Abstract
Age-related macular degeneration (AMD) is associated with chronic inflammation of the retinal pigment epithelium (RPE) and elevated cytokines including TNFα, TGF-β, IL-6, and IL-1β. As a metabolic intermediary supporting aerobic glycolysis in the adjacent photoreceptors, the RPE’s metabolic responses to inflammation and the optimal methods to study cytokine-driven metabolic programming remain unclear. We performed a rigorous comparison of ARPE-19 cells and rat eyecup metabolomes, revealing key distinctions. Rat eyecups exhibit higher levels of lactate and palmitate but depleted glutathione and high-energy nucleotides. Conversely, ARPE-19 cells are enriched with high-energy currency metabolites and the membrane phospholipid precursors phosphocholine and inositol. Both models showed contrasting responses to individual cytokines: ARPE-19 cells were more sensitive to TNFα, while eyecups responded more strongly to TGF-β2. Notably, a combined cytokine cocktail elicited stronger metabolic effects on ARPE-19 cells, more potently impacting both metabolite abundance (41 vs. 29) and glucose carbon flux (29 vs. 5), and influencing key RPE metabolites such as alanine, glycine, aspartate, proline, citrate, α-ketoglutarate, and palmitate. Overall, these findings position ARPE-19 cells as a more responsive platform for studying inflammatory cytokine effects on RPE metabolism and reveal critical RPE metabolites which may be linked with AMD pathogenesis.
Similar content being viewed by others
Introduction
The retinal pigment epithelium (RPE) is a polarized monolayer of cells coating the outer retina, acting as an essential metabolic intermediary. Metabolites from the choroid, the primary vascular layer of the retina, are transported across basal membrane of the RPE, en route to the neurons of the outer retina. Within the RPE, these metabolites either undergo processing or are exported to the apical side of the monolayer. Intercalated with the RPE’s apical membrane are the photoreceptors which, to satisfy their prodigious biosynthetic and energetic demands, have high rates of glucose consumption1,2. The prevailing paradigm is that the RPE has evolved to consume minimal amounts of glucose, ensuring maximal supply for the photoreceptors3,4. Lactate, a byproduct of photoreceptor metabolism, is thought to suppress glycolysis in the RPE4, and is instead employed to fuel oxidative phosphorylation (OXPHOS) to meet its energy demands.
Recent experiments indicate that RPE uses glucose, glutamine, and proline to synthesise a range of other metabolites, including citrate, malate, glutamate, and glutamine, preferentially exporting them to the apical side5,6. Moreover, the retina is believed to export succinate to transfer reducing power from the oxygen-poor retina to the oxygen-rich RPE7. Understanding the metabolic interactions that underpin the retinal metabolic ecosystem is of vital interest as imbalances in these processes have been identified as important factors in retinal diseases such as age-related macular degeneration (AMD)8.
In AMD, the RPE, adjacent Bruch’s membrane, and the choroid are sites of chronic inflammation9. Lipid- and protein-rich deposits known as drusen, coupled with RPE cell senescence, fuel these inflammatory conditions, driving elevated expression of cytokines such as TNFα, TGF-β, IL-6, and IL-1β by both infiltrating immune cells and the RPE itself10. Each of these cytokines has been shown to affect various aspects of glucose, mitochondrial, lipid, and amino acid metabolism in diverse non-retinal tissues and cell types11. Moreover, recent studies indicate that these cytokines, particularly TNFα and TGF-β2, can impact lactate production, glucose and oxygen consumption, and metabolic gene expression in multiple RPE models12,13. However, the effects of these cytokines, individually and in combination, on the broad RPE metabolic profile remain unknown.
RPE metabolism research has employed a range of in vitro and in vivo models. Among these, primary human RPE cells isolated from adult donors14,15 and RPE cells derived from induced pluripotent stem cells (iPSCs)16,17 are two common models. Another notable in vitro model of RPE metabolism is the human foetal RPE (hfRPE) cell5,12,18. While these cells exhibit many native RPE functions18, they present several drawbacks, such as ethical implications, limited availability, and discrepancies in emulating the metabolic and functional properties of mature, postnatal RPE cells.
The ARPE-19 cell line, spontaneously immortalized from the RPE of a 19-year-old male19, presents an alternative in vitro model. These cells are favoured for their ease of cultivation and have been employed in the study of diverse facets of RPE biology and disease such as autophagy20, growth factor signalling21, cytokine signalling22,23,24, oxidative stress25,26, epithelial-to-mesenchymal transition (EMT)27, as well as metabolism28,29. Once differentiated, ARPE-19 cells exhibit numerous characteristics of the native RPE, including cobblestone morphology, polarity, phagocytic capacity, and expression of RPE-specific markers19,30.
Rodent eyecups, which serve as an ex vivo model of the RPE/choroid, have also emerged as a tool for the study of RPE metabolism4,7,31,32. Eyecups consist of the tissue anterior to the cornea which overlays the neural retina. The outermost layer of the eyecup contains the sclera, which is comprised primarily of connective tissue interspersed with a sparse population of fibrocytes33. Directly anterior to the sclera lies the choroid, a complex tissue consisting of neurons, smooth muscle cells, melanocytes, fibroblasts, vascular endothelial cells, and various immune cells such as mast cells, macrophages, and lymphocytes34. The RPE, making up the innermost layer, has the highest cell density and is considered to be the most metabolically active cell population in the eyecup4,31,35.
This study aims to determine the influence of cytokines on RPE metabolism while comparing and contrasting ARPE-19 and eyecup as RPE models for metabolomic analysis. In addition to comparing the relative abundance of metabolites, we employ glucose carbon labelling to observe how glucose carbons are distributed across metabolites in both models. We find that while the overall metabolic profiles of both ARPE-19 cells and eyecups exhibit a considerable degree of overlap, they exhibit pronounced differential abundance of key metabolites including lactate, NAD+, glutathione, palmitate, and most amino acids. Likewise, metabolic pathway analysis highlighted differences in pathways involved in glutathione, ketone body, and amino acid metabolism between both RPE models. The metabolic differences between these models are reflected in the responses of individual metabolites and their overall sensitivity to specific cytokines.
Results
Metabolite abundance comparison between ARPE-19 cells and eyecups
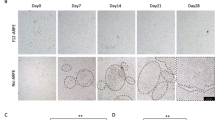
Both ARPE-19 cells and rodent eyecups are commonly employed as RPE models, however, they differ in their species of origin, cellular composition, and duration in culture. ARPE-19 cells were differentiated in 10 mM nicotinamide using previously described methods30, with differentiation confirmed by qRT-PCR analysis of the RPE marker genes RLBP1, RPE65, and BEST1 (Fig. S1A) and imaging to verify cobblestone morphology compared to undifferentiated cells (Fig. S1B,C). Differentiated ARPE-19 cells and rat eyecups were then incubated in medium containing 5.5 mM 1,2 13C glucose for 12 h before metabolites were extracted for targeted metabolomic analysis using liquid chromatography mass spectrometry (LC–MS). This provided metabolite abundance and isotopologue data for metabolic flux that formed the basis for our analysis. A full list of ARPE-19 and eyecup metabolite abundance values and 13C labelling data is available in Supplementary Table S1.
