
Colony MALDI-TOF mass spectrum of the bacteriocin producing strain L. lactis APC 3969. The highest intensity peak (6,081.45 Da) is highlighted by the red circle.
Genomic analysis of L. lactis APC 3969
The whole genome of L. lactis APC 3969 was sequenced to identify the genetic cluster in APC 3969 responsible for the observed inhibitory activity. The genome was assembled using Unicycler and then annotated using PGAP. The final assembly resulted in a complete genome with a CG content of 42.5% and a size of 1.952 Mbp (sequence coverage: short reads 83.5x, long reads 63.2x). The final assembly resulted in nine circular elements: one chromosome (1,758,865 bp) and eight plasmid-like elements (49,992 bp, 45,108 bp, 40,861 bp, 18,883 bp, 17,022 bp, 12,732 bp, 7,140 bp and 1,811 bp). The circular extrachromosomal DNA elements were assigned as plasmids based on the best hits when analyzed on the BLAST online platform using nt datbase.
Afterwards, the genome sequence was then submitted to the online platforms Antismash 7.0 and BAGEL4. Both tools revealed the presence of three putative bacteriocin gene clusters: (i) lactococcin 972-related peptide (chromosome), (ii) enteroxin X-related peptide (plasmid-like element pLeuC03), and (iii) leucocyclicin Q-related peptide (plasmid-like element pLeuC01). Other putative core peptides found showed homology to bacteriocin-encoding genes, however they were not considered due to the lack of a putative complete gene cluster (data not shown).
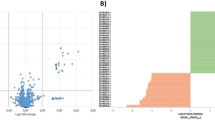
Three gene clusters were analyzed to determine if any encoded a peptide that would be consistent with the highest intensity mass spectrum peak (6,081.45 Da) from the colony mass spectrum. Each core peptide was aligned with its closest related amino acid sequence using Clustal Omega. The alignment of the lactococcin 972-related peptide with lactococcin 972 revealed that both peptides share a similar leader sequence with an Ala-X-Ala cleavage site (Fig. 2A)26. The predicted mature peptide of lactococcin 972 is 7,589.09 Da, which does not match any mass on the colony mass spectrum of APC 3969. The same was observed with the putative peptides found in the plasmid-like element pLeuC03, which were related to enterocin X27, a two-component class II bacteriocin. Neither predicted mass correlated with masses present in the mass spectrum (Fig. 2B, C).
Alignment of the precursor peptides of the putative bacteriocins with lactococcin 972 (A), enterocin Xα (B), enterocin Xβ (C). The leader sequences are highlighted in yellow and the recognition sites for cleavage are highlighted in bold.
CLUSTALW alignment of the leucocyclicin Q-related peptide with leucocyclicin Q revealed that both peptides have a high identity (98.41%) and the same two-amino acid (MF) leader sequence 24. The only difference between the mature peptides is a Threonine to Alanine substitution at position 48 of the leucocyclicin Q-related peptide (Fig. 3A). The predicted mass of the mature leucocyclicin Q-related peptide is 6,099.44 Da. However, since this putative peptide is a variant of leucocyclicin Q, we hypothesize that during the head-to-tail cyclization process, the peptide loses one H2O molecule (18 Da), resulting in a final theoretical mass of 6,081.44 Da. This mass matches precisely with that of the colony mass spectrum and is hereafter named leucocyclicin C. Upstream of the gene encoding the core peptide, the same ribosome binding site (GGAGGA) to leucocyclicin Q was observed in leucocyclicin C (Fig. 3B).
(A) Alignment of the precursor peptides of leucocyclicin Q and leucocyclicin C showing the leader sequences are highlighted in yellow and a single threonine to alanine substitution in the structural peptide highlighted in blue. Identical amino acids (*) and conservative substitutions (:). (B) The nucleotide sequence of the leucocyclicin C propeptide. The cleavage site is highlighted by the dashed line between residues 2 and 3. RBS, putative ribosome binding site – 7 bp upstream of the start codon. *—stop codon.
Next, we investigated the genes responsible for leucocyclicin C production by comparing the genes near the core peptide to the leucocyclicin Q genetic cluster (AB795997.1). Both bacteriocins exhibited the same genetic organization, and the proteins encoded by these genes exhibited ≥ 97.6% identity (Table 3; Fig. 4). In order to differentiate the two genes encoding the core peptide, the gene encoding leucocyclicin C is hereafter named llcA.
Gene clusters of leucocyclicin C and leucocyclicin Q. Data obtained from GenBank (AB795997.1) for leucocyclicin Q.
However, the leucocyclicin C genetic cluster appears to have five additional genes downstream of llcD (llcB2, llcI, llcE, llcF and llcG) in comparison to the leucocyclicin Q gene cluster. These genes may be involved in the biosynthesis of the novel variant (Fig. 4). The first gene, llcB2, encodes a putative membrane protein with low similarity to other membrane proteins found in other genetic clusters of circular bacteriocins including garvicin ML, circularin A, enterocin AS-48, uberolysin, pumilarin, among others. The second gene, llcI, is proposed to encode a putative dedicated immunity protein present in the majority of gene clusters as it shares the same biochemical properties, i.e., small, cationic, and hydrophobic14. Lastly, the following three genes, llcE, llcF and llcG, appear to encode a multi-component ABC transporter, which can be found in other circular bacteriocin genetic clusters (enterocin AS-48, carnocyclin A, circularin A, garvicin ML, cerecyclin and aureocyclicin 4185). The ~8.0 kb leucocyclicin C gene cluster contains ten genes, and the principal properties such as biochemical properties, intracellular localization and highest similarity hit are summarised in Table 4.
