Abstract
Cystic fibrosis (CF) lung disease is characterized by the presence of marked, neutrophil-dominant inflammation that contributes to tissue injury and the development of irreversible structural lung disease. Here, we describe a dysregulated, neutrophil-dominant inflammation and an accompanying pro-inflammatory airway epithelium in the pediatric CF lung through the application of single-cell RNA sequencing (scRNA-seq) to minimally invasive respiratory specimens collected during flexible bronchoscopy. These findings were present in both an infant and an adolescent with CF, the latter on cystic fibrosis transmembrane conductance regulator (CFTR) modulator therapy, suggesting a common pathological program that starts early in life and may be challenging to reverse once structural lung disease is established. Intercellular communication network analysis further revealed potential mechanisms whereby airway epithelial cells modulate the ongoing, destructive airway inflammation present in the CF lung. Importantly, the scRNA-seq workflow leveraged in this study provides a unique opportunity to investigate and monitor disease-related changes in the composition, function, and interaction of the immune and airway epithelial cell populations in CF and other respiratory diseases across the life course.
Similar content being viewed by others
Introduction
Cystic fibrosis (CF) is a multisystemic, autosomal recessive disorder caused by mutations in the cystic fibrosis transmembrane conductance regulator (CFTR) gene with the majority of morbidity and mortality extending from lung disease. Within the lung, defective or deficient CFTR protein in the airway epithelium leads to the characteristic cycle of mucus stasis and airway obstruction with the impaired clearance of pathogens, the development of airway inflammation, and ultimately the progression to irreversible structural lung disease with bronchiectasis. Early pathological studies of the CF lung1,2 reported that the majority of infants with CF as young as 0–4 months of age exhibited mucopurulent airway plugging and bronchial wall inflammation and that all children with CF had detectable bronchiectasis by two years of age. Subsequent clinical studies identified evidence of bacterial infection3,4,5; neutrophilic airway inflammation3,4,5; structural lung disease5,6,7,8, including bronchial dilation, bronchial wall thickening, and air trapping; and decreased lung function6,9,10,11,12 in infants with CF within the first year of life. The early application of CFTR modulator therapies – small molecules designed to target the underlying defects in the CFTR protein – in conjunction with newborn screening is expected to modify the trajectory of CF lung disease through the early restoration of CFTR protein functionality. An enhanced understanding of the early pathophysiology of CF lung disease, however, may reveal additional therapeutic targets that, if acted on early in life, may further improve future clinical outcomes, particularly for people with CF (pwCF) not currently eligible for or benefitting from CFTR modulator therapies.
Multiple single-cell studies have contributed to an enriched characterization of the immune13,14,15,16 and airway epithelial16,17 cell populations in the CF lung. Leveraging expectorated sputa13, bronchoalveolar lavage fluid (BALF)14,15, and bronchial forceps biopsies16, the single-cell studies characterizing the immune landscape have reported conflicting results, owing, in part, to methodological and technical variations between the studies. Schupp et al.13 described a neutrophil-dominant immunological profile in expectorated sputa from pwCF, consistent with previous clinical studies3,4,5, while the single-cell studies leveraging BALF14,15 and bronchial forceps biopsies16 from pwCF reported a macrophage-dominant immunological profile. Airway epithelial cells were detected in all three sample types – expectorated sputa13, BALF14,15, and bronchial forceps biopsies16 – with the most robust analysis coming from the biopsy specimens. In contrast to the immunological studies, the single-cell studies characterizing the CF airway epithelium, using bronchial forceps biopsies16 and explanted lung tissue17, have largely been in agreement. Both studies report altered airway epithelial cell composition, with reduced basal cell proliferation and skewed differentiation toward ciliated cells, as well as upregulation of immune and antimicrobial gene programs16,17. To date, Berg et al.16 provide the only single-cell study highlighting the interactions between the immune and airway epithelial cell populations in the CF lung, focusing on the lymphoid immune cell lineage. Further studies of immune-epithelial cell interactions in the CF lung are necessary, particularly those attentive to neutrophils given the critical role these cells play in CF pathophysiology.
Here, we provide a detailed characterization of the immune and airway epithelial cell populations in minimally invasive respiratory specimens, including tracheal and bronchial brush biopsies and BALF, collected from pediatric patients with CF and pediatric controls during flexible bronchoscopy.
Some of the data were previously reported in the form of an abstract18.
Results
Single-cell transcriptional profiling of the broad immune and airway epithelial cell populations in the pediatric CF lung
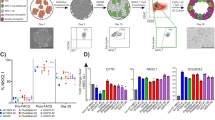
To characterize the immune and airway epithelial cell populations in the pediatric CF lung, we collected minimally invasive respiratory specimens from three distinct locations along the airway using two sampling methods (Fig. 1A) at the time of clinically indicated flexible bronchoscopy. Two pediatric patients with CF, including a 10-month-old male and a 16-year-old female, were included in this study (Table 1). Both patients underwent flexible bronchoscopy for persistent right upper lobe atelectasis, with the infant scheduled for a right upper lobectomy of a severely bronchiectatic and non-functional lobe and the adolescent requiring more aggressive airway clearance after failing conventional medical therapy. Both were colonized with Staphylococcus aureus and carried a diagnosis of pancreatic exocrine insufficiency. Neither had CF-related diabetes. The adolescent patient was receiving CFTR modulator therapy at the time of her flexible bronchoscopy. Further demographic and clinical information are available in Table 1. Age- and sex-matched pediatric patients undergoing flexible bronchoscopy for suspected anatomic large airway disease without CF or other suspected small airway or parenchymal disease served as the pediatric control group (n = 3). The control group was more heterogenous in underlying pulmonary diagnoses and included a 7-month-old male with bronchomalacia; a 4-year-old female with chronic cough, later diagnosed with asthma; and a 17-year-old female with shortness of breath and abnormal pulmonary function testing, which resolved and normalized, respectively, over time without specific intervention. None of the patients in the control group had systemic disease.
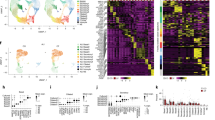
Single-cell transcriptional profiling of the broad immune and epithelial cell populations in minimally invasive respiratory specimens collected from pediatric patients with CF (n = 2) and pediatric controls (CO) (n = 3). (A) Schematic illustration of the study sampling sites and methods, including tracheal brush biopsy (2), bronchial brush biopsy (2), and BAL (1). (B) Uniform manifold approximation and projection (UMAP) visualization of the broad immune and epithelial cell populations divided by disease state and colored by cell population. (C) Stacked column chart visualization of the distributions of the broad immune and epithelial cell populations by disease state (left) and subsequently by pediatric patient and sampling site and method (right). (D) Pearson correlation matrix of the average gene expression profiles, based on all of the differentially expressed genes in all of the cell populations, by disease state and sampling site and method. DC: Dendritic cell; MΦ: Macrophage; Mo: Monocyte; NK: Natural killer cell.
All BALF specimens were sent for routine cell count and microbial culture as clinically indicated, with the results available in Supplementary Table S1. We then performed 3’ 10x single-cell RNA sequencing (scRNA-seq) of the tracheal brush biopsies, bronchial brush biopsies, and surplus BALF from the two pediatric patients with CF and three pediatric controls (total n = 15 specimens). The tracheobronchial brush biopsy specimens from one of the controls (CO3) were inadequate for further analysis and thus excluded from this study. After quality control, normalization, and integration of the single-cell datasets, 20,383 cells were available for analysis. Unsupervised graph-based clustering allowed for the identification of four transcriptionally distinct broad immune cell populations, including neutrophils; monocytes, macrophages, and dendritic cells; B cells; and CD4+ T cells, CD8+ T cells, and NK cells, as well as a broad epithelial cell population (Fig. 1B), with representation from all five pediatric patients and the majority of respiratory specimens (92.3%; 12/13 specimens) (Fig. 1C).
