Abstract
Electrocardiography is a cornerstone in the diagnosis of cardiovascular diseases; however, accurate interpretation demands expert knowledge and is often impeded by data scarcity and annotation costs. To address these challenges, we propose SimCardioNet, a hybrid self-supervised and supervised deep learning framework for multi-class electrocardiography image classification. SimCardioNet leverages a custom multi-scale convolutional neural network backbone enhanced with residual connections and multi-head self-attention, pretrained via a modified SimCLR contrastive learning strategy that integrates a hybrid loss combining InfoNCE and cosine similarity. Following self-supervised pretraining, the model undergoes supervised fine-tuning with progressive layer unfreezing to mitigate overfitting and preserve meaningful representations. We evaluate SimCardioNet across three distinct ECG image datasets: (1) a 4-class Pakistani clinical ECG dataset (Dataset I), (2) an external Kaggle electrocardiography dataset for out-of-distribution validation (Dataset II), and (3) the large-scale PTB-XL benchmark (Dataset III) covering five diagnostic superclasses. On Dataset I, SimCardioNet achieves 0.975 accuracy, 0.973 precision, 0.973 recall, and 0.972 F1-score under 3-fold cross-validation. On Dataset II, the model demonstrates perfect classification performance (1.00 accuracy, precision, recall, and F1-score), highlighting strong generalization. On the PTB-XL dataset (Dataset III), SimCardioNet attains 0.921 accuracy and 0.921 F1-score, outperforming current state-of-the-art models including dual-branch CNNs, entropy-enhanced CNNs, and Bi-GRU architectures. Ablation studies confirm the critical contributions of self-supervised pretraining, attention mechanisms, and domain-specific augmentations. Grad-CAM visualizations further validate the model’s focus on clinically relevant Electrocardiography regions. Our results underscore SimCardioNet’s potential to reduce reliance on labeled data while delivering robust, interpretable, and clinically viable Electrocardiography classification especially valuable in resource-constrained settings.
Similar content being viewed by others
Introduction
Cardiovascular diseases (CVDs) continue to be the primary cause of death worldwide and have placed an unprecedented burden on healthcare systems that need to effectively manage and diagnose a growing number of patients1,2. Electrocardiography (ECG) is a frequently used tool that is routinely used to assess cardiac electrical activity and detect a wide range of cardiovascular disorders and diseases3. However, accurately interpreting an ECG requires considerable clinical expertise and time, resulting in critical bottleneck effects on clinical workflows and delays in potentially life-saving timely interventions4,5. This is an even more critical issue in resource-limited settings, which typically have limited access to cardiologists6,7.
The development of artificial intelligence (AI) and machine learning (ML) has created exciting possibilities for automating ECG analysis. In particular, deep learning (DL) strategies have shown the ability to analyze ECGs with performance comparable to that of expert physicians8,9. Convolutional neural networks (CNNs) are a common architecture used for ECG classification tasks, and can learn both the temporal information and morphology of raw ECG signals10,11,12. Evidence supports the ability of AI to identify numerous cardiac issues, including arrhythmias, myocardial infarction, and structural heart disease, in diverse patient cohorts13,14. In fact, recent studies have shown classification accuracies above 95% for tasks on well-validated datasets, such as the MIT-BIH Arrhythmia Database and PTB-XL dataset, which emphasizes the downstream clinical utility of AI-enabled ECG interpretation15.
Despite these advances, a central limitation persists in traditional supervised learning paradigms: the requirement for extensive volumes of expertly annotated data16,17. In the medical imaging and signal domains, compiling sufficiently large and diverse labelled datasets is hampered by the labor-intensive demands of expert annotation, pronounced inter-observer variability, and the rarity of certain pathological conditions18,19. For ECG analysis, these barriers are further exacerbated by the necessity for cardiologist-level expertise in generating high-quality diagnostic labels, rendering the development of comprehensive training datasets both costly and logistically challenging20.
To overcome these data constraints, self-supervised learning (SSL) has emerged as a compelling paradigm that enables the extraction of meaningful representations from vast amounts of unlabeled data through innovative pretext tasks21,22. A cornerstone of this approach is contrastive learning, which operates by maximizing the similarity between differently augmented views of the same sample while minimizing the similarity among views of distinct samples, thereby facilitating robust feature learning in the absence of labelled data23,24. The contributions of this study are as follows:
-
Developed a custom ECG CNN backbone leveraging multi-scale convolutional blocks, residual connections, and attention mechanisms for enhanced feature extraction from ECG data.
-
Applied SimCLR for SSL to learn representations from ECG images, utilizing multi-scale augmentations for improved model robustness.
-
Introduced a hybrid SSL loss function combining InfoNCE and cosine similarity to stabilize contrastive learning and ensure effective representation learning.
-
Implemented a progressive unfreezing strategy during supervised fine-tuning to gradually unfreeze model layers and reduce overfitting.
-
Conducted 3-fold cross-validation to assess the model performance.
-
Demonstrated improved performance with SSL pre-training, achieving higher classification accuracy and robustness compared to existing models.
-
Highlighted the scalability of the approach for other medical imaging tasks with limited labelled data, ensuring minimal labeling effort in real-world deployments.
Literature review
Recent developments in DL have dramatically altered the landscape of automated ECG analysis and improved the precision of cardiovascular disease diagnosis. The process of interpreting an ECG has mostly been prescribed by clinicians to interpret the raw waveform signals. The advent of CNNs and hybrid architectures has enabled the automated classification of ECG, both as raw signals and images produced by the ECG, and has significantly enhanced diagnostic performance. In the study by Ao et al.25,26, CNNs were only trained on images of 12-lead ECGs and demonstrated effective classification of a larger variety of cardiac pathologies to a similar degree as the models based on the raw signals. This demonstrates the potential of image-based approaches, which are particularly beneficial in clinical care contexts when only images of ECGs are provided (e.g., printouts or scanned), leading to improved access to testing in resource-limited or remote conditions.
Despite these successes, established DL models still face the major limitation of operating on expensive, large-scale, and expertly labelled data. Producing significant volumes of labelled data from experts can be costly and time-consuming. Solutions are needed to tackle this, and researchers have explored SSL and domain adaptation techniques to build representation learning pipelines for large amounts of unlabeled electrocardiographic data. For example, Niu et al.27 suggested adversarial domain adaptation approaches to appropriately address data distribution differences between patient populations while reducing reliance on labelled samples and highlighted the necessary conditions for extracting strong and generalizable features to develop a generalizable classifier for ECG data across various clinical contexts.
Hybrid DL frameworks that integrate CNNs with RNNs such as LSTM units have exhibited capability to model the temporal properties of ECG signals. This type of modeling is effective because both CNNs and RNNs can model sequential dependencies while filtering noise and merging spatial and temporal features from raw data28. These hybrid approaches exemplify the powerful synergy between spatial and temporal modeling in comprehensive ECG analysis. Saranya et al.29 proposed DenseNet-ABiLSTM, leveraging DenseNet for spatial feature extraction and BiLSTM for temporal sequence learning. Their work on photoplethysmography (PPG) signals showed hybrid models’ capability to handle spatial and temporal complexities, improving accuracy in multiclass arrhythmia detection. The model highlighted modality-specific signal representation and integrated residual and recurrent connections to preserve features while minimizing information loss. This approach aligns with medical AI research where convolutional backbones combine with attention or recurrent modules to address local and long-range dependencies across clinical datasets.
In the context of arrhythmia detection, both CNN- and RNN-based models have been extensively benchmarked and shown to achieve reliable performance across a wide range of arrhythmia types. Enhancements, such as deep dictionary learning and wavelet transform pre-processing, further improve feature representation and reduce susceptibility to noise3,30. However, challenges remain in achieving a balance among model complexity and interpretability, especially when translating algorithmic output into clinically actionable information.
Attention mechanisms and multi-scale feature extraction have become essential components of sophisticated CNN architectures for ECG image analysis. These modules enable the models to focus on diagnostically significant waveform segments, such as ST-segment elevations or abnormal P-wave morphologies, while preserving crucial spatial and morphological features. Incorporating attention modules and residual connections has been shown to increase sensitivity to multi-scale ECG patterns and strengthen classification robustness25,31. Recent studies on cardiovascular disease prediction show a shift from classical classifiers to pipelines combining feature selection with stronger learners and deep models. Surveys across k-NN, decision trees, Naive Bayes, SVMs, and neural networks emphasize that selecting salient variables is as crucial as the model itself. Chi-square, correlation-based filters, and LASSO reduce redundancy and improve generalization, showing accuracy gains after selection versus full feature sets. Recent work explores hybrid architectures pairing feature selection with neural components, reporting higher predictive accuracy on Cleveland datasets, while noting caveats about tuning and dataset scope32. Cardiovascular disease remains a leading cause of mortality, and predicting its onset from clinical data is complex. Machine learning supports early risk assessment using k-nearest neighbor and ensemble models like XGBoost, AdaBoost, and Random Subspace. Feature relevance is quantified with a linear SVM measure, exploring diverse feature combinations to improve discrimination. Experiments on the UCI heart-disease dataset (80/20 train–test split) in MATLAB R2020b show strong results, with accuracy of 96%, precision of 97%, sensitivity of 95%, F-measure of 95%, Matthews correlation coefficient of 0.93, false-positive and false-negative rates of 4.53% and 3.10%, and true-positive rate of 96%. These findings indicate ensemble learning with feature selection yields robust heart-disease prediction33,34.
Large, diverse datasets, such as PTB-XL and the Chapman ECG database, have contributed significantly to the training of models that generalize well across different patient populations, providing the necessary breadth of labelled and unlabeled ECG data35,36. Recent studies have also explored hybrid CNN Variational Autoencoder (VAE) architectures and innovative models such as DRA-ECG, which employ strategies such as depthwise separable residual attention, adaptive focal loss, and 2D scalogram transformation, further advancing the robustness and reliability of automated ECG analysis37,38,39.
