Abstract
Multimodal Machine Learning (MML) methods address various efficient ways of driving insights from various data modalities, e.g., in healthcare settings, tabular electronic health records along with other modalities, such as medical imaging, electrocardiogram data (ECG), and textual doctors’ notes and reports. Using deep learning methods, we propose a novel MML approach for mortality prediction in healthcare settings that fuses tabular data, ECG, and written notes in various stages. To this end, this research addresses various challenges related to MML including (1) collecting and building comprehensive data representations from various modalities that may require different preprocessing steps to handle noise and distorted data, (2) ensuring data alignment across modalities, and (3) choosing the optimal fusion strategy (i.e., early, late, or hybrid). This study uses three distinct data modalities: tabular data (encompassing healthcare records, vital signs in real-time, laboratory test results, procedures, and diagnosis records), ECG data, and textual notes from doctors about patients. These modalities are obtained from the MIMIC-IV, MIMIC-ECG, and MIMIC-IV-Note datasets, which include comprehensive medical records, ECG reports, and textual doctors’ notes to explore and evaluate methods in all MML stages. The methodology includes data preprocessing to address noise, outliers, and missing values. It involves comparing fusion strategies (early, late, hybrid) for integrating multimodal data. In addition, novel deep learning models that use attention mechanisms are implemented for better data interaction. Model performance is evaluated with metrics like AUC-ROC, precision, recall, and F-score. The results of our proposed multimodal neural network model using multimodal information showed a substantial increase in performance, with an AUC of 0.96, surpassing the performance of previous single modality literature models. Using multimodal data, the aim is to make the proposed model obtain a holistic view of patient health similar to that of domain experts, resulting in better informed clinical decisions and potentially better clinical outcomes. Our promising results suggest the need to examine biases in training data, such as mortality class imbalances, to improve model performance. Future work should also address the interpretability of complex deep learning models for clinical adoption.
Similar content being viewed by others
Introduction
Multimodal machine learning (MML) leverages diverse data sources, or modalities, to enhance learning by capturing comprehensive data representations. A modality refers to any distinct data structure offering complementary information to the learning process. Integrating visual and linguistic modalities is common in fields such as computer vision and natural language processing1, while combining magnetic resonance imaging (MRI), biospecimen analysis, and clinical assessments is helpful in medical diagnostics2,3,4,5. Various modalities offer distinctive and additional information that allows multimodal machine learning to capture robust representations. Fusing these modalities has been a challenge, where multiple strategies can be used, including early fusion (i.e., all modal data are fused at the beginning), late fusion (i.e., each modality is processed individually and findings are combined at the end of the learning process), and hybrid strategies that include components of both6,7.
This research addresses MML in the context of predicting acute kidney injury (AKI) patient mortality in intensive care units (ICU). We focus on AKI in our study as AKI that requires hospitalization is estimated to impact more than 12% of males and 7% of females at some point in their lifetime8. AKI leads to around 100,000 fatalities per year, making up 15% of critically ill patients, with a fatality rate ranging from 40% to 50%. Identifying people at an early stage who are more likely to experience deterioration, or a subgroup with a high risk of AKI, by comprehensively comprehending various pathogenic proteins, may allow for more personalized and intensive treatment to extend healthy kidney function and enhance patient survival8.
This study uses three distinct data modalities: tabular data (e.g., healthcare records, vital signs, laboratory test, etc.), electrocardiogram (ECG) data, and textual notes from doctors about the patients. The data is obtained from the MIMIC-ECG9 MIMIC-IV-Note, and MIMIC IV10 datasets, which encompass various data sources, including ECG reports, textual reports, chart events, lab events, demographic information, procedure history, and diagnosis history. This work addresses the following challenges to build an efficient and adaptive MML framework. (1) Data preprocessing and alignment: Handling for the three modalities, i.e., tabular, ECG, and textual reports, from multiple sources requires careful consideration for preprocessing and alignment. Tabular data needs noise and outlier removal, along with time-series feature extraction and selection. ECG data and doctors’ reports must undergo several preprocessing steps for accurate representation. Proper temporal alignment of records is essential for prediction, and handling missing data is crucial due to the potential unavailability of certain modalities for some patients. (2) Multimodal fusion: We propose a novel multimodal fusion process that incorporates representation from all three modalities that we selected and processed. This research explores early, late, and hybrid fusion (i.e., incorporating early and late fusion methods) for the MML process. (3) Mortality prediction models: We propose using novel deep learning models to predict the mortality of AKI patients in the ICU. We provide a comprehensive analysis of the proposed model’s performance and how it compares to the performance of the state-of-the-art models. Our experiments demonstrate that the hybrid fusion approach achieves the best overall performance with an AUC of 96.19% and accuracy of 93.60%. The multimodal models consistently outperform both traditional machine learning baselines and unimodal approaches, validating the effectiveness of integrating ECG signals, structured clinical data, and textual notes for mortality prediction.
Contributions. We summarize the main contributions of this work as follows:
-
This research proposes a comprehensive MML framework that integrates tabular, ECG, and textual data for improved mortality prediction in AKI patients. We explore various fusion strategies to leverage the strengths of each data modality.
-
We develop novel deep learning models that utilize attention-based layers to fuse and enhance interactions among various modalities. We handle the unique challenges of aligning heterogeneous multimodal data representations for AKI mortality prediction and demonstrate improved performance compared to unimodal approaches.
-
We conduct extensive experiments and evaluations on large-scale clinical datasets of 21,469 AKI patients, providing valuable insights into the effectiveness of our proposed methods.
