Abstract
Lung cancer is the third most common cancer in the US with a 5-year survival rate of 17%. Non-small cell lung cancer, especially adenocarcinoma, prevails. Therefore, early detection and biomarker discovery are extremely important. This study uses deep learning to find new biomarkers for lung adenocarcinoma. RNA-Seq data from 522 samples, including 506 lung adenocarcinoma patients and 16 healthy controls, were analyzed. DEGs were identified after strict preprocessing, and deep learning algorithms predicted markers. Functional annotation, pathway, and protein interaction analyses elucidated the biological importance of DEGs. Clinical relevance was assessed by correlation with clinical parameters and survival analysis. External validation was carried out using GDAC and GEO datasets. Blood samples from 30 lung adenocarcinoma patients and 30 healthy people were analyzed by real-time PCR to validate the expression levels of key genes. Among 522 participants(506 cases, 16 controls), the mean age was 62.95 ± 15.71 years. Normalized data showed 3,513 DEGs. The deep learning model had a predictive accuracy of 98.44%, Brier score (probability MSE) = 0.0013, and AUC of 1.0. CYP3A7 had the highest effect size. ROC analysis found diagnostic genes A2M, CYP2C9, and SIRPD (Ensembl ID: 128646) with a sensitivity of 0.96. Real-time PCR showed upregulated CYP2C9, KRT14, and PECAM1 and downregulated A2M in patients compared to controls(P < 0.001). Bioinformatics-identified genes are potential markers for early lung adenocarcinoma detection and management. RT-PCR validation shows AI’s effectiveness in identifying biomarkers, enabling prompt treatment to halt disease progression.
Similar content being viewed by others
Introduction
According to the American Cancer Society (ACS), lung cancer remains one of the most widespread and deadliest forms of cancer globally, ranking as the third most common neoplasm in the United States. This is associated with a 5-year overall survival rate of approximately 17%. There are two primary types of lung cancer, encompassing both small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC). In general, 10% to 15% of all cases of pulmonary malignancy are identified as SCLC, while 80% to 85% are classified as NSCLC. The primary subtypes within NSCLC include adenocarcinomas, squamous cell carcinoma, and large cell carcinoma, with lung adenocarcinoma emerging as the predominant histological subtype, characterized by a distressing mortality rate of 80%1.
Acknowledgment of the critical role of early detection and diagnosis in enhancing patient outcomes and reducing lung cancer-related mortalities is widespread. Emphasis is placed on the vital nature of biomarker identification in lung carcinoma, recognizing that early detection and diagnosis significantly impact patient outcomes and mortality rates. Machine learning is a developing field of technology utilized to identify biomarkers in lung adenocarcinoma. ML, falling under the umbrella of artificial intelligence (AI), enables the absorption of information from past data and automatically executes assigned tasks. Deep Learning, a specific subfield within the realm of machine learning, functions similarly to machine learning but sets itself apart through distinct capabilities and approaches. It takes inspiration from the operations of neurons, leading to the formation of artificial neural networks. Within the domain of deep learning, models employ different filters to acquire knowledge and reveal insights from the given data.
Different approaches exist in the literature regarding the use of these techniques. Zhang et al. used the innovative one-class logistic regression (OCLR), an unsupervised machine learning algorithm, to identify lung adenocarcinoma stem cell biomarkers2. Another study in China identified a sixteen-gene biomarker for lung adenocarcinoma using a machine learning method Bhuvaneswari et al.3 investigated the lung adenocarcinoma stem cell biomarker expressions using machine learning approaches. (National C enter for Biotechnology Information. Homo Sapiens Gene LOC730668, 2024)
Despite all these efforts, there is little published data on using deep learning to identify biomarkers, so this study aims to identify novel biomarkers of lung adenocarcinoma using deep learning analysis and also experimentally verify the AI identified key biomarkers using RT-PCR method.
Methods
Data collection
The RNA-Seq data of patients diagnosed with lung adenocarcinoma, along with associated clinical parameters such as gender, tumor stage, TNM classification, and survival outcomes, were acquired from the gdac dataset (https://gdac.broadinstitute.org/). A comprehensive dataset comprising 522 samples and 20,531 genes was obtained. Subsequently, these samples were selected for subsequent RNA sequencing procedures.
Data preprocessing and the identification of DEGs
Deep learning techniques were then employed to discover novel genes, with normalization and filtering recognized as pivotal stages in data analysis. Preprocessing steps such as filtering and normalization were assessed to prepare RNA data for deep learning applications. Initially, duplicate genes and samples were eliminated using filtering code implemented in R programming. Subsequently, 20,531 genes underwent normalization using the Limma package within the R software environment. DEGs were identified based on specific features, including a log fold change |FC|≥ 2 and a P-value < 0.05. All analyses and plot visualizations in this phase were conducted using R software (version 4.1.2). Gene symbols were standardized according to the HGNC nomenclature. In cases where Ensembl identifiers were initially retrieved (e.g., SIRPD.128646), the approved gene symbol (SIRPD) was used consistently throughout the manuscript, with the original Ensembl ID provided here for clarity. Also, Gene expression values were normalized using limma and log-transformed before modeling; these transformations apply to inputs only and are not the target of the Brier score.
Identifying predictive markers
A bioinformatics analysis aimed at predicting DEGs as significant markers in lung adenocarcinoma was conducted using advanced deep-learning algorithms.
Machine learning algorithms
Deep learning, a subset of machine learning, represents a cutting-edge frontier in artificial intelligence, striving to achieve its ultimate objective of mimicking human intelligence. Deep neural networks, a key component of deep learning, have been extensively studied and applied across various domains, yielding impressive outcomes. In contrast to traditional statistical learning, where linear regression typically involves a single node, deep learning encompasses multiple classifiers collaborating.
Deep learning employs numerous neural nodes, forming a neural network, with each classifier node termed a neural unit or perceptron. Notably, deep learning incorporates multiple layers between input and output, each containing potentially hundreds or thousands of neural units. These intermediary layers, known as hidden layers, and their constituent nodes, referred to as hidden nodes, contribute to the network’s ability to uncover complex relationships autonomously. A distinguishing feature of deep neural networks is their capacity to generate complex hypotheses internally, contrasting with the manual formulation required in traditional machine learning classifiers. This inherent capability renders deep neural networks potent for effectively capturing nonlinear relationships. In many applications, deep learning models surpass the performance of shallow machine learning models and conventional data analysis methods.