We first examined whether the metabolic profiles of the two RPE models more closely resembled each other than a non-retinal control cell line, A549 lung carcinoma epithelial cells. Unsupervised hierarchical clustering was applied to both rows (metabolites) and columns (samples) to group metabolites with similar abundance profiles and sample groups with similar overall metabolite abundances, respectively (Fig. 1). Sample clustering grouped together the replicates for all three models with ARPE-19 cells and eyecups exhibiting more analogous metabolic profiles compared to the control A549 cells. In general, intermediates associated with amino acid and glucose metabolism were the most abundant metabolites across all three models while pyrimidine and pentose phosphate pathway (PPP) intermediates consistently marked the lower end of the abundance range.
Comparative metabolite abundance between A549, ARPE-19 and eyecups. Heatmap of heirarchical clustering of A549 (C1-C2), ARPE-19 (A1-A3), and eyecup (E1–E5) metabolites from LC–MS metabolomics analysis. Metabolite abundances are −log10 transformed and normalised to the total metabolite abundance for each sample. Blue indicates lower abundance and red indicates higher abundance. Analysis was performed and plot was produced using the R packages pheatmap and ggplot2, respectively.
Principal component analysis (PCA) of the abundance profiles of ARPE-19 cells and eyecups highlighted consistency between all of the ARPE-19 replicates and all but one of the eyecup replicates (Fig. S2A). The first two components captured the majority of the data variance, with 56% and 38.7% respectively. Subsequent components provided sharply diminishing contributions to the total variance, with the third component representing just 2.7% (Fig. S2B).
Linear regression analysis comparing ARPE-19 and eyecup metabolite abundances yielded an R2 value of 0.58, signifying a moderately strong relationship (Fig. 2A). Outlying metabolites included nicotinamide and glutathione which were highly abundant in ARPE-19 cells but not eyecups, and acetylcholine (Ac-choline) which exhibited higher expression in eyecups. The fold difference in ARPE-19 metabolite abundance relative to eyecups was calculated to identify metabolites with the most pronounced relative abundance differences between the two RPE models (Fig. 2B). Thresholds were set at log2 fold changes above or below 1 or −1 (equivalent to 2- or -twofold changes) and −log10 P-values exceeding 1.3 (P < 0.05). 12.9% of metabolites exceeded the positive fold change and significance thresholds, while 17.4% were below these thresholds, with the remainder (69.7%) of metabolites exhibiting similar levels of abundance. Relative to eyecups, ARPE-19 cells exhibited several-fold higher levels of ATP, UDP-glucose, phosphocholine (P-choline), and 6-phosphogluconate (6P-gluconate), but lower levels of xanthine, cysteine, sedohepulose 7-phosphate (S7P), 5-oxoproline, and uracil.
Comparative single-metabolite analysis between ARPE-19 cells and eyecups. (A) Linear regression analysis of mean single-metabolite abundance from ARPE-19 cells and eyecups from LC–MS metabolomics analysis. Metabolite abundances are −log10 transformed and normalised to the total metabolite abundance for each sample. Pink indicates high average abundance and blue indicates low average abundance. R2 = 0.577, P-value = 8.35 × 10−27. (B) Volcano plot of ARPE-19 metabolite fold-change relative to eyecups. 17.4% of metabolites with Log2 fold-changes < −1 and −Log10 P values > 1.3 (P < 0.05) were marked blue for the lower threshold, while 12.9% of metabolites with Log2 fold-changes > 1 and −Log10 P values > 1.3 (P < 0.05) were marked red for the upper threshold. Scatterplot and volcano plot were produced using the R package ggplot2.
Metabolic pathway comparison between ARPE-19 cells and eyecups
To compare the prominence of different metabolic pathways in each model, we aggregated the metabolite abundance for each pathway and compared their contribution to the total metabolite abundance (Fig. 3A & S3). The contribution of tricarboxylic acid (TCA) cycle intermediates, cysteine, and methionine metabolites, as well as glycine, serine, and threonine metabolites were similar between ARPE-19 cells and eyecups, while A549 cells resembled ARPE-19 cells in the contribution of metabolites involved in glutathione metabolism, valine, leucine and isoleucine metabolism, and fatty acid biosynthesis.
Comparative metabolic pathway analysis between models. (A) Stacked bar chart of summed contribution of metabolites in each pathway to total metabolite abundance in A549 cells, ARPE-19 cells, and rat eyecups. For each model, the mean percentage of total metabolite abundance for each metabolite was calculated, and metabolites were summed based on their Kyoto encyclopedia of genes and genomes (KEGG)75 pathway with some modifications (see supplementary data). (B) Scatterplot showing pathway analysis of metabolites with significantly different abundance between ARPE-19 cells and eyecups. KEGG pathway impact scores are plotted on the x-axis and pathway enrichment significance is plotted on the y-axis. Points are coloured according to Log10 P-values, with pink representing higher values. Larger points represent higher pathway impact values. Analysis was performed with MetaboAnalyst 5.0 and the plot was produced using the R package ggplot2.
Pathway analysis was performed on metabolites that showed statistically significant differences in abundance between ARPE-19 cells and eyecups to identify the metabolic pathways that are distinctly represented in each model. The pathway impact value, which represents the degree of influence of the chosen model on a metabolic pathway through pathway topology analysis, identified several pathways involved in amino acid metabolism as being most impacted (Fig. 3B). These pathways included phenylalanine, tyrosine and tryptophan biosynthesis, alanine, aspartate, and glutamate metabolism, and arginine biosynthesis.
Analysis of individual metabolite abundance revealed that within the glycolytic pathway, glucose and lactate were far more abundant in eyecups and A549 cells than ARPE-19 cells (Fig. 4). All models exhibited substantial levels of citrate and malate compared to other TCA cycle metabolites, and uniformly low abundance of PPP metabolites. ARPE-19 and A549 cells exhibited elevated, nicotinamide and NAD+, as well as both oxidised and reduced forms of glutathione, while eyecups contained high levels of 5-oxoproline, a derivative of glutathione breakdown.
Metabolite abundance comparison between A549 cells, ARPE-19 cells, and rat eyecups. For each model, the mean percentage of total metabolite abundance for each metabolite was calculated and metabolites were grouped according to their KEGG pathway with modifications (see supplementary data). Unpaired T-tests were performed on metabolite abundances between models and adjusted for multiple testing using the Benjamini–Hochberg method with a false discovery rate (FDR) of 5%. Error bars show mean ± SEM. *, q < 0.05; **, q < 0.01; ***, q < 0.001; ****, q < 0.0001.
Levels of most essential amino acids were comparable across all models, while ARPE-19 and A549 cells had significantly lower levels of cysteine, methionine, tyrosine, and lysine compared to eyecups. Among the non-essential amino acids, glutamate was prominent in ARPE-19 cells, glutamine in eyecups, and proline in A549 cells. ARPE-19 and A549 cells exhibited elevated concentrations of nucleoside polyphosphates relative to eyecups, while eyecups exhibited either comparable or higher levels of nucleoside monophosphates and non-phosphorylated nucleosides. Other notable differences between models were elevated levels of P-choline and palmitate in ARPE-19 cells and eyecups, respectively. Currency metabolite ratios in ARPE-19 cells favoured higher-energy phosphorylated nucleotides and reduced glutathione (Fig. S4). In contrast, eyecups exhibited elevated NADPH/NADP+ and P-creatine/creatine ratios compared to the other two models.
