Abstract
This study proposes a wrist radiography-based deep learning model for identifying radiographic features of metabolic bone disease (MBD) of prematurity and evaluate the impact of its decision support. This retrospective study included preterm infants with birth weights under 1500 g, born at Seoul National University Hospital (internal dataset: 814 subjects) and Seoul National University Bundang Hospital (external dataset: 261 subjects). Demographic and clinical information and wrist radiographs (postnatal ages: 4–8 weeks) were collected. An internal dataset was used to develop and train identification models and an external dataset was used for validation. Performance was evaluated using the area under the receiver operating characteristic curve (AUROC) and performance quality was compared based on paired t- and Wilcoxon signed-rank tests. The DenseNet-based model exhibited the highest performance quality with AUROC of 0.961 (sensitivity: 94.4%, specificity: 91.2%, accuracy: 92.0%). The external validation study yielded an AUROC of 0.927, indicating its potential applicability in a broad clinical setting. The reader study using external data demonstrated an improved reading performance, especially for non-radiologists (65.4% to 78.7% accuracy; P = 0.008, among pediatricians). The developed model can assist clinicians, especially non-radiologists, in identifying radiographic signs suggestive of MBD, enabling timely diagnosis and treatment to prevent disease progression.
Similar content being viewed by others
Introduction
Metabolic bone disease (MBD) of prematurity is also known as osteopenia or rickets of prematurity1. It is prevalent in preterm infants, especially those with a gestational age of < 28 weeks. Small for gestational age, birth weight < 1000 g, gestational age (GA) < 28 weeks, and total parenteral nutrition (TPN) period > 28 days1,2,3,4 are renowned risk factors for MBD of prematurity. Between 16 and 40% of preterm infants with very low birth weight (< 1500 g) and extremely low birth weight (< 1000 g) infants are diagnosed with MBD of prematurity, with a peak incidence at the postnatal age of 4–8 weeks1,5,6.
However, the exact incidence of MBD of prematurity among preterm infants remains unknown owing to the lack of consensus regarding its diagnostic definition7. Several screening protocols exist for selecting and evaluating infants who are at high risk of MBD due to prematurity using biochemical markers and wrist radiography7. Because it is attributable to calcium and phosphorus deficiency due to decreased intake or absorption in most cases1,2,8,9, clinicians suspect the presence of MBD if the serum levels of phosphorus and calcium are persistently low and the serum alkaline phosphatase level increases2,10. Wrist radiography is often performed for a more definitive diagnosis, which might demonstrate cupping or fraying at the radius metaphysis, the classical radiological finding of MBD of prematurity6,7,11,12. However, such radiological findings appear only after bone mineralization has been reduced by 20–40%; if not, identifying the early stage of MBD on radiographs can be challenging6,7.
A convolutional neural network is a complex computational model that uses multiple algorithm layers to achieve a high-level interpretation of data. Convolutional neural networks are widely applied to the classification of medical images13 and are generally based on a supervised approach14. In recent years, numerous studies on artificial intelligence (AI) have revealed that applying deep learning to medical images has improved the speed, accuracy, and quality of image interpretation and radiological diagnosis15,16. However, a diagnostic program for MBD of prematurity using data from preterm infants has not yet been developed. Because MBD of prematurity is a major complication of prematurity, this study proposes a deep learning model using wrist radiographs obtained from very low birth weight infants between 4 and 8 weeks of age and its clinical efficacy for identifying MBD of prematurity is evaluated. The proposed algorithm is expected to enhance the accuracy and efficacy of MBD diagnosis in preterm infants when integrated with clinical and biochemical assessment, thereby allowing earlier intervention and treatment for preventing disease progression.
Methods
Study design and dataset
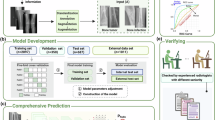
This retrospective study enrolled preterm infants with very-low birth weights born at Seoul National University Hospital (SNUH) and Seoul National University Bundang Hospital (SNUBH) from 2010 to 2020 and 2017 to 2020, respectively. The Institutional Review Board of SNUH approved the study, waiving the requirement for informed consent. This study was conducted and performed in accordance with relevant guidelines and regulations, including the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) and the Checklist for Artificial Intelligence in Medical Imaging (CLAIM) guidelines. Infants who had been discharged, transferred, or deceased before 28 days, those with associated anomalies, and those without wrist radiographs between 4 and 8 weeks postnatal age were excluded (Fig. 1). Demographic and clinical data were retrieved from electronic medical records. Wrist radiographs taken between 4 and 8 weeks of life were collected from SNUH and SNUBH to construct internal and external datasets, respectively.
Overall flowchart of the study design. This figure depicts the study population selection, imaging dataset distribution, augmentation process, and deep learning training and testing procedures.
