Abstract
TRIM56 plays a role in tumor development through the ubiquitination of several key substrate molecules. However, its relationship with tumor prognosis and immune infiltration remains unclear. The expression and localization of TRIM56 were analyzed from TCGA_GTEx, TCGA and HPA database. The effects of TRIM56 on the proliferation and migration of lung cancer cells A549 were evaluated by CCK-8 and wound healing assays. Correlations between TRIM56 expression and survival in patients were analyzed using the Kaplan-Meier Plotter and a nomogram model. Additionally, the relationship between TRIM56 and immune cell infiltration in tumors was explored via TIMER 2.0. Functional interactions and associated proteins of TRIM56 were examined using GEPIA 2.0 and the STING database. The signaling pathways influenced by TRIM56 were identified through GO and KEGG analyses. TRIM56 expression showed significant variation across 11 different tumor types when compared to normal tissues, with some tumors displaying high expression and others showing the opposite. TRIM56 inhibited the proliferation and migration of A549 cells. High TRIM56 expression was associated with shorter overall survival (OS) in patients with COAD, GBM, and LGG, but with longer OS in BLCA, KIRC, MESO, and SKCM. In BLCA and KIRC, high TRIM56 expression was closely linked to B cells, macrophages, and CD4(+) and CD8(+) T cell infiltration, contributing to a favorable prognosis. TRIM56 appears to affect tumor development through transcriptional regulatory complexes, transcriptional co-regulatory factor activity, and immune-related pathways.TRIM56 may play a critical role in tumor immunity and influence tumor prognosis. It holds potential as both a target for immunotherapy and a prognostic marker.
Similar content being viewed by others
Introduction
Globally, cancer has become a significant threat to public health, with both incidence and mortality rates rising each year1, among them, lung cancer, breast cancer, esophageal cancer, pancreatic cancer and liver cancer have high mortality rates worldwide. The rapid progression and poor prognosis of many cancers not only severely diminish patients’ quality of life but also place a heavy psychological and economic burden on society and governments. Consequently, there is an urgent need to discover new diagnostic and therapeutic approaches for cancer. In recent years, the use of biomarkers in certain cancers has shown promising results, leading to improved prognosis for affected patients2. However, many other tumors still lack effective biomarkers, particularly in the context of increasing emphasis on personalized and precision medicine. Therefore, identifying reliable biomarkers across various cancer types (pan-cancer) could significantly enhance treatment outcomes for a broader range of tumors.
TRIM proteins are involved in numerous physiological processes, including key roles in cell proliferation, apoptosis, DNA repair, signaling, innate immunity, and carcinogenesis3.Increasing evidence supports the role of the TRIM family in female genital tumors4, hepatocellular carcinoma5and lung cancer6. Ubiquitination enzymes primarily consist of ubiquitin-activating enzyme E1, ubiquitin-conjugating enzyme E2, and ubiquitin ligase E37. TRIM proteins form one of the largest subfamilies of RING (Really Interesting New Gene)-containing ubiquitin ligases. They belong to the highly conserved RING family of E3 ubiquitin ligases, also known as RBCC proteins—a large family of over 100 members found in all multicellular mammals, playing a pivotal role in ubiquitination8.Tripartite Motif-Containing Protein 56 (TRIM56), a member of the TRIM family, has been shown to play crucial roles in antiviral defense9and protection against nonalcoholic fatty liver disease10, however, its impact on pan-cancer prognosis and immune infiltration has not yet been reported. In this study, we found that TRIM56 was differentially expressed in most tumors. Survival analysis demonstrated that TRIM56 significantly impacted the prognosis of BLCA, COAD, GBM, KIRC, LGG, MESO, and SKCM. Given that alterations in the immune microenvironment can influence tumor prognosis, further analysis revealed that TRIM56 expression was associated with various immune cell populations. Additionally, enrichment analysis suggested that TRIM56 may contribute to tumor development through pathways such as lysine degradation, histone modification, peptidyl-lysine modification, transcriptional regulatory complexes, and transcriptional co-regulatory factor activity. Overall, TRIM56 plays diverse roles across multiple cancer types, influencing the prognosis of seven cancers (BLCA, COAD, GBM, KIRC, LGG, MESO, and SKCM), and modifying the immune microenvironment in 33 cancers from the TCGA database. Thus, our study identifies TRIM56 as a novel biomarker and a potential therapeutic target for cancer treatment.
