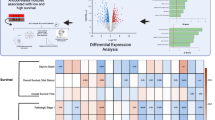
Abstract
The objective of this study was to explore the genetic link between moderate to severe asthma and early-stage lung adenocarcinoma (LUAD) using bioinformatic methods. The Cancer Genome Atlas gene-expression profiles for early-stage LUAD and GSE76225 data set for moderate to severe asthma were selected for weighted gene co-expression network analysis, and intersected with the relevant module genes and selected hub genes; the relevant network of hub genes was then determined through a protein–protein interaction network. In addition, gene-set enrichment analysis and gene-set variation analysis (GSVA) were conducted on differentially expressed genes between normal and tumor groups. Gene ontology and Kyoto Encyclopedia of Genes and Genomes pathway-enrichment analyses were applied to detect hub gene-related biological functions. Receiver operating characteristic (ROC) curves were employed to confirm the diagnostic value of hub genes. We identified four key genes, of which SFTPC exhibited relatively high value for areas under the ROC curves, indicating high diagnostic value for moderate to severe asthma. The clinical efficacy of SFTPC was thus consistent with GSVA results, indicating that moderate to severe asthma can inhibit the occurrence of early LUAD.
Similar content being viewed by others
Introduction
Asthma is a chronic inflammatory airway disease caused by specific immune mechanisms, with pathologic features such as airflow limitation and airway remodeling, while non-small cell lung cancer (NSCLC) is generally considered to be the most common type of pulmonary malignancy—particularly lung adenocarcinoma1. The combined effects of genes and the environment have now increased the incidence of both diseases.
According to statistics, the incidence of asthma continues to increase2. In 2014, there were approximately 24 million asthmatic patients in the United States, of whom over 70% were adults and 5–10% were severe or difficult-to-control asthma patients3,4. Severe asthmatics accounted for a significant proportion of medical expenses and exacerbated the burden of disease management5,6.
The Epidemiology and Natural History of Asthma: Outcomes and Treatment Regimens (TENOR I) study is currently the largest multicenter study conducted on severe or difficult-to-treat asthma, and results suggested that the risk of future deterioration increased for patients with a poor response to conventional medication and unstable control of asthma symptoms7.
The authors of the subsequent TENOR II study evaluated the clinical characteristics and risk factors related to severe or difficult to treat asthma, including total IgE, eosinophils, lung function, hormones, and FEV1 after bronchodilator use, demonstrating that nearly 50% of severe or refractory asthma patients still manifested persistent and poorly controlled asthma symptoms after more than 10 years of follow-up8. Most targeted biologics for asthma are currently monoclonal antibodies that specifically target type 2 inflammation; however, it is clear that novel therapies are also needed to treat non-type 2 asthma patients, as well as to further investigate asthma phenotypes and biomarkers9.
Lung cancer is the most common cancer worldwide and the leading cause of cancer deaths. In 2018, GLOBOCAN estimated that there were 2.09 million new cases of lung cancer and 1.76 million deaths from lung cancer worldwide10. Lung cancer is a multifactorial disease, and although smoking remains the primary factor in the development of lung cancer, the increase in non-smoking patients indicates that the occurrence of lung cancer is associated with other risk factors. According to statistics, approximately 25% of lung cancer worldwide is not related to smoking11.
An analysis based on the Southern Community Cohort Study (SCCS) revealed that asthma is associated with a high risk of lung cancer (HR, 1.25; 95% CI 1.00–1.57). From 2002 to 2011, 64,170 patients with a history of allergies and asthma were included in this study, and results showed that a total of 3628 people were diagnosed with tumors, including 667 cases of lung cancer12. A meta-analysis based on 18 studies also showed that asthma was positively associated with lung cancer risk13, and, intriguingly, lung cancer mortality related to asthma was higher among nonsmokers14.
The European Society of Allergy and Clinical Immunology (EAACI) has established a working group to conduct research and communication on the specific field of the relationship between asthma and NSCLC15, and authors have presented two different perspectives. A systematic review suggested that asthma might be positively associated with higher lung cancer risk, and that the mutation caused by oxidative stress and chronic inflammation may constitute a key mechanism in the development of lung cancer. In contrast, other work indicated that immune cells involved in the pathogenesis of asthma exerted an immune surveillance impact on tumor cells and inhibited tumorigenesis16.