Implementation and evaluation
Python version 3.7 served as the foundation for implementing machine learning techniques. Utilized packages included Pandas, NumPy, Matplotlib, and Scikit-learn. Model refinement involved optimization with training data and independent evaluation using test data. Train/test ratios with a 70/30 ratio were applied.
Model evaluation was performed using a hold-out validation strategy. The dataset was randomly divided into a training set (70%) and an independent test set (30%), with no overlap between samples. All performance metrics were calculated exclusively on the independent test set.
To implement the deep learning framework, we constructed a feed-forward neural network with three fully connected hidden layers containing 128, 64, and 32 neurons, respectively. Each hidden layer employed the rectified linear unit (ReLU) activation function, while the output layer used a sigmoid activation function to produce class probabilities. Model training was conducted using the Adam optimizer (learning rate 0.001) with binary cross-entropy loss. Dropout regularization was applied to the hidden layers, and early stopping was adopted based on validation loss to prevent overfitting. Hyperparameters, including the number of neurons, dropout rate, and learning rate, were tuned manually in an iterative fashion to maximize predictive performance on the validation set.
Performance was assessed with standard evaluation metrics including AUC, accuracy, F1 score, R² score, confusion matrix, and the Brier score. Accuracy measures the precision of classification by assessing true positives and negatives, F1 score accounts for imbalanced data, AUC evaluates discriminative capacity, and the confusion matrix summarizes classification outcomes (TN, TP, FN, FP). The Brier score corresponds to the mean squared error between predicted probabilities and true class labels, providing a measure of both accuracy and calibration. For a test set of size N, with predicted probabilities Pi∈[0,1] and true labels Yi∈{0,1}, the Brier score is defined as:
Brier Score (BS) = 1/N * Σ (Pi - Yi)2
Lower values indicate better probabilistic accuracy and calibration. In our study, all metrics, including the Brier score (reported as 0.0013), were calculated on the independent held-out test set obtained by a 70/30 train-test split, with no overlap between training and test samples.
Functional and pathway enrichment analyses
The analysis involved annotating and visualizing functional enrichment and critical pathways of the DEGs signature using the cluster Profiler package in R, with a stringent significance threshold of p-adjusted < 0.05. In addition, two common databases, Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG), were utilized as reference databases for functional enrichment. All figures represent original enrichment plots generated using the clusterProfiler R package, and no KEGG pathway maps were reproduced4,5.
Correlation between DEGs and demographic/clinical data
We examined 55 DEGs to investigate their connection with demographic and clinical factors such as age, tumor size, lymph node involvement, distant metastasis, and stage. This investigation involved utilizing correlation matrix and Spearman correlation techniques within the R programming environment, leveraging the ggcorrplot package and cor function.
Identifying prognostic markers
The Survmier and ggplot2 R packages were utilized to generate Kaplan-Meier survival curves and Cox analyses for the top-selected DEGs. All data met screening criteria, with a cut-off threshold of -1 < HR < 1 and P < 0.05. The identified genes are categorized as “prognostic genes”.
Combine ROC curve
A generalized linear model was utilized in conjunction with combined ROC curve analysis to identify diagnostic biomarkers. Multiple metrics, including sensitivity, specificity, cut-off value, positive predictive value, negative predictive value, and AUC curve, were assessed to gauge the discriminative capacity of individual or combined biomarkers. The entire process was executed using the combioROC package in R. The combioROC package provides sensitivity, specificity, AUC, positive predictive value (PPV), and negative predictive value (NPV) for biomarker combinations.
Validation of biomarkers gene expression
The expression levels of candidate genes in lung adenocarcinoma patients were conducted using data from the Global Data Assembly Centers (GDAC) at the Broad Institute (https://gdac.broadinstitute.org/) and the Gene Expression Omnibus (GEO) dataset (GSE19804, GSE75037, GSE131907, GSE37745, and GSE166720). Specifically, data from patients with lung adenocarcinoma were retrieved from these sources for validation purposes, and necessary preprocessing steps were carried out accordingly.
Experimental verification of biomarkers through RT-PCR method
Study samples
30 cases of lung adenocarcinoma and 30 healthy individuals entered the study. The cases were all in stage 1 to 3 of disease without any distant metastasis and no medical therapy including chemical or surgical had been yet initiated for them. Cancer diagnosis confirmed through pathological investigation. Also, cases and control groups were matched for age and sex. A consent form was signed by all individuals enrolled the study.
RNA extraction
5 milliliters (ml) of peripheral blood from all individuals were collected and the RNA extraction process was initiated immediately using the RNeasy Midi Kit (Qiagen Cat no.75144) based on the product protocols.
cDNA synthesis
cDNA synthesis was done using the Viva 2-step RT-PCR Kit (Cat No. RTPL12). The specific primers for biomarkers were designated and synthetized using the AlleleID6 software.
Real-time PCR using the HotTaq EvaGreen qPCR mix
Tables 1 and 2 show the parameters and their usage amounts in the final Real-time PCR reaction. (Tables 1 and 2.)
The kit of Sinacolon manufacture (Cat No. BT11101) was used, which had EvaGreen, as the fluorescent dye, becoming detectable when binding to double-stranded DNA. The master mix kit contained all necessary components for Real-time RT-PCR reaction as follows: 2 µL of Template sequence, 4 µL of Master mix, primers with optimal concentration and deionized distilled water (dH2O), which was added to bring the total reaction mixture to 20 µL volume. All stages are performed according to kit-specific instructions. Results were interpreted based on the amplification and melt curves.