Results
Expression of TRIM56 varies in different tumors
To investigate the expression of TRIM56 across pan-cancer types, we analyzed standardized TCGA_GTEx data from UCSC. The results revealed that TRIM56 was differentially expressed in most tumors. High TRIM56 expression was observed in normal tissues of BLCA (bladder urothelial carcinoma), BRCA (breast invasive carcinoma), CESC (cervical squamous carcinoma and adenocarcinoma), LUAD (lung adenocarcinoma), LUSC (lung squamous cell carcinoma), PRAD (prostate carcinoma), THCA (thyroid carcinoma), and UCEC (endometrial carcinoma). Conversely, high expression of TRIM56 was detected in tumor tissues of CHOL (cholangiocarcinoma), COAD (colon adenocarcinoma), ESCA (esophageal cancer), GBM (glioblastoma multiforme), HNSC (head and neck squamous cell carcinoma), KIRC (clear cell renal carcinoma), KIRP (renal papillary cell carcinoma), LIHC (hepatocellular carcinoma), PCPG (pheochromocytoma and paraganglioma), READ (rectal adenocarcinoma), SARC (sarcoma), STAD (gastric carcinoma), and THYM (thymic carcinoma). Additionally, TRIM56 was exclusively expressed in tumor tissues of MESO (mesothelioma) and UVM (uveal melanoma) (Fig. 1A), which aligned with the results from the TCGA dataset (Fig. 1B).
Differential expression of TRIM56 in ACC (adrenocortical carcinoma), DLBC (diffuse large B-cell lymphoma), KICH (kidney chromophobe), LAML (acute myeloid leukemia), LGG (low-grade glioma), OV (ovarian serous cystadenocarcinoma), PAAD (pancreatic adenocarcinoma), SKCM (skin cutaneous melanoma), TGCT (testicular germ cell tumors), and UCS (uterine carcinosarcoma) yielded varied results between the TCGA_GTEx and TCGA datasets.
We also analyzed TRIM56 expression in 23 paired tumor samples from the TCGA, and the findings were generally consistent with the results from the TCGA_GTEx and TCGA datasets (Fig. 1C). Moreover, survival analysis indicated that TRIM56 affected the prognosis of BLCA, COAD, GBM, KIRC, LGG, MESO, and SKCM. To further explore TRIM56 protein expression, we examined its presence in normal and tumor tissues across various organs using the Human Protein Atlas (HPA). We extracted representative immunohistochemical (IHC) images from bladder, brain, kidney, and skin tissues (Fig. 1D).
The expression of TRIM56 in pan-cancer tissues. (A-B) mRNA expression of TRIM56 in 33 tumors in the TCGA and TCGA_GTEx sample. (C) mRNA expression of tumor samples from the TCGA and TCGA_GTEx datasets. (D) Proteins expression of TRIM56 in normal tissues(upper) and cancer(lower), I: BLCA, II: GBM, III: KIRC, IV: SKCM. (ns, p > 0.05;*p < 0.05;**p < 0.01;***p < 0.001).
TRIM56 inhibits the proliferation and migration of lung cancer cells and is associated with overall survival (OS)
Analysis of the TCGA database revealed that TRIM56 expression in lung cancer tissues was lower than in normal tissues. We further analyzed TRIM56 expression in the CCLE database, which also demonstrated lower expression levels of TRIM56 in A549 lung cancer cells (Supplementary Figure S1). Based on these findings, we obtained a plasmid for TRIM56 overexpression and transfected it into A549 cells (Fig. 2A). The CCK-8 assay results showed that overexpression of TRIM56 significantly inhibited the proliferation of lung cancer cells (Fig. 2B). Additionally, the wound healing assay indicated that TRIM56 overexpression reduced the migration ability of lung cancer cells (Fig. 2C). Using the KM Plotter database, we analyzed TRIM56 expression in lung cancer and found that high TRIM56 levels were significantly associated with improved overall survival (p < 0.05) (Fig. 2D).
Relationship between TRIM56 and lung cancer. (A)The efficiency of transfection of TRIM56 in A549 cells was detected by Western blot. (B) The effect of TRIM56 on the growth of A549 cells was examined by CCK 8 at 24 h, 48 h, 72 h, 96 h and 120 h. (C) The migratory of A549 cells after up-regulation of TRIM56 was determined by wound healing assay.(**p < 0.01) (D) Survival analysis graph of TRIM56 and lung cancer patients.(p < 0.01).
Relationship between TRIM56 expression and prognosis in patients with pan-cancer
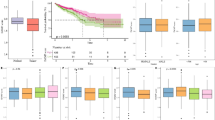
To investigate the prognostic value of TRIM56 across pan-cancer, we performed Kaplan-Meier survival analysis to evaluate the relationship between TRIM56 expression and clinical outcomes. First, we examined the correlation between TRIM56 expression and overall survival (OS) in 33 cancer types (Fig. 3A). The results showed that abnormal TRIM56 expression was significantly associated with OS in BLCA, COAD, GBM, KIRC, LGG, MESO, and SKCM. Specifically, high TRIM56 expression was linked to shorter OS in COAD, GBM, and LGG, while in BLCA, KIRC, MESO, and SKCM, high TRIM56 expression indicated longer OS (Fig. 3B-H).
Relationship between TRIM56 expression and OS in pan-cancer patients. (A) Forest plot showing the effect of TRIM56 expression on OS in pan-cancer. (B-H) Survival curves of TRIM56 expression on OS in patients with BLCA, COAD, GBM, KIRC, LGG, MESO and SKCM.
