Abstract
Neurodegenerative diseases of the eye such as optic neuritis (ON) are hallmarked by retinal ganglion cell (RGC) loss and optic nerve degeneration leading to irreversible blindness. Therapeutic interventions enhancing expression or activity of SIRT1, an NAD+-dependent deacetylase, support, at least in part, survival of RGCs in the face of injury. Herein, we used mice with experimental autoimmune encephalomyelitis (EAE) which recapitulates axonal and neuronal damages characteristic of ON to identify gene regulatory networks affected by constitutive ubiquitous Sirt1 expression in SIRT1 knock-in mice and wild-type mice upon targeted adeno-associated virus (AAV)-mediated SIRT1 expression in RGCs. RNA seq data analysis showed that the most upregulated genes in EAE mouse retinas include those involved in inflammation, immune response, apoptosis, and mitochondrial turnover. The latter includes genes regulating mitophagy (e.g., Atg4), mitochondrial transport (e.g., Ipo- 6, Xpo- 6), and mitochondrial localization (e.g., Chrna4, Scn9a). The constitutive or RGC-targeted SIRT1 overexpression in EAE mice upregulated the expression of non-mitochondrial genes such as Ecel1 and downregulated the expression of mitophagy genes (e.g., Atg2b, Arifip1) which were upregulated by EAE alone. Thus, SIRT1 induces neuroprotection by, at least in part, balancing mitochondrial biogenesis and mitophagy and/or enhancing mitochondrial self-repair to preserve the bioenergetic capacity of RGCs.
Similar content being viewed by others
Introduction
Retinal neurodegeneration characterized by optic nerve axonal damage and retinal ganglion cell (RGC) dysfunction and/or loss, is common to many optic neuropathies including optic neuritis, glaucoma, and genetic, traumatic optic, and ischemic optic neuropathies1,2. These diseases occur at various speeds and severity, and ultimately lead to irreversible loss of vision. There currently is no curative treatment available to prevent or reverse the eventual visual loss subsequent to retinal neuronal damage in most optic neuropathies. Causative factors that have been implicated in this process include inflammation, increased oxidative stress, reduction or suppression of neuronal survival factors, activation of neurodegenerative signaling pathways such as calcium influx in axons, and increased proteolytic activity prompting cytoskeletal protein degradation, axonal damage and RGC death3,4. However, although major progress has been made in understanding the mechanisms underlying the neurodegenerative process, much remains to be learned about therapeutic factors and the gene regulatory networks that can be targeted to potentially reverse or prevent neurodegeneration.
Sirtuins (SIRTs) form a class of histone deacetylase, with potential prophylactic/therapeutic effects in neurodegenerative and cardiovascular diseases5,6. SIRTs utilize NAD+as a cofactor to induce histone and non-histone protein deacetylation with or without mono-ADP-ribosylation. Although the catalytic mechanism is the same for all SIRTs, these enzymes have different substrate preferences and cellular distributions in the cytoplasm, nucleus and even mitochondria7. These highly conserved enzymes were suggested to regulate the lifespan of diverse organisms8.
Previous studies have shown that increased expression of the Sirt1 gene, the best characterized member of the Sirtuin family, led to beneficial phenotypes that may be relevant to human health and longevity9,10. Mice engineered to overexpress additional copies of Sirt1or treated with SIRT1 activators such as resveratrol (RSV) or NAD + precursors, have improved organ function, and increased disease resistance and longevity9,11,12,13. Interestingly, a transgenic mouse with high levels of Sirt1 exhibited beneficial metabolic effects including reduced blood lipid levels and improved glucose metabolism while another transgenic mouse with moderate overexpression of Sirt1 was resistant to inflammation, and was protected from liver cancer, diabetes and hepatic steatosis14,15. Yet, these Sirt1 transgenic mouse strains did not have a longer lifespan. Thus, it is possible that Sirt1 overexpression is most effective in restoring homeostasis in the context of a specific dysfunctional organ only. In addition, since SIRT1 activity regulates numerous pathways, the beneficial effects of SIRT1-based therapy could be contextual culminating in either improving or disimproving organ function.
Previous studies revealed that overexpression of Sirt1produced neuroprotective effects in several models of neurodegeneration in the eye16. Selective expression of SIRT1 in RGCs reduced RGC loss in models of optic neuritis, optic nerve crush, and glaucoma and in some models, it improved visual acuity as well17,18,19,20. Similarly, pharmacological activation of SIRT1 attenuated loss of RGCs associated with optic neuritis21,22. Understanding the molecular mechanisms of these SIRT1-based therapies requires a detailed molecular analysis of SIRT1-induced genetic reprogramming in the retina.
The aim of this study was to examine retinal transcriptomic changes associated with optic neuritis and identify those that are reversed upon targeted expression of SIRT1 in RGCs. Conceptually, identification of transcriptional differences between SIRT1-treated and non-treated retinas will help identify genes correlating with resilience and resistance to neurodegeneration and/or the molecular pathways affected by SIRT1 overexpression. To achieve this, we used a mouse model of experimental autoimmune encephalomyelitis (EAE)-induced optic neuritis, a commonly used model of multiple sclerosis, in which myelin oligodendrocyte-specific immune responses are induced by injecting myelin oligodendrocyte glycoprotein peptide (MOG35−55)19,23. EAE mice develop an autoimmune demyelinating reaction characteristic of multiple sclerosis and optic neuritis which include optic nerve inflammation, axon demyelination, and loss of RGCs and visual acuity24,25. Our previous studies have shown that AAV-mediated expression of SIRT1 in RGCs increased RGC survival and improved visual acuity compared to untreated mice19. Others have shown that genetic overexpression of Sirt1 in mice exerts similar effects of AAV-mediated expression of SIRT1 in EAE mice26. Therefore, we used an RNA-seq-based approach to determine the molecular bases of SIRT1-mediated neuroprotection in EAE mice modeling optic neuritis.
Results
Transcriptomic changes in retina of EAE mice
To examine the molecular effects of EAE and subsequent SIRT1 treatment, we set out to identify the core gene regulatory networks affected during the early stages of EAE. For this purpose, EAE was induced in four-week-old female mice through MOG35–55 immunization. The control mouse group was similarly treated as the EAE mouse group but without MOG35–55 injection. Retinas were harvested 4 weeks later. Total RNA was extracted and cDNA libraries from 3 control and 3 EAE retinas were generated and processed for RNA sequencing. To adjust for various factors such as variations of sequencing yield between samples, distribution of read counts in libraries was examined before and after normalization. The mapping rate of sample total reads to mouse reference genome ranged from 97.0 to 97.3%. We used Fragments Per Kilobase of transcript per Million mapped (FPKM) reads to represent relative gene expression abundance. Genes with an adjusted p-value < 0.05 were assigned as differentially expressed. Figure 1A is a heatmap representation of the top 50 genes that were upregulated or downregulated in EAE versus control group. Figure 1B shows the top 20 gene ontology (GO) terms that were affected in EAE mouse retinas. These include but are not limited to genes involved in inflammation and immune response, intrinsic apoptotic signaling pathway in response to endoplasmic reticulum (ER) stress, apoptotic mitochondrion changes, calcium ion transport, integrin-mediated signaling, negative regulation of metalloproteinase activity. Gene set enrichment analysis (GSEA) was then used to identify classes of genes that are over-represented in each group and that may be associated with specific pathways27. As shown in Fig. 1C-D, inflammation and apoptosis were among the top pathways with the highest normalized enrichment scores. We further validated differential expression of 4 mRNAs encoded by genes involved in various pathways. We found that C1aq, which is involved in the initiation and activation of the inflammatory and immune responses28, was significantly upregulated in EAE retinas compared to control (Fig. 1E). The gene expression of endothelin-converting enzyme-like 1 (Ecel1), a component of the plasma and ER membranes important in neural development and response to injury29, was increased in EAE retinas whereas Ppp1r1c, a marker of neuronal growth30 was significantly reduced in EAE versus control mouse retina (Fig. 1F-G). The expression of Brn3a, a marker of RGCs31 was not significantly affected by EAE (Fig. 1H).