Analysis of a putative composite transposon encoding leucocyclicin C
The genetic cluster encoding leucocyclicin C was not detected in 44 other L. lactis genomes deposited in the NCBI database. Sequences upstream and downstream of the cluster were analyzed for mobile genetic elements (MGEs), and the results revealed that the cluster is flanked by two insertion site (IS) 6 family elements, which fits the features of a composite transposon (Fig. 5). Two inverted repeats were identified upstream (GGTTCTGGTGCAAAGTTTA) and downstream (GGTTCTGGTGCAAAGTTGA) of the cluster. In addition, three other open read frames (ORFs) were identified within the borders of the putative composite transposon: (i) the one encodes a putative ATP-binding protein, (ii) the second encodes a DNA invertase protein, and (iii) encodes a DNA/RNA helicase domain-containing protein.
Putative composite transposon that encodes the leucocyclicin C genetic cluster. The leucocyclicin C gene cluster is highlighted by a dashed green box. The inverted repeats are highlighted by a dashed blue box.
Leucocyclicin C purification
To confirm that the strain’s antimicrobial activity is due to the production of leucocyclicin C, the peptide was purified from the cell extract by C18 Solid Phase Extraction and Reversed Phase HPLC fractionation. Well diffusion assays (WDA) with L. innocua DPC 3572 as the indicator strain were used to follow antimicrobial activity throughout purification. Following HPLC fractionation, the WDA assay revealed that the antimicrobial activity was present in fraction 51 (Fig. 6A), while MALDI-TOF mass spectrometry (Fig. 6B) confirmed the fraction contained a 6,081.06 Da mass that correlated with the mass from the colony mass spectrum (6,081.45 Da; Fig. 1), and the theoretical 6,081.44 Da mass of leucocyclicin C.
(A) HPLC analysis of the cell extract of Leuconostoc lactis APC 3969 with fraction 51, highlighted by the red circle. (B) MALDI-TOS MS of fraction 51 showing a mass of 6081.06 Da which correlates with the theoretical mass of leucocyclocin C (6081.44 Da). Note that the 3043 Da mass is doubly charged leucocyclin C.
The minimal inhibitory concentration of leucocyclicin C against C. perfringens EM 124
The minimum inhibitory concentration (MIC) of purified leucocyclicin C was assessed against C. perfringens EM124 in a 23-hour growth experiment (Fig. 7). The MIC value was determined to be 3.288 µM. The growth experiment also revealed that concentrations above 0.822 µM of leucocyclicin C increased the lag phase of the strain. In comparison, concentrations less than 0.822 µM did not affect the growth of the strain when compared with the control (Fig. 7).
Effect of different concentrations of leucocyclicin C on the growth of Clostridium perfringens EM124. The MIC value for the strain is highlighted by the blue circle.
Discussion
The alarming rise of antimicrobial resistance in clinical isolates in the past few years and increased consumer demand for food products without artificial preservatives has boosted research into discovering and developing new antimicrobials28,29. Bacteriocins are perhaps one of the most promising natural antimicrobial alternatives, especially those produced by LAB, due to the GRAS status of the strains.
When compared to other groups of bacteriocins, such as lantibiotics and pediocin-like bacteriocins, circular bacteriocins remain poorly understood14, yet offer particular promise as food biopreservatives due to their structural stability owing to their globular confirmation. In the present study, we identified and characterized leucocyclicin C, a new variant of the circular bacteriocin leucocyclicin Q, produced by the raw bovine milk collected in bulk isolate L. lactis APC 3969. The workflow of this study is presented in the Fig. S2.
In total, 16 of 26 indicator strains were inhibited by leucocyclicin C, and five exhibited reduced growth in its presence. Leucocyclicin C inhibited Enterococcus, Streptococcus, Lactococcus, Listeria and Escherichia, which was also reported for leucocyclicin Q24. In addition, leucocyclicin C could inhibit or reduce the growth of the four S. aureus strains tested24 unlike leucocyclicin Q, which did not inhibit S. aureus subsp. aureus ATCC 1260024. Further studies are needed to verify if the single amino acid substitution enhanced the antimicrobial activity of leucocyclicin C as only one Staphylococcus isolate was assessed for leucocyclicin Q. Leucocyclicin C is thus a broad-spectrum bacteriocin with activity against important food and clinical pathogens (Listeria, Clostridium, Staphylococcus, Enterococcus, Streptococcus, Escherichia, Pseudomonas), which makes this novel variant a promising candidate for future both food biopreservative and even human therapeutic applications.
