Abstract
This study explores the population structure, hybridization, and adaptation of Juniperus communis sensu lato in the context of its current habitat fragmentation, using inter-primer binding site markers and needle morphometry. Three native juniper taxa in Slovakia were analyzed: J. communis ssp. communis, J. communis ssp. nana, and their putative hybrid, J. communis nothovar. intermedia. The results reveal a moderate overall structure (ΦPT = 0.115, p = 0.001), indicating high gene flow within ssp. communis and low gene flow between ssp. communis and ssp. nana. However, after correcting for non-neutral loci, the inter-subspecies differentiation dropped significantly, suggesting adaptive divergence despite substantial gene flow. This conclusion was further supported using admixture regression of needle morphometry and ΦPT–QST comparison. We show that adaptive genetic differentiation may play a more significant role than phenotypic plasticity in these traits, a key consideration in preventing outbreeding depression in conservation management. Importantly, we also provide evidence for intermediate admixture and distinct selection pressures within two populations of nothovar. intermedia (Stolica, Kralova Studna), which may qualify their recognition as evolutionarily significant units (ESUs).
Similar content being viewed by others
Introduction
Habitat loss and fragmentation are important factors threatening biodiversity. This situation is usually associated with the isolation and decline of plant populations, leading to restricted gene flow, pollination failure, the Allee effect, genetic drift, and high inbreeding levels1. As a result, small populations may experience an increase in homozygosity and frequency of deleterious alleles, reducing overall population fitness and adaptability1,2,3,4. Preservation of these small populations is important as they may hold historical genetic diversity necessary for species resilience or speciation5,6.
Previously, most conservation studies of endangered species have focused on herbaceous and insect-pollinated species (e.g.5,7,8). Only a few studies have evaluated long-lived and wind-pollinated woody species with potentially high gene flow through pollen and seed dispersal4,9. For such species, the genetic effects of habitat fragmentation may not be apparent for a long time10. A good example is common juniper (Juniperus communis L. sensu stricto), a wind-pollinated and bird-dispersed evergreen shrub of the Northern Hemisphere. As a pioneer woodland species, it occupies natural rocky outcrops and other habitats with skeletal soils and abundant sunlight11. The species has the widest distribution range of all conifers but is strongly declining in North Atlantic and Central European countries, resulting in small and fragmented populations12. Probably, the most significant problem is the age-related decline in fertility, i.e., low regeneration by seeds13,14. However, the underlying mechanisms remain unresolved. Genetic studies did not reveal any genomic erosion that might explain the phenomenon well13,14,15,16; therefore, other explanations were suggested.
In reality, the genetic effects of habitat fragmentation are a long-term issue. During population decline, genetic drift and inbreeding cause deleterious mutations to be more frequent and homozygous, but this process can take many generations to become apparent after the cause of the decline17. This can be particularly important when considering the typical juniper generation time. Several authors, in this vein, have suggested that assisted gene flow interventions may be necessary for this species in the long term1,15,16. Additionally, species resilience could also be improved by utilizing subspecies hybridization. Because introgression can greatly surpass both mutation fitness effects and intra-subspecies gene flow, intentional admixture may be a powerful conservation tool for common juniper, which often exist in small and senescent populations. This strategy is, however, relevant only to systems where the risk of outbreeding depression is low18. It highly depends on a detailed genetic structure and admixture analysis, followed by fitness monitoring.
Our study is designed for the first stage of the process. It includes three native subtaxa: (i) the shrub-like J. communis L. var. communis (hereafter treated as ssp. communis, syn. ssp. eu-communis Syme, var. arborescens Gaud., var. montana Nielr.-non-Ait., var. vulgaris Ait.); (ii) the procumbent J. c. var. saxatilis Pall (hereafter treated as ssp. nana (Hook.) Syme, syn. ssp. alpina (Suter) Čelak., var. montana Ait., J. sibirica Burgsd.); and (iii) their putative hybrid J. communis nothovar. intermedia (Schur) Nyman (syn. J. c. ssp. communis var. intermedia Sanio). However, the hybrid nature of the latter was postulated based only on habitus and needle consistency19, and no attempts have been made to investigate the genetic structure of the three groups included. Although some needle morphometric differences were found between the parents, nothovar. intermedia did not show any major phenotypic differentiation from var. communis20. In general, the authors found a stronger association of the phenotype with ecology than phylogeny, suggesting that plasticity is more important than genetic ancestry for these traits.
Therefore, our questions are as follows: (i) does the study species exhibit high population genetic differentiation, low intrapopulation genetic diversity, and high inbreeding levels in Slovakia; (ii) are the populations isolated by distance; (iii) does population size affect genetic diversity and inbreeding; (iv) what is the extent of inter-subspecies genetic differentiation, admixture, and selection; and (v) is genetic ancestry less significant in determining phenotypic divergence in needles relative to the environment (if not, are these genetic differences for the traits adaptive or comparable to genetic drift)?
Material and methods
Sampling
In total, nine sampling sites of J. communis L. in Slovakia were subjected to population genetic analysis. These represented four pure reference stands of ssp. communis (Zahrada, Cervena Skala, Priechod, Dubniky) and one reference stand of ssp. nana (Kralova Hola). The remaining four locations contained several or few individuals of nothovar. intermedia (Sumiac, Besnik, Stolica, Kralova Studna). All (sub)populations are assumed to be natural according to the botanical mapping by Futák et al.19, Tocl21, and other relevant authorities. The location information of the sampling sites is given in Table 1.
As a source material, we used 1-year-old needles collected from May to August 2022. Special attention was devoted to standardizing the needles’ age for morphometric analysis. The sampling was performed randomly with respect to the individual phenotypes. During the sampling, the individual stands were also taxonomically identified based on their habitus and needle consistency (Fig. 1), providing a preliminary assessment of the genetic structure (Table 1). After harvesting, the collected needles were immediately measured and stored at − 81 °C until DNA extraction. A voucher specimen was deposited to the Slovak Academy of Sciences herbarium (SAV0019117, SAV0019118, SAV0019119, SAV0019120).
Typical habitus of sampled juniper individuals. (a) ssp. communis, (b) ssp. nana, and (c) nothovar. intermedia. Photo: Andrej Kormuťák.
Needle morphometry
Three phenotypic traits were selected for quantitative genetic analysis: needle length (L), needle width (W), and the L/W ratio. The data were analyzed for six locations (C/Za, C/Du, CI/Su, CI/Be, I/KS, N/KH), with N = 11 to 42 (133 in total, with ten replicates per individual). The needles were photographed using a Leica MZ10 F microscope equipped with Leica 10446275 PLAN 1.0× lens and a DFC 420 C camera. Leica Application Suite EZ (LAS EZ) 4.6.1 software (Leica, Switzerland; http://www.leica-microsystems.com) was used for this purpose, utilizing a zoom factor of 0.8x. The traits were subsequently measured by the image analysis software Fiji 2.15.122.
DNA extraction
DNA was extracted according to the CTAB protocol described by Murray and Thompson23.
PCR amplification and genotyping
Genotyping was performed by inter-primer binding site (iPBS) amplification with ten primers designed by Kalendar et al.24. These primers were chosen according to the PCR efficiency evaluated in that study. The (single-primer) reaction mixtures consisted of ~ 120 ng of DNA, 1 × B2 buffer, 2 mM MgCl2, 0.2 mM dNTP, 0.55 µM primer, 1 U of HOT FIREPol® DNA Polymerase (Solis BioDyne), and PCR Grade water (Solis BioDyne), for a total volume of 25 µL. The PCR program was initiated by a polymerase activation step at 95 °C for 15 min, followed by 35 cycles of 95 °C for 15 s, 51–63.3 °C (Supplementary Table S1) for 60 s, and 72 °C for 60 s, with a final extension at 72 °C for 5 min. Amplification was carried out in a TProfessional Gradient Thermocycler (Biometra). Products were analyzed in 1.7% agarose gels with 1 × TBE buffer and ethidium bromide (EtBr). The gels were run at 90 V for 7.5 h in a cold room and were scanned by a BioDoc-It (UVP). Bands (loci) were identified using a 100 bp DNA Ladder (Solis Biodyne) and scored for their presence (1) or absence (0).