Transcriptomic profiling of retina in the EAE model of optic neuritis. (A) Heatmap/hierarchical dendrogram clustering of 3 control (C) and 3 EAE mouse retinas four weeks after MOG immunization. (B) GO enrichment analysis of differentially expressed genes induced by EAE. The bar plot shows the highest enrichment of each category based on p adj value. (C-D) GSEA for pathway enrichment using gene expression data sets of inflammatory (C) and apoptotic (D) genes. (E-G) Differential gene expression of the C1qa, Ecel1, Ppp1r1c, and Brn3a genes was validated using qRT-PCR in control and EAE mouse retina. Data represented as mean ± SEM (n = 10). *p < 0.05; **p < 0.01.
Since mitochondrial alterations have been reported as the driver of the early damages that precede RGC degeneration in optic neuropathies32, we used GSEA to determine the enrichment of control and EAE retina transcriptomes with transcripts linked to regulation of mitochondrial homeostasis and life cycle. As shown in Table S1, EAE induced differential expression of numerous genes affecting mitochondrial biogenesis, transport, distribution and mitophagy when compared to control retinas. Of interest, mitophagy genes (e.g., Atg4b, Arfip2, Sqstm1, Vdac1) were consistently upregulated in EAE versus control healthy mouse retinas suggesting that EAE may tilt mitochondrial homeostasis towards clearance process of damaged mitochondria at the expense of mitochondrial biogenesis.
Transcriptomic changes in WT and SIRT1 KI mouse retinas after EAE
In the CNS, the Sirt1 gene has been shown to be predominantly expressed in neurons where the enzyme localizes mostly within the nucleus33,34. SIRT1 deacetylating activity regulates diverse biological processes related to axonal integrity35, cell differentiation36, apoptosis37, autophagy38, development39and metabolism36. Figure 2A shows that the steady state mRNA levels of the endogenous Sirt1 gene were not significantly altered in EAE mouse retina compared to controls. However, GSEA showed downregulation of SIRT1 targeted genes in EAE mouse retinas as their expression correlated negatively with the EAE disease state (Fig. 2B). Examples of the SIRT1 targeted genes affected are shown in Table S2. Genes affected are functionally involved in diverse biological processes including inflammation (e.g., Ccl28, Irf8), visual function (e.g., Tenm2, Cntn2), and cell growth (e.g., Ncor2).
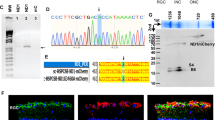
Expression of Sirt1 and its target genes in WT and SIRT1 KI mice. (A) Normalized endogenous Sirt1 mRNA levels presented as FPKM values as detected by RNA-Seq in WT mice with and without EAE. (B) GSEA comparison of SIRT1 target gene enrichment of EAE and control mouse retinas (used with permission from Kanehisa laboratories)81. Note the low abundance of SIRT1 target transcripts in EAE compared to control mouse retinas. (C) Genotype analysis of WT and SIRT1 KI mice. The Sirt1 KI allele (247-bp amplicon) and WT allele (547-bp amplicon) were detected by qPCR in SIRT1 KI mice while WT control mice expressed the WT allele (547-bp amplicon) only. Each lane represents an individual mouse. (D) Normalized Sirt1 mRNA levels presented as FPKM values as detected by RNA-Seq in WT and SIRT1 KI mice. *p < 0.05 (n = 3).
SIRT1 overexpression actively suppressed EAE clinical symptoms compared with wild-type (WT) EAE mice and provided significant neuroprotective effects, with fewer apoptotic cells found in SIRT1-overexpressing EAE mice26. To examine the molecular changes that occur in retina of SIRT1-overexpressing EAE mice, we used transgenic mice in which the Sirt1 cDNA was knocked into the β-actin locus. Previous studies have shown that these SIRT1 KI heterozygous mice express normal levels of β-actin mRNA but exhibit higher levels of SIRT1 protein in all tissues14. Figure 2C depicts genomic analysis results of WT and SIRT1 KI heterozygous mice. The latter express higher levels of Sirt1 transcripts than their WT counterparts (Fig. 2D).
We performed whole transcriptome RNA-Seq analysis on retinas and analyzed the transcriptomic changes between WT and SIRT1 KI mice after EAE. The overall similarity between the WT and SIRT1 KI mouse retina transcriptomes was assessed by the Euclidean distance between samples (Fig. 3A-B). The retina transcriptomes of the same group technical replicates (WT or SIRT1 KI) showed short distances among each other indicating that the retina samples fit the expectation of the experimental design. A total of 176 differentially expressed genes (DEGs) were identified, of which 41 were upregulated and 134 were downregulated in SIRT1 KI mice with EAE compared with WT mice with EAE. Figure 3C shows a bi-clustering heatmap of the expression profile of the top 30 differentially expressed genes sorted by their adjusted p-value. GO analysis showed that constitutive expression of Sirt1 in EAE mice reduced the transcript levels of genes involved in signaling pathways such as immune response, cell differentiation, proteolysis and cytoskeletal organization (Fig. 3D). We further validated by quantitative PCR differential expression of several genes. As shown in Fig. 3E, Sirt1 overexpression further increased Ecel1 mRNA levels which were already significantly elevated in EAE mice. Sirt1 overexpression returned Ppp1r1c mRNA levels, which were reduced by EAE, to levels found in control mice (Fig. 3F). In contrast, Sirt1 overexpression significantly reduced the expression of Elf3 (Fig. 3G), a transcription factor highly expressed in epithelial cells40.
Differential gene expression analysis in WT and SIRT1 KI mice with EAE. (A) Overall similarity among WT (n = 3) and SIRT1 KI (n = 3) retinal transcriptomes assessed by the Euclidean distance between samples. In the diagram, samples are clustered using the distance. The shorter the distance, the more closely related the samples are within each group. (B) Volcano plot showing DEGs in WT and SIRT1 KI mice with EAE. Each data point in the scatter plot represents a gene. The log2 fold change of each gene is represented on the x-axis and the log10 of its adjusted p-value is on the y-axis. Genes with an adjusted p-value less than 0.05 and a log2 fold change greater than 1 are indicated by red dots. These represent up-regulated genes. Genes with an adjusted p-value less than 0.05 and a log2 fold change less than − 1 are indicated by green dots. These represent down-regulated genes. (C) Bi-clustering heatmap for visualization of the expression profile of the top 30 DEGs sorted by their adjusted p-value. (D) Significantly differentially expressed genes clustered by their GO and the enrichment of GO terms was tested using Fisher exact test. GO terms significantly enriched with an adjusted P-value less than 0.05 in the differentially expressed gene sets are shown. (E-G) Differential gene expression of the Ecel1, Ppp1r1c, and Elf3 genes validated using qRT-PCR in WT and SIRT1 KI retinas of mice with or without EAE. Data represented as mean ± SEM (n = 10). Significance was analyzed by ANOVA of repeated measures followed by Tukey’s honest significant difference test. *p < 0.05; **p < 0.01.
Pathway analysis revealed that the transcriptome of the SIRT1 KI mice with EAE was less enriched with apoptotic, inflammatory and protein unfolding response genes than the retinal transcriptome of WT mice with EAE (Fig. 4A-C). Conversely, SIRT1 targeted genes were highly expressed/enriched in SIRT1 KI mice with EAE compared to WT mice with EAE (Fig. 4D). The expression of a set of 6 SIRT1 targeted genes was reversed in SIRT1 KI EAE mice compared to their WT counterparts with EAE (Tables S2 and S3). These include Adamtsl1, Zfhx4, Dennd3, Ptprm, Ccdc68, andCsmd2. However, the expression of a new set of SIRT1 targeted genes was upregulated as a result of Sirt1 overexpression in EAE mice. These include Bank1, Kcnt2, Fcmr, Tenm2 and Xkr4.