We first compared the relative proportions of the broad immune and epithelial cell populations within each of the three specimen types. The sampling method, itself, affected the resulting distribution of the cell populations with bronchoalveolar lavage (BAL) collecting more immune cells – both neutrophils and monocytes, macrophages, and dendritic cells – and the tracheobronchial brush biopsies collecting more epithelial cells, particularly at the more proximal tracheal location (Fig. 1C). Notably, the tracheobronchial brush biopsy specimens from the pediatric patients with CF had proportionally fewer epithelial cells than those from the controls (p = 0.002), likely owing to the early saturation of the cytology brushes with luminal immune cells and debris. Indeed, the majority of the immune cell populations in the combined tracheobronchial brush biopsy specimens were neutrophils and alveolar macrophages (Supplementary Fig. S1), reflecting the airway lumen microenvironment. While there were favorable gene expression correlations within the cell populations from the different specimen types (Fig. 1D), all subsequent analyses of the immune cell populations were performed on the BALF specimens and all subsequent analyses of the epithelial cell populations were performed on the combined tracheobronchial brush biopsy specimens unless otherwise indicated.
Clustering analysis allowed for the separation of 13,657 immune cells from the BALF specimens (n = 5) into eight clusters (Fig. 2A). The clusters were annotated based on marker gene analysis (Fig. 2B) with the eight clusters representing neutrophils (FCGR3B, IL1R2)19,20, monocytes (VCAN, CCL2)14,20, macrophages (FABP4)20, dendritic cells (FCER1A)14, B cells (CD79A, MS4A1)19,20,21, CD4 + T cells (CD40LG, TRAT1)19,20, CD8+ T cells (CD8A, CD8B)19,20, and NK cells (GNLY, KLRC1)20. All of the immune cell populations were identified in the BALF specimens from all of the pediatric patients with the exception of dendritic cells and NK cells in one pediatric patient with CF (CF1) and B cells in one pediatric control (CO2) (Fig. 2C).
Single-cell transcriptional profiling of the broad immune cell populations in the BALF specimens collected from pediatric patients with CF (n = 2) and pediatric controls (CO) (n = 3). (A) UMAP visualization of the broad immune cell populations divided by disease state and colored by cell population. (B) Dot plot visualization of the top differentially expressed genes in each broad immune cell population by expression level and frequency. (C) Stacked column chart visualization of the distributions of the broad immune cell populations by disease state and subsequently by pediatric patient. DC: Dendritic cell; NK: Natural killer cell.
Clustering analysis further allowed for the separation of 3,065 epithelial cells from the combined tracheobronchial brush biopsy specimens (n = 8) into three clusters (Fig. 3A). The clusters were annotated based on marker gene analysis (Fig. 3B) with the three clusters representing basal cells (KRT5)20,21,22, ciliated cells (RSPH1, PIFO)20,21,22, and secretory cells (SCGB1A1)21. All of the airway epithelial cell populations were identified in the combined tracheobronchial brush biopsy specimens from all of the pediatric patients (Fig. 3C).
Single-cell transcriptional profiling of the broad airway epithelial cell populations in the tracheobronchial brush biopsy specimens collected from pediatric patients with CF (n = 2) and pediatric controls (CO) (n = 2). (A) UMAP visualization of the broad airway epithelial cell populations divided by disease state and colored by cell population. (B) Dot plot visualization of the top differentially expressed genes in each broad airway epithelial cell population by expression level and frequency. (C) Stacked column chart visualization of the distributions of the broad airway epithelial cell populations by disease state and subsequently by pediatric patient. (D) Violin plot visualizations of thematically sorted differentially expressed genes in the secretory cells from the pediatric CF and control lungs. (E) Violin plot visualizations of thematically sorted differentially expressed genes in the ciliated cells from the pediatric CF and control lungs.
We detected a shift in the composition of the immune cell populations in the BALF specimens from a macrophage-dominant immunological profile in the pediatric control lung to a neutrophil-dominant immunological profile in the pediatric CF lung (Fig. 2C), consistent with previous findings in adult healthy control and CF sputa13. The overall distribution of the airway epithelial cell populations in the combined tracheobronchial brush biopsy specimens was similar between the pediatric CF and control lungs (Fig. 3C). Overall, our findings revealed a perceptible shift in immune cell composition in BALF specimens, but not in airway epithelial cell composition in tracheobronchial brush biopsy specimens, in the pediatric CF lung.
The pediatric CF lung contains a diverse landscape of neutrophils, including pro-inflammatory neutrophils
CF lung disease is characterized by the presence of marked, neutrophil-dominant inflammation that may be seen within the first few weeks of life, even in asymptomatic, culture-negative infants with CF3,23. Both pediatric patients with CF in this study had a neutrophil-dominant immunological profile (Fig. 2C) in contrast to the pediatric controls. Clustering analysis allowed for the separation of the previously isolated neutrophil population from the BALF specimens (n = 5) into five clusters (Fig. 4A). The clusters were annotated based on marker gene analysis (Fig. 4B) with reference to previous single-cell studies.
Neutrophil subsets in the pediatric CF and control lungs. (A) UMAP visualization of the five identified neutrophil subsets in the BALF specimens collected from pediatric patients with CF (n = 2) and pediatric controls (CO) (n = 3) divided by disease state and colored by subset. (B) Dot plot visualization of the top differentially expressed genes in each neutrophil subset by expression level and frequency. (C) Stacked column chart visualization of the distributions of the neutrophil subsets by disease state and subsequently by pediatric patient.
Of the five identified neutrophil subsets, one was predominantly found in the pediatric CF lung (Fig. 4C) and aligns with the heat shock response neutrophil archetype (HSPA1A, HSPH1, DNAJB1) previously reported by Schupp et al.13 in expectorated sputa from pwCF. Heat shock proteins (HSPs) are molecular chaperones with roles in cellular defense mechanisms against a variety of disease processes and environmental and metabolic stressors24,25,26,27. The upregulation of HSPs in neutrophils, specifically, has been associated with neutrophil chemotaxis, exocytosis, oxidation, and survival as well as macrophage activation, although HSPs can mediate both pro- and anti-inflammatory responses depending on the HSP, cell type, and overall cellular context28,29,30,31. In the CF lung, the presence of heat shock activated neutrophils may represent an adaptive response to chronic stress, including infection, hypoxia, and oxidative stress. Overexpressed HSPs, however, can stabilize pro-inflammatory signaling pathways and extend neutrophil survival, while extracellular HSPs can act as damage-associated molecular patterns (DAMPs); as such, overexpressed and extracellular HSPs in the CF lung may counterproductively sustain inflammation and contribute to the development of bronchiectasis, thus linking stress signaling and airway injury. Dissecting the protective and pathological roles of heat shock activated neutrophils will be critical to advancing our understanding of CF lung disease pathophysiology and may inform the development of targeted immunomodulatory therapies.
Three of the remaining four identified neutrophil subsets have also been characterized in previous single-cell studies, including immature neutrophils (CXCR4)13, mature neutrophils (CXCR2)13, and “hybrid” neutrophils (HLA-DRA, HLA-DRB1, CD74)19 with macrophage-like characteristics, including the expression of major histocompatibility complex (MHC) class II and complement activation genes. The final cluster was characterized as activated inflammatory neutrophils based on the expression of neutrophil recruitment and chemotaxis (PDE4B, FGD4, SSH2)32,33,34,35 and other pro-inflammatory (CREB5, PELI1)36,37 genes. The activated inflammatory neutrophils were also enriched for chromatin organization and inflammation-related signaling pathways, further supporting its annotation.
In contrast to the pediatric controls, particularly CO1 and CO3, where the majority of isolated neutrophils were identified as “hybrid” neutrophils, the pediatric patients with CF exhibited greater neutrophil subset diversity in their BALF specimens, including a greater proportion of neutrophils classified as immature, activated inflammatory, and heat shock activated neutrophils. Our findings are consistent with those by Schupp et al.13 in adult CF sputa, whereby they described an overall immature, pro-inflammatory neutrophil population.
Notably, many of the single-cell studies in the CF lung to date do not demonstrate the neutrophil-dominant inflammation characteristic of CF lung disease, with the two studies analyzing BALF specimens noting a paucity15 or complete absence14 of neutrophils in the pediatric and adult CF lungs, respectively. This is in contrast with both the present single-cell study and previous cytological studies38 that analyzed BALF specimens from pediatric patients with CF. Neutrophils are fragile, short-lived cells – highly sensitive to handling procedures – that express an exceptionally low number of mRNA transcripts39, rendering the application of scRNA-seq to the study of this population of immune cells challenging. The discrepancy in cellular composition between single-cell studies in the CF lung is most likely due to technical issue rather than biological phenomena, as Schupp et al.13 also reported a neutrophil-dominant immunological profile in expectorated sputa from adults with CF.