Recent research in CVD prediction highlights ensemble learning and meta-heuristic optimization to overcome limitations of traditional machine learning models, which often struggle with heterogeneous clinical datasets. While DL approaches achieve strong accuracy, they remain computationally intensive. Kumar and Rekha40 proposed an improved Hawks Optimizer-based ensemble framework for optimized feature selection and classification. Validated on the Kaggle CVD dataset, their method achieved 97% accuracy, 98% precision, and 96% sensitivity, outperforming classical and DL models. Recent advancements in deep learning have significantly improved ECG-based arrhythmia classification, particularly through the use of large-scale datasets like PTB-XL. Atwa et al.41 proposed two interpretable deep learning models–a custom dual-branch CNN and a modified VGG16–both integrating raw 12-lead ECG signals with demographic features (age and gender) to improve diagnostic accuracy. Evaluated across binary, 5-, 10-, and 15-class classification tasks, their CNN model achieved up to 97.78% accuracy in binary settings and 79.7% in multiclass scenarios, outperforming both the VGG16 variant and existing state-of-the-art approaches such as CNN-LSTM and entropy-augmented CNNs. Crucially, the study emphasized model interpretability using SHAP to quantify lead-specific contributions, aligning AI decisions with clinical knowledge, for instance, highlighting leads V2–V4 for myocardial infarction and aVR for ST-T changes. This work underscores the value of combining physiological signals with contextual patient data and explainable AI techniques to build clinically trustworthy and high-performing arrhythmia detection systems. Smigiel et al.42 evaluated three deep learning models on the PTB-XL dataset CNN, SincNet, and a CNN enhanced with entropy features (e.g., Shannon, Sample, Permutation entropy) for 2-, 5-, and 20-class ECG classification. The entropy-augmented CNN achieved the highest accuracy (89.2%, 76.5%, and 69.8%, respectively), outperforming the other architectures, while the baseline CNN offered the best computational efficiency due to its lightweight design.
Furthermore, Geng et al.43 proposed a multi-task deep learning model for ECG classification that combines a shared SE-ResNet feature extractor with task-specific classification heads enhanced by a Contextual Transformer (CoT) attention mechanism and a Bi-GRU layer. Evaluated on the PTB-XL and CPSC2018 datasets, their approach achieved macro F1-scores of 83.3% and 82.7%, respectively outperforming several state-of-the-art single-task baselines. The multi-task framework leveraged hierarchical label information (e.g., grouping bundle branch blocks into a common superclass) to improve generalization, while the CoT mechanism effectively captured both local and global temporal dependencies in 12-lead ECG signals. Ablation studies confirmed the critical contribution of both the CoT block and Bi-GRU module, with the CoT attention yielding the most significant performance gain.
Overall, the literature reflects a decisive shift toward advanced DL paradigms, including contrastive SSL, adversarial domain adaptation, hybrid CNN-RNN architectures, and attention-based CNNs for enhanced ECG classification. While much research remains focused on raw signal analysis, the clinical viability of image-based ECG interpretation has motivated the development of sophisticated pipelines that integrate robust data augmentation, multi-scale feature extraction, and comprehensive evaluation. By adopting such integrative approaches, researchers are increasingly equipped to address challenges related to data scarcity, model robustness, and the heterogeneity of ECG diagnostic categories, ultimately paving the way for clinically deployable AI-powered cardiovascular diagnostic systems.
Methodology
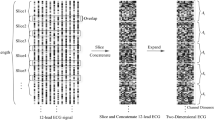
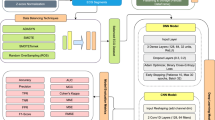
This section delineates the procedures employed in the development and evaluation of the ECG Classification Model through SSL and supervised fine-tuning. The process was structured into five stages: data preparation, model development, self-supervised pre-training, supervised fine-tuning, and performance evaluation. Figure 1 shows the structure of SimCardioNet, which serves as a complete depiction of the proposed model and the stages of ECG classification that we will go through.
Architecture of the SimCardioNet model for ECG classification, integrating pre-processing, CNN-based feature extraction, contrastive learning through SimCLR, and classification using ResNet blocks.
Figure 1 presents the overall configuration of the SimCardioNet model used for ECG image classification. The first stage in the pipeline is the pre-processing step, where the ECG image dimensions are changed to \(224 \times 224\) pixels. Pre-processing also applies several augmentations, such as random color adjustments, random grayscale, random rotations, random horizontal flips, and random resized crops to the modeled input information, which improves the robustness of the model.
After pre-processing, the image was passed to the CNN block for feature extraction. In the feature extraction block, a series of convolutional layers, pooling, and ReLU activations are applied to extract relevant image features.
Multi-head attention also allows the model to further refine its feature extraction as shown in Fig. 2
Attention blocks.
by learning relevant features at different scales, allowing it to focus on the most important parts of the ECG signal. Subsequently, the figure depicts the SimCLR block for feature extraction based on contrastive learning (indicated by a green block). The SimCLR block comprises two encoders, each equipped with a projection head that generates ECG signal representations. The model maximizes the agreement among augmented views of the same ECG signal while minimizing the agreement between all other samples. The fine-tuning process facilitates the transfer of weights from the pre-trained encoder, thereby assisting the model in effectively acquiring feature representations. The final block was the ResNet block for classification. The ResNet block employs a series of residual blocks to learn deep spatial features, which are subsequently processed through a fully connected layer that outputs the classification labels. The efficacy of the architecture is derived from its integration of CNN-based feature extraction, SSL for representation learning, and robust classification using ResNet.
Figure 3 illustrates the flowchart of the SimCardioNet model, delineating the process from pre-processing to both the self-supervised and supervised training stages. Initially, the model processes an input ECG image (\(224 \times 224 \times 3\)) and applies multi-scale SimCLR transformations to generate augmented views. These transformed images are subsequently passed through a series of convolutional blocks, each employing convolution layers, batch normalization (BN), rectified linear unit (ReLU), dropout, and residual connections. Following feature extraction, an attention mechanism refines the feature maps, and the outputs from global average pooling and max pooling are concatenated. These concatenated features are transformed by feature fusion, and the backbone features are directed to either the projection head for SSL pre-training or the classifier head for the supervised classification task. The projection head applies L2 normalization, followed by contrastive loss calculation during SSL pre-training, whereas the classifier head uses cross-entropy loss for supervised classification.
Architecture flowchart of the SimCardioNet model, outlining pre-processing, feature extraction, SSL pre-training, and supervised classification stages.
The final model underwent separate training loops: one for SSL pre-training and another for supervised training, with cross-validation employed for model evaluation.
Data preparation
The data preprocessing pipeline addresses ECG data diversity through multi-stage augmentation. For self-supervised learning, two transformation pipelines generate diverse views: one uses aggressive augmentations, including strong color jittering, cropping with a minimum scale of 0.1, and Gaussian blur, while the other employs moderate augmentations to accommodate acquisition variations. In supervised training, preprocessing includes rotation (\(20^{\circ }\)), color jittering, affine transformations, perspective distortion, and random erasing. These techniques simulate variations in electrode placement, acquisition systems, positioning, and viewing angles. All images are standardized to 224 × 224 using ImageNet normalization, ensuring robust feature learning across four ECG classes (Normal, MI, Abnormal Heartbeat, History of MI) and patient populations.
Model development
The model uses a Custom ECG CNN Backbone (denoted \(f_\textrm{ECG}(x)\)) to extract feature maps from ECG images:
where:
-
\(x \in \mathbb {R}^{C \times H \times W}\) is an ECG image with C channels and \(H \times W\) resolution,
-
\(\textrm{CNN}(x)\) applies to extract hierarchical features from x.
The backbone is followed by a Projection Head \(h(\cdot )\) for contrastive learning.
where:
-
z is the projected representation in a lower-dimensional embedding space.
Self-supervised learning (SSL) pre-training
The model underwent self-supervised pre-training using the SimCLR framework. The key objective is to learn useful feature representations from the unlabeled ECG data. This is achieved through contrastive learning, in which the model is trained to bring augmented views of the same ECG sample more closely aligned in the feature space while distancing those from different samples.
Contrastive loss (InfoNCE): Given a pair of augmented views \(x_0\) and \(x_1\), the contrastive loss (InfoNCE) is used to enhance the similarity of positive pairs while decreasing the distance for negative pairs. The loss is formulated as follows:
where:
-
\(\text {sim}(z_0, z_1) = \frac{z_0 \cdot z_1}{\Vert z_0\Vert \Vert z_1\Vert }\) is the cosine similarity.
-
\(\tau\) is a temperature scaling factor controlling the similarity’s sharpness.
-
N is the number of negative samples.
Hybrid loss function:
A hybrid loss combining InfoNCE with cosine similarity was introduced to stabilize the training:
where \(\lambda\) is a hyperparameter that balances the contrastive and cosine similarity components.
Optimizer:
The AdamW optimizer was used to optimize the model, with the learning rates for the backbone and projection head set separately. A Cosine Annealing Scheduler is employed to adapt the learning rate during model training:
where \(\eta _t\) is the learning rate at epoch t, \(\eta _\text {min}\) is the minimum learning rate, \(\eta _\text {max}\) is the maximum learning rate, and T is the total number of epochs.
Supervised fine-tuning
After the SSL pre-training phase, the model was fine-tuned for the classification task. The goal of fine-tuning is to classify ECG signals into a set number of classes. The output of the model is a set of class scores \(\hat{y}_i\) and we optimize the classifier using Cross-Entropy Loss. We evaluated the performance of the model using 3-fold cross-validation. In each fold, the dataset was split into training and validation sets, ensuring that every sample was used for both training and validation. The steps for each fold were as follows:
-
Split the dataset into training and validation subsets.
-
Apply augmentations to the training data and the same transformations to the validation data.
-
Fine-tune the model using a cross-entropy loss function, defined as:
Supervised loss (cross-entropy loss): Fine-tuning is performed to optimize the loss function for classification, specifically utilizing Cross-Entropy.
where:
-
N is the number of samples in the batch.
-
C is the number of classes.
-
\(y_{i}^{c}\) is the binary indicator (0 or 1) if class label c is the correct classification for sample i.