Patients were categorized into MBD and normal groups: those diagnosed with MBD at least once on wrist radiographs were classified as MBD. A pediatric radiologist and neonatologist (C.J.E. and K.E.K., 22 and 19 years of post-fellowship experience, respectively) who were blinded to the clinical conditions of the patients reviewed 2239 wrist radiographs of 814 patients. Images were labeled “0” if they look normal and “1” if they revealed MBD based on the screening criteria suggested by Koo et al.12 and used as the ground truth. Koo’s criteria of grade 2 and above is designated as MBD. Importantly, because the diagnosis of MBD requires comprehensive integration of clinical and radiographic features, these labels represent radiographic suspicion of MBD rather than confirmed clinical diagnoses. Discrepant categorizations were resolved through consensus. The healing state of the MBD showing radiolucent bands at the metaphysis was also classified as MBD.
Imaging dataset processing
Normal images were randomly down-sampled by 60% to balance the size of training data (1968 vs. 271 images). Images were split into train, tuning, and test sets at a ratio of 7:1:2 via stratified random sampling (Fig. 1).
Images were converted from DICOM to PNG using Python 3.7.11. Annotations were performed using roLabelImg version 1.8.0. Metaphyseal alterations of the radius are key features of MBD12; therefore, the region of interest (ROI) was drawn as a square surrounding the metaphyseal plate of the radius (Fig. 2), thus focusing on the defined area while minimizing input from unnecessary areas during training. Since this study aimed to develop an identification model rather than a screening tool, and given that metaphyseal alterations of the radius are the definitive radiological features of MBD12, we implemented precise manual ROI selection, focusing specifically on the metaphyseal plate of the radius (Fig. 2). This ROI selection ensures that the model learns from the exact anatomical regions that specialists use for MBD diagnosis. By methodically defining the ROI to include clinically important regions, our deep learning models not only concentrate on areas that are most relevant but also enhance both the accuracy and efficiency of the models.
Definition of ROI. Detailed guidelines for drawing the region of interest (ROI) on wrist X-rays.
Annotations were delegated to a neonatologist (P.S.G., 1 year of post-fellowship experience) for consistency and saved in an XML file. ROIs were cropped and resized to 300 × 300 pixels for EfficientNet-b3 and 224 × 224 pixels for the other algorithms. Our deep learning models, which were pretrained on the ImageNet17 dataset, were trained to identify image features indicative of MBD.
In the training phase, images were augmented using Albumentation version 1.1.018, adjusting rotation, brightness, contrast, GaussianBlur, and GaussNoise. Rotation by 1° was made randomly within the range of ± 15°. Random brightness and contrast were adjusted by −0.2 to + 0.2 using default settings. For GaussianBlur, a random-sized kernel with a center pixel of (3,7) was applied to obtain new values for the center pixels of (3,3), (4,4), (5,5), (6,6), and (7,7). GaussNoise was applied with a variance range for noise between 10 and 50.
Model development and external validation
Seven algorithms (AlexNet19, DenseNet-12120, ResNet-5021, ResNext-5022, ChexNet23, VGG-1924, and EfficientNet-B325 were used with ImageNet-pretrained models as the baseline26,27. The hyperparameters were consistent across models: Adam optimizer with a learning rate of 1e-4, early stopping, batch size of 16, and cross-entropy loss function. An Intel Core i5-11400 F CPU at 2·60 GHz and an NVIDIA RTX 3060 (12 GB) GPU were used for image processing and neural network development. Coding was performed using PyTorch version 1.8.228. An unbalanced number of MBD and normal images were used as the internal test dataset and external dataset to reflect the normal distribution of images obtained in clinical settings considering the prevalence of MBD. Each method was evaluated using evaluation metrics to select the most accurate model. External dataset validation was then performed.
Reader study
Model utility in a clinical setting was evaluated through reader studies planned as crossover designs. For the internal dataset reader study, 11 clinicians read 225 wrist radiographs (171 normal, 54 MBD images) from 83 patients twice (without and with AI prediction) with a minimum two-week washout period between readings. The test set of the internal dataset was used for this reader study (Fig. 1).
To validate the generalizability of our model’s clinical utility, an additional crossover reader study was conducted using the external dataset. A total of 11 clinicians read 309 wrist radiographs (276 normal, 33 MBD images) from 261 patients twice with a minimum two-week washout period between readings (Fig. 1).
The clinicians’ reading performance and time were assessed and compared to evaluate the effectiveness of the developed model across different datasets.
Data analysis
Statistical analyses were performed using IBM SPSS Statistics for Windows version 26.0. Demographic features and clinical information were analyzed using the chi-square test for categorical variables and the student’s t-test for continuous variables. Because infants born at a lower GA have a lower birth weight and Apgar scores, as well as longer TPN period, duration to reach full enteral feeding, and hospital stay29, multivariate logistic regression analysis was performed with adjustments for sex, GA, birth weight, TPN period, full enteral nutrition reaching period, length of hospital stay, and Apgar score. The area under the receiver operating characteristic curve (AUROC) was generated to define the test accuracy, and evaluation metrics such as sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), accuracy, and F1-score, were calculated to evaluate the model’s performance30,31. The AUROCs were compared using the DeLong test32 and the R plug-in package pROC33 (R version 4.3.0; pROC 1.18.2). For both internal and external reader studies, the median values of the participants’ performance metrics with or without the assistance of the developed model were compared using a paired t-test or Wilcoxon’s signed-rank test. Statistical significance was defined as P < 0.05.