Altogether, scRNA-seq revealed an overall diverse array of neutrophils in the pediatric CF lung, including heat shock activated neutrophils, which were rarely detected in the pediatric control lung.
The pediatric CF lung exhibits predominantly pro-inflammatory macrophages, while the pediatric control lung contains a diverse landscape of macrophages
While CF lung disease is characterized by the presence of neutrophil-dominant inflammation, macrophages are the dominant immune cell in the normal, healthy lung40,41. Indeed, macrophages in general – and alveolar macrophages (AMs) in particular – play important roles in immune regulation, pathogen clearance, and overall lung homeostasis. In concordance with the referenced cytological studies, the pediatric controls in this study, particularly CO1 and CO3, demonstrated a macrophage-dominant immunological profile in contrast to the pediatric patients with CF (Fig. 2C). Clustering analysis allowed for the separation of the previously isolated macrophage population from the BALF specimens (n = 5) into eight clusters (Fig. 5A), including the four superclusters of AMs (AM.S1, AM.S2, AM.S3, and AM.S4) initially reported by Li et al.14 and defined by the differential expression of two genes, IFI27 and APOC2 (Fig. 5B). Three of the remaining four identified macrophage subsets, all AMs, can be characterized by the differential expression of one or more functional genes, including CCL18-expressing AMs (CCL18), chemokine-expressing AMs (CCL4, CCL4L2, CCL20), and interferon-expressing AMs (ISG15, IFIT1, IFIT2), while the remaining macrophage subset represents transitioning monocyte-to-alveolar macrophages (FCN1, VCAN, CD14)14.
Macrophage subsets in the pediatric CF and control lungs. (A) UMAP visualization of the eight identified macrophage subsets in the BALF specimens collected from pediatric patients with CF (n = 2) and pediatric controls (CO) (n = 3) divided by disease state and colored by subset. (B) Dot plot visualization of the top differentially expressed genes in each macrophage subset by expression level and frequency. (C) Stacked column chart visualization of the distributions of the macrophage subsets by disease state and subsequently by pediatric patient. AM.S1: Alveolar macrophage supercluster 1; AM.S2: Alveolar macrophage supercluster 2; AM.S3: Alveolar macrophage supercluster 3; AM.S4: Alveolar macrophage supercluster 4; CCL18.AM: CCL18-expressing alveolar macrophage; CK.AM: Chemokine-expressing alveolar macrophage; IFN.AM: Interferon-expressing alveolar macrophage; Mono.AM: Transitioning monocyte-to-alveolar macrophage.
Although macrophages only represent a small fraction of the immune cells in the BALF specimens from the pediatric patients with CF (Fig. 2C), we identified a compositional shift in macrophage subsets in the pediatric CF lung relative to pediatric controls, illustrated by the increased relative abundance of CCL18-expressing AMs (Fig. 5C). CCL18, or C-C motif chemokine ligand 18, is a CC chemokine that is produced primarily by antigen-presenting cells, including AMs in the lung, and is chemotactic for both activated and non-activated T cells42. While CCL18 is constitutively expressed at high levels in the normal, healthy lung, largely in AMs, CCL18 expression increases with stimulation, suggesting that CCL18 and thus CCL18-expressing AMs may be involved in pro-inflammatory processes42. Indeed, other immune response genes, including NAMPT43,44 and NEAT145, are also upregulated in CCL18-expressing AMs.
In contrast to the pediatric CF lung, the four superclusters of AMs (AM.S1-AM.S4) were the dominant macrophage subsets in the pediatric control lung (Fig. 5C). These subsets are considered common macrophages and are predicted to have antimicrobial and metabolic functions14.
Altogether, scRNA-seq revealed a shift in the composition of the AM population from more common AMs in the pediatric control lung to more targeted, pro-inflammatory AMs in the pediatric CF lung.
The pediatric CF lung has a paucity of T cells in the airway lumen
While neutrophils were the dominant immune cell population in the pediatric CF lung and AMs were the dominant immune cell population in the pediatric control lung, both groups’ BALF specimens also contained other myeloid immune cell populations, including monocytes and dendritic cells, as well as multiple lymphoid immune cell populations, including B cells, CD4 + T cells, CD8+ T cells, and NK cells (Fig. 2C).
Clustering analysis allowed for the separation of the previously isolated CD4 + T cell, CD8+ T cell, and NK cell populations from the BALF specimens (n = 5) into five clusters (Fig. 6A). The clusters were annotated based on marker gene analysis (Fig. 6B) with reference to previous single-cell studies. The distribution of the five identified subsets, including CD4 + T cells, regulatory CD4 + T cells (CTLA4, FOXP3, IL2RA)20, naïve CD8+ T cells (LEF1)20, memory CD8+ T cells (CCL5, NKG7, CD8A)20, and NK cells, was similar between the pediatric patients with CF and the pediatric controls, with the exception of one pediatric patient with CF (CF1) whose BALF specimen did not contain either naïve or memory CD8+ T cells (Fig. 6C). Interestingly, naïve and memory CD8+ T cells were also underrepresented in the tracheobronchial brush biopsy specimens from the same patient (not shown). Whether the proportional increase in regulatory CD4 + T cells in this patient’s BALF specimen reflects the active suppression of CD8+ T cell-mediated immune responses or merely represents a byproduct of reduced naïve and memory CD8+ T cells warrants further investigation46.
Natural killer (NK) and T cell subsets in the pediatric CF and control lungs. (A) UMAP visualization of the five identified NK and T cell subsets in the BALF specimens collected from pediatric patients with CF (n = 2) and pediatric controls (CO) (n = 3) divided by disease state and colored by subset. (B) Dot plot visualization of the top differentially expressed genes in each NK and T cell subset by expression level and frequency. (C) Stacked column chart visualization of the distributions of the NK and T cell subsets by disease state and subsequently by pediatric patient. Treg: Regulatory CD4 + T cells.
While the majority of immunological studies in the CF lung to date have focused on neutrophils and macrophages, Ingersoll et al.47 demonstrated that the neutrophil-dominant inflammation characteristic of CF lung disease contributes to T cell function suppression – and ultimately T cell exclusion from the airway lumen – through the upregulation of arginase-1 (Arg1) function. Arg1 is stored in gelatinase neutrophil granules, and upon concomitant exocytosis with azurophil granules, Arg1 metabolizes extracellular arginine to form ornithine and urea48,49. Critically, arginine is necessary for the expression of the invariant ζ-chain of the T cell receptor complex, such that T cell function is inhibited in arginine-depleted environments, like the CF airway50,51. Previous immunohistochemical studies demonstrated a distinct compartmentalization of immune cell populations in the CF lung, with neutrophil-dominant inflammation in the airway lumens and lymphocyte-dominant inflammation in the airway walls, particularly within the submucosa52,53. This compartmentalized distribution of immune cell populations in the CF lung may contribute to its chronic pro-inflammatory and infection-tolerant state, promoting the development of bronchiectasis and the progression to end-stage lung disease46,54.
Neutrophil-driven T cell exclusion from the airway lumen may represent an underrecognized stage in the pathophysiological progression of CF lung disease, and scRNA-seq confirmed an overall paucity of T cells in the BALF specimens from the pediatric patients with CF.
The pediatric CF lung contains a pro-inflammatory airway epithelium
Leveraging explanted lung tissue from pwCF undergoing transplantation, Carrero et al.17 described disease-related changes in the airway epithelial cell populations in the adult CF lung relative to the adult healthy control lung, including a reduction in proliferating basal cells and an increase in ciliated and secretory cells assuming specialized molecular characteristics. In addition to the three major airway epithelial cell populations – basal cells, ciliated cells, and secretory cells – Carraro et al.17 also identified three rare airway epithelial cell populations: FOXN4 + cells, pulmonary ionocytes, and pulmonary neuroendocrine cells. Berg et al.16, using bronchial forceps biopsy specimens from adult patients with CF and adult healthy controls, identified the same three major airway epithelial cell populations as well as pulmonary ionocytes and a population of other rare cells, and Deprez et al.21, using less invasive bronchial brush biopsy specimens from healthy adults, also identified the three major airway epithelial cell populations as well as five rare populations: brush cells, multiciliating goblet cells, pulmonary ionocytes, pulmonary neuroendocrine cells, and “undefined” rare cells. In this study, we were only able to identify the three main airway epithelial cell populations in the combined tracheobronchial brush biopsy specimens from the pediatric patients with CF and the pediatric controls (Fig. 3A), likely due, in part, to the smaller equipment required for the bronchoscopy procedures.