Overall, while the metabolic profiles of the RPE models are more akin than A549 cells, several key metabolites are differentially expressed, including lactate, NAD+, glutathione, 5-oxoproline, glutamate, glutamine, and nucleoside polyphosphates. Moreover, the two RPE models differ in the representation of their amino acid, ketone, and glutathione metabolic pathways, with ARPE-19 cells also generally possessing a higher ratio of high-energy species of currency metabolites.
Differences in metabolite labelling between ARPE-19 cells and eyecups
Most glycolytic metabolites showed comparable 13C glucose carbon labelling in both models with metabolites downstream of F-1,6-BP cleavage into dihydroxyacetone phosphate (DHAP) and glyceraldehyde 3-phosphate (G3P) showing roughly 50% labelling in both models, implying saturation of 1,2 13C glucose labelling of these metabolites (Fig. 5). All TCA metabolites except acetyl-CoA exhibited greater labelling in ARPE-19 cells compared to eyecups. The labelling consistency of PPP intermediates varied between replicates, likely due to their universally low abundance in samples. Nicotinamide metabolite labelling in eyecups was markedly higher, with values around 70%, compared to 30% in ARPE-19 cells. Several amino acids including alanine, glutamate, proline, glycine, and serine showed less pronounced labelling in eyecups compared to ARPE-19 cells. However, both models had similar alanine, aspartate, and glutamate labelling at around 25%, and for proline at about 15%, while the remaining amino acids had minimal labelling. Purine labelling was predominantly higher in eyecups, with adenosine derivatives, in particular, displaying nearly twice the labelling observed in ARPE-19 cells. In contrast, fructose and sorbitol were almost completely labelled in ARPE-19 cells, whereas their labelling in eyecups was negligible. While most nucleotide sugar metabolites displayed more pronounced labelling in eyecups, glycerophospholipid metabolites exhibited comparable labelling in both models.
Metabolite labelling comparison between ARPE-19 cells and rat eyecups. For each model, the mean percentage of total metabolite with labelling was calculated and metabolites were grouped according to their KEGG pathway with modifications (see supplementary data). Unpaired T-tests were performed on labelling percentages between ARPE-19 cells and eyecups and adjusted for multiple testing using the Benjamini–Hochberg method with a false discovery rate (FDR) of 5%. Error bars show mean ± SEM. *, q < 0.05; **, q < 0.01; ***, q < 0.001; ****, q < 0.0001.
Comparative effects of cytokine treatment on ARPE-19 and eyecup metabolism
Investigating the impact of cytokines on RPE metabolism is an emerging research interest due to their elevated levels in age-related retinal diseases like AMD. TNFα, TGF-β2, IL-6, and IL-1β are among the most prominent of these cytokines, and our previous study showed that they each influence glucose metabolism in ARPE-19 cells to varying degrees, with a cytokine cocktail containing all four cytokines having the most potent effect13. The present study examined the metabolic response of differentiated ARPE-19 cells and rat eyecups to 20 ng/mL TNFα, TGF-β2, IL-6, IL-1β, or the cytokine cocktail in medium enriched with 5.5 mM 1,2 13C-labelled glucose for a duration of 12 h. The objective was to investigate cytokine-induced metabolic changes in the RPE and compare the responses between the RPE models. After 12 h, none of the cytokine treatments visibly affected the viability of ARPE-19 cells or eyecups, which is supported by the continuous production of lactate and other metabolites.
PCA revealed that the first two components captured approximately 50% of the metabolic variance post-cytokine treatment in both models (Fig. 6A,B), with the remaining variance accounted for by 16 and 27 components in APRE-19 cells and eyecups, respectively (Fig. S5A & Fig. S5B). Most cytokines produced clusters that were distinguishable from the control in both ARPE-19 cells and eyecups, with the cytokine cocktail notably exhibiting clustering distinctly separated from their respective controls in both models.
Comparative overview and pathway impact of cytokine treatment on ARPE-19 cells and eyecups. (A) PCA scores plot of LC–MS metabolite abundance values from ARPE-19 cells following treatment with vehicle (control, grey), TNFα (orange), TGF-β2 (green), IL-6 (pink), IL-1β (purple), and the cytokine cocktail (blue). Shading represents 95% confidence interval. (B) PCA scores plot of LC–MS metabolite abundance values from rat eyecups following treatment with vehicle (control, grey), TNFα (orange), TGF-β2 (green), IL-6 (pink), IL-1β (purple), and the cytokine cocktail (blue). Shading represents 95% confidence interval. PCA analysis was performed on Metaboanalyst 5.0 and the plots was produced with the R package ggplot2. (C) Linear regression analysis of ARPE-19 (x-axis) and eyecups (y-axis) KEGG pathway impact scores based on pathway analysis of metabolites with significantly changed abundance following cytokine cocktail treatment. Pink indicates high average impact score, and blue indicates low average impact score. R2 = 0.24, P-value = 0.00956. (D) Linear regression analysis of ARPE-19 (x-axis) and eyecups (y-axis) KEGG pathway enrichment P-values based on pathway analysis of metabolites with significantly changed abundance following cytokine cocktail treatment. Pink indicates high average enrichment P-value, and blue indicates low average enrichment P-value. R2 = 0.02, P-value = 0.481. Pathway analysis was performed with MetaboAnalyst 5.0, and the plots were produced using the R package ggplot2.
Linear regression analysis comparing the pathway impact and enrichment scores post-cytokine treatment between the two models did not show a statistically significant linear relationship with any single cytokine treatment (Fig. S6–10A–D). However, with cocktail treatment, pathway impact scores showed a statistically significant, moderately predictive relationship with an R2 of 0.24 and a P-value of 0.00956 (Fig. 6C), though this was not observed with enrichment scores (Fig. 6D). Cytokine cocktail treatment primarily affected metabolic pathways related to amino acid metabolism, as well as purine, ascorbate, and aldarate metabolism in both models. These results indicate that, when exposed to the cytokine cocktail, ARPE-19 cells and eyecups may regulate a common subset of metabolic pathways, notwithstanding their disparate responses to individual cytokines.
Cytokines elicited distinct changes in individual metabolite abundance in ARPE-19 cells compared to eyecups (Fig. 7). TNFα primarily suppressed glycolytic and TCA cycle metabolites in ARPE-19 cells, except for increasing α-ketoglutarate (α-KG) and succinyl-CoA, while minimally affecting eyecups. TNFα reduced amino acid levels in both models, decreased purines in eyecups, increased specific purines in ARPE-19 cells, while pyrimidine levels decreased in ARPE-19 cells but increased in eyecups. TGF-β2 and IL-6 similarly affected glycolytic and TCA cycle metabolites across both models but varied in α-KG and essential amino acid responses, with ARPE-19 cells showing increased levels of alanine, asparagine, cysteine, and proline. Both cytokines broadly increased purine and pyrimidine levels in ARPE-19 cells, though some nucleotides such as guanosine, inosine, IMP, hypoxanthine, and uridine decreased in eyecups.
Cytokine impacts on ARPE-19 and eyecup relative metabolite abundance. Heatmap of Log2 fold change of metabolite abundance following TNFα, TGF-β2, IL-6, IL-1β, and the cytokine cocktail treatment relative to vehicle-treated control. Blue indicates negative log2 fold change, red indicates positive log2 fold change, and grey indicates non-detected metabolites. Metabolites were grouped according to their KEGG pathway with some modifications (see supplementary data). Plots were produced using the R package ggplot2.
