Abstract
The clinical significance of phospholipase A2 receptor (PLA2R) antigen domain-specific antibodies has not been determined. We investigated the role of serum PLA2R domain antibodies and epitope spreading in risk stratification and proteinuria remission in primary membranous nephropathy (PMN). Overall, 101 patients with PLA2R-associated MN diagnosed at the Wuxi People’s Hospital between January 2019 and May 2021 were included and divided into low-to-medium risk (n = 12) and high-to-extremely-high risk (n = 89) groups. Serum PLA2R-IgG4 levels were significantly different between the groups, no statistical difference was found in PLA2R-IgG levels. The proportion of IgG4 epitope spreading was greater in the high-to-extremely-high risk group than in the low-to-medium risk group. The result of random forest machine learning method showed that combining the three variables (PLA2R-CTLD678-IgG4, PLA2R-CysR-IgG4, and IgG4 epitope-spreading) showed no significant difference in PMN risk stratification than that of PLA2R-IgG. For predicting proteinuria at 12 months, the ACC values were significantly greater for PLA2R-CTLD1-IgG4, PLA2R-CTLD678-IgG4, and PLA2R-IgG4 than for PLA2R-IgG. Combining IgG4 epitope-spreading and PLA2R-IgG may improve the ACC value of proteinuria prediction after 6 months of treatment compared with PLA2R-IgG alone. PLA2R-CTLD1-IgG4, PLA2R-CTLD678-IgG4, and PLA2R-IgG4 can predict proteinuria remission more accurately at 12 months of treatment than PLA2R-IgG.
Similar content being viewed by others
Introduction
Membranous nephropathy (MN) is an immune-mediated chronic kidney disease, accounting for approximately 30% of primary glomerular diseases, second only to immunoglobulin A nephropathy. The annual incidence of MN has been increasing. The clinical manifestation may be nephrotic syndrome with or without impaired renal function. According to etiological classification, MN can be divided into primary MN (PMN) (or IMN) and secondary MN (SMN). PMN accounts for approximately 80% of cases. PMN shows significant differences in the prognosis, with about one-third of patients in spontaneous remission without treatment, one-third in continuous proteinuria remission after treatment, and one-third showing continuous deterioration of renal function and eventually developing end-stage renal disease with a poor prognosis.
In 2009, Beck1 discovered the first podocyte autoantigen, the M-type phospholipase A2 receptor (PLA2R), in adults with IMN. They also detected PLA2R-specific antibodies in the serum of 70–80% of patients with IMN. Their study confirmed that IMN is an autoimmune disease limited to the kidney, opening a new era in the classification and diagnosis of MN based on autoantigens.
Enzyme-linked immunosorbent assays (ELISAs) are commonly used to detect the concentration of serum PLA2R-IgG antibodies, with a specificity of 99.6% but low sensitivity2. In 2016, Huang3 applied a time-resolved fluoroimmunoassay (TRFIA) to detect PLA2R-IgG antibodies quantitatively, with a diagnostic sensitivity of 74% and specificity of 100%, and could detect patients with weakly positive serum PLA2R. In 2019, Huang et al.4 applied the TRFIA method to detect serum PLA2R-IgG4 antibodies, setting a cut-off value of 161.2 ng/mL. The sensitivity (90%) and specificity (100%) of the diagnosis of PMN indicated that diagnoses based on serum PLA2R-IgG4 specific antibodies were more accurate in PMN. Liu et al.5 applied a random forest prediction model and found that the PLA2R-IgG4 antibodies were superior to the PLA2R-IgG antibodies for predicting disease progression in IMN. The machine learning algorithms have been widely used in pharmacy and biology6,7,8.
PLA2R autoantibodies may target only a specific structural region of the PLA2R protein on the surface of podocyte1,9,10. PLA2R is a type-I transmembrane protein belonging to the mannose receptor family. Its extracellular structure comprises one cysteine-rich domain (CysR), one fibronectin type II domain, and eight C-type lectin domains (CTLD). Seitz-Polski11 found that the CysR, CTLD1, and CTLD7 domains are independently recognized by different anti-PLA2R antibodies. PMN patients with only CysR domain antibodies have a good prognosis; the combined presence of CTLD1 and CTLD7 domain antibodies suggests a poor prognosis. However, the PLA2R epitope can change during follow-up, suggesting a spreading of the PLA2R epitope. As we all known, epitope spreading is an immunological mechanism described in many autoimmune diseases, which physiologically enables a pathogen to be eliminated more effectively12, and the study of epitope spreading in PMN started from the antibody-mediated pathogenesis of active Heymann nephritis13 The clinical significance of PLA2R antigen domain-specific antibodies has not been determined. Reinhard L et al.14 found that all patients with PMN at diagnosis recognized at least two epitope regions in the N- and C-terminals of PLA2R1, and total anti-PLA2R1 antibody levels, but not the epitope-recognition profiles at the time of diagnosis, were relevant for the clinical outcome of patients with PMN. Meanwhile, the reaearch results of Ruggenenti P et al. indicated that anti-PLA2R1 and anti-CysR antibodies similarly predicted rituximab efficacy independent of PLA2R1 domain recognition in patients with PMN15. Whether epitope spreading is related to risk stratification, proteinuria remission, or PMN prognosis is controversial. The answer will help to elucidate the pathogenesis of PMN, develop biomarker detection methods with greater accuracy (ACC), and provide a basis for the early identification of high-risk patients and the implementation of immunotherapy.