As no significant differences were observed in the distribution of the airway epithelial cell populations between the pediatric CF and control lungs based on the combined tracheobronchial brush biopsy specimens, we next evaluated for disease-related gene expression changes within each airway epithelial cell population (Supplementary Fig. S2 and Supplementary Tables S2-S4).
Within the secretory cells, the specimens from the pediatric patients with CF showed upregulation of immune response genes, including BPIFB117,55, LCN256,57,58, CXCL859, and TGM260 among others (Fig. 3D), with those listed described in previous studies of the CF lung17,55,57,59,60. Our findings are consistent with those of Carraro et al.17 in the adult CF lung, whereby they described an increase in secretory cell function with the upregulation of immune response genes, including BPIFB1, and antimicrobial defense genes, as well as with those of Berg et al.16, also in the adult CF lung, whereby they reported enrichment of immune response pathways.
Within the ciliated cells, the specimens from the pediatric patients with CF showed upregulation of both pro-inflammatory and immune response genes, including SAA1, SAA2, and LCN217, as well as ferroptosis-resistant genes, including LCN261, FTH162, GPX462,63, and HSBP163, the latter of which is also a heat shock response gene (Fig. 3E). Our findings are, once again, in line with those of Carraro et al.17 in the adult CF lung, whereby they described a subset of ciliated cells (Ciliated3) characterized by the upregulation of SAA1 and SAA2 and posited to either respond to or regulate the immune response. Berg et al.16 also noted upregulation of immune response genes, including HLA-DPA1 and HLA-DRB1, in their ciliated cells in the adult CF lung. The second set of upregulated genes in this study, the ferroptosis-resistant genes, is also interesting. Ferroptosis is a form of regulated cell death associated with iron accumulation and lipid peroxidation63 and has been described in the pathophysiological processes of many respiratory diseases, including asthma, chronic obstructive pulmonary disease (COPD), and idiopathic pulmonary fibrosis64. Maniam et al.65 demonstrated that CF airway epithelial cells were more susceptible to ferroptosis than their cognate wild-type cells in vitro. Interestingly, while GPX4 expression decreased significantly in CF airway epithelial cells following treatment with ferric ammonium citrate and erastin, a ferroptosis inducer, the initial levels of GPX4 expression in the CF airway epithelial cells were comparable to, if not greater than, their cognate wild-type cells. The role of ferroptosis in the pathophysiology of CF lung disease warrants further investigation, particularly through in vivo studies.
While our findings are limited by the small number of airway epithelial cells collected via tracheobronchial brush biopsy, scRNA-seq revealed an overall pro-inflammatory airway epithelium in the pediatric CF lung with a shift in predicted functionality, whereby secretory cells, in particular, are thought to be more active.
The airway epithelial cells regulate the neutrophilic inflammation in the pediatric CF lung
With evidence of both a pro-inflammatory immune landscape and a pro-inflammatory airway epithelium in the pediatric CF lung, we next sought to determine whether the airway epithelial cells were simply responding to environmental stimuli or actively partaking in and possibly regulating the overall immune response. We thus evaluated the intercellular communication networks between the airway epithelial cells in general and each of the five identified neutrophil subsets, as neutrophils were the most abundant immune cell population in the pediatric CF lung. Both BALF and tracheobronchial brush biopsy specimens were used in the analysis.
Through the evaluation of ligand-receptor mediated intercellular communication signals, we found that the airway epithelial cells in the pediatric CF lung were principally initiating communication signals while the various neutrophil subsets were largely in receipt of those signals. Overall, we found increased expression of the annexin A1 (ANXA1)-formyl peptide receptor (FPR) system, the serum amyloid A (SAA)-FPR system, and the complement component 3 (C3) system with multiple associated receptors (ITGAX+ITGB2, ITGAM+ITGB2, and C3AR1) between the airway epithelial cells and the five neutrophil subsets (Fig. 7). The dominant ligand-receptor pair mediating airway epithelial cell–neutrophil interactions was ANXA1-FPR1. ANXA1 serves, in part, to regulate the functions of neutrophils in the inflammatory response via its interactions with FPRs and is largely considered to serve a protective, anti-inflammatory role66,67,68. Expectedly, the more mature neutrophil subsets – mature neutrophils and “hybrid” neutrophils – exhibited increased engagement of this pathway (Fig. 7B). This is in contrast with the other highly expressed ligand-receptor pairs, including SAA1-FPR269,70 and C3-ITGAX+ITGB271, which have pro-inflammatory roles, including neutrophil activation and recruitment and neutrophil migration and phagocytosis, respectively. Taken together, the airway epithelial cells regulate, in part, the neutrophilic inflammation characteristic of CF lung disease, sending both anti- and pro-inflammatory signals.
Intercellular communication networks between airway epithelial cells (AECs) and neutrophils in the pediatric CF lung. (A) Violin plot visualizations of the contributing ligands and receptors involved in intercellular communication signaling between airway epithelial cells and each of the neutrophil subsets. (B) Heatmap visualization of the significant ligand-receptor pairs (p < 0.01) contributing to intercellular communication signaling from airway epithelial cells to each of the neutrophil subsets by communication probability.
The relative paucity of neutrophils in the pediatric control lung prohibited a comparative analysis of the intercellular communication networks between the pediatric CF and control lungs.
Minimally invasive respiratory specimens may be acceptable surrogates for lung tissue specimens in the study of respiratory diseases
Since its introduction, scRNA-seq has been regarded as a powerful tool for studying disease-related changes in cellular composition and gene expression across various tissues in the human body. Indeed, the available single-cell studies in the CF lung13,14,15,16,17, in conjunction with the present study, suggest a neutrophil-dominant, pro-inflammatory immune landscape and a pro-inflammatory airway epithelium.
As traditional scRNA-seq workflows require single-cell dissociation, they inherently lack spatial information that may prove indispensable for a comprehensive understanding of the pathophysiology of CF lung disease. Spatial transcriptomics serves, in part, to overcome this deficiency72 and has been widely applied in the study of lung diseases – most notably lung cancer73,74,75 – but not in CF lung disease.
As one of our pediatric patients with CF (CF1) underwent flexible bronchoscopy at the time of a scheduled lobectomy, we additionally collected resected lung tissue for both scRNA-seq and spatial transcriptomics to complement his set of minimally invasive respiratory specimens. Interestingly, while both the tissue scRNA-seq and spatial transcriptomic analyses revealed a predominance of neutrophils (Supplementary Figs. S3 and S4) – similar to the minimally invasive respiratory specimen scRNA-seq analyses (Fig. 1C) – this was less aligned with the histological review of the resected lung tissue (Supplementary Fig. S5). Upon histological review, neutrophils were identified within the airway lumens as well as at the center of histiocytic clusters within the lung tissue but were not the predominant cell population. The lung tissue was, instead, marked by a preponderance of lymphoid immune cell populations as well as histiocytes from the myeloid lineage. Although multiple lung tissue specimens from the same resected lobe were evaluated histologically and transcriptionally, sampling bias undoubtedly contributed to the conflicting results. The lobectomy specimen necessarily went to clinical pathology prior to the research laboratory, such that the lung tissue specimens used for scRNA-seq and spatial transcriptomics were spatially closer to each other than to the specimens used for histological analysis. Furthermore, as evident on the latter, neutrophils were regionally clustered within the lung, such that their representation in each specimen could vary depending on the area sampled. Notably, as the clinical BALF cell counts confirm a neutrophil-dominant immunological profile in the pediatric CF lung (Supplementary Table S1), there is histological and transcriptional congruence in the minimally invasive respiratory specimens, such that the neutrophil annotation is justified. While the histological and transcriptional incongruence in the lung tissue specimens is hypothesized to be secondary to sampling bias, further investigation with more, and ideally adjacent, histology and spatial gene expression slides is needed.
Importantly, the scRNA-seq analyses from the minimally invasive respiratory specimens aligned with the scRNA-seq analyses from the resected lung tissue specimens obtained from the same pediatric patient with CF, suggesting that minimally invasive respiratory specimens and presumably more invasive bronchoscopically acquired respiratory specimens, like endobronchial and transbronchial biopsies, may be acceptable surrogates for lung tissue specimens in the study of various respiratory diseases. The addition of scRNA-seq and spatial transcriptomics to traditional histological review may provide new pathophysiological insights and open novel paths for future investigations.