-
\(p_{i}^{c}\) is the predicted probability for class c for sample i.
Progressive unfreezing:
To allow the model to retain the features learned during SSL pre-training, we applied a progressive unfreezing strategy. Initially, only the final layers were fine-tuned, and progressively, the earlier layers of the backbone were unfrozen and trained. Specifically:
-
In epoch 333, we unfreeze the final convolutional block.
-
In epoch 666, we unfreeze the middle layers.
-
In epoch 101010, we unfreeze the entire backbone, allowing the model to fine-tune all parameters.
Fine-tuning procedure: During fine-tuning, the model uses the AdamW optimizer with a learning rate \(\alpha = 2\times 10^{-3}\) and weight decay \(\lambda = 10^{-4}\). The model was trained for hundred epochs, and early stopping was adopted to prevent overfitting if the validation accuracy stagnated.
Experiments and results
The dataset utilized for this research consisted of ECG images obtained from publicly available datasets with varying heart conditions. The dataset was additionally preprocessed and augmented with various transformations to enhance model generalization and robustness, specifically for the task of ECG classification.
Dataset I Description: The dataset44 consists of 12-lead ECG images obtained from different healthcare institutions in Pakistan. The ECG recordings were acquired at a sampling rate of 500 Hz, with each record corresponding to an individual patient’s data. This diverse patient cohort enhances the generalization capabilities of the DL models. The dataset was classified into four distinct categories: one representing normal ECGs and the remaining three corresponding to various cardiac conditions. This dataset is publicly accessible and can be retrieved via the following link: https://data.mendeley.com/datasets/gwbz3fsgp8/2.
Dataset II Description: We utilized the “ECG Images Dataset of Cardiac Patients” from Kaggle for out-of-distribution evaluation. This dataset contains ECG images for cardiovascular research and was used only for external validation, without training or hyperparameter tuning. Using data from an alternative source introduces variations in acquisition conditions and noise patterns, testing the model’s robustness to domain shift. We followed the dataset’s license terms and retained original labels. Images were converted to grayscale, resized to match model input, and normalized, without relabeling or augmentation. Quality screening removed unreadable files. Per-class and per-split counts are reported below from the downloaded files. This dataset is publicly accessible and can be retrieved via the following link: https://www.kaggle.com/datasets/evilspirit05/ecg-analysis.
Dataset III Description: Dataset III: The PTB-XL41 dataset is a large, publicly available clinical electrocardiography dataset released on PhysioNet, consisting of 21,837 standardized 12-lead ECG recordings, each with a duration of 10 s. The signals are provided in WFDB format at two sampling rates, 500 Hz (high resolution) and 100 Hz (downsampled), facilitating both detailed signal analysis and efficient machine-learning model training. Each ECG record is accompanied by rich metadata, including patient demographics (such as age and sex) and acquisition information, with all personal identifiers pseudonymized. Importantly, the dataset includes comprehensive diagnostic annotations based on the SCP-ECG standard, covering diagnostic, rhythm, and morphological statements, resulting in a multilabel classification setting with 71 possible ECG statements that can be grouped into clinically meaningful super classes (e.g., normal ECG, myocardial infarction, conduction disturbances, ST/T-wave changes, and hypertrophy). PTB-XL also provides recommended patient-wise stratified train–test splits, ensuring reproducibility and preventing data leakage, which makes it a benchmark dataset for ECG analysis and cardiovascular machine-learning research. This dataset is publicly accessible and can be retrieved via the following link: https://physionet.org/content/ptb-xl/1.0.3/.
Hardware and training setup details:
All experiments were performed on a workstation equipped with an Intel Core i7 3rd generation processor, 16 GB of random access memory, and an NVIDIA GTX 1660 Super GPU with 6 GB of video random access memory. This hardware was chosen to ensure computational efficiency and accessibility by including a general standard hardware configuration that represents a typical mid-range device.
Computation time tracking: Computation time is now comprehensively tracked throughout the code using time.time() measurements. The implementation monitors SSL pretraining with per-epoch timing and total training duration, classification training with fold-level and epoch-level timing across cross-validation, and inference evaluation time for each fold. All timing metrics are displayed in real-time during training, for example, the format:
“SSL Epoch 1/50, Loss: 0.4521, Time: 45.32s”
Additionally, these timing metrics are saved to JSON files, which include:
-
fold_training_time_seconds
-
average_epoch_time_seconds
-
total_cv_time_seconds
-
evaluation_time_seconds
At the end of training, a comprehensive timing summary is displayed, which includes:
-
SSL pretraining time
-
Cross-validation time
-
Total training time
-
Average per-fold time
This provides a complete computational efficiency analysis for deployment considerations.
Training time:
The model was trained for hundred epochs and the data were subjected to 3-fold cross-validation. On average, it took approximately 126 s to train each epoch, which provided an estimate of the total training time as a function of the epochs and folds. The timing represents a real-world experience of the expected computational resources that can be consumed, assuming similar hardware.
Model configuration and training details
The ResNet parameters are configured through multiple layers in the implementation. The backbone uses ResNet50, initialized with weights=’IMAGENET1K_V2’ (pretrained on ImageNet), with the final classification layer replaced by torch.nn.Identity(). Additional CNN layers follow with channel dimensions of \(2048 \rightarrow 1024 \rightarrow 512\), using \(1 \times 1\) convolutions, BatchNorm, and ReLU activations.
SSL phase configuration: For the Self-Supervised Learning (SSL) phase, the AdamW optimizer is employed with differential learning rates. The parameters of the backbone are configured with a learning rate \(\text {lr} = 3 \times 10^{-4}\) and weight decay \(\text {weight}\_\text {decay} = 1 \times 10^{-4}\), while the projection head uses a learning rate \(\text {lr} = 1 \times 10^{-3}\) and weight decay \(\text {weight}\_\text {decay} = 1 \times 10^{-5}\). The CosineAnnealingLR scheduler is applied with \(T_{\text {max}} = 50\) and \(\eta _{\text {min}} = 1 \times 10^{-6}\), running for 50 SSL epochs.
Classification phase configuration: For the classification phase, the AdamW optimizer is again used with a learning rate \(\text {lr} = 2 \times 10^{-3}\) and weight decay \(\text {weight}\_\text {decay} = 1 \times 10^{-4}\). The progressive unfreezing strategy is applied to the convolutional blocks: conv_block3 is unfrozen at epoch 3, conv_block2 at epoch 6, and all layers are unfrozen at epoch 10.
Batch size and dropout configuration: The batch size is set to 16 for SSL and 8 for classification. The dropout layers have varying rates across the layers: \(0.05 \rightarrow 0.1 \rightarrow 0.2\).
Loss functions: The loss functions used during training include InfoNCE loss with temperature \(\tau = 0.1\), Cosine loss, Triplet loss with margin \(\gamma = 0.1\), and Hybrid loss, which combines InfoNCE and cosine components with a weight for cosine \(\text {cosine}\_\text {weight} = 0.1\). These loss functions are carefully designed to ensure the model learns meaningful representations for both self-supervised pretraining and classification tasks.
Model assessment and performance analysis
This section considers the performance of the SimCardioNet ECG Classification Model using qualitative and quantitative evaluation metrics. This section provides an overview of the evaluation techniques used to assess the performance of SimCardioNet in both SSL pre-training and supervised fine-tuning. In particular, the model was assessed using 3-fold cross-validation, which increased the overview of established generalizability to further construct data attributes of accuracy and reliability. In addition to focusing on the quantitative aspects of performance evaluations, we also undertook a detailed comparison of the performance of SimCardioNet relative to a baseline to promote comparisons that regarded the proposed feature of SSL applied to supervised fine-tuning as being effective if it improved parts of the ECG classification performance. The quantitative results were visualized using training curves, loss plots, and confusion matrices to illustrate the varying areas of potential strengths and weaknesses of the model.
SSL performance
The performance of the SSL method will be assessed using various visualizations and statistical techniques. SSL has demonstrated to be a very powerful approach for our learning of discriminative features from ECG images, when no labels were provided. Employing contrastive learning procedures allows the model to learn rich representations of ECGs, recognizing performance-rich patterns and abnormal characteristics needed for performance rich classification problems.
Backbone feature statistics: Figure 4 represents the Mean Features per Class on the backbone features of ECG images of Dataset I. Each cell on the heatmap represents the mean of one of the feature dimensions for different ECG classes: myocardial infarction patients, patients with a history of MI, patients with irregular heartbeats, and normal heartbeats. The brighter the color in the heatmap, the greater the mean feature dimension values. The heatmap shows how disparate or homogeneous the learned features are across each class, while also illustrating how well the model has potentially segregated ECG patterns based on feature extraction.
Mean features per class, indicating the values for each feature dimension in the backbone using Dataset I.
Projection Feature Statistics: The Mean Features per Class for the projection head features is displayed in the heatmap, where each column represents a feature dimension (1–128), and rows represent ECG classes: MI patients, patients with a history of MI, patients exhibiting abnormal heartbeats, and normal individuals. The brightness of the color signifies the mean feature values; a brighter hue represents a higher mean value. As such, Fig. 5 visualizes these feature statistics and provides a perspective on how the projection head was able to discriminate between the different ECG classes and how well it was able to distinguish discriminative features in the reduced feature space.
Mean features per class for the projection head.
This Fig. 6 illustrates the class-wise statistical characteristics of extracted features from the Dataset III. The upper heatmap presents the mean feature values across feature dimensions for four ECG diagnostic classes (NORM, MI, STTC, and CD), highlighting subtle but consistent differences in feature distributions that reflect class-specific ECG patterns. The lower heatmap shows the corresponding standard deviations, indicating the degree of intra-class variability for each feature dimension. Warmer colors in the standard deviation map reveal features with higher dispersion, suggesting greater morphological or temporal variability within those classes, while cooler regions indicate more stable features. Together, these visualizations demonstrate that although mean feature values are relatively close across classes, distinct variability patterns exist, supporting the discriminative potential of the extracted feature set for ECG classification.
Mean features per class for the projection head. using Dataset III.
