Abstract
Hereditary hearing loss is highly heterogeneous. SLC12A2 is linked to autosomal dominant nonsyndromic hearing loss, DFNA78, with all the pathogenic variants affecting the exon 21. The gene encodes a cotransporter NKCC1 crucial for regulating intracellular osmotic pressure and producing endolymph in the cochlea. We generated two mouse strains with heterologous Slc12a2 variants in the splice site of the exon 21 (Em1: NM_009194.3:c.2912-2 A > G and Em2: c.2912-4_2913del). Slc12a2Em2/Em2 mice with complete skip of the exon 21 showed reduced endolymph on postnatal day 1 (P1), reduced stria vascularis (StV) and no auditory brainstem responses at 4 weeks. Reduced StV size was considered to be due to rebalance osmotic pressure, and upregulation of Cldn9 revealed by RNA-seq was considered as tissue response to repair the gaps from reduced cell sizes in the Slc12a2Em2/Em2 cochlea. Female Slc12a2Em2/+ mice also exhibited mild elevation of ABR thresholds in several sound frequencies. Slc12a2Em1/Em1 mice showed normal hearing, presumably due to sufficient cotransporter activity from the 9 bases shorter transcript by cryptic splicing. Minigene assays indicated that a single nucleotide difference between humans and mice at the 5’ end of the exon 21 affects exon 21 splicing. Slc12a2Em2 mouse is proposed as a model for studying DFNA78 pathology.
Similar content being viewed by others
Introduction
Genetic causes of hearing loss are highly heterogeneous; to date, more than 150 genes have been reported to be responsible for nonsyndromic hearing loss1. An analysis of four families identified SLC12A2 as a novel gene responsible for nonsyndromic autosomal dominant hearing loss, DFNA78 (MIM: 619081)2. Since then, several other DFNA78 families have been reported globally3,4,5,6, with all of these patients appearing to show congenital, severe to profound hearing loss. SLC12A2 encodes the Na+, K+, 2Cl− cotransporter NKCC1, which is expressed in almost all cell types throughout the body and has been known to regulate cell volume by balancing osmotic pressure7. In mammalian cochlea, SLC12A2 is predominantly expressed in the marginal cells of the stria vascularis (StV), a three-layered tissue in the lateral wall2,8,9,10 and subset of spiral ligament fibrocytes (SLF), another component of the lateral wall. The StV plays major roles in the production and homeostasis of K+- enriched endolymph that fills Scala Media (SM), which is essential for depolarization of sensory hair cells of the organ of Corti11,12. In contrast to SM, Scala Vestibuli (SV) and Scala Tympani (ST) are filled with perilymph which is connected to cerebrospinal fluid through cochlear aqueduct13. The primary endolymph without endocochlear potential (EP) at early postnatal stage is known to be replaced with secondary endolymph with high K+ content and elevated EP14,15which allows mouse pups to start responding to sound at approximately postnatal day 12 (P12)16. SM is surrounded by tight junction proteins such as Claudins that form a barrier inhibiting diffusion between the extracellular environment and epithelial or endothelial cells17. Among tight junction protein-encoding genes, CLDN9 has been known to be responsible for autosomal recessive nonsyndromic hearing loss DFNB116 (OMIM 619093)18. In addition to SLC12A2 and CLDN9, multiple deafness genes with diverse functional properties are also distributed to cells surrounding auditory epithelium, such as transcription factor GATA3 responsible for Hypoparathyroidism, Sensorineural Deafness, And Renal Dysplasia Syndrome (14255), anion transporter SLC26A4 responsible for Pendred Syndrome (274600) and DFNB4 (600791), and transmembrane serine protease TMPRSS3 responsible for DFNB8 (601072).
To date, all reported pathogenic variants of SLC12A2 associated with hearing loss have been heterozygous missense variants in exon 21 or its splice acceptor site. recombinant SLC12A2 proteins with heterologous missense variants or omission of exon 21 were found unable to incorporate Cl− in vitro2. However, whether the pathogenic mechanisms by which SLC12A2 variants cause hearing loss, is due solely to haploinsufficiency resulting from reduced ion transporter activity is unclear, because multiple loss-of-function variants of SLC12A2 have been detected in population databases such as gnomAD19. In addition, loss-of-function alleles of SLC12A2 have been associated with the autosomal recessive inheritance disorder Kilquist syndrome (OMIM 619080)20,21,22, but the parents of patients with Kilquist syndrome have not exhibited noticeable symptoms.
We have speculated that SLC12A2 protein without the exon 21-encoded region (Δex21) not only deteriorates cotransporter activity, but also mediates an unrecognized function2. The exon 21-encoded region has been proposed to mediate the trafficking of SLC12A2 protein to the basolateral membrane in polarized cells, based on the report that replacement of the partial cytoplasmic region, including exon 21, with the corresponding SLC12A1 region resulted in trafficking of the SLC12A2 to the apical membrane of MDCK cells23. However, this is inconclusive because of the contradictory result showing that SLC12A2Δex21 was retained in the basolateral membranes of MDCK cells24. Multiple molecules interacting with SLC12A2 have been reported25, but whether they bind to the exon 21-encoded region are unkonown.
While all the clinical evidence strongly suggests that SLC12A2 is responsible for hereditary nonsyndromic hearing loss, experimental animal models that harbor the Slc12a2 variant equivalent to the variant detected in patients are required to prove the gene-disease correlation and understand its molecular pathology. Based on the loss-of-function alleles of SLC12A2 associated with Kilquist syndrome, Sc12a2-knockout mice26,27,28 and mice deficient in Sc12a229, all of which exhibit hearing loss associated with severe reduction of endolymph production, are considered as inappropriate models for DFNA78. This study generated two mouse models by knocking in (KI) the splice variant at the acceptor site considered to be equivalent to the one found in a patient with DFNA78 (SLC12A2: NM_001046.2:c.2930‒2 A > G) which results in complete skip of exon 21 in vitro2. Hearing properties and morphological changes of the cochleae were assessed in these mice, as were the distinct characteristics of exon 21 splicing based on a one nucleotide difference between humans and mice.
Results
Generation of mouse strains with mutations knocked in at the splice acceptor site of Slc12a2
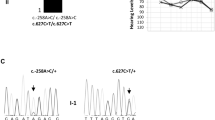
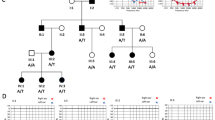
Knocking in variant into 330 fertilized eggs of FVB/N mice using the CRISPR/Cas9 system yielded two strains of mice: Slc12a2Em1, carrying a heterozygous variant at the splice acceptor site of exon 21 of Slc12a2 (NM_009194.3:c.2912‒2 A > G), and Slc12a2Em2, carrying a heterozygous six-base deletion of Slc12a2 (c.2912-4_2913del) (Fig. 1A).
Generation of Slc12a2 knock in mouse strains. (A) Nucleotides of wild-type (+), Em1 and Em2 alleles at the acceptor site of the exon 21 region of Slc12a2. (B) RT-PCR of cochlear and cerebellar samples of Slc12a2Em2 mice at age 4 weeks (4 W). Genotypes and the sizes of the transcripts are shown on the top and right. M; molecular weight marker. (C) ABR thresholds of Slc12a2Em2 mice at 4 W. Arrows indicate no response to the maximum sound. ***; p < 10− 6, **; p < 0.01, *; p < 0.05.