Discussion
In this study, we leveraged single-cell transcriptional analyses to characterize the immune and airway epithelial cell populations in the pediatric CF and control lungs using minimally invasive respiratory specimens collected at the time of clinically indicated flexible bronchoscopy. We focused on the pediatric population in an effort to better understand the early pathophysiology of CF lung disease. While other single-cell studies in the CF lung have largely investigated the composition of the immune landscape13,14,15 or airway epithelium17 individually, these interacting cell populations warrant study in concert16. Therefore, we complemented routine BALF specimens with tracheobronchial brush biopsy specimens in an effort to characterize the immune and airway epithelial cell populations as well as their intercellular communication networks. Through our approach, we provide insights into the composition of, functional changes within, and intercellular communication signals between the immune and airway epithelial cell populations in the pediatric CF lung.
Single-cell RNA sequencing revealed a dysregulated, neutrophil-dominant inflammation and an accompanying pro-inflammatory airway epithelium in the pediatric CF lung. More specifically, we observed an increase in the relative proportion of neutrophils and in the diversity of neutrophil subsets, including the presence of a disease-related neutrophil subset, heat shock activated neutrophils, with a concomitant decrease in the relative proportion of AMs and in the diversity of AM subsets, with a shift from more common to more targeted, pro-inflammatory AM subsets. Furthermore, we observed evidence of suspected neutrophil-driven T cell exclusion from the airway lumen with a relative paucity of T cells in general and of naïve and memory CD8+ T cells in one pediatric patient with CF in particular. While we did not detect a change in the airway epithelial cell composition in the pediatric CF lung, we observed pro-inflammatory functional changes, whereby both secretory and ciliated cells appear to become more active. Interestingly, the airway epithelial cells were found to regulate, in part, the neutrophilic inflammation characteristic of CF lung disease, sending both anti-inflammatory and pro-inflammatory signals through the ANXA1-FPR system and the SAA1-FPR and C3 systems, respectively.
Infancy and early childhood represent a critical period in the pathophysiology of CF lung disease such that the well-timed introduction of targeted therapies, including CFTR modulator therapies, may serve to delay or even prevent the development of lung damage and, in turn, alter the longer-term clinical trajectory of pwCF. The scRNA-seq workflow leveraged in this study provides an opportunity to investigate and monitor disease-related changes in the composition, function, and interaction of the immune and airway epithelial cell populations in the CF lung, although serial sampling from the same cohort of pwCF is required for the latter. Indeed, Loske et al.76, leveraging a similar workflow with nasal swabs, were able to detect reductions in pro-inflammatory gene signatures and elevations in innate mucosal immunity within the immune and nasal epithelial cell populations, respectively, in children with CF following the initiation of the triple combination CFTR modulator therapy, elexacaftor-tezacaftor-ivacaftor (ETI). Conceptually, our scRNA-seq workflow may thus be used to not only better understand the pathophysiology of CF lung disease but also monitor disease progression and evaluate therapy response, particularly as exploratory endpoints in the research setting77. While the youngest patient included in this study was seven months old at the time of flexible bronchoscopy, we have performed the same sampling methods, including tracheal and bronchial brush biopsies, in pediatric patients less than one month of age without complications. The major impediments to the widespread adoption of such a workflow is the invasiveness of the procedure and the need for anesthesia.
While epidemiological data boast improving survival rates with preserved lung function throughout childhood and often into late adolescence78, the majority of children with CF not receiving CFTR modulator therapies have evidence of bronchiectasis by school age79,80. Thus, despite the tremendous advancements in the clinical care of pwCF, some pwCF are still developing irreversible structural lung disease early in life. It is anticipated that the early application of CFTR modulator therapies will modify the trajectory of CF lung disease, but recent clinical studies have shown that chronic bacterial infections81,82 and immune cell functional deficits83 both persist following the initiation of CFTR modulator therapies, such that novel anti-microbial and anti-inflammatory therapies may be additionally required to prevent the development and progression of structural lung disease. Furthermore, recent animal studies have suggested that the early initiation of CFTR modulator therapies containing tezacaftor may negatively impact nervous system development84, such that the initiation of such therapies will require thoughtful timing.
It is notable and perhaps disheartening that one of the two pediatric patients with CF included in this study (CF2) was on ETI at the time of flexible bronchoscopy and for over eighteen months prior to the procedure. Despite being on targeted therapy, her immune and airway epithelial cell cellular and molecular phenotype was more closely aligned with the other pediatric patient with CF not on CFTR modulator therapy than the pediatric controls. While Loske et al.76 were able to detect the partial restoration of a normalized transcriptional signature in the immune and nasal epithelial cells from children with CF following the initiation of ETI, some disease-related gene expression changes were unaffected. The pediatric patient with CF on ETI in this study (CF2) was a teenager with established bronchiectasis at the time of ETI initiation and later study inclusion. We thus suspect that the severity of her lung disease prohibited the restoration of a normalized cellular and molecular phenotype following the initiation of ETI despite signs of clinical improvement. Indeed, Berg et al.16 also report changes in immune and airway epithelial cell cellular and molecular phenotypes in adults with CF on CFTR modulator therapies, also concluding that alterations in these populations persist even after restoring CFTR function. Ultimately, longitudinal studies throughout the life course, with disease exacerbations, and pre- and post-treatment initiation are needed. These studies will be critical to understanding the early pathophysiology of CF lung disease, including the changes present in the early cycles of airway infection, inflammation, and tissue injury, as well as the potential for specific therapies, including CFTR modulator therapies, to mitigate the aforementioned changes.
This study has several important limitations. First, the results are based on specimens from a small cohort of pediatric patients with CF and pediatric controls. The introduction and widespread adoption of CFTR modulator therapies have improved the clinical outcomes for many pwCF, such that flexible bronchoscopy procedures are less often clinically indicated, particularly in the pediatric population. Second, this study lacks a true asymptomatic pediatric healthy control group, as all of the pediatric patients included in this study had clinical indications for flexible bronchoscopy. We sought to mitigate this limitation as much as possible by specifically recruiting pediatric patients with suspected anatomic large airway disease and without suspected small airway or parenchymal disease, acknowledging that these children cannot be considered “normal”. Third, there were significant differences in the distribution of the immune cell populations between the pediatric patients with CF and the pediatric controls, such that we could not reliably evaluate disease-related gene expression changes within the immune cell populations or subsets. We thus confined our analyses to understanding the immune cell population and subset distributions in the pediatric CF lung as compared to the pediatric control lung. Fourth, airway epithelial cells represented only a small fraction of the analyzed cells, such that rare airway epithelial cell populations, like pulmonary ionocytes, were not able to be identified. Despite these limitations, our findings are in accordance with many of the other single-cell studies in the CF lung and demonstrate the feasibility and utility of multiple site and method sampling during flexible bronchoscopy, even in infants.
In summary, through the application of a scRNA-seq workflow to minimally invasive respiratory specimens collected during flexible bronchoscopy, we describe a dysregulated, neutrophil-dominant inflammation and an accompanying pro-inflammatory airway epithelium in the pediatric CF lung. These findings were present in an infant with CF as well as a teenager with CF on ETI, suggesting a common pathological program that starts early in life and may be challenging to reverse once structural lung disease is established. Furthermore, we provide new insight into the interplay between immune and airway epithelial cell populations in the pediatric CF lung, including the identification of ligand-receptor signals originating from airway epithelial cells and modulating airway inflammation. Further mechanistic studies are necessary to understand when and how these findings contribute to the pathophysiology of CF lung disease and whether they can facilitate the design and application of new anti-inflammatory therapies.
Methods
Pediatric patients and specimens
Minimally invasive respiratory specimens (n = 15), including tracheal brush biopsies (n = 5), bronchial brush biopsies (n = 5), and surplus BALF (n = 5), were collected from pediatric patients with CF (n = 2) undergoing clinically indicated flexible bronchoscopy for any indication and the next chronologic age- and sex-matched pediatric patient without CF or other suspected small airway or parenchymal disease (n = 3) undergoing clinically indicated flexible bronchoscopy for suspected anatomic large airway disease (e.g. tracheomalacia, bronchomalacia, bronchial stenosis) at the Icahn School of Medicine at Mount Sinai (ISMMS) in New York, New York. All procedures were performed under general anesthesia.
