Abstract
Oxford Nanopore Technologies (ONT) offers an efficient and effective solution for studying DNA methylation. Both R9.4.1 and R10.4.1 ONT chemistries have been widely used to generate numerous DNA methylation data, making it inevitable to conduct cross-ONT-chemistry methylation analysis. However, the two ONT chemistries have unique designs and may cause potential bias in methylation detection, complicating cross-chemistry methylation investigation. In this study, we sequenced two pairs of wild-type and their knockout samples using R9.4.1 and R10.4.1 chemistries and investigated the concordance and bias of the two pairs of ONT methylation data. Although we confirmed high concordances of methylation data generated by the two ONT chemistries and the improvement of R10 chemistry in repeat regions, we found that both chemistries possess detection bias for methylation. Thus, cross-ONT-chemistry methylation studies identified hundreds of thousands of differential methylation sites caused by chemistry variabilities. We also explored different methods to calculate coverage and methylation percentages. Our evaluations provide valuable recommendations for cross-ONT-chemistry methylation analysis and suggest better practices for robust methylation investigation.
Similar content being viewed by others
Introduction
DNA methylation is an epigenetic mechanism that adds a methyl group to a nucleotide base without modifying DNA sequences. The most common DNA methylation in the human genome is 5-methylcytosine (5mC). 5mC plays important roles in various necessary biological processes1, such as gene silencing2, safeguarding repetitive elements3, and maintaining genomic stability4. What is more important is that abnormal 5mC methylation levels have been found in a myriad of human diseases5,6,7,8. To reliably investigate DNA methylation and its roles in biological processes and human diseases, accurate detection of DNA methylation is needed9.
DNA methylation can be detected using approaches like methylation microarray10, bisulfite sequencing11, bisulfite-free methods (such as TAPS12 and EM-seq13, and long-read sequencing14. However, methylation array is limited to detecting 5mC at specific known methylation sites, while bisulfite and other short-read sequencing cannot reliably detect methylation in repeat regions, which account for about half of the human genome15. In contrast, Oxford Nanopore Technologies (ONT) and PacBio SMRT sequencing16 offer long-read solutions to detect DNA methylation, while ONT sequencing offers a cheaper long-read solution to overcome the struggles of methylation detection in repeat regions17.
ONT has two main chemistries, i.e., R10.4.1 flowcells with kit 14 chemistry (called R10 chemistry in this work) and R9.4.1 flowcells with kit 10 chemistry (called R9 chemistry in this work). R9.4.1 flowcell has one sensor on each pore18, and its basecalling is much more accurate compared to R7 and R8 chemistries, which were discontinued several years ago19. Thus, R9 chemistry was widely used to study DNA methylation20,21,22, generating a lot of methylation data and leading to the development of methylation detection tools23,24,25,26,27. R10.4.1 flowcells were released in 2023, containing two sensors per pore to improve basecalling errors18. As R9 chemistry was discontinued in the middle of 2024, R10 chemistry has begun to produce abundant methylation data as well28, and cross-ONT-chemistry methylation analysis is inevitable. However, few studies directly investigate the concordance and potential bias of cross-ONT-chemistry methylation data, making ONT methylation studies complex without a straightforward procedure for handling the data analysis.
To provide valuable recommendations for researchers to conduct ONT methylation analysis, we investigated the concordance of methylation detection by R10 and R9 chemistries as well as their inherent biases to remove potential false-positive analyses. To do this investigation, we used R9.4.1 flowcells to sequence a pair of an HCT116 wild-type (WT) sample and the paired sample after knocking out (KO) inositol polyphosphate multi-kinase (IPMK), while additionally using R10.4.1 flowcells to sequence another pair. We used modbam2bed29 to summarize the methylation and analyzed the Pearson correlation30,31,32,33,34 of different samples to study the concordance of R10 and R9 chemistries as well as chemistry-biased methylation positions. We further investigated different ways to calculate coverage and methylation percentages based on modbam2bed outputs and suggested a better practice to analyze ONT methylation data to reduce potential false dissection.
Results
Summary of ONT sequencing data
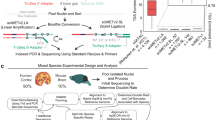
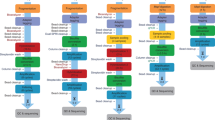
We cultured two pairs of HCT116 WT and the KO samples (after knocking out IPMK), and then extracted DNA for sequencing using either R9.4.1 ONT flowcells or R10.4.1 flowcells. Each sample were sequenced with > 30x coverage for robust analysis. We used ONT’s Dorado basecaller (version 7.2.13 + fba8e8925) for basecalling, minimap235 for long-read alignment, and modbam2bed29 for summarizing whole-genome methylation profiling, as shown in Fig. 1. The sequencing data is summarized in Table 1.