Testing of their genotypes at 2 weeks of age showed that approximately 19% of the pups (33 out of 172) resulting from the mating of Slc12a2Em2/+ mice were Slc12a2Em2/Em2 mice. RT-PCR (Fig. 1B) of RNA from the cochlea and cerebellum of these Slc12a2Em2/Em2 mice, followed by Sanger sequencing, showed the absence of the entire exon 21 (48 nucleotides) from Slc12a2 transcripts (Supplemental Fig. 1A, B). Quantitative RT-PCR (qRT-PCR) demonstrated that even though there was significant change in the composition of the exon21-including or skipped Slc12a2 transcripts, overall gene expression levels of Slc12a2 did not differ significantly between Slc12a2+/+ and Slc12a2Em2/Em2 cochleae (Supplemental Fig. 1C). At age 4 weeks (4 W), Slc12a2Em2/Em2 mice exhibited circling as well as head-bobbing behavior, typical of an inner ear disorder (Supplemental Movie 1), whereas Slc12a2Em2/+ mice behaved normally (Supplemental Movie 2). Auditory brainstem response (ABR) tests showed that Slc12a2Em2/Em2 mice did not respond to any of the frequencies of pure tone sounds tested (99, 106, 112 dB SPL at 8, 16, and 32 kHz, respectively). Sound threshold levels between Slc12a2Em2/+ mice (23.55 ± 7.21, 29.18 ± 12.48, 47.45 ± 9.88 dB SPL, respectively) and those of Slc12a2+/+ mice (16.50 ± 7.50, 16.83 ± 10.17, 34.50 ± 10.31 dB SPL, respectively) were similar at 8 kHz (p = 0.0821) but showed significant differences at 16 kHz and 32 kHz (p = 0.0060 and 0.030, Fig. 1C). Mild but significant elevation of ABR thresholds at 16 kHz and 32 kHz in Slc12a2Em2/+ mice appeared to be the phenotype in female but not in male (Supplemental Fig. 2A, B). However, due to variable ABR thresholds among individuals, significant differences between female and male Slc12a2Em2/+ mice were found only at 32 kHz (Supplemental Fig. 2C, D). Because no response to the measured sound frequencies observed in Slc12a2Em2/Em2 mice resembled severe to profound hearing loss in DFNA78 patients, we sought to characterize pathology of Slc12a2Em2/Em2 cochleae in this study. Since ABR threshold levels did not differ between male and female Slc12a2Em2/Em2 mice, both sexes were included in subsequent experiments.
Histopathology of Slc12a2 Em2/Em2 cochlea
The cochlear morphology of postnatal Slc12a2Em2 mice was assessed using paraffin sections. At postnatal day 1 (P1), the space of SM was readily visible in the cochleae of Slc12a2Em2/+ mice (Fig. 2A–C, Supplemental Fig. 3A), but was almost absent in the cochleae of Slc12a2Em2/Em2 mice (Fig. 2D–F, Supplemental Fig. 3B). This was shown as significantly reduced proportion of the areas of SM/SV at the basal turn of the Slc12a2Em2/Em2 cochleae (Supplemental Fig. 4A, C). These findings indicate that Slc12a2 with the exon 21-encoded region contributes to endolymph production before mouse starts hearing.
Histological study of Slc12a2Em2 cochlea. Hematoxylin-eosin staining of cochlear ducts of Slc12a2Em2/+ (A-C, G-I, M-O) and Slc12a2Em2/Em2 (D-F, J-L, P-R) mice at P1 (A-F), P12 (G-L) and 4 W (M-R). Arrows indicate the stria vascularis. Asterisc indicates partial loss of the lateral wall. Scale bars = 50 μm. (S-U) Estimated cell density in the eosinophilic StV areas at 4 W. N = 4. GER, greater epithelial ridge; RM, Reissner’s membrane; SM, scala media; StV, stria vascularis.
At P12, the space of SM was clearly visible in both Slc12a2Em2/+ and Slc12a2Em2/Em2 cochleae (Fig. 2G–L, Supplemental Fig. 3C, D). In the basal turn, the StV appeared to be smaller and curled up toward the direction of spiral prominence (Fig. 2L, Supplemental Fig. 3D). In the apical turn, the tectorial membrane was disoriented, but still in contact with the StV in the Slc12a2Em2/Em2 cochlea (Fig. 2J), suggesting that this turn is in the process of expansion of the SM. At 4 W, the smaller and thicker sizes of the StV in the apical and middle turns and the shrunken StV in the basal turns were evident in the Slc12a2Em2/Em2 cochleae (Fig. 2M–R, Supplemental Fig. 3E, F). Proportion of the SM/SV in the basal turn was significantly lower in the Slc12a2Em2/Em2 cochleae than that in the Slc12a2Em2/+ cochleae at 4 W (Supplemental Fig. 4B, D. These findings indicate importance of Slc12a2 with the exon 21-encoded region for endolymph production after the onset of hearing as well.
The cell densities in the eosinophilic StV areas were approximately 2-fold higher in Slc12a2Em2/Em2 than in Slc12a2Em2/+ cochleae at the all three turns (N = 4, Fig. 2S–U). Due to tissue fragility, surface preparations of auditory epithelium could not be separated from Slc12a2Em2/Em2 cochleae fixed in 4% paraformaldehyde. Cochleae embedded in paraffin blocks were further subjected to nanoscale computed tomography (nano-CT) at low energy X-ray anode, allowing membranous tissues to be observed without damaging them during sectioning and dissection.
Nano-CT images showed reduced-sized StVs without vacuolization of spiral prominence in the lateral wall in Slc12a2Em2/Em2 (Fig. 3A; Supplemental Movie 3) compared with those in Slc12a2Em2/+ (Fig. 3B; Supplemental Movie 4) cochleae. These findings suggest that the occasional degeneration of part of the lateral wall in Slc12a2Em2/Em2 cochleae (asterisk in Fig. 2Q) was an artefact of paraffin sectioning. Whether the partial absence of spiral ganglion neurons (SGNs) in Slc12a2Em2/Em2 cochleae (Supplemental Fig. 3D, F) indicates degeneration of SGNs awaits further precise measurement. The heights of StVs lined with spiral ligament from the apical to the basal turns appeared to be significantly shorter in Slc12a2Em2/Em2 than the Slc12a2Em2/+ cochleae (Fig. 3C-G), confirming the reduction in StV. In addition, CT image signals were brighter at the basal turn of the StV area in Slc12a2Em2/Em2 cochleae (Supplemental Fig. 5), indicating that the subcellular organelles of the StV in Slc12a2Em2/Em2 cochleae are condensed, especially in the basal turn.
Reconstructed nano-CT images of Slc12a2Em2 cochlea. Slc12a2Em2/+ (A, C) and Slc12a2Em2/Em2 (B, D) cochleae at P28. (C, D) Visualization of StVs at basal turns (magenta). (E-G) Lengths of StV lined with spiral ligament at the apical (E), middle (F), and basal (G) turns (N = 3, two images per turn). Small arrows indicate non-homogenous brightness in the areas of spiral ganglion neurons. Scale bars = 100 μm.