Results
Patient demographics and clinical characteristics
To provide an overview of the study population and establish the clinical context for subsequent analyses, the demographic and clinical characteristics of the enrolled participants are summarized in Table 3. This study involved 522 participants (53.6% females), including 506 cases and 16 healthy controls with a mean age of 62.95 ± 15.71 years at initial pathologic diagnosis. The racial distribution was predominantly White (75.3%). The average pack-years smoked were 28.48 ± 29.66. Moreover, neoplasm cancer status revealed diverse stages, with most patients at Stage 1 (53.6%). Tumor invasion depth also varied, with T2 being the most common (53.8%). In addition, lymph node involvement was observed in 82.9%, and most samples were free of metastasis (67.6%, M0). Furthermore, the overall mean days to the last follow-up were 337.73 ± 737.85. It indicated a 24.5% mortality rate.
Data preprocessing and the identification of DEGs
As an initial analytical step, the RNA-seq dataset was systematically preprocessed to identify differentially expressed genes associated with lung adenocarcinoma. The dataset, obtained from GDAC, encompassed 522 samples and 20,531 genes. Key features were identified by extracting those with a correlation coefficient > 0.5. Finally, 3513 genes were normalized and identified based on our criteria (a standard log and P < 0.05). These were visualized using volcano plots and PCA analysis (Figs. 1, 2, 3 and 4).
Visualization of the dataset, illustrating preprocessing steps and the distribution of differentially expressed genes (DEGs) after normalization and selection criteria application.
Visualization of the dataset, illustrating preprocessing steps and the distribution of differentially expressed genes (DEGs) after normalization and selection criteria application.
Visualization of the dataset, illustrating preprocessing steps and the distribution of differentially expressed genes (DEGs) after normalization and selection criteria application.
Visualization of the dataset, illustrating preprocessing steps and the distribution of differentially expressed genes (DEGs) after normalization and selection criteria application.
Identifying predictive markers
Model performance was evaluated to assess the predictive accuracy of the deep learning framework and to identify key genes associated with lung adenocarcinoma. The deep learning model exhibited excellent classification performance, with a Brier score of 0.0013 (on test set probabilities), an AUC of 1.0, and an overall accuracy of 98.44%. To further assess the model fit in terms of regression-style prediction of expression-derived features, we also calculated the Root Mean Squared Error (RMSE = 0.0363) and the coefficient of determination (R² = 0.9856), which confirmed the consistency of the model’s predictive capacity. After evaluation, the deep learning framework identified 20 key genes, as presented in Table 4. Among these, CYP3A7 exhibited the highest effect size with a value of 1, followed by LDLR (0.946212), KCNA4 (0.920194), SSTR4 (0.917751), and CD101 (0.908966).
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses
Functional enrichment analyses were performed to characterize the biological processes and pathways associated with the identified differentially expressed genes. Genes for KEGG pathway analysis and gene ontology were analyzed using R software. These genes were found to influence various molecular and cellular pathways and several congenital and acquired conditions (Figs. 5, 6, 7, 8, 9, 10, 11 and 12). A significance threshold of P < 0.05 was applied.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
Gene ontology analyses of DEGs, functional annotation, and pathway enrichment analyses.
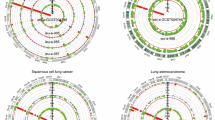
Investigation of the correlations of demographic/clinical with disease
Correlation analysis was conducted to examine the relationships between candidate genes and key demographic and clinical variables. The relationships between candidate genes and demographic/clinical variables are illustrated in Figs. 13, 14, 15 and 16. Data analysis revealed a significant positive correlation between tumor invasion depth and stage (r = 0.9), tumor invasion and metastasis (r = 0.4), and metastasis and stage (r = 0.6), as well as a negative correlation between metastasis and race (r = − 0.4). Correlation values below 0.3 were considered weak, between 0.3 and 0.6 moderate, and above 0.6 substantial.
Investigation of the correlations of demographic/clinical with Disease.
Investigation of the correlations of demographic/clinical with Disease.
Investigation of the correlations of demographic/clinical with Disease.
Investigation of the correlations of demographic/clinical with Disease.
Identifying diagnostic markers
Diagnostic performance was evaluated to identify gene combinations with optimal discriminative ability for lung adenocarcinoma. Tables 5 and 6 illustrate the sensitivity and specificity of different combinations of genes in diagnosing lung adenocarcinoma. The combination of A2M.2, CYP2C9.1559 and SIRPD (Ensembl ID: 128646) attained the highest sensitivity with the best AIC value in GLM analysis. Table 5 represents cut-off, sensitivity, specificity, AUC, negative and positive predictive values, and GLM analysis of 16 combinations with sensitivity > 90%.
Identifying prognostic markers
Prognostic relevance was assessed using survival-based analyses to identify genes associated with patient outcomes in lung adenocarcinoma. Our cox regression model identified genes CYP2C9, GATA1, GPBAR1, KCNV1, KRT24, LOC286467, NKX2, and SIRPD as the prognostic markers. While employing deep learning, gene LOC730668 was recognized as a prognostic marker. Besides, based on weight correlation, ACVRL1.94, C10orf67.256815, C14orf132.56967, CDH5.1003, LIMS2.55679, PECAM1.5175, PTPN21.11099, STX11.8676, and WWC2.80014 are potential prognostic markers.
Validation
External validation was performed to assess the robustness and reproducibility of the identified candidate genes across independent datasets. Candidate genes were validated in 80% of datasets.
Analysis of identified biomarkers using RT-PCR
Positivity rates of biomarkers
Biomarker positivity rates were compared between lung adenocarcinoma patients and healthy controls to evaluate their diagnostic potential in peripheral blood. The positivity rates of biomarkers were assessed in the peripheral blood of cancer patients and healthy individuals (Fig. 17A–D). CYP2C9 was positive in 20 out of 30 cancer patients, compared to 5 out of 30 healthy individuals, with a significant difference between the groups (P < 0.001).
(A–D) Sensitivity and specificity of gene combinations for lung adenocarcinoma diagnosis. The A2M.2, CYP2C9.1559, and SIRPD (Ensembl ID: 128646) combination showed the highest sensitivity with the best AIC in GLM analysis.
A2M showed a positive result in 8 out of 30 cancer patients, compared to 25 out of 30 healthy individuals, showing a significant difference between groups (P < 0.001).