The International Lung Cancer Alliance (ILCCO) also conducted a summary analysis of 16 studies on asthma and lung cancer, and ascertained that the increase in the incidence of lung cancer was principally observed in small cell and squamous cell lung cancer, and mainly within two years after the diagnosis of asthma. Patients with a 10-year history of asthma do not have an increased risk of lung cancer, and their symptoms are often severe or difficult to treat17.
Asthma is a complex and diverse disease18, and genetic research can facilitate the identification of specific subtype markers19. Lung adenocarcinoma is a disease with gene- driven characteristics, and gene-mutation analysis and targeted therapy have developed into significant detection and treatment modalities20. Although the authors of one study exploited differential analysis to screen key genes between asthma and lung cancer and identified them as biomarkers of asthma that can promote the development of cancer, the relationship between moderate to severe or refractory asthma and early LUAD is yet to be elucidated21.
We herein selected genetic data from patients with moderate to severe asthma and early-stage LUAD in order to explore associations between genes reflecting asthma severity and tumor characteristics of early-stage LUAD. Early-stage LUAD is the initial stage of cancer development and principally refers to stage I disease22,23.
Methods
Identification of gene expression data sets
From the The Cancer Genome Atlas (TCGA) (https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga), we downloaded a total of 322 gene profiles (30 normal samples and 292 tumor samples) for weighted gene co-expression network analysis (WGCNA). The GSE76225 data set was downloaded from the GEO database and used for WGCNA of moderate to severe asthma. GSE76225 contained 35 normal samples and 56 asthma samples, and the platform adopted was GPL13158 ([HT_HG-U133_Plus_PM] Affymetrix HT HG-U133 + PM Array Plate).
Data preprocessing
After obtaining the raw data sets for the gene-expression and platform files, we determined the quality of the gene-expression profiles with the R packages “affPLM” and “RcolorBrewer.” The Robust multichip averaging (RMA) method was then implemented to preprocess the data, including background correction, normalization, and expression calculation; the R package “impute” was employed to replenish the missing data.
Implementation of WGCNA
We executed the R package “WGCNA” was used for WGCNA. This analysis was based on the overall information of genes, with the aim of ascertaining the relationship between gene sets and clinical phenotype. The PickSoftThreshold function was employed to determine the appropriate soft-thresholding power to construct scale-free networks of genes, and module eigengenes were calculated for the combination of modules. Hierarchical clustering analysis was subsequently performed and the clustering results were segmented according to the set criteria to obtain different gene modules (i.e., of highly interconnected gene sets). We lastly analyzed and visualized the correlation between gene modulesand clinical phenotypes.
Screening for differentially expressed genes
We prepared gene-expression files for the normal and tumor groups, and applied the “limma” package in R to screen for differentially expressed genes (DEGs) between the two groups. The p-correction threshold was < 0.05 and |log2FC| was > 1. Finally, we exploited the R package “ggplot2” to generate a volcano map.
Gene-set enrichment analysis (GSEA)
GSEA was deployed to assess whether the expression of a specific gene set was closer to the experimental group or to the control group. The results were used to determine whether a certain pathway was activated or inhibited in a certain group of samples, and we focused on the expression trend of the entire gene set. For this study, the R package “clusterProfiler” was utilized as the analysis tool, and “c2. cp. kegg. cedivus. v2024.1.Hs. symbols” were downloaded as the reference gene set. The log2FC values were sorted from high to low, and genomes with a P-adjustment < 0.05 were considered to be significantly enriched.
Gene-set variation analysis (GSVA)
GSVA analysis was used to convert the expression-level matrix of genes between different samples into the expression-level matrix of gene sets between samples, the enrichment score of specific gene sets in each sample was calculated, and the gene sets with significant differences between samples were searched. Gene sets possess greater biological significance than genes do. We employed the R packages “GSVA” and “ggplot2” for analysis and plotting.
