Abstract
Ultraviolet (UV) radiation is a key environmental factor contributing to photoaging, inflammation, and carcinogenesis. While sunscreen is known to prevent sunburn and lower cancer risk, its effectiveness in limiting UV-induced molecular alterations in human skin is not fully defined. In this study, 32 female volunteers belonging to four Fitzpatrick phototypes underwent repeated moderate UV exposures, with skin samples collected from three distinct treatment sites on each participant, including untreated, UV-exposed without protection, and UV-exposed with prior application of sunscreen. Comprehensive profiling revealed that UV exposure markedly altered gene expression and DNA methylation, impacting pathways involved in DNA repair, immune response, and cell cycle regulation. Mapping these molecular signatures to the Hallmarks of Health revealed broad disturbance of the skin’s core functional attributes. The application of sunscreen effectively prevented most UV-driven disruptions, including the acceleration of epigenetic age, and maintained the stability of hallmark-associated pathways, with minor residual molecular changes remaining. This study demonstrates the importance of regular photoprotection in preventing visible skin damage and maintaining molecular skin health. By deepening our understanding of the mechanisms underlying photoprotection, these findings also indicate a new scope for further advancements in the efficacy of future sunscreen formulations.
Similar content being viewed by others
Introduction
Ultraviolet (UV) radiation is a potent environmental stressor that has a profound effect on the health of human skin, leading to DNA damage, inflammation, oxidative stress, and premature skin aging1,2. Beyond the well-known effects of UV exposure, there is increasing evidence that repeated low-dose and sub-erythemal exposures, which do not result in visible erythema, can also disrupt skin homeostasis and promote accelerated photoaging3,4. Such exposures trigger a cascade of molecular stress responses including the activation of DNA repair mechanisms, the induction of inflammatory signaling and alterations of cellular differentiation and proliferation5. Over time, these processes result in cumulative structural and immunological changes that impair barrier function and contribute to the functional decline of the skin6. Chronic low-level UV exposure has also been associated with immune dysregulation and an increased risk of photocarcinogenesis7,8.
To counteract the detrimental effects of UV radiation, topical sunscreens are routinely used as effective physical or chemical filters, reducing the penetration of solar radiation into the skin9,10. Their protective effects have been demonstrated to extend beyond the prevention of sunburn, involving the reduction of DNA damage, clinical signs of photoaging, and UV-induced immunosuppression3,11. A seminal Australian study demonstrated that regular use of sunscreen with a sun protection factor (SPF) of 15 or higher significantly lowers the incidence of melanoma12 and squamous cell carcinoma13 underscoring the public health value of consistent photoprotection. The aforementioned benefits are achieved through multiple mechanisms, as sunscreen limits the activation of stress pathways, minimizes oxidative damage, and preserves the structural integrity of epidermal cells14. By attenuating inflammation and supporting the natural repair processes, sunscreens help maintain barrier function and immune competence15. Despite theoretical concerns, regular sunscreen use has not been shown to compromise vitamin D status in healthy individuals16.
Nevertheless, sunscreens do not entirely obstruct UV radiation. For instance, SPF 30 permits approximately 3% of UV rays to reach the skin, while SPF 50 reduces this to about 2%. Consequently, some residual molecular damage may still occur, particularly in cases of repeated or prolonged exposure17.
Advancements in omics technologies, particularly transcriptomics and epigenomics, have contributed to a more comprehensive understanding of the skin response to UV radiation at the molecular level. UV exposure has been demonstrated to trigger extensive transcriptional changes in epidermal keratinocytes, activating stress-response pathways involved in DNA repair, apoptosis, inflammation and differentiation18, while also inducing epigenetic modifications such as alterations in DNA methylation (DNAm)19. Despite these insights, most SPF assessments remain focused on the prevention of erythema and UV-induced damage, and there remains a notable lack of studies evaluating the efficacy of sunscreens in mitigating the skin’s molecular response to UV exposure. As a result, the extent to which sunscreen alleviates UV-driven molecular stress and shapes long-term biological outcomes is not fully understood. Integrating transcriptomic and epigenomic analyses in the context of sunscreen use may help clarify the underlying biological mechanisms of photoprotection at the molecular level.