Organization. The rest of this paper is organized as follows. Section Related work reviews related work on multimodal machine learning and mortality prediction for acute kidney injury. Section Methodology presents the methodology, including dataset descriptions, preprocessing steps, and details of the proposed multimodal fusion architectures. Section Result and discussion reports the experimental results and provides a discussion of the findings. Section Conclusion concludes the paper and outlines potential directions for future research.
Related work
This section reviews the related works to this research, focusing on multimodal machine learning and AKI mortality prediction.
Multimodal machine learning in healthcare
The study by Yan et al.11 reviews how MML models can represent heterogeneous medical data, including clinical features and time-series data such as ECG signals, and imaging modalities like MRI, toward better accuracy, robustness, and interpretability in diagnostics. Yan et al.11 emphasized that the multimodal techniques are superior to conventional or unimodal methods by pointing out that the integration of several sources of data substantially increases the diagnostic results while providing a holistic perspective on patient health. Similarly, Xu et al.12 proposed a new methodology for predicting ICD-10 diagnostic codes using an MML model. Several models that process unstructured texts, semi-structured texts, and structured tabular data were developed and further combined with ensemble techniques for better performance in accuracy and interpretability. Their approach is based on the MIMIC-III dataset, outperforming baseline models such as TF-IDF and Text-CNN, with results shown by a micro-F1 score of 0.7633 and a micro-AUC of 0.9541. In a study by Dcouto and Pradeepkandhasamy13, the researchers review the progress and challenges of applying several state-of-the-art deep multimodal learning methods to accomplish the early detection of autism spectrum disorder. Various data modalities are identified for use in improving the accuracy of neuroimaging, genomics, and behavioral data. Furthermore, a different research project by Joo et al.14 aims to explore whether a complex deep learning model could be developed using clinical data and pretreatment MR images to predict pathologic complete response to neoadjuvant chemotherapy in patients with breast cancer. Another study by Venugopalan et al.15 proposed to classify various stages of Alzheimer’s disease by combining imaging, genetic, and clinical information.
In the work by Ektefaie et al.16, the study investigates the integration of different data modalities with graph-based techniques for learning and inference over complex structures. The authors proposed a methodological way of enhancing the tasks of classification, segmentation, and prediction by integrating multiple modalities into coherent representations. The work puts into perspective a range of methods that could be applied in disease prediction, analysis of images, and the construction of knowledge graphs. Huang et al.17 presented a fundamental point of view by systematically reviewing the application of various multimodal deep learning fusion techniques. Their contribution accentuated the importance of making consistent terminologies and categorizations regarding fusion strategies, proving that no approach is universally superior to others across all medical domains. In another study by Liu et al.18, the challenges and perspectives related to multimodal research on electronic health records were explored, emphasizing the need for a combination of structured and unstructured data to leverage all dimensions of the available data. Amal et al.19 discusses multimodal data fusion in cardiovascular disease care and proposes the use of normalization techniques to better integrate disparate data formats.
Acute kidney injury and mortality prediction
Several recent works have proposed different models for identifying high-risk AKI patients using both structured and unstructured data. These models improve their prediction accuracy by adopting advanced techniques either in feature extraction and selection or by focusing on particular therapeutic applications31. In Alfieri et al.32, the research explores the development and validation of a deep-learning model developed to predict severe AKI in ICU patients, using hourly urine output data to predict AKI events, especially stages 2 and 3, by retrospectively analyzing two large datasets: eICU and MIMIC-III. Their approach successfully predicted 88% of AKI cases at least 12 hours before their onset. Another study by Yue et al.31 aimed to develop ML models to predict AKI in patients with sepsis. The highest performance was achieved using XGBoost, with an AUC value of 82.10%. In another approach by Liu et al.33, the study presents a deep learning model that makes continuous predictions of severe AKI, considering the changes in urine lab results. Their deep learning model correctly predicted 0.88 of AKI events at a lead time of greater than 12 hours. In Le et al.34, the study focuses on developing and validating a convolutional neural network model for predicting AKI in ICU patients, achieving an AUROC of 86%. Another study by Chang et al.35 investigated the performance of machine learning models in predicting 30-day mortality among AKI patients who require acute renal replacement therapy (RRT) in the ICU. The authors suggest that the implementation of XGBoost in clinical practice might considerably improve patient outcomes in this population. In Hu et al.36, the research studies the early prediction of hospital death among patients admitted with AKI in intensive care units using explainable techniques of artificial intelligence. In Le et al.34, the research introduces a CNN-based machine learning approach to predict AKI in ICU patients up to 48 hours in advance, achieving an AUC of 86%. In hai Bai et al.37, the research proposes a model to predict the risk of mortality at 28 days among elderly patients undergoing continuous RRT for AKI. Their approach achieved an AUC of 80.90%.
Current models in the literature typically depend on individual data modalities, which are insufficient to obtain all aspects of patient information required for precise predictions. This is similar to the approach healthcare professionals take to analyze various data before making their clinical decisions. Multimodal fusion ensures that the advantages of each modality complement one another in enhancing predicted accuracy and resilient model performance. Advanced fusion strategies, such as attention-based mechanisms, improve the interplay among various modalities, leading to a deeper understanding and more precise predictions38. Table 1 lists a comparative summary of the literature focused on mortality prediction in AKI patients.
Methodology
End-to-end MML framework for prediction of AKI mortality using MIMIC-IV tabular data, MIMIC-ECG, and MIMIC-Note datasets. Data cleaning and preprocessing is done per modality. AKI-labeled cohorts are split into stratified K train/validation/test sets. Multiple baseline ML and proposed MML models are built and evaluated using various metrics. Baseline ML: Random Forest (RF), AdaBoost, Decision Tree (DT), KNN, Logistic Regression, SVM, Gradient Boosting (GB).