Construction of Venn diagrams and a protein–protein interaction (PPI) network
The Venny 2.1.0 online tool (https://bioinfogp.cnb.csic.es/tools/venny/index.html) was implemented to construct Venn diagrams, and the database STRING (STRING, https://string-db.org/) was adopted to construct a PPI network. Each node represented a protein and the line between the nodes represented the interaction between two proteins.
Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses
DAVID online tools (DAVID, https://david.ncifcrf.gov/home.jsp) were adopted to perform GO and KEGG pathway analysis24,25, with P < 0.05 generally indicating a statistically significant difference.
Analysis of ROC curves
The horizontal and vertical coordinates of the ROC curves were designated as specificity and sensitivity; and we regarded the areas under the ROC curves (AUCs) as the evaluation standard. The AUC values typically ranged from 0.5 to 1.0, indicating favorable performance. We adopted the R package “pROC” for ROC analysis.
Results
Selection of suitable samples for LUAD and asthma for WGCNA
All genes were ranked according to their standard deviations and 25 percent of the genes with the highest expression levels were selected for WGCNA. Early-stage LUAD genes (n = 14,856) and moderate to severe asthma genes (n = 1926) were retained. We used the “WGCNA” package to cluster the samples and outliers were deleted. We ultimately obtained 289 early LUAD samples and 84 moderate to severe asthma samples (Fig. 1).
Sample dendrogram and clinical trait heatmap. (A) The sample clustering and clinical traits of early-stage LUAD. (B) The sample clustering and clinical traits of moderate to severe asthma. Clinical characteristic values were converted into continuous colors, with white indicating low and red indicating high.
Soft-thresholding power
The soft threshold was the lowest power at which the scale topology fitting index curve tended to flatten out when it reached a higher value. Different power values were used to determine a suitable soft-thresholding power and R2 = 0.9 was set as the screening criterion. We adopted a soft threshold to establish an adjacency matrix and allowed the gene distribution to conform to a scale-free network based on connectivity. In addition, we noted and analyzed the mean connectivity of different power values (Fig. 2).
Soft-thresholding power and mean connectivity. (A, B) The soft-thresholding power of early-stage LUAD was 9. (C, D) The soft-thresholding power of moderate to severe asthma was 2.
Weighted co-expression network and modules
The topological overlap matrix was constructed using soft-thresholding power and the genes were were clustered based on dissimilarity between them. In addition, eigengenes of modules was calculated and similar modules were calculated and similar modules were merged based on MEDissThres = 0.25 (Fig. 3).
Gene cluster tree and module diagram. (A) Module cluster dendrogram of early-stage LUAD. (B) Gene cluster tree and its corresponding modules of early-stage LUAD. (C) Module cluster dendrogram of moderate to severe asthma. (D) Gene cluster tree and its corresponding modules of moderate to severe asthma.
Identification of key modules
A module-trait relationship heatmap was generated to analyze the relationship between modules and clinical features. The upper portion of the space was the correlation coefficient between the module and the clinical traits, and the lower part of the space was the p-value that represented the significance of the correlation coefficient. Red represents a positive correlation and blue a negative correlation; the darker the color, the stronger the correlation between the module and clinical features (the brown module might have an impact on inhibiting tumors). As for asthma, the black module was identified to be associated with disease progression (Fig. 4A,B). The gene module memberships (correlation between all gene-expression profiles and module eigengenes) and gene significance (the absolute correlation between gene-expression values and clinical phenotypes) were calculated to verify the modules that were most relevant to trait (Fig. 4C,D) (the numbers of genes in the modules are displayed in Supplementary Table S1).
The relationship between modules and clinical traits. (A) In the brown module, the correlation coefficient was − 0.69, and the p-value was 1e−41. (B) In the black module, the correlation coefficient was 0.98, and the p-value was 4e−61. (C) The module membership and gene significance of the brown module. (D) The module membership and gene significance of the black module.
Differentially expressed genes (DEG) identification
A total of 59,427 statistically significant DEGs with a correction of < 0.05 were identified between the healthy control group and early lung adenocarcinoma samples |Log2 fold-change [FC] |> 1), comprising 12,495 upregulated genes and 3063 downregulated genes. We generated a volcano map for visualization (Fig. 5).