Workflow for data analysis. (1) HCT 116 WT and IPMK KO cell culture; (2) Cell lysis and total genomic DNA extraction; (3) ONT library preparation and adapter ligation; (4) DNA library loading; (5) ONT sequencing; (6) Basecalling with 5mC methylation, and alignment; (7) Whole-genome methylation summary; (8) Concordance and bias analysis of DNA methylation.
We then filtered out non-CpG or low-coverage (< 10) sites and investigated the concordance and potential bias of the methylation profiles between R9.4.1 and R10.4.1 flowcells. Since modbam2bed29 is a powerful tool for analyzing ONT methylation data, we also tested different methods to calculate the coverage and methylation percentages based on modbam2bed outputs, as described in the “Methods” section.
Concordance analysis for methylation detection using R9.4.1 and R10.4.1 flowcells
Abundant methylation data has been generated using ONT R9.4.1 flowcells and R10.4.1 flowcells, and thus, there is an increasing need to compare ONT methylation data generated across ONT chemistries. To assist the investigation of the effect of chemistry-biased methylation on real-world applications, we investigate the concordance and discordance of methylation data generated by R9.4.1 flowcells (R9 methylation data for short) and those generated by R10.4.1 flowcells (R10 methylation data for short).
We first assessed the reliability of methylation levels detected by ONT sequencing and bisulfite sequencing. We downloaded bisulfite sequencing data of WT HCT11636, and calculated the Pearson correlation coefficients of detected methylation between bisulfite data and R9 methylation and between bisulfite and R10 methylation (refer to Fig. 2a, b). The bisulfite sequencing data has 0.868 correlation against R10-Nanopore data and 0.839 correlation against R9-Nanopore data. This higher correlation suggests that Nanopore methylation data is very reliable, as demonstrated by numerous existing studies. Also, R10-Nanopore methylation data has a higher correlation than R9-Nanopore methylation data, suggesting the improvement in ONT R10 chemistries.
We then checked the concordance of R9 methylation data and R10 methylation data on different replicates of WT (or KO) samples, and the scatter plots are shown in Fig. 2. It can be seen from Fig. 2c–f that the replicates sequenced by different ONT sequencing chemistries have a higher correlation of methylation percentage. For example, the two WT replicates have a Pearson correlation of 0.9185, while the two KO replicates correlate with a value of 0.9194. In particular, R9 WT and R10 WT methylation data have 24,577,147 out of 34,132,876 (72.00%) methylation sites with ≤ 10% difference in methylation percentage, while R9 KO and R10 KO methylation data have 29,211,649 out of 40,200,383 (72.67%) methylation sites with ≤ 10% difference in methylation percentage. The higher correlation values and larger percentage of methylation sites with a smaller methylation difference demonstrate a high concordance of methylation data generated by R9 and R10 chemistries.
Concordance of detected methylation by bisulfite sequencing, R9.4.1, and R10.4.1 flowcells. (a) R9 vs. Bisulfite (b) R10 vs. Bisulfite; (c) WT pair with a log scale of the density; (d) KO pair with a log scale of the density; (e) WT HCT-116 pair without log scale; (f) IPMK-KO HCT-116 pair without log scale. R is the Pearson correlation coefficient. The shade of color from light to dark represents an increasing number of sites.
Besides using Pearson correlation coefficients, we also calculated the discordant percentage of methylations across the different chemistries. We used three thresholds of methylation difference (15%, 20%, 25%) to investigate the changes of discordant percentages. We found that the WT pair had 4.78% (1,632,048 out of 34,132,876) discordant sites, while the KO pair had 4.45% (1,788,722 out of 40,200,383) discordant sites with a ≥ 15% difference in methylation percentage. When the threshold increased to 20%, the WT pair had 2.94% (1,002,430 out of 34,132,876) discordant sites, while the KO pair had 2.70% (1,084,143 out of 40,200,383) discordant sites. The discordant percentage further decreased to 1.49% (507,515 out of 34,132,876) for the WT sample pair and to 1.92% (772,071 out of 40,200,383) for the KO pair when using 25% as the threshold of methylation difference. These lower discordant percentages clearly demonstrate the reliable detection of methylation across chemistries, as suggested by higher Pearson correlation coefficients.
Chemistry-biased methylation detection using R9.4.1 and R10.4.1 flowcells
To check the chemistry-biased effect of different ONT chemistries on methylation detection, we calculated the Pearson correlation of R9 WT against R9 KO or R10 KO, and of R10 WT against R9 KO or R10 KO, and showed the results in Table 2. We found that cross-chemistry (R9 and R10) WT and KO comparisons always show lower correlation values. For example, R9 WT against R10 KO correlates with 0.8432, lower than the correlation of 0.8612 for R9 WT against R9 KO. Similarly, R10 WT against R9 KO correlates to 0.8502, lower than the correlation of 0.8925 for R10 WT against R10 KO. This comparison demonstrates that methylation differences across ONT sequencing chemistries substantially affect differential methylation investigations across conditions.