Leucocyclicin C activity was retained when treated with 0.2M NaOH, confirming that the antimicrobial activity is not due to organic acids. However, the activity was unaffected by carboxypeptidase A and trypsin, and it was sensitive to proteinase K, confirming the proteinaceous nature of the antimicrobial compound. Carboxypeptidases cleave the C terminal amino acid of peptides and so cannot act on circular bacteriocins as they have undergone head-to-tail cyclization. Resistance to carboxypeptidase has previously been used to confirm the circular nature of other bacteriocins, such as enterocin AS-48RJ30 and leucocyclicin Q24.
Lastly, the resistance to trypsin was expected as leucocyclicin Q is also trypsin-resistant. However, this resistance phenotype is not specific to all circular peptides, pumilarin presented partial sensitivity31 and carnocyclicin A antimicrobial activity was eradicated in the presence of trypsin32. Resistance to trypsin is presumably due to the globular conformation of the peptide, making any cleavage sites unavailable to the protease14,33,34.
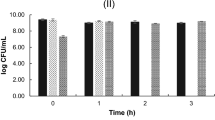
Leucocyclicin C retained all inhibitory activity after 2 h exposure to a range of pH values (4.0 to 9.0) and after 15 minutes exposure to different temperatures (60° to 121°C). This data agrees with the findings for leucocyclicin Q24 and lactocyclicin Q35. This ultra-stability to thermal and pH stresses is a feature of all representatives of this group14.
The genome analysis revealed that leucocyclicin C is encoded by a large bacteriocinogenic cluster of ten genes. The first five genes of leucocyclicin C share the same genetic organization as those encoding leucocyclicin Q, with high levels of identity between them (≥ 97.6%). The other five genes encode a putative membrane protein (llcB2), a putative dedicated immunity protein (llcI) and a putative multi-component ABC transporter (llcEFG). The gene clusters of most circular bacteriocins encode a small, cationic and hydrophobic dedicated immunity protein. The same biochemical characteristics were found for the encoded protein of the gene llcI14. It is important to highlight that a previous study with leucocyclicin Q, Mu and collaborators revealed that a lcyD knockout mutant of the bacteriocinogenic strain L. mesenteroides TK41401 still presents the same high level of immunity against leucocyclicin Q as the wildtype strain. Even if the presence of the gene lcyD alone confers immunity against the circular peptide, the results together suggest that another protein(s) could also confer immunity, in this case, a possible llcI-like gene not identified in the sequenced region of L. mesenteroides TK4140125. Three genes encode a multi-component ABC transporter downstream of the immunity gene, which is potentially related to an accessory function. This hypothesis is supported by studies performed with two other circular bacteriocins, enterocin AS-48 (as-48EFGH) and carnocyclin A (cclEFGH), where the deletion of the corresponding genes in those gene clusters led to a reduction of the immunity of the strain and a concomitant decrease in bacteriocin production36,37. The strain L. mesenteroides TK41401 was not fully sequenced, generating only a fragment of 3.7 kb flanking the core peptide. This may explain why the other five genes downstream of the gene lcyD were not found, as the fragment length did not encompass the entire genetic cluster25. The five extra genes in the leucocyclicin C gene cluster are most likely also present in the genetic cluster of leucocyclicin Q. Future studies are necessary in order to prove the contribution of these five genes in the biosynthetic and immunity of leucocyclicin C.
To determine if the leucocyclicin C genetic cluster is present in other L. lactis genomes, 44 unique genomes from this species were analyzed using the online platform BAGEL4. The results show that none of the 44 strains harbour the bacteriocinogenic cluster found in APC 3969. Thus, the putative function of the ORFs flanking the cluster was investigated against the NCBI database using the BLAST online platform (https://www.ncbi.nlm.nih.gov/)38. Downstream of the genetic cluster, three ORFs can be found: one encoding an ATP-binding protein (WP_216988100.1), the second encoding a DNA invertase Pin (recombinase family protein; WP_228934490.1) and the third encoding a DNA/RNA helicase domain-containing protein (WP_224132774.1). In addition, flanking those 13 genes are two insertion sequences (ISs) 6 family transposases (WP_064520495.1; WP_055307866.1) with respective inverted repeats (GGTTCTGGTGCAAAGTTTA and GGTTCTGGTGCAAAGTTGA). The presence of two ISs flanking other proposed genes is typically associated with a composite transposon39. These transposable elements are known for carrying antimicrobial resistance genes and other types of genes, such as ones related to antimicrobial resistance, in this case, a bacteriocin gene cluster40. However, more studies are necessary to elucidate if this putative MGE is responsible for disseminating this uncommon genetic cluster.
Initially, the colony mass spectrum obtained for L. lactis APC 3969 revealed a mass of 6,081.45 Da, which agrees with the predicted 6081.44 Da mass for leucocyclicin C. The bacteriocin was purified using HPLC to confirm that the antimicrobial activity produced by the strain was indeed due to leucocyclicin C production. Mass spectrometry analysis of the active HPLC fraction verified that the mass of the purified bacteriocin (6,081.06 Da) matches with the mass obtained for the bacterial colony (6,081.44 ± 0.5 Da) and correlates with the theoretical mass of leucocylcin C. Comparison of the mass for leucocyclin C with the theoretical mass for the structural gene reveals a two amino acid leader sequence (MF). In contrast, the 18 Da difference between the theoretical and detected mass confirms that the peptide (6,099 Da) goes through head-to-tail cyclization (6,099 – 18 = 6,081 Da).