Reliability of the iPBS data
Before the data analyses, several precautions were taken to ensure the data reliability. First, samples of poor DNA integrity, i.e., fragmented DNA, were identified and discarded from genotyping based on 1% agarose gels with EtBr (Supplementary Fig. S1). Second, individual PCRs were tested based on intragel replicates for the repeatability of the DNA profiles (Supplementary Fig. S2–S4). Third, samples exhibiting odd profiles, i.e., with many extra or poorly visible/absent bands, were run multiple times as intergel replicates. If these replicates were not repeatable, the samples were also discarded. At the locus level, information from intra- and intergel replicates was used to clean up the binary data matrix from unstable (non-repeatable) marker loci. Fourth, negative controls were utilized in each fourth gel to check for exogenous contamination. Fifth, the whole genotyping process was performed with gel images rendered in GIMP 2.10.825 according to Fattal et al.26 to reduce potential bias associated with intergel differences in overall intensity (Supplementary Fig. S5–S9). Additionally, to remove scoring subjectivity, band presence/absence was scored semiautomatically in GelAnalyzer 19.127 using the rolling ball background subtraction. No other manipulations, such as removing monomorphic loci, were performed.
Data analysis
Since common juniper can reproduce clonally, we first tested the dataset for clonality using the method ‘Find clones’ in GenAlEx 6.51b228. However, as only two putative clones from different locations were suggested, they were not removed from the analyses.
Genetic structure and gene flow were quantified in several ways. First, population differentiation was calculated with Weir and Cockerham’s FST analog (ΦPT) using 999 AMOVA permutations in GenAlEx. This value was converted according to Wright’s formula Nm = (1 − ΦPT)/(4ΦPT) to provide an indirect estimate of gene flow. Alternatively, we used the Maximum Likelihood population reallocation test for individuals in FAMD 1.3129, with a minimum log-likelihood difference (MLD) of 1 and 2. Second, we calculated Cavalli-Sforza and Edwards’s chord distances between locations (DCE) using Phylip 3.69830, as they provide more reliable measures for both isolation-by-distance and clustering analyses31,32,33,34,35. The isolation-by-distance effect was tested by regressing DCE on geographic distances (ArcMap 9.336) using the Mantel test with 9,999 permutations (GenAlEx). Third, we assessed inter-subspecies differentiation, or marker informativeness, for phylogenetic clustering analysis. To do this, we calculated null allele frequencies using the Bayesian approach with a nonuniform among-population prior (correction factor = 0.01) (FAMD). Inter-subspecies differentiation was subsequently quantified as Gregorius and Roberds’s absolute allele frequency difference (Dj) between the weighted means of ssp. communis and ssp. nana. Following Halder et al.37, markers with a Dj value (i.e., Shriver’s δC) higher than 0.3 were considered ancestry informative (AIMs). The data distribution was tested for normality by the Shapiro‒Wilk test38.
For clustering analyses, we used the galled network algorithm in Dendroscope 3.8.1039. The network was based on Neighbor-Joining midpoint-rooted gene trees (from Phylip) estimated for AIMs only, with a consensus threshold of 24%. At the individual level, the samples were clustered in STRUCTURE 2.3.440 according to the admixture model (LOCPRIOR) with correlated allele frequencies (all loci were included). The number of assumed clusters was K = 1 to 10, with ten replicates for each K. The burn-in length and Markov chain Monte Carlo iterations were set to 100,000 and 500,000, respectively. The optimal K was determined by the Evanno41 and Puechmaille methods42 in StructureSelector43, and the consensus was provided by the LargeKGreedy algorithm (CLUMPAK)44.
Next, we conducted admixture regression analysis using a partially linear semiparametric model (the npplreg function from the np package, R45), as suggested by Connor and Fuerst (‘Model 3’)46. The aim was to quantify the genetic component of phenotypic variation in the needle L/W ratio across different ancestries. It is important to note that this characteristic should not be confused with heritability, which is a different measure. To check for significant phenotypic differences among ancestry groups, we performed a mixed model nested ANOVA, followed by Tukey’s post hoc test (the lmer and glht functions, R47,48).
Intrapopulation genetic diversity was analyzed in AFLP-SURV 1.049, using the Bayesian method with non-uniform prior distribution of allele frequencies. The parameters included Nei’s gene diversity (Hj, or expected heterozygosity) and the percentage of polymorphic loci (PPL) at the 5% level corrected for sample size. Moreover, we estimated the inbreeding coefficient (F) with the dominant-marker-based program FAFLPcalc50. To investigate the genetic hypothesis of habitat fragmentation, all these measures (Hj, PPL, F) were tested for Spearman’s rank correlation51 with census population size. As for selection, we carried out two neutrality tests: Tajima’s D and Fu’s FS statistics, using 16,000 simulations in Arlequin 3.552. The first test compares the average pairwise difference between haplotypes within a sample (e.g., subpopulation) with the same measure but is expected based on the number of segregating loci assuming neutrality (both should be the same unless selection, population nonstationarity, or heterogeneity of mutation rates among loci occur). The second test is based on the probability of observing k or more alleles in a sample, conditioned on the observed average pairwise difference. It is especially sensitive to population expansion or departure from equilibrium and is significant only if p < 0.02. To identify putatively non-neutral loci, we used the FST approach with the Shapiro‒Wilk test.
Natural selection was also tested explicitly for the three phenotypic traits of interest (L, W, and the L/W ratio). This was done by ΦPT–QST comparison, where genetic differentiation in a phenotypic trait (QST) was compared with genetic differentiation based on neutral markers (ΦPT). To estimate QST, we used an approximation by PST (phenotypic differentiation) calculated with Pstat53. The accuracy with which PST approximates QST depends on the ratio of narrow-sense heritability among the groups (c) to the average narrow-sense heritability within the groups (h2), i.e., c/h2. These parameters were estimated using the rrBLUP method, a genomic prediction approach with a linear mixed model54. In this model, the phenotypic values (P) were regressed on ssp. communis ancestry (Q) as fixed effects, while accounting for additive relatedness (A matrix) as random effects. The A matrix was calculated by multiplying the REAP kinship coefficient55 by two, assuming an inbreeding level of 0.1 for the kinship calculation. Finally, the resulting variance components were used to estimate the additive genetic proportion of phenotypic variation, σu2/(σu2 + σe2), both within and between subpopulations (via the mean P, Q, and A).
Results
Genetic structure and gene flow
The final dataset included 116 marker loci and nine samples of J. communis (298 individuals). Overall, there was significant and moderate genetic structure (ΦPT = 0.115, p = 0.001). Most differences were found between the reference ssp. nana (N/KH) and the ssp. communis samples (C/Za, C/CS, C/Pr, C/Du) (ΦPT = 0.229–0.234), with pairwise ΦPT values corresponding to low differentiation within the latter (ΦPT = 0.014–0.053) (Supplementary Table S2).