Then, we investigated the methylation sites that have a larger difference (> 30%) between R9 and R10 methylation data. We found that a small number of methylation sites have a large methylation difference between R9 and R10 methylation data when comparing two WT replicates or two KO replicates. As shown in Fig. 2c, d, methylation sites located in the top-left corners suggest that the R9 chemistry detected few methylated positions while the R10 chemistry identified a higher percentage of methylation positions, or vice versa for the methylation sites located in the bottom-right corners. We referred to the sites in the top-left corners as R10-prefered methylation sites and those in the bottom-right corners as R9-prefered methylation sites. To reduce the effect of the variability from individual samples, we investigated overlapped R10-prefered methylation sites of R9 WT versus R10 WT and R9 KO versus R10 KO, as well as overlapped R9-prefered methylation sites of R9 WT versus R10 WT and R9 KO versus R10 KO. We found that there are 58,753 R10-prefered methylation sites and 33,215 R9-preferred methylation sites.
We then annotated these chemistry-biased methylation sites with repeat regions in hg38, and we found that 51.46% of R10-prefered methylation sites (i.e., 30,232 sites) and 40.00% of R9-preferred methylation sites (i.e., 13,253 sites) were from repeat regions. Given that 44.07% of the hg38 human genome is within repeat regions, R10 chemistry seems better than R9 chemistry in detecting methylation levels in repeat regions.
We further checked various repeat subcategories of chemistry-biased overlapped methylation, as shown in Fig. 3 and Table 3. The results suggest that (1) R10 chemistry can detect 9 times more methylation sites from simple repeat regions. This is reasonable because R10.4.1 flowcells have two sensors per nanopore to improve the detection of homopolymer regions; (2) R10 chemistry also improves the methylation detection in SINE, retro-transposon, and satellite regions; (3) R10 and R9 chemistries have similar detection rates of other repeat regions. In summary, R10 chemistry improves methylation detection in repeat regions.
Overlap of chemistry-biased methylation sites found within repeat regions of WT and KO samples. (a) Raw counts of repeat region annotations; (b) Proportion of repeat region types relative to total number of sites.
The effect of chemistry-biased methylation on differential analysis
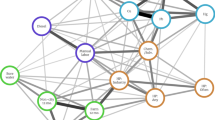
We also wanted to investigate how different ONT chemistries affect the detection of differential methylation positions (DMPs). We conducted three groups of differential analysis: the first group is cross-condition analysis, i.e., we compared R9 WT against R9 KO, and R10 WT against R10 KO, and then obtained the overlap of methylation sites with > 30% methylation percentage difference; the second group is cross-chemistry analysis, i.e., we compared R9 WT against R10 WT, and R9 KO against R10 KO, and generated the overlap of methylation sites with > 30% methylation percentage difference; the third group is cross-condition and cross-chemistry analysis, i.e., we compared R9 WT against R10 KO, and R9 KO against R10 WT, to find the overlap of methylation sites with > 30% methylation percentage difference. We plotted the Venn diagram of the three groups of differential methylated positions in Fig. 4.
Venn Diagram depicting overlap of inconsistent methylation across groups. Yellow: cross-condition (WT/KO) comparison; blue: cross-chemistry and cross-condition comparison; red: cross-chemistry comparison. ∩: overlap; -: comparison.
We found that cross-condition analysis detected 858,012 DMPs, of which 78.88% (676,821 DMPs) overlap with DMPs detected in cross-condition and cross-chemistry analysis. Cross-condition and cross-chemistry analysis detected 298,184 DMPs (i.e., 30.58%) that were not detected by cross-condition analysis. Interestingly, cross-chemistry analysis detected 180,614 DMPs, which is reasonably less than the number of DMPs detected by cross-condition analysis and cross-condition cross-chemistry analysis. Among the DMPs detected by cross-chemistry analysis, 72.85% (131,586 DMPs) were detected by cross-condition and cross-chemistry analysis, while 3.13% (5,661 DMPs) were detected by cross-condition analysis, suggesting that cross-condition and cross-chemistry analysis detected 23 times more methylation patterns caused by chemistry variability rather than by experimental knockouts. Thus, cross-chemistry differential methylation analyses require more attention to those methylation sites biased in different ONT chemistries.