KRT14 was positive in 25 out of 30 cancer patients and 6 out of 30 healthy individuals, with a statistically significant difference (P < 0.001).
PECAM1 was positive in 26 out of 30 in cancer group, and 7 out of 30 in healthy group, exhibiting a significant difference observed (P < 0.001).
Analysis of expression levels of biomarkers (fold changes)
Quantitative expression analysis was performed to compare biomarker expression levels between lung adenocarcinoma patients and healthy controls. The relative expression levels of biomarkers between patients and healthy individuals were measured using the ΔΔCt method for VEGF-mRNA and CK20-mRNA. The results are presented in Fig. 18.
The fold change of CYP2C9, A2M, KRT14, and PECAM1 in two groups of lung adenocarcinoma patients and healthy individuals showed a statistically significant difference between groups (P < 0.001). CYP2C9 expression was 4.1 times higher in cancer patients compared to healthy individuals. A2M expression in cancer patients was 0.5 times or half thet of healthy individuals. KRT14 expression was 8.1 times higher in cancer patients than in healthy individuals. PECAM1 expression was 2.2 times higher in cancer patients compared to healthy individuals.
Discussion
In the current study, bioinformatic analysis and deep learning approaches were combined to identify novel biomarkers of lung adenocarcinoma. Our data illustrated the potential value of the combination of A2M, CYP2C9, KCNV1, KRT24, and SIRPD as a diagnostic marker in lung adenocarcinoma. This combination of biomarkers significantly enhances the accuracy of early detection and diagnosis, providing a robust tool for identifying lung adenocarcinoma at its nascent stages. The bioinformatics analyses provide inferential outcomes of numerous statistical computations based on chip or sequencing data. Although the obtained results are somewhat reliable, they still require experimental confirmation. Beyond biomarker discovery, our study also contributes methodologically by integrating a deep learning framework into the conventional DEG–TCGA/GEO analysis pipeline. In contrast to traditional models such as logistic regression, random forest, and SVM, our neural network achieved superior discriminative performance, as reflected by higher accuracy, AUC, and calibration metrics. This highlights that deep learning not only improves the robustness of predictive modeling but also enhances the translational relevance of biomarker identification in lung adenocarcinoma. By combining computational modeling with RT-PCR validation, our workflow offers an original and integrative strategy that distinguishes this study from previously published bioinformatics-based approaches. Recent studies have increasingly applied artificial intelligence and multi-omics integration to cancer biomarker discovery. For example, Yu et al.6 introduced a deep-learning model based on immune-related features such as M1 macrophages and APOBEC3G to predict immunotherapy efficacy, while Yu et al.7 developed a multimodal framework combining pathology images, transcriptomics, and genomic data for papillary thyroid carcinoma prognosis. Similarly, Ouyang et al.8 identified cuproptosis-related lncRNA signatures in lymphoma using machine learning and transformer-based models, and Zhang et al.9 highlighted the emerging role of RNA ac⁴C modifications in tumor biology. In another recent study, transformer-based machine learning models were applied to integrate TCGA and GEO transcriptomic data for identifying prognostic lncRNA signatures associated with regulated cell death pathways in diffuse large B-cell lymphoma, demonstrating the growing application of advanced deep learning architectures in cancer biomarker discovery8. Moreover, recent comprehensive reviews have emphasized the emerging importance of RNA modifications, including N4-acetylcytidine (ac⁴C), in tumor biology and biomarker research, further expanding the landscape of RNA-based cancer biomarkers9. Advanced integrative computational frameworks combining complex biological data with data-driven analytical strategies have also been applied across diverse disease contexts, demonstrating the versatility and scalability of modern modeling approaches in biomedical research10. Compared with these studies, our work is innovative in its focus on early-stage lung adenocarcinoma and the integration of a multi-hidden-layer deep-learning model with external dataset validation and RT-PCR confirmation of peripheral blood biomarkers. This approach not only improves diagnostic and prognostic accuracy beyond conventional machine learning models but also emphasizes clinical applicability through minimally invasive testing.
The results of bioinformatics analysis helped us to identify key genes, which we could validate their expression level in vitro through RT-PCR for further confirmation. to replicate and experimentally verify. After experimental verification through RT-PCR, we identified four biomarkers including CYP2C9, A2M, KRT14 and PECAM1, that were significantly differentially expressed in the plasma of lung adenocarcinoma patients when compared to healthy individulas.
Furthermore, the results of the analysis revealed the prognostic role of LOC730668, CYP2C9, GATA1, GPBAR1, KCNV1, KRT2, LOC286467, NKX2.5, SIRPD, ACVRL1, ACADL, C14orf132, CDH5, LIMS2, PECAM1, PTPN21, STX11, and WWC2 biomarkers. These biomarkers collectively offer insights into disease progression and patient outcomes, helping to tailor more effective and personalized treatment plans.
Recent genomic studies suggest that LOC730668, a pseudogene located on chromosome 22q13.31, may play a regulatory role in cancer-related biological processes, particularly through epigenetic mechanisms. Although LOC730668 does not encode a functional protein, emerging evidence indicates that genetic loci associated with this gene may influence DNA methylation patterns and gene expression regulation, which are critical contributors to tumorigenesis. In the present study, LOC730668 was identified as a prognostic marker in lung adenocarcinoma, suggesting that its potential involvement in epigenetic regulation may be relevant to disease progression and patient outcomes. Further functional investigations are warranted to clarify the mechanistic role of LOC730668 in lung adenocarcinoma.
Another diagnostic biomarker is A2M, one of the large glycoproteins (720 kDa) present in bodily fluids, which primarily functions as a protease inhibitor. A2M plays diverse and complex roles by binding to different hormones and regulating their activity; several studies on the association between A2M and cancer have indicated that A2M might play an important anti-tumor role. For instance, Qiu et al. investigated the possible role of A2M-AS1 in pancreatic cancer, showing that its expression levels decreased in pancreatic cancer and were positively correlated with the survival time of patients. Moreover, A2M-AS1 can also be used as a diagnostic and prognostic biomarker for pancreatic cancer. Serum A2M levels have been also been inversely correlated with PSA levels in men with advanced prostate cancer11. This multifaceted role of A2M in various biological processes highlights its significance and potential in both clinical diagnostics and therapeutic applications.