Results
Demographic and clinical data and imaging dataset Preparation
Between 2010 and 2020, 1110 preterm infants with very-low birth weights were born at SNUH and admitted to the neonatal intensive care unit (NICU). Among them, 296 were excluded, of which 257 were discharged, transferred to other hospitals, or were deceased in less than 28 days, and 39 did not undergo wrist radiography between 4 and 8 weeks of postnatal age (25–59 days, with a window period of 3 days). None of the patients had any major congenital anomalies, metabolopathies, or chromosomal abnormalities. A total of 814 patients were enrolled in this study (Fig. 1). The study population comprised 51.1% (416/814) males and 48.9% (398/814) females. The mean (± standard deviation) GA and birth weight were 28.97 (± 2.69) weeks and 1033.48 (± 270.32) grams, respectively. MBD was present in 14.3% (116/814) of the population. Table 1 summarizes the demographic and basic clinical features of each group. Biochemical data collected at the time of wrist radiography showed higher ALP levels (536.90 vs. 410.53 IU/L, P < 0.001) and lower calcium levels (9.645 vs. 9.967 mg/dL, P = 0.015) when patients showed radiographic features suggestive of MBD (Table 2). The MBD group showed lower GA (OR 0.751, 95% CI 0.658–0.857, P < 0.001), lower birth weight (OR 0.997, 95% CI 0.996–0.999, P < 0.001), longer TPN period (OR 1.022, 95% CI 1.001–1.044, P = 0.04), and longer hospital stay (OR 1.007, 95% CI 1.000–1.015.000.015, P = 0.046) were significantly associated with a higher risk of MBD prematurity (Table 3) as reported in previous studies6,7.
Wrist radiographs from SNUH were used as an internal dataset to train the model. The dataset comprised 2239 wrist images obtained from 814 patients, with 271 images being categorized as MBD and 1968 as normal. After down-sampling the normal images, the final internal dataset contained 833 normal images from 298 patients and 271 MBD images from 116 patients. The dataset was then divided into train (776 images; 585 normal, 191 MBD), tuning (103 images; 77 normal, 26 MBD), and test (225 images; 171 normal, 54 MBD) sets at a ratio of 7:1:2 by stratified random sampling to avoid data leakage, considering our research involved multiple scans of the same patient34.
Deep learning model training using the internal dataset and external validation study
Seven deep learning algorithms were developed in our study, and their performances were evaluated in terms of the previously mentioned metrics (Table 4). The AUROCs of the models developed with AlexNet, DenseNet-121, ResNet-50, and ResNext-50 were 0.837, 0.961, 0.960, and 0.941, respectively. Model developed using CheXNet showed the AUROC of 0.954, while that of EfficientNet-B3 was 0.954 and VGG-19 was 0.870 (Fig. 3).
Receiver operating characteristic (ROC) curves of the developed models using various algorithms. DenseNet-121 (orange line) exhibits the highest area under the ROC curve, suggesting the highest performance in terms of quality in both (a) the internal data training and (b) the external validation study.
To ensure robustness and generality, model performance was compared using AUROCs with DeLong’s test (P < 0.05)32. Considering the importance of accurately diagnosing patients with this disease, we chose DenseNet-121, which showed the highest AUROC, to aid interpretation during the reader study. DenseNet-121 had a sensitivity of 94.4% (51/54), specificity of 91.2% (151/171), PPV of 77.2% (51/66), NPV of 98.1% (156/159), F1-score of 84.9% (102/120), accuracy of 92.0% (207/225), and AUROC of 0.961.
Images for the external validation study were collected from 261 patients at SNUBH and comprised 276 normal images and 33 MBD images. An external validation study performed on DenseNet-121 resulted in a sensitivity of 75.7% (25/33), specificity of 88.0% (243/276), PPV of 43.1% (25/58), NPV of 96.8% (243/251), F1-score of 54.9% (50/91), accuracy of 86.7% (268/309), and AUROC of 0.927.
Evaluation of the model’s effectiveness in clinical settings
Eight pediatricians and three radiologists participated in the study using internal dataset to evaluate the clinical effectiveness of the model (four resident pediatricians in training, four first-year neonatologist fellows, two resident radiologists in training, and one pediatric radiologist with 1 year of post-fellowship experience). We compared the metrics achieved while detecting MBD from wrist radiographs with and without the proposed model’s prediction. The reading time of one pediatrician was omitted from the data analysis because the reader study was disrupted by a medical emergency. When provided with our model’s prediction (predicted percentage of the given image being MBD) using internal dataset, clinicians experienced significant improvement in their performance (without AI vs. with AI: sensitivity (%) 80.4 vs. 91.5 (P = 0.02), specificity (%) 59.8 vs. 80.2 (P < 0.001), PPV (%) 42.1 vs. 64.2 (P < 0.001), NPV (%) 88.6 vs. 96.9 (P = 0.006), accuracy (%) 64.8 vs. 82.9 (P < 0.001), and F1-score (%) 52.5 vs. 73.3 (P < 0.001)) (Table 5; Fig. 4a). The mean time taken to complete the reading was 1740.5 s and 1522.0 s without and with the support of our model (P = 0.08), respectively.