The tracheal brush biopsies were obtained from the distal trachea. The bronchial brush biopsies were obtained from the right lower lobe and more specifically the anterior segment of the right lower lobe where technically feasible. For the bronchial brush biopsies, the cytology brush was advanced until gentle resistance was encountered, at which point the brush was slightly retracted to avoid airway trauma. The tracheal and bronchial brush biopsy specimens, separated by location, were placed on ice in MACS Tissue Storage Solution (Miltenyi Biotec) and processed immediately after the procedure. All tracheobronchial brush biopsies were performed by the same pediatric pulmonologist. BAL was performed with three aliquots of 1 mL/kg – maximum 30 mL/aliquot – sterile 0.9% sodium chloride in the right middle lobe and more specifically the medial segment of the right middle lobe where technically feasible unless another location was clinically indicated. BALF specimens were placed on ice and processed immediately after the procedure. Clinically indicated deviations from the aforementioned are noted in Supplementary Table S5.
Ethical approval
This study was approved by the Institutional Review Board at the Ichan School of Medicine at Mount Sinai (STUDY-20-01845) and was conducted in accordance with relevant guidelines and regulations. Written informed consent was obtained from all of the pediatric patients’ parents or legal guardians, and written or verbal informed assent was obtained from all of the pediatric patients over six years of age at the time of study enrollment.
Preparation of single-cell suspensions from respiratory specimens
The tracheal and bronchial brush biopsies were collected and stored in MACS Tissue Storage Solution (Miltenyi Biotec) for transport. The tracheal and bronchial brush biopsy and BALF specimens were processed within one hour of flexible bronchoscopy. Briefly, the tracheal brush biopsies, bronchial brush biopsies, and BALF were spun down at 500 x g for 5 min at 4 °C, and the cytology brushes were removed. The specimens were treated with 0.5 mL eBioscience 1X RBC Lysis Buffer (Invitrogen) for 3 min. The Buffer was then removed and replaced with 1.5 mL PBS supplemented with 1% UltraPure BSA (Invitrogen). The cells were pelleted at 500 x g for 5 min at 4 °C and resuspended in 1 mL PBS supplemented with 1% UltraPure BSA (Invitrogen). The cell resuspensions were filtered through a 30 μm pre-separation filter (Miltenyi Biotec). Specimen cell count and viability were confirmed with an EVOS M7000 Imaging System (Invitrogen).
Resected lung tissue was collected by the Mount Sinai Biorepository at the ISMMS. The lung tissue specimens were processed within two hours of surgical resection. Briefly, the lung tissue specimens were cleaned, minced, and ultimately digested in Hanks’ Balanced Salt Solution (Gibco) supplemented with elastase (Worthington Biochemical Corporation) and collagenase A (Roche) for 1 h at 37 °C. The resulting cell suspensions were filtered through a 30 μm pre-separation filter (Miltenyi Biotec). Further specimen processing was performed as described above.
10x Genomics single-cell RNA sequencing
The processed specimens, including the tracheal brush biopsies, bronchial brush biopsies, BALF, and resected lung tissue, were provided to the Genomics Core Facility at the ISMMS for 10x Genomics scRNA-seq. The Chromium Next GEM Single Cell 3’ Reagents Kit (10x Genomics) was used to generate the scRNA-seq libraries. Briefly, the cell suspensions were spun down and resuspended in PBS supplemented with 1% UltraPure BSA (Invitrogen) to a final concentration of 1,000 cells/µL with a target recovery of 10,000 cells per specimen. The Master Mix was prepared with an adjusted volume of nuclease-free water as per the Chromium Single Cell 3’ Reagent Kits User Guide (v3.1 Chemistry Dual Index; 10x Genomics). The cells were loaded onto the Chromium Next GEM Chip G to generate the gel beads-in-emulsion (GEMs). The GEMs subsequently underwent GEM-RT incubation, cDNA amplification, and library construction as per the Chromium Single Cell 3’ Reagent Kits User Guide (v3.1 Chemistry Dual Index; 10x Genomics). Quality controls for cDNA amplification and the final barcoded libraries were performed using the High Sensitivity DNA Kit (Agilent Technologies) with the Agilent 2100 Bioanalyzer to assess quantity and fragment size, respectively. The sequencing libraries were then loaded onto a NovaSeq 6000 System (Illumina).
Tissue handling for spatial transcriptomics
Resected lung tissue was collected by the Mount Sinai Biorepository at the ISMMS. The lung tissue specimens were provided to the Genomics Core Facility at the ISMMS for Visium spatial gene expression (10x Genomics) within two hours of surgical resection. Briefly, the fresh lung tissue specimens were snap frozen in a bath of isopentane (Millipore Sigma) and liquid nitrogen and then embedded in Tissue-Tek O.C.T. Compound (Sakura). The tissue block was then cryosectioned in a pre-cooled cryostat, and the tissue section was transferred to the 6.5 mm x 6.5 mm capture area on the Visium Spatial slide (10x Genomics). The tissue section was fixed, H&E stained, and imaged as per the Methanol Fixation, H&E Staining, and Imaging for Visium Spatial Protocols (10x Genomics). The tissue section subsequently underwent permeabilization, reverse transcription, second strand synthesis, denaturation, cDNA amplification, and library construction as per the Visium Spatial Gene Expression Reagent Kits User Guide (10x Genomics). Quality controls for cDNA amplification and the final barcoded libraries were performed using the High Sensitivity DNA Kit (Agilent Technologies) with the Agilent 2100 Bioanalyzer to assess quantity and fragment size, respectively. The sequencing libraries were then loaded onto a NovaSeq 6000 System (Illumina).
Single-cell RNA sequencing data analysis
The 10x Cell Ranger toolkit (v6.0.0) was used to filter and align the reads onto the human genome hg38 and to count the barcodes and the unique molecular identifiers (UMIs). Seurat (v5.0.1)85 was used for the subsequent downstream scRNA-seq data analyses. To ensure data quality, cells with less than 200 detected genes or more than 6,000 detected genes and cells with a mitochondrial gene percentage greater than 0.2 were excluded from further analysis. DecontX (v1.0.0)86 was used to estimate the ambient RNA contamination.
In brief, each specimen was individually preprocessed prior to the subsequent integration by sampling method. Normalization through log transformation and canonical correlation were used to uncover anchors among the specimens. Iterative pairwise integration was used to define the distance between the specimen datasets and to cluster the distance matrix. Following the integration of the specimen datasets, unsupervised K-nearest neighbors graph-based clustering was performed using the significant principal components. The major cell populations were annotated based on their expression of canonical marker genes and their reference-based integration with published datasets. The major cell populations were then renormalized and reintegrated for the annotation and analysis of population subsets. The Seurat function FindAllMarkers with a minimum detection percent of 25 and a logFC threshold of 0.25 was used to identify cluster-specific marker genes. The Wilcoxon rank-sum test was used for differential expression analysis. The Benjamini-Hochberg correction was used to adjust for multiple comparisons.
Ligand-receptor mediated intercellular communication analysis
CellChat (v.1.6.1)87 was used to quantitatively infer and analyze intercellular communication networks between the airway epithelial cell population and the neutrophil subsets in the pediatric CF lung. In brief, to predict significant intercellular communication signals, differentially overexpressed signaling genes were identified for each cell population or subset, and the resulting gene expression profiles were projected onto a high confidence, experimentally validated protein-protein interaction network. The probability of intercellular communication was then calculated by modeling ligand-receptor mediated signaling interactions using the law of mass action.
Spatial transcriptomics data analysis
The 10x Space Ranger toolkit (v1.3.1) was used to filter and align the reads onto the human genome hg38 to estimate gene expression for each of the barcoded spots. Seurat (v5.0.1)85 was used for the subsequent downstream spatial transcriptomic analysis.
Data availability
Raw FASTQ and processed matrix data that support the findings of this study have been deposited in the National Center for Biotechnology Information’s Gene Expression Omnibus at the National Institutes of Health with the accession number GSE271984. Additional data and code required to analyze the results and generate the figures included in this manuscript are available from the corresponding authors upon reasonable request.
References
Bedrossian, C. W. et al. The lung in cystic fibrosis. A quantitative study including prevalence of pathologic findings among different age groups. Hum. Pathol. 7, 195–204 (1976).