PCA and t-SNE visualization for backbone features: The PCA visual shows the distribution of the projection head features mapped along its first two principal components, capturing the maximum variance of the data. The measure of separation of the different ECG classes along these two principal components demonstrates how well the model is classifying and distinguishing based on the learned features. The clearer the separation, the better the success of the classification. In this case, the model has learned the differences in classification between normal and abnormal ECG signals, as well as specific anomalies including myocardial infarction, arrhythmias etc. Figure 7 is an example of this visual.
PCA visualization of the 128-dimensional projection head features, showing the separation of ECG classes along the first two principal components.
This Fig. 8 presents a two-dimensional PCA projection of SSL features extracted from the Dataset III, where each point represents an ECG recording and colors indicate diagnostic classes (NORM, MI, STTC, and CD). The visualization shows substantial overlap among classes, reflecting the intrinsic complexity and inter-class similarity of ECG morphologies, particularly between pathological categories. Nevertheless, regions of partial clustering and varying point densities suggest that the SSL features capture meaningful latent structure related to cardiac conditions, even without explicit supervision. The spread along the first two principal components indicates that these components explain a significant portion of the feature variance, while the observed overlaps highlight the need for higher-dimensional decision boundaries or supervised fine-tuning for improved class separability.
PCA visualization of self-supervised ECG feature representations from the Dataset III, illustrating class-wise distributions and overlap among normal and pathological conditions.
The t-SNE visualization provides a more robust analysis of projection head features by visualizing clustering in two-dimensions. Where PCA is concerned with variance, t-SNE is concerned with local relationships and allows us to better see how together pretty much identical samples from the same class cluster together. Tight and distinct clusters suggest the model has learned strong discriminative features. A large overlap between clusters would suggest that the model had difficulties in distinguishing between classes. See Fig. 9 for the t-SNE plot.
t-SNE visualization of the 128-dimensional projection head features, illustrating the clustering and separation of ECG classes in a 2D space.
This Fig. 10 shows a t-SNE visualization of SSL feature representations extracted from the Dataset III, with points colored according to diagnostic classes (NORM, MI, STTC, and CD). Unlike PCA, t-SNE emphasizes local neighborhood structure, revealing more distinct clusters and subclusters that correspond to different ECG patterns. Normal recordings form relatively compact and well-defined groups, while pathological classes exhibit greater dispersion and partial overlap, reflecting heterogeneity within disease categories and shared morphological characteristics across conditions. The presence of multiple localized clusters suggests that the SSL model captures meaningful nonlinear relationships in the ECG signals, providing a strong basis for downstream supervised classification despite remaining inter-class overlap.
t-SNE projection of self-supervised ECG feature embeddings from the Dataset III, illustrating local clustering behavior and class-wise distribution of normal and pathological recordings.
Conv1 activation maps: The Conv1 activation maps (Figs. 11, 12) represent the low-level features of the ECG signal, highlighting the essential features of the ECG, primarily rhythm, edges (notable discontinuities), and periodicity (recurring patterns). These are the fundamental features on which all other complex characteristics of ECG signal can be learned. The activation maps describe the regions of the ECG waveform that are spatially relevant for the low-level features we are observing, particularly in the horizontal regions. Each channel represents a feature map with a different degree of activation for various portions of the signal.
Conv1 activation maps for sample (Class: 0), showing the extraction of low-level features such as rhythm and shape from the ECG signal, with activation focused on specific regions of the waveform of Dataset I.
Conv1 activation maps for sample (Class: 0), showing the extraction of low-level features such as rhythm and shape from the ECG signal, with activation focused on specific regions of the waveform of Dataset II.
The Conv2 activation maps (Figs. 13, 14) capture more intermediate features of the ECG signal and focus on patterns that are recognizable such as the P-QRS-T waveforms. These are functions at a higher level of abstraction than Conv1 and allow the model to detect critical components of the ECG waveform. The Conv2 activation maps depict concentrated areas that emphasized the regions of the ECG signal that were most relevant for distinguishing these important features, and therefore were more focused on the overall shape of the waveform.
Conv2 activation maps for sample (Class: 0), highlighting intermediate-level features such as the P-QRS-T wave patterns and focusing on the shape of the ECG waveform Dataset I.
Conv2 activation maps for sample (Class: 0), highlighting intermediate-level features such as the P-QRS-T wave patterns and focusing on the shape of the ECG waveform Dataset II.
The Conv3 activation maps (Figs. 15, 16) represent higher-order features of the ECG signal as well as advanced-level features of disease-specific abnormality, which includes specific arrhythmias and myocardial infarctions. At this level of advancement, the model learns to locate and show complex abnormality specific features. The activation maps display more complex patterns than earlier layers of convolutions. They also concentrate in areas of the ECG signal that demonstrate areas in which the ECG display features from differentiating these higher-order features.
Conv3 activation maps for sample (Class: 0), highlighting deep features and advanced patterns in the ECG signal, such as arrhythmias and myocardial infarctions Dataset I.
Conv3 activation maps for sample (Class: 0), highlighting deep features and advanced patterns in the ECG signal, such as arrhythmias and myocardial infarctions of Dataset II.
This Fig. 17 illustrates the learned activation patterns of the first convolutional layer (conv1) for a representative normal (NORM) ECG recording from the Dataset III. The left panel shows the original multi-lead ECG signal, while the remaining panels depict activation maps from individual convolutional channels. Each activation map highlights regions of the input signal that strongly stimulate a specific filter, indicating that different channels respond to distinct temporal and morphological ECG characteristics. Some channels emphasize localized high-energy segments corresponding to prominent waveform components, whereas others exhibit more distributed activations, reflecting sensitivity to broader signal patterns across leads and time. Together, these activation maps demonstrate how early convolutional layers decompose raw ECG signals into diverse low-level representations that form the foundation for higher-level feature learning.
Visualization of conv1 activation maps for a normal ECG recording, showing channel-wise responses of the first convolutional layer to different temporal and morphological signal patterns.
To assess the model’s learning progress, the SimCLR SSL Training Loss was monitored over 100 epochs during the pre-training phase. The training loss, shown in Figs. 18, 19 and 20 depicts the contrastive loss during training. Initially, the contrastive loss was relatively high, indicating that the model embeddings were not yet effective at distinguishing between similar and dissimilar ECG signals. As training progressed, sharp drops in the loss were observed, signifying that the model was learning to better distinguish between positive and negative pairs. The loss stabilizes at certain points, suggesting that the model has reached a certain level of optimal-feature representation.
SimCLR SSL training loss on Dataset I.
SimCLR SSL training loss on Dataset II.
SimCLR SSL training loss on Dataset III.
These periodic drops in loss are consistent with the ability of the SimCLR framework to learn discriminative embeddings through contrastive learning. As the loss approaches zero in the later epochs, the model representations become more refined. This improvement in the training loss correlates with the strong performance metrics observed across the per-class results, where the model achieved high precision, recall, and F1-scores in classifying various ECG signals. The reduction in contrastive loss directly contributes to the model’s increased ability to generalize and ensure prediction accuracy, as shown in the confusion matrix and accuracy metrics.
Multi-class classification
The assessment of the model on the multi-class classification task was conducted on four distinct classes of Myocardial Infarction (ECG Images), History of MI (ECG Images), Abnormal Heartbeat (ECG Images), and Normal Person (ECG Images). The model’s effectiveness was evaluated using the following main evaluation metrics. These performance measures provide a broad view of how the model performs on distinct types of ECG signals. A summary of the model performance for each class is presented in Table 1.
As detailed in Table 1, ECG MI Patients achieve precision \(0.9795 \pm 0.0058\), meaning that nearly 98% of predictions labeled as MI are correct, and recall \(1.0000 \pm 0.0000\), indicating no MI cases were missed (no false negatives). The resulting F1-score \(0.9896 \pm 0.0029\) reflects a very strong and well-balanced performance for this class.
For History of MI, precision is \(0.9598 \pm 0.0147\) and recall is \(0.9485 \pm 0.0090\). While both are high, the slightly lower recall compared to MI Patients suggests a small number of History-of-MI cases were missed; the F1-score \(0.9541 \pm 0.0115\) confirms effective performance with most errors likely being false negatives.
In Abnormal Heartbeat, precision reaches \(0.9868 \pm 0.0105\), indicating very reliable positive predictions, but recall is comparatively lower at \(0.9404 \pm 0.0234\), implying some abnormal beats were overlooked (misclassified as other classes). The F1-score \(0.9630 \pm 0.0167\) shows strong yet slightly less balanced performance driven by those missed positives–consistent with the clinical difficulty of distinguishing certain abnormal patterns from near-normal rhythms.
For Normal ECG, precision is \(0.9724 \pm 0.0138\) with perfect recall \(1.0000 \pm 0.0000\). The model correctly identifies all normal cases while keeping false positives low, yielding an F1-score \(0.9860 \pm 0.0071\).
On Dataset II (external validation), all four classes achieve \(1.000 \pm 0.000\) for precision, recall, and F1-score, indicating perfect separability on the evaluated split(s). While highly encouraging, such ceiling performance should be interpreted cautiously, as it may reflect an easier distribution or smaller sample sizes relative to Dataset I.
This Table 2 reports the class-wise classification performance of the proposed model on the Dataset III using precision, recall, and F1-score as evaluation metrics. The model achieves the strongest performance on the NORM class, with high precision (0.945) and recall (0.93), indicating reliable discrimination of normal ECG signals. For pathological classes, CD and MI demonstrate robust and balanced performance, suggesting effective detection of conduction disturbances and myocardial infarction patterns. The STTC class also shows stable results, reflecting consistent sensitivity to ST/T-wave abnormalities. In contrast, the HYP class exhibits noticeably lower performance across all metrics, likely due to higher intra-class variability and class imbalance commonly observed in hypertrophy-related ECG patterns. Overall, the results indicate strong classification capability for most classes while highlighting hypertrophy as a challenging category requiring further feature refinement or data augmentation.