Machine learning has revolutionized the healthcare domain, offering powerful tools to analyze medical data and make accurate predictions16,17. By leveraging algorithms and statistical modeling techniques, machine learning can uncover hidden patterns, identify potential diseases, and improve clinical decision-making. Moreover, machine learning algorithms can be used to predict future health outcomes and forecast disease trends. By analyzing historical patient data, these models can estimate the likelihood of disease recurrence, disease progression, and therapeutic outcomes. This predictive capability allows healthcare providers to tailor treatment plans, allocate resources efficiently, and intervene proactively to prevent adverse outcomes.
This retrospective cohort study aimed to investigate the role of serum PLA2R domain antibodies and epitope spreading in disease risk stratification and proteinuria remission in patients with PMN to provide a theoretical basis for early identification of high-risk patients.
Methods
Ethics statements
This study was reviewed and approved by the Ethics Committee of Wuxi People’s Hospital (Kyl2016001). We confirm that our research was performed in accordance with relevant guidelines and regulations. All patients provided written informed consent to participate in the study.
Patients
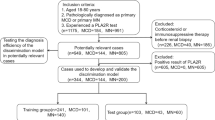
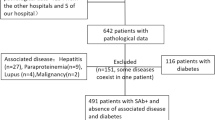
This study enrolled 101 patients diagnosed with PLA2R-associated MN by kidney biopsy at Wuxi People’s Hospital between January 2019 and May 2021, including 50 females and 51 males, with an average age of 51.78 years. This retrospective cohort study excluded patients with tumors, chronic viral hepatitis B or C, or SMN caused by drugs or infections. Among the 101 enrolled patients, 61 completed the 6-month (M6) follow-up and specimen collection, and 27 completed the 12-month (M12) follow-up and specimen collection (Table 1). All patients received angiotensin-converting enzyme inhibitor (ACEI) and angiotensin receptor blocker (ARB)–based integrated renal support before immunosuppressive therapy and risk stratification. All patients treated with glucocorticoids and immunosuppressants were selected according to the Kidney Disease: Improving Global Outcomes (KDIGO) guidelines for managing glomerular diseases. Among them, 8 patients received the RTX(1.0 g q2w × 2), 62 patients chose the CTX (0.8–1.0 g/m) combined with prednisone (0.6–0.8 mg/Kg/d) regimen, and 31 patients chose the tacrolimus (0.05 mg/Kg/d) combined with prednisone (0.6–0.8 mg/Kg/d) (maintaining concentration of tacrolimus was approximately 7 ng/dl).The clinical and laboratory indicators of the 101 patients with PMN were used as independent variables (Table 2).
Materials
The study used the following materials: recombinant expression of PLA2R antigen (Wuxi AORui Dongyuan Biotechnology Co., Ltd., China), goat anti-human IgG antibodies (Jackson ImmunoResearch Co., Ltd., USA), monoclonal human IgG antibodies (Hytest Co., Ltd., China) Finland), recombinant expression of PLA2R-CysR domain antigen (Zhejiang Boshi Biotechnology Co., Ltd., China), recombinant expression of PLA2R-CTLD1 domain antigen (Zhejiang Boshi Biotechnology Co., Ltd., China), recombinant expression of PLA2R-CTLD678 domain antigen (Zhejiang Boshi Biotechnology Co., Ltd., China), China), and the PLA2R-IgG Antibody ELISA kit (Euroimmun, Germany).