Esterly, J. R. & Oppenheimer, E. H. Observations in cystic fibrosis of the pancreas. 3. Pulmonary lesions. Johns. Hopkins Med. J. 122, 94–101 (1968).
Khan, T. Z. et al. Early pulmonary inflammation in infants with cystic fibrosis. Am. J. Respir Crit. Care Med. 151, 1075–1082 (1995).
Armstrong, D. S. et al. Lower airway inflammation in infants and young children with cystic fibrosis. Am. J. Respir Crit. Care Med. 156, 1197–1204 (1997).
Sly, P. D. et al. Lung disease at diagnosis in infants with cystic fibrosis detected by newborn screening. Am. J. Respir Crit. Care Med. 180, 146–152 (2009).
Martinez, T. M. et al. High-resolution computed tomography imaging of airway disease in infants with cystic fibrosis. Am. J. Respir Crit. Care Med. 172, 1133–1138 (2005).
Stick, S. M. et al. Bronchiectasis in infants and preschool children diagnosed with cystic fibrosis after newborn screening. J. Pediatr. 155, 623–628 (2009).
Sly, P. D. et al. Risk factors for bronchiectasis in children with cystic fibrosis. N Engl. J. Med. 368, 1963–1970 (2013).
Ranganathan, S. C. et al. Airway function in infants newly diagnosed with cystic fibrosis. Lancet 358, 1964–1965 (2001).
Ranganathan, S. C. et al. The evolution of airway function in early childhood following clinical diagnosis of cystic fibrosis. Am. J. Respir Crit. Care Med. 169, 928–933 (2004).
Linnane, B. M. et al. Lung function in infants with cystic fibrosis diagnosed by newborn screening. Am. J. Respir Crit. Care Med. 178, 1238–1244 (2008).
Belessis, Y. et al. Early cystic fibrosis lung disease detected by Bronchoalveolar lavage and lung clearance index. Am. J. Respir Crit. Care Med. 185, 862–873 (2012).
Schupp, J. C. et al. Single-cell transcriptional archetypes of airway inflammation in cystic fibrosis. Am. J. Respir Crit. Care Med. 202, 1419–1429 (2020).
Li, X. et al. ScRNA-seq expression of IFI27 and APOC2 identifies four alveolar macrophage superclusters in healthy BALF. Life Sci. Alliance. 5, e202201458 (2022).
Maksimovic, J. et al. Single-cell atlas of bronchoalveolar lavage from preschool cystic fibrosis reveals new cell phenotypes. Preprint at bioRxiv 2022.2006.2017.496207 (2022).
Berg, M. et al. Evidence for altered immune-structural cell crosstalk in cystic fibrosis revealed by single cell transcriptomics. J Cyst. Fibros (2025).
Carraro, G. et al. Transcriptional analysis of cystic fibrosis airways at single-cell resolution reveals altered epithelial cell States and composition. Nat. Med. 27, 806–814 (2021).
Januska, M. N. et al. Moving toward personalized cystic fibrosis care: a workflow for single-cell RNA sequencing. Am. J. Respir Crit. Care Med. 205, A5179 (2022).
Wauters, E. et al. Discriminating mild from critical COVID-19 by innate and adaptive immune single-cell profiling of Bronchoalveolar lavages. Cell. Res. 31, 272–290 (2021).
Sikkema, L. et al. An integrated cell atlas of the lung in health and disease. Nat. Med. 29, 1563–1577 (2023).
Deprez, M. et al. A single-cell atlas of the human healthy airways. Am. J. Respir Crit. Care Med. 202, 1636–1645 (2020).
Liu, Y. Identification and comprehensive analysis of super-enhancer related genes involved in epithelial-to-mesenchymal transition in lung adenocarcinoma. PLoS One. 18, e0291088 (2023).
Thursfield, R. et al. S82 Airway inflammation is present by 4 months in CF infants diagnosed on newborn screening. Thorax 67, A40–A41 (2012).
Rozhkova, E. et al. Exogenous mammalian extracellular HSP70 reduces endotoxin manifestations at the cellular and organism levels. Ann. N Y Acad. Sci. 1197, 94–107 (2010).
Vinokurov, M. et al. Recombinant human Hsp70 protects against Lipoteichoic acid-induced inflammation manifestations at the cellular and organismal levels. Cell. Stress Chaperones. 17, 89–101 (2012).
Yao, Y. W. et al. Lipopolysaccharide pretreatment protects against ischemia/reperfusion injury via increase of HSP70 and Inhibition of NF-κB. Cell. Stress Chaperones. 16, 287–296 (2011).
Afrazi, A. et al. Intracellular heat shock protein-70 negatively regulates TLR4 signaling in the newborn intestinal epithelium. J. Immunol. 188, 4543–4557 (2012).
Jog, N. R. et al. Heat shock protein 27 regulates neutrophil chemotaxis and exocytosis through two independent mechanisms. J. Immunol. 178, 2421–2428 (2007).
Hashiguchi, N. et al. Enhanced expression of heat shock proteins in activated polymorphonuclear leukocytes in patients with sepsis. J. Trauma. 51, 1104–1109 (2001).
Gupta, S. et al. Heat-shock protein-90 prolongs septic neutrophil survival by protecting c-Src kinase and caspase-8 from proteasomal degradation. J. Leukoc. Biol. 103, 933–944 (2018).
Zheng, L. et al. Pathogen-induced apoptotic neutrophils express heat shock proteins and elicit activation of human macrophages. J. Immunol. 173, 6319–6326 (2004).
Ariga, M. et al. Nonredundant function of phosphodiesterases 4D and 4B in neutrophil recruitment to the site of inflammation. J. Immunol. 173, 7531–7538 (2004).
Cao, M. et al. Mechanisms of impaired neutrophil migration by MicroRNAs in myelodysplastic syndromes. J. Immunol. 198, 1887–1899 (2017).
Xu, X. et al. GPCR-mediated PLCβγ/PKCβ/PKD signaling pathway regulates the Cofilin phosphatase slingshot 2 in neutrophil chemotaxis. Mol. Biol. Cell. 26, 874–886 (2015).
Tang, W. et al. A PLCβ/PI3Kγ-GSK3 signaling pathway regulates Cofilin phosphatase slingshot2 and neutrophil polarization and chemotaxis. Dev. Cell. 21, 1038–1050 (2011).
Liu, X. et al. Single-cell RNA transcriptomic analysis identifies Creb5 and CD11b-DCs as regulator of asthma exacerbations. Mucosal Immunol. 15, 1363–1374 (2022).
Chang, M. et al. Peli1 facilitates TRIF-dependent Toll-like receptor signaling and Proinflammatory cytokine production. Nat. Immunol. 10, 1089–1095 (2009).
Alexis, N. E. et al. Attenuation of host defense function of lung phagocytes in young cystic fibrosis patients. J. Cyst. Fibros. 5, 17–25 (2006).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334 (2019).
Ratjen, F. et al. Differential cytology of Bronchoalveolar lavage fluid in normal children. Eur. Respir J. 7, 1865–1870 (1994).
Midulla, F. et al. Bronchoalveolar lavage studies in children without parenchymal lung disease: cellular constituents and protein levels. Pediatr. Pulmonol. 20, 112–118 (1995).
Hieshima, K. et al. A novel human CC chemokine PARC that is most homologous to macrophage-inflammatory protein-1 alpha/LD78 alpha and chemotactic for T lymphocytes, but not for monocytes. J. Immunol. 159, 1140–1149 (1997).
Zhan, T. et al. Nicotinamide phosphoribose transferase facilitates macrophage-mediated pulmonary fibrosis through the Sirt1-Smad7 pathway in mice. Eur. J. Pharmacol. 967, 176355 (2024).
Hong, S. M. et al. NAMPT mitigates colitis severity by supporting redox-sensitive activation of phagocytosis in inflammatory macrophages. Redox Biol. 50, 102237 (2022).
Liu, Y. et al. Long non-coding RNA NEAT1 participates in ventilator-induced lung injury by regulating miR-20b expression. Mol. Med. Rep. 25, 66 (2022).
Chen, K. & Kolls, J. K. T cell-mediated host immune defenses in the lung. Annu. Rev. Immunol. 31, 605–633 (2013).
Ingersoll, S. A. et al. Mature cystic fibrosis airway neutrophils suppress T cell function: evidence for a role of arginase 1 but not programmed death-ligand 1. J. Immunol. 194, 5520–5528 (2015).