Volcanic map of DEGs distribution. Red dot: upregulated expression; Blue dots: downregulated expression; Gray dots: meaningless expressions.
GSEA for signaling pathways
We conducted GSEA in order to explore the potential mechanisms of DEG function, with the molecular feature database (MSigDB) set used to select the signaling pathways with the most significant enrichment. GSEA revealed that CXCR4-Gn α q-PLC β (NES standardized enrichment score, NES) = − 2.212, P adjustment = 0.012), PRNPPI3K-NOX2 (NES = − 2.157, P adjustment = 0.0083), RTK-RAS-ERK (NES = − 2.391, P adjustment = 0.002), TLR2 4-MAPK (NES = − 2.039, P adjustment = 0.011), and SOD1 (NES = − 2.366, P adjustment = 0.002) were significantly enriched in the normal and downregulated groups (Fig. 6A–E).
GSEA and GSVA results. (A) CXCR4-Gnαq-PLCβ. (B) PRNP-PI3K-NOX2. (C) RTK-RAS-ERK. (D) TLR2/4-MAPK. (E) SOD1. (F) Heat map of GSVA analysis results.
GSVA for signaling pathways
We applied GSVA to evaluate the relative expression differences of pathways between the normal group and LUAD, and visualized them using heatmaps. LUAD showed significantly attenuated expression of KKEGG_MEDICAUS-ENV_FACTOR-NICOTINE-TO_JAKRENT_SIGNALING-PATHWAY, as well as significantly elevated expression of KEGG.MEDICUS-PATHOGEN-EBV-EBNA3C_TO_CELL_CYCCLE_1_SN00484 pathway (Fig. 6F).
Venn diagram and PPI network
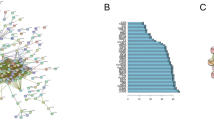
We utilized the Venny 2.1.0 online tool (https://bioinfogp.cnb.csic.es/tools/venny/index.html) to construct Venn diagrams and determined the genes between the brown and black modules (Fig. 7A). Machine learning models, including RF, SVM, GLM, GBM, KNN, NNET, LASSO, and DT, were used to screen for hub genes. Absolute residual values of each machine learning model were observed through reverse cumulative distribution curve of residues, and ROC curves were used to evaluate the diagnostic value of the machine learning model (Fig. 7B,C). Two genes (SFTPC and GPRC5A) were obtained by taking the intersection (Fig. 7D). We exploited the online database STRING (STRING, https://string-db.org/) to construct a PPI network for LUAD and asthma module genes. We then determined the connectivity of overlapping genes FEZ1, GPRC5A, SFTPC, and SMAD6 with other genes, and further constructed a PPI network (Fig. 7E). Overlapping genes were considered to be hub genes connecting LUAC and asthma.
Venn diagram and PPI network. (A) Venn diagram of the brown and black module genes. (B) Reverse cumulative distribution curve of residues. (C) ROC curve showing the diagnostic capability of eight machine learning model. (D) Gene Venn diagram of machine learning models. (E) Overlapping gene PPI network.
Enrichment analysis (GO and KEGG)
To investigate the biologic functions of genes in PPI maps, GO and KEGG pathway-enrichment analyses were conducted. According to GO term analysis, these genes exhibited robust enrichment in positive regulation of SMAD protein signal transduction, outflow tract morphogenesis, and transforming growth factor beta receptor signaling pathway (biological processes [BP]); lamellar body, extracellular space, and clathrin-coated endocytic vesicle (cellular components [CC]); and transforming growth factor beta receptor activity, type I transforming growth factor beta receptor binding, and transforming growth factor beta receptor activity type III (molecular function [MF]) (Fig. 8A–C). The enriched KEGG pathways included TGF-beta signaling pathway, Hippo signaling pathway, and AGE-RAGE signaling pathway in diabetic complications (Fig. 8D).
GO and KEGG enrichment analysis. (A) Biological process. (B) Cellular components. (C) Molecular function. (D) KEGG.