IL-1β induced consistent effects across models on TCA intermediates and non-essential amino acids, except for asparagine and cysteine levels increasing in ARPE-19 cells and aspartate and glutamate rising in eyecups. Nucleotide levels responded similarly, with increases in adenosine, ATP, CTP, and UTP, and decreases in IMP and xanthine in both models. On the other hand, the cytokine cocktail produced markedly different effects on glycolytic intermediates (e.g., PEP), TCA intermediates (e.g., α-KG), and amino acids, with non-essential amino acids like alanine, arginine, cystine, and proline increasing in ARPE-19 cells but either decreasing or remaining stable in eyecups, while aspartate and glutamate levels increased in eyecups only.
Cytokine treatment also influenced metabolite labelling differently between ARPE-19 cells and eyecups (Fig. S11). TNFα increased nucleotide labelling in both models, with a greater effect in ARPE-19 cells, while decreasing labelling of most TCA metabolites in ARPE-19 and increasing it in eyecups. Amino acid labelling varied: eyecups showed increased labelling of alanine, asparagine, glutamine, and proline, while ARPE-19 cells had higher glycine labelling. TGF-β2 and IL-6 produced similar trends within each model but different labelling profiles between them; glycolytic labelling stayed mostly constant, except for PEP labelling which increased in ARPE-19 cells and decreased in eyecups. TCA labelling rose in eyecups but decreased in ARPE-19 cells, with alanine and glycine labelling higher in ARPE-19 cells and asparagine and glutamine elevated in eyecups. Purine and pyrimidine labelling increased in ARPE-19 cells but showed modest changes in eyecups.
IL-1β had minimal impact on glycolytic labelling but elevated TCA and non-essential amino acid labelling in eyecups, with only glycine increasing in ARPE-19 cells. UTP labelling rose in eyecups, while UMP increased in ARPE-19 cells. The cytokine cocktail slightly decreased glycolytic labelling across both models, increased α-KG and succinate labelling in eyecups, and elevated alanine and glycine labelling in both models. In ARPE-19 cells, it reduced labelling of asparagine, aspartate, and glutamine, while purine and pyrimidine labelling generally increased in both models.
Overall, ARPE-19 cells responded more strongly to TNFα treatment than eyecups, showing a greater number of up- and downregulated metabolites in both metabolite abundance (24 vs. 10, Fig. 8A) and 13C labelling (14 vs. 4, Fig. 8B). Conversely, TGF-β2 had a greater impact on abundance in eyecups than ARPE-19 cells (24 vs. 11), while altering 13C labelling to a similar degree in both models. IL-6 and IL-1β impacted the abundance and 13C labelling in a similar number of metabolites in both models, however both cytokines were more likely to upregulate metabolites in ARPE-19 cells while downregulating them in eyecups. In ARPE-19 cells, the cytokine cocktail induced changes in a much larger set of metabolites compared to eyecups, more potently affecting both abundance (41 vs. 29) and 13C labelling (29 vs. 5).
Cytokine cocktail treatment has the most potent effects on ARPE-19 metabolism. (A) Stacked bar chart showing the number of upregulated (red) or downregulated (blue) metabolites in ARPE-19 cells and eyecups with cytokine treatment. Thresholds were defined as > 1.5 and < 0.75 fold-change abundance relative to controls for upregulated and downregulated metabolites, respectively. (B) Stacked bar chart showing the number of metabolites with increased (red) or decreased (blue) 13C labelling in ARPE-19 cells and eyecups with cytokine treatment. Thresholds were defined as > 1.5 and < 0.75 fold-change 13C labelling relative to controls for metabolites with increased or decreased labelling, respectively. (C) Proposed model for the effects of cytokines on the RPE in vivo based on cytokine cocktail treatment in ARPE-19 cells. The cytokine cocktail promotes glucose consumption and increased flux of glucose carbons into 1,3-BPG, alanine, glycine, and through the PPP to purine and pyrimidine metabolites. Cytokine cocktail treatment also promotes palmitate consumption, decreases proline utilisation, and influences the abundance of key metabolites of exchange between the RPE and photoreceptors including aspartate, citrate, and α-KG. 1,3-BPG, 1,3-bisphosphoglycerate; α-KG, alpha ketoglutarate; Asp, aspartate; OAA, oxaloacetate; PPP, pentose phosphate pathway.
Given that multiple cytokines are present in the inflamed RPE and choroid, the cytokine cocktail most closely represents the conditions that the RPE is exposed to in vivo. ARPE-19 cells were much more responsive to cytokine cocktail treatment than eyecups, with the cocktail inducing changes in a range of metabolites that are key components of the retinal metabolic ecosystem (Fig. 8C). The cocktail promoted glucose consumption in the RPE which may restrict glucose availability for the photoreceptors. Moreover, cocktail treatment increased consumption of the fatty acid palmitate and decreased consumption of proline, a critical amino acid for RPE health36. The cytokine cocktail also altered the abundance of several metabolites that are exchanged between the RPE and photoreceptors including aspartate, citrate, and α-KG.
Discussion
ARPE-19 cells have been employed to investigate diverse aspects of RPE biology, and while they adopt many native RPE characteristics when differentiated, they lack the multicellular context of living organisms. On the other hand, ex vivo eyecups more accurately capture the structural and physiological environment of the RPE but introduce the challenge of maintaining tissue viability and discerning the metabolic signature of specific cell populations. In this study, we rigorously compared the metabolomic profile of these two models and assessed their responses to cytokine treatment. The metabolic profiles of ARPE-19 cells and eyecups were found to be more akin to each other than to A549 cells, exhibiting strikingly similar TCA cycle intermediate profiles, with citrate and malate as the most abundant TCA metabolites. High relative citrate levels are characteristic of a bioenergetic profile that is reliant on OXPHOS, rather than glycolysis37, which supports the prevailing metabolic ecosystem model4.
Both RPE models also exhibited similarities in the abundance of glycine, serine, and threonine pathway components. The retina and RPE depend on serine for the synthesis of key metabolites including sphingolipids, phosphatidylserine, as well as glycine—all of which are vital for retina and RPE physiology38. Notably, the RPE/choroid exhibits relatively high expression of serine biosynthesis enzymes31, and in vitro radiolabelling experiments show that cultured RPE cells readily synthesise serine from glucose5,39. However, our observations of low isotopic incorporation into serine and glycine, especially in eyecups, suggest a modest contribution of glucose carbons for synthesising these amino acids. Notably, proline, an important metabolite in retinal health and disease5,32,36, was present at levels approximately sevenfold lower in ARPE-19 cells and eyecups than A549 cells. This may indicate high proline consumption, concurring with current models that suggest the RPE uses proline to synthesise other metabolites to support the photoreceptors5.
However, in contrast to eyecups, the ARPE-19 metabolic profile indicated a relatively energy-rich state, with high ATP/ADP and ATP/AMP ratios comparable to proliferating cells like macrophages40, islets41, and cancer cell lines42,43. On the other hand, eyecups exhibited lower 13C labelling of TCA cycle intermediates and low ATP/ADP ratio is indicative of impaired mitochondrial respiration and ATP synthesis, while the higher concentrations of nucleoside monophosphates such as AMP, IMP, and UMP indicate insufficient ATP for synthesising higher-energy nucleotides44,45. Instead, the eyecups exhibited elevated nucleotide breakdown products, including inosine and hypoxanthine.