Methods
Blood samples were collected from patients with MN at baseline, M6 treatment, and M12 treatment. After resting, the blood samples were centrifuged at 3000 repetitions/min for 4 min, the upper serum was taken, and the data were recorded into the biological sample bank (MN queue) and stored in the refrigerator at − 80 °C. The patients’ clinical data were collected, including sex, age, underlying diseases, blood pressure, serum albumin, renal function (estimated glomerular filtration rate [eGFR] calculated by the CKD-EPI formula), lipid profile, blood cell analysis, C-reactive protein, erythrocyte sedimentation rate, 24-h urinary protein measurement, PLA2R antibodies (ELISA), and other laboratory test results. TRFIA was used to quantify the detection of serum PLA2R-IgG and PLA2R-IgG4 antibodies5. The specific IgG antibodies of serum PLA2R domain antibodies (CysR, CTLD1, and CTLD678) were monitored quantitatively by TRFIA18. The process of the research is shown in Fig. 1.
The research process: Step 1, 101 PLA2R-associated MN patients were recruited, and clinical and laboratory data were collected. Step 2, blood samples were obtained at baseline, 6 months (M6), and 12 months (M12), and patients were categorized into four risk groups. Step 3, a random forest model was employed, with permutation variable importance assessed using the Out-of-Bag (OOB) score.
The main steps of this study are outlined as follows. In step 1, we enrolled 101 patients diagnosed with PLA2R-associated MN by kidney biopsy at Wuxi People’s Hospital between January 2019 and May 2021. Clinical and laboratory indicators were collected (Tables 1 and 2). In step 2, blood samples were collected from patients at baseline (M0), M6 treatment, and M12 treatment. The specific methods and steps of processing blood samples are described later. The patients were divided into four groups according to risk stratification of PMN, which is also described later. In step 3, we used the random forest model as a statistical analysis tool and defined the permutation variable importance measure based on the Out-of-Bag (OOB) score.
Risk stratification of PMN
According to the 2021 KDIGO Guidelines for the Management of Glomerular Diseases, the risk stratification of PMN was as follows:
-
1)
Low risk was defined as normal eGFR, proteinuria < 3.5 g/day, serum albumin > 30 g/L, or proteinuria decreased by > 50% at M6 conservative ACEI/ARB treatment.
-
2)
Medium risk was defined as normal eGFR, proteinuria > 3.5 g, urinary protein > 50% at M6 conservative ACEI/ARB treatment, and other indicators that do not meet the high-risk criteria.
-
3)
High risk was defined as eGFR < 60 mL/min/1.73 m2, and/or proteinuria > 8 g/day for more than 6 months; normal eGFR, proteinuria > 3.5 g/day, proteinuria not decreasing > 50% at M6 conservative treatment with ACEI/ARB, and any of the following conditions were met: serum albumin < 25 g/L; PLA2R-Ab > 50 RU/mL; Urinary ɑ1 microglobulin > 40 µg/mL; urinary IgG > 1 µg/mL; urinary β2 microglobulin > 250 mg/day; IgG clearance/albumin clearance > 0.2.
-
4)
Extremely high risk was defined as life-threatening nephrotic syndrome or a sharp decline in kidney function that cannot be explained by other causes.
Definition of epitope-free spreading
Lack of epitope spreading is defined as the presence of only CysR-specific antibodies in the serum; CTLD1 and CTLD678 specific antibodies test negative. We determined epitope spreading according to each domain’s IgG/IgG4 antibody levels. Epitope spreading was defined as having serum PLA2R-domain antibodies that were positive for CTLD1 and/or CTLD678 in addition to CysR-specific antibodies.
Statistical analysis
Measurement data with a normal distribution are expressed as the mean ± standard deviation, and a comparison between groups was performed using the t-test. Statistical data were compared between groups using the chi-squared test. P < 0.05 was considered statistically significant.
The random forest (RF) algorithm19 is a popular machine learning algorithm for solving tasks such as regression and classification. In general, a RF is constructed by initializing with \(n_{tree}\) subsamples \(\left\{ {{\Theta }_{k} } \right\}\left( {k = 1,2, \ldots ,n_{tree} } \right)\) drawn from the training data set \({\text{M}} = \left( {{\text{M}}_{X} ,{\text{M}}_{Y} } \right)\) using bootstrapping resampling with replacement. These subsamples \(\left\{ {\Theta_{k} } \right\}\) are then used to create individual trees \(h\left( {{\text{X}},{\Theta }_{k} } \right)\) employing either the Classification and Regression Tree or the Conditional Inference Tree algorithm. Accompanying OOB data were obtained for each subsample \({\Theta }_{k}\) by \({\text{B}}_{k} = {\text{M}} - {\Theta }_{k}\) (Fig. 2).
Random forest machine learning algorithm: An overview of the Random Forest workflow, illustrating the ensemble of multiple decision trees and the aggregation of their predictions for robust output.