Immunostaining of the cochlear specimens revealed that Slc12a2 signals were very low and almost indistinguishable from background at P1 (Fig. 4A, B), except at the apical surface of part of the GER in the Slc12a2Em2/Em2 cochleae (Fig. 4B). At 4 W, Slc12a2 signals were detectable at the StV and in a subset of spiral ligament fibrocytes (SLF) from the apical to basal turns in both the Slc12a2Em2/+ and Slc12a2Em2/Em2 cochleae (Fig. 4C–H). Although the signals of the marginal cells of the StV in the Slc12a2Em2/+ cochleae were limited to the basolateral membrane (Fig. 4C–E), an intense Slc12a2 signal was observed at the apical membrane of the marginal cells in Slc12a2Em2/Em2 cochleae (Fig. 4F–H), suggesting that the Slc12a2Δex21 is mistrafficked. Immunostaining of the cochlear specimens with anti-Kcnj10 antibody, a marker for intermediate cells of the StV, showed intermediate cells beneath the marginal cell layer in both Slc12a2Em2/+ and Slc12a2Em2/Em2 cochleae (Supplemental Fig. 6). The Kcnj10 signals at the basal turn of the Slc12a2Em2/Em2 cochleae were found to have accumulated at the superior and inferior ends (Supplemental Fig. 6D), as if the cells of the StV were interfolded.
Slc12a2 signals in the Slc12a2Em2 cochlea. Paraffin sections of Slc12a2Em2/+ (A, C-E) and Slc12a2Em2/Em2 (B, F-H) cochleae at P1 (A, B) and 4 W (C-H) were immunostained with anti-SLC12A2 antiserum. Arrows indicate StV. SLF; spiral ligament fibrocytes. Scale bars = 50 μm (A-C, E,F, H) and 10 μm (D, G).
Transcriptomic analysis of Slc12a2 Em2/Em2 cochleae
The pathology of Slc12a2Em2/Em2 cochleae at the level of gene expression may be better understood by RNA-seq analysis using whole cochlear tissues dissected from three Slc12a2+/+ and three Slc12a2Em2/Em2 mice each at 4 W. Visualization of mapped transcript sequences on an Integrative Genomics Viewer30 demonstrated that sequencing reads mapped to exon 21 of Slc12a2 (NM_009194.3) were predominantly detected in Slc12a2+/+ but not in Slc12a2Em2/Em2 cochleae (Supplemental Fig. 7). The small amount of exon 21-skipped reads in the Slc12a2+/+ cochleae were likely expressed in the SGNs31. RNA-seq analysis showed that 142 and 554 differentially expressed genes were up- or down-regulated more than 2-fold in Slc12a2Em2/Em2 cochleae, respectively (Fig. 5A, B; Supplemental Table 2). Gene ontology enrichment analysis demonstrated that multiple genes upregulated in the Slc12a2Em2/Em2 cochlea were associated with cell adhesion, including tight junction formation (Supplemental Table 3). Of the six Cldns more highly expressed in Slc12a2Em2/Em2 than in Slc12a2+/+cochleae by RNA-seq, three (Cldn6, Cldn9, and Cldn24) were found to show significantly higher levels of expression (p < 0.05) in Slc12a2Em2/Em2 than in Slc12a2+/+ cochleae by qRT-PCR (Supplemental Fig. 8A–F), as were Gata3, Mcoln3, Slc26a4, and Fat2 mRNAs (Supplemental Fig. 8G–J). Interestingly, ontology analysis also demonstrated that multiple enriched genes associated with inner ear including Gata3, Slc26a4, and Tmprss3 were upregulatd in Slc12a2Em2/Em2 cochlea (Supplemental Table 4). Among the 554 genes down-regulated in Slc12a2Em2/Em2 relative to Slc12a2+/+ cochleae, several were associated with cell signaling, the nervous system, and ion transport (Supplemental Table 3), with downregulation of Mbp and Prph in Slc12a2Em2/Em2 cochleae confirmed by qRT-PCR (Supplemental Fig. 8K–N).
Changes of gene expression in the whole cochleae of Slc12a2Em2/Em2 mice. (A) Heatmap of genes differentially expressed in the whole cochleae of Slc12a2Em2/Em2 and Slc12a2+/+ mice. Red and green indicate higher or lower expression levels in the Slc12a2Em2/Em2 cochlea. (B) Volcano plot of the differentially expressed genes. Genes subjected to qRT-PCR (Supplemental Fig. 7) are indicated. Red and blue dots indicate genes with higher- or lower-expression in Slc12a2Em2/Em2 cochleae. (C-E) Levels of expression of Cldn9 in the developing Slc12a2Em2 cochleae. N = 4.
qRT-PCR showed that the gene most highly upregulated in Slc12a2Em2/Em2 relative to Slc12a2+/+ cochleae was Cldn9. The expression of Cldn9 was significantly higher at 4 W, but not at P1 nor P12, when the reduction in StV size was already evident (Fig. 5C–E). Immunostaining of cochleae at 4 W with anti-Cldn9 antiserum detected signals at between the surfaces of marginal cells in both Slc12a2Em2/+ and Slc12a2Em2/Em2 cochleae, as previously reported32 (Fig. 6A–D). These signals were weak despite the apparent morphological change of the StV in the Slc12a2Em2/Em2 cochlea. More intense Cldn9 signals were detected in the organ of Corti of Slc12a2Em2/Em2 than of Slc12a2Em2/+ cochleae (Fig. 6E–J). In addition, the number of nuclei corresponding to the positions of inner and outer hair cells appeared to be reduced in Slc12a2Em2/Em2 cochleae (asterisks in Fig. 6F, I). Because the hair cell marker Myo7a could not be detected in samples fixed with 10% trichloroacetic acid for Cldn9 immunostaining, the exact numbers of hair cells in Slc12a2Em2/Em2 cochleae could not be counted.
Cldn9 signals in the Slc12a2Em2 cochlea at 4 W. Cldn9 signals in the lateral wall (A-D) and strial surface preparations (E-J) of the cochleae of Slc12a2Em2/+ (A, B,E-G) and Slc12a2Em2/Em2 (C, D,H-J) mice. The signals in (A-D) are enhanced and not comparable to those in (E-J). (E-J) Cldn9 signals counterstained with Hoechsst 33,258 (Nucleus) in the auditory epithelium. Asterisks indicate predicted positions of hair cells. Scale bar = 50 μm.