Cytochrome P450 2C9 (CYP2C9) has been implicated as a cancer-related biomarker due to its role in drug metabolism and regulation of tumor-associated biological processes. Extrahepatic expression of CYP2C9 has been reported in neoplastic tissues, and the enzyme has been linked to vascular endothelial cell proliferation, suggesting a role in tumor angiogenesis12.
CYP2C9 functions as a major epoxygenase involved in endothelial cell migration and angiogenesis under hypoxic conditions. Altered metabolism of arachidonic acid by CYP2C9 variants has been shown to influence epoxyeicosatrienoic acid (EET) production, thereby contributing to non-small cell lung cancer (NSCLC) progression through modulation of angiogenic and hypoxia-related pathways13. Genetic variants of CYP2C9 have been associated with lung cancer susceptibility, disease progression, and therapeutic response, highlighting its potential prognostic relevance14,15,16. Experimental studies further indicate that modulation of CYP2C9 expression affects cell-cycle progression and proliferation, supporting its functional involvement in tumor growth and treatment response17.
This suggests that pre-therapeutic genotyping could be advantageous for planning treatment strategies for affected patients18. Differential expression of CYP2C9 has also been reported across tumor subtypes, underscoring its context-dependent role in cancer biology19.
Research on the KCNV1 gene has indicated its potential role as a biomarker for cancer, particularly in the context of gene expression and tumor progression. While direct studies on KCNV1 as a biomarker in cancer are limited, references suggest its involvement in cancer-related pathways and potential utility in cancer diagnostics and prognosis.
KRT14 (Keratin 14) has been investigated as a potential cancer-related biomarker, as alterations in keratin expression are commonly associated with malignant transformation in epithelial tissues. Changes in KRT14 expression have been reported among the molecular alterations detected during early stages of tumor development20. Overexpression of KRT14 has been observed in esophageal adenocarcinoma, supporting its potential diagnostic relevance in epithelial malignancies. Transcriptomic analyses have further demonstrated alterations in KRT14-related regulatory pathways during disease progression21. KRT14 expression has also been detected in biological fluids of cancer patients and has been associated with prognostic indicators in transitional cell carcinoma of the bladder22. In lung adenocarcinoma, KRT14 expression has been linked to patient outcomes, potentially through interactions with FSTL3-related signaling pathways23. Experimental evidence from lung cancer models indicates that KRT14-high tumor cell subpopulations exhibit enhanced metastatic potential by resisting anoikis via upregulation of Gastrokine 1 (Gkn1), which correlates with poor survival24. Collectively, these findings support a mechanistic and clinical role for KRT14 as a diagnostic and prognostic biomarker, particularly in the context of lung adenocarcinoma.
Moreover, in cutaneous squamous cell carcinoma (cSCC), decreased degradation of KRT14, facilitated by the overexpression of transglutaminase 3 (TGM3), results in the dysregulation of epidermal differentiation in cSCC cells. TGM3 inhibits tumor progression through the p13k/Akt signaling pathway, which mediates the degradation of KRT1425.
PECAM-1, also known as platelet-endothelial cell adhesion molecule 1 (PECAM-1), is a member of the immunoglobulin superfamily and functions as an adhesion receptor expressed on endothelial cells, platelets, and leukocytes. CD31 plays an important role in regulating endothelial cell functions and the tumor microenvironment, contributing to angiogenesis and lymphangiogenesis, which are key determinants of tumor growth and metastasis26,27. Studies utilizing mouse models have demonstrated that PECAM-1 knockdown inhibits tumor development and metastasis, further establishing its role in tumor biology27.
Evidence indicates that PECAM-1 (CD31) is differentially expressed in non-small cell lung cancer (NSCLC) and is associated with tumor angiogenesis and disease progression26,27,28. In non-small cell lung cancer (NSCLC) patients, circulating endothelial cells with surface CD31 have revealed a great predictive value for the efficacy of anti-angiogenic therapeutic, such as anlotinib. Anlotinib downregulates CD31 through enhancing the PI3K/AKT signaling pathway29. CD31 overexpression has been linked to enhanced angiogenic activity through its association with δ-catenin, a regulator of tumor invasion and migration in NSCLC30. However, Cao et al. found high PECAM-1 expression was associated with a survival advantage suggests that it may serve as a protective prognostic factor maybe through regulating T-cell mediated immune responses31. CD31 functions as a multifunctional receptor involved in signaling pathways that regulate cell migration and tumor-related cellular behavior. Its dysregulated expression has been associated with aggressive pathological features in cancer, underscoring the context-dependent role of CD31 in tumor progression and its potential therapeutic relevance32,33.
In our study, CYP2C9 emerged as both a diagnostic and prognostic biomarker, consistent with its role as a cytochrome P450 epoxygenase that metabolizes arachidonic acid into epoxyeicosatrienoic acids (EETs). EETs are known to promote angiogenesis, endothelial cell proliferation, and tumor progression across several cancer types34. Moreover, recent evidence indicates that functional variants of CYP2C9 (CYP2C92/*3) directly modulate tumorigenesis by altering EET production, which may influence cancer development and vascularization in lung tissues35. These findings resonate with our data, suggesting that the elevated expression of CYP2C9 in lung adenocarcinoma may reflect enhanced angiogenic and metabolic activity that contributes to tumor growth and progression.
A2M, identified in our study as a key diagnostic marker, is a broad-spectrum protease inhibitor implicated in immune modulation and tumor–stromal interactions. Recent proteogenomic analyses have shown that downregulation of ADAMTS1, which interacts with A2M, contributes to tumorigenesis and alters the immune microenvironment in lung adenocarcinoma36. In parallel, a pan-cancer analytical framework established that the LINC00987/A2M axis correlates with immune infiltration, hypoxia, and stemness characteristics, suggesting that A2M may serve not only as a biomarker but also as a functional regulator of the immune contexture in LUAD37. These observations align with our data and support a mechanistic hypothesis: altered A2M expression in lung adenocarcinoma may influence immune cell recruitment and microenvironmental remodeling, thereby linking biomarker potential with biological relevance.