The overall level of gene flow was estimated to be Nm ≈ 1.9. Similarly, the assignment test showed that two individuals (0.7%) were at least 100 times (i.e., MLD = 2) more likely to originate from a location different from their sampling site, which increased to 24 individuals (8.1%) when using only a tenfold difference (i.e., MLD = 1). Within ssp. communis, the number of migrants varied from Nm ≈ 4.5 to 17.6(ΦPT method) or from 0(MLD = 2) to 11(MLD = 1) (11.2%). We found no evidence for isolation by distance in this group using the Mantel test (r2 = 0.36, p = 0.164) (J. communis sensu lato was not tested due to potential bias from inter-subspecies genetic differentiation).
The distribution of inter-subspecies differentiation across loci (Dj) is shown in Fig. 2. The histogram indicates that the data deviated from normality significantly (Shapiro–Wilk test: W(116) = 0.69, p < 0.001). Of the nine outliers identified, eight markers were also ancestry informative (Dj > 0.3) and thus used for phylogenetic network analysis.
Distribution of marker information content for ancestry across iPBS loci. The X-axis shows Gregorius and Roberds’s allele frequency difference (Dj) between four ssp. communis locations and one ssp. nana location. The dashed line represents our threshold for considering markers to be ancestry informative.
The resulting network demonstrated a clear distinction between the two subspecies, separating N/KH (the most divergent) from the reference samples of ssp. communis. These samples clustered together, however, forming a paraphyletic group. The clade also included CI/Be with a close relationship to CI/Su. Only CIN/St and I/KS were found to originate from hybridization between the main clusters (Fig. 4).
The observed pattern was further corroborated by the STRUCTURE results. The ΔK approach, often considered limited to the uppermost structure, strongly supported two clusters present in the data (ΔK > 53; see56) (Fig. 3a), which clearly differentiated between ssp. communis and ssp. nana. The average membership varied between Q = 0.967–0.994 in the reference ssp. communis locations, while the corresponding Q = 0.008 in N/KH (Fig. 3c). A fine-scale population structure was revealed by the Puechmaille estimators (0.5 threshold). These statistics favored K = 3 (MedMed K, MedMea K) or K = 4 as the less robust solution (MaxMed K, MaxMea K) (Fig. 3b), showing further structuring within ssp. communis. All scenarios were highly consistent among clustering runs (Fig. 3c). Notably, K = 3 was supported as the next possible option also by the ΔK method, due to highest mean log-likelihood. However, because the study focuses primarily at the taxonomic (subspecies) level, not population membership within, K = 2 was selected for further discussion. In other words, these ancestry estimates should better reflect the proportion of genes inherited from ssp. communis and ssp. nana.
Determination of the optimal K value. (a) The Evanno method: ΔK and mean log-likelihood with standard deviation. (b) The Puechmaille method: Median-of-Median (MedMed K), Median-of-Mean (MedMea K), Max-of-Median (MaxMed K), and Max-of-Mean (MaxMea K). The optimal K is indicated by red lines. (c) Best clustering results from CLUMPAK for all outcomes (K = 2, 3, and 4), with consistency among runs.
The highest amount of admixture was observed in the locations of nothovar. intermedia. Unlike the galled network, the STRUCTURE analysis detected admixture also in CI/Be and CI/Su. The frequency of intermediates, defined by a range of Q = 0.4–0.6, was highest in I/KS (0.800). Conversely, individuals from CI/Be and CIN/St primarily represented the introgressive form of ssp. communis (Q = 0.6–0.8), with the frequencies of 0.964 and 0.673, respectively. The least admixture among the nothovar. intermedia locations was found in CI/Su (Fig. 4).
Genetic structure of juniper in Slovakia as inferred by galled network and STRUCTURE analysis. The galled network shows a generalization of Neighbor-Joining gene trees estimated for eight ancestry informative iPBS markers (Dj > 0.3) using Cavalli-Sforza and Edwards’s chord distances. The STRUCTURE plot shows individual admixture proportions for the uppermost structure detected by the ΔK approach. For samples information, see Table 1. The map was created in ArcMap 9.3 (https://www.arcgeo.sk/produkty/gis-pre-desktop/).
The admixture regression results are shown in Fig. 5. The purpose of this analysis was to model the total needle area, as measured by the L/W ratio, and to examine the relationship and explanatory power of genetic ancestry for this trait. As indicated by the plot, we found a substantial genetic component to the phenotypic variation (R2 = 0.502). The ancestry-phenotype relationship was mostly linear among the genotypes connecting ssp. nana and intermediates. However, from the intermediate form to pure ssp. communis, the predicted phenotypic values plateau nonlinearly, with no systematic increase in the trait.
Admixture regression of needle length/width ratio in two juniper taxa and their hybrid. The parental forms can be defined by the intervals of 0.0–0.2 (ssp. nana) and 0.8–1.0 (ssp. communis), while the rest corresponds to nothovar. intermedia. The red line represents predicted values of the trait based on a partially linear semiparametric model.
This was also supported by nested ANOVA combined with Tukey’s post hoc test using all four ancestry groups (C, CI, I, N). For the two traits that were significantly or almost significantly differentiated (p(L) = 0.003, p(L/W) = 0.098; but p(W) = 0.491), the lowest p values were found in the N–C and N–I comparisons, while the I–C was highly non-significant (p(L) = 0.001, p(L/W) = 0.115; p(L) = 0.137, p(L/W) = 0.262; vs. p(L) = 0.630, p(L/W) = 0.999). In addition, CI was more similar to I than to C for these two traits (p(L) = 0.938, p(L/W) = 0.662; vs. p(L) = 0.132, p(L/W) = 0.405).
Intrapopulation diversity and selection
Out of all iPBS loci, 50.9% were polymorphic in both ssp. nana and, on average, in ssp. communis (36.2–73.3%). Similar values were observed across the studied locations, except for I/KS, where PPL reached 82.8%. No such variations were noted in the Hj and F parameters. The average inbreeding level ranged from − 0.086 in I/KS to 0.087 in C/Pr (Table 2). We found no significant correlation between any of these variables and subpopulation size (p(PPL) = 0.746, p(Hj) = 0.948; but p(F) = 0.063 for r(7) = 0.641).
Tajima’s D provided no evidence of selection or demographic nonstationarity (p(D) = 0.528–0.861). In contrast, Fu’s FS values, which are more powerful statistics57, were all negative and significant (p(Fs) < 0.02). The most negative values were found in N/KH, CI/Su, and, unexpectedly, in CIN/St, a location characterized by the most even admixture distribution (FS = − 24.137, − 24.022, − 24.196). For these subpopulations, the census size was estimated at 500 (CI/Su, CIN/St) or 1,000 individuals (N/KH). Smaller subpopulations (C/CS, C/Du, CI/Be, I/KS) had the least negative FS values (from FS = − 6.325 to − 14.487) (Table 3).
In conclusion, the tests indicate demographic expansion and/or selective sweeps. Locus-specific ΦPT analysis revealed 13 outliers that deviated from the other 103 loci (Shapiro–Wilk test: W(116) = 0.65, p < 0.001). The outliers varied between ΦPT = 0.183–0.645, with eight corresponding to the AIMs identified earlier (Fig. 2).
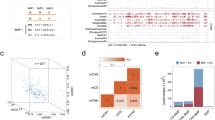
Apart from the population genetic approach, evidence for adaptive genetic differentiation also comes from the ΦPT–QST comparison (Fig. 6). In these plots, we compared the neutral ΦPT (Supplementary Table S3) with the PST calculated for the three needle traits, each of which was presented as a function of c/h2 (rather than just a single value; this is because how well PST approximates QST depends on accurate heritability estimates, which are often difficult to obtain in natural environments). The plots illustrate the potential QST values and 95% confidence intervals, depending on different assumptions of c/h2. All traits, especially the needle length and the L/W ratio, were found to be genetically highly differentiated relative to the neutral ΦPT baseline. The PST(L) and PST(L/W) were significantly high across almost all the plotted c/h2 values (from ≈ 0.1; according to our rrBLUP heritability estimates, c/h2(L) = 0.972/0.459, c/h2(W) = 1.000/0.522, and c/h2(L/W) = 0.951/0.436) (Fig. 6).