Confusion matrix and cross-validation insights
The Mean Confusion Matrix in Figs. 21 and 22 presents a more detailed illustration of the model’s classification results of the four ECG signal classes: Myocardial Infarction (ECG Images), History of MI (ECG Images), Abnormal Heartbeat (ECG Images), and Normal Person (ECG Images). Every row in the matrix indicates the true class labels, and every column indicates the model’s predicted class labels. The diagonal elements indicate the number of true classified samples, or true positives (TP), and the off-diagonal elements show incorrectly predicted samples across classes.
Mean confusion matrix for cross-validation (CV) in multi-class ECG classification of Dataset I.
Mean confusion matrix for cross-validation (CV) in multi-class ECG classification of Dataset II.
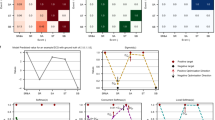
The mean confusion matrix from cross-validation in Fig. 23 using Dataset III, showing how well a classifier distinguishes five classes: NORM, MI, STTC, CD, and HYP. The diagonal cells (e.g., NORM\(\rightarrow\)NORM = 1693±12) show correct predictions, while off-diagonal cells show misclassifications (e.g., 42±5 NORM cases wrongly predicted as MI). The ± values indicate variability across CV folds. Darker blue = higher counts. Overall, the model performs best on NORM and worst on HYP, with some confusion between similar classes like NORM/MI or STTC/CD.
Mean confusion matrix for cross-validation in multi-class ECG classification of Dataset III.
For Myocardial Infarction (ECG Images), the model achieved a very high number of true positives at 79.7, which means that it classified very well for this class. There were no misclassifications into other classes, which means that the model could accurately distinguish Myocardial Infarction cases.
The Normal Person (ECG Images) class was indicated to have 94.7 as a true positive with no misclassifications, meaning that for this class the model has perfect recall. The History of MI (ECG Images) class did have some misclassifications: for the History of MI class there were 2.3 objects that were misclassified as Abnormal Heartbeat (ECG Images), and 0.7 objects that were misclassified as Normal Person (ECG Images). This indicates that the model has difficulty classifying the history of MI, and struggles to differentiate between history of MI / abnormal heartbeat with normal persons. Additionally, the Abnormal Heartbeat (ECG Images) class had misclassifications of its own, which were (2.7) objects misclassified as History of MI, and another (2.7) as normal. Overall, this confirms that the model has trouble distinguishing between Abnormal Heartbeat and history of MI, which is understandable because of ECG patterns that are very similar between both classes.
The confusion matrix also includes the standard deviation of the predictions for each class, indicating variability across the cross-validation folds. For example, Myocardial Infarction has a standard deviation of \(\pm 8.2\), and a history of MI shows a standard deviation of \(\pm 5.0\), suggesting some variability in model performance across multiple subsets of the data. As illustrated by the cross-validated mean confusion matrix (Fig. 22), predictions are essentially perfectly diagonal, with all off-diagonal entries equal to 0.0 ± 0.0 0.0±0.0, indicating no systematic class confusions across folds. The model consistently identifies each category with high stability: ECG MI Patients 318.7 ± 9.4 318.7±9.4 correct per fold, History of MI 172.0 ± 2.9 172.0±2.9, Abnormal Heartbeat 233.0 ± 11.3 233.0±11.3, and Normal ECG 284.0 ± 7.8 284.0±7.8. The small standard deviations reflect limited between-fold variability, and the larger ± ± for Abnormal Heartbeat appears driven by fold composition rather than misclassification, since counts remain strictly on the diagonal. Row annotations (e.g., 240 \(\times\) 12 = 2880 240\(\times\)12=2880) denote per-fold counts \(\times\) \(\times\) number of folds, providing sample-size context. Overall, the matrix corroborates near-perfect separability of the four ECG classes under cross-validation, with robust and consistent performance across splits.
The cross-validation results averaged over 3-folds, presented in Table 3, provide a comprehensive view of the model performance across the four classes of ECG signals. The model attained an accuracy of 0.9731 (\(\pm 0.0093\)), indicating that, on average, it correctly classified \(97.31\%\) of the instances across all classes during the cross-validation process. This high accuracy reveals the model’s overall competence in classifying ECG signals correctly, which aligns with the exceptional precision and recall performance observed in the per-class analysis, particularly for the Myocardial Infarction and Normal Person classes, where the model performed exceptionally well.
Additionally, As summarized in Table 3, the model shows strong discrimination on Dataset I, with accuracy 0.9752 ± 0.0085 0.9752±0.0085 and macro F1 0.9732 ± 0.0085 0.9732±0.0085. Macro-averaged precision/recall/F1 closely match their weighted counterparts, indicating consistent performance across classes rather than dominance by a majority class. Ranking ability is likewise high (AUC-ROC macro 0.9841 ± 0.0026 0.9841±0.0026; weighted 0.9855 ± 0.0036 0.9855±0.0036; AUPRC macro 0.9698 ± 0.0065 0.9698±0.0065; weighted 0.9713 ± 0.0063 0.9713±0.0063). Calibration on Dataset I is reasonable, with ECE 0.0340 ± 0.0121 0.0340±0.0121 and Brier 0.0514 ± 0.0161 0.0514±0.0161, suggesting probability estimates that are broadly aligned with empirical frequencies. On Dataset II (external validation), discrimination metrics are reported as 1.000 ± 0.000 1.000±0.000 for accuracy, macro/weighted precision, recall, F1, AUC-ROC, and AUPRC, implying perfect separability on the evaluated split(s). Despite this, calibration is mixed: the Brier score improves markedly ( 0.0053 ± 0.0004 0.0053±0.0004), while ECE increases to 0.0593 ± 0.0031 0.0593±0.0031, indicating slightly less well-calibrated probabilities (mild over/under-confidence) even though the hard decisions are flawless. Taken together, the near-identical macro and weighted scores on Dataset I argue for balanced per-class performance, and the perfect external results are promising but should be interpreted with caution, as they can arise from an easier distribution, smaller sample sizes, or other dataset-specific factors. The perfect alignment between the true and predicted labels for all the displayed samples confirms the model’s high capability in detecting ECG signal patterns across different classes. This visual confirmation, as shown in Fig. 24, supports the earlier results, highlighting that the model successfully learns the distinct features of each class and can reliably predict heart conditions based on ECG data.
ECG waveform samples with true and predicted labels.
To evaluate the model’s training progress for classification, we examined the training accuracy and loss curves for the training and validation sets for 100 epochs. The curves shown in Figs. 25 and 26 provide insight into how well the model learns and generalizes.
Classification training curves for loss and accuracy Dataset I.
Classification training curves for loss and accuracy Dataset II.
In the training loss curve (left plot), the training loss (in blue) decreases steadily, demonstrating that the model efficiently minimizes the loss function and refines its predictions on the training data. However, the validation loss (in orange) exhibits some fluctuations, initially decreasing and then increasing after approximately the 60th epoch. This behavior suggests that the model starts to overfit the training data as it becomes less capable of generalizing to the validation set.
The accuracy curve (right plot) exhibits a similar trend. The training accuracy (in blue) increases consistently, reflecting the model’s improvement in the training set. The validation accuracy ( orange) follows a similar pattern but with more significant fluctuations, particularly after the 60th epoch. This inconsistency in the validation accuracy further supports the presence of overfitting, where the model performance on the validation set no longer improves and starts to degrade as it becomes more specialized to the training data.
These training curves, as shown in Figs. 25 and 26, highlight the model’s ability to learn over time, but they also point to potential overfitting after a certain number of epochs. This issue can be addressed using techniques such as early stopping, regularization, or more diverse data augmentation to improve the model’s generalization to unseen data.
The classification model trained on Dataset III demonstrates robust convergence and generalization, as evidenced by Fig. 27. Both training and validation loss decrease steadily over 100 epochs, with minimal divergence, indicating low overfitting. Correspondingly, training and validation accuracy rise in tandem to approximately 92% and 90%, respectively, reflecting strong discriminative performance. The stability of validation metrics confirms the model’s capacity to generalize effectively to unseen instances within Dataset III.
Classification training curves for loss and accuracy Dataset III.
Explainability: grad-CAM heatmaps
To enhance interpretability, we employ Grad-CAM heatmaps over convolutional feature maps to localize regions influential for predictions. Grad-CAM computes class-specific gradients and projects them onto feature activations, highlighting discriminative signal patterns like ST-segment deviations or rhythm irregularities. We apply Grad-CAM to correctly and incorrectly classified samples and overlay heatmaps on original inputs. These visual explanations confirm the model’s focus on clinically plausible regions and reveal failure modes where attention shifts to artifacts. We report representative examples and provide qualitative consistency across cross-validation folds. As shown in Fig. 28, the model focuses on waveform regions rather than page artifacts.
Grad-CAM heatmaps highlighting regions driving the model’s predictions. Warm colors indicate higher contribution; overlays show attention focused on ECG waveforms rather than page artifacts od Dataset I.
Grad-CAM heatmaps highlighting regions driving the model’s predictions. Warm colors indicate higher contribution; overlays show attention focused on ECG waveforms rather than page artifacts od Dataset II.
Also, Fig. 29 shows the raw ECG, the Grad-CAM map from the attention_conv layer, and the overlay. Unlike block-level maps that focus on waveform regions, these saliency maps assign the highest importance to the header and border regions (red/yellow), with low emphasis on the ECG trace itself (blue). This pattern, consistently observed across examples, suggests a potential shortcut: the model attends to non-physiologic artifacts rather than the underlying ECG morphology. While such behavior may yield high accuracy if the plot layout correlates with labels, it reduces model robustness. Possible mitigation include cropping headers and margins, standardizing plot layouts, applying border and brightness augmentations, and retraining while monitoring saliency maps to ensure proper attention to QRS complexes, ST segments, and T-waves. Quantitative checks can then verify that predictions no longer depend on layout artifacts.
Grad-CAM heatmaps highlighting regions driving the model’s predictions. Warm colors indicate higher contribution; overlays show attention focused on ECG waveforms rather than page artifacts od Dataset III.