In ARPE-19 cells, the most abundant metabolites in ARPE-19 cells were phosphocholine and inositol, which are precursors for the synthesis of the membrane phospholipids phosphatidylinositol and phosphatidylcholine, respectively46,47. Although both are constituents of the culture medium, choline and myo-inositol are present at lower concentrations than most other metabolites (11966-025, Thermo Fisher Scientific-Gibco, Grand Island, NY). The terminally differentiated status of ARPE-19 cells may obviate the requirement of new membrane formation, causing these metabolites to accumulate intracellularly due to arrested proliferation and long culture duration.
On the other hand, the fatty acid palmitate was highly abundant in eyecups, representing approximately 8% of total metabolites compared to 1% in ARPE-19 and A549 cells. Vitamin A (retinol) in the RPE is typically stored in the form of retinyl esters, such as retinyl palmitate, which can be hydrolysed to facilitate the visual cycle, releasing all-trans-retinol and palmitate48. Moreover, palmitic acid accounts for approximately 20% of the fatty acid composition of POS49. Thus, vitamin A uptake from circulation and POS phagocytosis may provide abundant sources of palmitate in the RPE in vivo, a condition that may not be recapitulated in cultured cells. This discrepancy should be considered in metabolic studies involving in vitro models.
Unexpectedly, eyecups contained very low levels of the tripeptide glutathione, which is typically one of the most abundant non-protein molecules in the cell50 and is credited with playing a vital role in protecting against oxidative stress in the RPE26. The accumulation of glutathione breakdown product, 5-oxoproline, as well as cysteine in the eyecups in the context of low ATP and glutamate levels suggests a stalling of the gamma-glutamyl cycle, perhaps due to insufficient ATP to fuel the 5-oxoprolinase and glutamate-cysteine ligase (GCL) reactions51. While these findings may reflect compromised glutathione regeneration due to insufficient ATP, the reasons for elevated glutathione breakdown in eyecups are unclear.
Despite consuming less glucose, lactate abundance in eyecups was roughly eight-fold higher than in ARPE-19 cells. This observation was surprising given that the RPE, regarded as the most metabolically active cells in the eyecup, are believed to favour mitochondrial OXPHOS over glycolysis for ATP generation, while producing minimal lactate4. The elevated lactate in eyecups may be attributable to several factors, including highly glycolytic non-RPE cells such as endothelial cells52 or fibroblasts53, a hypoxic response to ischemia following surgical removal54, or impaired lactate export causing accumulation.
Taken together, comparison of the metabolic profiles of ARPE-19 cells and eyecups suggests that ARPE-19 cells may offer a more appropriate model of RPE metabolism. This cell line possesses a more active TCA cycle and is bioenergetically enriched with high ATP/ADP and ATP/AMP ratios, as well as relatively high NADH/NAD+. On the other hand, the eyecups exhibit a bioenergetically compromised state, with low ATP synthesis, as well as low glutathione, high lactate production, and high purine, pyrimidine, and nicotinamide metabolite turnover. The metabolism of eyecups is likely heavily influenced by surgical removal from the eye and is therefore not in a state of metabolic homeostasis.
The impact of cytokines on RPE metabolism is an emerging area of research interest due to the prevalence of chronic inflammation during aging and diseases such as AMD11. Topological analysis identified several pathways that were impacted by the cytokine cocktail in both models, including the metabolism of purines, glutathione, and several amino acids. However, this analysis identifies pathways based on the centrality of significantly changed metabolites55, regardless of direction of change. Individual metabolites within the identified pathways exhibited contrasting changes in abundance and glucose carbon labelling; increases in ARPE-19 cells typically corresponded with decreases in eyecups, or vice versa.
Several factors could explain the contrasting responses between models. This study used ARPE-19 cells, derived from a male human, and eyecups from female rats, introducing differences in both sex and species. Recent studies have highlighted sex-specific differences in the murine RPE and retinal metabolomes56, as well as the regulatory role of the metabolic transcriptional co-activator, peroxisome proliferator-activated receptor gamma coactivator-1 alpha (PGC-1α)57. While rat and human retinas share a conserved layered structure58 and vascular architecture59 and exhibit similar distributions of metabolic enzymes such as PKM2 and LDHA60, sex- and species-specific differences may contribute to the distinct metabolic responses of these models to cytokine treatment.
The eyecup consists of a complex, heterogeneous population of cells, including the RPE, endothelial cells, pericytes, Schwann cells, melanocytes, fibroblasts, and immune cells34,61. This cellular diversity means that cytokines might elicit distinct responses due to cell-specific differences in cytokine receptor expression, metabolic enzyme abundance, and basal metabolic state. Cell types within a tissue can exhibit distinct mitochondrial dynamics, influencing factors including morphology, spatial organisation, internal structure, and protein composition, reflecting the functional and bioenergetic requirements of each cell type62. In their naïve, unstimulated state, immune cells such as macrophages63, dendritic cells64, and helper T cells65 generally rely on OXPHOS for energy production and switch to aerobic glycolysis when exposed to proinflammatory stimuli. Endothelial cells primarily satisfy their energy demands through glycolysis52, producing large amounts of lactate66. Likewise, quiescent fibroblasts exhibit high glycolytic flux53, which is intensified upon activation67. This metabolic heterogeneity within the eyecup, combined with varied expression of cytokine receptors and signalling components61, may ultimately give rise to a mixed metabolic response to cytokine treatment.
The heterogenous nature of eyecups may also enable nutrient exchange between different cell populations. The community of cells within a tissue engage in extensive crosstalk, involving the exchange of metabolites, cytokines, growth factors, and other signals to maintain tissue homeostasis68,69. For example, in the heart, lactate is shuttled from cardiac fibroblasts to cardiomyocytes to support their metabolism70 and astrocytes synthesise and release glutamine and citrate to aid neuronal function in the central nervous system71. In cancer, the cells of the tumour microenvironment participate in extensive exchanges of metabolites including lactate, pyruvate, glutathione, as well as various lipids and amino acids72. Thus, the multicellular composition of eyecups may enable them to adapt to the metabolic stress imposed by cytokine treatment in a manner that is unavailable to a homogenous cell line.
While each individual cytokine induced unique effects on ARPE-19 metabolism, the cytokine cocktail most closely represents the ensemble of inflammatory factors expected to act on the RPE in AMD. The cocktail influenced numerous important components of ARPE-19 metabolism, such as increasing glucose consumption and lactate production, as well as decreasing palmitate abundance while increasing Ac-carnitine, suggesting increased fatty acid oxidation. Moreover, cytokine cocktail treatment increased proline abundance, suggesting inhibition of proline consumption. While aspartate and citrate were decreased with cocktail treatment, alanine, glycine and α-KG were increased. These changes may have consequences in AMD as many of these metabolites are either vital for RPE health or participate in metabolic exchanges between the RPE and outer retina.
Overall, the metabolomes of ARPE-19 cells and eyecups exhibit significant metabolic differences stemming from factors including culture duration, viability, and the heterogeneous composition of eyecups. Due to these inherent disparities, their metabolic responses to stimuli such as cytokine treatment are markedly different. The emergence of single cell metabolomics techniques may offer means of parsing the responses of individual cell populations in complex tissues like eyecups73. Alternatively, NaIO3 may be used to selectively induce RPE necroptosis in eyecups74, providing a means of controlling for their metabolic response. While these techniques are developed and refined, the considerable metabolic variance between these models should be considered when interpreting results and drawing definitive conclusions about RPE metabolism.