Next, we analyzed the relationship between TRIM56 expression and disease-specific survival (DSS) (Supplementary Fig. 2 A), which revealed associations in GBM, KIRC, and LGG. High TRIM56 expression was linked to better DSS in KIRC, while in GBM and LGG, higher TRIM56 expression was associated with poorer DSS (Supplementary Fig. 2B-D). Additionally, we explored the correlation between TRIM56 expression and progression-free interval (PFI) (Supplementary Fig. 3 A). The analysis showed that TRIM56 expression was associated with PFI in GBM, KIRC, LGG, and SKCM. High TRIM56 expression indicated better PFI in KIRC and SKCM, but was linked to poorer PFI in GBM and LGG (Supplementary Fig. 3B-E). Furthermore, ROC curves were generated for six tumors where TRIM56 expression correlated with prognosis, illustrating the diagnostic potential of TRIM56 in these cancers (Supplementary Fig. 4 A-E).
Correlation of TRIM56 expression with clinicopathologic features
Among the 33 tumors in the TCGA database, the prognosis of 7 tumors—BLCA, COAD, GBM, KIRC, LGG, MESO, and SKCM—was associated with TRIM56 expression. We further investigated the relationship between TRIM56 expression and clinicopathological features in these seven tumors. The results indicated that TRIM56 expression was correlated with age in LGG, BLCA, and COAD (Supplementary Fig. 5 A-C). Additionally, TRIM56 expression was associated with tumor size and invasion (T stage) in KIRC, BLCA, and SKCM (Supplementary Fig. 5D-F). Furthermore, TRIM56 expression was correlated with body weight in SKCM (Supplementary Fig. 5G).
Construction and evaluation of nomogram models for colon adenocarcinoma and renal clear cell carcinoma
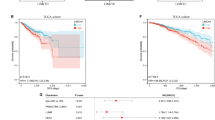
To further explore the impact of TRIM56 expression on tumor prognosis, we conducted univariate Cox regression analyses for overall survival (OS) in seven tumors where TRIM56 influences prognosis (Supplementary Tables S1-S6). Based on univariate Cox regression results from tumors with sample sizes exceeding 500, nomogram models were constructed for COAD and KIRC to validate their prognostic values. Calibration curves were used to assess the predictive accuracy of these nomogram models at 1, 3, and 5 years. The results indicated that TRIM56 significantly contributed to prognosis within the nomogram model, demonstrating strong predictive ability for OS in both COAD (Fig. 4A) and KIRC (Fig. 4C). The calibrated survival prediction curves at 1, 3, and 5 years confirmed that the nomogram model had high predictive accuracy for OS (Fig. 4B, D).
Nomogram models of COAD and KIRC were built and evaluated. (A) A nomogram model containing the expression of TRIM56 in COAD was developed. (B) The nomogram model of 1-, 3- and 5-year COAD was evaluated using calibration curves.(C) The nomogram model containing the expression of TRIM56 in KIRC was constructed. (D) Evaluation of the prediction accuracy of the nomogram model in KIRC using 1-, 3- and 5-year calibration curves.
Relationship between TRIM56 expression and tumor immune microenvironment
The immune microenvironment plays a critical role in tumor development. To explore the relationship between TRIM56 and the immune microenvironment across pan-cancer, we conducted a correlation analysis between TRIM56 expression and immune cell populations using the GEPIA2 database. Heatmaps were generated to illustrate the correlation between TRIM56 expression and B cells (Fig. 5A), macrophages (Fig. 5B), CD4(+) T cells (Fig. 5C), and CD8(+) T cells (Fig. 5D). The results indicated that TRIM56 expression in pan-cancer was generally positively correlated with the high expression of various immune cells, particularly in CD4(+) T cells (Th1 XCELL), CD8(+) T cells (naive_XCELL), and M2 macrophages (M2_TIDE), which showed relatively low TRIM56 expression in pan-cancer overall.
Correlation of TRIM56 expression with immune cell infiltration. (A-D) Heatmap of TRIM56 expression in the TIMER2 database correlating with B cells, macrophages, CD4(+) T cells and CD8(+) T cells.
Effect of TRIM56 expression combined with immune infiltration on overall survival rate
To evaluate the impact of immune cell infiltration on tumor prognosis, we selected four types of immune cells and divided them into high and low expression groups, then generated Kaplan-Meier (KM) survival curves for BLCA and KIRC. The results indicated that high expression of TRIM56 in B cell, macrophage, CD4(+) T cell, and CD8(+) T cell infiltration was associated with a favorable prognosis, consistent with the previous KM curve analysis without considering immune infiltration. Notably, the survival curves differed based on the levels of immune cell enrichment. In BLCA patient samples, the survival was higher when B cells, CD8(+) T cells, and macrophages were enriched compared to when they were reduced, whereas CD4(+) T cells showed the opposite trend. In KIRC patient samples, survival was higher when all four immune cell types were reduced in infiltration, compared to when they were enriched. These findings suggest that the role of immune cells in prognosis may depend on the expression levels of TRIM56 (Supplementary Fig. 6).