In this study, we investigated the molecular consequences of repetitive UV exposure on human skin in healthy volunteers, simulating sun exposure and evaluating the protective efficacy of a topically applied sunscreen (SPF 30) prior to irradiation. By integrating a genome-wide analysis of gene expression and DNAm profiling, we aimed to characterize how cumulative UV stress alters gene expression and epigenetic pattern in the skin, and to assess the extent to which sunscreen application may mitigate these molecular changes. These findings could contribute to a deeper understanding of the photoprotective effects, providing a foundation for more comprehensive evaluation of sunscreen efficacy and advancing our knowledge of the biological mechanisms underlying skin photoprotection at the molecular level. This might allow for the development of more effective sunscreen formulations.
Results
Distinct molecular signatures of UV stress across phototypes
To identify UV-stress signatures shared across phototypes under repetitive exposure and assess sunscreen protection efficacy, 32 female volunteers (40–65 years) were enrolled and treated defined lower-back areas for three consecutive days with: untreated control, UV (0.9 minimal erythema dose (MED)), or UV with SPF 30 pre-application (Fig. 1a). Participants represented Fitzpatrick skin phototypes I to IV (Fig. 1b). MED was determined individually for each participant and distribution of MED over the phototypes is visualized in Supplementary Figure S1, showing that higher MED correlates with higher phototype. To assess molecular responses, suction blisters were collected from each treatment area, followed by transcriptomic and DNAm profiling.
Study design, volunteer distribution, and PCA according to skin phototype. [a] Schematic representation of the study design: 32 female volunteers received three treatments over three consecutive days: untreated control, UV exposure at 0.9 MED, and UV exposure at 0.9 MED after pretreatment with SPF 30, [b] Participant distribution across Fitzpatrick phototypes I–IV, [c] PCA of normalized gene expression by treatment (control, green; SPF 30 + UV, purple; UV, orange), shown separately by phototype. From left to right: phototypes I&II, III, and IV, illustrating phototype‑dependent clustering and treatment separation.
To summarize global transcriptomic variation across different conditions, we performed principal component analysis (PCA) across all samples (Fig. 1c). Due to low sample numbers in phototype I, phototypes I and II were combined into a single category. This analysis showed a UV‑induced shift away from control across phototypes in unprotected skin, with the most pronounced shift observed in phototype I/II, and III. In contrast, samples of phototype IV remained closer to their control. Notably, the application of sunscreen SPF 30 before exposure minimized the effect of UV-induced shifts, indicating that SPF 30 works as a protective agent against UV-induced molecular alterations. To account for individual differences in UV sensitivity, each participant’s MED was included as a covariate in subsequent analyses, allowing for the identification of universal UV-induced stress response signatures.
UV Radiation induces significant transcriptomic and epigenetic alterations
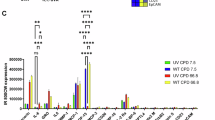
To quantify transcriptomic changes after repeated UV exposure, differential gene expression analysis was conducted, resulting in the identification of differentially expressed genes (DEGs). To interpret biological programs, we ranked DEGs by log2 fold change (log2FC) and performed pathway enrichment, which identified upregulated cell cycle and barrier pathways, and downregulated collagen and immunoregulatory pathways (Fig. 2a). To validate these signals, we analyzed a transcriptomic dataset containing skin samples exposed to acute high UV (2 MED)20, which confirmed enrichment of barrier (formation of the cornified envelope, keratinization) and cell cycle pathways (kinetochore regulation, cell cycle checkpoints), and downregulation of extracellular matrix (ECM) formation (collagen synthesis) and immune pathways.