Methods
ARPE-19 cell culture
The ARPE-19 human retinal pigment epithelial cell line was obtained from the American Type Cell Collection (Manassas, VA). ARPE-19 cells at passage 12 were plated at a density of 50,000 cells per cm2 and cultured for a duration of 6 weeks. The culture medium consisted of an equal mixture of Dulbecco’s Modified Eagle Medium (DMEM) (11996-065, Thermo Fisher Scientific-Gibco, Grand Island, NY) and F12 medium (11765-065, Thermo Fisher Scientific-Gibco, Grand Island, NY), enriched with 10% foetal calf serum (FCS) (Merck-Sigma Aldrich, St Louis, MO), 1% Glutamax (Thermo Fisher Scientific-Gibco, Grand Island, NY), 100 U/mL penicillin/streptomycin (Thermo Fisher Scientific-Gibco, Grand Island, NY), as well as 10 mM nicotinamide (Merck-Sigma Aldrich, St Louis, MO) to promote differentiation as previously described30. The cells were maintained in a humidified incubator at 37 °C with 5% CO2, and the medium was replaced every five days.
Eyecup removal
The animal experiment was approved by the Animal Ethics Committee of the University of Adelaide and follows the recommendations in the ARRIVE guidelines. All methods were carried out in accordance with the relevant guidelines and regulations. All rats used in this study were female and between 7 and 13 weeks old and were humanely euthanized via CO2 asphyxiation. Following this, whole eyes were quickly enucleated and placed in ice-cold Hank’s Balanced Salt Solution (HBSS) (Thermo Fisher Scientific-Gibco, Grand Island, NY), supplemented with 100 U/mL penicillin/streptomycin. Any excess connective tissue was gently removed from the sclera, and a small incision, about 1–2 mm from the cornea, was made using a scalpel, allowing for the removal of the cornea, lens, and vitreous, revealing the posterior eyecup. After carefully removing the retina, the remaining eyecup was immediately placed in a petri dish filled with DMEM (Dulbecco’s Modified Eagle Medium) containing 5.5 mM glucose and 100 U/mL penicillin/streptomycin.
1,2 13C glucose labelling and cytokine treatment
ARPE-19 cells were pre-equilibrated in DMEM (11966-025, Thermo Fisher Scientific-Gibco, Grand Island, NY) supplemented with 5.5 mM glucose and 10% FCS, dialysed using a 14 kDa molecular weight cutoff membrane, for 24 h prior to the experiment. ARPE-19 cells in 6-well tissue culture plates (Corning, NY) and eyecups in individual wells of 12-well tissue culture plates (Corning, NY) were incubated for 12 h in DMEM (11966-025, Thermo Fisher Scientific-Gibco, Grand Island, NY) supplemented with 5.5 mM 1,2 13C glucose (453188, Merck-Sigma Aldrich, St. Louis, MO) and 10% dialysed FCS in a humidified incubator at 37 °C with 5% CO2. For cytokine treatments, 20 ng/mL TNFα (Merck-Sigma Aldrich, St Louis, MO), TGF-β2 (Merck-Sigma Aldrich, St Louis, MO), IL-6 (Thermo Fisher Scientific-Gibco, Grand Island, NY), IL-1β (Merck-Sigma Aldrich, St Louis, MO), or a cytokine cocktail containing all four cytokines were included in the ARPE-19 and eyecup culture medium.
Metabolite extraction
For metabolite extraction from ARPE-19 cells, 6-well plates of cells were placed on ice, media was aspirated, and cells were washed with ice-cold 150 mM ammonium acetate pH 7.3. I mL of 80% MeOH, pre-cooled at − 80 °C, was added per well. Cells were scraped and pipetted into 1.5 mL Eppendorf tubes on dry ice. Cells were sonicated for 90 s per tube in short bursts to homogenise the cells. Samples were then incubated at − 80 °C for 90 min. Following incubation, samples were vortexed for 10 s and centrifuged at 16,000 g for 15 min at 4 °C. The supernatant was transferred to new Eppendorf tubes and lyophilised using an Alpha 2–4 Lo Plus freeze dryer (Christ).
Following treatment, eyecups were washed with ice-cold 150 mM ammonium acetate pH 7.3, blotted dry and then snap frozen in LN2. Eyecups were crushed with a stainless-steel tissue pulveriser cooled with LN2 and transferred to Eppendorf tubes on dry ice. 1 mL of 80% MeOH, pre-cooled at −80 °C, was added to each tube and the samples were vortexed for 30 s and then incubated at − 80 °C for 90 min. Samples were warmed on ice for 5 min, vortexed again for 30 s, and then centrifuged at 16,000 g for 15 min at 4 °C. Supernatants were transferred to new tubes on ice. Any remaining MeOH was decanted from the original tubes and the pellets were resuspended in 500 µL 0.2 M NaOH and heated at 95 °C for 20 min. Samples were then cooled to room temperature, vortexed briefly, and then centrifuged at 16,000 g for 15 min. The supernatant was removed, and protein concentration was determined for each sample using a Pierce BCA kit (Thermo Fisher Scientific, Grant Island, NY). Methanol extracts equivalent to 500 µg of protein were transferred to new Eppendorf tubes and and lyophilised using an Alpha 2–4 Lo Plus freeze dryer (Christ).
Metabolomics analysis
Dried metabolites were resuspended in 50% ACN:water and 5 µl was loaded onto a Luna NH2 3 µm 100A (150 × 2.0 mm) column (Phenomenex, Torrance, CA). The chromatographic separation was performed using a Vanquish Flex UPLC (Thermo Fisher Scientific, Grant Island, NY) with mobile phases A (5 mM ammonium acetate pH 9.9) and B (ACN) at a flow rate of 200 µL/min. A linear gradient from 15 to 95% A over 18 min was followed by a 7 min isocratic flow at 95% A and re-equilibration to 15% A. Metabolites were detected with a Q Exactive mass spectrometer (Thermo Fisher Scientific, Grant Island, NY) run with polarity switching (+ 3.5 kV/− 3.5 kV) in full scan mode using a range of 70–975 m/z and 70.000 resolution. Maven (v 8.1.27.11) was used to quantify the targeted polar metabolites by AreaTop, using expected retention time and accurate mass measurements (< 5 ppm) for identification.
Data availability
Data is provided within the manuscript or supplementary information files.
References
Ng, S. K. et al. Cancer-like metabolism of the mammalian retina. Clin. Exp. Ophthalmol. 43, 367–376 (2015).
Chinchore, Y., Begaj, T., Wu, D., Drokhlyansky, E. & Cepko, C. L. Glycolytic reliance promotes anabolism in photoreceptors. Elife 6, e25946 (2017).
Hurley, J. B. Retina metabolism and metabolism in the pigmented epithelium: A busy intersection. Annu. Rev. Vis. Sci. 7, 665–692 (2021).
Kanow, M. A. et al. Biochemical adaptations of the retina and retinal pigment epithelium support a metabolic ecosystem in the vertebrate eye. Elife 6, e28899 (2017).
Chao, J. R. et al. Human retinal pigment epithelial cells prefer proline as a nutrient and transport metabolic intermediates to the retinal side. J. Biol. Chem. 292, 12895–12905 (2017).