Effect of various ways of calculating coverage and methylation percentages
Although different tools can be used to call methylation from ONT data, modbam2bed developed by ONT is a widely used tool. However, there are various ways to summarize the methylation percentages generated by modbam2bed. To facilitate methylation analysis using modbam2bed in future, we compared three different ways of calculating coverage and methylation percentage described in the “Methods” section (i.e., modbam2bed-calculation, method-1 calculation, and method-2 calculation), and investigated their effect on methylation analysis.
We first calculated the Pearson correlation coefficient for the modbam2bed-calculation and the method-1 calculation for the pairwise comparison of the four samples, and the results are shown in Table 2. The method-1 calculation has a higher correlation for both the two WT replicates and the two KO replicates. For example, the correlation of the method-1 calculation is 0.9185 for the two WT replicates (i.e., R10-WT versus R9-WT), higher than the correlation generated by the modbam2bed-calculation (0.9173), as shown in Table 2; Fig. 5a, b. Similarly, the correlation of methylation for the two KO replicates is 0.9194 for the method-1 calculation and 0.9181 for the modbam2bed-calculation (Table 2; Fig. 5d, e). However, when we investigated the correlation of cross-condition and cross-chemistry pairs, we found that the modbam2bed-calculation usually generated higher correlations than the method-1 correlations (Table 2). Since we have already observed that different ONT chemistries (i.e., R10, and R9) usually have more methylation sites with larger methylation differences, a higher correlation of the WT-KO pairs should not be expected. In summary, the method-1 calculation seems more reasonable than the modbam2bed-calculation for methylation analysis.
Density plots of methylation generated by R9 and R10 ONT chemistries and three different methylation calculations. (a–c): Density heatmap for WT replicates; (d–f): Density heatmap for KO replicates; (a, d): Method-1 calculation; (b, e): Modbam2bed-calculation; (c, f): Method-2 calculation.
To understand the methylation difference for the method-1 calculation and the modbam2bed-calculation, we detected the number of overlapped methylation sites of two WT replicates and the two KO replicates with the same parameters of coverage filtering for the two methods. We found that the method-1 calculation generated 34,132,876 overlapped sites in WT and 40,200,383 overlapped sites in the KO, while the modbam2bed-calculation output more overlapped sites (45,553,841 for WT replicates and 48,643,954 for KO replicates). We investigated those methylation sites that are only detected in the modbam2bed output for the WT and KO pairs, i.e., 11,420,965 sites for the WT pair (Fig. 6a, c), and 8,443,571 sites for the KO pair (Fig. 6b, d). We calculated the correlation for the modbam2bed-unique sites and found that the correlations decreased, compared against the correlation with all detected sites of the modbam2bed-calculations: WT correlated at 0.9024 and the KO pair was at 0.8996, indicating that methylation positions of modbam2bed-unique sites provide a less reliable methylation evaluation than the sites detected by the method-1 calculation.
Investigation of modbam2bed-unique methylation sites. (a, c): WT; (b, d): KO; (a, b): Raw counts; (c, d): Log scale.
We also investigated the correlation of methylation data generated by the method-2 calculation. As shown in Fig. 5c, f, the method-2 calculation has a much lower correlation: 0.8592 for the two WT replicates and 0.8541 for the two KO replicates, which are worse than the correlation of the other two methods (Fig. 5a, b, d, e). Thus, the method-2 calculation is not a good solution for methylation investigation in real-world applications.
Discussion
ONT sequencing has become a widely used technique in methylation investigation. As ONT discontinued R9.4.1 flowcells, more methylation data has been generated using R10.4.1 flowcells. To maximize the use of existing methylation data, ONT methylation research will need to use data generated by both R9 and R10 chemistries. Although R10.4.1 flowcells demonstrate the improvement of basecalling errors, it is important to understand the concordance and bias of methylation data generated by the two chemistries. Our analysis suggests that the methylation data generated by the two chemistries has very high concordance, while R10 chemistry can detect more reliable methylation in repeat regions than R9 chemistry. This is reasonable since ONT claimed that R10.4.1 flowcells are designed to improve the sequencing in low-complexity regions. However, tens of thousands of methylation sites that were reliably detected in R9 chemistry failed to be detected in R10 chemistry, suggesting that both chemistries possess sequencing bias. This observation strongly shows that cross-ONT-chemistry methylation studies need more caution to reduce the number of methylation differences caused by various chemistries.
Since modbam2bed offers an easy and efficient solution to detect methylation, it is worth figuring out proper methylation calculations based on modbam2bed outputs. We investigated three methods to calculate coverage and methylation percentages, and we found that the method-1 calculation is better for methylation studies when using modbam2bed outputs.