ROC of hub genes
We generated ROC curves to further verify the diagnostic value of the hub genes, and uncovered SFTPC (AUC = 0.750), SMAD6 (AUC = 0.491), FEZ1 (AUC = 0.633), and GPRC5A (AUC = 0.593) (Fig. 9A–D) as hub genes. We used five-fold cross-validation to validate the diagnostic efficacy of the AUC model and discerned that the AUC values for SFTPC and FEZ1 were 0.746 and 0.636, respectively (Fig. 9E–H), indicating that the AUC model reflected favorable diagnostic efficacy. These results suggested that our hub genes manifested the ability to assess the severity of moderate to severe asthma, and that they were negatively correlated with early LUAD.
ROC of hub genes. (A–D) ROC curves fo SFTC, SMAD6, FEZ1 and GPRC5A. (E–H) AUC plots of fivefold cross-validation.
Discussion
As a disease generated by the interaction between genes and environment, early biomarkers of LUAD have constituted an attractive area of research on tumor tissues26,27. In our study, WGCNA was performed for the first time on tumor staging, as tumor staging significantly reflects the tumor burden and potential mechanisms underlying cancer, while asthma is also a disease caused by the combined effects of genes and environment28,29. However, moderate to severe asthma can undergo a series of molecular biological changes as the disease progresses, and, therefore, the chief foci of this study were to identify specific biomarkers associated with the severity of asthma and to analyze their relationship with LUAD.
The WGCNA analysis results indicated that the black module was positively correlated with the severity of asthma, while the brown module was negatively correlated with early LUAD. After intersecting the two modules, we obtained four overlapping genes (FEZ1, GPRC5A, SFTPC, and SMAD6), as described earlier.
The loss of FEZ1 expression was positively correlated with high-grade tumor grading—particularly in NSCLC—indicating that FEZ1 may be involved in the early development of different histologic types of lung cancer30; the upregulation of GPRC5A levels in fibroblasts and blood of patients with severe asthma indicated that this gene was critical in the development of asthma31. GPRC5A is a member of the GPRC superfamily and is preferentially expressed in lung tissue, with the highest expression observed in normal bronchial epithelium. GPRC5A exerts tumor-suppressive effects in lung cancer, and this may be related to its regulation of NF-κB- and EGFR/STAT3-signaling pathways32. In patients with severe asthma, structural cells in the airway play an important role, and airway remodeling was more severe. The number of fibroblasts was also significantly higher than in general asthma patients, causing irreversible airflow limitation, a progressive decline in lung function, and difficulty in controlling respiratory distress. As a gene closely related to fibroblast function, abnormally low expression of SFPTC can promote the proliferation of lung adenocarcinoma by enhancing the PI3K/AKT pathway32. TGF-β is important in allergic asthma, and the activated TGF-β receptor complex signal is transmitted by SMAD protein. SMAD6 is an inhibitory SMAD that participates in the negative feedback regulation of the TGF-β signal33, and researchers have found that SMAD6 was downregulated in both human lung adenocarcinoma and lung squamous cell carcinoma tissues, and that its low expression was associated with poor overall survival and progression-free survival—which can be used to predict the prognosis of lung cancer34.
Our ROC curve analysis of the four genes showed that SFTPC had the highest AUC value among the genes (with an AUC = 0.750). We used five-fold cross-validation to verify the diagnostic efficacy of the AUC model and ascertained that the SFTPC AUC value was 0.746, which indicated that the SFTPC gene was useful in the diagnosis and evaluation of asthma severity.
Our GO annotation results indicated the enrichment of functions such as “positive regulation of SMAD protein signaling transduction” and “activity of transforming growth factor beta receptor,” while the KEGG annotation results revealed enrichment in the “TGF-β signaling pathway, Hippo signaling pathway, and AGE-RAGE signaling pathway.”
The expression of STAT in lung cancer is primarily concentrated in adenocarcinoma, and STAT3 can self-secrete IL-6 to activate EGFR. However, in the case of a KRAS mutation, STAT3 also harbors tumor-suppressive activity. The inhibitory effect under similar mutation backgrounds was also manifested in the STAT1 and STAT5 genes; and the role of the JAK-STAT pathway in lung cancer is therefore complex and requires further exploration. Our GSVA results showed that the JAK-STAT pathway was expressed at low levels in lung adenocarcinoma tissues, and thus the presence of a mutation background necessitates further validation35.