Normal hearing of Slc12a2 Em1/Em1 mice and species-specific splicing property of exon 21
The second mouse strain generated in this study, Slc12a2Em1, was also characterized. The NM_009194.3:c.2912‒2 A > G variant in Em1 allele was initially considered equivalent to the splice variant of SLC12A2 (NM_001046.3:c.2930–2 A > G) detected in a patient with DFNA782. Unexpectedly, RT-PCR of mRNA isolated from Slc12a2Em1/Em1 cochleae detected two PCR products: one with a size similar to that of the transcript that included exon 21, and the other similar in size to the Slc12a2Em2/Em2 transcript (Fig. 7A). ABR thresholds of Slc12a2Em1/Em1 mice (21.50 ± 10.41, 30.50 ± 9.88, and 38.25 ± 6.25 db SPL at 8, 16, and 32 kHz, respectively) and those of Slc12a2Em1/+ mice (11.86 ± 7.56, 17.43 ± 9.00, and 27.00 ± 9.13 db SPL) did not show significant differences (p = 0.16628, 0.06212, and 0.05406, respectively, Fig. 7B) and normal cochlear morphology at 4 W (Supplemental Fig. 9A, B) with proper Slc12a2 and Kcnj10 signals in the cochlea (Supplemental Fig. 9C, D). Sanger sequencing demonstrated that the longer PCR product detected in Slc12a2Em1/Em1 cochleae included an incomplete exon 21, with deletion of its first nine bases (Fig. 7C, D). We presume that the resulting Slc12a2 protein, truncated by three amino acids (NP_033220.2:p.D971_E973del), mediated proper trafficking of the protein to the basolateral membrane of marginal cells and retained sufficient functional activity as an ion cotransporter for Slc12a2Em1/Em1 mice to have normal hearing. Complete omission of exon 21 was shown in cerebellar transcripts of both Slc12a2Em1/Em1 mice (Fig. 7A) without abnormal behavior and Slc12a2Em2/Em2 mice (Fig. 1B) with head-bobbing behavior. Therefore, omission of exon 21 of the Slc12a2 transcript in the brain may not be related to the behavioral phenotypes. Further investigation of the transcripts in other brain subregions in Slc12a2Em2/Em2 mice will clarify this assumption.
Characterization of the Slc12a2Em1 mouse. (A) RT-PCR of samples from the cochleae and cerebella of Slc12a2Em1 mice at 4 W. Genotypes are shown at the top. (B) ABR thresholds of Slc12a2Em1/+ and Slc12a2Em1/Em1 mice at 4 W. (C, D) Partial nucleotide sequence of Slc12a2 (NM_009194.3) (C) and partial electropherogram of the longer RT-PCR transcript (D) in Slc12a2Em1/Em1 cochleae (asterisk in A). Nucleotides mapped to exons 20, 21, and 22 are in green, red, and blue, respectively. Spliced out nucleotides in the Em1 allele are highlighted in gray. Predicted amino acid residues are shown below the sequences.
Comparison of the genomic sequences of SLC12A2 orthologues demonstrated that exon 21 sequences were highly conserved among Placentalia33 (Supplemental Fig. 10). The nucleotide at the 5’ end of exon 21 was shown to be thymine in the parvorder Catarrhini (old world monkeys and Hominoidea including human) but adenine in the parvorders Platyrrhini (new world monkeys) and Haplorhini (such as tarsier), the suborder Strepsirrhini (such as lemurs, galagos, and loris), and all other mammalian species including mouse. In addition, nucleotide 24 of exon 21 was found to be guanine in the parvorder Catarrhini, but cytosine in all other placental mammals.
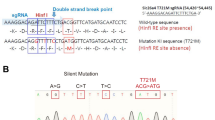
The effect of the acceptor site variant on exon 21 splicing was investigated by a minigene assay using 1,518 bases of the exon 21 region of mouse Slc12a2 genomic DNA (Fig. 8A). Following transfection of the heterologous minigene vector into HEK293T cells, the wild-type allele resulted in a reporter transcript of the expected size, including 48 bp from exon 21 (Fig. 8B, lane 3). The Em1 allele resulted in the reporter transcript of approximately the predicted size including exon 21 and the shorter reporter transcript without exon 21 (Fig. 8B, lane 4). Cloning and Sanger sequencing revealed that the longer PCR products included not only the nine bases shorter transcript corresponding to the transcript in the Slc12a2Em1/Em1 cochlea but also the other transcript with distinct cryptic acceptor site, consisting of seven bases upstream of the original 5’ end of exon 21 (Supplemental Fig. 11). Utilization of this additional cryptic acceptor site distinct from the transcript of Slc12a2Em1/Em1 cochleae could be due to the differences in splicing factors between mouse cochlear tissues and HEK293T cells. Replacing of the adenine at the 5’ end of exon 21 by thymine (c.2912 A > T) had no effect on the inclusion of exon 21 in the wild-type allele but resulted in complete exclusion of exon 21 from the Em1 allele (Fig. 8B, lanes 5, 6). These results indicated that the Em1 allele was not equivalent to the splice site variant detected in a DFNA78 patient due to a single nucleotide difference at the 5’ end of exon 21.
Minigene assay of exon 21 of Slc12a2. (A) Minigene structures. Raus sarcoma virus long terminal repeat (LTR), exon2 and 3 of rat insulin-2 (rnIns2), M13 forward sequencing primer (M13F) and M13 reverse primer (M13R), partial genomic region of wild-type or Em2 allele of Slc12a2 including exon 21, and polyA signal are shown. Numbers in blue indicate the nucleotide lengths (bp) of the Slc12a2 genomic region inserted into the vector. Gray arrows indicate primer sets to amplify reporter transcripts. (B) RT-PCR to detect reporter transcripts in HEK293T cells. Inserted alleles with nucleotide at c.2912 are indicated above the image of the RT-PCR transcripts. Lane 7; 100 bp ladder marker.
Discussion
Hearing loss and complete omission of exon 21 from transcripts of Slc12a2 in Slc12a2Em2/Em2 mice were similar to findings in patients with severe to profound hearing loss and omission of exon 21 in the SLC12A2 transcript in the splice site variant2. Abnormal behavior of Slc12a2Em2/Em2 mice also resembled vestibular defect occasionally observed in DFNA78 patients2,4. These phenotypic similarities are additional evidence of functional importance of the exon 21-encoded region of SLC12A2 for inner ear function. This study did not examine the mild but significant elevation of ABR thresholds in female Slc12a2Em2/+ mice. Further investigation including possible differential expression levels of Slc12a2 transcripts associated with sex34 and increasing the number of data acquisition would provide additional insight on the mechanisms of hearing loss in female Slc12a2Em2/+ mice.