Transcriptomic and epigenetic dynamics in human skin in response to UV exposure. [a] GSEA based on differential gene expression showing the top eight up‑ and downregulated Reactome pathways ranked by NES. Dot size denotes the number of leading‑edge genes; color encodes Bemjamini-Hochberg (BH) False Discovery Rate (FDR) (significant at FDR < 0.05). Pathways validated by the independent acute UV (2 MED)20 dataset are highlighted in red. [b] Venn diagram showing the overlap of significantly enriched pathways (DNAm–based GSEA) between our repeated UV dataset (0.9 MED) and an independent chronic UV study21. Overlapping segments indicate pathways concordantly enriched in both datasets (full lists in Supplementary Table S1).
To assess epigenetic alterations, we performed differential methylation analysis and identified Differentially methylated probes (DMPs) associated with repeated UV exposure (0.9 MED). To contextualize these findings within the chronic effects of sun exposure, a comparison was made between our results and an independent dataset profiling epidermal DNA methylation in chronically sun exposed (outer arm) and sun protected (inner arm) skin21. This comparison revealed 30 biological pathways that were consistently altered in both datasets (Fig. 2b), including regulation of GTPase activity, collagen synthesis, vascular endothelial growth factor (VEGF) related signaling, and cell–cell communication (Supplementary Table S1). Further pathway enrichment analysis of DEGs and DMPs revealed 22 shared pathways, including those involved in cell–cell communication and collagen biosynthesis. These pathways are detailed in Supplementary Table S2. Collectively, these analyses demonstrate that repeated UV exposure induces coordinated transcriptomic and epigenetic alterations in pathways central to skin structure and function, and that many of these molecular changes are consistently observed in both acute and chronically sun exposed skin, underscoring their relevance to long-term photodamage.
Sunscreen protection attenuates UV-induced molecular disruptions
To assess the physical effects of SPF 30 sunscreen, standardized images were captured across phototypes and treatments after 24 h from the last UV exposure (Fig. 3a). The repeated exposure of unprotected skin areas with 0.9 MED resulted in clear signs of reddening in phototypes I and II, and a combination of reddening and tanning in III and IV. In contrast, the SPF 30 pretreated areas showed no visible skin reaction at the same repeated UV-dose. Furthermore, the quantification of transcriptomic perturbation was achieved through the computation of DEGs, yielding 150 DEGs for the comparison of SPF 30 pretreated group with the control group, resulting in a substantial attenuation compared to 3,618 DEGs in the UV-exposed group (Fig. 3b, Supplementary Fig. S2ab). Similarly, differential DNAm analysis revealed 314 DMPs in the SPF 30 pretreated group compared to the control, representing a strong reduction relative to the 83,594 DMPs identified in the UV-exposed group (Fig. 3c, Supplementary Fig. S2cd). There was no significant correlation between DEGs and DMPs (Supplementary Figure S3).
SPF 30 protects against UV-induced visible reactions and prevents molecular perturbations in human skin. Colors: control (green), sunscreen protected UV (purple), unprotected UV (orange). [a] Representative photographs of each treatment site, taken 24 h after the final irradiation and immediately before suction blister sampling; examples shown for Fitzpatrick phototypes I–IV. [b] Number of DEGs identified for UV exposed vs. control and sunscreen protected UV vs. control comparisons (adjusted p < 0.05, |log2FC|> 0.5). [c] Number of DMPs identified for UV exposed vs. control and sunscreen protected UV vs. control comparisons (adjusted p < 0.05). [d] Heatmap of pathway activity for the 16 pathways most enriched in UV vs control by GSEA. Group-average GSVA scores are displayed. Colors show deviation from control (red, higher; blue, lower), enabling direct comparison of unprotected UV and sunscreen protected UV relative to the same baseline. Pathway regulation in sunscreen protected group did not reach significant level. [e] Boxplot of DNAm GrimAge across groups; pairwise within‑donor differences tested by two‑sided paired Wilcoxon signed‑rank tests (** p < 0.01; *** p < 0.001).