Through differential analysis, then, we secured a list of DEGs between the tumor group and the normal group, and through GSEA analysis we obtained pathway-enrichment information. We noted that CXCR4-Gnαq-PLCβ-, PRNP-PI3K-NOX2-, RTK-RAS-ERK-, and TLR2/4-MAPK-pathway-related genes were downregulated in our normal group, and that the pathways were inhibited. CXCR4 is associated with augmented cancer risk and a lower survival rate of NSCLC patients36, and its binding to chemokines can activate downstream pathways such as PI3K/AKT, PLC/PIP2, ERK1/2, and MAPK so as to promote tumor proliferation and angiogenesis37.
In an extant study on the relationship between asthma and NSCLC, the authors identified the key genes FKBP5, HSPD1, IL7R, FCER1A, and YY1, but due to the presence of both adenocarcinoma and squamous cell carcinoma in the NSCLC, the expression data from these genes in the two types of lung cancer were not consistent. This result suggests that we need to treat different pathologic types of lung cancer separately, and this is one of the reasons why we chose early lung adenocarcinoma for the present study21.
Conclusions
We herein analyzed the genes related to moderate to severe asthma and early lung adenocarcinoma. Due to the complex mechanisms underlying asthma, different phenotypes exhibited different characteristics. Similarly, the multifactorial risk background of lung cancer led to different gene expressions at different stages of development or of disparate pathological types. Therefore, in this study, we focused on the specific phenotypes and stages of two diseases, analyzed their relationship, and minimized the influence of confounding factors in order to obtain relatively objective results.
We identified four key genes, of which SFTPC showed relatively high AUC, indicating high diagnostic value for moderate to severe asthma. The clinical efficacy of SFTPC was also consistent with GSVA results, indicating that moderate to severe asthma can inhibit the onset of early LUAD.
Data availability
The raw data used for analysis are available in the Zenodo repository. https://sandbox.zenodo.org/records/146,471.
Abbreviations
- LUAD:
-
Lung adenocarcinoma
- TCGA:
-
The Cancer Genome Atlas
- WGCNA:
-
Weighted gene co-expression network analysis
- PPI:
-
Protein–protein interaction
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- EAACI:
-
European Academy of Allergy and Clinical Immunology
- RMA:
-
Robust multichip averaging
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under curve
- DEG:
-
Differentially expressed genes
References
Hong, Q. Y. et al. Prevention and management of lung cancer in China. Cancer 121(Suppl 17), 3080–3088. https://doi.org/10.1002/cncr.29584 (2015).
Zahran, H. S., Bailey, C. & Garbe, P. Vital signs: Asthma prevalence, disease characteristics, and self-management education: United States 2001–2009. MMWR: Morb. Mortal. Wkly. Rep. 60, 547–552 (2011).
Chipps, B. E. et al. More than a decade follow-up in patients with severe or difficult-to-treat asthma: The epidemiology and natural history of asthma: Outcomes and treatment regimens (TENOR) II. J. Allergy Clin. Immunol. 141, 1590-1597.e1599. https://doi.org/10.1016/j.jaci.2017.07.014 (2018).
Chung, K. F. et al. International ERS/ATS guidelines on definition, evaluation and treatment of severe asthma. Eur. Respir. J. 43, 343–373. https://doi.org/10.1183/09031936.00202013 (2014).
Yaghoubi, M., Adibi, A., Safari, A., FitzGerald, J. M. & Sadatsafavi, M. The projected economic and health burden of uncontrolled asthma in the United States. Am. J. Respir. Crit. Care Med. 200, 1102–1112. https://doi.org/10.1164/rccm.201901-0016OC (2019).
Nurmagambetov, T. A. & Krishnan, J. A. What will uncontrolled asthma cost in the United States?. Am. J. Respir. Crit. Care Med. 200, 1077–1078. https://doi.org/10.1164/rccm.201906-1177ED (2019).