The marked reduction of endolymph in the cochleae of Slc12a2Em2/Em2 at P1 indicates that a functional Slc12a2, acting as an ion cotransporter, is required for the production of primary endolymph without EP14,15. The association between a reduced StV and the presence of fluid in the SM observed as early as P12 in Slc12a2Em2/Em2 cochleae suggests that the Slc12a2Δex21 protein mediates production of incomplete secondary endolymph with presumably elevated K+ concentration, at least to some extent, even if the protein is devoid of cotransporter activity. Severe reduction of endolymph observed in adult Slc12a2-deficient cochleae26,27,28,29 would likely be explained by continuously inhibited production of primary as well as secondary endolymph due to absence of Slc12a2 protein. Because SLC12A2 balances the osmotic pressure of cells35,36,37,38,39, hearing loss in the cochleae of Slc12a2Em2/Em2 mouse can be due to two factors. First, an increase in K+ concentration in the StV for production of the secondary endolymph with elevated EP alters the environment in the cochlea as early as in P12. Second, a reduction of cell sizes in cochlear tissues is required to rebalance the osmotic pressure in response to increased ionic flow and in the presence of Slc12a2Δex21, which lacks cotransporter activity2. This may result in the leakage of extracellular fluid into the SM through gaps between smaller cells, eliminating the EPs required for hair cell excitation. Increased expression of Cldn9 in Slc12a2Em2/Em2 cochleae at 4 W would therefore be a secondary reaction of cochlear tissues to repair the gaps in intercellular barriers facing the SM, especially in the auditory epithelium. Cldn9 was shown important in controlling intercellular Na+ and K+ permeability40, with tight junction proteins during wound healing being important in skin41, intestine42, and lung43 injury. Upregulation of multiple genes related to inner ear function44 in the Slc12a2Em2/Em2 cochlea might also imply that cochlear tissues were responding to alleviate the Slc12a2Δex21-associated loss of cotransporter activity. Evidence of fluid leakage through the SM, precise determination of endolymph composition and measure of EP in the cochleae would be required to confirm this hypothesized pathology of hearing loss in Slc12a2Em2 mice, which is ongoing. Quantification of K⁺, Na⁺, and Cl⁻ levels in the cochlear fluids would provide additional physiological insight. Whether genes downregulated in Slc12a2Em2/Em2 cochleae such as those related to nervous system or ion transporters also contribute to elevated ABR thresholds will also be investigated in the future. This study is almost entirely focused on the study of Slc12a2Em2/Em2 mouse with profound hearing loss. Additional study to assess if Slc12a2Em2/Em2 as well as Slc12a2Em2/+ mice exhibit phenotypes other than hearing loss such as those reported in Slc12a2-deficient mice (reviewed by Gagnon and Delpire45, together with splicing alteration of Slc12a2 transcripts in other organs/tissues including subregions of the brain will be required to verify the biological roles of the exon 21-encoded region. Measurement of Slc12a2 and Slc12a2Δex21 protein levels and their changes in the cochleae associated with genotypes and sex, and rescue experiment such as gene editing the Em2 allele back to wild type allele in the Slc12a2Em2/Em2 cochlea would further confirm that Slc12a2Em2 mouse is a suitable model for DFNA78.
Among the genes up- and down-regulated in the cochleae of Slc12a2Em2/Em2 mice, Mcoln3 was the only gene whose encoded protein has been shown to interact with SLC12A225. Although a constitutively active variant of Mcoln3, with a semi-dominant mode of inheritance, has been shown to cause progressive hearing loss in varitint-waddler mice46,47,48,49, direct evidence of an association between Mcoln3 and Slc12a2Δex21 proteins in the cochlea is required to determine whether these proteins are associated with the pathology of hearing loss in Slc12a2Em2/Em2 mice.
The detection of Slc12a2Δex21 signals at the apical membrane of marginal cells of the StV in the Slc12a2Em2/Em2 cochleae supports the previous report that the exon 21-encoded region mediates the trafficking of Slc12a2 protein to the basolateral membrane in polarized cells23. It can be speculated that Slc12aΔex21 may have neomorphic and adverse effects at the apical membrane of marginal cells. A comprehensive study to identify molecules interacting with Slc12a2Δex21 as well as those associated with the exon 21-encoded region in cochlear tissues would clarify these possibilities. It would also be important to investigate whether the exon 21-encoded regions with two distinct amino acid residues between Catarrhini and the other mammalian species share orthologous or distinct binding partners that could result in distinct functional properties. These studies would reveal molecular mechanisms of hearing loss of Slc12a2Em2 mouse proposed as an animal model of DFNA78.
Species-specific nucleotide sequences that affect exon 21 splicing based on the study of Em1 allele provided an intriguing example to consider primate evolution. Evaluation of nucleotides in exon 21 specific to species of Catarrhini showed species-specific differences at the 5’ end and at nucleotide 24 of exon 21 that resulted in amino acid changes (977V and 985A, respectively, in humans to 971D and 979P, respectively in mice), suggesting positive selection of primate species that split from Platyrrhini species50. Results of the present study also suggested that the change at the 5’ end was selected to affect the robustness of exon 21 splicing, which seems to be more easily spliced out in Catarrhini than in other mammalian species. The first seven nucleotides of mouse exon 21 (ATGAGGA) would be recognized by the splicing factor SRSF151, whereas this sequence in human exon 21 (TTGAGGA) would not. The presence of the identical heptanucleotide sequence (ATGAGGA) in the Em2 allele suggests that cryptic splicing of exon 21 derived from the Em1 allele, which is distinct to humans, is affected not only by SRSF1 but also by other splicing factors. Endogenous exon 21- skipping SLC12A2 transcripts are detectable predominantly in central nervous system of both human52 and mouse53, suggesting that mechanism of alternative splicing is not related to the species-specific nucleotide changes on the exon 21.
In conclusion, the present study described the generation and partial characterization of a mouse strain Slc12a2Em2, which we propose as an animal model of DFNA78. Further investigation of the molecular and electrophysiological properties associated with the pathology of hearing loss may confirm whether SLC12A2 acts as a cotransporter as well as a mediator of secondary endolymph production in the cochlea.
Methods
Sex as a biological variable
Our study included both male and female animals, and the data are derived from both sex except ABR measurements.
Generation of Tg mice and genotyping
Slc12a2 KI mice were created on an FVB/NJcl (FVB/N, Japan CLEA, Japan) genetic background using the CRISPR/Cas9 system with Alt-R CRISPR-Cas9 tracrRNA (Integrated DNA Technologies). Briefly, single-guide RNA and custom-designed single-stranded mouse Slc12a2 genomic sequence donor oligonucleotide (Supplemental Table 1) were injected into fertilized eggs from FVB/N mice. Surviving eggs (n = 330) were transduced into 11 pseudo-pregnant FVB/N females. After genotyping, a male carrying the Em1 allele (c.2912–2 A > G) and two females carrying the Em2 allele (c.2912-4_2913del) were subjected to in vitro fertilization. Both Slc12a2Em1 and Slc12a2Em2 strains were backcrossed with FVB/N mice for at least four generations to reduce the possibility of an off-target effect resulting from genome editing. ABR were measured as described54. ABR thresholds on the left ears were measured at 8, 16, and 32 kHz.
Histochemical and nano-computed tomography (nano-CT) analyses
Mice at P12 and 4 W were anesthetized by injection of ketamine (10 mg/kg) and xyladine (10 mg/kg). Mice at P1 were anesthetized on ice. The mice were decapitated, and their inner ears were dissected and subjected to RNA extraction or fixed in 4% paraformaldehyde (PFA) overnight at 4 °C. The latter samples were decalcified with 0.1 M EDTA/phosphate buffer at 4 °C for at least 2 weeks and cryo- or paraffin-embedded. To measure cell densities of the StV, right inner ears were paraffin-embedded and cut into 5-µm sections, which were stained with hematoxylin-eosin, and viewed under a microscope (×20 objective lens). The numbers of hematoxylin-positive nuclei in the StV were counted using ImageJ software. Areas of the StV were defined as eosinophilic regions of the lateral wall. For measurement of areas of SM and SV, a midmodiolar section of each cochlea was selected, and both scala at the basal turn were outlined and the areas were measured using ImageJ software. Proportion of areas of SM to SV were estimated.