Differential expression of genes involved in apoptosis, inflammation, and mitochondrial turnover in EAE mice with constitutive expression of Sirt1. (A-E) GSEA performed with signatures for apoptosis, inflammation (TNF-alpha signaling), unfolded protein, SIRT1 target, and mitochondrial turnover-associated genes (used with permission from Kanehisa laboratories). (F) Heatmap of mitochondria-associated genes with highest enrichment scores in WT and SIRT1 KI mice with EAE. (G-I) Differential gene expression of the Atg4b, Arfip2, and Slc25a46genes validated using qRT-PCR in WT and SIRT1 KI retinas of mice with or without EAE. P values indicate the significance compared to EAE untreated mice (n = 10). Data represented as mean ± SEM. Significance was analyzed by ANOVA of repeated measures followed by Tukey’s honest significant difference test. *p < 0.05.
Since EAE caused major changes in mitochondrial turnover, we examined the transcriptomic changes associated with genes functionally important in mitochondrial dynamics (Fig. 4E-F). Our data showed that the expression of genes like Hk2 and Vsp13 d, predicted to be involved in mitochondrion organization and positive regulation of mitophagy41 were upregulated in SIRT1 KI mice with EAE. We further used qPCR to validate the differential expression of several genes including Atg4b which induces mitophagy by promoting phagophore formation42, Arfip2, an Atg4bproximity partner that promotes mitophagy through the PINK-PARK pathway43, and Slc25a46, a mitochondrial outer member protein-encoded gene, which when mutated in patients induces peripheral neuropathy and optic atrophy44. As shown in Fig. 4G-I, Sirt1 overexpression in EAE mice significantly reduced the expression of Atg4b and Arfip2 and tended to normalize the levels of Slc25a46, suggesting that Sirt1 overexpression reduced, at least in part, the mitochondrial changes induced by EAE.
Transcriptomic changes associated with AAV-mediated selective expression of SIRT1 in RGCs in EAE mice
Since RGCs are the retinal cells foremost susceptible to injury and loss in optic neuritis45, we employed a more targeted approach based on AAV-mediated selective expression of human SIRT1 in RGCs and examined the subsequent effects of EAE on the retina transcriptome. Accordingly, female C57Bl6/J mice received at postnatal week 4 intravitreal injections of AAV-SIRT1 vector containing the RGC-selective g-synuclein (SNCG) promoter46 to target SIRT1 expression to RGCs without affecting other retinal cell types (Fig. 5A). EAE was then induced in these mice two weeks later through MOG35–55 immunization. We compared the transcriptomic changes between EAE mouse retinas injected with the vehicle (EAE group) and the AAV2.SNCG.SIRT1 vector (EAE-AAV-SIRT1 group) four weeks after EAE induction. We first performed a principal component analysis (PCA) across all samples to assess the overall relationship between the transcriptomes of the replicates in the EAE and EAE-AAV-SIRT1 groups. PCA of gene expression profiles separated EAE from EAE + AAV-SIRT1 groups along both components suggesting that most variance of gene expression was between AAV-SIRT1- and vehicle-treated EAE mice (Fig. 5B). A total of 468 DEGs with an adjusted p-value < 0.05 were identified (Fig. 5C-D) of which 255 were upregulated and 213 were downregulated. GO analysis showed that more than 35 biological processes were affected (Fig. 5E). Together with other immune-related pathways, phagocytosis, inflammatory and metabolic processes were among the terms associated with AAV-mediated SIRT1 expression in RGCs of EAE mice. Quantitative PCR analysis further validated differential expression of Ecel1, Ppp1r1c, and Elf3. As shown in Fig. 6A-C, targeted expression of SIRT1 to RGCs of EAE mice increased the expression of Ecel1 and Ppp1r1c and decreased the expression of Elf3 compared to untreated EAE mice. In addition, GSEA showed that AAV-mediated SIRT1 expression induced gene expression changes associated with mitochondrial turnover (Fig. 6D). Of these, we validated the expression of Arfip2, Atg4b, and Slc254a in the control, EAE and EAE + AAV-SIRT1 groups. As shown in Fig. 6E-G, the expression of Atg4b was significantly downregulated in AAV-SIRT1-treated EAE mice while the expression of both Arfip2 and Slc254a was not affected. Thus, Atg4b expression is dependent on SIRT1 activity in RGCs.
Transcriptomic changes in EAE mouse retinas with targeted expression of SIRT1 to RGCs. (A) Schematic representation of the AAV2-SNCG-SIRT1 used to target SIRT1 expression to RGCs. The AAV2 capsid was designed to contain codon optimized SIRT1 cDNA under the control of hSNCG promoter combined with different regulatory sequences (eCMV: CMV enhancer; iSV40: SV40 intron; WPRE: woodchuck hepatitis post-transcriptional regulatory element; ITR: inverted terminal repeats; hGH Poly A: growth hormone polyadenylation signal). (B) PCA showing the similarity between samples of EAE and EAE AAV-SIRT1 groups based on the distance between samples. (C) Volcano plot showing DEGs in EAE mice with and without prior intravitreal injection with AAV-SIRT1. Each data point in the scatter plot represents a gene. Genes with an adjusted p-value less than 0.05 and a log2 fold change greater than 1 are indicated by red dots. These represent up-regulated genes. Genes with an adjusted p-value less than 0.05 and a log2 fold change less than − 1 are indicated by green dots. These represent down-regulated genes. (D) Bi-clustering heatmap for visualization of the expression profile of the top 30 differentially expressed genes sorted by their adjusted p-value in EAE and EAE with AAV-SIRT1 treatment. (E) Significantly differentially expressed genes clustered by their GO and the enrichment of GO terms was tested using Fisher exact test. GO terms significantly enriched with an adjusted P-value less than 0.05 in the differentially expressed gene sets are shown.
Differential gene expression analysis of EAE mouse retinas with targeted expression of SIRT1 to RGCs. (A-C) Differential gene expression of the Ecel1, Ppp1r1c, and Elf3 genes validated using qRT-PCR in WT and AAV-SIRT1 treated retinas of mice with or without EAE. (D) GSEA comparison of mitochondria-associated gene enrichment of EAE mice treated or untreated with AAV-SIRT1. Note the low abundance of SIRT1-targeted transcripts in EAE AAV-SIRT1 group compared to EAE group. (E-G) Differential gene expression of theAtg4b, Arfip2, and Slc25a46 genes validated using qRT-PCR in WT and AAV-SIRT1 treated mouse retinas with and without EAE. (H-J) GSEA comparison of SIRT1 target gene enrichment of EAE mice treated or untreated with AAV-SIRT1 or overexpressing SIRT1. Note the low abundance of SIRT1-targeted transcripts in EAE AAV-SIRT1 compared to EAE SIRT1 KI mouse group. Data represented as mean ± SEM (n = 7–10). Significance was analyzed by ANOVA of repeated measures followed by Tukey’s honest significant difference test. *p < 0.05; **p < 0.01; ***p < 0.001.
We further determined and contrasted the profile of SIRT1 targeted genes in the EAE + AAV-SIRT1-treated group versus the EAE group. As shown in Fig. 6H, RGC-selective expression of SIRT1 was associated with enrichment of retina transcriptome with SIRT1-targeted genes in mice with EAE. The expression of a subset of SIRT1-targeted genes was reversed in EAE mouse retinas upon SIRT1 expression in RGCs (Table S4). Interestingly, AAV-SIRT1 injected retinas were comparatively more enriched with SIRT1-targeted genes than retinas of SIRT1 KI mice (Fig. 6I-J). The expression of genes like Adamtsl1, Zfhx4, Dennd3, and Csmd2 was reversed in EAE mice upon either overexpression of Sirt1 in SIRT1 KI mice or AAV-mediated expression of SIRT1 in RGCs (data not shown). This suggests that these genes could be selectively targeted by SIRT1 in RGCs.