Comparison between the clinicians’ reading performance without (blue) and with (orange) AI assistance. The boxes show the interquartile ranges between the first and third quartiles, and the whiskers are extended to the ± 1.5 interquartile range. Circles represent outliers. For each performance metric, the mean values are calculated (standard deviation within parentheses) among the clinicians, and the comparison is made. A paired t-test is performed for the data following a normal distribution; if not, the Wilcoxon signed-rank test is used. (a) Performance comparison among 11 clinicians after internal dataset reader study shows significant improvement in terms of sensitivity, specificity, PPV, NPV, accuracy, and F1-score when assisted by our model. (b) External dataset reader study also demonstrates significant improvement in terms of specificity, PPV, accuracy, and F1-score. Reading time is also reduced in both reader studies, but no statistical significance is observed.
Additional reader study using external dataset involved eight pediatricians and three radiologists (one first-year and four second-year neonatologist fellows, two neonatologists with 1 year of post-fellowship experience, one neonatologist with 1 year of post-fellowship experience, one first-year pediatric radiologist fellow, one pediatric radiologist with 1 year of post-fellowship experience, one general radiologist with 1 year of post-fellowship experience) to assess our model’s effect on clinical decision even with the external dataset. When provided with our model’s prediction using external dataset, clinicians experienced significant improvement in their performance (without AI vs. with AI: specificity (%) 62.3 vs. 78.8 (P = 0.001), PPV (%) 24.3 vs. 34.5 (P = 0.001), accuracy (%) 64.8 vs. 79.6 (P = 0.001), and F1-score (%) 36.6 vs. 48.7 (P < 0.001)). However, sensitivity did not show significant improvement in their performance (without AI vs. with AI: 85.6% vs. 87.0%) (Table 5; Fig. 4b).
AI assistance significantly improved the diagnostic performance of pediatricians with limited experience in wrist radiograph interpretation. In the internal dataset reader study, the accuracy of pediatricians increased from 63.2% to 82.0% (P = 0.008), while radiologists also showed an improvement from 69.0% to 85.4%, though not statistically significant (P = 0.14) (Table 5; Fig. 5a). Similar trends were observed in the external dataset, where pediatricians’ accuracy improved significantly from 65.4% to 78.7% (P = 0.008), while radiologists showed a non-significant increase from 63.1% to 82.1% (P = 0.20) (Table 5; Fig. 5b). These improvements were more pronounced among pediatricians in both studies. The specificity, PPV, and F1-score also showed statistically significant enhancement among pediatricians, whereas improvements among radiologists were not statistically significant (Supplementary Fig. 1 and Supplementary Fig. 2). In terms of sensitivity and specificity, both pediatricians and radiologists demonstrated improvements with AI support in the internal dataset. In the external dataset, AI assistance led to notable gains in both sensitivity and specificity for pediatricians, while radiologists showed sustained sensitivity and improved specificity (Supplementary Fig. 3).
Accuracy comparison of metabolic bone disease diagnosis between pediatricians (green) and radiologists (blue). Improvement in accuracy with the assistance of our model is more evident among pediatricians only in both (a) internal dataset reader study and (b) external dataset reader study.
AI assistance corrected 51.6% of false positives and 65.5% of false negatives in the internal reader study, and 43.7% and 34.6% in the external reader study, respectively, thereby demonstrating its utility in mitigating both overdiagnosis and missed diagnoses. Overall, the utility of our model for clinicians in identifying MBD was well demonstrated, especially by non-radiologists.
Discussion
MBD is a preterm complication that can lead to pathological fractures5,8. However, radiologists’ readings can be delayed, and non-radiologists’ reading accuracy may be limited. Therefore, for a more efficient and accurate radiographic identification of MBD of prematurity in clinical settings, we developed a deep learning model that can improve clinicians’ reading performance, especially that of non-radiologists.
Of the 814 patients in the internal dataset, 14.3% (116/814) were found to have MBD based on wrist radiographs. The observed incidence of MBD in our study population was close to the reported incidence of 16–40% among infants with very- and extremely-low birth weights5. Male sex is a known risk factor for MBD of prematurity6, which is consistent with our population’s sexual characteristics (61.2% male in the MBD group vs. 49.4% in the normal group, P = 0.021). The clinical findings and risk factor analysis showed that lower GA (OR 0.751, 95% CI 0.658–0.857, P < 0.001), lower birth weight (OR 0.997, 95% CI 0.996–0.999, P < 0.001), and longer TPN period (OR 1.022, 95% CI 1.001–1.044, P = 0.04) were associated with a higher risk of MBD, which is consistent with the known risk factors for MBD of prematurity. These statistical findings indicate that our study population appropriately represented the general preterm population, thereby supporting the reliability of the ground truth.