As C. perfringens is an important pathogen, the present study focused on the effectiveness of leucocyclicin C against C. perfringens EM124. To our knowledge, the inhibitory activity of circular bacteriocins such as plantacyclin B21AG18, circularin A9 and garvicin ML33 against C. perfringens is unknown since not all bacteriocins were assessed against this strain, and no MIC values have been reported. Thus, the MIC obtained for leucocyclicin C (3.288 µM) against C. perfringens EM124 was compared to published values for bacteriocins from other classes and other antimicrobial peptides. Some show comparable results, such as plectasin, a defensin-like fungal peptide, with an MIC value of 3.64 µM41; sublancin, a glycocin, with an MIC value of 8 µM42; nisin, a lantibiotic, (0.29 – 1.14 µM), MP1102 – 0.91 µM41; thuricin A5, a leaderless bacteriocin, (2.0 µM)43; and ruminococcin C1, a sactibiotic, (0.4 – 0.8 µM)44. This comparison is somewhat limited as these studies assayed different C. perfringens strains, and thus, genetic variations between them could justify the MIC range. However, circular bacteriocins stand out from other bacteriocin classes due to their highly stable nature and desirable biotechnological properties. These desirable characteristics and its potent inhibitory activity make leucocyclicin C a possible new candidate for further investigation as a food biopreservative against C. perfringens and potentially other pathogens and spoilage organisms.
Conclusion
The genus Leuconostoc constitutes a large proportion of the LAB group of bacteria and possesses many desirable biotechnological properties, especially the capacity to produce bacteriocins. This work evaluated and characterized the antimicrobial activity of the strain L. lactis APC 3969, which was isolated from raw bovine milk collected in bulk and can produce leucocylicin C, a novel single-amino acid variant of leucocylicin Q. The circular peptide presented a broad spectrum of activity, inhibiting important pathogens (Listeria, Staphylococcus and Clostridium and gram-negative E. coli and Pseudomonas aeruginosa), high thermal and pH stability, and resistance to trypsin and carboxypeptidase A. Most of these characteristics are shared with leucocyclicin Q. The genetic cluster of leucocyclicin C is bigger than expected, being composed of 10 genes, which makes it, as far we are aware, the biggest genetic cluster described to date for a circular bacteriocin. Interestingly, the bacteriocinogenic cluster also seems to be part of a putative MGE, which may justify why only this genome of L. lactis encodes this bacteriocin. However, future studies will investigate the contribution of each of these genes to bacteriocin production and immunity and determine if the composite transposon can be mobilised to other strains. Lastly, the MIC obtained for this peptide against C. perfringens is compatible with the MIC values found for other antimicrobial peptides. Further investigation is necessary to assess the feasibility of the strain as a novel food biopreservative against C. perfringens and other foodborne pathogens in a food model.
Materials and methods
Antimicrobial activity
The antimicrobial activity of the strain L. lactis APC 3969 was assayed on MRS medium by the agar-spot assay as described by Giambiagi-deMarval et al. (1990) with minor modifications45. Five microliter of an overnight growth culture of the bacteriocinogenic strain was spotted on the surface of MRS agar, allowed to dry at room temperature, and then incubated for 20-24 h at 30°C. After incubation, chloroform was applied to inactivate the cells and the plate was overlaid with 3 mL of soft agar (0.75 % w/v) inoculated with the indicator strain (∼7.0 log CFU/mL). The plates were incubated overnight at optimal growth conditions, as shown in Table 1. The inhibitory activity effect of the producer strain against the indicator was assessed by measuring the true halo of the inhibitory zone (total halo – spot size = true halo) and divided into five categories: growth reduction (hazy zone with reduction of cell density); weak inhibition (0.5–5 mm); moderate inhibition (>5–≤10 mm); strong inhibition (>10 mm) and no inhibition (0 mm). The experiment was conducted in triplicate.
Due to the lack of growth of the Clostridium indicators (C. perfringens EM124 and C. tyrobutyricum DSM 663) on the agar-spot assay, the well diffusion assay was applied as described by Twomey et al. (2021) with minor modifications46. The indicator strain was inoculated (∼7.0 log CFU/mL) in 20 mL of molten agar and then poured into sterile petri dishes. After solidification of the agar, wells of 6 mm width were created in the agar and 50 μL of the neutralized (using NaOH) cell-free supernatant of L. lactis APC 3969 (24 h incubation) was added. Finally, the plates were incubated at the optimal growth conditions, as described in Table 1. The inhibitory effect of the strain was determined by the presence or absence of inhibitory halos around the wells where supernatants were transferred.