Discussion
Neurodegeneration, a key component of optic neuritis, is refractive to anti-inflammation therapies which, though they reduce the immune response, do not prevent RGC death and progressive vision loss suggesting that neurodegeneration is not entirely dependent on ongoing inflammation47. RGCs have an abundant number of mitochondria to keep up with their high metabolic demands. Mitochondrial abnormalities have been implicated in the pathogenesis of many neurodegenerative diseases which led to the hypothesis that rescue and/or preservation of mitochondrial function could increase neuronal survival and preserve axonal function and integrity48,49. Mitochondria undergo fusion to temper damages through genetic complementation or fission to meet increased energy demands. Their number at any given time is reflective of balanced degradation and formation through mitophagy and biogenesis processes50. Interestingly, SIRT1 was shown to regulate some aspects of energy metabolism through the mitochondria, by, at least in part increasing mtDNA copy number51. Paradoxically, other studies have implicated SIRT1 in accelerating autophagic degradation of mitochondria and promoting the destruction of damaged or aged mitochondria52. By analyzing the molecular pathways associated with SIRT1-based therapy in the EAE model of retinal neurodegeneration, our study identified a potential role of SIRT1 in balancing mitochondrial turnover.
First, our results demonstrated that the retinal transcriptome of EAE mice was highly enriched with gene transcripts involved largely in neuroinflammation, apoptotic mitochondrial changes and ER calcium concentration and stress pathways, consistent with previous studies of DEGs in optic neuritis53. GO analyses showed that Timp1 and Atp2a1 (aka Serca1) represented hub genes orchestrating different cellular processes linked to EAE including calcium binding and transport, negative regulation of catalytic activity, response to ER stress, and negative regulation of metalloproteinase activity. Our GSEA further showed that several genes functionally associated with mitochondrial damages were upregulated in EAE mouse retinas. The list of these genes included, but was not limited to, Slc4a5, which induces abnormal mitochondrial distribution54; Slc25a5which is linked to mitochondria metabolism, ADP/ATP ratio and oxidative stress55; Sybu, which regulates mitochondrial transport56; Dynll1, which promotes the release of mitochondrial reactive oxygen species57; and Sqstm1, an autophagy-cargo receptor which promotes mitophagy58,59. Thus, results suggest oxidative mitochondrial damages are an important contributor to RGC degeneration and functional visual deficits associated with EAE.
In addition, other genes previously shown to be associated with optic nerve damage were strongly upregulated in EAE mouse retinas. In particular, Ecel1 was one of the top 10 most upregulated genes in EAE compared to control retinas. Previous studies have shown that Ecel1 (aka damage-induced neuronal endopeptidase [Dine]) is a nerve regeneration-associated molecule, whose expression was found to be upregulated in response to nerve injury in both the peripheral and central nervous systems60. The Ecel1 gene is transcriptionally regulated by activating transcription factor 3 (ATF3), known as an axonal injury marker and key transcription factor for nerve regeneration61,62. The reason why Ecel1 expression was upregulated in EAE mouse retinas and in other models of neuronal injury despite the fact that neurons like RGCs have a limited ability for axonal regeneration remains a conundrum. One potential explanation is that Ecel1 expression was only transiently elevated and does not last long enough to allow axonal regeneration. The observation by Nakagomi et al. that peripheral nerve neurons, which exhibit prolonged Ecel1 expression after nerve injury, were able to survive and regenerate after injury is in support of this hypothesis63. As an orphan protease, Ecel1 cleaves unidentified substrates, presumably peptides, on regenerating axons or their cell body to elicit intercellular signaling between injured neurons and surrounding glial cells or neighboring neurons64. Further studies are needed to assess potential therapeutic effects of sustained expression of Ecel1 in optic neuritis and other optic neuropathy models.
Another major goal of our study design was to determine how SIRT1 overexpression preempts, at least in part, some of the deleterious effects of EAE on the neural retina. Our gene enrichment data showed that, even though Sirt1 gene expression was not significantly affected in retinas of EAE mice, the expression of numerous SIRT1-targeted genes was reduced in EAE versus control mouse retinas. GO analysis showed that differentially expressed SIRT1 targeted genes included those functionally associated with mitochondrial function and behavior. The list of these genes included autophagy-related protein 2 homolog A (Atg2a) which is involved in mitophagy and phagophore expansion65; importin 8 (Ipo8) and exportin 6 (Xpo6), both involved in mitochondrial transport66; broad substrate specificity ATP-binding cassette transporter (Abcg2a) involved in monocarboxylic acid transport and establishment of protein localization to mitochondrion67; Chrna4 and Scn9a which are involved in mitochondrial membrane depolarization68; sperm-associated antigen 6 (Spag6) involved in regulation of mitochondrial membrane polarity69; and adhesion G protein-coupled receptor B3 (Adgrb3) involved in synapse pruning70. These genetic alterations lead to dysregulation of mitochondrial function further exacerbating both neuroinflammation and neurodegeneration. RGCs have their soma in the ganglion cell layer of the retina and long axons that extend through the optic nerve to form distal terminals and connections in the brain. Because of this unique cytoarchitecture and highly compartmentalized energy demand in the soma and axons, mitochondrial dysfunction ineluctably depletes the bioenergetic capacity of RGCs and compromises their survival.
To better understand the underlying mechanisms of SIRT1 overexpression and its impact on mitochondrial function in the retina as whole, we analyzed and contrasted the transcriptomic changes associated with EAE in WT and SIRT1 KI mice which constitutively overexpress Sirt1. As expected, the transcriptome of the SIRT1 KI mouse retina after EAE was highly enriched with SIRT1-targeted genes compared to WT EAE mouse retinas. Functional analysis showed that only two DEGs were associated with mitochondrial function. These include neuronal acetylcholine receptor subunit alpha- 4 (Chrna4) and Sodium channel protein type 9 subunit alpha (Scn9a); both were upregulated in SIRT1 KI mouse retinas with EAE compared to WT with EAE. The overexpression of Scn9a, which encodes a voltage-gated sodium channel, has been reported to be associated with small fiber neuropathy and affects mitochondrial membrane depolarization68. Conversely, the expression of pro-mitophagy factors like Atg4b and Arfip2 was significantly reduced in retinas of SIRT1 KI mice with EAE compared to WT with EAE. As well, the expression of Slc25a46, which regulates mitochondrial dynamic and morphology44 was significantly reduced as a result of overexpression of Sirt1 in EAE mice. Thus, constitutive expression of Sirt1 tended to reverse the mitochondrial dynamics linked to EAE and mitigated the mitophagy process. However, the ubiquitous expression of the Sirt1 gene in the SIRT1 KI mice could manifest its effects in the retina in several other ways. For instance, SIRT1 has been shown to exert an anti-inflammatory role in many disease models71. In particular, SIRT1 can directly and indirectly regulate NF-kB, a well-known regulator of the transcription of proinflammatory genes including Tnfa, Il6 and Il1b72. As a deacetylase, SIRT1 suppresses NF-kB signaling through interaction with and deacetylation of RELA/p65 subunit of NF-kB to suppress inflammatory responses73. Additionally, constitutive expression of Sirt1 may exert its effects through signaling pathways that are essential to retinal homeostasis. Parker et al. have studied the neuroprotective effects of SIRT1 in C. elegansand showed that constitutive overexpression of SIRT1 in these nematodes reduced the vulnerability of neurons to cell death through the longevity-promoting factor the Forkhead box protein O (FOXO) pathway74. FOXO is indeed a substrate for SIRT1, and together they form a complex with b-catenin in the nucleus to modulate the expression of genes like Ucp2, which promotes mitochondrial biogenesis and elevated ATP levels in the cells. This is consistent with Andrews et al. study that showed that Ucp2 overexpression has a neuroprotective effect in mouse models of Parkinson’s disease75. The expression of other genes like Ecel1, which promotes neuronal survival, was significantly increased in EAE mouse retinas of SIRT1 KI mice compared to EAE WT mice. Conversely, the expression of Elf3, was significantly decreased in the EAE – SIRT1 KI mouse retina compared to either WT EAE or SIRT1 KI mouse retinas. This effect could be detrimental to the health of the neural retina. Indeed, loss of function of Elf3 affected brain morphology and resulted in microphthalmia in zebrafish76. It was suggested that Elf3 expression was critical for optic nerve fasciculation/arborization and spinal motor neuron projections. Thus, decreased expression of Elf3 in WT and SIRT1 KI mouse retinas of EAE mice could underlies some persistent alterations that SIRT1 overexpression could not overcome. Additional studies focusing on Elf3’s role during neurodegeneration are needed to understand its function and relevance to the pathogenesis of neurodegenerative diseases.