Grad-CAM images of true-positive and true-negative cases showed that our model focused mainly on the area of the radius metaphysis (Supplementary Fig. 4a and b). However, false positives and false negatives showed that the model focused on the ulna or intravenous fluid line more than the metaphysis area of the radius (Supplementary Fig. 4c and d); this may have decreased the model’s accuracy because the MBD of prematurity radiological definition should not be applied to isolated cupping of the distal ulna35. Although attention to artifacts could impair performance, we deliberately retained images containing intravenous line, rotated or tilted wrists, and variable positioning. Because strict image acquisition is challenging in preterm infants, training the model with such heterogeneous images may enhance its applicability to real NICU settings.
Although a relatively low PPV in external validation test is observed, this is attributed to the class imbalance in our external dataset (276 normal images vs. 33 MBD images), reflecting the real-world prevalence of MBD among preterm infants. Such relationship between disease prevalence and PPV is a fundamental principle in diagnostic medicine. Lower disease prevalence exacerbates data imbalance, leading to increased false positives, thereby inevitably resulting in a lower PPV. To demonstrate the model’s intrinsic discriminative ability under balanced conditions, we conducted additional analysis with balanced datasets using random down sampling of normal cases (Supplementary Table S1). In this balanced condition analysis, our model showed substantially improved performance metrics, with DenseNet-121 achieving a PPV of 90.3%. The model maintains high NPV and AUROC, which suggests that our model’s performance is clinically meaningful when interpreted in the context of the natural disease prevalence.
In real-world settings, wrist radiographs are usually obtained when MBD is suspected based on risk factors, biochemical markers, or clinical symptoms. Thus, the observed PPV represents meaningful diagnostic information for confirming or excluding suspected MBD. The high NPV (96.8%) is particularly valuable for safely ruling out MBD in clinically suspected cases, while the moderate PPV still provides useful confirmatory evidence when integrated with clinical judgment. Furthermore, precision (PPV) – recall (sensitivity) curve analysis (Supplementary Fig. 5) demonstrates that our model’s performance threshold can be adjusted based on clinical priorities, allowing optimization for either higher precision (up to 83.0% internal, 85.0% external) when specificity is crucial, or higher recall (up to 96.3% internal, 81.8% external) when sensitivity is prioritized for not missing cases.
Reader study using internal dataset showed that clinicians’ diagnostic performance significantly improved with our model’s assistance, especially for non-radiologists. Accuracy also showed a significant improvement with AI assistance in the reader study using external dataset, suggesting our model’s general applicability in various clinical settings. Despite the excellent performance of our model, PPV results are also relatively low in both internal (without AI 42.1% vs. with AI 64.2%) and external (without AI 24.3% vs. with AI 34.5%) dataset reader studies. Again, this can be explained with the imbalanced dataset (internal reader study used 171 normal and 54 MBD images, and external reader study used 276 normal and 33 MBD images) provided for the reader study, which reflects real prevalence of MBD among preterm infants. When using balanced dataset, the PPV values increased substantially in both internal (without AI 73.0% vs. with AI 88.1%) and external (without AI 71.7% vs. with AI 80.5%) dataset reader studies (Supplementary Table ).
Studies applying deep learning approaches to MBD diagnosis are extremely limited in the literature. Meda et al. developed an AI diagnostic tool for MBD using 104 MBD and 264 normal medical images from patients aged under seven years36. However, to our knowledge, there have been no previous studies developing a deep learning model specifically for MBD diagnosis in preterm infants. Our model, trained solely on wrist radiographs from preterm infants, may be more appropriate for use in NICU settings. While we utilized established deep learning architectures to ensure reliable clinical implementation, our work addresses a significant clinical need by providing the first specialized AI tool for this vulnerable population.
This advancement is particularly critical for MBD in preterm infants, a prevalent disease that can benefit significantly from accurate diagnosis to provide prompt, accurate intervention. Implementing this model in a clinical decision support system (CDSS) not only enables healthcare providers to perform better diagnoses and interventions but also to receive diagnostic assistance to enhance their capability, especially when access to professional radiologists is limited. The integration of our deep learning model into a CDSS could be transformative for regions with limited medical resources, such as developing countries, or primary care settings, where specialized medical expertise is scarce. The deployment of such advanced diagnostic tools through mobile and tablet apps not only makes medical care more accessible but can also be extended to areas such as ubiquitous and pervasive computing, wearable technology, and primary healthcare, thus broadening the impact of healthcare delivery and improving patient outcomes. The threshold adjustability demonstrated in our precision (PPV) - recall (sensitivity) analysis (Supplementary Fig. 5) further enhances clinical applicability by allowing healthcare providers to customize the diagnostic sensitivity based on their specific clinical context and resource availability.