The permutation variable importance measure (PVIM)20 is defined based on OOB data. The error rate of the \(k\) tree was calculated from the OOB data \({\text{B}}_{k}\) before and after random permuting the values of \(X_{i}\) shown in Eq. (1) and Eq. (2), respectively.
Here, \(\hat{y}_{j}^{\left( k \right)}\) and \(\hat{y}_{j,i}^{\left( k \right)}\) denote the predicted values of the OOB data by the \(k\) tree before and after permuting \(X_{i}\), respectively. The PVIM for \(X_{i}\) in the \(k\) tree is defined as \(PVIM_{i}^{\left( k \right)} = ER_{k,i} - ER_{k}\); the overall PVIM of \(X_{i}\), denoted \(PVIM_{i}\), is determined by averaging \(PVIM_{i}^{\left( k \right)}\) across all trees. \(PVIM_{i}\) measures the average difference between the error rates of the OOB data computed before and after permuting the value of \(X_{i}\). Considering its interaction with other inputs, it can also be interpreted as a contribution of \(X_{i}\) to the model’s prediction ACC. T-distributed stochastic neighbor embedding (T-SNE) is a dimensionality reduction technique commonly used to visualize high-dimensional data in a low-dimensional space while preserving the topological structure of the data. Here, we used T-SNE to visualize the distribution of PMN data (Fig. 3). The data was well distributed, but there were differences in data distribution between groups.
T-distributed stochastic neighbor embedding of variables’ distribution in PMN. PMN: primary membranous nephropathy.
Results
General information of the patients with MN
The study included 101 patients with MN, including 51 males and 50 females, with an average age of 51.78 ± 12.85 years; 60 patients (59.4%) had hypertension and 19 (18.8%) had diabetes. According to the KDIGO guidelines21, the 101 patients with MN were risk-stratified, including 6 patients with low risk, 6 with medium risk, 85 with high risk, and 4 with extremely high risk. The 101 patients were divided into two risk groups: low-to-medium (n = 12) and high-to-extremely-high (n = 89). The 24-h urinary protein level of the low-to-medium group (3.41 ± 1.81 g) was significantly lower than that of the high-to-extremely-high risk group (5.61 ± 1.9 g, P = 0.000). Serum albumin levels were significantly greater in the former group than in the latter group (30.2 ± 4.27 vs. 20.89 ± 3.86 g/L, P = 0.000). The eGFR level in patients with low-to-medium risk group (94.95 ± 14.77 ml/min) showed no statistical difference from patients with high-to-extremely-high risk group (88.52 ± 25.76 ml/min)(P = 0.401) (Table 3).
PLA2R domain antibodies and epitope spreading in patients with different risk levels
Analysis of serum PLA2R-IgG and PLA2R-IgG4 levels (Table 3) showed no significant difference between the level of serum PLA2R-IgG in low-to-medium (65.15 ± 96.37) and high-to-extremely-high (158.83 ± 199.37) risk groups (P = 0.113). However, the levels of serum PLA2R-IgG4 were significantly different between the two groups (2295.52 ± 3774.05 vs. 6503.96 ± 6940.21, P = 0.043). Further analysis showed that the levels of serum PLA2R-CTLD678-IgG4 (2548.87 ± 6198.85 vs. 273.12 ± 597.78, P = 0.001) and domain IgG (65/89 vs. 5/12, P = 0.027) and the proportion of IgG4 epitope spreading (74/89 vs. 6/12, P = 0.001) were greater in the high-to-extremely-high risk group. No significant differences between the groups were found in serum PLA2R-CysR-IgG (30.39 ± 38.83 vs. 87.74 ± 150.49, P = 0.194) PLA2R-CysR-IgG4 (725.45 ± 1380.93 vs. 2194.48 ± 6495.32, P = 0.068), PLA2R-CTLD1-IgG (16.17 ± 23.60 vs. 43.09 ± 84.39, P = 0.276) and PLA2R-CTLD1-IgG4 (305.11 ± 852.08 vs. 478.83 ± 1181.35, P = 0.628).
Evaluation of PMN risk stratification
The RF-OOB algorithm was used to evaluate the effect of clinical features (31 variables) on the classification ACC of PMN risk stratification. OOB scores quantified the importance of each variable in PMN risk stratification (Fig. 4).
Predictive value of each parameter for risk stratification. PVIM: permutation variable importance measure.
The analysis showed that the ACCs of CTLD678-IgG4, CysR-IgG4, and IgG4 epitope spreading for PMN risk stratification were less than that of PLA2R-IgG. When the three variables were combined, no significant difference was evident between the combination and PLA2R-IgG levels (P = 0.8863) (Table 4).