Building on our previous experimental findings that TRIM56 inhibits the proliferation and migration of lung cancer cells and that high TRIM56 levels are significantly linked to improved overall survival, we utilized bioinformatics methods to analyze the relationship between TRIM56 and macrophage polarization. The results showed that TRIM56 is positively correlated with M1 macrophage polarization and negatively correlated with M2 polarization. Notably, high expression of CD80, a marker of M1 - polarized macrophages, is associated with a poorer survival prognosis for lung cancer patients, while high expression of CD163, a marker of M2 - polarized macrophages, is associated with a better survival prognosis (Supplementary Figure S7).
Functional enrichment analysis of TRIM56-related genes and protein interaction analysis
To further elucidate the biological functions of TRIM56 in tumors, we identified the 100 genes most closely related to TRIM56 from the GEPIA2 database (Supplementary Table S7). Gene Ontology (GO) analysis (Fig. 6A) suggested that TRIM56-related genes may be involved in biological processes such as “histone modifications,” “peptidyl-lysine modification,” “histone methylation,” and “peptidyl-lysine trimethylation.” These genes are also associated with cellular components including “transcriptional regulatory complexes,” “nuclear speckles,” “transcriptional repressor complexes,” and the “INO80-type complex.” In terms of molecular functions, these genes are linked to “transcriptional co-regulatory factor activity,” “histidine-rich calcium-binding protein,” “histone methyltransferase activity,” and “histone-lysine N-methyltransferase activity.” KEGG pathway analysis11,12(Fig. 6B) indicated that TRIM56-related genes may be associated with pathways such as “lysine degradation,” “hepatitis B,” “TGF-β signaling pathway,” and “Notch signaling pathway.” Additionally, a PPI (protein-protein interaction) network was generated for the 100 TRIM56-related genes using the STRING database (Supplementary Figure S8).
To further investigate the function of TRIM56, we performed Gene Set Enrichment Analysis (GSEA) based on differential TRIM56 expression to elucidate its biological roles in the seven tumors where TRIM56 expression was associated with prognosis: BLCA, COAD, GBM, KIRC, LGG, MESO, and SKCM (Fig. 6C-I).
Functional enrichment analysis of TRIM56-related genes. (A) GO enrichment analysis based on 100 TRIM56-related genes, including BP, CC and MF.(B) KEGG pathway analysis based on 100 TRIM56-related genes. (C-I) Differential expression analysis of GSEA based on BLCA, COAD, GBM, KIRC, LGG, MESO and SKCM, respectively.
Discussion
Cancer continues to spread in the 21 st century, with an estimated 28.4 million cancer cases projected worldwide by 2040, representing a 47% increase from 202013. While recent research has identified effective biomarkers for certain tumors, the limitations of current therapeutic targets, the toxic side effects of drugs, and the body’s resistance mechanisms14, contribute to a poor prognosis for many cancers. As a result, identifying effective biomarkers across a wider range of tumors and developing treatments for cancers lacking viable targets have become primary focuses for current researchers.
Recent studies suggest that the E3 ubiquitin ligase TRIM56 may be involved in the development of breast cancer15, glioblastoma16and lung adenocarcinoma17. We also confirmed this in our experiments with A549 cells, whereas current lung cancer analyses mainly reflect p53 wild-type contexts, with p53-mutant scenarios remaining unexplored. Given the limited research on TRIM56, we explored its differential expression across pan-cancer for the first time to investigate its potential role in various cancers. We analyzed TRIM56 expression in normal and tumor tissues from different organs using TCGA_GTEx samples, TCGA samples, and TCGA paired samples. The results revealed that TRIM56 was highly expressed in some tumors compared to normal tissues, while showing lower expression in others.
After initially clarifying TRIM56 expression across various tumors, we further analyzed its relationship with prognosis. Interestingly, in GBM, LGG, and COAD, low TRIM56 expression was associated with a better prognosis compared to high expression, suggesting a pro-carcinogenic role. In contrast, TRIM56 played an anticancer role in BLCA, KIRC, MESO, and SKCM, where higher expression correlated with improved prognosis. Previous studies support these findings, showing that TRIM56 promotes the malignant progression of glioblastoma by stabilizing the cIAP1 protein16, while in hepatocellular carcinoma, TRIM56 inhibits malignant development by targeting RBM24 and inactivating the Wnt signaling pathway18. Additionally, TRIM56 has been reported to suppress multiple myeloma progression by activating the TLR3/TRIF signaling pathway19.