Limitations and future directions
Our study has several limitations. Our model’s prediction represents radiographic interpretation and should be integrated with clinical biochemical assessments for a definitive diagnosis. Although expert-defined, the ground truth is a subjective radiographic interpretation and may not represent a definitive clinical diagnosis. Therefore, potential errors or biases inherent in labeling may have influenced both the model’s training and its performance evaluation. Our model’s performance can be improved with larger datasets. Our training dataset was collected from a single center, though external validation was performed using the dataset obtained from a different center. Therefore, the applicability of the model to the general preterm population at any postnatal age might be limited. Using images from multiple centers at any postnatal age for training could enhance our model’s generalizability.
Additionally, while data augmentation techniques were applied to reflect real-world NICU acquisition conditions, the model can be further strengthened by more comprehensively addressing acquisition variability (e.g., rotation, flexion and beam angles). Future studies could explore more comprehensive augmentation strategies available in the Albumentations library, particularly those that simulate the diverse positional and angular variations encountered in clinical neonatal radiography. Implementation of such enhanced augmentation techniques may further improve the model’s robustness and clinical applicability in varied NICU settings.
From a methodological perspective, while we chose established deep learning architectures to ensure clinical reliability, future studies could explore state-of-the-art approaches to potentially enhance model performance. This could include advanced feature extraction methods, attention mechanisms, sophisticated ensemble approaches, as well as interactive diagnostic systems with dialogue capabilities. Such systems could provide more interpretable results and facilitate better collaboration between AI and clinicians, potentially enhancing the diagnostic process while maintaining clinical reliability. Additionally, implementation of automated ROI detection methods could enhance clinical practicality while maintaining diagnostic accuracy, particularly in resource-limited settings. To this end, we have developed a YOLOv1137-based automated ROI detection model that addresses inter-observer variability concerns (Supplementary Fig. 6).
Although clinical efficacy testing based on a reader study showed significant improvement in reading performance, the comprehensive analysis was limited by the small number of participants. Additionally, while the proposed model improved the reading performance among non-radiologists more than among radiologists, this result might lack statistical power considering that only three radiologists participated in the reader study. The reading time decreased when supported by our model; however, its statistical significance was not proven owing to the small number of participants. Improvements in performance and reading time could be more appropriately examined with more participants, and comparisons of clinical experience across physicians might be established.
Moreover, our model aims to radiographically diagnose MBD in preterm infants, thereby allowing for earlier intervention. However, its prediction is currently limited to radiographic interpretation rather than confirmed clinical diagnosis, which requires comprehensive integration of clinical and biochemical information. Inclusion of biochemical indicators in the multi-modal deep learning model should be considered to establish a more useful and accurate diagnostic model. Furthermore, our model was designed to distinguish MBD from normal cases, without differentiating between disease stages (e.g., early or healing). To enable the categorization of MBD phases and facilitate the development of more suitable and tailored medical therapies, the proposed model should be trained for various MBD stages. While our study focused on the 4–8 week period where peak incidence typically occurs, we observed that radiological signs of MBD emerged at various time points rather than following fixed patterns, likely due to individual variations in disease progression. With prospective data collection and larger datasets, future studies could potentially develop more sophisticated models that analyze the progression of radiological findings over time, which could help identify optimal time points for diagnosis and enable earlier intervention.
Conclusions
MBD of prematurity is an important complication of premature birth. However, no deep learning diagnostic model for infants with MBD of prematurity has been developed based on data obtained entirely from preterm infants. To enhance the quality of diagnosis and medical interventions, we developed an identification deep learning model for MBD of prematurity. An external validation study revealed its applicability in clinical settings, and reader studies confirmed that it can facilitate a significant improvement in identifying radiographic signs of MBD for clinicians, especially for non-radiologists. This model can assist clinicians in detecting MBD more accurately and conveniently, thereby enabling timely treatment and prevention of disease progression in preterm infants. Moreover, this model can help detect MBD among preterm infants in developing countries where access to professional radiologists is limited.
Data availability
De-identified participant data and the dataset used in this study can be made available upon reasonable request, which must include an appropriate protocol, analysis plan, and data exchange with institutional approvals. This request needs to be formally addressed to the corresponding author via email.
Abbreviations
- AI:
-
Artificial intelligence
- AUROC:
-
Area under the ROC curve
- CDSS:
-
Clinical decision support system
- GA:
-
Gestational age
- MBD:
-
Metabolic bone disease
- NICU:
-
Neonatal intensive care unit
- NPV:
-
Negative predictive value
- PPV:
-
Positive predictive value
- ROC:
-
Receiver operating characteristic
- ROI:
-
Region of interest
- TPN:
-
Total parenteral nutrition
References
Rustico, S. E., Calabria, A. C. & Garber, S. J. Metabolic bone disease of prematurity. J. Clin. Transl Endocrinol. 1 (3), 85–91. https://doi.org/10.1016/j.jcte.2014.06.004 (2014).