Prediction of proteinuria remission in PMN
At M6 treatment, 18 patients had a complete response, 56 had a partial response, and 37 had no response, with a response rate of 73.3%. The OOB algorithm was used to analyze the predictive significance of each variable for proteinuria remission at M6 treatment. The OOB score of IgG4 epitope spreading (0.261) was not significantly different from that of PLA2R-IgG (0.194) at M6 (P = 0.689) (Fig. 5). Further, we used RF to perform a leave-one-out-cross-validation (LOOCV)22 and report the receiver operating characteristic curve (AUC) value. LOOCV involves training a model on one iteration of the dataset and validating it on a separate iteration, leaving out one observation or feature at a time. This repeated process provides an estimate of the model’s performance under unseen data. The area under the receiver operating characteristic curve (AUC) indicates a variable’s performance. No significant difference was found between the AUCs of IgG4 epitope spreading (0.58) and PLA2R-IgG (0.62) in proteinuria remission at M6 (Fig. 6).
Predictive value of each parameter for proteinuria remission at the M6 follow-up. PVIM: permutation variable importance measure, M6: 6-month.
Comparison of AUC values for predicting proteinuria remission at the M6 follow-up. Scr: serum creatinine, X22: IgG4 Epitope spreading; X24: PLA2R-IgG, AUC, area under the curve, M6: 6-month.
At M12 treatment, 26, 21, and 10 patients showed complete, partial, and no remission, respectively. The RF-OOB algorithm analysis showed that PLA2R-CTLD1-IgG4 (ACC = 0.405, P = 0.010), CTLD678-IgG4 (ACC = 0.407, P = 0.011), and PLA2R-IgG4 (ACC = 0.444, P = 0.011) were significantly better than PLA2R-IgG in predicting proteinuria remission at M12 treatment (Table 5, Fig. 7).
The predictive value of each parameter for proteinuria remission at the. M12 follow-up. Scr: serum creatinine, M12: 12-month.
For PLA2R-IgG, PLA2R-IgG4, CTLD1-IgG4, and CTLD678-IgG4, the AUC values for predicting the proteinuria remission at M12 treatment were 0.74, 0.76, 0.73, and 0.78 respectively (Fig. 8).
Comparison of AUC values for predicting proteinuria remission at the M12 follow-up. X24: PLA2R-IgG, X25: PLA2R-IgG4, X29: CTLD1-IgG4, X31: CTLD678-IgG4, AUC: area under the curve, M12: 12-month.
Comparison with other classifiers
We employed RF as a statistical analysis tool because compared with other classifiers, RF has a higher classification ACC. Here, we evaluated the performance of RF and other well-known classifiers, including support vector classification (SVC)23, K-nearest neighbor (KNN)24, and logistic regression (LR)25. The ACC and area under the curve (AUC) of LOOCV are used to evaluate performance. To calculate the AUC for three-class classification of the M6 and M12 dataset, we used the one-versus-rest approach. The results are displayed in Table 6 and Fig. 9 (Table 6, Fig. 9).
Comparison of AUCs achieved by RF, SVC, KNN, and LR using the M6 dataset under LOOCV. LOOCV: leave-one-out-cross-validation, RF: random forest, SVC: Support vector classification, KNN: K-nearest neighbor, LR: logistic regression, AUC: area under the curve, M6: 6-month.
Table 6 showed that RF achieved the best ACC using the three datasets. RF shows ACC values of 6.05%, 3.7%, and 0.99% over the second best SVC on M6 treatment (Fig. 9), M12 treatment (Fig. 10) and M0 (Fig. 11), respectively. Figures 9, 10, and 11, RF also achieves the best AUC using the three datasets. This further emphasizes the superior performance of the RF model in terms of both ACC and AUC. In summary, RF offered robustness, interpretability, and scalability advantages, making it suitable for our study.
Comparison of AUCs achieved by RF, SVC, KNN, and LR using the M12 dataset under LOOCV. LOOCV: leave-one-out-cross-validation, RF: random forest, SVC: support. vector classification, KNN: K-nearest neighbor, LR: logistic regression, AUC: Area under the curve, M6: 6-month.
Comparison of AUCs achieved by RF, SVC, KNN, and LR using the MN. dataset under LOOCV. LOOCV: leave-one-out-cross-validation, RF: random forest, SVC: support. vector classification, KNN: K-nearest neighbor, LR: logistic regression, AUC: Area under the curve.