Furthermore, targeting the expression of SIRT1 to RGCs allowed us to pinpoint the transcriptomic changes associated with SIRT1 overexpression in these cells and the subsequent changes to the entire transcriptome in the retina. Using the AAV vector and an RGC-selective promoter to deliver SIRT1 to RGCs, we found that this approach resulted in inducing molecular changes similar to those observed in mice with constitutive overexpression of Sirt1, including significant downregulation of Atg4b suggesting reduced mitophagy effects in EAE mice. The effects on other genes (e.g., Arfip1 and Slc25a46) linked to ribosomal function were not significantly altered compared to WT and WT with EAE suggesting that the Arfip1 and Slc25a46 pathways may be independent of SIRT1 activity in RGCs. The expression of other genes like Ecel1 and Elf3 was similarly affected after either AAV-mediated SIRT1 or constitutive expression of Sirt1 in EAE mouse retinas.
In conclusion, our molecular analyses showed that SIRT1 overexpression regulates mitochondrial quality through modulation of the expression of genes involved in mitochondrial turnover. Excessive production of ROS impairs the structure and function of RGC mitochondria which become unable to carry out self-homeostasis/repair, including the inability to complete normal mitochondrial biogenesis. This is consistent with previous studies showing that SIRT1 activity reduced the levels of damaging ROS77. Known targets of SIRT1 deacetylase activity include p53, the myocyte-specific enhancer factor 2 (MEF2), FOXO and peroxisome proliferator-activated receptor-γ coactivator- 1α (PGC- 1α) all of which promote gene expression programs that increase mitochondrial function78. In particular, SIRT1 physically interacts with and deacetylates PGC- 1a at 13 lysine residues, which triggers an increase in PGC- 1a transcriptional activity and subsequently reduced ROS production and increased cell survival51. The SIRT1/PGC- 1a signaling axis could be playing a key role in the neuroprotective effects of SIRT1 in not only optic neuritis but also other ocular and non-ocular neurodegenerative diseases.
Methods
Mice. We used four-week-old female C57Bl/6 J mice designated as WT and SIRT1 knock in (SIRT1 KI) mice (Jackson Laboratory stock #013080). Like human susceptibility to multiple sclerosis, female mice are significantly more susceptible to EAE disease following immunization with myelin peptides. Mice were housed in the animal facility at the University of Pennsylvania in a 12-hour light/dark cycle. Experiments were carried out according to the institutional ethical guidelines and following an animal protocol approved by the Institutional IACUC and in compliance with federal regulations. All authors followed he ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines.
AAV.SNCG. SIRT1 vector and intravitreal injection
AAV vector (AAV2.SNCG.SIRT1)46used in this study consists of an AAV2 capsid, the CMV enhancer/CBA promoter derived from Invivogen pDRIVE CAG plasmid (San Diego, CA, USA), the codon optimized SNCG (gamma-synuclein) promoter79, first SV40 intron, the cDNA encoding a codon optimized human SIRT1cDNA, the woodchuck hepatitis post-transcriptional regulatory element (WPRE) (i.e., GenBank accession no. J04514), and the bovine growth hormone (bGH) polyadenylation signal. AAV vectors were generated as previously described46 and stored in 0.001% Pluronic F- 68 at − 80 °C until just before use, and then diluted to the appropriate concentration. All vectors used have been produced by the Research Vector Core at the Center for Advanced Retinal and Ocular Therapeutics at the University of Pennsylvania.
Four-week-old mice were anesthetized with isoflurane inhalation, and their pupils were dilated by topical application of tropicamide 1% for 1 min, followed by application of phenylephrine 2.5%. A 10-µL Hamilton syringe attached to a 33-gauge blunt-end needle was inserted ~ 1.5 mm into the vitreous cavity. The needle was positioned directly above the optic nerve head with its opening oriented toward the temporal retina. Two microliters of AAV vector preparation (1 × 1010vector genomes) were intravitreally injected into each eye of six-week old mice. Vehicle-treated eyes were injected with an equivalent volume of vector dilution buffer (0.001% Pluronic F68 in PBS). Mice were kept warm while they were awakening from anesthesia. We have previously shown that the AAV vectors used had high transduction efficiency of RGCs at varying locations from the site of injection in the eye (45–75%)46.
MOG immunization for EAE induction
As in prior studies22, eight-week-old mice were immunized subcutaneously at two sites on the back, each with 150 µg myelin oligodendrocyte glycoprotein (MOG35–55, GenScript, Piscataway, NJ, USA) emulsified in complete Freund’s adjuvant (CFA; Difco Laboratories) containing 2.5 mg/ml heat-killed Myobacterium tuberculosis. Mice were further injected intraperitoneally with 200 ng pertussis toxin (PTX) (List Biological, Campbell, CA, USA) in 0.1 mL of PBS at 0 h and 48 h postimmunization with MOG35–55. Control mouse groups were injected with the same volume of Freund’s adjuvant emulsion and with PTX at day 0 and day 2 as the EAE mouse group but without MOG33−55. Mice were weighed daily to ensure weight loss did not exceed 20% of starting weight. EAE clinical manifestations of progressive ascending paralysis were assessed daily using a standard scoring system22. Both WT and SIRT1 KI mice showed typical features of EAE that worsened over time (4 weeks) including loss of tail tone, waddling gait, partial or total paralysis of one or two limbs, etc.
RNA isolation, library preparation and sequencing, and data analysis
Four weeks after MOG immunization, SIRT1 KI and AAV-SIRT1-injected WT mice and their respective controls were euthanized with isoflurane followed by cervical dislocation, and retinas were rapidly isolated and snap frozen in liquid nitrogen. Total RNA was extracted from homogenized mixtures of retinas using RNeasy purification kit according to the manufacturer’s instructions (Qiagen). Sample RNA concentrations and purity were measured using the Thermo Scientific NanoDrop 8000 UV-Vis Spectrophotometer.
RNA sequencing was performed with three biological replicates per group of mice by Genewiz/Azenta (New Jersey, USA) where the cDNA libraries were prepared and sequenced using Illumina Hiseq, 2 × 150 bp paired-end chemistry. Briefly, quality control of RNA integrity was confirmed followed by rRNA depletion. Prior to cDNA library preparation, samples were treated for initial polyA selection, mRNA fragmentation and random priming with first and second strand cDNA synthesis. Thereafter, adaptor ligation, polymerase chain reaction (PCR) enrichment, and Illumina NovaSeq technology–based sequencing with 2 × 150–base pair (bp) read length were performed. The library preparation was specific to each strand. Library quality was assessed on the Agilent Bioanalyzer 2100 system. To remove adaptor sequence and poor-quality nucleotide reads, the trimmomatic v.0.36 software was used. A minimum of 25 M reads were generated from each RNA sample, which resulted in a sequencing depth sufficient for transcriptome analysis. The trimmed reads were mapped to the Mus musculus GRCm38 reference genome available on ENSEMBL using the STAR aligner v.2.5.2b. The STAR aligner is a splice aligner that detects splice junctions and incorporates them to help align the entire read sequences. Unique gene hit counts were calculated with the feature counts from the Subread package v.1.5.2. The hit counts are reported based on the gene ID feature in the annotation file. Only unique reads that fall within exon regions were considered. BAM files were generated as a result of this step.