Kavurt, S. et al. Evaluation of radiologic evidence of metabolic bone disease in very low birth weight infants at fourth week of life. J. Perinatol. 41 (11), 2668–2673. https://doi.org/10.1038/s41372-021-01065-y (2021).
Gaio, P. et al. Incidence of metabolic bone disease in preterm infants of birth weight < 1250 g and in those suffering from bronchopulmonary dysplasia. Clin. Nutr. ESPEN. 23, 234–239. https://doi.org/10.1016/j.clnesp.2017.09.008 (2018).
Avila-Alvarez, A. et al. Metabolic bone disease of prematurity: risk factors and associated short-term outcomes. Nutrients 12 (12), 3786. https://doi.org/10.3390/nu12123786 (2020).
Chacham, S., Pasi, R., Chegondi, M., Ahmad, N. & Mohanty, S. B. Metabolic bone disease in premature neonates: an unmet challenge. J. Clin. Res. Pediatr. Endocrinol. 12 (4), 332–339. https://doi.org/10.4274/jcrpe.galenos.2019.2019.0091 (2020).
Faienza, M. F. et al. Metabolic bone disease of prematurity: diagnosis and management. Front. Pediatr. 7, 143. https://doi.org/10.3389/fped.2019.00143 (2019).
Moreira, A., Jacob, R., Lavender, L. & Escaname, E. Metabolic bone disease of prematurity. NeoReviews 16 (11), e631–641. https://doi.org/10.1542/neo.16-11-e631 (2015).
Chinoy, A., Mughal, M. Z. & Padidela, R. Metabolic bone disease of prematurity: causes, recognition, prevention, treatment and long-term consequences. Arch. Dis. Child. Fetal Neonatal Ed. 104 (5), F560–F566. https://doi.org/10.1136/archdischild-2018-316330 (2019).
Stewart, K., Rittenhouse, M., Gloeckner, J. & Torisky, D. Screening for metabolic bone disease in preterm infants. Infant Child. Adolesc. Nutr. 7 (5), 229–232. https://doi.org/10.1177/1941406415601482 (2015).
Rayannavar, A. & Calabria, A. C. Screening for metabolic bone disease of prematurity. Semin Fetal Neonatal Med. 25 (1), 101086. https://doi.org/10.1016/j.siny.2020.101086 (2020).
Chang, C. Y. et al. Imaging findings of metabolic bone disease. Radiographics 36 (6), 1871–1887. https://doi.org/10.1148/rg.2016160004 (2016).
Koo, W. W., Gupta, J. M., Nayanar, V. V., Wilkinson, M. & Posen, S. Skeletal changes in preterm infants. Arch. Dis. Child. 57 (6), 447–452. https://doi.org/10.1136/adc.57.6.447 (1982).
Erickson, B. J., Korfiatis, P., Akkus, Z. & Kline, T. L. Machine learning for medical imaging. Radiographics 37 (2), 505–515. https://doi.org/10.1148/rg.2017160130 (2017).
Moore, M. M., Slonimsky, E., Long, A. D., Sze, R. W. & Iyer, R. S. Machine learning concepts, concerns and opportunities for a pediatric radiologist. Pediatr. Radiol. 49 (4), 509–516. https://doi.org/10.1007/s00247-018-4277-7 (2019).
Kahn, C. E. Jr. From images to actions: opportunities for artificial intelligence in radiology. Radiology 285 (3), 719–720. https://doi.org/10.1148/radiol.2017171734 (2017).
Chartrand, G. et al. Deep learning: A primer for radiologists. Radiographics 37 (7), 2113–2131. https://doi.org/10.1148/rg.2017170077 (2017).
Russakovsky, O. et al. ImageNet large scale visual recognition challenge. Int. J. Comput. Vis. 115, 211–252. https://doi.org/10.1007/s11263-015-0816-y (2015).
Buslaev, A. et al. Albumentations: fast and flexible image augmentations. Information 11 (2), 125. https://doi.org/10.3390/info11020125 (2020).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 25 (2), 1097–1105. https://doi.org/10.1145/3065386 (2012).
Iandola, F. et al. DenseNet: implementing efficient ConvNet descriptor pyramids. Preprint at. https://doi.org/10.48550/arXiv.1404.1869 (2014).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. 2016 IEEE Conf. Comput. Vis. Pattern Recognit. (CVPR). 770-778 https://doi.org/10.1109/CVPR.2016.90 (2016).
Xie, S. et al. Aggregated residual transformations for deep neural networks. IEEE Conference on Computer Vision and (CVPR), 5987–5995; (2017). https://doi.org/10.1109/CVPR.2017.634 (2017).
Rajpurkar, P. et al. Radiologist-level pneumonia detection on chest X-rays with deep learning. Preprint at. https://doi.org/10.48550/arXiv.1711.05225 (2017).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. Preprint at. https://doi.org/10.48550/arXiv.1409.1556 (2015).