Figure 30 presents Grad-CAM visualizations derived from the first conv1 of the trained classification model, applied to four representative ECG leads (II, V1, V2, V5) from Dataset III. Each row juxtaposes the original ECG signal (left), the corresponding Grad-CAM intensity heatmap (center), and an overlay highlighting regions of high activation (right). The Grad-CAM heatmaps color-coded by intensity (0.2 to 1.0) reveal spatiotemporal patterns within the input signal that the model identifies as discriminative for classification. Notably, regions of elevated activation consistently align with morphologically salient features such as QRS complexes and ST-segment deviations across all leads, suggesting that the model leverages clinically relevant waveform characteristics for decision-making. The consistent spatial correspondence between high-intensity regions and diagnostic waveforms across multiple leads supports the interpretability and physiological plausibility of the model’s learned representations. This visualization provides critical insight into the model’s internal reasoning, reinforcing its potential utility in clinical decision support systems where transparency is paramount.
State-of-the-art comparison
Various models have been proposed for ECG classification, ranging from traditional ML to advanced DL techniques. This section compares several state-of-the-art models, including MobileNet V2, VGG16, AlexNet, YOLOv8, and LSTM with SSL, using fundamental metrics. Table 4 summarizes their performance, emphasizing the strengths and limitations of each method.
Table 5 presents a comparative evaluation of recent state-of-the-art methods for multi-class ECG classification on Dataset III, alongside our proposed model, SimCardioNet. Our approach achieves a balanced performance across all metrics, yielding a precision of 0.922, recall of 0.921, F1-score of 0.921, and overall accuracy of 0.921. Notably, SimCardioNet outperforms prior architectures including a dual-branch CNN42, a CNN enhanced with entropy features52, an LSTM53, and a Bi-GRU network43 in both F1-score and accuracy, while maintaining near-perfect harmony between precision and recall. This uniformity indicates minimal bias toward any specific class and reflects robust generalization across the five diagnostic categories (NORM, MI, STTC, CD, HYP). The consistent superiority of SimCardioNet underscores the efficacy of its architectural design in capturing discriminative morphological and temporal patterns in 12-lead ECG signals.
Ablation study
In this section, we conduct an ablation analysis to assess the impact of various architectures and configurations on ECG classification performance. In particular, we investigate the combination of CNN with ResNet, the combination of SimCLR and ResNet, and a standalone ResNet framework. We will investigate the performance of these models, in order to understand the contributions of the various approach, which could lead to the appropriate architecture design in order to yield accurate ECG classifications.
In Table 6, the metrics from a CNN + ResNet model are used for the classification of ECG data into four categories: MI Patients, History of MI, Abnormal Heartbeat, and Normal. When evaluating MI Patients, the model showed excellent metrics, with a precision of 0.95 and a recall of 0.94, resulting in a balanced F1 score of 0.94. This suggests that our model has a strong ability to correctly identify patients with myocardial infarction. The model performs exactly like this for the History of MI, with precision and recall both equal to 0.93, resulting in an F1 score of 0.93, indicating that it has a strong performance in identifying patients who had a heart attack in the past.
For the Abnormal Heartbeat class, the model shows a precision of 0.93, along with a recall value of 0.95, which produces an F1-score of 0.94. These results indicate that the model is quite accurate in detecting abnormal beats. The model has a performance extremely close to that for the normal class, where the precision and recall values are 0.94 and 0.95, respectively, providing an overall F1-score for the Normal class of 0.94. That the F1-score is consistently high (0.94) suggests that the model is reasonably accurate overall and fairly consistent across the classes. This gives us confidence in concluding that our model can effectively and consistently detect both healthy and diseased heart conditions.
The results presented in Table 7 for the SimCLR + ResNet model provide a summary of the model’s performance across four categories of ECG data: MI Patients, History of MI, Abnormal Heartbeat, and Normal. For the MI category, the model achieved a precision and recall of 1.00, producing an F1-score of 1.00, demonstrating perfect identification of patients with myocardial infarction. For the History of MI category, the model attained a precision of 1.00, with a slightly lower recall of 0.90, yielding an F1-score of 0.95. This suggests that, although the model is highly effective in identifying patients with a history of myocardial infarction, some cases may be missed.
The classifying model for Abnormal Heartbeat showed a precision of 0.92 and recall of 0.94, resulting in an F1-score of 0.93, further suggesting good performance in terms of the true understanding of the rate of abnormal heart rhythms. The Normal class model showed a strong precision of 0.94, 1.00 recall, and an F1-score of 0.97. Hence, the overall accuracy was 0.96, which is noteworthy; it demonstrates that the model was reasonably successful in classifying normal instances across the classes. It can be concluded, given the overall performance as demonstrated by the addressed performance metrics, that the SimCLR + ResNET model was exceptional when applied to diagnosis, especially for indicating MI patients and normal instances, and in-class distributions maneuvering good rates of precision and recall evenly across the entire set of classes.
The performance metrics for the Simple ResNet model, as presented in Table 8, underscore its capability to classify various ECG conditions. For the MI class, the model attained a precision and recall of 0.92 and 0.86, respectively, resulting in an F1-score of 0.89. This suggests that the model is relatively effective in identifying myocardial infarction cases, although there is a slight reduction in recall, indicating that it misses some instances. For the History of MI class, the model’s precision was 0.84, with a recall of 0.75 and an F1-score of 0.79, suggesting that it encountered more difficulty in accurately identifying patients with a history of MI, with a higher incidence of false negatives.
For the abnormal class, the model demonstrated a precision of 0.81 and a recall of 0.93, culminating in an F1-score of 0.87. Although the model effectively identified a substantial number of abnormal heartbeats, it exhibited a higher incidence of false positives, as indicated by the comparatively lower precision. Regarding the Normal class, the model exhibited commendable performance, with a precision of 0.92, recall of 0.93, and an F1-score of 0.92, signifying robust efficacy in recognizing healthy ECGs. The model accuracy was 0.92, indicating a well-maintained classification balance across all classes. The Simple ResNet model performed proficiently overall, particularly in detecting Normal and Abnormal classes; however, it could benefit from enhancements in identifying MI and History of MI patients. These findings suggest that the Simple ResNet model is dependable for general ECG classification tasks, although it encounters challenges with specific cardiac conditions.
The ablation studies provide insight into how configurations in the SimCardioNet model affect the results, with the results illustrating model outcomes in a CNN+ResNet configuration, SimCLR+ResNet configuration, and Simple ResNet baseline/weak learner. The CNN + ResNet configuration performed strongly in terms of precision, recall, and F1-scores, which were relatively consistent across all classes, demonstrating an effective model for ECG classification. The SimCLR+ResNet configuration was the overall best-performing model in these evaluations based on precision, recall, and F1-scores as a result of the contrastive learning integrated framework, thus enhancing the precision and recall when determining abnormal heartbeats and myocardial infarction, while continuing to show strong precision, recall, and F1-scores for all classes. The Simple ResNet baseline model performed well; however, its lower precision and recall for certain detection classes, such as History of MI, imply that this configuration does not sufficiently classify more complicated cardiac abnormal detections.
To assess the contribution of the attention mechanism, the model was trained without integrating any attention block. As shown in Table 9, the overall accuracy dropped to 0.92, accompanied by consistent reductions in precision and recall across all ECG classes. The absence of attention hindered the model’s ability to emphasize diagnostically relevant waveform segments, resulting in less discriminative feature representation. Notably, the recall for History of MI and Normal ECG decreased to 0.90 and 0.94, respectively, underscoring that the attention block enhances the model’s sensitivity to subtle morphological differences in ECG signals.
The pre-trained model was evaluated using only the SSL framework, excluding both the attention mechanism and any supervised fine-tuning of the encoder. As summarized in Table 10, the overall accuracy declined further to 0.86, with moderate reductions across all performance metrics. This result indicates that SSL pretraining enhances general feature representation but cannot independently achieve strong class discrimination.
The model demonstrates adequate generalization yet reduced precision for classes with overlapping ECG morphologies. These findings suggest that SSL provides a meaningful initialization advantage; however, its full diagnostic potential is realized only when combined with supervised fine-tuning and attention-driven feature weighting, as explored in subsequent ablations.
To assess the impact of fine-tuning strategies, the learning rates of both the backbone and projection head were deliberately reduced to achieve more stable and gradual optimization during supervised training. Specifically, the backbone learning rate was decreased from 3e−4 to 1e−4 (a 3-fold reduction), while the projection head learning rate was lowered from 1e−3 to 3e−4 (approximately a 3.3 × reduction).
As shown in Table 11, the reduced learning rates improved the model’s convergence stability and produced an overall accuracy of 0.91. The more conservative step size enabled smoother fine-tuning of pre-trained features, preventing abrupt gradient updates that could otherwise disrupt learned representations. This configuration maintained a balanced precision–recall profile across all ECG classes, confirming that moderate learning-rate adjustments enhance task-specific adaptation while retaining the representational strength gained through self-supervised pretraining.
To assess the influence of domain-specific augmentations, the model was trained under two configurations: without any augmentation and with physiologically inspired signal augmentations, including random time scaling (0.9–1.1\(\times\)), additive Gaussian jitter (\(\sigma = 0.02\)), and synthetic baseline drift (up to 0.1 mV amplitude). As shown in Table 12, the inclusion of these augmentations improved the overall accuracy from 0.94 to 0.96, along with consistent gains in F1-scores across all ECG classes.
These improvements confirm that domain-specific augmentations enable the network to generalize better by simulating realistic physiological variations and acquisition noise typically present in ECG signals. The temporal scaling and jittering mimic natural heart rate fluctuations and sensor disturbances, while baseline drift augmentation enhances robustness to low-frequency noise. Consequently, the augmented model demonstrates stronger stability and discrimination between cardiac classes. To assess the contribution of different attention mechanisms, the existing Attention block was replaced with a temporal attention module that explicitly models time-dependent dependencies within the ECG signal. As presented in Table 13, the modified network maintained balanced performance across all classes, achieving an overall accuracy of 0.95. The temporal attention block effectively captured temporal correlations between successive waveform segments, enhancing the model’s ability to interpret the dynamic evolution of cardiac cycles.