KRT14 emerged in our study as a strong diagnostic biomarker and its potential mechanistic link with invasion and metastasis is supported by recent evidence. Prior research in lung cancer models demonstrated that subpopulations of KRT14-high cells possess elevated metastatic capability by resisting anoikis via upregulation of Gastrokine-1 (Gkn1)38. Additionally, broader analyses across multiple tumor types revealed that KRT14 overexpression is associated with poorer survival and may be involved in epithelial–mesenchymal transition (EMT) and collective cell invasion39. These observations align with our findings of upregulated KRT14 in lung adenocarcinoma samples and suggest a mechanistic hypothesis: elevated KRT14 expression in LUAD may facilitate invasive and metastatic behavior by enhancing anoikis resistance and EMT-related processes.
Looking ahead, the biomarkers identified in our study, namely A2M, CYP2C9, and KRT14, may extend beyond diagnostic and prognostic roles to serve as targets or modulators in RNA-based therapeutic strategies. Emerging advances in RNA therapeutics, including mRNA cancer vaccines and microRNA-focused treatments, offer promising avenues for leveraging such biomarkers for therapy. For instance, recent reviews highlight that mRNA cancer vaccines can reprogram immunity by targeting tumor-specific transcripts, and over 60 candidates are currently in clinical development40. In parallel, microRNAs are increasingly recognized as both potential therapeutic agents and targets, capable of modulating oncogenic pathways in vivo41. Conceivably, our identified markers, if mechanistically linked to tumor progression pathways, could be incorporated into personalized RNA-directed interventions, such as RNA vaccines encoding neoantigens or miRNA-based inhibitory/activating constructs, thereby bridging biomarker discovery with translational oncology.
Recent evidence suggests that dysregulated RNA expression profiles in tumors might not only serve as effective diagnostic and prognostic biomarkers but also provide actionable targets for RNA-based therapeutics. For example, a pan-cancer analysis identified tumor antigens and immune subtypes as potential candidates for mRNA vaccine development42, and a recent review has outlined the emerging progress and mechanistic basis of tumor-targeted mRNA vaccines43. These findings complement our current work by positioning RNA biomarkers in lung adenocarcinoma within a broader translational framework that includes potential RNA-directed therapeutic strategies.
The SIRP family of proteins, including SIRPD, plays significant roles in immune response modulation, which is crucial for cancer progression and treatment. Although specific research articles directly linking SIRPD as a biomarker for cancer are limited, the protein family it belongs to, particularly SIRPα, has been studied extensively in cancer immunotherapy. SIRPα interacts with CD47, a “don’t eat me” signal on cancer cells, allowing these cells to evade phagocytosis by macrophages. Blocking this interaction has been shown to enhance the immune system’s ability to attack cancer cells, making it a target for therapeutic interventions. Each biomarker exhibited a significant difference in positive rates between the patient and healthy groups, highlighting their potential relevance in disease detection.
The dataset used for model development comprised 522 samples, including 506 lung adenocarcinoma cases and 16 healthy controls, reflecting a substantial class imbalance that is inherent to the structure of the publicly available dataset.
To mitigate and monitor the potential bias introduced by this imbalance, we employed evaluation metrics that are recommended for imbalanced classification tasks, including the area under the receiver operating characteristic curve (AUC) and the F1 score, rather than relying solely on overall accuracy. These metrics provide a more informative assessment of model performance under skewed class distributions.
Although no resampling techniques (such as oversampling or undersampling) were applied, the consistently high AUC and F1 score values observed on the independent test set suggest that the model performance was not driven exclusively by the majority class. Nevertheless, future studies using larger and more balanced cohorts are warranted to further validate the robustness and generalizability of the proposed biomarkers.
Conclusion
We concluded that the combination of A2M, CYP2C9, KCNV1, KRT24, and SIRPD has significant potential as a diagnostic marker in lung adenocarcinoma. which was confirmed by RT-PCR method. This combination of biomarkers significantly enhances the accuracy of early detection and diagnosis, providing a robust tool for identifying lung adenocarcinoma at its nascent stages. Early and precise diagnosis is critical for improving patient outcomes, as it allows for timely intervention and tailored treatment strategies.
Further research is needed to fully emphasize this potential. Continued studies should focus on validating these biomarkers in larger, more diverse populations and exploring their mechanisms in lung adenocarcinoma progression. This will help solidify their role in clinical practice and maximize their impact on patient care. Bioinformatics methods have the potential to identify key biomarkers involved in disease pathogenesis and serve as a valuable tool to recognize key genes and biomarkers before confirmation of these genes through RT-PCR using patients’ samples.
Data availability
The data that support the findings of this study are available upon request through corresponding authors.
Abbreviations
- A2M:
-
Alpha-2-macroglobulin
- ACS:
-
American Cancer Society
- AI:
-
Artificial intelligence
- AUC:
-
Area under curve
- cDNA:
-
Complementary DNA
- CT:
-
Computed tomography
- DEGs:
-
Differentially expressed genes
- EAC:
-
Esophageal adenocarcinoma
- ER:
-
Endoplasmic reticulum
- FC:
-
Fold change
- GDAC:
-
Global data assembly centers
- GEO:
-
Gene expression omnibus
- GLM:
-
Generalized linear model
- GO:
-
Gene ontology
- HR:
-
Hazard ratio
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- KRT14:
-
Keratin 14
- LOC730668:
-
Long non-coding RNA 730668
- MSE:
-
Mean squared error
- miRNA:
-
Micro RNA
- NSCLC:
-
Non-small cell lung cancer
- OCLR:
-
One-class logistic regression
- PCR:
-
Polymerase chain reaction
- PECAM1:
-
Platelet-endothelial cell adhesion molecule 1
- PPI:
-
Protein–protein interaction
- PSA:
-
Prostate-specific antigen
- R2:
-
Coefficient of determination
- RNA-Seq:
-
RNA sequencing
- ROC:
-
Receiver operating characteristic
- RT-PCR:
-
Real-time polymerase chain reaction
- SCLC:
-
Small cell lung cancer
- SIRPD:
-
Signal-regulatory protein delta
- SPSS:
-
Statistical package for the social sciences
- STRING:
-
Search tool for the retrieval of interacting genes/proteins
- TCGA:
-
The cancer genome atlas
- TNM:
-
Tumor, node, metastasis
References
Li, Y. et al. A large cohort study identifying a novel prognosis prediction model for lung adenocarcinoma through machine learning strategies. BMC Cancer 19, 1–14 (2019).