Discussion
PMN is an autoimmune disease mediated by different autoantigens. PLA2R accounts for 70–80% of the pathogenic antigens26. Circulating PLA2R autoantibodies do not bind to the admissible PLA2R antigen to form immune complexes, indicating that PLA2R autoantibodies may only produce immune responses against specific conformational regions of PLA2R antigens on the surface of kidney podocytes27. PLA2R-specific IgG4 antibodies, the major subtype of PLA2R-IgG antibodies, are central to PMN pathogenesis28. By analyzing four risk models, Liu et al.5 confirmed that PLA2R-IgG4 was superior to PLA2R-IgG in PMN risk stratification. Our analysis found the serum titers of PLA2R-IgG4 (P = 0.043), and PLA2R-CTLD678-IgG4 (P = 0.001) were greater in the high-to-extremely-high risk group than in the low-to-medium risk group. The incidence of IgG4 epitope spreading was greater, whereas the antibody titers of PLA2R-IgG, PLA2R-CysR-IgG, and PLA2R-CTLD678-IgG were not significantly different between the groups. Besides, in our study, we found differences in PLA2R-IgG4 antibodies between the two groups, indicating that PLA2R-IgG4 is more sensitive than PLA2R-IgG in differential expression. The same is true CTLD678-IgG4 and IgG4 epitope spreading also could be potential biomarkers for risk stratification in PMN patients. Therefore,we believe that serum PLA2R-IgG4 antibodies have a greater clinical value than serum PLA2R-IgG in assessing risk stratification in PLA2R-MN.
Seitz-Polski29 discovered that the CysR, CTLD1, and CTLD7 domains of the PLA2R antigen extracellular segments are independent domains recognizable by different anti-PLA2R antibodies and associated with different clinical prognoses. However, the PLA2R domain-specific antibodies have inconclusive value, and whether epitope spreading is associated with risk stratification and PMN prognosis is controversial by conventional statistical analysis methods. The RF-OOB algorithm was used to screen for clinical and laboratory indicators that were significant for risk stratification. Their analysis showed that PLA2R-IgG (OOB score: 0.2816) ranked below serum urea nitrogen and serum creatinine in PMN risk stratification, followed by PLA2R-CTLD678-IgG4 (OOB score: 0.2202), PLA2R-CysR-IgG4 (OOB score: 0.1977), and IgG4 epitope spreading (OOB score: 0.1690). The immunodominant epitope of the PLA2R antigen in patients with PMN is located in the extracellular CysR domain30. PLA2R epitopes spread from the N-terminal CysR domain to the C-terminal CTLD7 domain; patients with PMN diseases targeting different epitopes have different prognoses31. Besides PLA2R-IgG and PLA2R-IgG4, which statistical values in risk stratification and prognosis previously more documented, PLA2R-CTLD678-IgG4 and epitope spreading also have clinical value in risk stratification in patients with PMN. As shown in the results of this study, compared with those in the low-to-medium risk group, the serum titer of PLA2R-CTLD678-IgG4 antibodies and the incidence of IgG4 epitope spreading in the high-to-extremely-high risk group were greater. In addition, the combined variables PLA2R-CTLD678-IgG4, PLA2R-CysR-IgG4, and IgG4 epitope spreading had similar efficacy in assessing risk stratification in PMN compared with that of PLA2R-IgG alone. Therefore, we have reasons to believe that combining together these three variables (PLA2R-CTLD678-IgG4, PLA2R-CysR-IgG4 and IgG4 epitope spreading) will probably bring potential value in assessing risk stratification in PMN.
Seitz-Polski discovered29 that epitope spreading beyond the CysR epitope was an independent risk factor for poor renal prognosis. Epitope spreading during follow-up was associated with disease worsening, which was consistent with our findings. Therefore, we speculated that epitope spreading occurs from CysR to CTLD1 and CTLD678, indicating that autoantibodies of CTLD1 or CTLD678 may indicate epitope spreading. Consequently, PLA2R-domain antibodies are potential novel biomarkers for PMN risk assessment.
For the PLA2R-IgG, PLA2R-IgG4, and domain antibodies, we applied the RF-OOB algorithm to predict proteinuria remission at M6 and M12 PMN treatment. At M6 treatment, IgG4 epitope spreading was similar to PLA2R-IgG for predicting proteinuria remission (OOB score: 0.2614 vs. 0.1492, P = 0.6897). We speculated that the predictive effectiveness of the combination (epitope spreading and PLA2R-IgG) would be more accurate than that of PLA2R-IgG alone. Huang et al.32 showed that the PLA2R-IgG4 antibody levels at initial treatment had the best performance for predicting complete proteinuria remission at M6 in PMN. PLA2R-CTLD1-IgG4 was better at predicting proteinuria remission at M12 treatment (OOB score: 0.3605) than PLA2R-IgG4 (OOB score: 0.2642), PLA2R-CTLD678-IgG4 (OOB score: 0.1769), and PLA2R-IgG (OOB score: 0.0781). In our study, PLA2R-CTLD1-IgG4, PLA2R-IgG4, and PLA2R-CTLD678-IgG4 were significantly better than PLA2R-IgG at predicting the proteinuria remission at M12 treatment (Table 5). Seitz-Polski33 also reported that a low epitope spreading rate at the initial treatment was associated with the albuminuria remission rate at M6 treatment in the GEMRITUX and NICE cohorts.