Tan, M., Le, Q. V. & EfficientNet Rethinking model scaling for convolutional neural networks. Preprint at. https://doi.org/10.48550/arXiv.1905.11946 (2019).
Sehar, N., Krishnamoorthi, N. & Vinoth Kumar, C. Deep learning Model-Based detection of anemia from conjunctiva images. Healthc. Inf. Res. 31 (1), 57–65. https://doi.org/10.4258/hir.2025.31.1.57 (2025).
Shermin, T. et al. Enhanced transfer learning with ImageNet trained classification layer in Image and Video Technology PSIVT Lecture notes in Computer Science, vol. 11854 (Springer, Cham., 2019); (2019). (ed. Lee, C., Su, Z. & Sugimoto, A.) https://doi.org/10.1007/978-3-030-34879-3_12
Paszke, A. et al. PyTorch: an imperative style, high-performance deep learning library. Adv. Neural Inf. Process. Syst. 32, 8024–8035. https://doi.org/10.48550/arXiv.1912.01703 (2019).
Walsh, M. C. et al. Neonatal outcomes of moderately preterm infants compared to extremely preterm infants. Pediatr. Res. 82 (2), 297–304. https://doi.org/10.1038/pr.2017.46 (2017).
Handelman, G. S. et al. Peering into the black box of artificial intelligence: evaluation metrics of machine learning methods. AJR Am. J. Roentgenol. 212 (1), 38–43. https://doi.org/10.2214/AJR.18.20224 (2019).
Jang, B. K. & Park, Y. R. Development and validation of adaptable skin cancer classification system using dynamically expandable representation. Healthc. Inf. Res. 30 (2), 140–146. https://doi.org/10.4258/hir.2024.30.2.140 (2024).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: A nonparametric approach. Biometrics 44 (3), 837–845 (1988).
Robin, X. et al. pROC: an open-source package for R and S + to analyze and compare ROC curves. BMC Bioinform. 12, 77. https://doi.org/10.1186/1471-2105-12-77 (2011).
Dao, B. T., Nguyen, T. V., Pham, H. H. & Nguyen, H. Q. Phase recognition in contrast-enhanced CT scans based on deep learning and random sampling. Med. Phys. 49 (7), 4518–4528. https://doi.org/10.1002/mp.15551 (2022).
Oestreich, A. E. Concave distal end of ulna metaphysis alone is not a sign of rickets. Pediatr. Radiol. 45 (7), 998–1000. https://doi.org/10.1007/s00247-014-3268-6 (2015).
Meda, K. C., Milla, S. S. & Rostad, B. S. Artificial intelligence research within reach: an object detection model to identify rickets on pediatric wrist radiographs. Pediatr. Radiol. 51 (5), 782–791. https://doi.org/10.1007/s00247-020-04895-8 (2021).
Khanam, R. & Hussain YOLOv11: An overview of the Key Architectural Enhancements. Preprint at (2024). https://doi.org/10.48550/arXiv.2410.17725
Acknowledgements
We thank Seul Bi Lee, June Young Seo, Kanghwi Lee, Jae Won Choi, and Seok Young Koh from the Department of Radiology, Seoul National University Hospital, Seoul, Korea; Chang Hyun Ryoo from the Department of Radiology, Nowon Eulji Medical Center, Seoul, Korea; Jae Hui Ryu, Hye Su Hwang, Ki Teak Hong from the Department of Pediatrics, Ewha Womans University Medical Center, Seoul, Korea; Eun Woo Nam from the Department of Pediatrics, Dankook University Hospital, Cheonan, Chung Nam, Korea; Seung Hoon Lee, Baek Sup Shin, Gyeong Eun Yeom, Hae Min Kang, Dabin Kim, Jihye Yoon, Jieun Jeong, and Yeong Seok Lee from the Department of Pediatrics, Seoul National University Children’s Hospital, Seoul, Korea; and Ho Jung Choi from the Department of Pediatrics, SMG-SNU Boramae Medical Center, Seoul, Korea, for participating in our reader study.This work was supported by the Institute of Information & Communications Technology Planning & Evaluation (IITP) - Global Data-X Leader HRD program (IITP-2024-RS-2024-00441407).
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Contributions
Park SG, Jeong S, Cho MW, Cheon J-E, Kong H-J, Kim E-K, Choi CW conceptualized and designed the study. Park SG, Jeong S, and Kim MJ collected and verified the data. Cheon J-E and Kim E-K reviewed the images. Park SG and Jeong S contributed to the preprocessing of the images. Jeong S developed the deep learning model under Cho MW and Kong H-J’s supervision. Park SG wrote the first draft of the manuscript. All authors edited the manuscript and agreed to the final version before submission and had access to all the raw datasets. All authors had final responsibility for the decision to submit for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, S.G., Jeong, S., Cho, M. et al. Deep learning model for identification of metabolic bone disease of prematurity using wrist radiographs. Sci Rep 16, 7885 (2026). https://doi.org/10.1038/s41598-026-37116-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37116-7