Zhang, R. et al. Inhibition of mitochondrial respiration impairs nutrient consumption and metabolite transport in human retinal pigment epithelium. J. Proteome Res. 20, 909–922 (2021).
Bisbach, C. M. et al. Succinate can shuttle reducing power from the hypoxic retina to the O2-rich pigment epithelium. Cell Rep. 31, 107606 (2020).
Fisher, C. R. & Ferrington, D. A. Perspective on AMD pathobiology: A bioenergetic crisis in the RPE. Investig. Ophthalmol. Vis. Sci. 59, 41–47 (2018).
Ambati, J., Atkinson, J. P. & Gelfand, B. D. Immunology of age-related macular degeneration. Nat. Rev. Immunol. 13, 438–451 (2013).
Lee, K. S., Lin, S., Copland, D. A., Dick, A. D. & Liu, J. Cellular senescence in the aging retina and developments of senotherapies for age-related macular degeneration. J. Neuroinflamm. 18, 32 (2021).
Hansman, D. S., Du, J., Casson, R. J. & Peet, D. J. Eye on the horizon: The metabolic landscape of the RPE in aging and disease. Prog. Retin. Eye Res. 104, 101306 (2025).
Shu, D. Y. et al. Dimethyl fumarate blocks tumor necrosis factor-alpha-driven inflammation and metabolic rewiring in the retinal pigment epithelium. Front. Mol. Neurosci. 15, 896786 (2022).
Hansman, D. S. et al. Metabolic reprogramming of the retinal pigment epithelium by cytokines associated with age-related macular degeneration. Biosci. Rep. 44, BSR20231904 (2024).
Zhang, M. et al. Dysregulated metabolic pathways in age-related macular degeneration. Sci. Rep. 10, 2464 (2020).
Golestaneh, N., Chu, Y., Xiao, Y., Stoleru, G. L. & Theos, A. C. Dysfunctional autophagy in RPE, a contributing factor in age-related macular degeneration. Cell Death Dis. 8, e2537 (2017).
Golestaneh, N. et al. Repressed SIRT1/PGC-1α pathway and mitochondrial disintegration in iPSC-derived RPE disease model of age-related macular degeneration. J. Transl. Med. 14, 344 (2016).
Senabouth, A. et al. Transcriptomic and proteomic retinal pigment epithelium signatures of age-related macular degeneration. Nat. Commun. 13, 4233 (2022).
Adijanto, J. & Philp, N. J. Cultured primary human fetal retinal pigment epithelium (hfRPE) as a model for evaluating RPE metabolism. Exp. Eye Res. 126, 77–84 (2014).
Dunn, K. C., Aotaki-Keen, A. E., Putkey, F. R. & Hjelmeland, L. M. ARPE-19, a human retinal pigment epithelial cell line with differentiated properties. Exp. Eye Res. 62, 155–170 (1996).
Bhattacharya, S. et al. Prominin-1 is a novel regulator of autophagy in the human retinal pigment epithelium. Investig. Ophthalmol. Vis. Sci. 58, 2366–2387 (2017).
Slomiany, M. G. & Rosenzweig, S. A. Autocrine effects of IGF-I-induced VEGF and IGFBP-3 secretion in retinal pigment epithelial cell line ARPE-19. Am. J. Physiol. Cell Physiol. 287, 746–753 (2004).
An, E., Gordish-Dressman, H. & Hathout, Y. Effect of TNF-α on human ARPE-19-secreted proteins. Mol. Vis. 14, 2292–2303 (2008).
Dvashi, Z., Goldberg, M., Adir, O., Shapira, M. & Pollack, A. TGF-β1 induced transdifferentiation of RPE cells is mediated by TAK1. PLoS One 10, e0122229 (2015).
Kutty, R. K. et al. Proinflammatory cytokines decrease the expression of genes critical for RPE function. Mol. Vis. 22, 1156–1168 (2016).
Chen, X. D. et al. Oxidative stress affects retinal pigment epithelial cell survival through epidermal growth factor receptor/AKT signaling pathway. Int. J. Ophthalmol. 10, 507–514 (2017).
Sun, Y., Zheng, Y., Wang, C. & Liu, Y. Glutathione depletion induces ferroptosis, autophagy, and premature cell senescence in retinal pigment epithelial cells. Cell Death Dis. 9, 753 (2018).
Kim, S. J. et al. Activation of ERK1/2-mTORC1-NOX4 mediates TGF-β1-induced epithelial-mesenchymal transition and fibrosis in retinal pigment epithelial cells. Biochem. Biophys. Res. Commun. 529, 747–752 (2020).
Iacovelli, J. et al. PGC-1α induces human RPE oxidative metabolism and antioxidant capacity. Investig. Opthalmol. Vis. Sci. 57, 1038–1051 (2016).
Jadeja, R. N. et al. Loss of NAMPT in aging retinal pigment epithelium reduces NAD+ availability and promotes cellular senescence. Aging (Albany NY). 10, 1306–1323 (2018).
Hazim, R. A., Volland, S., Yen, A., Burgess, B. L. & Williams, D. S. Rapid differentiation of the human RPE cell line, ARPE-19, induced by nicotinamide. Exp. Eye Res. 179, 18–24 (2019).
Sinha, T., Naash, M. I. & Al-Ubaidi, M. R. The symbiotic relationship between the neural retina and retinal pigment epithelium is supported by utilizing differential metabolic pathways. iScience 23, 101004 (2020).
Yam, M. et al. Proline mediates metabolic communication between retinal pigment epithelial cells and the retina. J. Biol. Chem. 294, 10278–10289 (2019).
Boote, C. et al. Scleral structure and biomechanics. Prog. Retin. Eye Res. 74, 100773 (2020).
Nickla, D. & Wallman, J. The multifunctional choroid. Prog. Retin. Eye Res. 29, 144–168 (2010).
Hass, D. T. et al. Succinate metabolism in the retinal pigment epithelium uncouples respiration from ATP synthesis. Cell Rep. 39, 110917 (2022).
Du, J., Zhu, S., Lim, R. R. & Chao, J. R. Proline metabolism and transport in retinal health and disease. Amino Acids 53, 1789–1806 (2021).
Icard, P. et al. Understanding the central role of citrate in the metabolism of cancer cells and tumors: An update. Int. J. Mol. Sci. 22, 6587 (2021).
Sinha, T., Ikelle, L., Naash, M. I. & Al-Ubaidi, M. R. The intersection of serine metabolism and cellular dysfunction in retinal degeneration. Cells 9, 674 (2020).
Li, B. et al. Metabolic features of mouse and human retinas: Rods versus cones, macula versus periphery, Retina versus RPE. iScience 23, 101672 (2020).
Magnes, C. et al. Efficient phagocytosis requires triacylglycerol hydrolysis by adipose efficient phagocytosis requires triacylglycerol hydrolysis by adipose triglyceride lipase. J. Biol. Chem. 285, 20192–20201 (2010).
Joseph, J. W. et al. Free fatty acid-induced-cell defects are dependent on uncoupling protein 2 expression free fatty acid-induced β-cell defects are dependent on uncoupling protein 2 expression. J. Biol. Chem. 279, 51049–51056 (2004).
Xia, Z. et al. Titanium dioxide nanoparticles induce mitochondria-associated apoptosis in HepG2 cellls. RSC Adv. 8, 31764–31776 (2018).