Comparison of phenotypic and neutral genetic differentiation in juniper subpopulations. Needle length (PST(L)), needle width (PST(W)), and the needle L/W ratio (PST(L/W)) were measured. The green line represents neutral ΦPT, the red line represents PST as a function of the between-subpopulation/within-subpopulation heritability ratio (c/h2), and the brown lines depict 95% confidence intervals. The blue dots show the values corresponding to the assumed c/h2 estimated by ridge regression-Best Linear Unbiased Prediction (rrBLUP), thus the approximations of QST.
Discussion
Population structure and differentiation
Small fragmented populations are generally assumed to have restricted gene flow and, consequently, high genetic differentiation1,2,3,4. For J. communis, it was first reported in Great Britain58 based on amplified fragment length polymorphism (AFLP) data. Although no summary statistics were provided (e.g., FST, ΦPT), the authors suggested ‘clear structuring’ and restricted gene flow between the studied populations. The same conclusion was drawn by Provan et al.59, who used nuclear microsatellite (nSSR) markers and found ‘significant differences’ between J. communis populations in Ireland. However, the level of differentiation was only moderate (ΦST = 0.096), which is relatively comparable with the average estimate for biparentally inherited markers in conifers (FST = 0.11660). A similar structure was also found in our study of J. communis sensu lato from Slovakia (ΦPT = 0.115). For J. ssp. communis subpopulations, there was low or no differentiation (ΦPT = 0.014–0.053, p = 0.001–0.115), and only two genetic populations were detected with STRUCTURE: one corresponding to ssp. communis (with a minor cluster within) and the other to ssp. nana. These results are consistent with other nuclear marker data on fragmented populations from continental Europe.
For instance, low to moderate differentiation was observed in the Netherlands (allozymes ΦPT = 0.02613), Germany (nSSR ΦPT = 0.02515), Belgium (nSSR ΦPT = 0.03116), Northwestern Europe (AFLP ΦPT = 0.10014), and between Slovakia, Italy, and Norway (nSSR FST = 0.021–0.03115). Using the same markers, moderate differentiation was found even across the entire Eurasian distribution of J. communis sensu lato from Middle Europe to the far east of Russia and Alaska (nSSR ΦPT = 0.09861). Thus, there seems to be no indication of increased isolation among fragmented European populations compared to large populations of Asia. In fact, it is this wide scale where genetic distances were found to be significantly correlated with geography (r = 0.574, p = 0.0161). We found no spatial structure of ssp. communis in Slovakia, and there is also no isolation by distance according to geographically more extensive studies from continental Europe13,14,15,16. The same applies to the Irish study, where the authors nonetheless concluded restricted gene flow59.
Gene flow between populations
In population genetics, the common rule of thumb predicts that the average level of gene flow necessary for maintaining intrapopulation diversity while allowing divergence among populations (i.e., drift-migration equilibrium) is one migrant per generation62. However, even though the model does not hold for real-world populations and much higher values are suggested to be appropriate for conservation management purposes (e.g., Nm = 1–1063), the connectivity in J. c. ssp. communis appears to be still high, despite fragmentation. In Germany, for example, Reim et al.15 reported the average Nm to be 9.9. Similarly, our estimates between individual ssp. communis populations were 4.5–17.6 migrants per generation. These values are also comparable with the average norm estimated for conifers64, Nm > 3. On the other hand, the population group of J. communis sensu lato (i.e., shrub-like, procumbent, and intermediate forms included) is much less connected, as indicated by the Nm = 1.9. The actual connection is probably even lower given that these estimates are indirect and do not separate between historical and contemporary gene flow. Consequently, because the FST approach assumes many unrealistic conditions, such as constant population size, random migration, no selection, mutation, or spatial structure, the Nm values are advised to be interpreted with caution65,66.
A more direct approach is the population assignment test considering individuals’ age. In their study, Vanden-Broeck et al.14 detected from 3%(MLD=2) to 14%(MLD=1) migrants, with the majority having a shrub height smaller than 2 m, suggesting a quite high rate of recent dispersal in northwestern Europe (ssp. communis). According to our estimates, we may postulate a similar gene flow intensity for populations in Slovakia (up to 11.2%), but it is approximately half that value in J. communis sensu lato (0.7–8.1%). It is worth mentioning that 0.7–5.4% of migrants in this larger group represent dispersal events with some contact zone, and none of them directly connects the two different allopatric stands. In other words, most gene flow occurs within the taxonomic boundaries of these groups or, when hybridizing, in specific habitats.
Intrapopulation diversity
Genetic variation within populations also seems relatively high. Microsatellite estimates of allelic/genotypic diversities (i.e., HE/HO) in Europe are, on average, directly comparable to those in Asia, where most populations do not experience anthropogenic fragmentation. This trend also holds for the inbreeding coefficient, which is high in all cases (HE/HO/FIS = 0.602/?/?59, 0.84/0.59/0.2915, 0.751/0.451/0.38616; vs. 0.708/0.440/0.355 in Eurasia61). Interestingly, Hantemirova and Bessonova61 reported high F values even for the most numerous, undisturbed populations of Yamal (0.337) and Polar Urals (0.313), suggesting the presence of null alleles and/or the Wahlund effect. However, we believe that all SSR-based inbreeding values are likely overestimated by null alleles only, as the Wahlund effect relates to FST or FIT, not to F calculated for individual sampling sites. For instance, Vanden-Broeck et al.14 estimated the average Nei’s gene diversity (Hj) and inbreeding (FIS) for AFLP markers to be 0.361 and 0.172 in Northwestern Europe, which are about half the values mentioned above. In Slovakia, we observed similar diversity (Hj = 0.193) and no inbreeding (FIS = − 0.019) using iPBS. These findings indicate that dominant-marker based FIS may be even more informative than those from SSR studies, despite certain limitations in their estimation (such as violation of FAFLPcalc assumptions, potential band scoring errors, or non-independence between loci; see50).
Natural selection and the causes of population decline
Overall, most studies provide no support for genomic erosion in J. communis, and the species, due to its adaptability and large global population, is not included on the IUCN Red List11. However, recent population fragmentation in many European countries (e.g., Britain67,68, the Netherlands13, Germany70, Spain71) has led to its inclusion in Annex I of the EU Habitat Directive (code 5130). The species is now considered locally threatened, which is frequently attributed to various factors such as climate change72, urbanisation, and intensification of agriculture68,73. These factors are assumed to be directly responsible for population fragmentation, resulting in a highly skewed age structure toward old individuals in some areas74. Like many other conifers, juniper produces a large portion of empty seeds, which requires an appropriate quantity75 and diversity of pollen76 for successful fertilization and cone development. With a decrease in the size and number of young individuals, populations may further suffer from limited sexual regeneration75. There were only 1.73–18.98% filled seeds and 0.10–5.49% seed viability reported from northwestern Europe14. In Belgium, seed viability ranges from 0 to 30%, with the highest germination rate of only 8.9%16. It is believed that overgrazing by livestock combined with limited sexual reproduction has worsened the impacts on juniper68.