For nano-CT analysis, decalcified right inner ears dissected at P28 were embedded in paraffin. Biopsy samples 3 mm in diameter were punched out (KAI Medical, Japan) and analyzed by nano3DX (RIGAKU, Japan) using a copper X-ray anode (8 keV) to collect 2250 projections of voxel size 0.636 μm (L0540, bin1, XD1) at an exposure time of 30 s. The height of each StV lining with a spiral ligament and the brightness of each StV area excluding background were measured using ImageJ software.
Immunohistochemical analyses
Right inner ears embedded in paraffin were cut into 5-µm sections. These sections were rehydrated, pretreated with boiling 10 mM sodium citrate buffer for 30 min, and blocked in PBS containing 5% fetal bovine serum (FBS), 1% bovine serum albumin, and 0.05% Tween-20, and incubated with primary antibodies at 4 °C overnight. For Cldn9 staining, dissected inner ears were fixed with 10% trichloroacetic acid for 1 h and decalcified for 1 week. The lateral wall, StV and auditory epithelium dissected by surface preparation were subjected to immunostaining. The primary antibodies used in this study included rabbit anti-SLC12A2 (Abcam, ab59791, 1:100), rabbit anti-KCNJ10 (allomone, APC-035, 1:800), and rabbit anti-Cldn9 (1:1000, a gift from Dr. Mikio Furuse of the National Institute of Physiological Sciences)32. Binding was visualized by incubation with appropriate secondary antibodies conjugated to either Alexa 488 or Alexa 568 (ThermoFisher) followed by counterstaining with Hoechst 33,258 (DOJINDO, Japan). Images were captured by fluorescence microscopy (DM2500, Leica Microsystems) and confocal microscopy (Axio 700, Carl Zeiss Microscopy).
Bulk RNA-seq analysis
Left whole cochlear tissues from 4-week-old Slc12a2+/+ and Slc12a2Em2/Em2 mice (n = 3 each) were dissected after perfusion of ice-cold saline and homogenized in acidic phenol solution (Isogen II, NipponGene, Japan). Extracted total RNA was treated with DNase I and cleaned up using RNeasy mini kits (Qiagen, Germany). Total RNA (100 ng) samples, each having a RIN value ≥ 7.8, were subjected to polyA selected RNA-seq library preparation using NEBNext Ultra II directional RNA library prep kit (E7760S, New England Biolabs, USA). The samples were subjected to 100 × 2 bp paired-end sequencing using the Hiseq 2500 platform (Illumina). Approximately 40 million reads per sample were subjected to quality checking by Trimmomatic55, mapped to reference mouse genome (GRCm39) using STAR56, and quantified using RSEM57.
RT-PCR and qRT-PCR
After anesthetization and perfusion with saline, left inner ears were dissected and immediately homogenized in Isogen II. Total RNA was extracted and purified using RNeasy mini kits (QIAGEN) and subjected to RT-PCR and RNA-seq analyses. For RT-PCR and qRT-PCR, total RNA samples were reverse transcribed using SuperScript III kits (ThermoFisher). Slc12a2 transcripts were amplified using a Type-iT Multi Detection PCR kit (QIAGEN) and analyzed using Bioanalyzer 2100. Amplification consisted of denaturation at 95 °C for 5 min, followed by 40 cycles of denaturation at 95 °C for 30 s, annealing at 54 °C for 90 s, and extension at 72 °C for 40 s, and a final extension at 68 °C for 5 min. Amplified PCR products from Slc12a2+/+ and Slc12a2Em2/Em2 cochleae were subjected to direct sequencing. The longer PCR product from Slc12a2Em1/Em1 mouse cochleae was gel-extracted and cloned into the pCRII vector (zero Blunt TOPO PCR cloning kit, Invitrogen/ThermoFisher). Levels of expression of genes of interest were measured using PowerUpSybrGreen kits (Applied Biosystems) and QuantStudio 3 (Applied Biosystems) according to the manufacturer’s protocols. Nucleotides sequences of the primers are shown in Supplemental Table 1.
Minigene assay
Briefly, a splicing reporter was prepared by amplifying the genomic region of Slc12a2 consisting of part of intron 20 (854 bp), exon 21 (48 bp), and part of intron 21 (616 bp). The resulting product was inserted between M13 forward and M13 reverse sequencing primer regions in the pSpliceExpress vector (Addgene #32485)58. Heterologous reporters with nucleotide changes (c.2912 A > T) were also prepared. Each vector was transfected into HEK293T cells (RIKEN Bio Resource Center) using Lipofectamine 2000 (Thermo Fisher Scientific) and incubated for 2 days (three independent experiments). Total RNA was extracted using TRIZOL reagent (Thermo Fisher Scientific), reverse transcribed with ReverTra Ace qPCR RT Master (TOYOBO, Japan), and subjected to RT-PCR using Quick Taq HS DyeMix (TOYOBO). Longer PCR product of the reporter transcripts derived from the Slc12a2Em1 allele was gel-extracted, cloned into pGEM-T Easy vector (Promega), and sequenced.
Statistics
Statistical significance was determined by one-way ANOVA followed by the Tukey-Kramer test.
Data availability
RNA-sequencing data are deposited to DNA Data Bank of Japan (DDBJ, BioProject ID: PSUB023824). Slc12a2Em1 and Slc12a2Em2 mouse strains generated in this study are deposited to RIKEN BioResource Research Center.
Change history
11 July 2025
The original online version of this Article was revised: The original version of this Article contained an error in the Acknowledgements section. It now reads: “The authors thank Dr. Atsuo Ogura at Bioresource Engineering, RIKEN, for help with in vitro fertilization of Slc12a2Em1 mice; Dr. Atsushi Toyoda at Comparative Genomics Laboratory, National Institute of Genetics, for technical support with RNA-seq; and Dr. Mikio Furuse of the National Institute for Physiological Sciences for providing rabbit anti-Claudin 9 antiserum. This study was supported by grants to HM from the Japan Society of Promotion of Sciences (KAKENHI; grant numbers 18K0933, 21K09598, and 24K12688), and from the JSPS Platform for Advanced Genome Science (grant number 212118), by a grant to TM from the Uehara Memorial Foundation (grant number 201820479), and by a grant to SI from the Amgen Foundation (Undergraduate Summer Research Program in Science and Biotechnology 2022).”
References
Van Camp, G. & Smith, R. A. Hereditary Hearing Loss Homepage. http://hereditaryhearingloss.org. Accessed 14 May 2024 (2024).
Mutai, H. et al. Variants encoding a restricted carboxy-terminal domain of SLC12A2 cause hereditary hearing loss in humans. PLoS Genet. 16, e1008643. https://doi.org/10.1371/journal.pgen.1008643 (2020).
Adadey, S. M. et al. Further confirmation of the association of SLC12A2 with non-syndromic autosomal-dominant hearing impairment. J. Hum. Genet. 66, 1169–1175. https://doi.org/10.1038/s10038-021-00954-6 (2021).
McNeill, A. et al. SLC12A2 variants cause a neurodevelopmental disorder or cochleovestibular defect. Brain 143, 2380–2387. https://doi.org/10.1093/brain/awaa176 (2020).
Vanniya, S. P. et al. PNPT1, MYO15A, PTPRQ, and SLC12A2-associated genetic and phenotypic heterogeneity among hearing impaired assortative mating families in Southern India. Ann. Hum. Genet. 86, 1–13. https://doi.org/10.1111/ahg.12442 (2022).