Clinicopathologic features are key indicators of cancer prognosis, yet there are no reports on the clinicopathologic characteristics of TRIM56 across pan-cancer. In our study, we found that TRIM56 expression was correlated with T stage in BLCA, KIRC, and SKCM, with age in BLCA, COAD, and LGG, and with body weight in SKCM. Based on these findings, we hypothesize that TRIM56 may be linked to tumor size and the degree of infiltration. Why does TRIM56 play such varied roles in different cancers? To explore this further, we conducted functional enrichment analysis in pan-cancer. The results suggest that TRIM56 is involved in histone and lysine-related biological processes. Previous studies have shown that histone-lysine N-methyltransferases are frequently overexpressed in various cancers, including hepatocellular carcinoma20. Histone methylation is also one of the most widely studied epigenetic markers in gastric cancer, with numerous histone methyltransferases and demethylases found to be altered in this disease21.
Since TRIM56 is associated with antiviral immunity22and and innate cellular immunity23, and cellular immunity plays a key role in tumor prognosis24,25, immune cell infiltration, which alters the tumor microenvironment, is a major factor influencing tumor progression26. Therefore, we analyzed the relationship between TRIM56 and immune cells across pan-cancer. We stratified TRIM56 expression into high and low groups and assessed the prognostic impact of immune cell infiltration, suggesting that immune cell function may be dependent on TRIM56 expression. Increased CD8 + T cell infiltration and cytolytic activity may represent an anti-tumor pathway driven by STING-dependent type I IFN signaling27. Additionally, CD4 + T cells must interact with CD8 + T cells via the same dendritic cell during the effector phase, forming a three-cell-type cluster (triad) to activate CD8 + T cell cytotoxicity and eliminate cancer cells28. Tumor-associated macrophages, the most abundant immune cells in the ovarian cancer microenvironment, exhibit heterogeneous subpopulations with either protumor or antitumor functions29. Tumor-associated macrophages exhibit polarization, with M2 macrophages promoting tumor progression and M1 macrophages suppressing it. Our findings reveal that TRIM56 negatively correlates with M2 polarization and positively correlates with M1 polarization, consistent with previous in vitro experiments. This aligns with survival analyses of M1 marker CD160 and M2 marker CD80 in lung cancer patients. Based on these findings, we speculate that TRIM56 may modulate the immune microenvironment and could be a potential target for immunotherapy in certain cancers, paving the way for the development of new targeted drugs that may benefit a large number of cancer patients.
Our study analyzed various aspects of pan-cancer, focusing on both its immune infiltration and prognosis. Despite our best efforts to make our findings accurate and reliable, some limitations are inevitable. On the one hand, the small sample size of some cancers or the insufficient sample size of the control group occasionally led to divergent or even ambiguous results in some analyses. On top of that, our study is mainly based on bioinformatics analysis, and the conclusions drawn need to be confirmed by further in vivo and in vitro experiments. In summary, we found that TRIM56 is prognostically relevant in a variety of tumors and acts by regulating the immune microenvironment. Mechanistically, it may affect tumorigenesis and progression through lysine degradation, histone modification, peptidyl-lysine modification, transcriptional regulatory complexes and transcriptional co-regulatory activity. However, the roles of TRIM56 vary in different tumors, further experimental validation is still required for its functions and mechanisms in specific tumors.
Materials and methods
Data collection and processing
[37] Expression profiling and clinical data for 33 tumor types were obtained from The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/). The TCGA_GTEx dataset, which includes TCGA tumor samples and normal samples from the University of California, Santa Cruz (UCSC) Xena platform (https://xenabrowser.net/datapages/), was used for analysis. Immunohistochemical images of normal and tumor tissues were sourced from the Human Protein Atlas (HPA, https://www.proteinatlas.org/). Additionally, the 100 genes most correlated with TRIM56 were identified using the Gene Expression Profiling Interaction Analysis database (GEPIA2, http://gepia2.cancer-pku.cn/#index) from the TCGA dataset. This study followed the published guidelines of TCGA and UCSC, and as a result, ethical approval and informed consent from patients were not required.
Expression analysis of TRIM56
The expression of TRIM56 mRNA in normal and tumor tissues was compared using TCGA_GTEx samples, TCGA samples, and paired TCGA samples. Additionally, TRIM56 protein levels in normal and tumor tissues were analyzed through the Human Protein Atlas (HPA, https://www.proteinatlas.org/), which provides detailed information on protein distribution in human tissues and cells. Based on survival analysis, which demonstrated the prognostic significance of TRIM56 in BLCA (bladder urothelial carcinoma), COAD (colon adenocarcinoma), GBM (glioblastoma multiforme), KIRC (clear cell renal carcinoma), LGG (low-grade glioma), MESO (mesothelioma), and SKCM (skin melanoma), we further downloaded immunohistochemical images of representative tumor tissues, along with corresponding normal tissues, from the HPA database. These included tissues from the bladder, colon, brain, kidney, mesothelium, and skin.
The mRNA expression matrix for tumor cell lines was obtained from the CCLE dataset (https://portals.broadinstitute.org/ccle). The analysis was performed using the R software (v4.0.3) with the ggplot2 package (v3.3.3).