The assessment of potential functional consequences was conducted through the execution of gene set variation analysis (GSVA) on gene expression data for 16 pathways previously identified as strongly regulated upon UV exposure (Fig. 3d). This analysis revealed a substantial regulation of cell-cycle, keratinization, and immune programs in the presence of unprotected UV, whereas these pathways were not significantly regulated following sunscreen pre-application (p > 0.05). The results presented above for GSVA reflect group-level averages. Supplementary Figure S4 provides a sample-level view, further stratified by phototype, suggesting that sunscreen and phototype IV exhibit attenuated differential gene expression signals in some pathways. These results demonstrate that sunscreen application effectively prevents visible skin reactions and markedly reduces both transcriptomic and epigenetic alterations induced by repeated UV exposure.
In order to assess the impact of UV radiation on epigenetic aging, we estimated biological age using GrimAge clock22, which measures age based on DNAm profiles. Within each donor, we compared GrimAge estimates across conditions (Fig. 3e). Unprotected UV exposure was found to significantly accelerate GrimAge progression in comparison to the untreated control group (Wilcoxon signed-rank, p < 0.05). In contrast, SPF 30, administered pre-application before UV exposure, yielded DNAm age estimates that were not different from the control group (Wilcoxon signed-rank, p ≥ 0.05) and were significantly lower than the unprotected UV group (Wilcoxon signed-rank, p < 0.05). These findings suggest that SPF 30 application prevented the acceleration of the epigenetic age induced by UV. Supplementary Figure S5 provides a detailed visualization with paired samples for each participant included in the GrimAge analysis, highlighting the individual-level comparison within the study.
Residual UV effects under SPF 30 reveal potential for next-generation photoprotective strategies
To explore potential avenues for enhancing photoprotection, we examined differential analyses of SPF 30 protected samples versus controls. While residual DEGs and DMPs were still detectable under sunscreen protected UV exposure, they represented only a very small fraction of the overall gene expression and methylation landscape. Building on this, we performed promoter-variance analyses based on DNAm. Unprotected UV exposure led to a substantial increase in promoter DNAm variance (PERMANOVA, p < 0.05), while promoter variance stayed at control-like levels for sunscreen protected samples (PERMANOVA, p ≥ 0.05) (Fig. 4a). Extending this analysis to investigate methylation plasticity in enhancer regions and assess epigenetic regulation beyond promoters, revealed a smaller but significant shift in the sunscreen protected group relative to controls (PERMANOVA, p < 0.05), while unprotected UV exposure produced more pronounced alterations compared to both control and sunscreen protected samples (PERMANOVA, p < 0.05) (Fig. 4b).
SPF 30 pre-application preserves epigenetic stability and maintains Hallmarks of Health pathways following UV exposure, with minor residual molecular changes detectable. Colors: control (green), sunscreen protected UV (purple), unprotected UV (orange). In all panels, * p < 0.05; ** p < 0.01; *** p < 0.001. [a] Boxplot of promoter DNAm variance across groups; differences assessed by PERMANOVA. [b] Boxplot of enhancer DNAm variance across groups; differences assessed by PERMANOVA. [c] Circular plot provides an overview of the Hallmarks of Health pathway activity, which were found to be significantly regulated across treatments (red, upregulated; blue, downregulated; grey, not significant). Boxplots show sample‑level GSVA scores for each hallmark in control, SPF 30 + UV, and UV. Pairwise differences were tested with two‑sided paired Wilcoxon signed‑rank tests. Panel c adapted from the original Hallmarks of Health publication23.