Data modalities: processing and handling
Datasets. This research uses three large-scale datasets: i.e., MIMIC-IV, MIMIC-ECG, and MIMIC-IV Note. (1) The MIMIC-IV has a variety of medical records, including patient demographics, chart events, lab results, procedure history, and diagnosis history. This acts as a source of primary tabular healthcare records. It manages a total of 16,081,680 chart event records, 122,103,657 lab event data, and a large number of records of demographic, procedure, and diagnostic information belonging to more than 53,000 patients. (2) The MIMIC-ECG provides 800,035 ECG recordings, totaling 12,598,583 data points. These include 10-second segments of 12 signal leads for 268,688 patients. (3) MIMIC-IV Note dataset is a subset of the MIMIC-IV database. It consists of deidentified clinical notes for more than 380,000 unique ICU patients between the years 2008 and 2019. It includes discharge summaries, progress notes, radiology reports, and nursing documentation that provide a source of rich unstructured text data. Figure 1 illustrates the major steps of our proposed framework for multimodal AKI mortality prediction.
Data Preprocessing and feature extraction. For the MIMIC-IV, we analyze chart events, lab events, demographic information, diagnoses information, and the procedural history of patients. From the chart events, we extract the minimum, maximum, mean, and standard deviation for all recorded items during patients’ ICU stays. Considering 2,226 items, we extracted 8,904 features. Similarly, considering 532 lab events, we extract the minimum, maximum, mean, and standard deviation of all items. Lab data contributed 2,128 features to our set of features. For diagnostic and procedure items, MIMIC-IV has 53,134 and 47,868 unique diagnoses and procedures, respectively. We use only items that appear at least once in the target subjects in our research, i.e.,, subjects with AKI. We assign a binary value to each item to indicate whether a patient has a particular diagnosis or procedure. For demographic information, we use only gender information in the features. MIMIC-IV contributes 41,512 features to our data collection. Out of 53,150 subjects included in MIMIC-IV, and after handling data alignments and cleaning, our records include 26,005 subjects identified with AKI, of whom 4,047 are dead and 21,958 are alive.
For the MIMIC-ECG, records are collected and analyzed for our subjects with AKI. Of the 268,688 total subjects in the datasets, we extracted ECG records from 21,962 subjects who were identified with AKI. We use the entire 10-second dataset of 12 leads from the latest record. For subjects with multiple ECG records, we choose to use the most recent recording. This generates data records of 5,000 data points for each lead in 12 leads, i.e., totaling 60,000 temporal data points per subject. The raw dataset underwent a comprehensive preprocessing pipeline to ensure signal quality and consistency for downstream analysis. This included denoising using wavelet transform to suppress noise while preserving critical signal features, baseline wander removal to eliminate low-frequency drift that can obscure key ECG components, and correction for irregular time intervals to maintain uniform temporal resolution. The data is then normalized using min-max normalization.
For the MIMIC-IV-note, we obtain notes for subjects with AKI who also have ECG records. This allows for the collection of notes, ECG records, and tabular data from 21,469 subjects with AKI (3,349 dead and 18,120 alive), who are included in our multimodal fusion analysis. To analyze notes, we use pre-trained large language models (LLM), e.g., Bert39, to obtain embeddings and representations of notes. We use a maximum of 512 tokens to represent the extracted notes. The preprocessing included the removal of non-informative content, such as headers, timestamps, and physician identifiers; conversion of text to lowercase for normalization; and elimination of special characters, digits, and irrelevant punctuation. Tokenization is performed to break down the text, and the [‘UKN’] token is used to represent out-of-vocabulary tokens for the used LLM. The final output comprised BERT-compatible token sequences suitable for generating rich contextual embeddings for downstream tasks, such as mortality prediction. We use the representation of the [‘CLS’] token to represent the input notes.
Feature selection. Feature selection is conducted only on the MIMIC-IV tabular features. Robust representations of data from MIMIC-ECG and MIMIC-Notes are obtained using deep learning-based methods. We utilize mutual information to identify and retain the most important 500 tabular features (i.e., MIMIC-IV features) from a large dataset. Preliminary experiments with feature sizes (e.g., 50, 100, 500, and 1,000) indicate that 500 features yield the most consistent results across folds and settings.
Evaluation and data splitting. We follow a 10-fold stratified cross-validation approach to examine the generalization of the models. 8 folds (training), 1 fold (validation), and 1 fold (testing) are built for each cross-validation split in the ten total splits. Experiments are conducted for 15 evaluation rounds, each including 10-fold cross-validation process. The results from the test set are averaged across folds and are reported in the results. Feature extraction/selection and model training are conducted using training folds in each cross-validation round. This is to ensure that there is no data leakage from the validation/test folds.
Handling data imbalance. Since the dataset is highly imbalanced, with a mortality rate of 15.6% (i.e., 3,349 dead and 18,120 alive subjects), we used class weights to balance the training process. The class weights are calculated as the inverse of the class frequencies, ensuring that the model pays more attention to the minority class during training. This helps mitigate the impact of class imbalance and improves the model’s ability to predict mortality accurately. The class weights are calculated as follows: \(\text {class\_weight} = {\text {total number of samples}} \div ({\text {number of classes} \times \text {number of samples in each class}})\), where the total number of samples is 21,469, the number of classes is 2 (alive and dead), and the number of samples in each class is the number of alive and dead subjects, respectively.