However, the comparison indicates that MHSA provides slightly superior spatial–channel contextual integration for two-dimensional ECG representations. Temporal attention, while proficient at modeling temporal continuity, lacks the broader global dependency modeling inherent to MHSA.
The SimCardioNet model demonstrated superior performance compared with all ablation studies, achieving an accuracy of 97.31%, precision of 97.33%, recall of 97.31%, and F1-score of 97.28%. These results validate the efficacy of the proposed architecture and indicate that the integration of DL techniques, including CNNs, SimCLR, and ResNet, enhances the classification of ECG data. The findings from the ablation studies underscore the improvements facilitated by the SimCLR method and the layered structure of the ResNet architecture, further substantiating SimCardioNet as a reliable, reproducible, and robust solution for cardiac disease detection.
Discussion
CVDs remain a leading cause of morbidity and mortality worldwide; although ECG is ubiquitous and inexpensive, expert interpretation is time-consuming and not uniformly available. Our findings show that SimCardioNet a CNN backbone with multi-scale feature extraction, attention mechanisms, and SimCLR-based self-supervised pretraining can help close this gap by learning transferable ECG representations from unlabeled data and then specializing them via supervised fine-tuning. In 3-fold cross-validation on Dataset I, SimCardioNet achieved high discrimination (accuracy, precision, recall, and F1 all \(\approx 0.973\) ), with macro and weighted scores closely aligned, indicating balanced per-class performance rather than dominance by the majority class. Per-class analysis further showed near-perfect identification of ECG MI Patients and Normal ECG, with modest recall reductions in History of MI and Abnormal Heartbeat, consistent with the known clinical ambiguity of these phenotypes. Crucially, SimCardioNet generalizes beyond local clinical data. On Dataset II, an out-of-distribution external validation set, it achieved perfect discrimination (1.000±0.0 across all metrics), reinforcing its robustness to domain shifts though such ceiling performance warrants cautious interpretation due to potential dataset simplicity or layout-based shortcuts. More rigorously, on the large-scale, clinically diverse Dataset III, SimCardioNet attained 0.921 accuracy and 0.921 F1-score across five diagnostic super classes (NORM, MI, STTC, CD, HYP), outperforming recent state-of-the-art approaches including dual-branch CNNs, entropy-augmented CNNs, LSTMs, and Bi-GRU architectures. Performance was strongest for Normal and Conduction Disturbance classes, while Hypertrophy exhibited lower accuracy, likely due to intra-class heterogeneity and label sparsity, a known challenge in Dataset III. Two ingredients appear particularly beneficial. First, contrastive pretraining with SimCLR stabilizes and enriches feature representations under limited labels, improving downstream separability and ranking metrics (macro AUC-ROC/AUPRC). Second, progressive unfreezing during supervised fine-tuning preserves useful pretraining features while allowing task-specific adaptation, mitigating overfitting, and supporting generalization. These design choices are reflected in the confusion matrices, which are effectively diagonal under cross-validation on Dataset I and show meaningful class separation on Dataset III despite inherent ECG morphological overlap. Calibration analysis reveals a nuanced picture: while Dataset I shows well-calibrated probabilities (ECE = 0.034), Dataset II exhibits slightly higher Expected Calibration Error (ECE = 0.059) despite perfect hard predictions suggesting overconfident SoftMax outputs. This underscores the need for post-hoc calibration (e.g., temperature scaling) in safety-critical deployments, especially when operating under distributional shift. Explainability analyses using Grad-CAM provided face-valid insights into the model’s decision process. Heatmaps taken from convolutional blocks generally emphasized waveform regions, particularly QRS complexes and ST-segment deviations–indicating reliance on physiologic morphology. However, maps from the attention layer sometimes highlighted page headers and borders, revealing a potential shortcut. This observation motivates practical mitigation before deployment: cropping or masking non-physiologic regions, standardizing plotting areas, and applying augmentations that randomize borders, grid intensity, and brightness. We recommend stress tests–such as masking headers at inference and using deletion/insertion curves to verify that predictions depend on P–QRS–T structure rather than layout artifacts. Despite strong results, limitations remain. Small but persistent errors in the History of MI and Abnormal Heartbeat on Dataset I and reduced HYP performance on Dataset III indicate that subtle or heterogeneous phenotypes remain challenging. Class-aware thresholding, focal or cost-sensitive losses, and augmentation tailored to ECG image idiosyncrasies (e.g., grid perturbation, scan artifacts, or synthetic ST-T changes) may improve recall where clinical risk is asymmetric. While Dataset II provides an initial out-of-distribution check, broader evaluation across institutions, devices, export formats, and acquisition settings including raw 1D signal benchmarks is required to establish robustness. Finally, although we operate on ECG images, many clinical systems store raw multi-lead time series; extending SimCardioNet to 1D signals (or to joint image–signal learning) is a promising avenue to improve physiologic fidelity and reduce sensitivity to presentation artifacts. In summary, SimCardioNet leverages SimCLR and progressive unfreezing to deliver high, balanced performance for ECG classification under limited labels, with encouraging external validation on both small-scale image datasets and large-scale clinical benchmarks. Grad-CAM analyses, calibration metrics, and cross-dataset consistency highlight concrete steps–artifact mitigation, probability calibration, and broader validation–that will further advance reliability and readiness for clinical integration.
Conclusion
We presented SimCardioNet, a self-supervised–then–supervised ECG image classification framework that combines contrastive pretraining (SimCLR), a multi-scale CNN backbone with attention, and progressive unfreezing during fine-tuning. On the internal clinical dataset (Dataset I), the model achieved strong and balanced performance (0.975 accuracy, 0.973 F1-score), with effectively diagonal confusion matrices. External evaluation on Dataset II yielded perfect discrimination, suggesting promising generalization under domain shift, while results on the Dataset III demonstrated competitive performance (0.921 accuracy, 0.921 F1-score), surpassing multiple state-of-the-art baselines in a standardized multi-class setting. Qualitative explanations with Grad-CAM largely confirmed that decisions are driven by waveform regions, although occasional attention to page headers and borders revealed a potential shortcut, motivating straightforward mitigation (cropping/masking non-physiologic regions and layout-randomizing augmentations). Taken together, these results show that self-supervised representation learning–paired with a carefully designed fine-tuning schedule can significantly reduce label dependency while delivering high-quality, generalizable ECG classification across diverse clinical and research settings. Future work will focus on prospective, multi-site validation; explicit domain generalization and test-time adaptation; routine calibration and drift monitoring; and extension of the approach to raw multi-lead time series or joint image–signal architectures. Addressing these directions should strengthen robustness, interpretability, and clinical reliability, paving the way for safe integration of automated ECG interpretation into real-world cardiovascular care pathways.
Data availability
The datasets analyzed during the current study are available in the Mendeley Data repository, under the title “ECG Images Dataset of Cardiac Patients” (DOI: 10.17632/gwbz3fsgp8.2), published on March 19, 2021. The dataset includes ECG images of cardiac patients and is created under the auspices of Ch. Pervaiz Elahi Institute of Cardiology, Multan, Pakistan. The dataset can be accessed publicly for research purposes. If required, further details or supplementary data can be provided upon reasonable request to the corresponding author.
References
Gustafsson, S. et al. Development and validation of deep learning ECG-based prediction of myocardial infarction in emergency department patients. Sci. Rep. 12, 19615. https://doi.org/10.1038/s41598-022-24254-x (2022).
Xiong, P., Lee, S.-Y. & Chan, G. Deep learning for detecting and locating myocardial infarction by electrocardiogram: A literature review. Front. Cardiovasc. Med. 9, 860032. https://doi.org/10.3389/fcvm.2022.860032 (2022).
Ansari, Y., Mourad, O., Qaraqe, K. & Serpedin, E. Deep learning for ECG arrhythmia detection and classification: an overview of progress for period 2017–2023. Front. Physiol. 14, 1246746. https://doi.org/10.3389/fphys.2023.1246746 (2023).
Herman, R. et al. Validation of deep learning system for comprehensive 12-lead ECG interpretation. Circulation 148, A17037. https://doi.org/10.1161/circ.148.suppl_1.17037 (2023).
Oke, O. & Cavus, N. A systematic review on the impact of artificial intelligence on electrocardiograms in cardiology. Int. J. Med. Inform. 195, 105753. https://doi.org/10.1016/j.ijmedinf.2024.105753 (2025).
Kashou, A. et al. A comprehensive artificial intelligence-enabled electrocardiogram interpretation program. Cardiovasc. Digit. Health J. 1, 62–70. https://doi.org/10.1016/j.cvdhj.2020.08.005 (2020).
Alhussan, A. A. et al. Classification of breast cancer using transfer learning and advanced al-biruni earth radius optimization. Biomimetics 8, 270. https://doi.org/10.3390/biomimetics8030270 (2023).
Lu, L. et al. Decoding 2.3 million ECGs: Interpretable deep learning for advancing cardiovascular diagnosis and mortality risk stratification. Eur. Heart J. Digit. Health 5, 247–259. https://doi.org/10.1093/ehjdh/ztae014 (2024).
Zhang, X. et al. Automated detection of cardiovascular disease by electrocardiogram signal analysis: A deep learning system. Cardiovasc. Diagn. Ther. 10, 227–235. https://doi.org/10.21037/cdt.2019.12.10 (2020).
Khan, F., Yu, X., Yuan, Z. & Rehman, A. ECG classification using 1-d convolutional deep residual neural network. PLoS ONE 18, e0284791. https://doi.org/10.1371/journal.pone.0284791 (2023).
Katal, N., Gupta, S., Verma, P. & Sharma, B. Deep-learning-based arrhythmia detection using ECG signals: A comparative study and performance evaluation. Diagnostics 13, 3605. https://doi.org/10.3390/diagnostics13243605 (2023).
Katal, N., Gupta, S., Verma, P. & Sharma, B. Deep-learning-based arrhythmia detection using ECG signals: A comparative study and performance evaluation. Diagnostics (Basel) 13, 3605. https://doi.org/10.3390/diagnostics13243605 (2023).