Read counts were normalized using DESeq2 software80. Transcripts with normalized counts less than 5 were removed and this resulted in a dataset of less than 20,000 transcripts. To capture small set of “principal components” that reflect a large proportion of the variance in the original data, PCA was performed using regularized log-transformed data. The proportion of the variance captured by each of the principal components was determined. Weighted gene co-expression network analysis was conducted to identify modules (or sets) of transcripts that are co-expressed. Comparison of gene expression levels between the different sample groups was also obtained using DESeq2. P values and log2 fold changes were calculated using the Wald test. Genes with an adjusted P value (Padj) of < 0.05 were considered to be DEG. The Kyoto Encyclopedia of Genes and Genomes aka KEGG (Kanehisa Laboratories, Kyoto, Japan) pathway analysis (Pathview Web) was used for pathway-based data integration and visualization59.
Quantitative real-time PCR.
Eyes were removed, then retinas were dissected and frozen in liquid nitrogen. RNA was extracted following the RNeasy Mini Kit (Qiagen, Cat# 74004). Sample RNA concentrations and purity were measured using the Thermo Scientific NanoDrop 8000 UV-Vis Spectrophotometer. To validate the expression patterns revealed by RNAseq, the expression of a selected set of genes was measured using quantitative RT- PCR. For reverse transcription, RNA (800 ng) was converted to cDNA using Advantage® RT-for-PCR Kit (Takara, Cat# 639506) protocol. Expression of selected DEGs was determined using qRT-PCR. Master mix of primers specific to DEGs were obtained from Integrated DNA Technologies. Gapdh-specific primers were used to account for variations in baseline RNA levels between samples. RT-qPCR was performed using TaqMan Fast Advanced Master Mix (Applied Biosystems, Vilnius, Lithuania, Cat# 4444554). Gene expression was calculated using the 2−ΔΔCT method.
Statistical analysis
Data were presented as the mean ± SEM. Groups were compared using an unpaired two-tailed student’s t-test. For multiple group analyses, statistical significance was determined using either the Mann–Whitney U test, one-way ANOVA, or ANOVA of repeated measures followed by Tukey’s honest significant difference test as indicated in each figure legend. The GraphPad Prism 5.0 (GraphPad Software, Inc., La Jolla, CA, USA) was used.
Data availability
RNA seq data deposited in NCBI Gene Expression Omnibus under the series record GSE275359.
References
Casini, G., Ola, M. S., Koulen, P. & Editorial Neurodegeneration and neuroprotection in retinal disease, II. Front. Neurosci. 16, 1009228. https://doi.org/10.3389/fnins.2022.1009228 (2022).
Carelli, V., La Morgia, C. & Yu-Wai-Man, P. Mitochondrial optic neuropathies. Handb. Clin. Neurol. 194, 23–42. https://doi.org/10.1016/B978-0-12-821751-1.00010-5 (2023).
Kang, E. Y. et al. Role of oxidative stress in ocular diseases associated with retinal ganglion cells degeneration. Antioxid. (Basel). 10 https://doi.org/10.3390/antiox10121948 (2021).
Wong, K. A. & Benowitz, L. I. Retinal ganglion cell survival and axon regeneration after optic nerve injury: role of inflammation and other factors. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms231710179 (2022).
Hong, J. Y. & Lin, H. Sirtuin modulators in cellular and animal models of human diseases. Front. Pharmacol. 12, 735044. https://doi.org/10.3389/fphar.2021.735044 (2021).
Wu, Q. J. et al. The Sirtuin family in health and disease. Signal. Transduct. Target. Ther. 7, 402. https://doi.org/10.1038/s41392-022-01257-8 (2022).
Singh, V. & Ubaid, S. Role of silent information regulator 1 (SIRT1) in regulating oxidative stress and inflammation. Inflammation 43, 1589–1598. https://doi.org/10.1007/s10753-020-01242-9 (2020).
You, Y., Liang, W. & SIRT1 and SIRT6. The role in aging-related diseases. Biochim. Biophys. Acta Mol. Basis Dis. 1869, 166815. https://doi.org/10.1016/j.bbadis.2023.166815 (2023).
Bonkowski, M. S. & Sinclair, D. A. Slowing ageing by design: the rise of NAD(+) and sirtuin-activating compounds. Nat. Rev. Mol. Cell. Biol. 17, 679–690. https://doi.org/10.1038/nrm.2016.93 (2016).
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464. https://doi.org/10.1038/366461a0 (1993).
Satoh, A. et al. Sirt1 extends life span and delays aging in mice through the regulation of Nk2 homeobox 1 in the DMH and LH. Cell. Metab. 18, 416–430. https://doi.org/10.1016/j.cmet.2013.07.013 (2013).
Liu, B. et al. Resveratrol rescues SIRT1-dependent adult stem cell decline and alleviates progeroid features in laminopathy-based Progeria. Cell. Metab. 16, 738–750. https://doi.org/10.1016/j.cmet.2012.11.007 (2012).
Navarro, S., Reddy, R., Lee, J., Warburton, D. & Driscoll, B. Inhaled Resveratrol treatments slow ageing-related degenerative changes in mouse lung. Thorax 72, 451–459. https://doi.org/10.1136/thoraxjnl-2016-208964 (2017).
Bordone, L. et al. SIRT1 Transgenic mice show phenotypes resembling calorie restriction. Aging Cell. 6, 759–767. https://doi.org/10.1111/j.1474-9726.2007.00335.x (2007).
Pfluger, P. T., Herranz, D., Velasco-Miguel, S., Serrano, M. & Tschop, M. H. Sirt1 protects against high-fat diet-induced metabolic damage. Proc. Natl. Acad. Sci. U S A. 105, 9793–9798. https://doi.org/10.1073/pnas.0802917105 (2008).
Camacho, D. K., Go, C. C., Chaqour, B., Shindler, K. S. & Ross, A. G. Emerging gene therapy technologies for retinal ganglion cell neuroprotection. J. Neuroophthalmol. 43, 330–340. https://doi.org/10.1097/WNO.0000000000001955 (2023).
Zuo, L. et al. SIRT1 promotes RGC survival and delays loss of function following optic nerve crush. Invest. Ophthalmol. Vis. Sci. 54, 5097–5102. https://doi.org/10.1167/iovs.13-12157 (2013).
McDougald, D. S., Dine, K. E., Zezulin, A. U., Bennett, J. & Shindler, K. S. SIRT1 and NRF2 gene transfer mediate distinct neuroprotective effects upon retinal ganglion cell survival and function in experimental optic neuritis. Invest. Ophthalmol. Vis. Sci. 59, 1212–1220. https://doi.org/10.1167/iovs.17-22972 (2018).
Ross, A. G. et al. Selective upregulation of SIRT1 expression in retinal ganglion cells by AAV-Mediated gene delivery increases neuronal cell survival and alleviates axon demyelination associated with optic neuritis. Biomolecules 12 https://doi.org/10.3390/biom12060830 (2022).
Yue, J. et al. Cell-Specific expression of human SIRT1 by gene therapy reduces retinal ganglion cell loss induced by elevated intraocular pressure. Neurotherapeutics https://doi.org/10.1007/s13311-023-01364-6 (2023).