Evaluation of the effects of proteases and NaOH on the antimicrobial activity of L. lactis APC 3969
The sensitivity of the antimicrobial activity of the strain L. lactis APC 3969 was evaluated against proteinase K (1 mg/mL; Sigma-Aldrich, St Louis, USA), carboxypeptidase A (1 mg/mL; Sigma-Aldrich, St Louis, USA), trypsin (1 mg/mL; Merck KgaA, Darmstad, DE) and 0.2 M NaOH on agar by the methodology previously described45. Briefly, an overnight growth culture of the strain L. lactis APC 3969 was spotted (5 μL) five times on an MRS (de Man, Rogosa and Sharpe) plate and incubated for 20-24 h at 30°C. Subsequently, the spotted cells were inactivated with chloroform and 40 μL of each treatment (0.2M NaOH or protease) was added around the bacteriocinogenic strain. For the control, 10 mM PBS (pH 7.4) was used as treatment. The plates were air-dried and incubated at 37°C for 4 hours. Lastly, the plates were overlaid with L. lactis HP and incubated overnight at 30°C. The resistance (R) or the sensitivity (S) of the antimicrobial activity against each treatment was determined by measuring the true halo and comparison to the control.
Evaluation of the stability of the antimicrobial activity of the strain L. lactis APC 3969 at different temperatures and pH
The stability of the antimicrobial activity of the strain L. lactis APC 3969 against different temperatures and pH was evaluated by WDA against L. lactis HP as described previously with minor modifications47. For this, a 24 h neutralized (pH 7) CFSN of the producer strain was treated with two different protocols. For the temperature evaluation, the CFSN was exposed to different temperatures (60°C, 70°C, 80°C, 90°C, 100°C and 121°C) for 15 min and then cooled to room temperature and immediately assayed. For the pH evaluation, the CFSN pH was changed to target conditions (pH 4, pH 5, pH 6, pH 8, pH 9, pH 10), using 1M HCl and 1M NaOH, and incubated at room temperature for 2 h. The sample’s pH was then returned to its initial value (pH 7) and immediately assayed using well diffusion assay46. The effect of each treatment was determined by the maintenance or reduction of the size of the halo when compared to the control (neutralized CFSN without any thermic treatment). All the treatments were evaluated on the same Petri dish plate and the assay was performed in duplicate.
Colony MALDI-TOF mass spectrometry
In order to assess the molecular mass profile of peptides associated with L. lactis APC 3969 cells, colony MALDI-TOF (matrix-assisted laser desorption/ionization coupled to time-of-flight) mass spectrometry was applied as previously described by Field et al. (2012) with minor modifications48. Briefly, the strain was streaked on an MRS plate and incubated overnight at 30°C. Fresh colonies from the plate were mixed with 50 µL 70% propan-2-ol + 0.1% trifluoroacetic acid (TFA), vortexed and centrifuged at 21,000 g for 20 seconds. The supernatant was used for the MALDI-TOF mass spectrometry analysis, which was performed with an Ultraflex MALDI-TOF mass spectrometer (Bruker, Bremen, Germany). A 0.5-µL aliquot of matrix solution [α-cyano 4-hydroxy cinnamic acid, 10 mg/mL in acetonitrile-0.1 % (v/v) TFA] was placed onto the target and left for approximately 20 seconds before being removed. The sample supernatant (0.5 µL) was applied to the pre-coated target spot. Afterwards, the matrix solution (0.5 µL) was added to the samples and air-dried. Sample was analyzed in positive-ion reflectron mode, and molecular masses from the spectrum were compared to antimicrobial databases, especially bacteriocin databases.
Genome sequencing and bioinformatics analysis
The bacterial genome of the bacteriocinogenic strain L. lactis APC 3969 was extracted using the GenEluteTM Bacterial Genomic DNA Kit (Sigma-Aldrich, St Louis, USA) as described by the manufacturer. A Qubit 2.0 fluorometer (ThermoFisher Scientific, Waltham, USA) was applied to quantify the genomic DNA. A combination of Illumina and Oxford Nanopore platforms were used to obtain short and long reads. The genome was assembled de novo using the Unicycler 0.5.0v49 from the short reads and long reads. The final assembly was annotated using NCBI Prokaryotic Genome Annotation Pipeline (PGAP)50. In order to assess the coverage of the short and long reads, Bowtie2 (v2.3.4.1), minimap2 (v2.17) and Samtools (v1.7) were utilized51,52,53. The online platforms antiSMASH v.7.0 (https://antismash.secondarymetabolites.org)54 and BAGEL4 (http://bagel.molgenrug.nl)55 were used to identify the presence of genetic clusters related to antimicrobial compounds. To predict the putative function of the ORFs from the antimicrobial genetic clusters, the sequences were analyzed against the NCBI database using the BLAST online platform (https://www.ncbi.nlm.nih.gov/)38. The core gene of each genetic cluster was aligned with its closest relative using Clustal Omega56. The predicted protein and peptides had their molecular mass, isoelectric point (pI), grand average of hydropathicity (GRAVY), and aliphatic index (AI) determined using the online ProtParam (https://web.expasy.org/protparam/)57. The putative cellular localization of the identified protein and peptide was predicted using the online platform PSORTb v3.0.358. Genomic data is available at GenBank/EMBL under accession no. CP157073-CP157081.