Uni-modal baseline models
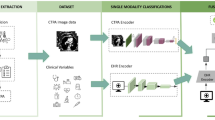
To evaluate the individual contribution of each data modality to mortality prediction, we examine baseline models for tabular data, ECG signals, and clinical notes. For MIMIC-IV data, the models include Random Forest, AdaBoost, Decision Tree, K-Nearest Neighbors (KNN), Logistic Regression, Support Vector Machine with Radial Basis Function kernel (SVM-RBF), and Gradient Boosting. For Random Forest, an ensemble learning method, we use 100 decision trees grown to the maximum extent. For AdaBoost (Adaptive Boosting), another ensemble learning method, we use 100 boosting stages. For KNN, we use \(k=5\). For Decision Tree, Logistic Regression, SVM-RBF, and Gradient Boosting, we follow the default and standard settings to build the classifiers. These models represent a diverse set of approaches to classification, ranging from simple linear models to complex ensemble methods, providing a good baseline for comparison. For MIMIC-ECG, we use a deep learning-based model similar to the one used in the early fusion MML architecture designed specifically for single modality input (see Fig. 2a). For MIMIC-Note modeling, we build a classification head on top of the pre-trained Bert-large39 model.
Proposed MML fusion strategies and model architectural designs. Early Fusion (a) integrates data representations from various modalities in the early stages of the pipeline. Late Fusion (b) processes modalities through branches and combines the representations at late stage. Hybrid Fusion (c) combines various integrations of modalities at various stages in the pipeline.
Multimodal fusion strategies
Early fusion MML. Figure 2a shows the architecture designed to process each modality separately and then fuse their representations into one unified feature vector for subsequent predictive tasks. (1) MIMIC data is provided as tabular data with (1, 500) dimensions representing the selected features of the structured clinical data, processed through a dense neural layer with 5,000 units and ReLU to capture non-linear relationships among clinical variables. (2) ECG data has a shape of (5000, 12), representing the 12 leads with 10-second data recordings. (3) MIMIC-Note data are tokenized into sequences of 512 tokens that pass through a pre-trained BERT-large model, producing a (512, 768) embedding. We use the [‘CLS’] token embeddings to represent the input, i.e., (1, 768) representation, and then pass them to a dense layer of size 5,000 and ReLU.
Representations from the three modalities are then fused using an attention-based module with a multi-head self-attention mechanism (i.e., 4 attention heads) to capture inter-modal interactions, producing a fused representation of shape (5000, 14). We use a CNN architecture to process fused data. The architecture consists of five modules, i.e., one Conv-BatchNorm-ReLU block with 64 filters and four ResNet-SE blocks with 64, 128, 256, and 256 filters. The Conv-BatchNorm-ReLU block applies a 1D Convolution with 64 filters, followed by Batch Normalization (BatchNorm) to stabilize the learning process and speed up convergence, then ReLU activation. This transforms the input feature space from 14 channels to 64 channels, resulting in an output shape of (5000, 64). The ResNet-SE blocks are designed to capture complex patterns in the fused data while maintaining efficient gradient flow through residual connections. Each block consists of two convolutional layers with BatchNorm, followed by a Squeeze-and-Excitation (SE) module that adaptively recalibrates channel-wise feature responses. The first ResNet-SE block has 64 filters and maintains the temporal resolution, while the subsequent blocks increase the number of filters to 128, 256, and 256, respectively, while reducing the temporal resolution through strided convolutions. The output shapes after each ResNet-SE block are (5000, 64), (2500, 128), (1250, 256), and (625, 256), respectively. The final output is globally averaged and passed to the output dense layers to produce the predictive probability values.
Late fusion MML. As shown in Fig. 2a, the late fusion model processes each modality independently before combining their feature representations at a later stage, preserving the unique characteristics of each modality while enabling meaningful interactions during fusion. (1) MIMIC data is processed using a dense layer of 128 units and ReLU activation, followed by a dropout regularization with a dropout rate of 0.3. (2) ECG data is processed using a CNN architecture that includes two Conv-BatchNorm-ReLU blocks (similar to the one described in Early Fusion MML) using 64 and 128 filters, respectively. Max-pooling with a pool size of 2 is applied after the first block, and global average pooling is applied after the second block to generate a representation vector of size 128. (3) MIMIC-Note data is processed similarly to the Early Fusion MML (using a pre-trained BERT model, then a dense layer of size 128 with ReLU and dropout). The outputs from the three modalities are fused to form a unified representation vector of size 384. The fused representation is refined through one dense layer with 128 neurons and ReLU activation and dropout regularization with a dropout rate of 0.5, then passed to the final output layer that produces the probability of mortality.
Hybrid fusion MML. As shown in Fig. 2c, (1) MIMIC data and (2) ECG data are processed similarly to the Late Fusion MML, and their outputs are fused through a fusion layer to create a unified representation of size 256. The combined ECG and tabular data representation is passed through a dense layer of size 128 with ReLU activation and dropout regularization with a dropout rate of 0.3. (3) MIMIC-Note data is processed similarly as in the Early and Late Fusion MMLs then passed through a dense layer of size 128 with ReLU activation and dropout regularization with a dropout rate of 0.3. The output from the combined ECG and tabular data is then fused with the clinical notes’ output through another fusion, producing a unified representation of size 256. The fused representation is passed to one dense layer with 128 neurons with ReLU and dropout with a dropout rate of 0.5, then passed to the final output layer to predict mortality.