Apart from the short-term threats discussed above, fragmented juniper populations may suffer from genomic erosion due to inbreeding and drift in subsequent generations. However, the process usually takes many generations and may not be apparent for a long time. According to a pink pigeon simulation study17, this time lag of decline in neutral diversity was estimated to be approximately 100 years, and changes in the genetic load took even twice as long to register. The average generation time of this bird was assumed to be 3.6 years. Common juniper, by contrast, has an average generation span of 20 years11, suggesting a delay of over 550 and 1,100 years, respectively. Worth noting is that junipers are dioecious and wind-pollinated species with a tendency toward high gene flow and intrapopulation diversity4,9,77. It is therefore not surprising that research on common juniper has not shown the genetic impact of habitat fragmentation yet. Our data also indicate no correlation between population size and genetic diversity, as evidenced by other studies as well13,14,16. Only one study revealed a significant positive correlation between population size and inbreeding16. Moreover, both genetic diversity and inbreeding appear unrelated to the seed germination rate16 or seed viability14, i.e., fitness, indicating no ‘realized’ load of deleterious mutations.
Indeed, the situation in common juniper can best be explained by a time lag of evolutionary genetic effects of habitat fragmentation, sometimes referred to as ‘drift debt’78. Basically, this process leads recently bottlenecked populations to lose genetic variation due to the absence of mutation-drift-selection equilibrium. Because the level of diversity at balancing selection genes unlikely reflects the actual selection regime in such populations, this variation will be partially lost while moving toward a new equilibrium under a 'no-change’ scenario. This is likely why our FS values are all negative and significant (p < 0.02), suggesting demographic expansion and/or selective sweeps (i.e., selection removing genetic diversity). We conclude that the drift-debt phenomenon is the most plausible explanation for these seemingly contradicting results in the fragmentation context.
Conservation strategies and management
A good understanding of the genetic dynamics of declining populations is of particular interest to conservation biologists. The growing awareness of time-delayed extinction (extinction debt) associated with habitat fragmentation also highlights the urgency of various conservation measures79,80. The consensus is to push against drift and inbreeding in a process called genetic rescue. For successful rehabilitation of common juniper, several authors have suggested bridging large distances between stands by ‘stepping stone’ populations or individuals in favorable open areas using different source populations15,16. These measures are intended for both the short- and long-term issues but should only be considered ‘first aid’. It would be more ideal to use genomics-informed management, that is, to identify individuals with the lowest masked load for translocations and help populations counteract both inbreeding depression (realized load) and potential inbreeding depression (masked load)81.
With respect to assisted gene flow interventions, we further suggest considering both genetic admixture and local adaptation in natural J. communis populations. According to our study, adaptive differentiation may play an important role in the evolution of species’ infraspecifics, including ssp. communis and ssp. nana. We found no direct gene flow between the pure reference stands, and there is also evidence for directional or divergent selection according to both population (the FS values) and quantitative genetic analyses. In woody plants, needle morphology is crucial for differentiating congeneric taxa82 and is widely recognized as an altitudinal adaptation associated with more compact leaves in harsher environments83; however, there is little information on how much this variation in J. communis is explained by genetic ancestry20. Our study revealed a substantial genetic component of needle morphometry variation and provided evidence for high genetic differentiation for these traits relative to neutral differentiation. Consequently, we propose that intentional admixture of ssp. communis and ssp. nana would be suitable only in the typical habitat of nothovar. intermedia to avoid potential outbreeding depression.
Taxonomic Aspect
Finally, we would like to address a few concluding remarks regarding taxonomy. While the main goal of the study was to contribute to the discussion on the conservation status of common juniper from the perspective of the Slovakian territory, delineating species boundaries is a key component of wildlife conservation84. Not only the listing of what deserves the attention but also the very management decisions depend on taxonomy–for example, whether gene flow should be prevented or facilitated85,86. This question fully relies on the accepted taxonomy, as intraspecific gene flow is often regarded as beneficial in contrast to the interspecific. Unfortunately, species boundaries are arbitrary, and because there is no unique species definition or delineation procedure, different conservationists may come to different recommendations based on the same data87. Therefore, we feel the taxonomic aspect requires a comment.
Juniperus communis L. is a variable species with not very consensual infraspecifics. According to Thomas et al.12, it includes the following:
-
1.
J. communis ssp. communis—a spreading shrub or small tree, rarely procumbent, needles mostly 8–20 × 1–1.5 mm, loosely set, sharp-pointed; usually occurring on calcareous soils but largely indifferent to the soil type
-
2.
J. communis ssp. nana—a procumbent shrub (ca. 10 cm high), needles mostly 4–12 × c. 1.5 mm, closely set, blunt-pointed; restricted to well-drained bogs or rocky outcrops in higher altitudes
-
3.
J. communis ssp. hemisphaerica (J. & C. Presl) Nyman—intermediate; Mediterranean and North-West African mountains
-
4.
J. communis ssp. depressa (Pursh) Franco—intermediate; North America
Some authors also refer to ssp. nana by its original name, J. sibirica, separating it as a distinct species. This name was first mentioned by Burgsdoff in 1787 but was later accepted as a synonym of the subspecies nana (Hook.) Syme 186888. In addition, substantial overlap between ssp. communis and ssp. nana in both phenotypic traits (wood anatomy89, needle tip90, monoterpene content91, volatile distillates92) and genetic variation (chloroplast DNA91, allozymes93, RAPD94) led some specialists to further downgrade all infraspecific taxa into varieties. Plants of the World Online database, for instance, introduces J. communis var. saxatilis Pall. as the correct name for ssp. nana; however, some differentiation from the nominate was also observed95, and Flora Europaea lists the taxon as J. communis ssp. alpina (Suter) Čelak.
So, which taxonomic treatment should we follow? Galtier87 suggested that one way toward a more robust, objective, and reproducible taxonomy could be a kind of reference system. Species delineation might be achieved by comparing a particular group with reference taxa, where species boundaries are consensual and large amounts of data are available. In conifers, this might be Pinus sylvestris and Pinus mugo sensu lato, which have allways attracted the attention of European taxonomists. For instance, Łabiszak and Wachowiak96 sequenced 48 nuclear loci (794 SNPs) and reported that the level of neutral divergence between P. sylvestris and P. mugo sensu stricto, which are closely related but distinct species, was ΦPT = 0.260 (the authors found no signatures of selection or reproductive isolation at these markers). Between even closer entities, this value reached ΦPT = 0.068–0.088 for P. mugo vs. P. uncinata and ΦPT = 0.056 for P. uncinata ssp. uncinata vs. P. uncinata ssp. uliginosa. The population-level differentiation, calculated for a wide distribution of P. sylvestris using nSSR, varied between 0.028 and 0.03397.
Therefore, our estimates suggest that J. communis ssp. communis and J. communis ssp. nana indeed correspond to the subspecies ranks, rather than mere varieties (neutral ΦPT = 0.081–0.103, Supplementary Table S3). Our data also confirms that J. communis nothovar. intermedia (e.g., the populations Kralova Studna and Stolica) is a hybrid between these taxa and should be considered an evolutionarily significant unit (ESU). This is supported by the STRUCTURE results and the galled network that illustrates differences in selection in the absence of neutral differentiation. Generally speaking, gene flow appears sufficient to prevent genetic differentiation at neutral loci but insufficient to prevent genetic differentiation for adaptive phenotypic traits. However, common garden or reciprocal transplant experiments would be critical for further examination of this observation.