Bacteriocin purification
One litre of Leuconostoc lactis APC 3969 culture was grown overnight at 30°C in MRS broth. The culture was then centrifuged at 8,280 g for 20 minutes at 10°C, and cells were separated from the supernatant. The cells were mixed with 250 ml 70% IPA and stirred at room temperature for 3-4 hours with the objective to extract the bacteriocin associated with the cell mass. This cell suspension was then centrifuged, and the supernatant was retained. A rotary evaporator (Buchi Labortechnik AG, Flawil, Switzerland) was used to remove all the IPA from the supernatant before applying it to a 2 g, 12 ml C18 solid phase extraction column pre-equilibrated with methanol and water. The column was washed with 20 ml 25% ethanol and 20 ml 70% IPA. The IPA was removed from an aliquot of the C18 IPA eluent and applied to an analytical Jupiter Proteo (C12, 4.6 x 250 mm, 4 µ, 90 Å) high-pressure liquid chromatography column (Phenomenex, Cheshire, UK) running a 30-60% gradient at 1 ml/min where mobile phase A is H2O + 0.1% TFA and mobile phase B is 100% acetonitrile + 0.1% TFA. The HPLC eluent (2.5 ml/min) was monitored at 214 nm, and fractions were collected at 1-minute intervals. Fractions were assayed on L. innocua DPC3572 using well diffusion assays46, as previously described, and active fractions were evaluated for the antimicrobial mass of interest using MALDI-TOF mass spectrometry.
Minimal inhibitory concentration of leucocyclicin C against Clostridium perfringens EM124
The MIC of the pure leucocyclicin C against approximately 1 × 105 CFU/ml of the target strain C. perfringens EM124 was assayed in a 96-well microtiter plates (Sarstedt, Co. Wexford, Ireland) using a Stratus Microplate Reader (Cerillo, Virginia, USA) to measure the optical density at 600 nm (O.D.600) as previously described48. Initially, 4 times the test concentration (13.155 μM) solution of the peptide was resuspended in 70% propan-2-ol. Then, 100 μL of CRM (Clostridial Reinforced Medium) was added to all wells, followed by 100 μL of the peptide solution to the first well. Afterwards, a two-fold serial dilution was carried out from the first well until the twelfth well. Optical densities at 600nm were collected every hour for 23 h at 37°C in an anaerobic environment. The MIC was determined as the lowest concentration necessary to inhibit the target strain completely. The assay was done in triplicate.
Data availability
The accession number of the L. lactis APC 3969 complete genome is CP157073-CP157081. All other data generated or analyzed in this project are included in this published article. Any additional information or data can be addressed to the corresponding author upon reasonable request.
References
Schneider, K. R., Goodrich-Schneider, R., Hubbard, M. A. & Richardson, S. Preventing Foodborne Illness Associated with Clostridium perfringens: FSHN035/FS101, rev. 1/2014. EDIS, (2014).
Robertson, S., Li, J. & McClane, B. A. Bacteria: Clostridium perfringens. In Encyclopedia of Food Safety 395–402. https://doi.org/10.1016/B978-0-12-378612-8.00092-5 (Elsevier, 2014).
García, S., Vidal, J. E., Heredia, N. & Juneja, V. K. Clostridium perfringens. In Food Microbiology (eds. Doyle, M. P. et al.) 513–540 (ASM Press, Washington, DC, USA, 2019). https://doi.org/10.1128/9781555819972.ch19.
Grenda, T. et al. Clostridium perfringens—Opportunistic foodborne pathogen, its diversity and epidemiological significance. Pathogens 12, 768 (2023).
Rood, J. I. et al. Expansion of the Clostridium perfringens toxin-based typing scheme. Anaerobe 53, 5–10 (2018).
Talukdar, P. K., Udompijitkul, P., Hossain, A. & Sarker, M. R. Inactivation strategies for Clostridium perfringens spores and vegetative cells. Appl. Environ. Microbiol. 83, e02731-e2816 (2017).
Heo, S., Kim, M. G., Kwon, M., Lee, H. S. & Kim, G.-B. Inhibition of Clostridium perfringens using bacteriophages and bacteriocin producing strains. Korean J. Food Sci. Anim. Resour. 38, 88–98 (2018).
Noor Mohammadi, T. et al. Characterization of Clostridium perfringens bacteriophages and their application in chicken meat and milk. Int. J. Food Microbiol. 361, 109446 (2022).
Liu, F., Van Heel, A. J., Chen, J. & Kuipers, O. P. Functional production of clostridial circularin A in Lactococcus lactis NZ9000 and mutational analysis of its aromatic and cationic residues. Front. Microbiol. 13, 1026290 (2022).
Sugrue, I., Ross, R. P. & Hill, C. Bacteriocin diversity, function, discovery and application as antimicrobials. Nat. Rev. Microbiol. https://doi.org/10.1038/s41579-024-01045-x (2024).
Field, D., Fernandez de Ullivarri, M., Ross, R. P. & Hill, C. After a century of nisin research - where are we now?. FEMS Microbiol. Rev. 47, 1023 (2023).
Perez, R. H., Zendo, T. & Sonomoto, K. Circular and leaderless bacteriocins: Biosynthesis, mode of action, applications, and prospects. Front. Microbiol. 9, 2085 (2018).