Result and discussion
This section presents the experimental results of the proposed MML framework for mortality prediction in AKI patients using MIMIC-IV, MIMIC-ECG, and MIMIC-Note datasets. We first present the metrics and experimental setup used in our study. This is followed by presenting the performance of unimodal baseline models for each data modality, followed by the results of the proposed multimodal fusion strategies. A comprehensive statistical analysis and discussion of the findings is also provided.
Evaluation metrics and experiments setup
Metrics. We use the following evaluation metrics to assess the performance of our models. \(\text {Precision} = \text {TP} / \text {(TP + FP)}\), \(\text {Recall} = \text {TP} / \text {(TP + FN)}\), \(\text {F-score} = 2 \times {(\text {Precision} \times \text {Recall})} /\) \({\text {(Precision + Recall)}}\), and \(\text {AUC} = \int _{0}^{1} \text {TPR} \cdot \text {FPR} \, d\text {(FPR)}\), where \(\text {FPR} = \text {FP} / \text {(FP + TN)}\), \(\text {TPR} = \text {TP} / \text {(TP + FN)}\), \(d\text {(FPR)}\) the change in FPR, FP, FN, TP, and TN represent False Positives, False Negatives, True Positives, and True Negatives, respectively. We also measure the Brier score, which provides information on the accuracy of probabilistic predictions, with lower values indicating a better calibration of predicted probabilities.
Deep learning training hyperparameters. For mortality prediction, all models are trained to minimize the binary cross-entropy loss with the Adam optimizer. The learning rate for the optimizer follows exponential decay with an initial value of \(1e-3\), decay steps 10, 000, and a decay rate of 0.9. Training runs for up to 100 epochs with a batch size of 128 on the training dataset and validates on the validation dataset with class weights applied. We implemented an early stopping technique that monitors the validation accuracy with a patience of 20 and restores the best weights if no improvements are observed. We adopt L2 regularization on all layer parameters (i.e., weights) with a coefficient of \(1e-4\) to reduce and combat overfitting.
Uni-model results
Selected features from MIMIC-IV data. In our tabular data used for the experiments, feature selection identified the top highly correlated features and grouped them into clinically meaningful categories that are aligned with standard clinical assessments. The liver function group included markers such as AST, ALT, LDH, and INR, which indicate hepatic stress and metabolic activity. Renal function was represented by creatinine, BUN, and phosphate-related values, capturing kidney performance. Respiratory parameters included respiratory rate, inspired oxygen, \(pO_{2}\), and pH, reflecting ventilation and gas exchange. Acid-base balance was assessed using lactate, base excess, bicarbonate, and total \(CO_{2}\), offering insight into metabolic compensation. Neurological status combined GCS components, Richmond-RAS scores, and strength measures to reflect consciousness and motor function. Skin and pressure ulcer risk relied on Braden scale subcomponents like mobility and nutrition. Cardiovascular function included heart rate and both invasive and non-invasive blood pressure measurements. These groupings provide a structured view of patient physiology and support modeling while maintaining clinical relevance.
Uni-modal model performance. Table 2 presents the performance metrics for models trained on different data modalities from the MIMIC datasets, i.e., tabular data, ECG data, and textual clinical notes. Accuracy, AUC, precision, recall, and F1-score are reported for each model. The tabular data models achieve the highest accuracy of 92.92 (mean of 91.90±0.60% using Gradient Boosting) and a strong AUC of 80.37% (77.21±1.60 using AdaBoost). In terms of F1-score, Gradient Boosting attains the highest value of 73.02% (mean of 68.65±2.67%). This indicates a gap between precision and recall, with precision being significantly higher than recall in most models (except DT and SVM), suggesting that while the models are good at identifying true positives, they miss a substantial number of actual positive cases. Generally, the tabular data models show that tabular clinical data contains valuable information for mortality prediction. Regarding other modalities, the ECG-based model performs achieves the highest AUC of 94.83% (mean of 79.12±9.55%) and F1-score of 87.87% (mean of 55.76±15.13%), making it the best at identifying positive cases, with an accuracy of 90.77% (mean of 74.60±9.27%) and recall of 89.28% (mean of 43.45±19.50%). Considering multiple evaluation runs, the ECG model shows high variability in performance, particularly in recall, indicating sensitivity to training conditions or data splits. The textual clinical notes model achieves an AUC of 86.91% (mean of 78.67±7.47%) and a precision of 87.50% (mean of 77.05±5.89%), indicating strong performance in identifying true positives among predicted positives, with an accuracy of 82.40% (mean of 72.10±7.47%). The textual model struggles with recall (i.e., 41.94% (mean of 35.32±5.64%)), and consequently F1-score, as it misses a significant number of actual positive cases. While unimodal models provide demonstrate data relevance from different modalities, their limitations in capturing the full complexity of patient data highlight the need for multimodal approaches to enhance predictive performance.
Multi-model results and statistical analysis of results
Performance comparison of multi-models. (1) Early fusion model As shown in Table 3, the model evaluation shows strong performance, with an accuracy of 92.20% shows that the model correctly classifies most cases. The high AUC of 94.60% reflects excellent ability to distinguish between positive and negative cases. Regarding classification, the model achieves a recall of 76.02%, identifying most positive cases. The precision of 75.36% indicates moderate confidence in positive predictions. Furthermore, the model correctly identifies a high number of true negatives (with an average of 96.47%), reflecting its ability to detect positive outcomes. While the model prioritizes recall, ensuring most positive cases are captured, its precision is slightly lower, indicating room for improvement in reducing false positives. These metrics suggest a well-rounded model that is effective in distinguishing between classes, with a strong focus on minimizing missed positive cases, which is critical in applications such as mortality prediction in AKI patients.