The conclusion of this study is based on two pairs of replicates sequenced by R10.4.1 flowcells and R9.4.1 flowcells. Thus, there may be some limitations. First, we only used two replicate pairs that originated from the same human cell line. Although the choice of the same cell line reduces cell-type specific methylation bias, more samples might increase the analysis power. Second, we do not compare results against methylation levels detected by methylation microarrays and do only a simple comparison for bisulfite sequencing. This is because (1) existing studies have demonstrated a high concordance of direct ONT methylation detection against bisulfite sequencing data20,23,34,37,38, and thus long-read sequencing has been used independently in various applications of methylation studies; (2) methylation detection via long-read sequencing can reliably identify DNA methylation in repetitive regions (~ 50% of the human genome)39, suggesting long-read sequencing is a better choice than bisulfite sequencing to study DNA methylation. Thus, our conclusion would not be affected without this comparison.
Conclusion
In this study, we investigated methylation concordance and bias caused by two ONT chemistries and various methylation calculation methods. Although both R10 and R9 chemistries have their own bias of methylation detection, we found a high concordance of methylation data generated by different chemistries, and R10 chemistry improved the methylation detection. We also determined a better choice to investigate methylation data generated by modbam2bed. Our investigation offers an applicable practice of how to conduct robust cross-chemistry investigations of ONT methylation data in real-world applications, avoiding false positive methylation differences that are not caused by the factor of interest.
Methods
Sample preparation
The tested samples in this study are DNA extracted from the human colon tumor cell line, HCT116 (WT)40, and its knockout (KO). The KO was generated by Biocytogen utilizing an extreme genome editing version of CRISPR/Cas9 technology to delete a part of exon 5 and exon 6 of inositol polyphosphate multi-kinase (IPMK) in a turnaround time of about 2 weeks. Exons 5 and 6 are necessary for IPMK to be translated into a functional protein, and their deletion leads to KO cells without the expression of IPMK. After clonal selection, the company used a southern blot to confirm the knockout of IPMK and to verify that no random insertions were present.
Then, both WT and KO cell lines were cultured in McCoy’s 5 A medium at 37 °C for 12 days with four passages, while supplemented with 10% Fetal Bovine Serum, 100 U/mL penicillin, and 100 mg/mL streptomycin. The KO cells were further validated by a western blot and a qPCR to check the expression of IPMK. The western blot showed no presence of IPMK proteins, and the qPCR showed only partial transcripts being generated, demonstrating the success of the knockout. To generate robust results, we sequenced two pairs of WT and KO samples.
Please note that (1) the HCT116 colon cancer cell line was chosen because it is extensively used in scientific research and (2) IPMK-knockout cells were used, because our companion study suggested that IPMK deletion substantially affects global DNA methylation levels. Thus, the use of both WT cells and IPMK-KO cells would benefit our investigation: tens of thousands of CpG sites exhibit significant methylation changes after IPMK-KO, but the remaining sites (roughly ~ 27 million CpG sites) maintain similar methylation levels across both WT and IPMK-KO cells. The remaining sites were used to evaluate both the reliability of ONT methylation detection and cross-chemistry bias, while differential sites were used to assess how cross-chemistry bias affects differential pattern analysis.
DNA extraction and library preparation
DNA was extracted from WT and KO cells separately by using the GenElute mammalian genomic DNA Miniprep Kit (Sigma-Aldrich). Cells were lysed and digested by Proteinase K at 70 °C. Then, the cell lysate was poured into a DNA binding column to eliminate excess reagents and prevent contamination. Extracted DNA was eluted in pure water, and its concentration and purity were determined using a NanoDrop (ThermoFisher). As shown in Table 1, all four samples have higher concentration and quality, according to the A260/A280 and A260/A280 ratios. The DNA was stored at -20 °C for further analysis.
The extracted DNA was used for ONT library preparation. For R9.4.1 flowcells, 1.5 ug DNA was used per the protocol of the SQK-LSK110 (Kit 10 Chemistry). DNA was processed with end-repair, adapter ligation, and bead-based isolation. The final prepared library was quantified using a Qubit fluorometer (ThermoFisher) to ensure there was sufficient DNA to sequence. Then, 100-200ng of prepared DNA was loaded onto an R9.4.1 (FLO-PRO002) flow cell for sequencing. The sequencing was conducted on a PromethION 2 Solo (ONT) connected with a GPU A100 server and ran for about 72 h. During sequencing, when there was a significant drop in available pores or sequencing rate, whichever came first, we washed the flowcells per the ONT washing protocol. We reloaded another 100-200ng DNA library to continue sequencing—the washing and sample reloading significantly increased yield.
The library preparation and sequencing on R10.4.1 flowcells were similar to the process above for R9 flowcells. The difference for R10.4.1 sequencing is that: (1) 4 ug DNA and SQK-LSK114 were used for library preparation and (2) 200 ng prepared DNA was loaded to R10.4.1 (FLO-PRO114) flowcells for sequencing.