Sugrue, I., O’Connor, P. M., Hill, C., Stanton, C. & Ross, R. P. Actinomyces produces defensin-like bacteriocins (Actifensins) with a highly degenerate structure and broad antimicrobial activity. J. Bacteriol. 202, e00529–e00619 (2020).
Ladjouzi, R., Dussert, E., Teiar, R., Belguesmia, Y. & Drider, D. A review on enterocin DD14, the leaderless two-peptide bacteriocin with multiple biological functions and unusual transport pathway. Antibiotics 12, 1188 (2023).
Montalbán-López, M. et al. Characterization of linear forms of the circular enterocin AS-48 obtained by limited proteolysis. FEBS Lett. 582, 3237–3242 (2008).
Stevens, C. A. et al. Peptide backbone circularization enhances antifreeze protein thermostability: Peptide Backbone circularization enhances AFP thermostability. Protein Sci. 26, 1932–1941 (2017).
Cebrián, R. et al. The bacteriocin AS-48 requires dimer dissociation followed by hydrophobic interactions with the membrane for antibacterial activity. J. Struct. Biol. 190, 162–172 (2015).
Golneshin, A. et al. Discovery and characterisation of circular bacteriocin plantacyclin B21AG from Lactiplantibacillus plantarum B21. Heliyon 6, e04715 (2020).
Mills, S., Ross, R. P. & Hill, C. Bacteriocins and bacteriophage; a narrow-minded approach to food and gut microbiology. FEMS Microbiol. Rev. 41, S129–S153 (2017).
Mills, S. et al. A multibacteriocin cheese starter system, comprising nisin and lacticin 3147 in Lactococcus lactis, in combination with plantaricin from Lactobacillus plantarum. Appl. Environ. Microbiol. 83, e00799-e817 (2017).
McAuliffe, O. Genetics of Lactic Acid Bacteria. in Cheese 227–247 (Elsevier, 2017). https://doi.org/10.1016/B978-0-12-417012-4.00009-0.
Papathanasopoulos, M. A. et al. Sequence and structural relationships of leucocins A-, B- and C-TA33a from Leuconostoc mesenteroides TA33a. Microbiology 144, 1343–1348 (1998).
Sawa, N. et al. Identification and characterization of novel multiple bacteriocins produced by Leuconostoc pseudomesenteroides QU 15. J. Appl. Microbiol. 109, 282–291 (2010).
Masuda, Y. et al. Identification and characterization of leucocyclicin Q, a novel cyclic bacteriocin produced by Leuconostoc mesenteroides TK41401. Appl. Environ. Microbiol. 77, 8164–8170 (2011).
Mu, F. et al. Biological function of a DUF95 superfamily protein involved in the biosynthesis of a circular bacteriocin, leucocyclicin Q. J. Biosci. Bioeng. 117, 158–164 (2014).
Martı́nez, B., Fernández, M., Suárez, J. E. & Rodrı́guez, A,. Synthesis of lactococcin 972, a bacteriocin produced by Lactococcus lactis IPLA 972, depends on the expression of a plasmid-encoded bicistronic operon The GenBank accession number for the sequence reported in this paper is AJ002203. Microbiology 145, 3155–3161 (1999).
Hu, C.-B., Malaphan, W., Zendo, T., Nakayama, J. & Sonomoto, K. Enterocin X, a Novel Two-Peptide Bacteriocin from Enterococcus faecium KU-B5, has an antibacterial spectrum entirely different from those of its component peptides. Appl. Environ. Microbiol. 76, 4542–4545 (2010).
Yemeke, T., Chen, H.-H. & Ozawa, S. Economic and cost-effectiveness aspects of vaccines in combating antibiotic resistance. Human Vacc. Immunother. 19, 2215149 (2023).
Zapaśnik, A., Sokołowska, B. & Bryła, M. Role of lactic acid bacteria in food preservation and safety. Foods 11, 1283 (2022).
Abriouel, H. et al. Enterocin AS-48RJ: a variant of enterocin AS-48 chromosomally encoded by Enterococcus faecium RJ16 isolated from food. Syst. Appl. Microbiol. 28, 383–397 (2005).
van Heel, A. J., Montalban-Lopez, M., Oliveau, Q. & Kuipers, O. P. Genome-guided identification of novel head-to-tail cyclized antimicrobial peptides, exemplified by the discovery of pumilarin. Microbial Gen. https://doi.org/10.1099/mgen.0.000134 (2017).
Martin-Visscher, L. A. et al. Isolation and characterization of carnocyclin A, a novel circular bacteriocin produced by carnobacterium maltaromaticum UAL307. Appl. Environ. Microbiol. 74, 4756–4763 (2008).
Borrero, J. et al. Characterization of Garvicin ML, a novel circular bacteriocin produced by Lactococcus garvieae DCC43, isolated from mallard ducks (Anas platyrhynchos). Appl. Environ. Microbiol. 77, 369–373 (2011).
Borrero, J. et al. Plantaricyclin A, a novel circular bacteriocin produced by Lactobacillus plantarum NI326: purification, characterization, and heterologous production. Appl. Environ. Microbiol. 84, e01801-e1817 (2018).