(2) Late fusion model As shown in Table 3, the model with a late fusion strategy performs exceptionally well compared to the early fusion model, achieving a high accuracy of 93.50% and an excellent AUC of 96.80%. The results reflect highly calibrated predictions, with an effective trade-off between precision of 78.43% and recall of 81.87%.
(3) Hybrid fusion model The evaluation results of the hybrid model evaluation show the strong performance of the model with an accuracy of 93.60%, an AUC of 96.16%, a precision of 82.33%, a recall of 76.32%, and an F1-score of 79.21%. Table 3 summarizes the performance of the proposed models with different fusion strategies, indicating the outperformance of the hybrid model compared to early and late fusions.
Distribution of performance metrics (AUC, accuracy, precision, recall, and F1-score) across early, late, and hybrid models for AKI mortality prediction after 15 evaluation runs. The box plots illustrate the variability and central tendency of each metric, providing insight into the comparative performance of different modeling approaches.
Statistical analysis of results. We assess the stability and variability of metrics across the multiple evaluation runs of the models. Figure 3 illustrates the distribution of metrics across early, late, and hybrid models after 15 evaluation runs. The AUC and accuracy distributions indicate that hybrid models achieve more stable and consistently high performance, while early models exhibit greater variability. Precision is highest in early models but with substantial spread, suggesting inconsistency, whereas late and hybrid models demonstrate more stable precision values. Recall is notably higher in early models but at the cost of lower precision, highlighting a trade-off between sensitivity and specificity. The F1-score distribution shows that late and hybrid models balance precision and recall more effectively than early models. These results suggest that hybrid models provide a more reliable trade-off among metrics, making them a strong candidate for AKI mortality prediction.
One-way ANOVA and HSD analysis. For further analysis of results, a one-way ANOVA was conducted to compare the performance of early, late, and hybrid models across metrics. The results indicated significant differences in AUC (p = 0.00445), Accuracy (p = 0.0114), Recall (p = 0.0293), and F1-score (p = 0.00443), while Precision (p = 0.287) did not show statistical significance. A post-hoc Tukey’s Honest Significant Difference (HSD) test was performed to identify specific model comparisons. The results showed that for AUC, the early model was significantly different from both the hybrid (p = 0.0080) and late models (p = 0.0156). Accuracy was significantly different between the early and hybrid models (p = 0.0133). Recall exhibited a significant difference between the early and late models (p = 0.0347). For F1-score, significant differences were observed between the early and hybrid models (p = 0.0084) and between the early and late models (p = 0.0145). Considering the ANOVA and HSD results and performance shown in Table 3 and Fig. 3, the hybrid model emerges as the best overall choice. It achieves the highest AUC, accuracy, and competitive F1-score while maintaining a reasonable recall. Although the late model exhibits the best recall, the hybrid model provides a better balance across multiple evaluation metrics, making it the optimal choice for robust mortality prediction in AKI patients. The hybrid model demonstrates a well-balanced performance across all confusion matrix components. It correctly identifies 261 true positive cases and 1742 true negatives, misclassifying only 81 false negatives and 56 false positives. Compared to the early fusion model, which records more false positives (85) and slightly more false negatives (82), the hybrid model shows clear improvement in both precision and recall. On the other hand, the late fusion model achieves the highest number of true positives (280), indicating superior recall performance. However, this comes at the cost of a higher number of false positives (77), which may lead to over-alerting in clinical settings. However, the hybrid model offers a more stable trade-off (i.e., fewer false alarms than the late model and fewer missed deaths than the early model). This confirms that the hybrid fusion strategy maintains robustness in real mortality prediction tasks.
Comparison with related work. Finally, the reviewed studies, shown in Table 1, vary in focus, from evaluating clinical scoring systems like RIFLE and KDIGO, to using statistical features such as RDW, to machine learning models including logistic regression, CNNs, and XGBoost. Most studies use the MIMIC-III or MIMIC-IV datasets. Reported results range from AUROCs of 0.76 to 0.89 in traditional models, with few exceeding 0.90. The last row highlights our proposed multimodal learning (MML) approach, which integrates structured and unstructured data from MIMIC-IV, ECG, and clinical notes, achieving a higher accuracy (93.6%) and AUC (0.961), reflecting superior performance over prior models.
Limitations and future directions
We acknowledge several limitations in our study that warrant consideration and suggest avenues for future research. We discuss these limitations in terms of dataset generalizability, model interpretability, and class imbalance/fairness considerations.
Dataset and generalizability limitations. While the MIMIC datasets provide comprehensive clinical data, they represent a single-center dataset, which may limit generalizability across different healthcare systems, populations, and clinical practices. The 15.6% mortality rate in our AKI cohort, while substantial, may not reflect mortality rates across all healthcare institutions with varying patient demographics, severity distributions, and care protocols. Additionally, the retrospective nature of the data may introduce selection bias, as patients with complete multimodal data (ECG, structured data, and notes) may represent a specific subset of the AKI population. In terms of modality availability challenge, our study requires patients to have all three data modalities available, reducing the cohort from 26,005 to 21,469 subjects. This requirement may introduce bias toward patients with more comprehensive monitoring, potentially excluding those with less severe conditions or shorter ICU stays. The reliance on structured data extraction may miss subtle clinical nuances captured in free-text documentation, and the quality of ECG recordings and clinical notes may vary significantly across different time periods and clinical contexts. Future work should develop robust strategies for missing modalities and evaluate model performance when certain modalities are unavailable.