To assess Hallmarks of Health23 perturbations, we applied GSVA to curated hallmark‑specific gene sets across all treatment groups (Fig. 4c). This revealed significant dysregulation with unprotected UV in homeostatic resilience, recycling and turnover, repair and regeneration, containment of perturbations, integration of circuitries, and integrity of barriers (Paired Wilcoxon signed-rank, p < 0.05), while hormetic regulation and rhythmic oscillations were unchanged (Paired Wilcoxon signed-rank, p ≥ 0.05). To evaluate residual effects under protection, we applied the same analysis to sunscreen pre-applied, UV exposed samples (Fig. 4c). As a result, mild yet significant dysregulation was identified, predominantly affecting homeostatic resilience, recycling and turnover, and repair and regeneration, with markedly smaller effects than in unprotected UV. These results indicate that while SPF 30 sunscreen provides strong protection against UV-induced molecular and functional changes, minor residual effects persist, highlighting the potential for further advancement of photoprotective strategies.
Discussion
In this study, three consecutive UV doses (0.9 MED) were delivered to healthy human skin to test whether the pre-application of an SPF 30 sunscreen could mitigate the resulting molecular perturbations. The findings indicate that the most substantial alterations in gene profiles are observed in lighter phototypes (I, II, and III), attenuated in phototypes IV, consistent with melanin mediated photoprotection24,25, although melanin is not the sole determinant of UV sensitivity26,27. To account for inter-individual UV sensitivity, each participant’s MED was included as a covariate. This granted the identification of universal UV-induced stress-response signatures.
The study’s findings revealed that exposure of unprotected skin to repeatedly 0.9 MED resulted in cumulative visible erythema across three days, underscoring the detrimental accumulative effects of UV radiation on the skin’s appearance28,29. The application of SPF 30 sunscreen not only prevented erythema but also markedly reduced the number and magnitude of UV-responsive DEGs14, and, for the first time to our knowledge, DMPs. This novel observation demonstrates that sunscreen can mitigate UV-induced epigenetic changes, extending its protective role beyond previously recognized indicators. Despite this, a minor molecular imprint persisted, indicating that while sunscreen provides substantial protective benefits, minor alterations could accumulate over time due to ongoing UV exposure30,31. This observation underlines that long-term strategies for sun protection could emphasize a regular reapplication of sunscreen, as despite clinically proven immediate protection, a remaining potential for cumulative molecular damage endures32,33,34.
The examination of pathways mapped to the Hallmarks of Health 23, complemented by conventional pathway enrichment of both DEGs and DMPs, revealed coherent effects across the Hallmarks of Health. These effects included barrier integrity, containment of perturbations, repair and regeneration, and recycling and turnover23. The study demonstrated broad cross-study and population-level generalizability. In an independent acute, higher-dose (2 MED) model20, gene-based enrichment was validated to ascertain its robustness across dose and sampling. Furthermore, DNAm-based enrichment was examined against a chronic exposed-versus-protected dataset21 to determine whether pathways perturbed after three days of repeated exposure already bear a long-term molecular imprint. In the context of the barrier integrity hallmark, including keratinization and epidermal differentiation, unprotected skin exhibited the anticipated upregulation, consistent with compensatory reinforcement and restoration. This aligns with prior research associating UV stress with enhanced cornification and stratum corneum lipid processing35,36. This pattern was replicated in the high UV dose dataset20, thereby emphasizing the robustness and generalizability of the findings. With SPF 30 application, these differentiation signals were closer to levels of unexposed control, indicating maintenance of barrier homeostasis under UV stress37,38.
In accordance with the hallmark of containment of perturbations, the collected data demonstrates a significant divergence in immune regulation responses between sunscreen protected and unprotected skin. Specifically, unprotected skin exhibited a significant downregulation of immune signaling pathways, consistent with the phenomenon of photoimmunosuppression and its associated risks, including an increased susceptibility to skin cancer39,40. The validity of this finding is further substantiated by the independent dataset involving a high UV dose20, which reproduced the immune suppression signature observed in unprotected tissues. In stark contrast, sites protected by SPF 30 did not show statistical difference to control sites, suggesting an effective preservation of non–UV exposed baseline immune tone despite ongoing daily UV exposure41. All in all, these findings suggest that sunscreen can attenuate UV-induced immune suppression, which may have implications for maintaining skin health.