Khan, R. S., Dine, K., Das Sarma, J. & Shindler, K. S. SIRT1 activating compounds reduce oxidative stress mediated neuronal loss in viral induced CNS demyelinating disease. Acta Neuropathol. Commun. 2 https://doi.org/10.1186/2051-5960-2-3 (2014).
Fonseca-Kelly, Z. et al. Resveratrol neuroprotection in a chronic mouse model of multiple sclerosis. Front. Neurol. 3, 84. https://doi.org/10.3389/fneur.2012.00084 (2012).
Ballerini, C. Experimental autoimmune encephalomyelitis. Methods Mol. Biol. 2285, 375–384. https://doi.org/10.1007/978-1-0716-1311-5_27 (2021).
Quinn, T. A., Dutt, M. & Shindler, K. S. Optic neuritis and retinal ganglion cell loss in a chronic murine model of multiple sclerosis. Front. Neurol. 2, 50. https://doi.org/10.3389/fneur.2011.00050 (2011).
Redler, Y. & Levy, M. Rodent models of optic neuritis. Front. Neurol. 11, 580951. https://doi.org/10.3389/fneur.2020.580951 (2020).
Nimmagadda, V. K. et al. Overexpression of SIRT1 protein in neurons protects against experimental autoimmune encephalomyelitis through activation of multiple SIRT1 targets. J. Immunol. 190, 4595–4607. https://doi.org/10.4049/jimmunol.1202584 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U S A. 102, 15545–15550. https://doi.org/10.1073/pnas.0506580102 (2005).
Cho, K. Emerging roles of complement protein C1q in neurodegeneration. Aging Dis. 10, 652–663. https://doi.org/10.14336/AD.2019.0118 (2019).
Sato, K. et al. Ecel1 knockdown with an AAV2-Mediated CRISPR/Cas9 system promotes optic nerve Damage-Induced RGC death in the mouse retina. Invest. Ophthalmol. Vis. Sci. 59, 3943–3951. https://doi.org/10.1167/iovs.18-23784 (2018).
Luo, W. et al. Protein phosphatase 1 regulatory subunit 1A regulates cell cycle progression in ewing sarcoma. Oncotarget 11, 1691–1704. https://doi.org/10.18632/oncotarget.27571 (2020).
Jain, V., Ravindran, E. & Dhingra, N. K. Differential expression of Brn3 transcription factors in intrinsically photosensitive retinal ganglion cells in mouse. J. Comp. Neurol. 520, 742–755. https://doi.org/10.1002/cne.22765 (2012).
Sadun, A. A. Mitochondrial optic neuropathies. J. Neurol. Neurosurg. Psychiatry. 72, 423–425. https://doi.org/10.1136/jnnp.72.4.423 (2002).
Ramadori, G. et al. Brain SIRT1: anatomical distribution and regulation by energy availability. J. Neurosci. 28, 9989–9996. https://doi.org/10.1523/JNEUROSCI.3257-08.2008 (2008).
Sakamoto, J., Miura, T., Shimamoto, K. & Horio, Y. Predominant expression of Sir2alpha, an NAD-dependent histone deacetylase, in the embryonic mouse heart and brain. FEBS Lett. 556, 281–286. https://doi.org/10.1016/s0014-5793(03)01444-3 (2004).
Guo, W. et al. Sirt1 overexpression in neurons promotes neurite outgrowth and cell survival through Inhibition of the mTOR signaling. J. Neurosci. Res. 89, 1723–1736. https://doi.org/10.1002/jnr.22725 (2011).
Fulco, M. et al. Glucose restriction inhibits skeletal myoblast differentiation by activating SIRT1 through AMPK-mediated regulation of Nampt. Dev. Cell. 14, 661–673. https://doi.org/10.1016/j.devcel.2008.02.004 (2008).
Li, J. et al. Neuroprotective effects of oxymatrine via triggering autophagy and inhibiting apoptosis following spinal cord injury in rats. Mol. Neurobiol. 60, 4450–4471. https://doi.org/10.1007/s12035-023-03364-1 (2023).
Alam, F. et al. Interplay between oxidative stress, SIRT1, reproductive and metabolic functions. Curr. Res. Physiol. 4, 119–124. https://doi.org/10.1016/j.crphys.2021.03.002 (2021).
Kojima, J., Dai, Y., Suzuki, T., Ono, M. & Nishi, H. Sirtuin 1 is a potential therapeutic candidate gene for fetal growth restriction via insulin-like 4. J. Matern Fetal Neonatal Med. 36, 2253486. https://doi.org/10.1080/14767058.2023.2253486 (2023).
Jobling, A. I., Fang, Z., Koleski, D. & Tymms, M. J. Expression of the ETS transcription factor ELF3 in the retinal pigment epithelium. Invest. Ophthalmol. Vis. Sci. 43, 3530–3537 (2002).
Sultan, T. et al. Clinical and molecular heterogeneity of VPS13D-related neurodevelopmental and movement disorders. Gene 899, 148119. https://doi.org/10.1016/j.gene.2023.148119 (2024).
Sedlackova, L., Kelly, G. & Korolchuk, V. I. The pROS of autophagy in neuronal health. J. Mol. Biol. 432, 2546–2559. https://doi.org/10.1016/j.jmb.2020.01.020 (2020).
Nguyen, T. N., Padman, B. S. & Lazarou, M. ATG4s: above and beyond the Atg8-family protein lipidation system. Autophagy 17, 2648–2650. https://doi.org/10.1080/15548627.2021.1953263 (2021).
Li, Z. et al. Loss of SLC25A46 causes neurodegeneration by affecting mitochondrial dynamics and energy production in mice. Hum. Mol. Genet. 26, 3776–3791. https://doi.org/10.1093/hmg/ddx262 (2017).
Soucy, J. R. et al. Retinal ganglion cell repopulation for vision restoration in optic neuropathy: a roadmap from the RReSTORe consortium. Mol. Neurodegener. 18, 64. https://doi.org/10.1186/s13024-023-00655-y (2023).
Chaqour, B. et al. AAV2 vector optimization for retinal ganglion cell-targeted delivery of therapeutic genes. Gene Ther. https://doi.org/10.1038/s41434-023-00436-8 (2024).
Bastakis, G. G., Ktena, N., Karagogeos, D. & Savvaki, M. Models and treatments for traumatic optic neuropathy and demyelinating optic neuritis. Dev. Neurobiol. 79, 819–836. https://doi.org/10.1002/dneu.22710 (2019).
Su, B. et al. Abnormal mitochondrial dynamics and neurodegenerative diseases. Biochim. Biophys. Acta. 1802, 135–142. https://doi.org/10.1016/j.bbadis.2009.09.013 (2010).
Witte, M. E. et al. Enhanced number and activity of mitochondria in multiple sclerosis lesions. J. Pathol. 219, 193–204. https://doi.org/10.1002/path.2582 (2009).
Brooks, C. D., Kodati, B., Stankowska, D. L. & Krishnamoorthy, R. R. Role of mitophagy in ocular neurodegeneration. Front. Neurosci. 17, 1299552. https://doi.org/10.3389/fnins.2023.1299552 (2023).
Tang, B. L. Sirt1 and the mitochondria. Mol. Cells. 39, 87–95. https://doi.org/10.14348/molcells.2016.2318 (2016).
Kang, M. H., You, S. Y., Hong, K. & Kim, J. H. RNA sequencing data of mouse 2-cell embryos treated with DMSO. Data Brief. 28, 105025. https://doi.org/10.1016/j.dib.2019.105025 (2020).
Chen, X. et al. Different immunological mechanisms between AQP4 antibody-positive and MOG antibody-positive optic neuritis based on RNA sequencing analysis of whole blood. Front. Immunol. 14, 1095966. https://doi.org/10.3389/fimmu.2023.1095966 (2023).
Kao, L. et al. Severe neurologic impairment in mice with targeted disruption of the electrogenic sodium bicarbonate cotransporter NBCe2 (Slc4a5 gene). J. Biol. Chem. 286, 32563–32574. https://doi.org/10.1074/jbc.M111.249961 (2011).