Zeineh, N. et al. The TSPO ligands MGV-1 and 2-Cl-MGV-1 differentially inhibit the cigarette smoke-induced cytotoxicity to H1299 lung cancer cells. Biology (Basel) 10, 395 (2021).
Panayiotou, C., Solaroli, N. & Karlsson, A. The many isoforms of human adenylate kinases. Int. J. Biochem. Cell Biol. 49, 75–83 (2014).
Ballut, L., Violot, S., Kumar, S. & Aghajari, N. GMP synthetase : Allostery, structure, and function. Biomolecules 13, 1379 (2023).
Kent, C. Regulatory enzymes of phosphatidylcholine biosynthesis: A personal perspective. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1733, 53–66 (2005).
Croze, M. L. & Soulage, C. O. Potential role and therapeutic interests of myo-inositol in metabolic diseases. Biochimie 95, 1811–1827 (2013).
Mata, J. R., Mata, N. L. & Tsin, A. T. C. Substrate specificity of retinyl ester hydrolase activity in retinal pigment epithelium. J. Lipid Res. 39, 604–612 (1998).
Acar, N. et al. Lipid composition of the human eye: Are red blood cells a good mirror of retinal and optic nerve fatty acids?. PLoS One 7, e35102 (2012).
Balendiran, G. K., Dabur, R. & Fraser, D. The role of glutathione in cancer. Cell Biochem. Funct. 22, 343–352 (2004).
Kageyama, S. et al. Mechanisms of tumor growth inhibition by depletion of γ-glutamylcyclotransferase (GGCT): A novel molecular target for anticancer therapy. Int. J. Mol. Sci. 19, 2054 (2018).
De Bock, K. et al. Role of PFKFB3-driven glycolysis in vessel sprouting. Cell 154, 651–663 (2013).
Lemons, J. M. S. et al. Quiescent fibroblasts exhibit high metabolic activity. PLoS Biol. 8, e1000514 (2010).
Zhang, Z., Yao, L. I., Yang, J., Wang, Z. & Du, G. PI3K/Akt and HIF-1 signaling pathway in hypoxia-ischemia (Review). Mol. Med. Rep. 18, 3547–3554 (2018).
Xia, J., Wishart, D. S. & Valencia, A. MetPA: A web-based metabolomics tool for pathway analysis and visualization. Bioinformatics 27, 2342–2344 (2011).
Saravanan, M. et al. Tissue-specific sex difference in mouse eye and brain metabolome under fed and fasted states. Investig. Ophthalmol. Vis. Sci. 64, 18 (2023).
Taskintuna, K., Bhat, M. A., Shaikh, T., Hum, J. & Golestaneh, N. Sex-dependent regulation of retinal pigment epithelium and retinal function by Pgc-1α. Front. Cell. Neurosci. 18, 1442079 (2024).
Galindo-Romero, C. et al. The retina of the lab rat: Focus on retinal ganglion cells and photoreceptors. Front. Neuroanat. 16, 994890 (2022).
Liu, K., Akula, J. D., Falk, C., Hansen, R. M. & Fulton, A. B. The retinal vasculature and function of the neural retina in a rat model of retinopathy of prematurity. Investig. Ophthalmol. Vis. Sci. 47, 2639–2647 (2006).
Casson, R. J. et al. M-type pyruvate kinase isoforms and lactate dehydrogenase a in the mammalian retina: Metabolic implications. Investig. Ophthalmol. Vis. Sci. 57, 66–80 (2016).
Voigt, A. P. et al. Single-cell transcriptomics of the human retinal pigment epithelium and choroid in health and macular degeneration. Proc. Natl. Acad. Sci. U. S. A. 116, 24100–24107 (2019).
Kuznetsov, A. V., Hermann, M., Saks, V., Hengster, P. & Margreiter, R. The cell-type specificity of mitochondrial dynamics. Int. J. Biochem. Cell Biol. 41, 1928–1939 (2009).
Koo, S. & Garg, N. J. Metabolic programming of macrophage functions and pathogens control. Redox Biol. 24, 101198 (2019).
Krawczyk, C. M. et al. Toll-like receptor-induced changes in glycolytic metabolism regulate dendritic cell activation. Blood 115, 6–8 (2010).
Michalek, R. D. et al. Cutting edge: Distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Parra-bonilla, G., Alvarez, D. F., Alexeyev, M. & Stevens, T. Critical role for lactate dehydrogenase A in aerobic glycolysis that sustains pulmonary microvascular endothelial cell proliferation. Am. J. Physiol. Lung Cell. Mol. Physiol. 299, 513–522 (2010).
Xie, N. et al. Glycolytic reprogramming in myofibroblast differentiation and lung fibrosis. Am. J. Respir. Crit. Care Med. 192, 1462–1474 (2015).
Chen, Y. & Tang, L. The crosstalk between parenchymal cells and macrophages : A keeper of tissue homeostasis. Front. Immunol. 13, 1050188 (2022).
Berumen Sánchez, G., Bunn, K. E., Pua, H. H. & Rafat, M. Extracellular vesicles: Mediators of intercellular communication in tissue injury and disease. Cell Commun. Signal. 19, 104 (2021).
Wei, T. et al. Fibroblast-to-cardiomyocyte lactate shuttle modulates hypertensive cardiac remodelling. Cell Biosci. 13, 151 (2023).
Waagepetersen, H. S., Sonnewald, U., Larsson, O. M. & Schousboe, A. Multiple compartments with different metabolic characteristics are involved in biosynthesis of intracellular and released glutamine and citrate in astrocytes. Glia 35, 246–252 (2001).
Li, F. & Simon, M. C. Cancer cells don’t live alone: Metabolic communication within tumor microenvironments. Dev. Cell 54, 183–195 (2020).
Wei, Z. et al. Single cell metabolism methods and protocols. Methods Mol. Biol. 2064, 31–59 (2020).
Hanus, J., Anderson, C., Sarraf, D., Ma, J. & Wang, S. Retinal pigment epithelial cell necroptosis in response to sodium iodate. Cell Death Discov. 2, 16054 (2016).
Kanehisa, M. & Gto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Acknowledgements
This work was supported by the National Health and Medical Research Council of Australia [grant number 1099932] and the University of Adelaide Biochemistry Trust Fund. D. Thomas is supported by a Commonwealth Serum Laboratories Centenary Fellowship, Australian Medical Research Future Fund and National Health and Medical Research Council and The Leukemia & Lymphoma Society (www.lls.org), Snowdome Foundation (www.snowdome.org) and the Leukemia Foundation (www.leukemia.org.au) grants 6619-21 and 6650-23 and The Leukemia & Lymphoma CMML Special Initiative Award.
Author information
Authors and Affiliations
Contributions
D.S.H.: Conceptualization, Formal analysis, Validation, Investigation, Visualization, Methodology, Writing—original draft, Data curation. K.L.: Investigation, Methodology. D.T.: Resources, Funding acquisition, Writing—review & editing. R.J.C.: Conceptualization, Supervision, Funding acquisition, Writing—review & editing. D.J.P.: Conceptualization, Resources, Supervision, Funding acquisition, Project administration, Writing—review & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hansman, D.S., Lim, K., Thomas, D. et al. Distinct metabolome and flux responses in the retinal pigment epithelium to cytokines associated with age-related macular degeneration: comparison of ARPE-19 cells and eyecups. Sci Rep 15, 13012 (2025). https://doi.org/10.1038/s41598-025-93882-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-93882-w