Model interpretability and clinical adoption. The complexity of our multimodal deep learning approach presents significant challenges for clinical interpretability, which is crucial for healthcare adoption. While our model achieves high predictive performance, clinicians require an understanding of why specific predictions are made. Future work should incorporate explainable AI techniques, including attention visualization mechanisms to highlight important ECG segments, clinical features, and note excerpts that contribute to mortality predictions. Additionally, developing feature importance analysis and providing uncertainty quantification would enhance clinical trust and support evidence-based decision-making.
In terms of temporal dynamics, our current approach uses static snapshots of patient data rather than modeling dynamic temporal trajectories of patient deterioration. This limitation overlooks the temporal evolution of AKI and mortality risk over the course of ICU stay. Future research should incorporate time-series analysis and dynamic risk assessment capabilities that can continuously update predictions as new data becomes available. Additionally, the computational requirements for real-time inference, particularly for ECG processing and BERT-based text analysis, present challenges for clinical deployment that require optimization for edge computing environments.
Class imbalance and fairness considerations. Despite using class weights to address the 15.6% mortality rate imbalance, the model may still exhibit bias toward the majority class, potentially affecting performance in subgroups with different baseline mortality rates. Future work should examine model performance across demographic subgroups (age, gender, ethnicity) and AKI severity stages to ensure equitable predictions. Additionally, investigating the impact of socioeconomic factors and healthcare access patterns on model performance would enhance understanding of potential algorithmic bias.
Conclusion
Many studies show that early identification of high-risk patients allows timely medical interventions, which can reduce AKI-related mortality and optimize the allocation of ICU resources. This study introduced a multimodal machine learning framework to enhance mortality prediction for AKI patients in ICUs. By integrating structured clinical data, ECG signals, and radiology notes, the model demonstrated superior predictive accuracy compared to unimodal approaches. The findings show that the proposed hybrid fusion model outperforms the early and late fusion methods, achieving 93.60%, 96.19%, 82.33%, 76.32%, and 79.21%, with the highest accuracy of 93.60%. The early fusion model achieves 92.20%, 94.60%, 75.36%, 76.02%, and 75.69%, while late fusion achieves 93.50%, 96.80%, 78.43%, 81.87%, and 80.11%, for accuracy, AUC, precision, recall, and F1-score, respectively. Several key insights emerge from this work: (1) the complementary nature of different data modalities significantly enhances prediction accuracy, suggesting that a holistic view of patient data is crucial for reliable mortality prediction; (2) hybrid fusion approaches provide a better balance between local and global feature interactions compared to early or late fusion alone; (3) the statistical analysis reveals that model performance is not only superior in terms of accuracy but also more stable and consistent across different evaluation metrics; and (4) while the model shows promising results, future work should focus on improving interpretability and addressing potential biases in training data to ensure clinical applicability. These findings highlight the potential of multimodal learning in advancing clinical decision support systems in critical healthcare settings.
Data availability
MIMIC-IV dataset is available by requesting it on https://physionet.org/content/mimiciv/2.2/. The MIMIC-ECG dataset is available; you can request it on https://physionet.org/content/mimic-iv-ecg/1.0/. MIMIC-IV Note dataset is available by requesting it on https://www.physionet.org/content/mimic-iv-note/2.2/.
References
Yu, G., Qin, L., Geng, Y. & Wan, Q. Computer vision and natural language processing. In Practical Machine Learning Illustrated with KNIME, 275–304. (Springer, 2024).
Adrienne, K. et al. Multimodal machine learning in precision health: A scoping review. npj Digital Medicine, 5, 171 (2022).
Yang, H., Kuang, L. & Xia, F. Q. Multimodal temporal-clinical note network for mortality prediction. J. Biomed. Semant. 12, 1–14 (2021).
Fatemeh, B. & Mohammad, S. A. An overview of deep learning methods for multimodal medical data mining. Expert Syst. Appl. 200, 117006 (2022).
George, K. K. et al. Development and external validation of multimodal postoperative acute kidney injury risk machine learning models. JAMIA open 6, ooad109 (2023).
Sören, R. S., Benjamin, U. & Jane, S. Multimodal deep learning for biomedical data fusion: a review. Briefings in Bioinformatics, 23, bbab569 (2022).
Rahate, A., Walambe, R., Ramanna, S. & Kotecha, K. Multimodal co-learning: challenges, applications with datasets, recent advances and future directions. Inf. Fusion 81, 203–239 (2022).
John, A. K. et al. Acute kidney injury. Nat. Rev. Dis. Primers, 7, 1–17 2021.
Brian, G. et al. Mimic-iv-ecg-diagnostic electrocardiogram matched subset. Type: dataset 6, 13–14 (2023).
Alistair, J. et al. PhysioNet. Available online at: https://physionet.org/content/mimiciv/1.0/, pages 49–55, (2020). (accessed August 23, 2021).
Keyue, Y., Tengyue, L., João, A. L. M., Juntao, G., & Simon, J. F. A review on multimodal machine learning in medical diagnostics. Math. Biosci. Eng, 20, 8708–8726 (2023).
Keyang, X. et al. Multimodal machine learning for automated icd coding. In Machine learning for healthcare conference, 197–215, (2019).
Sheril, S. D. & Jawahar, P. Multimodal deep learning in early autism detection—recent advances and challenges. Eng. Proc. 59, 205 (2024).
Sunghoon, J. et al. Multimodal deep learning models for the prediction of pathologic response to neoadjuvant chemotherapy in breast cancer. Sci. Rep., 11, 18800 (2021).