Clemencon, B., Babot, M. & Trezeguet, V. The mitochondrial ADP/ATP carrier (SLC25 family): pathological implications of its dysfunction. Mol. Aspects Med. 34, 485–493. https://doi.org/10.1016/j.mam.2012.05.006 (2013).
Frederick, R. L. & Shaw, J. M. Moving mitochondria: Establishing distribution of an essential organelle. Traffic 8, 1668–1675. https://doi.org/10.1111/j.1600-0854.2007.00644.x (2007).
Jung, Y., Kim, H., Min, S. H., Rhee, S. G. & Jeong, W. Dynein light chain LC8 negatively regulates NF-kappaB through the redox-dependent interaction with IkappaBalpha. J. Biol. Chem. 283, 23863–23871. https://doi.org/10.1074/jbc.M803072200 (2008).
Cazzaro, S., Zhao, X., Zhao, V. K., Kim, Y. K. & Woo, J. A. Slingshot homolog-1 amplifies mitochondrial abnormalities by distinctly impairing health and clearance of mitochondria. Hum. Mol. Genet. 32, 1660–1672. https://doi.org/10.1093/hmg/ddad006 (2023).
Luo, W., Pant, G., Bhavnasi, Y. K., Blanchard, S. G. Jr. & Brouwer, C. Pathview web: user friendly pathway visualization and data integration. Nucleic Acids Res. 45, W501–W508. https://doi.org/10.1093/nar/gkx372 (2017).
Kaneko, A., Kiryu-Seo, S., Matsumoto, S. & Kiyama, H. Damage-induced neuronal endopeptidase (DINE) enhances axonal regeneration potential of retinal ganglion cells after optic nerve injury. Cell. Death Dis. 8, e2847. https://doi.org/10.1038/cddis.2017.212 (2017).
Chiha, W. et al. Retinal genes are differentially expressed in areas of primary versus secondary degeneration following partial optic nerve injury. PLoS One. 13, e0192348. https://doi.org/10.1371/journal.pone.0192348 (2018).
Huang, Y. et al. Brachial plexus root avulsion Injury-Induced Endothelin-Converting Enzyme-Like 1 overexpression is associated with injured motor neurons survival. Mol. Neurobiol. https://doi.org/10.1007/s12035-023-03887-7 (2024).
Nakagomi, S., Suzuki, Y., Namikawa, K., Kiryu-Seo, S. & Kiyama, H. Expression of the activating transcription factor 3 prevents c-Jun N-terminal kinase-induced neuronal death by promoting heat shock protein 27 expression and Akt activation. J. Neurosci. 23, 5187–5196. https://doi.org/10.1523/JNEUROSCI.23-12-05187.2003 (2003).
Matsumoto, S., Kiryu-Seo, S. & Kiyama, H. Motor nerve arborization requires proteolytic domain of Damage-Induced neuronal endopeptidase (DINE) during development. J. Neurosci. 36, 4744–4757. https://doi.org/10.1523/JNEUROSCI.3811-15.2016 (2016).
Tang, Z. et al. TOM40 Targets Atg2 to Mitochondria-Associated ER Membranes for Phagophore Expansion. Cell Rep 28, 1744–1757 e1745, (2019). https://doi.org/10.1016/j.celrep.2019.07.036
Corum, D. G., Tsichlis, P. N. & Muise-Helmericks, R. C. AKT3 controls mitochondrial biogenesis and autophagy via regulation of the major nuclear export protein CRM-1. FASEB J. 28, 395–407. https://doi.org/10.1096/fj.13-235382 (2014).
Kobuchi, H. et al. Mitochondrial localization of ABC transporter ABCG2 and its function in 5-aminolevulinic acid-mediated protoporphyrin IX accumulation. PLoS One. 7, e50082. https://doi.org/10.1371/journal.pone.0050082 (2012).
Lee, S. I. et al. The small fiber neuropathy NaV1.7 I228M mutation: impaired neurite integrity via bioenergetic and mitotoxic mechanisms, and protection by Dexpramipexole. J. Neurophysiol. 123, 645–657. https://doi.org/10.1152/jn.00360.2019 (2020).
He, X. et al. Bi-allelic Loss-of-function variants in CFAP58 cause flagellar axoneme and mitochondrial sheath defects and asthenoteratozoospermia in humans and mice. Am. J. Hum. Genet. 107, 514–526. https://doi.org/10.1016/j.ajhg.2020.07.010 (2020).
Kakegawa, W. et al. Anterograde C1ql1 signaling is required in order to determine and maintain a single-winner climbing fiber in the mouse cerebellum. Neuron 85, 316–329. https://doi.org/10.1016/j.neuron.2014.12.020 (2015).
Tian, J. et al. Intrathecal injection of SIRT1-modified human mesenchymal stem cells alleviates neuropathic pain in rat. J. Mol. Neurosci. 71, 972–980. https://doi.org/10.1007/s12031-020-01717-2 (2021).
Rasouri, S., Lagouge, M. & Auwerx, J. [SIRT1/PGC-1: a neuroprotective axis?]. Med. Sci. (Paris). 23, 840–844. https://doi.org/10.1051/medsci/20072310840 (2007).
Yeung, F. et al. Modulation of NF-kappaB-dependent transcription and cell survival by the SIRT1 deacetylase. EMBO J. 23, 2369–2380. https://doi.org/10.1038/sj.emboj.7600244 (2004).
Parker, J. A. et al. Integration of beta-catenin, Sirtuin, and FOXO signaling protects from mutant Huntingtin toxicity. J. Neurosci. 32, 12630–12640. https://doi.org/10.1523/JNEUROSCI.0277-12.2012 (2012).
Andrews, Z. B. et al. Uncoupling protein-2 is critical for nigral dopamine cell survival in a mouse model of Parkinson’s disease. J. Neurosci. 25, 184–191. https://doi.org/10.1523/JNEUROSCI.4269-04.2005 (2005).
Sarmah, S., Hawkins, M. R., Manikandan, P., Farrell, M. & Marrs, J. A. Elf3 deficiency during zebrafish development alters extracellular matrix organization and disrupts tissue morphogenesis. PLoS One. 17, e0276255. https://doi.org/10.1371/journal.pone.0276255 (2022).
Khan, R. S. et al. SIRT1 activating compounds reduce oxidative stress and prevent cell death in neuronal cells. Front. Cell. Neurosci. 6, 63. https://doi.org/10.3389/fncel.2012.00063 (2012).
Andreux, P. A., Houtkooper, R. H. & Auwerx, J. Pharmacological approaches to restore mitochondrial function. Nat. Rev. Drug Discov. 12, 465–483. https://doi.org/10.1038/nrd4023 (2013).
Chaffiol, A. et al. A new promoter allows optogenetic vision restoration with enhanced sensitivity in macaque retina. Mol. Ther. 25, 2546–2560. https://doi.org/10.1016/j.ymthe.2017.07.011 (2017).
Wang, Q. et al. A novel method to efficiently highlight nonlinearly expressed genes. Front. Genet. 10, 1410. https://doi.org/10.3389/fgene.2019.01410 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Acknowledgements
The authors thank the CAROT research vector core for production of AAV vectors.
Author information
Authors and Affiliations
Contributions
B.C.: Planning the project, supervision, execution, writing and editing manuscript; J.B.R, MM, and K.E.D.: experimental investigation and formal analysis of data; A.G.R and K.S.S.: planning, supervision, editing manuscript and funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chaqour, B., Rossman, J.B., Meng, M. et al. SIRT1-based therapy targets a gene program involved in mitochondrial turnover in a model of retinal neurodegeneration. Sci Rep 15, 13585 (2025). https://doi.org/10.1038/s41598-025-97456-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-97456-8