Sawa, N. et al. Identification and characterization of lactocyclicin Q, a novel cyclic bacteriocin produced by Lactococcus sp. Strain QU 12. Appl Environ Microbiol 75, 1552–1558 (2009).
Diaz, M. et al. Characterization of a new operon, as-48EFGH, from the as-48 gene cluster involved in immunity to enterocin AS-48. Appl. Environ. Microbiol. 69, 1229–1236 (2003).
van Belkum, M. J., Martin-Visscher, L. A. & Vederas, J. C. Cloning and characterization of the gene cluster involved in the production of the circular bacteriocin carnocyclin A. Probiot. Antimicro. Prot. 2, 218–225 (2010).
Altschul, S. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl. Acids Res. 25, 3389–3402 (1997).
Lipszyc, A., Szuplewska, M. & Bartosik, D. How do transposable elements activate expression of transcriptionally silent antibiotic resistance genes?. IJMS 23, 8063 (2022).
Razavi, M., Kristiansson, E., Flach, C.-F. & Larsson, D. G. J. The association between insertion sequences and antibiotic resistance genes. mSphere 5, e00418–e00420 (2020).
Zong, L. et al. Mechanism of action of a novel recombinant peptide, MP1102, against Clostridium perfringens type C. Appl. Microbiol. Biotechnol. 100, 5045–5057 (2016).
Wang, S. et al. The antimicrobial peptide sublancin ameliorates necrotic enteritis induced by Clostridium perfringens in broilers12. J. Animal Sci. 93, 4750–4760 (2015).
Deng, S. et al. Thuricins: Novel leaderless bacteriocins with potent antimicrobial activity against gram-positive foodborne pathogens. J. Agric. Food Chem. 70, 9990–9999 (2022).
Roblin, C. et al. The unusual structure of Ruminococcin C1 antimicrobial peptide confers clinical properties. Proc. Natl. Acad. Sci. U.S.A. 117, 19168–19177 (2020).
Giambiagi-Marval, M., Mafra, M. A., Penido, E. G. C. & Bastos, M. C. F. Distinct groups of plasmids correlated with bacteriocin production in Staphylococcus aureus. J. General Microbiol. 136, 1591–1599 (1990).
Twomey, E., Hill, C., Field, D. & Begley, M. Recipe for success: Suggestions and recommendations for the isolation and characterisation of Bacteriocins. Int. J. Microbiol. 2021, 1–19 (2021).
Wang, Z. et al. A novel bacteriocin isolated from Lactobacillus plantarum W3–2 and its biological characteristics. Front. Nutr. 9, 1111880 (2023).
Field, D. et al. Bioengineered Nisin A derivatives with enhanced activity against both gram positive and gram negative pathogens. PLoS ONE 7, e46884 (2012).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. https://doi.org/10.1101/096412 (2016)
Li, W. et al. RefSeq: Expanding the prokaryotic genome annotation pipeline reach with protein family model curation. Nucl. Acids Res. 49, D1020–D1028 (2021).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Blin, K. et al. antiSMASH 70: New and improved predictions for detection, regulation, chemical structures and visualisation. Nucl. Acids Res. 51, W46–W50 (2023).
van Heel, A. J. et al. BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucl. Acids Res. 46, W278–W281 (2018).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Duvaud, S. et al. Expasy, the Swiss Bioinformatics Resource portal, as designed by its users. Nucl. Acids Res. 49, W216–W227 (2021).
Yu, N. Y. et al. PSORTb 3.0: Improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26, 1608–1615 (2010).
Acknowledgements
We would like to thank Jim Flynn (Teagasc) for providing the bovine milk collected in bulk samples for this project.
Funding
Co-funded by a research grant from Kraft-Heinz Company, Science Foundation Ireland (SFI) under Grant Number SFI/ 12/RC/2273_P2 and the European Union (ERC, BACtheWINNER, 101054719). Views and opinions expressed are however those of the author(s) only and do not necessarily reflect those of the European Union or the European Research Council. Neither the European Union nor the granting authority can be held responsible for them. This project has received funding from the European Union’s Horizon 2020 Research and Innovation Programme under the INSPIRE COFUND Marie Skłodowska-Curie grant agreement No. 101034270.
Author information
Authors and Affiliations
Contributions
Conceptualization, F.M.F., M.C.S., R.P.R., C.H. and C.S.; Analysis collection, F.M.F., P.M.O., X.H., C.B., E.K., D.H. and M.C.S.; Data analysis, F.M.F., P.M.O. and C.B.; Supervision, R.P.R., C.H., C.S., L.D., J.W., A.D. and O.F.; Project administration, R.P.R., C.H., C.S. and L.D.; Funding acquisition, R.P.R., C.H. and C.S.; Writing, F.M.F. and X.H.; Review and editing, R.P.R., C.H., C.S., P.M.O., C.B., L.D., J.W., A.D. and O.F. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
de Farias, F.M., Soria, M.C., O’Connor, P.M. et al. Leuconostoc lactis strain APC 3969 produces a new variant of cyclic bacteriocin leucocyclicin Q and displays potent anti-Clostridium perfringens activity. Sci Rep 15, 6372 (2025). https://doi.org/10.1038/s41598-025-89450-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-89450-x