One pair of WT and KO samples were sequenced using R9.4.1 flowcells, and the other pair was sequenced using R10.4.1 flowcells. The four samples were sequenced with a depth of at least 30X coverage for robust analysis, as shown in Table 1.
Basecalling and alignment
After sequencing, raw ONT files were transferred to a GPU server where ONT’s Dorado basecaller (version 7.2.13 + fba8e8925) was used for super-accuracy rebasecalling with both 5mC and 5hmC calling configuration. 5hmC was called to avoid misclassification of 5hmC to 5mC.
The sequencing data were aligned against the well-studied human reference genome (i.e., hg38) via minimap234 during the time of basecalling. The alignment BAM files were filtered via SAMtools41 to remove low-quality alignments (mapping quality < 20) or shorter alignments (alignment length < 1000 bp). The statistics of the filtered data for each sequencing sample are shown in Table 1, where all samples had > 5 million reads and > 30X coverage.
Methylation summary
After that, modbam2bed29 was used to summarize 5mC calling. Whole-genome methylation profiles were summarized in BED files where each row of a cytosine site was defined by a chromosome, a position, and a strand, as well as the counts of unmodified bases (Ncanon), of modified bases (Nmod), of filtered bases (Nfilt), of reads with an absent modification call (Nno call), of reads with a substitution concerning the reference (\(\:{N}_{\text{sub}})\), of reads with a deletion concerning the reference (\(\:{N}_{\text{del}}\)), and of reads with an alternative modification call (Nalt mod). After that, modbam2bed uses the equation below to calculate coverage and methylation percentage for each site, which was called “modbam2bed-calculation” to distinguish it from other calculation methods.
This coverage calculation includes absent modification support, substitution, and deletion support, and might mislead the coverage filter for methylation analysis. We thus tested the equations below to calculate coverage and methylation percentages, and called it “method-1 calculation”:
Besides the two calculation methods above, we also tested the effect of including absent modification and failing modification probability on the calculation of methylation percentage. We tested the equations below for the calculation of coverage and methylation percentage, and called this method “Method-2 calculation”:
Methylation analysis
To reliably investigate methylation data generated by ONT techniques, we focused only on CpG sites and filtered out methylation sites with coverage < 10. Then, given two methylation profiles, we used the Pearson correlation coefficient (r) to assess the concordance of methylation calling using different ONT techniques or various methylation calculations. Please note that the Pearson correlation coefficient is a metric widely used to assess the relationship between two methylation datasets of replicates, even if generated by various sequencing techniques, including bisulfite sequencing14,23,24,26,28.
The distribution of pairwise comparison profiles was visualized in 2D histograms using the Matplotlib Python library. Also, discordant subsets of methylation sites were annotated against repeat regions in hg38 for further analysis.
Discordance analysis
Besides using the Pearson correlation coefficient, we also estimated the level of discordance between different ONT chemistries within each condition. A site is considered to be discordant if the difference of its methylation within two samples is larger than a threshold (i.e., 15% or 20% or 25%). Since highly methylated sites have larger methylation differences, a site with > 75% methylation level in both chemistries was excluded. We then counted discordant sites and calculated the discordant percentage by dividing the number of discordant sites by that of available sites. The discordant percentage indicates overall variability.
Bisulfite sequencing comparison
We downloaded a bisulfite sequencing data of WT HCT11636 from GEO with the accession number GSM1465024. We converted the whole genome methylation data from the human genome hg19 to hg38 via the LiftOver tool42 and filtered those sites with a minimum coverage of 5. We then compared methylation levels of whole-genomic sites against R10- and R9-methylation data to additionally show the reliability of methylation detected via ONT sequencing.
Data availability
The codes are available on GitHub via https://github.com/qgenlab/R9R10Methyl/. The ONT datasets were deposited in NCBI SRA (BioProject accession: PRJNA1255515), and the datasets will be openly available when the companion study, which investigated differential methylations between WT and IPMK-KO cells, is published.
References
Moore, L. D., Le, T. & Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 38(1), 23–38. https://doi.org/10.1038/npp.2012.112 (2013).
Jones, P. A. & Takai, D. The role of DNA methylation in mammalian epigenetics. Science 293(5532), 1068–1070. https://doi.org/10.1126/science.1063852 (2001).
Bestor, T. H. The DNA methyltransferases of mammals. Hum. Mol. Genet. 9(16), 2395–2402. https://doi.org/10.1093/hmg/9.16.2395 (2000).
Ming, X. et al. Kinetics and mechanisms of mitotic inheritance of DNA methylation and their roles in aging-associated methylome deterioration. Cell. Res. 30(11), 980–996. https://doi.org/10.1038/s41422-020-0359-9 (2020).