Plasmid and cell culture
The TRIM56 expression plasmid (Addgene) was purchased from the MiaoLing Plasmid Platform (Wuhan, China). Transient transfections were carried out using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s protocol. A549 cells were obtained from Pricella Biotechnology (Wuhan, China) and cultured in Ham’s F-12 K medium supplemented with 10% fetal bovine serum.
Cell counting Kit-8 (CCK-8) assay
After transfecting A549 cells with the TRIM56 plasmid for 24 h, the cells were seeded into 96-well plates at a density of 5,000 cells per well. Cell viability was then assessed using the Cell Counting Kit-8 (MedChemExpress, USA) according to the manufacturer’s instructions.
Wound healing assay
After transfecting A549 cells with the TRIM56 plasmid for 24 h, the cells were seeded into 6-well plates without fetal bovine serum (FBS). Once the cells reached 100% confluency, straight scratches were made using yellow pipette tips. The wound distance was measured at the indicated time points and normalized to the initial time point. The cell migration rate (%) was calculated as: (scratch width at 0 h - wound width at 24 h)/scratch width at 0 h × 100%.
Western blot
Cells were harvested and lysed using NP40 buffer. Proteins were separated by SDS polyacrylamide gel electrophoresis (SDS-PAGE) and then electrotransferred onto a PVDF membrane. The membranes were blocked with milk and incubated overnight at 4 °C with TRIM56 antibody (1:500, Zenbio, China) and β-actin antibody (1:2000, ABclonal, China). After three washes with PBS, the membranes were incubated with secondary antibodies for 1 h at room temperature. Protein bands were visualized using an ECL kit (Tanon, China) and imaged with the ChemiDoc™ Gel Imaging System (Bio-Rad).
Prognostic analysis
Kaplan-Meier analysis and log-rank tests were performed using the Kaplan-Meier Plotter website (https://kmplot.com/analysis) to assess the association of TRIM56 expression with overall survival (OS), progression-free interval (PFI), and disease-specific survival (DSS) in patients with pan-cancer from the TCGA dataset. Survival curves with p-values < 0.05 were generated. The diagnostic value of TRIM56 in pan-cancer was evaluated using ROC curves for the seven tumors where TRIM56 impacted prognosis. ROC curves were calculated using the R software package “pROC” (version 1.17.0.1) and plotted with “ggplot2” (v3.3.3). The Area Under the Curve (AUC), critical value, sensitivity, specificity, positive predictive value, negative predictive value, and Youden Index (YI) were calculated. AUC values closer to 1 indicate higher diagnostic accuracy: 0.5–0.7 suggests low accuracy, 0.7–0.9 indicates good accuracy, and 0.9 or higher represents high accuracy. The Youden Index (YI) reflects the effectiveness of a screening method in distinguishing true patients from non-patients, with a higher YI indicating a more accurate and reliable screening method.
Correlation analysis between TRIM56 expression and clinical features
In tumors where TRIM56 influences prognosis, the correlation between TRIM56 expression and key clinical parameters, such as T-stage, age, and weight in TNM staging, was analyzed. The main variables were grouped, and appropriate statistical methods were selected based on the data format characteristics using the stats and car packages. Data visualization was performed with the ggplot2 package (v3.3.3).
Nomogram modeling and evaluation
A one-way Cox regression analysis was performed on overall survival (OS) for tumors where TRIM56 affects prognosis, including OS, progression-free interval (PFI), and disease-specific survival (DSS). The survival package was used for proportional hazard hypothesis testing and Cox regression analysis. For tumors with p < 0.05 and a sample size greater than 500, a nomogram model was constructed. The nomogram assigns scores based on the magnitude of the regression coefficients of the independent variables, with each value level receiving a score. A total score is then calculated for each patient, and a conversion function between the score and the probability of occurrence is used to predict the likelihood of the outcome, providing an effective and convenient method for predicting individual patient OS. Proportional hazard hypothesis testing was conducted using the survival package, and Cox regression results were visualized with the rms package for column line plots. Calibration curves were generated to assess the accuracy of the 1-, 3-, and 5-year nomogram predictions. The calibration analysis was performed using the survival and rms packages. The closer the actual curve is to the ideal curve, the better the calibration, indicating that the model’s predictions closely match actual outcomes.
Immune infiltration analysis
The correlation between TRIM56 expression and various immune cells, including B cells, macrophages, CD4(+) T cells, and CD8(+) T cells, was analyzed across all TCGA tumors using Tumor Immune Estimation Resource 2.0 (TIMER2.0, http://timer.cistrome.org/). Immune cell infiltration and the immune microenvironment were estimated using multiple algorithms such as TIMER, CIBERSORT, quanTIseq, xCell, MCP-counter, and EPIC. Additionally, we investigated the impact of immune cell infiltration on overall survival (OS) after stratifying tumors by TRIM56 expression.