Conclusions
To our knowledge, no genetic study has yet analyzed the population structure and hybridization of J. communis var. communis, J. communis var. saxatilis, and J. communis nothovar. intermedia. Therefore, this study represents the first investigation into these important biological aspects of the species for conservation management. Consistent with other European studies, we found that var. communis is likely well connected by gene flow in Slovakia, and the populations are not genetically eroded compared to large, undisturbed populations in Asia. The time since habitat fragmentation began was probably too brief to have any genetic consequences for the species; however, the genetic load may still compromise population viability in the future. Furthermore, our findings provide the first evidence for both adaptive and non-adaptive divergence between var. communis and var. saxatilis (or rather ssp. communis and ssp. nana), including the support for the hybrid nature of nothovar. intermedia. It is, therefore, essential to adopt appropriate conservation measures that consider both local adaptation and taxonomic boundaries. The question of source material for plant translocation can be addressed through genomics-informed management, by identifying specific individuals for interbreeding, such as those with the lowest genetic load. Since only SSR and PCR-based markers have been employed thus far, further research utilizing population genomics is imperative.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Frankham, R., Ballou, J. D. & Briscoe, D. A. A Primer of Conservation Genetics (Cambridge University Press, 2004).
Aguilar, R., Ashworth, L., Galetto, L. & Aizen, M. A. Plant reproductive susceptibility to habitat fragmentation: review and synthesis through a meta-analysis. Ecol. Lett. 9, 968–980. https://doi.org/10.1111/j.1461-0248.2006.00927.x (2006).
Angeloni, F., Ouborg, N. J. & Leimu, R. Meta-analysis on the association of population size and life history with inbreeding depression in plants. Biol. Conserv. 144, 35–43. https://doi.org/10.1016/j.biocon.2010.08.016 (2011).
Vranckx, G., Jacquemyn, H., Muys, B. & Honnay, O. Meta-analysis of susceptibility of woody plants to loss of genetic diversity through habitat fragmentation. Conserv. Biol. 26, 228–237. https://doi.org/10.1111/j.1523-1739.2011.01778.x (2012).
Van Geert, A., Van Rossum, F. & Triest, L. Perspectives for genetic rescue of the extremely fragmented Primula vulgaris populations in The Netherlands: reflecting the future of Belgian populations?. Plant. Ecol. Evol. 148, 329–334. https://doi.org/10.5091/plecevo.2015.1101 (2015).
De Vriendt, L. et al. Population isolation shapes plant genetics, phenotype and germination in naturally patchy ecosystems. J. Plant Ecol. 10, 649–659. https://doi.org/10.1093/jpe/rtw071 (2017).
Betz, C., Scheuerer, M. & Reisch, C. Population reinforcement—A glimmer of hope for the conservation of the highly endangered Spring Pasque flower (Pulsatilla vernalis). Biol. Conserv. 168, 161–167. https://doi.org/10.1016/j.biocon.2013.10.004 (2013).
Rascle, P. et al. Identification of success factors for the reintroduction of the critically endangered species Eryngium viviparum J. Gay (Apiaceae). Ecol. Eng. 122, 112–119. https://doi.org/10.1016/j.ecoleng.2018.07.021 (2018).
Aguilar, R., Quesada, M., Ashworth, L., Herrerias-Diego, Y. & Lobo, J. Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Mol. Ecol. 17, 5177–5188. https://doi.org/10.1111/j.1365-294X.2008.03971.x (2008).
Bowles, M. L., McBride, J. L. & Bell, T. J. Long-term processes affecting restoration and viability of the federal threatened Mead’s milkweed (Asclepias meadii). Ecosphere 6, 11. https://doi.org/10.1890/ES14-00240.1 (2015).
IUCN. The IUCN Red List of Threatened Species. Version 2023–1. https://www.iucnredlist.org. Accessed 4 Feb 2024.
Thomas, P. A., El-Barghathi, M. & Polwart, A. Biological flora of the British Isles: Juniperus communis L. J. Ecol. 95, 1404–1440. https://doi.org/10.1111/j.1365-2745.2007.01308.x (2007).
Oostermeijer, J. G. B. & De Knegt, B. Genetic population structure of the wind-pollinated, dioecious shrub Juniperus communis in fragmented Dutch heathlands. Plant Species Biol. 19, 175–184. https://doi.org/10.1111/j.1442-1984.2004.00113.x (2004).
Vanden Broeck, A. et al. Genetic structure and seed-mediated dispersal rates of an endangered shrub in a fragmented landscape: a case study for Juniperus communis in northwestern Europe. BMC Genet. 12, 73. https://doi.org/10.1186/1471-2156-12-73 (2011).
Reim, S., Lochschmidt, F., Proft, A., Tröber, U. & Wolf, H. Genetic structure and diversity in Juniperus communis populations in Saxony, Germany. Biodiv. Res. Conserv. 42, 9–18. https://doi.org/10.1515/biorc-2016-0008 (2016).
Jacquemart, A.-L., Buyens, C., Delescaille, L.-M. & Van Rossum, F. Using genetic evaluation to guide conservation of remnant Juniperus communis (Cupressaceae) populations. Plant Biol. 23, 193–204. https://doi.org/10.1111/plb.13188 (2021).
Pinto, A. V., Hansson, B., Patramanis, I., Morales, H. E. & Van Oosterhout, C. The impact of habitat loss and population fragmentation on genomic erosion. Conserv. Genet. 25, 49–57. https://doi.org/10.1007/s10592-023-01548-9 (2023).
Zecherle, L. J. et al. Subspecies hybridization as a potential conservation tool in species reintroductions. Evol. Appl. 14, 1216–1224. https://doi.org/10.1111/eva.13191 (2021).
Futák, J., Dostál, J. & Novák, F. A. Flóra Slovenska I—Všeobecná časť (Vydavateľstvo Slovenskej akadémie vied, 1966).
Lakušić, B. & Lakušić, D. Anatomy of four taxa of the Genus Juniperus sect. Juniperus (Cupressaceae) from the Balkan peninsula. Bot. Serb. 35, 145–156 (2011).
Tocl, K. O ceste po Slovensku (Vesmír, 1898).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682. https://doi.org/10.1038/nmeth.2019 (2012).
Murray, M. G. & Thompson, W. F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4326. https://doi.org/10.1093/nar/8.19.4321 (1980).
Kalendar, R., Antonius, K., Smykal, P. & Schulman, A. H. iPBS: A universal method for DNA fingerprinting and retrotransposon isolation. Theor. Appl. Genet. 121, 1419–1430. https://doi.org/10.1007/s00122-010-1398-2 (2010).
The GIMP Development Team. GIMP 2.8.10. www.gimp.org (1997–2014). Retrieved 12 June 2022.
Fattal, R., Lischinski, D. & Werman, M. Gradient domain high dynamic range compression. ACM Trans. Graph. https://doi.org/10.1145/566570.566573 (2002).
Lazar, I. & Lazar, I. GelAnalyzer 19.1. www.gelanalyzer.com. Retrieved 15 Feb 2022.
Peakall, R. & Smouse, P. E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research–An update. Bioinformatics 28, 2537–2539. https://doi.org/10.1093/bioinformatics/bts460 (2012).
Schlüter, P. M. & Harris, S. A. Analysis of multilocus fingerprinting data sets containing missing data. Mol. Ecol. Notes 6, 569–572. https://doi.org/10.1111/j.1471-8286.2006.01225.x (2006).
Felsenstein, J. PHYLIP (Phylogeny Inference Package) version 3.7a. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle (2009).
Takezaki, N. & Nei, M. Genetic distances and reconstruction of phylogenetic trees from microsatellite DNA. Genetics 144, 389–399. https://doi.org/10.1093/genetics/144.1.389 (1996).