Dolan, C. M. et al. Design and baseline characteristics of the epidemiology and natural history of asthma: Outcomes and treatment regimens (TENOR) study: A large cohort of patients with severe or difficult-to-treat asthma. Ann. Allergy Asthma Immunol. 92, 32–39. https://doi.org/10.1016/s1081-1206(10)61707-3 (2004).
Katial, R. K. et al. Changing paradigms in the treatment of severe asthma: The role of biologic therapies. J. Allergy Clin. Immunol. Pract. 5, S1–S14. https://doi.org/10.1016/j.jaip.2016.11.029 (2017).
Wenzel, S. E. et al. Evidence that severe asthma can be divided pathologically into two inflammatory subtypes with distinct physiologic and clinical characteristics. Am. J. Respir. Crit. Care Med. 160, 1001–1008. https://doi.org/10.1164/ajrccm.160.3.9812110 (1999).
Bade, B. C., Dela Cruz, C. S. & Cancer, L. Epidemiology, etiology, and prevention. Clin. Chest Med. 41(1–24), 2020. https://doi.org/10.1016/j.ccm.2019.10.001 (2020).
Vavalà, T., Giaj-Levra, M. & Novello, S. Lung cancer in never smokers: A different disease. Curr. Respir. Care Rep. 3, 26–34. https://doi.org/10.1007/s13665-013-0071-z(2014) (2014).
Kantor, E. D., Hsu, M., Du, M. & Signorello, L. B. Allergies and asthma in relation to cancer risk. Cancer Epidemiol. Biomark. Prev. 28, 1395–1403. https://doi.org/10.1158/1055-9965.EPI-18-1330 (2019).
Qu, Y. L. et al. Asthma and the risk of lung cancer: A meta-analysis. Oncotarget 8, 11614–11620. https://doi.org/10.18632/oncotarget.14595 (2017).
Brown, D. W., Young, K. E., Anda, R. F. & Giles, W. H. Asthma and risk of death from lung cancer: NHANES II mortality study. J. Asthma 42, 597–600. https://doi.org/10.1080/02770900500216234 (2005).
Jensen-Jarolim, E. et al. AllergoOncology—The impact of allergy in oncology: EAACI position paper. Allergy 72, 866–887. https://doi.org/10.1111/all.13119 (2017).
Tirado-Rodriguez, B. & Huerta-Yepez, S. Allergies: Diseases closely related to cancer. Bol. Med. Hosp. Infant. Mex. 73, 432–445. https://doi.org/10.1016/j.bmhimx.2016.11.006 (2016).
Rosenberger, A. et al. Asthma and lung cancer risk: A systematic investigation by the international lung cancer consortium. Carcinogenesis 33, 587–597. https://doi.org/10.1093/carcin/bgr307 (2012).
Binia, A. & Kabesch, M. Respiratory medicine—Genetic base for allergy and asthma. Swiss Med. Wkly. 142, w13612. https://doi.org/10.4414/smw.2012.13612 (2012).
Moffatt, M. F. et al. A large-scale, consortium-based genomewide association study of asthma. N. Engl. J. Med. 363, 1211–1221. https://doi.org/10.1056/NEJMoa0906312 (2010).
Zhang, D., Huang, Y. & Wang, H. Advances of driver gene and targeted therapy of non-small cell lung cancer. Zhongguo Fei Ai Za Zhi 17, 750–754. https://doi.org/10.3779/j.issn.1009-3419.2014.10.07 (2014).
Yang, W., Li, Z., Wang, W., Wu, J. & Ye, X. Five-hub genes identify potential mechanisms for the progression of asthma to lung cancer. Med. Baltim. 102, e32861. https://doi.org/10.1097/md.0000000000032861 (2023).
Ghysen, K. & Vansteenkiste, J. Immunotherapy in patients with early stage resectable nonsmall cell lung cancer. Curr. Opin. Oncol. 31, 13–17. https://doi.org/10.1097/cco.0000000000000497 (2019).