Functional enrichment analysis and protein-protein interaction network analysis
The 100 genes with expression patterns most similar to TRIM56 were obtained from the GEPIA2 database (GEPIA2, http://gepia2.cancer-pku.cn/#index). Based on these TRIM56-related genes, Gene Ontology (GO) analysis—focusing on biological processes (BP), cellular components (CC), and molecular functions (MF)—and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis were performed to further investigate the potential functions of TRIM56. Additionally, the 100 TRIM56-related genes were used to construct a protein-protein interaction (PPI) network using the STRING database (https://cn.string-db.org/), with a minimum interaction threshold of 0.4.
Differential expression analysis and gene set enrichment analysis
Data for the corresponding molecules were extracted from the TCGA database and divided into high and low expression groups based on TRIM56 expression. Variance analysis of the original counts matrix was performed using the DESeq2 package, following standard procedures, and normalization was achieved using the Variance Stabilizing Transformations (VST) method provided by DESeq2. Subsequently, based on the results of differential expression analysis of TRIM56 in various tumors, molecule IDs in the input data were converted using the MSigDB Collections gene set database (https://www.gsea-msigdb.org/gsea/msigdb/index.jsp). Gene set enrichment analysis (GSEA) was then conducted using the clusterProfiler R package.
Statistical analysis
All results were analyzed using GraphPad Prism 9. Experimental data were presented as means ± standard deviations (SDs), and unpaired Student’s t-tests were used to compare continuous variables between groups. The Wilcoxon rank sum test was applied to compare differences between two groups, while the Spearman rank correlation test was used to calculate correlations between variables. Univariate and multivariate Cox proportional hazard regressions were performed to identify factors affecting prognosis. Survival analysis was conducted using Kaplan-Meier curves and log-rank tests. Statistical analysis was performed using R software (version 4.0.2), with p < 0.05 considered statistically significant. (*p < 0.05, **p < 0. 01, ***p < 0.001, ****p < 0.0001).
Data availability
Data related to the current study are available from the corresponding author on reasonable request.
Change history
09 May 2025
The original online version of this Article was revised: In the original version of this Article Yunpeng Cao, Lingwei Kong and Yulu Zhai were omitted as equally contributing authors. The original Article has been corrected.
Abbreviations
- TCGA:
-
The Cancer Genome Atlas
- GEO:
-
Gene Expression Omnibus
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Cancer J. Clin. 74, 229–263. https://doi.org/10.3322/caac.21834 (2024).
Cao, W., Chen, H. D., Yu, Y. W., Li, N. & Chen, W. Q. Changing profiles of cancer burden worldwide and in China: a secondary analysis of the global cancer statistics 2020. Chin. Med. J. 134, 783–791. https://doi.org/10.1097/cm9.0000000000001474 (2021).
Huang, N. et al. TRIM family contribute to tumorigenesis, cancer development, and drug resistance. Experimental Hematol. Oncol. 11 https://doi.org/10.1186/s40164-022-00322-w (2022).
Liu, T. et al. Role of the tripartite motif (TRIM) family in female genital neoplasms. Pathol. Res. Pract. 250, 154811. https://doi.org/10.1016/j.prp.2023.154811 (2023).
Lu, K. et al. TRIM proteins in hepatocellular carcinoma. J. Biomed. Sci. 29, 69. https://doi.org/10.1186/s12929-022-00854-7 (2022).
Liang, M. et al. E3 ligase TRIM28 promotes anti-PD-1 resistance in non-small cell lung cancer by enhancing the recruitment of myeloid-derived suppressor cells. J. Experimental Clin. cancer Research: CR. 42, 275. https://doi.org/10.1186/s13046-023-02862-3 (2023).
Gu, X. et al. The midnolin-proteasome pathway catches proteins for ubiquitination-independent degradation. Sci. (New York N Y). 381, eadh5021. https://doi.org/10.1126/science.adh5021 (2023).
Wang, L. & Ning, S. TRIMming type I Interferon-Mediated innate immune response in antiviral and antitumor defense. Viruses 13 https://doi.org/10.3390/v13020279 (2021).
Fu, L., Zhou, X., Jiao, Q. & Chen, X. The functions of TRIM56 in antiviral innate immunity and tumorigenesis. Int. J. Mol. Sci. 24 https://doi.org/10.3390/ijms24055046 (2023).
Xu, S. et al. TRIM56 protects against nonalcoholic fatty liver disease by promoting the degradation of fatty acid synthase. J. Clin. Investig. 134 https://doi.org/10.1172/jci166149 (2024).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Ge, W. et al. Potential biomarkers: identifying powerful tumor specific T cells in adoptive cellular therapy. Front. Immunol. 13 https://doi.org/10.3389/fimmu.2022.1003626 (2022).
Xue, M. et al. Regulation of Estrogen signaling and breast cancer proliferation by an ubiquitin ligase TRIM56. Oncogenesis 8, 30. https://doi.org/10.1038/s41389-019-0139-x (2019).
Yang, X. et al. TRIM56 promotes malignant progression of glioblastoma by stabilizing cIAP1 protein. J. Experimental Clin. cancer Research: CR. 41, 336. https://doi.org/10.1186/s13046-022-02534-8 (2022).