Janani, V., Li, T., Hamid, R. H. & May, D. W. Multimodal deep learning models for early detection of alzheimer’s disease stage. Sci. Rep., 11, 3254, (2021).
Ektefaie, Y., Dasoulas, G., Noori, A., Farhat, M. & Zitnik, M. Multimodal learning with graphs. Nat. Mach. Intell. 5, 340–350 (2023).
Shih-Cheng, H., Anuj, P., Saeed, S., Imon, B., & Matthew, P. L. Fusion of medical imaging and electronic health records using deep learning: a systematic review and implementation guidelines. NPJ digital medicine, 3, 136, (2020).
Ziyi, L. et al. Machine learning for multimodal electronic health records-based research: Challenges and perspectives. In China Health Information Processing Conference 135–155, (2022).
Saeed, A. et al. Use of multi-modal data and machine learning to improve cardiovascular disease care. Front. Cardiovasc. Med., 9, 840262, (2022).
Ramazan, E. E. et al. Comparison between rifle, akin, and kdigo: acute kidney injury definition criteria for prediction of in-hospital mortality in critically ill patients. Iran. J. Kidney Dis., 14(5), 365, (2020).
Joana, G., José, A. F., Sofia, J., João, G. & José, A. L. Neutrophil, lymphocyte and platelet ratio as a predictor of mortality in septic-acute kidney injury patients. Nefrologia (English Edition), 40(4), 461–468, (2020).
Jia, L. et al. Red blood cell distribution width predicts long-term mortality in critically ill patients with acute kidney injury: a retrospective database study. Sci. Rep. 10(1), 4563 (2020).
Zheng-hai, B. et al. A nomogram to predict the 28-day mortality of critically ill patients with acute kidney injury and treated with continuous renal replacement therapy. Am. J. Med. Sci., 361(5), 607–615 (2021).
Francesca A. et al. A deep-learning model to continuously predict severe acute kidney injury based on urine output changes in critically ill patients. J. Nephrol., 34(6), 1875–1886 (2021).
Sidney, L. et al. Convolutional neural network model for intensive care unit acute kidney injury prediction. Kidney Int. Rep., 6(5), 1289–1298 (2021).
Yue, S. et al. Construction and validation of a risk prediction model for acute kidney injury in patients suffering from septic shock. Disease markers 2022(1), 9367873 (2022).
Chang, H. et al. Application of interpretable machine learning for early prediction of prognosis in acute kidney injury. Comput. Struct. Biotechnol. J. 20, 2861–2870 (2022).
Hsin-Hsiung, C. et al. Predicting mortality using machine learning algorithms in patients who require renal replacement therapy in the critical care unit. J. Clin. Med., 11(18), 5289 (2022).
Javier, A. N. et al. Prediction of mortality and major adverse kidney events in critically ill patients with acute kidney injury. Am. J. Kidney Dis., 81(1), 36–47 (2023).
Gao, T. et al. Machine learning-based prediction of in-hospital mortality for critically ill patients with sepsis-associated acute kidney injury. Ren. Fail. 46(1), 2316267 (2024).
Yue, S. et al. Construction and validation of a risk prediction model for acute kidney injury in patients suffering from septic shock. Disease markers 2022, 9367873 (2022).
Francesca, A. et al. A deep-learning model to continuously predict severe acute kidney injury based on urine output changes in critically ill patients. J. Nephrol., 34, 1875–1886 (2021).
Liu, W., Qiu, J.-L., Zheng, W.-L. & Bao-Liang, L. Comparing recognition performance and robustness of multimodal deep learning models for multimodal emotion recognition. IEEE Trans. Cogn. Dev. Syst. 14, 715–729 (2021).
Sidney, L. et al. Convolutional neural network model for intensive care unit acute kidney injury prediction. Kidney Int. Rep., 6, 1289–1298 (2021).
Hsin-Hsiung, C. et al. Predicting mortality using machine learning algorithms in patients who require renal replacement therapy in the critical care unit. J. Clin. Med. 11, 5289 (2022).
Chang, H. et al. Application of interpretable machine learning for early prediction of prognosis in acute kidney injury. Comput. Struct. Biotechnol. J. 20, 2861–2870 (2022).
Zheng hai, B. et al. A nomogram to predict the 28-day mortality of critically ill patients with acute kidney injury and treated with continuous renal replacement therapy. Am. J. Med. Sci., 361, 607–615 (2021).
Junwei, D., Jiaqi, X., Yinghui, L. & Weiping, D. Deep learning based multimodal biomedical data fusion: An overview and comparative review. Inf. Fusion 112, 102536 (2024).
Jacob, D., Ming-Wei, C., Kenton. L. & Kristina, T. Bert: Pre-training of deep bidirectional transformers for language understanding. In Proceedings of the 2019 conference of the North American chapter of the association for computational linguistics: human language technologies, volume 1 (long and short papers), 4171–4186, (2019).
Funding
We acknowledge the Fundamental Research Grant Scheme grant number FRGS/1/2022/ICT02/UKM/02/7, funded by the Ministry of Higher Education (MOHE) Malaysia.
Author information
Authors and Affiliations
Contributions
Husam Abuhamad: Writing - original draft, Visualization, Methodology, Formal analysis. Suhaila Zainudin: Writing review & editing, Formal analysis, Investigation, Supervision. Azuraliza Abu Bakar: Writing - review & editing, Formal analysis, Supervision, Resources, Investigation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abuhamad, H., Zainudin, S. & Abu Bakar, A. Integrative multimodal hybrid data fusion for mortality prediction. Sci Rep 16, 5803 (2026). https://doi.org/10.1038/s41598-026-36296-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36296-6