Yamamoto, N. et al. Comprehensive gene panel testing for hearing loss in children: Understanding factors influencing diagnostic yield. J. Pediatr. 262, 113620. https://doi.org/10.1016/j.jpeds.2023.113620 (2023).
Haas, M. The Na-K-Cl cotransporters. Am. J. Physiol. 267 (C869-885). https://doi.org/10.1152/ajpcell.1994.267.4.C869 (1994).
Crouch, J. J., Sakaguchi, N., Lytle, C. & Schulte, B. A. Immunohistochemical localization of the Na-K-Cl co-transporter (NKCC1) in the gerbil inner ear. J. Histochem. Cytochemistry: Official J. Histochem. Soc. 45, 773–778. https://doi.org/10.1177/002215549704500601 (1997).
Kikuchi, T., Adams, J. C., Miyabe, Y., So, E. & Kobayashi, T. Potassium ion recycling pathway via gap junction systems in the mammalian cochlea and its interruption in hereditary nonsyndromic deafness. Med. Electron. Microscopy: Official J. Clin. Electron. Microscopy Soc. Japan. 33, 51–56. https://doi.org/10.1007/s007950000009 (2000).
Schulte, B. A. & Adams, J. C. Distribution of immunoreactive Na+,K+-ATPase in gerbil cochlea. J. Histochem. Cytochem. 37, 127–134 (1989).
Hibino, H., Nin, F., Tsuzuki, C. & Kurachi, Y. How is the highly positive endocochlear potential formed? The specific architecture of the stria vascularis and the roles of the ion-transport apparatus. Pflug, Arch.: Eur. J. Physiol. 459, 521–533. https://doi.org/10.1007/s00424-009-0754-z (2010).
Wangemann, P. Supporting sensory transduction: cochlear fluid homeostasis and the endocochlear potential. J. Physiol. 576, 11–21. https://doi.org/10.1113/jphysiol.2006.112888 (2006).
Mathiesen, B. K. et al. Delivery of gene therapy through a cerebrospinal fluid conduit to rescue hearing in adult mice. Sci. Transl Med. 15, eabq3916. https://doi.org/10.1126/scitranslmed.abq3916 (2023).
Li, Y., Liu, H., Zhao, X. & He, D. Z. Endolymphatic potential measured from developing and adult mouse inner ear. Front. Cell. Neurosci. 14, 584928. https://doi.org/10.3389/fncel.2020.584928 (2020).
Yamasaki, M., Komune, S., Shimozono, M., Matsuda, K. & Haruta, A. Development of monovalent ions in the endolymph in mouse cochlea. ORL J. Oto-Rhino-Laryngol. Relat. Spec. 62, 241–246. https://doi.org/10.1159/000027753 (2000).
Ehret, G. Development of absolute auditory thresholds in the house mouse (Mus musculus). J. Am. Audiol. Soc. 1, 179–184 (1976).
Furuse, M. & Tsukita, S. Claudins in occluding junctions of humans and flies. Trends Cell. Biol. 16, 181–188. https://doi.org/10.1016/j.tcb.2006.02.006 (2006).
Sineni, C. J. et al. A truncating CLDN9 variant is associated with autosomal recessive nonsyndromic hearing loss. Hum. Genet. 138, 1071–1075. https://doi.org/10.1007/s00439-019-02037-1 (2019).
Chen, S. et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 625, 92–100. https://doi.org/10.1038/s41586-023-06045-0 (2024).
Bilal Shamsi, M. et al. Clinical characterization and further confirmation of the autosomal recessive SLC12A2 disease. J. Hum. Genet. 66, 689–695. https://doi.org/10.1038/s10038-021-00904-2 (2021).
Macnamara, E. F. et al. Kilquist syndrome: A novel syndromic hearing loss disorder caused by homozygous deletion of SLC12A2. Hum. Mutat. 40, 532–538. https://doi.org/10.1002/humu.23722 (2019).
Stodberg, T. et al. SLC12A2 mutations cause NKCC1 deficiency with encephalopathy and impaired secretory epithelia. Neurol. Genet. 6, e478. https://doi.org/10.1212/NXG.0000000000000478 (2020).
Carmosino, M., Gimenez, I., Caplan, M. & Forbush, B. Exon loss accounts for differential sorting of Na-K-Cl cotransporters in polarized epithelial cells. Mol. Biol. Cell. 19, 4341–4351. https://doi.org/10.1091/mbc.E08-05-0478 (2008).
Koumangoye, R., Omer, S. & Delpire, E. A Dileucine motif in the COOH-terminal domain of NKCC1 targets the cotransporter to the plasma membrane. Am. J. Physiol. Cell Physiol. 316, C545–C558. https://doi.org/10.1152/ajpcell.00023.2019 (2019).
Huttlin, E. L. et al. The bioplex network: A systematic exploration of the human interactome. Cell 162, 425–440. https://doi.org/10.1016/j.cell.2015.06.043 (2015).
Delpire, E., Lu, J., England, R., Dull, C. & Thorne, T. Deafness and imbalance associated with inactivation of the secretory Na-K-2Cl co-transporter. Nat. Genet. 22, 192–195. https://doi.org/10.1038/9713 (1999).
Flagella, M. et al. Mice lacking the basolateral Na-K-2Cl cotransporter have impaired epithelial chloride secretion and are profoundly deaf. J. Biol. Chem. 274, 26946–26955 (1999).
Pace, A. J. et al. Failure of spermatogenesis in mouse lines deficient in the Na(+)-K(+)-2Cl(-) cotransporter. J. Clin. Investig. 105, 441–450. https://doi.org/10.1172/JCI8553 (2000).
Dixon, M. J. et al. Mutation of the Na-K-Cl co-transporter gene Slc12a2 results in deafness in mice. Hum. Mol. Genet. 8, 1579–1584 (1999).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26. https://doi.org/10.1038/nbt.1754 (2011).
Kurki, S. N. et al. Expression patterns of NKCC1 in neurons and non-neuronal cells during cortico-hippocampal development. Cereb. Cortex. 33, 5906–5923. https://doi.org/10.1093/cercor/bhac470 (2023).
Kitajiri, S. I. et al. Expression patterns of Claudins, tight junction adhesion molecules, in the inner ear. Hear. Res. 187, 25–34. https://doi.org/10.1016/s0378-5955(03)00338-1 (2004).
Foley, N. M. et al. A genomic timescale for placental mammal evolution. Science 380, eabl8189. https://doi.org/10.1126/science.abl8189 (2023).
Jukneviciene, M. et al. Valproic acid inhibits NA-K-2CL cotransporter RNA expression in male but not in female rat thymocytes. Dose Response. 17, 1559325819852444. https://doi.org/10.1177/1559325819852444 (2019).
Chou, C. L. et al. Roles of basolateral solute uptake via NKCC1 and of myosin II in vasopressin-induced cell swelling in inner medullary collecting duct. Am. J. Physiol. Renal. Physiol. 295, F192–201. https://doi.org/10.1152/ajprenal.00011.2008 (2008).