Gao, J. et al. Exosomal circZNF451 restrains anti-PD1 treatment in lung adenocarcinoma via polarizing macrophages by complexing with TRIM56 and FXR1. J. Experimental Clin. cancer Research: CR. 41, 295. https://doi.org/10.1186/s13046-022-02505-z (2022).
Yang, Y., Mao, F. F., Guo, L. & Guo, W. X. TRIM56 suppresses the malignant development of hepatocellular carcinoma via targeting RBM24 and inactivating the Wnt signaling. Eur. Rev. Med. Pharmacol. Sci. 25, 722–730. https://doi.org/10.26355/eurrev_202101_24633 (2021).
Chen, Y. et al. TRIM56 suppresses multiple myeloma progression by activating TLR3/TRIF signaling. Yonsei Med. J. 59, 43–50. https://doi.org/10.3349/ymj.2018.59.1.43 (2018).
Thng, D. K. H. et al. Histone-lysine N-methyltransferase EHMT2 (G9a) Inhibition mitigates tumorigenicity in Myc-driven liver cancer. Mol. Oncol. 17, 2275–2294. https://doi.org/10.1002/1878-0261.13417 (2023).
De Marco, K., Sanese, P., Simone, C. & Grossi, V. Histone and DNA methylation as epigenetic regulators of DNA damage repair in gastric cancer and emerging therapeutic opportunities. Cancers 15 https://doi.org/10.3390/cancers15204976 (2023).
Yang, L. et al. UBXN3B positively regulates STING-mediated antiviral immune responses. Nat. Commun. 9, 2329. https://doi.org/10.1038/s41467-018-04759-8 (2018).
Yang, D. et al. The E3 ligase TRIM56 is a host restriction factor of Zika virus and depends on its RNA-binding activity but not MiRNA regulation, for antiviral function. PLoS Negl. Trop. Dis. 13, e0007537. https://doi.org/10.1371/journal.pntd.0007537 (2019).
Liu, T. et al. Comprehensive Pan-Cancer analysis of KIF18A as a marker for prognosis and immunity. Biomolecules 13 https://doi.org/10.3390/biom13020326 (2023).
Laumont, C. M., Banville, A. C., Gilardi, M., Hollern, D. P. & Nelson, B. H. Tumour-infiltrating B cells: immunological mechanisms, clinical impact and therapeutic opportunities. Nat. Rev. Cancer. 22, 414–430. https://doi.org/10.1038/s41568-022-00466-1 (2022).
Xia, X. et al. Reshaping the tumor immune microenvironment to improve CAR-T cell-based cancer immunotherapy. Mol. Cancer. 23, 175. https://doi.org/10.1186/s12943-024-02079-8 (2024).
Maxwell, M. B. et al. ARID1A suppresses R-loop-mediated STING-type I interferon pathway activation of anti-tumor immunity. Cell 187, 3390–3408e3319. https://doi.org/10.1016/j.cell.2024.04.025 (2024).
Espinosa-Carrasco, G. et al. Intratumoral immune triads are required for immunotherapy-mediated elimination of solid tumors. Cancer Cell. 42, 1202–1216e1208. https://doi.org/10.1016/j.ccell.2024.05.025 (2024).
Jazwinska, D. E., Kulawiec, D. G. & Zervantonakis, I. K. Cancer-mesothelial and cancer-macrophage interactions in the ovarian cancer microenvironment. Am. J. Physiol. Cell Physiol. 325, C721–c730. https://doi.org/10.1152/ajpcell.00461.2022 (2023).
Acknowledgements
We sincerely thank the TCGA, HPA, GEPIA2, CCLE, Kaplan-Meier, Timer2.0 and STRING projects for use of their data.
Funding
This study was supported by Chengde Medical University Discipline Construction Funds, Chengde Science and Technology Bureau(202204 A164), Research Start-up Fund for High-level Talents of Chengde Medical University(202409) and the Cultivating Programme Plan for Clinical Excellent Talent of Hebei in 2023 (ZF2023249).
Author information
Authors and Affiliations
Contributions
Conceptualization and Writing-original draft, Yunpeng Cao and Yulu Zhai; Validation, Yulu Zhai and Lingwei Kong; Formal analysis, Weiyan Hou; Funding acquisition, Peiyuan He, Lingwei Kong and Hairu Ji; Methodology, Jiayuan Wang; Data curation, Yulu Zhai, Yuxi Liu and Chengru Wang; Software, Wei Zhao; Project administration, Supervision, Writing-review & editing, Hairu Ji and Lingwei Kong.
Corresponding author
Ethics declarations
Consent for publication
All authors have approved the publication of this study.
Competing interests
The authors declare no competing interests.
Supplementary Information
Supplementary Material 1.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cao, Y., Kong, L., Zhai, Y. et al. Comprehensive analysis of TRIM56’s prognostic value and immune infiltration in Pan-Cancer. Sci Rep 15, 13673 (2025). https://doi.org/10.1038/s41598-025-97856-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-97856-w