This study was limited by a small sample size, which may have caused insignificant differences between groups; these differences require confirmation by prospective cohort studies with larger sample sizes. However, we believe that PLA2R epitope spreading can clinically predict proteinuria remission at M6 treatment. Serum PLA2R-CTLD1-IgG4 level is a potential novel biomarker for predicting proteinuria remission at M12 treatment.
Conclusions
Compared to PLA2R-IgG alone, the combination of PLA2R-IgG4, PLA2R domain antibodies and PLA2R-IgG could bring more hints for precise risk stratification in PMN. IgG4 epitope spreading, PLA2R-CTLD678-IgG4, and PLA2R-IgG4 are potential biomarkers for predicting proteinuria remission in PMN and could help in early identification of high-risk patients with PMN and provide effective interventions.
Data availability
The original contributions presented in this study are included in the article. Further inquiries can be directed at the corresponding author.
References
Beck, L. H. et al. M-type phospholipase A2 receptor as target antigen in idiopathic membranous nephropathy. N. Engl. J. Med. 361, 11–21 (2009).
Cheng, G. et al. Serum phospholipase A2 receptor antibodies and immunoglobulin G subtypes in adult idiopathic membranous nephropathy: Clinical value assessment. Clin. Chim. Acta. 490, 135–141 (2019).
Huang, B. et al. Improvement of idiopathic membranous nephropathy diagnosis with ultrasensitive quantitative detection of anti-phospholipase A2 receptor. J. Allergy Clin. Immunol. 139, 1988-1990.e2 (2017).
Huang, B. et al. Phospholipase A2 receptor antibody IgG4 subclass improves sensitivity and specificity in the diagnosis of idiopathic membranous nephropathy. Kidney Blood Press. Res. 44, 848–857 (2019).
Liu, X. et al. A PLA2R-IgG4 Antibody-based predictive model for assessing risk stratification of idiopathic membranous nephropathy. J. Healthc. Eng. https://doi.org/10.1155/2021/1521013 (2021).
Ding, Y., Guo, F., Tiwari, P. & Zou, Q. Identification of drug-side-effect association via multiview semisupervised sparse model. IEEE Trans. Artif. Intell. 5, 2151–2162 (2024).
Ding, Y., Tiwari, P., Guo, F. & Zou, Q. Multi-correntropy fusion based fuzzy system for predicting DNA N4-methylcytosine sites. Inf. Fus. 100, 101911 (2023).
Ding, Y., Tiwari, P., Zou, Q., Guo, F. & Pandey, H. M. C-Loss based higher order fuzzy inference systems for identifying DNA n4-methylcytosine sites. IEEE Trans. Fuzzy Syst. 30, 4754–4765 (2022).
Kao, L., Lam, V., Waldman, M., Glassock, R. J. & Zhu, Q. Identification of the immunodominant epitope region in phospholipase A2 receptor-mediating autoantibody binding in idiopathic membranous nephropathy. J. Am. Soc. Nephrol. 26, 291–301 (2015).
Gu, Y., Xu, H. & Tang, D. Mechanisms of primary membranous nephropathy. Biomolecules. 11, 513 (2021).
Seitz-Polski, B. et al. Epitope spreading of autoantibody response to pla2r associates with poor prognosis in membranous nephropathy. J. Am. Soc. Nephrol. 27, 1517–1533 (2016).
Li, N., Aoki, V., Hans-Filho, G., Rivitti, E. A. & Diaz, L. A. The role of intramolecular epitope spreading in the pathogenesis of endemic pemphigus foliaceus (fogo selvagem). J. Exp. Med. 197(11), 1501–1510 (2003).
Shah, P., Tramontano, A. & Makker, S. P. Intramolecular epitope spreading in heymann nephritis. J. Am. Soc. Nephrol. 18, 3060–3066 (2007).
Reinhard, L. et al. Clinical relevance of domain-specific phospholipase a2 receptor 1 antibody levels in patients with membranous nephropathy. J. Am. Soc. Nephrol. 31, 197–207 (2020).