Zhang, Y. et al. mRNAsi index: machine learning in mining lung adenocarcinoma stem cell biomarkers. Genes (Basel) 11 (3), 257 (2020).
National Center for Biotechnology Information. Homo sapiens gene LOC730668. (2024).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51 (D1), D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Kanehisa, M. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28 (1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Yu, Y. et al. Unraveling the role of M1 macrophage and CXCL9 in predicting immune checkpoint inhibitor efficacy through multicohort analysis and single-cell RNA sequencing. MedComm 5 (3), e471. https://doi.org/10.1002/mco2.471 (2024).
Yu, Y. et al. Artificial intelligence-based multi-modal multi-tasks analysis reveals tumor molecular heterogeneity, predicts preoperative lymph node metastasis and prognosis in papillary thyroid carcinoma: a retrospective study. Int. J. Surg. 111 (1), 839–856. https://doi.org/10.1097/JS9.0000000000001875 (2025).
Ouyang, W., Lai, Z., Huang, H. & Ling, L. Machine learning-based identification of cuproptosis-related lncRNA biomarkers in diffuse large B-cell lymphoma. Cell Biol. Toxicol. 41 (1), 72. https://doi.org/10.1007/s10565-025-10030-w (2025).
Ouyang, W. et al. RNA ac4C modification in cancer: Unraveling multifaceted roles and promising therapeutic horizons. Cancer Lett. 601, 217159. https://doi.org/10.1016/j.canlet.2024.217159 (2024).
Ouyang, W. et al. Traditional Chinese medicine in cerebral infarction: Integrative strategies and future directions. Phytomedicine 143, 156841. https://doi.org/10.1016/j.phymed.2025.156841 (2025).
Kostova, M. B. et al. Degrade circulating peptide hormones. Prostate 78 (11), 819–829. https://doi.org/10.1002/pros.23539.PSA-alpha-2-macroglobulin (2021).
Schmelzle, M. et al. Esophageal cancer proliferation is mediated by cytochrome P450 2C9 (CYP2C9). Prostaglandins Other Lipid Mediat. 94 (1–2), 25–33 (2011).
Myung, S. J., Yoon, J. & Yu, S. J. STAT3 & Cytochrome P450 2C9: A novel signaling pathway in liver cancer stem cells. Biomed. Pharmacother. 66 (8), 612–616. https://doi.org/10.1016/j.biopha.2012.08.011 (2012).
Garcia-Martin, E. et al. Influence of cytochrome P450 CYP2C9 genotypes in lung cancer risk. Cancer Lett. 180 (1), 41–46 (2002).
Yadav, S. S. et al. Association of polymorphism in cytochrome P450 2C9 with susceptibility to head and neck cancer and treatment outcome. Appl. Transl. Genom. 3 (1), 8–13 (2014).
Yadav, S. S. et al. Applied & translational genomics association of polymorphism in cytochrome P450 2C9 with susceptibility to head and neck cancer and treatment outcome ☆. Appl. Trans. Genom. 3, 8–13 (2014).
Narjoz, C. et al. Genomic consequences of cytochrome P450 2C9 overexpression in human hepatoma cells. Chem. Res. Toxicol. 22 (5), 779–787 (2009).
Paul, S., Chaudhuri, T., Pant, M. C., Parmar, D. & Srivastava, K. Association of cytochrome P450 2C9 polymorphism with locally advanced head and neck squamous cell carcinoma and response to concurrent cisplatin-based radical chemoradiation. South Asian J. Cancer 3 (03), 154–158 (2014).
Jiang, Z. et al. CYP2C9 inhibits the invasion and migration of esophageal squamous cell carcinoma via downregulation of HDAC. Mol. Cell. Biochem. 476 (5), 2011–2020. https://doi.org/10.1007/s11010-021-04050-3 (2021).
Das, S., Devireddy, R. & Gartia, M. R. Surface plasmon resonance (SPR) sensor for cancer biomarker detection. Biosensors 13 (3), 396 (2023).
Fu, Y. et al. Transcriptomic changes and gene fusions during the progression from Barrett’s esophagus to esophageal adenocarcinoma. Biomark. Res. 12 (1), 78 (2024).
Rasti, A. et al. Evaluation of IGF2, KRT14, and KRT20 as urinary biomarkers in patients with bladder cancer. Rep. Biochem. Mol. Biol. 11 (4), 710 (2023).
Meng, X. et al. FSTL3 is associated with prognosis and immune cell infiltration in lung adenocarcinoma. J. Cancer Res. Clin. Oncol. 150 (1), 17 (2024).
Yao, S. et al. Keratin 14-high subpopulation mediates lung cancer metastasis potentially through Gkn1 upregulation. Oncogene 38 (36), 6354–6369 (2019).
Zhou, K. et al. Transglutaminase 3 regulates cutaneous squamous carcinoma differentiation and inhibits progression via PI3K-AKT signaling pathway-mediated Keratin 14 degradation. Cell Death Dis. 15 (4), 252 (2024).
Lee, S. et al. Relationships between lymph node metastasis and expression of CD31, D2–40, and vascular endothelial growth factors A and C in papillary thyroid cancer. Clin. Exp. Otorhinolaryngol. 5 (3), 150–155 (2012).
Guo, Y. L. X. B. & Kang, Y. H. W. X. L. Serum CXCL13 and PECAM - 1 can be used as diagnostic and prognostic markers in elderly patients with gastric cancer. Clin. Trans. Oncol. https://doi.org/10.1007/s12094-020-02403-w (2020).