Jacobsen, L. C. et al. Arginase 1 is expressed in myelocytes/metamyelocytes and localized in gelatinase granules of human neutrophils. Blood 109, 3084–3087 (2007).
Rotondo, R. et al. Exocytosis of azurophil and arginase 1-containing granules by activated polymorphonuclear neutrophils is required to inhibit T lymphocyte proliferation. J. Leukoc. Biol. 89, 721–727 (2011).
Munder, M. et al. Suppression of T-cell functions by human granulocyte arginase. Blood 108, 1627–1634 (2006).
Zea, A. H. et al. L-Arginine modulates CD3zeta expression and T cell function in activated human T lymphocytes. Cell. Immunol. 232, 21–31 (2004).
Regamey, N. et al. Distinct patterns of inflammation in the airway lumen and bronchial mucosa of children with cystic fibrosis. Thorax 67, 164–170 (2012).
Hubeau, C. et al. Quantitative analysis of inflammatory cells infiltrating the cystic fibrosis airway mucosa. Clin. Exp. Immunol. 124, 69–76 (2001).
Hynes, J. et al. Innate immunity in cystic fibrosis: varied effects of CFTR modulator therapy on cell-to-cell communication. Int. J. Mol. Sci. 26, 2636 (2025).
Bingle, L. et al. BPIFB1 (LPLUNC1) is upregulated in cystic fibrosis lung disease. Histochem. Cell. Biol. 138, 749–758 (2012).
Cowland, J. B. & Borregaard, N. Molecular characterization and pattern of tissue expression of the gene for neutrophil gelatinase-associated Lipocalin from humans. Genomics 45, 17–23 (1997).
Cowland, J. B. et al. Neutrophil gelatinase-associated Lipocalin is up-regulated in human epithelial cells by IL-1 beta, but not by TNF-alpha. J. Immunol. 171, 6630–6639 (2003).
Zughaier, S. M. et al. Peripheral monocytes derived from patients with cystic fibrosis and healthy donors secrete NGAL in response to Pseudomonas aeruginosa infection. J. Investig Med. 61, 1018–1025 (2013).
Oglesby, I. K. et al. miR-17 overexpression in cystic fibrosis airway epithelial cells decreases interleukin-8 production. Eur. Respir J. 46, 1350–1360 (2015).
Maiuri, L. et al. Tissue transglutaminase activation modulates inflammation in cystic fibrosis via PPARgamma down-regulation. J. Immunol. 180, 7697–7705 (2008).
Gao, J. et al. Gene interfered-ferroptosis therapy for cancers. Nat. Commun. 12, 5311 (2021).
Luo, T. et al. Ferroptosis assassinates tumor. J. Nanobiotechnol. 20, 467 (2022).
Tang, D. et al. The molecular machinery of regulated cell death. Cell. Res. 29, 347–364 (2019).
Li, Y. et al. Multifaceted roles of ferroptosis in lung diseases. Front. Mol. Biosci. 9, 919187 (2022).
Maniam, P. et al. Increased susceptibility of cystic fibrosis airway epithelial cells to ferroptosis. Biol. Res. 54, 38 (2021).
Sugimoto, M. A. et al. Annexin A1 and the resolution of inflammation: modulation of neutrophil recruitment, apoptosis, and clearance. J. Immunol. Res. 2016, 8239258 (2016).
Bensalem, N. et al. Down-regulation of the anti-inflammatory protein Annexin A1 in cystic fibrosis knock-out mice and patients. Mol. Cell. Proteom. 4, 1591–1601 (2005).
D’Acquisto, F. et al. Pro-inflammatory and pathogenic properties of Annexin-A1: the whole is greater than the sum of its parts. Biochem. Pharmacol. 85, 1213–1218 (2013).
Abouelasrar Salama, S. et al. Serum amyloid A1 (SAA1) revisited: restricted leukocyte-activating properties of homogeneous SAA1. Front. Immunol. 11, 843 (2020).
Abouelasrar Salama, S. et al. Acute-serum amyloid A and A-SAA-derived peptides as formyl peptide receptor (FPR) 2 ligands. Front. Endocrinol. 14, 1119227 (2023).
Dustin, M. L. Complement receptors in myeloid cell adhesion and phagocytosis. Microbiol Spectr 4, 56 (2016).
Stahl, P. L. et al. Visualization and analysis of gene expression in tissue sections by Spatial transcriptomics. Science 353, 78–82 (2016).
Zhu, J. et al. Delineating the dynamic evolution from preneoplasia to invasive lung adenocarcinoma by integrating single-cell RNA sequencing and Spatial transcriptomics. Exp. Mol. Med. 54, 2060–2076 (2022).
Wang, Y. et al. Spatial transcriptomics delineates molecular features and cellular plasticity in lung adenocarcinoma progression. Cell. Discov. 9, 96 (2023).
De Zuani, M. et al. Single-cell and Spatial transcriptomics analysis of non-small cell lung cancer. Nat. Commun. 15, 4388 (2024).
Loske, J. et al. Pharmacological improvement of cystic fibrosis transmembrane conductance regulator function rescues airway epithelial homeostasis and host defense in children with cystic fibrosis. Am. J. Respir Crit. Care Med. 209, 1338–1350 (2024).
Januska, M. N. & Walsh, M. J. Single-cell RNA sequencing reveals new basic and translational insights in the cystic fibrosis lung. Am. J. Respir Cell. Mol. Biol. 68, 131–139 (2022).
Cystic Fibrosis Foundation. Cystic Fibrosis Foundation Patient Registry 2022 Annual Data Report (2023).
Wainwright, C. E. et al. Effect of Bronchoalveolar lavage-directed therapy on Pseudomonas aeruginosa infection and structural lung injury in children with cystic fibrosis: a randomized trial. JAMA 306, 163–171 (2011).
Owens, C. M. et al. Lung clearance index and HRCT are complementary markers of lung abnormalities in young children with CF. Thorax 66, 481–488 (2011).
Hisert, K. B. et al. Restoring cystic fibrosis transmembrane conductance regulator function reduces airway bacteria and inflammation in people with cystic fibrosis and chronic lung infections. Am. J. Respir Crit. Care Med. 195, 1617–1628 (2017).
Harris, J. K. et al. Changes in airway Microbiome and inflammation with Ivacaftor treatment in patients with cystic fibrosis and the G551D mutation. Ann. Am. Thorac. Soc. 17, 212–220 (2020).
Zhang, S. et al. Cystic fibrosis macrophage function and clinical outcomes after elexacaftor/tezacaftor/ivacaftor. Eur. Respir J. 61, 2102861 (2023).
Ciobanu, D. Z. et al. Tezacaftor is a direct inhibitor of sphingolipid delta-4 desaturase enzyme (DEGS). J. Cyst. Fibros (2024).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Yang, S. et al. Decontamination of ambient RNA in single-cell RNA-seq with decontx. Genome Biol. 21, 57 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using cellchat. Nat. Commun. 12, 1088 (2021).
Acknowledgements
We would like to thank our pediatric patients and their families for their generous participation in this study, and we would like to thank the Cystic Fibrosis Center at Lenox Hill Hospital/Northwell Health for graciously entrusting us with the care of their pediatric patients with CF. We would additionally like to thank the Genomics Core Facility and the Mount Sinai Biorepository at the Icahn School of Medicine at Mount Sinai for their essential contributions.
Funding
This work was supported by the Cystic Fibrosis Foundation (JANUSK21D0; awarded to MNJ) as well as the National Center for Advancing Translational Sciences (KL2TR004421; awarded to MNJ) and the National Institute of Diabetes and Digestive and Kidney Diseases (R01DK118946; awarded to MJW) at the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
YS: Methodology, investigation, data curation, formal analysis, visualization, and writing – review and editing; AGV: Writing – review and editing and resources; MBB: Writing – review and editing; MJW: Supervision, methodology, writing – review and editing, funding acquisition, and resources; and MNJ: Conceptualization, methodology, investigation, visualization, writing – original draft, writing – review and editing, funding acquisition, and resources.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, Y., Vicencio, A.G., Beasley, M.B. et al. The single-cell transcriptional landscape of the pediatric cystic fibrosis lung from minimally invasive respiratory specimens. Sci Rep 16, 8113 (2026). https://doi.org/10.1038/s41598-026-36125-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36125-w