O’Brien, J. A., Walters, R. J., Valverde, M. A. & Sepulveda, F. V. Regulatory volume increase after hypertonicity- or vasoactive-intestinal-peptide-induced cell-volume decrease in small-intestinal crypts is dependent on Na(+)-K(+)-2Cl- cotransport. Pflug, Arch.: Eur. J. Physiol. 423, 67–73. https://doi.org/10.1007/BF00374962 (1993).
Gregoriades, J. M. C., Madaris, A., Alvarez, F. J. & Alvarez-Leefmans, F. J. Genetic and Pharmacological inactivation of apical Na(+)-K(+)-2Cl(-) cotransporter 1 in choroid plexus epithelial cells reveals the physiological function of the cotransporter. Am. J. Physiol. Cell Physiol. 316, C525–C544. https://doi.org/10.1152/ajpcell.00026.2018 (2019).
Demian, W. L. et al. The ion transporter NKCC1 links cell volume to cell mass regulation by suppressing mTORC1. Cell. Rep. 27, 1886–1896e1886. https://doi.org/10.1016/j.celrep.2019.04.034 (2019).
Kahle, K. T. et al. K-Cl cotransporters, cell volume homeostasis, and neurological disease. Trends Mol. Med. 21, 513–523. https://doi.org/10.1016/j.molmed.2015.05.008 (2015).
Nakano, Y. et al. A claudin-9-based ion permeability barrier is essential for hearing. PLoS Genet. 5, e1000610. https://doi.org/10.1371/journal.pgen.1000610 (2009).
Volksdorf, T. et al. Tight junction proteins Claudin-1 and occludin are important for cutaneous wound healing. Am. J. Pathol. 187, 1301–1312. https://doi.org/10.1016/j.ajpath.2017.02.006 (2017).
Slifer, Z. M. & Blikslager, A. T. The integral role of tight junction proteins in the repair of injured intestinal epithelium. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21030972 (2020). 21.
Wray, C. et al. Claudin-4 augments alveolar epithelial barrier function and is induced in acute lung injury. Am. J. Physiol. Lung Cell. Mol. Physiol. 297, L219–227. https://doi.org/10.1152/ajplung.00043.2009 (2009).
Sajan, S. A., Warchol, M. E. & Lovett, M. Toward a systems biology of mouse inner ear organogenesis: gene expression pathways, patterns and network analysis. Genetics 177, 631–653. https://doi.org/10.1534/genetics.107.078584 (2007).
Gagnon, K. B. & Delpire, E. Physiology of SLC12 transporters: lessons from inherited human genetic mutations and genetically engineered mouse knockouts. Am. J. Physiol. Cell Physiol. 304, C693–714. https://doi.org/10.1152/ajpcell.00350.2012 (2013).
Cable, J. & Steel, K. P. Combined cochleo-saccular and neuroepithelial abnormalities in the Varitint-waddler-J (VaJ) mouse. Hear. Res. 123, 125–136. https://doi.org/10.1016/s0378-5955(98)00107-5 (1998).
Di Palma, F. et al. Mutations in Mcoln3 associated with deafness and pigmentation defects in varitint-waddler (Va) mice. Proc. Natl. Acad. Sci. U.S.A. 99, 14994–14999. https://doi.org/10.1073/pnas.222425399 (2002).
Grimm, C. et al. A helix-breaking mutation in TRPML3 leads to constitutive activity underlying deafness in the varitint-waddler mouse. Proc. Natl. Acad. Sci. U.S.A. 104, 19583–19588. https://doi.org/10.1073/pnas.0709846104 (2007).
Xu, H., Delling, M., Li, L., Dong, X. & Clapham, D. E. Activating mutation in a Mucolipin transient receptor potential channel leads to melanocyte loss in varitint-waddler mice. Proc. Natl. Acad. Sci. U.S.A. 104, 18321–18326. https://doi.org/10.1073/pnas.0709096104 (2007).
Shao, Y. et al. Phylogenomic analyses provide insights into primate evolution. Science 380, 913–924. https://doi.org/10.1126/science.abn6919 (2023).
Liu, H. X., Zhang, M. & Krainer, A. R. Identification of functional exonic splicing enhancer motifs recognized by individual SR proteins. Genes Dev. 12, 1998–2012. https://doi.org/10.1101/gad.12.13.1998 (1998).
Vibat, C. R., Holland, M. J., Kang, J. J., Putney, L. K. & O’Donnell, M. E. Quantitation of Na+-K+-2Cl- cotransport splice variants in human tissues using kinetic polymerase chain reaction. Anal. Biochem. 298, 218–230. https://doi.org/10.1006/abio.2001.5398 (2001).
Randall, J., Thorne, T. & Delpire, E. Partial cloning and characterization of Slc12a2: the gene encoding the secretory Na+-K+-2Cl- cotransporter. Am. J. Physiol. 273, C1267–1277 (1997).
Mutai, H., Miya, F., Fujii, M., Tsunoda, T. & Matsunaga, T. Attenuation of progressive hearing loss in DBA/2J mice by reagents that affect epigenetic modifications is associated with up-regulation of the zinc importer Zip4. PloS One. 10, e0124301. https://doi.org/10.1371/journal.pone.0124301 (2015).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323. https://doi.org/10.1186/1471-2105-12-323 (2011).
Kishore, S., Khanna, A. & Stamm, S. Rapid generation of splicing reporters with pSpliceExpress. Gene 427, 104–110. https://doi.org/10.1016/j.gene.2008.09.021 (2008).
Acknowledgements
The authors thank Dr. Atsuo Ogura at Bioresource Engineering, RIKEN, for help with in vitro fertilization of Slc12a2Em1 mice; Dr. Atsushi Toyoda at Comparative Genomics Laboratory, National Institute of Genetics, for technical support with RNA-seq; and Dr. Mikio Furuse of the National Institute for Physiological Sciences for providing rabbit anti-Claudin 9 antiserum. This study was supported by grants to HM from the Japan Society of Promotion of Sciences (KAKENHI; grant numbers 18K0933, 21K09598, and 24K12688), and from the JSPS Platform for Advanced Genome Science (grant number 212118), by a grant to TM from the Uehara Memorial Foundation (grant number 201820479), and by a grant to SI from the Amgen Foundation (Undergraduate Summer Research Program in Science and Biotechnology 2022).
Author information
Authors and Affiliations
Contributions
HM contributed to the generation of mouse strains; to physiological and histochemical experiments; to RNA-seq analysis; and to designing, drafting, and finalizing the manuscript. YK, SN, and KM contributed to nanoscale CT analyses and drafting the manuscript. SI, ST, and NK contributed to minigene assays and drafting the manuscript. MF and TM assessed the contents and provided valuable comments to finalize the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics declarations and approval for animal experiments
All experimental protocols were approved by Institutional Animal Care and Use Committee of NHO Tokyo Medical Center (23-animal-01, 18-animal-02). All experiments were performed in accordance with relevant guidelines and regulations. Authors complied with the ARRIVE guidelines (https://arriveguidelines.org).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1
Supplementary Material 2
Supplementary Material 3
Supplementary Material 4
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mutai, H., Kuroda, Y., Noji, S. et al. Complete omission of exon 21 from Slc12a2 transcripts in mice results in hearing loss. Sci Rep 15, 14790 (2025). https://doi.org/10.1038/s41598-025-99827-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99827-7