Paetkau, D., Waits, L. P., Clarkson, P. L., Craighead, L. & Strobeck, C. An empirical evaluation of genetic distance statistics using microsatellite data from bear (Ursidae) populations. Genetics 147, 1943–1957. https://doi.org/10.1093/genetics/147.4.1943 (1997).
Kalinowski, S. T. Evolutionary and statistical properties of three genetic distances. Mol. Ecol. 11, 1263–1273. https://doi.org/10.1046/j.1365-294X.2002.01520.x (2002).
Séré, M., Thévenon, S., Belem, A. M. G. & De Meeûs, T. Comparison of different genetic distances to test isolation by distance between populations. Hered. 119, 55–63. https://doi.org/10.1038/hdy.2017.26 (2017).
Reif, J. C., Melchinger, A. E. & Frisch, M. Genetical and mathematical properties of similarity and dissimilarity coefficients applied in plant breeding and seed bank management. Crop. Sci. 45, 1–7. https://doi.org/10.2135/cropsci2005.0001 (2005).
ESRI. ArcGIS Desktop: Release 9.3. (Environmental Systems Research Institute, 2011).
Halder, I. et al. Measurement of admixture proportions and description of admixture structure in different U.S. populations. Hum. Mutat. 30, 1299–1309. https://doi.org/10.1002/humu.21045 (2009).
Statistics Kingdom. Shapiro-Wilk Test Calculator. https://www.statskingdom.com/shapiro-wilk-test-calculator.html (2017). Accessed 12 Jan 2024.
Huson, D. H. & Scornavacca, C. Dendroscope 3: An interactive tool for rooted phylogenetic trees and networks. Syst. Biol. 61, 1061–1067. https://doi.org/10.1093/sysbio/sys062 (2012).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959. https://doi.org/10.1093/genetics/155.2.945 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14, 2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x (2005).
Puechmaille, S. J. The program structure does not reliably recover the correct population structure when sampling is uneven: subsampling and new estimators alleviate the problem. Mol. Ecol. Resour. 16, 608–627. https://doi.org/10.1111/1755-0998.12512 (2016).
Li, Y. L. & Liu, J. X. StructureSelector: A web based software to select and visualize the optimal number of clusters using multiple methods. Mol. Ecol. Resour. 18, 176–177. https://doi.org/10.1111/1755-0998.12719 (2018).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191. https://doi.org/10.1111/1755-0998.12387 (2015).
Hayfield, T. & Racine, J. S. Nonparametric kernel smoothing methods for mixed data types. R package np documentation (2020).
Connor, G. & Fuerst, G. R. Linear and partially linear models of behavioral trait variation using admixture regression. Preprint at https://doi.org/10.1101/2021.05.14.444173 (2021).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Soft. 67, 1–48. https://doi.org/10.18637/jss.v067.i01 (2015).
Hothorn, T., Bretz, F. & Westfall, P. Simultaneous inference in general parametric models. Biom. J. 50, 346–363. https://doi.org/10.1002/bimj.200810425 (2008).
Vekemans, X. AFLP-SURV Version 1.0. Distributed by the Author. Laboratoire De Génétique Et Ecologie Végétale, Université Libre De Bruxelles, Belgium (2002).
Dasmahapatra, K. K., Lacy, R. C. & Amos, W. Estimating levels of inbreeding using AFLP markers. Hered. 100, 286–295. https://doi.org/10.1038/sj.hdy.6801075 (2008).
Statistics Kingdom. Correlation Coefficient Calculator. https://www.statskingdom.com/correlation-calculator.html (2017). Accessed 12 Jan 2024.
Excoffier, L. & Lischer, H. E. L. Arlequin suite ver 35: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x (2010).
Da Silva, S. B. & Da Silva, A. Pstat: An R package to assess population differentiation in phenotypic traits. R J. 10, 447–454. https://doi.org/10.32614/RJ-2018-010 (2018).
Endelman, J. B. Ridge regression and other kernels for genomic selection with R package rrBLUP. TPG 4, 250–255. https://doi.org/10.3835/plantgenome2011.08.0024 (2011).
Thornton, T. et al. Estimating kinship in admixed populations. Am. J. Hum. Genet. 91, 122–138. https://doi.org/10.1016/j.ajhg.2012.05.024 (2012).
Cullingham, C. I. et al. Confidently identifying the correct K value using the ΔK method: When does K = 2?. Mol. Ecol. 29, 862–869. https://doi.org/10.1111/mec.15374 (2020).
Ramírez-Soriano, A., Ramos-Onsins, S. E., Rozas, J., Calafell, F. & Navarro, A. Statistical power analysis of neutrality tests under demographic expansions, contractions and bottlenecks with recombination. Genetics 179, 555–567. https://doi.org/10.1534/genetics.107.083006 (2008).
Van der Merwe, M., Winfield, M. O., Arnold, G. M. & Parker, J. S. Spatial and temporal aspects of the genetic structure of Juniperus communis populations. Mol. Ecol. 9, 379–386. https://doi.org/10.1046/j.1365-294x.2000.00868.x (2000).
Provan, J. et al. Restricted gene flow in fragmented populations of a wind-pollinated tree. Conserv. Genet. 9, 1521–1532. https://doi.org/10.1007/s10592-007-9484-y (2008).
Petit, R. J. et al. Comparative organization of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol. Ecol. 14, 689–701. https://doi.org/10.1111/j.1365-294X.2004.02410.x (2005).
Hantemirova, E. V. & Bessonova, V. A. Genetic diversity of Juniperus communis L. in Eurasia and Alaska inferred from nuclear microsatellite markers. Russ. J. Genet. 59, 271–280. https://doi.org/10.1134/S1022795423030055 (2023).
Wright, S. Evolution in mendelian populations. Genetics 16, 97–159. https://doi.org/10.1093/genetics/16.2.97 (1931).
Mills, L. S. & Allendorf, F. W. The one-migrant-per-generation rule in conservation and management. Conserv. Biol. 10, 1509–1518. https://doi.org/10.1046/j.1523-1739.1996.10061509.x (1996).
Ledig, F. T. Genetic variation in Pinus. In Ecology and Biology of Pinus (ed. Richardson, D. M.) 251–280 (Cambridge University Press, 1998).
Whitlock, M. C. & McCauley, D. E. Indirect measures of gene flow and migration: FST≠1/(4Nm+1). Hered 82, 117–125. https://doi.org/10.1038/sj.hdy.6884960 (1999).
Neigel, J. E. Is FST obsolete?. Conserv. Genet. 3, 167–173. https://doi.org/10.1023/a:1015213626922 (2002).
Ward, L. K. The conservation of Juniper. I. Present Status of Juniper in Southern England. J. Appl. Ecol. 10, 165–188. https://doi.org/10.2307/2404724 (1973).
Clifton, S. J., Ward, L. K. & Ranner, D. S. The status of juniper Juniperus communis L. in northeast England. Biol. Conserv. 79, 67–77. https://doi.org/10.1016/S0006-3207(96)00101-2 (1997).
Frankard, P. Évolution de la population de Juniperus communis L. dans la réserve naturelle domaniale de la genévrière de Cour pendant ces vingt dernières années et impact des mesures de gestion appliquées. Parcs et Réserves 59, 32–37 (2004).
Hüppe, J. Zur problematik der Verjüngung des Wacholders (Juniperus communis) unter dem Einfluβ von Wildkaninchen in Hudegebieden pleistozäner Sandlandschaften. Z. Ökol. Nat.schutz 4, 1–8 (1995).
García, D., Zamora, R., Hódar, J. A. & Gómez, J. M. Age structure of Juniperus communis L. in the Iberian Peninsula: Conservation of remnant populations in Mediterranean mountains. Biol. Conserv. 87, 215–220. https://doi.org/10.1016/S0006-3207(98)00059-7 (1999).