Padda, S. K., Burt, B. M., Trakul, N. & Wakelee, H. A. Early-stage non-small cell lung cancer: Surgery, stereotactic radiosurgery, and individualized adjuvant therapy. Semin. Oncol. 41, 40–56. https://doi.org/10.1053/j.seminoncol.2013.12.011 (2014).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Li, C. et al. Tumor-derived exosomal lncRNA GAS5 as a biomarker for early-stage non-small-cell lung cancer diagnosis. J. Cell. Physiol. 234, 20721–20727. https://doi.org/10.1002/jcp.28678 (2019).
Balata, H. et al. Prevention and early detection for NSCLC: Advances in thoracic oncology 2018. J Thorac. Oncol. 14, 1513–1527. https://doi.org/10.1016/j.jtho.2019.06.011 (2019).
Lee, J. U., Kim, J. D. & Park, C. S. Gene-environment interactions in asthma: genetic and epigenetic effects. Yonsei Med. J. 56, 877–886. https://doi.org/10.3349/ymj.2015.56.4.877 (2015).
Yang, I. V., Lozupone, C. A. & Schwartz, D. A. The environment, epigenome, and asthma. J. Allergy Clin. Immunol. 140, 14–23. https://doi.org/10.1016/j.jaci.2017.05.011 (2017).
Nonaka, D. et al. Reduced FEZ1/LZTS1 expression and outcome prediction in lung cancer. Cancer Res. 65, 1207–1212. https://doi.org/10.1158/0008-5472.Can-04-3461 (2005).
Hachim, M. Y. et al. Derangement of cell cycle markers in peripheral blood mononuclear cells of asthmatic patients as a reliable biomarker for asthma control. Sci. Rep. 11, 11873. https://doi.org/10.1038/s41598-021-91087-5 (2021).
Jiang, X. et al. GPRC5A: An emerging biomarker in human cancer. Biomed. Res. Int. 2018, 1823726. https://doi.org/10.1155/2018/1823726 (2018).
Groneberg, D. A., Witt, H., Adcock, I. M., Hansen, G. & Springer, J. Smads as intracellular mediators of airway inflammation. Exp. Lung Res. 30, 223–250. https://doi.org/10.1080/01902140490276320 (2004).
Pan, S., Zhou, G., Hu, W. & Pei, H. SMAD-6, -7 and -9 are potential molecular biomarkers for the prognosis in human lung cancer. Oncol. Lett. 20, 2633–2644. https://doi.org/10.3892/ol.2020.11851 (2020).
Erdogan, F. et al. JAK-STAT core cancer pathway: An integrative cancer interactome analysis. J. Cell. Mol. Med. 26, 2049–2062. https://doi.org/10.1111/jcmm.17228 (2022).
Zhang, C., Li, J., Han, Y. & Jiang, J. A meta-analysis for CXCR4 as a prognostic marker and potential drug target in non-small cell lung cancer. Drug Des. Dev. Ther. 9, 3267–3278. https://doi.org/10.2147/dddt.S81564 (2015).
Cheng, Z. H. et al. Chemokines and their receptors in lung cancer progression and metastasis. J. Zhejiang Univ. Sci. B. 17, 342–351. https://doi.org/10.1631/jzus.B1500258 (2016).
Acknowledgements
The authors would like to thank Dr. Puyu Wang of the Nanjing University of Science and Technology who provided many useful suggestions in expressions and wording of this manuscript. We thank LetPub (https://www.letpub.com.cn) for its linguistic assistance during the preparation of this manuscript.
Funding
This study was supported by the Special Research Project of the National Clinical Research Base of Traditional Chinese Medicine(No. 2022JDZX126). The funders had no involvement in the design of the study or in the collection, analysis, or interpretation of data or in the writing of this manuscript.
Author information
Authors and Affiliations
Contributions
Jiaqian Xue analyzed the data and wrote the paper and Qingwei Zhou revised the manuscript. All authors have read and approved the final submitted manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xue, J., Zhou, Q. Identification of hub genes between moderate to severe asthma and early lung adenocarcinoma through bioinformatics analysis. Sci Rep 15, 9243 (2025). https://doi.org/10.1038/s41598-025-94270-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-94270-0