Chen, X. et al. Acute myocardial infarction detection using deep learning-enabled electrocardiograms. Front. Cardiovasc. Med. 8, 654515. https://doi.org/10.3389/fcvm.2021.654515 (2021).
Makimoto, H. et al. Performance of a convolutional neural network derived from an ECG database in recognizing myocardial infarction. Sci. Rep. 10, 8445. https://doi.org/10.1038/s41598-020-65105-x (2020).
Śmigiel, S., Pałczyński, K. & Ledziński, D. ECG signal classification using deep learning techniques based on the ptb-xl dataset. Entropy 23, 1121. https://doi.org/10.3390/e23091121 (2021).
Diamant, N. et al. Patient contrastive learning: A performant, expressive, and practical approach to electrocardiogram modeling. PLoS Comput. Biol. 18, e1009862. https://doi.org/10.1371/journal.pcbi.1009862 (2022).
Alqaraleh, M., Khleifat, K. M., Abu Hajleh, M. N., Farah, H. S. & Ahmed, K.A.-A. Fungal-mediated silver nanoparticle and biochar synergy against colorectal cancer cells and pathogenic bacteria. Antibiotics 12, 597. https://doi.org/10.3390/antibiotics12030597 (2023).
Sangha, V. et al. Biometric contrastive learning for data-efficient deep learning from electrocardiographic images. J. Am. Med. Inform. Assoc. 31, 855–865. https://doi.org/10.1093/jamia/ocae002 (2024).
Dave, D. et al. Diagnostic test accuracy of Ai-assisted mammography for breast imaging: A narrative review. PeerJ Comput. Sci. 11, e2476. https://doi.org/10.7717/peerj-cs.2476 (2025).
Kandaswamy, M., Vijayakumar, P. R. & Rajagopal, R. K. Detection of cardiovascular diseases in ECG images using deep learning. In AIP Conference Proceedings, Vol. 3279 020164 (AIP Publishing LLC, 2025).
Lee, B., Kong, S., Song, Y. & Lee, Y. Self-supervised learning with electrocardiogram delineation for arrhythmia detection. In Annual International Conference of the IEEE Engineering in Medicine and Biology Society 591–594. https://doi.org/10.1109/EMBC46164.2021.9630364 (2021).
Soltanieh, S., Hashemi, J. & Etemad, A. In-distribution and out-of-distribution self-supervised ECG representation learning for arrhythmia detection. IEEE J. Biomed. Health Inform. 28, 789–800. https://doi.org/10.1109/JBHI.2023.3201956 (2023).
Chen, W. et al. Temporal and spatial self-supervised learning methods for electrocardiograms. Sci. Rep. 15, 6029. https://doi.org/10.1038/s41598-025-90084-2 (2025).
Hussain, S. et al. A swin transformer and CNN fusion framework for accurate Parkinson disease classification in MRI. Sci. Rep. 15, 15117. https://doi.org/10.1038/s41598-025-93671-5 (2025).
Ao, R. & He, G. Image based deep learning in 12-lead ECG diagnosis. Front. Artif. Intell. 5, 1087370. https://doi.org/10.3389/frai.2022.1087370 (2023).
Rustam, F. et al. Enhanced detection of diabetes mellitus using novel ensemble feature engineering approach and machine learning model. Sci. Rep. 14, 23274. https://doi.org/10.1038/s41598-024-74357-w (2024).
Niu, L., Chen, C., Liu, H., Zhou, S. & Shu, M. A deep-learning approach to ECG classification based on adversarial domain adaptation. Healthcare (Basel) 8, 437. https://doi.org/10.3390/healthcare8040437 (2020).
Kolhar, M. & Al Rajeh, A. Deep learning hybrid model ECG classification using alexnet and parallel dual branch fusion network model. Sci. Rep. 14, 26919. https://doi.org/10.1038/s41598-024-78028-8 (2024).
Santhakumar, D., Shree, K. D., Buvanesvari, M., Kumar, A. S. & Salau, A. O. Hd-mvcnn: High-density ECG signal based diabetic prediction and classification using multi-view convolutional neural network. Egypt. Inform. J. 28, 100573 (2024).
Ebrahimi, Z., Loni, M., Daneshtalab, M. & Gharehbaghi, A. A review on deep learning methods for ECG arrhythmia classification. Expert Syst. Appl. X 7, 100033. https://doi.org/10.1016/j.eswax.2020.100033 (2020).
Narotamo, H. et al. Deep learning for ECG classification: A comparative study of 1d and 2d representations and multimodal fusion approaches. Biomed. Signal Process. Control 93, 106141. https://doi.org/10.1016/j.bspc.2024.106141 (2024).
Kumar, A. S. & Rekha, R. A dense network approach with Gaussian optimizer for cardiovascular disease prediction. N. Gener. Comput. 41, 859–878 (2023).
Arunachalam, S. K. & Rekha, R. A novel approach for cardiovascular disease prediction using machine learning algorithms. Concurr. Comput. Pract. Exp. 34, e7027. https://doi.org/10.1002/cpe.7027 (2022).
Rajagopal, R. & Ranganathan, V. Evaluation of effect of unsupervised dimensionality reduction techniques on automated arrhythmia classification. Biomed. Signal Process. Control 34, 1–8 (2017).
Marwa, S. R. & Shakir, M. A. Improving ECG signals classification by using deep learning techniques: A review. In ITM Web of Conferences, Vol. 64 01023 (EDP Sciences, 2024).
Eleyan, A. & Alboghbaish, E. Electrocardiogram signals classification using deep-learning-based incorporated convolutional neural network and long short-term memory framework. Computers 13, 55. https://doi.org/10.3390/computers13020055 (2024).
Selvam, I. J., Madhavan, M. & Kumarasamy, S. K. Detection and classification of electrocardiography using hybrid deep learning models. Hellenic J. Cardiol. 81, 75–84 (2024).
Monday, H. N. et al. Enhancing ECG classification in cardiac diagnostics: A novel approach using adaptive focal cross-entropy loss function. IEEE J. Biomed. Health Inform. https://doi.org/10.1109/JBHI.2025.3205134 (2025).
Saranya, K., Karthikeyan, U., Kumar, A. S., Salau, A. O. & Tin Tin, T. Densenet-abilstm: revolutionizing multiclass arrhythmia detection and classification using hybrid deep learning approach leveraging ppg signals. Int. J. Comput. Intell. Syst. 18, 1–19 (2025).
Kumar, A. S. & Rekha, R. An improved hawks optimizer based learning algorithms for cardiovascular disease prediction. Biomed. Signal Process. Control 81, 104442 (2023).
Wagner, P. et al. Ptb-xl, a large publicly available electrocardiography dataset. Sci. Data 7, 1–15 (2020).
Atwa, A. E. M. et al. Interpretable deep learning models for arrhythmia classification based on ECG signals using ptb-x dataset. Diagnostics 15, 1950 (2025).
Geng, Q. et al. An ECG classification method based on multi-task learning and cot attention mechanism. In Healthcare, Vol. 11 1000 (MDPI, 2023).
Khan, A. H. & Hussain, M. ECG images dataset of cardiac patients. https://doi.org/10.17632/gwbz3fsgp8.2 (2021).
Mhamdi, L., Dammak, O., Cottin, F. & Dhaou, I. B. Artificial intelligence for cardiac diseases diagnosis and prediction using ECG images on embedded systems. Biomedicines 10, 2013 (2022).
Ozdemir, M. A., Ozdemir, G. D. & Guren, O. Classification of covid-19 electrocardiograms by using hexaxial feature mapping and deep learning. BMC Med. Inform. Decis. Mak. 21, 170. https://doi.org/10.1186/s12911-021-01510-7 (2021).
Alsayat, A. et al. Enhancing cardiac diagnostics: A deep learning ensemble approach for precise ECG image classification. J. Big Data 12, 7 (2025).
Sattar, S. et al. Cardiac arrhythmia classification using advanced deep learning techniques on digitized ECG datasets. Sensors 24, 2484. https://doi.org/10.3390/s24082484 (2024).
Aversano, L., Bernardi, M. L., Cimitile, M., Montano, D. & Pecori, R. Characterization of heart diseases per single lead using ECG images and cnn-2d. Sensors 24, 3485 (2024).
Rautela, D., Bajeli, M., Kumar, A. & Vaidya, H. Identifying cardiovascular disorders through ECG image analysis. In 2024 International Conference on Cognitive Robotics and Intelligent Systems (ICC-ROBINS) 607–612 (IEEE, 2024).
Micheal, A. A., Gupta, K., Singh, R., Singh, V. & Mohanty, B. Hbtrackr: Ai-based heart attack prediction through ECG monitoring on wearable devices. In 2024 First International Conference on Pioneering Developments in Computer Science & Digital Technologies (IC2SDT) 557–562 (IEEE, 2024).
Śmigiel, S., Pałczyński, K. & Ledziński, D. ECG signal classification using deep learning techniques based on the ptb-xl dataset. Entropy 23, 1121 (2021).
Rath, A., Mishra, D. & Panda, G. Lstm-based cardiovascular disease detection using ECGn signal. In Cognitive Informatics and Soft Computing: Proceeding of CISC 2020 133–142 (Springer, 2021).
Funding
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2601).
Author information
Authors and Affiliations
Contributions
The original research idea and first draft of the manuscript were conceived and written by Muhammad Dawood Majid. The manuscript was revised by Muhammad Anwar, Syed Fakhar Bilal and Muhammad Zubair to address the technical comments of the reviewers. The revised manuscript was reviewed and edited by Muzammil Hussain for accuracy and clarity. Jabeen Sultana contributed to the manuscript design and data analysis. Muhammad Asif Habib supervised the project and provided the guidance for critical details regarding the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Majid, M.D., Anwar, M., Bilal, S.F. et al. A hybrid learning framework for automated multiclass electrocardiogram classification with SimCardioNet. Sci Rep 16, 7621 (2026). https://doi.org/10.1038/s41598-026-36932-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36932-1