Su, Z., Han, L. & Zhao, Z. Conservation and divergence of DNA methylation in eukaryotes. Epigenetics 6(2), 134–140. https://doi.org/10.4161/epi.6.2.13875 (2011).
Wang, Q., Jia, P., Cheng, F. & Zhao, Z. Heterogeneous DNA methylation contributes to tumorigenesis through inducing the loss of coexpression connectivity in colorectal cancer. Genes Chromosomes Cancer 54(2), 110–121. https://doi.org/10.1002/gcc.22224 (2015).
Nishiyama, A. & Nakanishi, M. Navigating the DNA methylation landscape of cancer. Trends Genet. 37(11), 1012–1027. https://doi.org/10.1016/j.tig.2021.05.002 (2021).
Ballestar, E., Sawalha, A. H. & Lu, Q. Clinical value of DNA methylation markers in autoimmune rheumatic diseases. Nat. Rev. Rheumatol. 16(9), 514–524. https://doi.org/10.1038/s41584-020-0470-9 (2020).
Zhang, H., Liu, L. & Li, M. Mini-review of DNA methylation detection techniques and their potential applications in disease diagnosis, prognosis, and treatment. ACS Sens. 9(3), 1089–1103. https://doi.org/10.1021/acssensors.3c02328 (2024).
Heiss, J. A. et al. Battle of epigenetic proportions: Comparing Illumina’s EPIC methylation microarrays and TruSeq targeted bisulfite sequencing. Epigenetics 15(1–2), 174–182. https://doi.org/10.1080/15592294.2019.1656159 (2020).
Li, Y. & Tollefsbol, T. O. DNA methylation detection: Bisulfite genomic sequencing analysis. Methods Mol. Biol. Clifton NJ 791, 11–21. https://doi.org/10.1007/978-1-61779-316-5_2 (2011).
Liu, Y. et al. Bisulfite-free direct detection of 5-methylcytosine and 5-hydroxymethylcytosine at base resolution. Nat. Biotechnol. 37(4), 424–429. https://doi.org/10.1038/s41587-019-0041-2 (2019).
Vaisvila, R. et al. Enzymatic Methyl sequencing detects DNA Methylation at single-base resolution from picograms of DNA. Genome Res. 31(7), 1280–1289 (2021).
Sigurpalsdottir, B. D. et al. A comparison of methods for detecting DNA methylation from long-read sequencing of human genomes. Genome Biol. 25(1), 69. https://doi.org/10.1186/s13059-024-03207-9 (2024).
Zheng, Y. et al. Prediction of genome-wide DNA methylation in repetitive elements. Nucleic Acids Res. 45(15), 8697–8711. https://doi.org/10.1093/nar/gkx587 (2017).
Ardui, S., Ameur, A., Vermeesch, J. R. & Hestand, M. S. Single molecule real-time (SMRT) sequencing comes of age: Applications and utilities for medical diagnostics. Nucleic Acids Res. 46(5), 2159–2168. https://doi.org/10.1093/nar/gky066 (2018).
Giesselmann, P. et al. Analysis of short tandem repeat expansions and their methylation state with nanopore sequencing. Nat. Biotechnol. 37(12), 1478–1481. https://doi.org/10.1038/s41587-019-0293-x (2019).
R10.3: The newest nanopore for high accuracy nanopore sequencing – now available in store. Oxford Nanopore Technologies. January 13. Accessed November 14, 2024. (2020). https://nanoporetech.com/news/news-r103-newest-nanopore-high-accuracy-nanopore-sequencing-now-available-store
Wang, Y., Zhao, Y., Bollas, A., Wang, Y. & Au, K. F. Nanopore sequencing technology, bioinformatics and applications. Nat. Biotechnol. 39(11), 1348–1365. https://doi.org/10.1038/s41587-021-01108-x (2021).
Sakamoto, Y. et al. Long-read whole-genome methylation patterning using enzymatic base conversion and nanopore sequencing. Nucleic Acids Res. 49(14), e81. https://doi.org/10.1093/nar/gkab397 (2021).
Tourancheau, A., Mead, E. A., Zhang, X. S. & Fang, G. Discovering multiple types of DNA methylation from bacteria and Microbiome using nanopore sequencing. Nat. Methods 18(5), 491–498. https://doi.org/10.1038/s41592-021-01109-3 (2021).
Kuschel, L. P. et al. Robust methylation-based classification of brain tumours using nanopore sequencing. Neuropathol. Appl. Neurobiol. 49(1), e12856. https://doi.org/10.1111/nan.12856 (2023).
Simpson, J. T. et al. Detecting DNA cytosine methylation using nanopore sequencing. Nat. Methods 14(4), 407–410. https://doi.org/10.1038/nmeth.4184 (2017).