Virman, J. P., Bono, P., Luukkaala, T. H., Sunela, K. L. & Kujala, P. M. Combined angiogenesis and proliferation markers’ expressions as long-term prognostic factors in renal cell cancer. Clin. Genitourin. Cancer 14 (4), e283–e289. https://doi.org/10.1016/j.clgc.2015.12.014 (2016).
Liu, Z. et al. CD31-labeled circulating endothelial cells as predictor in anlotinib-treated non-small-cell lung cancer: analysis on ALTER-0303 study. Cancer Med. 7 (7), 3011–3021 (2018).
Liu, X. L. et al. Correlation between expression and significance of δ-catenin, CD31, and VEGF of non-small cell lung cancer. Genet. Mol. Res. 14 (4), 13496–13503 (2015).
Cao, S. et al. Prognostic implication of the expression level of PECAM-1 in non-small cell lung cancer. Front. Oncol. 11, 1–12. https://doi.org/10.3389/fonc.2021.587744 (2021).
Righi, L. et al. Role of CD31 / platelet endothelial cell adhesion molecule-1 expression in in vitro and in vivo growth and differentiation of human breast cancer cells. Am. J. Pathol. 162 (4), 1163–1174. https://doi.org/10.1016/S0002-9440(10)63912-0 (2003).
Wu, Q. et al. PECAM - 1 drives β - catenin - mediated EndMT via internalization in colon cancer with diabetes mellitus. Cell Commun. Signal. https://doi.org/10.1186/s12964-023-01193-2 (2023).
Jernström, H., Bågeman, E., Rose, C., Jönsson, P.-E. & Ingvar, C. CYP2C8 and CYP2C9 polymorphisms in relation to tumour characteristics and early breast cancer related events among 652 breast cancer patients. Br. J. Cancer 101 (11), 1817–1823. https://doi.org/10.1038/sj.bjc.6605428 (2009).
Sausville, L. N. et al. The cytochrome P450 slow metabolizers CYP2C9*2 and CYP2C9*3 directly regulate tumorigenesis via reduced epoxyeicosatrienoic acid production. Cancer Res. 78 (17), 4865–4877. https://doi.org/10.1158/0008-5472.CAN-17-3977 (2018).
Sun, Z., Du, T., Yang, G., Sun, Y. & Xiao, X. Identification of exosome-related genes in NSCLC via integrated bioinformatics and machine learning analysis. Sci. Rep. 15 (1), 22962. https://doi.org/10.1038/s41598-025-04485-4 (2025).
Ma, J. et al. Reconstruction and analysis of the immune-related LINC00987/A2M axis in lung adenocarcinoma. Front. Mol. Biosci. https://doi.org/10.3389/fmolb.2021.644557 (2021).
Yao, S. et al. Keratin 14-high subpopulation mediates lung cancer metastasis potentially through Gkn1 upregulation. Oncogene 38 (36), 6354–6369. https://doi.org/10.1038/s41388-019-0889-0 (2019).
Han, W., Hu, C., Fan, Z.-J. & Shen, G.-L. Transcript levels of keratin 1/5/6/14/15/16/17 as potential prognostic indicators in melanoma patients. Sci. Rep. 11 (1), 1023. https://doi.org/10.1038/s41598-020-80336-8 (2021).
Magoola, M. & Niazi, S. K. Current progress and future perspectives of RNA-based cancer vaccines: a 2025 update. Cancers (Basel) 17 (11), 1882. https://doi.org/10.3390/cancers17111882 (2025).
Di Martino, M. T., Tagliaferri, P. & Tassone, P. MicroRNA in cancer therapy: breakthroughs and challenges in early clinical applications. J. Exp. Clin. Cancer Res. 44 (1), 126. https://doi.org/10.1186/s13046-025-03391-x (2025).
Lin, Y. et al. Pan-cancer analysis for identification of tumor antigens and immune subtypes in mRNA vaccine development. iCell https://doi.org/10.71373/FEHU5094 (2024).
Gao, Y., Yang, L., Li, Z., Peng, X. & Li, H. mRNA vaccines in tumor targeted therapy: mechanism, clinical application, and development trends. Biomark. Res. 12 (1), 93. https://doi.org/10.1186/s40364-024-00644-3 (2024).
Author information
Authors and Affiliations
Contributions
Dr. Shadi Shafaghi, Dr. Abdolreza Mohamadnia, and Dr. Elham Nazari had full access to all of the data in the study and took full responsibility for the integrity of the data and the accuracy of the data analysis. Substantial contributions to the conception or design of the study: Rahem Rahmati, Elham Nazari, Rasoul Hossein Zadeh, Fatemeh Zarimeidani, Reza Hossein Zadeh, Maryam Hajimoradi, Muhammad Islampanah. Data acquisition and analysis: Rahem Rahmati, Mahdi Ahmadinia, Naghmeh Bahrami, Abdolreza Mohamadnia, Shadi Shafaghi. Study data Interpretation: Rahem Rahmati, Elham Nazari, Rasoul Hossein Zadeh, Reza Hossein Zadeh, Fatemeh Zarimeidani, Naghmeh Bahrami, Abdolreza Mohamadnia, Shadi Shafaghi. Drafting the study manuscript: All authors. Critical revision for important intellectual content: Elham Nazari, Rasoul Hossein Zadeh, Reza Hossein Zadeh, Fatemeh Zarimeidani, Naghmeh Bahrami, Abdolreza Mohamadnia, Shadi Shafaghi. Final approval of the version to be published: All authors. Agreement to be accountable for all aspects of the work: All authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent for publication
The study was approved by the ethical committee of Shahid Beheshti University of Medical Sciences (Ethics code: IR.SBMU.NRITLD.REC.1403.061). An informed consent was signed by all participants. The study was conducted in accordance with Helsinki’s declaration.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hossein Zadeh, R., Hossein Zadeh, R., Hajimoradi, M. et al. Identification of diagnostic and prognostic biomarkers in lung adenocarcinoma through integrated bioinformatics analysis and real time PCR validation. Sci Rep 16, 6679 (2026). https://doi.org/10.1038/s41598-026-35971-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-35971-y