Ruggenenti, P. et al. Anti-phospholipase a2 receptor 1 and anti-cysteine rich antibodies, domain recognition and rituximab efficacy in membranous nephropathy: A prospective cohort study. Am. J. Kidney Dis. 83, 588–600 (2024).
Qian, Y., Ding, Y., Zou, Q. & Guo, F. Identification of drug-side effect association via restricted Boltzmann machines with penalized term. Br. Bioinform. 23, bbac458 (2022).
Qian, Y., Ding, Y., Zou, Q. & Guo, F. Multi-view kernel sparse representation for identification of membrane protein types. IEEE/ACM Trans. Comput. Biol. Bioinform. 20, 1234–1245 (2023).
Zhang, Q. et al. Ultrasensitive quantitation of anti-phospholipase A2 receptor antibody as a diagnostic and prognostic indicator of idiopathic membranous nephropathy. Sci. Rep. 7, 12049 (2017).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Carrizosa, E., Molero-Río, C. & Romero Morales, D. Mathematical optimization in classification and regression trees. Top (Berl). 29, 5–33 (2021).
Rovin, B. H. et al. KDIGO 2021 clinical practice guideline for the management of glomerular diseases. Kidney Int. 100, S1–S276 (2021).
Wong, T. T. Performance evaluation of classification algorithms by k-fold and leave-one-out cross validation. Pattern. Recognit. 48, 2839–2846 (2015).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297 (1995).
Guo, G., et al. KNN model-based approach in classification. In on the move to meaningful internet systems 2003: CoopIS, DOA, and ODBASE. LNCS. 2888, 986–996 (2003).
Lever, J., Krzywinski, M. & Altman, N. Logistic regression. Nat. Methods. 13, 541–542 (2016).
Jullien, P. et al. Anti-phospholipase A2 receptor antibody levels at diagnosis predicts spontaneous remission of idiopathic membranous nephropathy. Clin. Kidney J. 10, 209–214 (2017).
Li, Y., Yu, J., Wang, M., Cui, Z. & Zhao, M. H. Anti-phospholipase A2 receptor antibodies directly induced podocyte damage in vitro. Ren. Fail. 44, 304–313 (2022).
Haddad, G. et al. Altered glycosylation of IgG4 promotes lectin complement pathway activation in anti-PLA2R1-associated membranous nephropathy. J. Clin. Invest. 131, e140453 (2021).
Seitz-Polski, B. et al. Phospholipase A2 Receptor 1 Epitope spreading at baseline predicts reduced likelihood of remission of membranous nephropathy. J. Am. Soc. Nephrol. 29, 401–408 (2018).
Fresquet, M. et al. Identification of a major epitope recognized by PLA2R autoantibodies in primary membranous nephropathy. J. Am. Soc. Nephrol. 26, 302–313 (2015).
Qin, Y. et al. Quantitative detection of anti-PLA2R antibodies targeting different epitopes and its clinical application in primary membranous nephropathy. Clin. Chem. Lab. Med. 61, 251–259 (2023).
Huang, Y. et al. PLA2R-IgG4 antibody as a predictive biomarker of treatment effectiveness and prognostic evaluation in patients with idiopathic membranous nephropathy: a retrospective study. PeerJ. 10, e14193 (2022).
Seitz-Polski, B. et al. High-dose rituximab and early remission in pla2r1-related membranous nephropathy. Clin. J. Am. Soc. Nephrol. 14, 1173–1182 (2019).
Funding
This work was supported by the Wuxi Medical Innovation Team Project (CXTD2021010), the Top Talent Support Program for young and middle-aged people of the Wuxi Health Committee (HB2023009), and the Cohort and Clinical Research Program of Wuxi Medical Center, Nanjing Medical University (WMCC202316).
Author information
Authors and Affiliations
Contributions
X. L. and L. W. designed the experiments. X. Z., F. Y., and B. H. performed most experiments. X. L., Y. F., and J. X. analyzed the experimental data. B. L., Z. Z., W. S., and H. C. revised the manuscript. X. L., L. W., L. Z., and T. C. performed clinical analyses. All the authors contributed to the manuscript and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
This study was reviewed and approved by the Ethics Committee of Wuxi People’s Hospital (Kyl2016001). All patients provided written informed consent to participate in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, X., Yang, F., Fan, Y. et al. Role of PLA2R domain antibodies and epitope spreading in risk stratification and prediction of proteinuria remission in primary membranous nephropathy. Sci Rep 15, 6868 (2025). https://doi.org/10.1038/s41598-025-91366-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-91366-5