Liu, Q. et al. Detection of DNA base modifications by deep recurrent neural network on Oxford nanopore sequencing data. Nat. Commun. 10(1), 2449. https://doi.org/10.1038/s41467-019-10168-2 (2019).
Liu, Q., Georgieva, D. C., Egli, D. & Wang, K. NanoMod: A computational tool to detect DNA modifications using nanopore long-read sequencing data. BMC Genom. 20(1), 78. https://doi.org/10.1186/s12864-018-5372-8 (2019).
Ni, P. et al. DeepSignal: Detecting DNA methylation state from nanopore sequencing reads using deep-learning. Bioinformatics 35(22), 4586–4595. https://doi.org/10.1093/bioinformatics/btz276 (2019).
Rand, A. C. et al. Mapping DNA methylation with high-throughput nanopore sequencing. Nat. Methods 14(4), 411–413. https://doi.org/10.1038/nmeth.4189 (2017).
Liu, Y. et al. DNA methylation-calling tools for Oxford nanopore sequencing: A survey and human epigenome-wide evaluation. Genome Biol. 22(1), 295. https://doi.org/10.1186/s13059-021-02510-z (2021).
epi2me-labs/modbam2bed. Published online November 6. Accessed November 14, 2024. (2024). https://github.com/epi2me-labs/modbam2bed
Akalin, A. et al. MethylKit: A comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 13(10), R87. https://doi.org/10.1186/gb-2012-13-10-r87 (2012).
Okae, H. et al. Genome-Wide analysis of DNA methylation dynamics during early human development. PLOS Genet. 10(12), e1004868. https://doi.org/10.1371/journal.pgen.1004868 (2014).
Bock, C. et al. Quantitative comparison of DNA methylation assays for biomarker development and clinical applications. Nat. Biotechnol. 34(7), 726–737. https://doi.org/10.1038/nbt.3605 (2016).
Spainhour, J. C., Lim, H. S., Yi, S. V. & Qiu, P. Correlation patterns between DNA methylation and gene expression in the Cancer genome atlas. Cancer Inf. 18, 1176935119828776. https://doi.org/10.1177/1176935119828776 (2019).
Yuen, Z. W. S. et al. Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing. Nat. Commun. 12(1), 3438. https://doi.org/10.1038/s41467-021-23778-6 (2021).
Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 34(18), 3094–3100. https://doi.org/10.1093/bioinformatics/bty191 (2018).
Blattler, A. et al. Global loss of DNA methylation uncovers intronic enhancers in genes showing expression changes. Genome Biol. 15(9), 469. https://doi.org/10.1186/s13059-014-0469-0 (2014).
Gombert, S. et al. Comparison of methylation estimates obtained via minion nanopore sequencing and Sanger bisulfite sequencing in the TRPA1 promoter region. BMC Med. Genomics 16(1), 257. https://doi.org/10.1186/s12920-023-01694-6 (2023).
López-Catalina, A., Costes, V., Peiró-Pastor, R., Kiefer, H. & González-Recio, O. Oxford nanopore sequencing as an alternative to reduced representation bisulphite sequencing for the identification of CpGs of interest in livestock populations. Livest. Sci. 279, 105377. https://doi.org/10.1016/j.livsci.2023.105377 (2024).
Udine, E. et al. Targeted long-read sequencing to quantify methylation of the C9orf72 repeat expansion. Mol. Neurodegener 19(1), 99. https://doi.org/10.1186/s13024-024-00790-0 (2024).
Brattain, M. G., Fine, W. D., Khaled, F. M., Thompson, J. & Brattain, D. E. Heterogeneity of malignant cells from a human colonic Carcinoma1. Cancer Res. 41(5), 1751–1756 (1981).
Li, H. et al. The sequence alignment/map format and samtools. Bioinformatics 25(16), 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
Hinrichs, A. S. et al. The UCSC genome browser database: Update 2006. Nucleic Acids Res. 34(suppl_1), D590–D598. https://doi.org/10.1093/nar/gkj144 (2006).
Acknowledgements
The authors would like to thank the lab members for their valuable discussion. This research was funded by the National Institute of General Medical Sciences grant number P20GM121325 and a UNLV startup grant to Dr. Qian Liu. The funders had no role in the study design, data collection and analysis, the decision to publish, or the preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Q.L. conceived and supervised the study, R.D. and E.K. developed algorithms, R.D., E.K. and Q.L. conducted data analysis, S.C. and P.G. cultured cells and extracted DNA, R.D. conducted ONT library preparation and sequencing, R.D. and Q.L. wrote the manuscript, E.K., S.C. and P.G. revised the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Doshi, R., Kinnear, E., Chatterjee, S. et al. Reliable investigation of DNA methylation using Oxford nanopore technologies. Sci Rep 15, 15900 (2025). https://doi.org/10.1038/s41598-025-99882-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99882-0