Sanz-Elorza, M., Dana, E. D., González, A. & Sobrino, E. Changes in the high-mountain vegetation of the Central Iberian Peninsula as a probable sign of global warming. Ann. Bot. 92, 273–280. https://doi.org/10.1093/aob/mcg130 (2003).
Verheyen, K., Schreurs, K., Vanholen, B. & Hermy, M. Intensive management fails to promote recruitment in the last large population of Juniperus communis (L.) in Flanders (Belgium). Biol. Conserv. 124, 113–121. https://doi.org/10.1016/j.biocon.2005.01.018 (2005).
Preston, S. J., Wilson, C., Jennings, S., Provan, J. & McDonald, R. A. The status of juniper (Juniperus communis) in Northern Ireland in 2005. Ir. Nat. J. 28, 372–378 (2007).
McCartan, S. A. & Gosling, P. G. Guidelines for seed collection and stratification of common juniper (Juniperus communis L.). TPN 26, 24–29 (2013).
Breed, M. F. et al. Pollen diversity matters: Revealing the neglected effect of pollen diversity on fitness in fragmented landscapes. Mol. Ecol. 21, 5955–5968. https://doi.org/10.1111/mec.12056 (2012).
Hamrick, J. L. & Godt, M. J. W. Effects of life history traits on genetic diversity in plant species. Philos. Trans. R. Soc. B. Biol. 351, 1291–1298. https://doi.org/10.1098/rstb.1996.0112 (1996).
Gilroy, D. L., Phillips, K. P., Richardson, D. S. & Van Oosterhout, C. Toll-like receptor variation in the bottlenecked population of the Seychelles warbler: computer simulations see the ‘ghost of selection past’ and quantify the ‘drift debt’. J. Evol. Biol. 30, 1276–1287. https://doi.org/10.1111/jeb.13077 (2017).
Tilman, D., May, R. M., Lehman, C. L. & Nowak, M. A. Habitat destruction and the extinction debt. Nature 371, 65–66. https://doi.org/10.1038/371065a0 (1997).
Kuussaari, M. et al. Extinction debt: a challenge for biodiversity conservation. Trends Ecol Evol. 24, 564–571. https://doi.org/10.1016/j.tree.2009.04.011 (2009).
Dussex, N., Morales, H. E., Grossen, C., Dalén, L. & Van Oosterhout, C. Purging and accumulation of genetic load in conservation. Trends Ecol. Evol. 38, 961–969. https://doi.org/10.1016/j.tree.2023.05.008 (2023).
Elias, T. S. The complete trees of North America (Van Nostrand Reinhold Co., 1980).
Körner, C. Alpine Plant Life: Functional Plant Ecology of High Mountain Ecosystems (Springer, 2003). https://doi.org/10.1007/978-3-642-18970-8.
Hey, J., Waples, R. S., Arnold, M. L., Butlin, R. K. & Harrison, R. G. Understanding and confronting species uncertainty in biology and conservation. Trends Ecol. Evol. 18, 597–603. https://doi.org/10.1016/j.tree.2003.08.014 (2003).
Allendorf, F. W., Hohenlohe, P. A. & Luikart, G. Genomics and the future of conservation genetics. Nat. Rev. Genet. 11, 697–709. https://doi.org/10.1038/nrg2844 (2010).
Frankham, R. et al. Implications of different species concepts for conserving biodiversity. Biol. Conserv. 153, 25–31. https://doi.org/10.1016/j.biocon.2012.04.034 (2012).
Galtier, N. Delineating species in the speciation continuum: A proposal. Evol. Appl. 12, 657–663. https://doi.org/10.1111/eva.12748 (2018).
Christensen, K. I. Juniperus communis subsp. alpina (Smith) Čelakovský (Cupressaceae). A Nomenclatural Comment. Taxon 34, 686–688. https://doi.org/10.2307/1222215 (1985).
Miller, H. J. Anatomical studies of Juniperus communis L. spp. communis and J. communis L. ssp. nana Syme. Acta Bot. Neerl. 23, 91–98 (1974).
Sullivan, G. Prostrate juniper heath in north-west Scotland: Historical, ecological, and taxonomic issues. Ph.D. Thesis (University of Aberdeen, 2001).
Filipowicz, N., Piotrowski, A., Ochocka, J. R. & Asztemborska, M. The phytochemical and genetic survey of common and dwarf juniper (Juniperus communis and Juniperus nana) identifies chemical races and close taxonomic identity of the species. Planta Med. 72, 850–853. https://doi.org/10.1055/s-2006-941543 (2006).
Guerra Hernández, E., López Martínez, M. C. & García-Villanova, R. Componentes volatiles identificados por cromatografia en fase gaseosa de macerados de bayas de Junperus en etanol. An. Bromatol. 39, 229–237 (1987).
Khantemirova, E. V. & Semerikov, V. L. Genetic variation of some varieties of common Juniper Juniperus communis L. inferred from analysis of allozyme loci. Russ. J. Genet. 46, 546–554. https://doi.org/10.1134/S1022795410050066 (2010).
Adams, R. P. & Pandey, R. N. Analysis of Juniperus communis and its varieties based on DNA fingerprinting. Biochem. Syst. Ecol. 31, 1271–1278. https://doi.org/10.1016/S0305-1978(03)00036-X (2003).
Hantemirova, E. V., Berkutenko, A. N. & Semerikov, V. L. Systematics and gene geography of Juniperus communis inferred from isoenzyme data. Russ. J. Genet. 48, 920–926. https://doi.org/10.1134/S1022795412090050 (2012).
Łabiszak, B. & Wachowiak, W. Molecular signatures of reticulate evolution within the complex of european pine taxa. Forests 12, 489. https://doi.org/10.3390/f12040489 (2021).
Żukowska, W. B., Wójkiewicz, B., Lewandowski, A., László, R. & Wachowiak, W. Genetic variation of Scots pine (Pinus sylvestris L.) in Eurasia: impact of postglacial recolonization and human-mediated gene transfer. Ann. For. Sci. https://doi.org/10.1186/s13595-023-01207-6 (2023).
Acknowledgements
We are grateful to Prof. Cock Van Oosterhout (University of East Anglia) for his valuable consultation on genetic load and the study’s hypotheses. We also appreciate Dr. Drahoš Blanár (Muránska Planina National Park, Slovakia) for collecting samples in Besnik and Stolica, and Dr. Radovan Ostrovský (Institute of Forest Ecology, Slovak Academy of Sciences) for designing the location map.
Funding
The work was supported by: Scientific Grant Agency of the Ministry of Education, Science, Research and Sport of the Slovak Republic [2/0005/23]; and the Operational program Integrated Infrastructure within the project: Demand-driven research for the sustainable and inovative food [313011V336], cofinanced by the European Regional Development Fund.
Author information
Authors and Affiliations
Contributions
M. G. and A. K. harvested the plant material and isolated the DNA samples. J. J. conducted the measurement of needle characteristics. M. K. helped with the DNA isolation, conceived the idea, designed and performed the analyses, and wrote the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Approval for collecting samples
The Juniperus communis L. samples were collected with the necessary permissions from the District Office Banská Bystrica under the file number OU-BB-OSZP1-2022/015915-010. All sampling procedures complied with relevant regulations and guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Klobučník, M., Kormuťák, A., Jurčík, J. et al. Conservation genetic evaluation of Juniperus communis sensu lato in Slovakia. Sci Rep 15, 8398 (2025). https://doi.org/10.1038/s41598-025-92792-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-92792-1