In the context of the repair and regeneration hallmark, unprotected, UV exposed skin exhibits checkpoint activation and a temporary cell-cycle arrest in response to DNA damage, facilitating the initiation of cellular repair mechanisms. This finding is supported by additional data focusing on acute UV exposure20, where DEGs linked to DNA damage checkpoints were notably abundant25,42. Complementing transcriptomic analyses, DNAm-based pathway enrichment highlighted dysregulation in signal transduction and cell-cycle regulatory pathways43, reflecting a delicate balance of cellular functions following exposure. This pattern also persisted in the chronic exposed-versus-protected dataset21, indicating that three days of repeated exposure can leave an early epigenetic imprint. On the other hand, the pre-application of an SPF 30-containing product led to a substantial mitigation of these cellular responses, which is consistent with a significantly lower DNA damage burden44. However, the levels did not stay completely at the unexposed control baseline, indicating that there were residual alterations in the repair and regeneration hallmark.
As part of the recycling and turnover hallmark, significant alterations were observed in collagen synthesis pathways, particularly in unprotected UV exposed skin. These alterations reflected notable shifts in ECM composition45,46. The same pattern was reproduced in an independent acute, high UV dose dataset20 and aligned with signatures of chronically sun exposed skin21. Together, these findings indicate an incipient photoaging trajectory47,48. Importantly, the application of sunscreen with an SPF 30 prior to UV exposure appeared to mitigate those alterations. Furthermore, the integration of DNAm-based enrichment analyses pointed to a differentially regulated pathway associated with VEGF, which aligns with microvascular adaptation. These epigenetic signals were similarly detected in the study concerning chronic UV exposure21, thereby reinforcing the evidence of persistent alterations in vascular signaling in response to UV damage49. Despite the substantial protection provided by sunscreen pre-application across recycling and turnover pathways dysregulated by UV exposure, it is still slightly dysregulated compared to the unexposed control. This finding suggests that minor differential responses may persist, thereby emphasizing the complexity of the skin’s adaptive responses to environmental stressors.
At the epigenome-transcriptome interface, UV exposure has been implicated in compromising regulatory integrity and disrupting DNAm patterns at promoter- and enhancer-linked Cytosine-phosphate-Guanine (CpG) sites50,51. SPF 30 successfully mitigated overall DNAm variability relative to untreated skin. Nevertheless, DNAm levels at enhancer-associated sites were not fully preserved, which suggests that the distal regulatory architecture has been partially stabilized. This finding supports the hypothesis of a layered organization of the human epigenome. In this model, promoters and gene bodies form a stable core, and enhancers constitute a more plastic and less protected mantle52. UV primarily perturbs the mantle, while SPF 30 preserves core regulation yet leaves some residual mantle variability. In line with this layer-specific sensitivity, repeated UV exposure has been associated with visible signs of aging, where frequent sunbathing habits correlated with an “older” looking appearance53. Extending these findings, our data show that SPF 30 effectively prevents UV-driven molecular aging. The DNAm age estimates obtained using the GrimAge clock22 in sunscreen protected skin were not significantly different from unexposed controls, whereas unprotected UV exposed skin showed clear acceleration of DNAm age. Collectively, these findings suggest that preserving core epigenomic stability and limiting enhancer perturbations may mitigate the effects of UV-driven biological aging.
Achieving optimal photoprotection necessitates the implementation of three fundamental principles: adequate dosing, uniform coverage, and regular reapplication54,55,56. Our findings demonstrate that SPF 30 provides reliable and substantial protection, although it still allows a small fraction of incident UV (~ 3.3%) to reach the skin. Using sunscreen formulations with higher SPFs can further reduce this residual UV transmission57,58, potentially enhancing molecular level photoprotection. In addition, the integration of validated active ingredients alongside highly protective sunscreen filter systems offers a promising strategy to further strengthen protection capabilities. A compelling example, proving that active ingredients can contribute substantially to photoprotection is the plant extract Licochalcone A, which significantly reduced UV-induced formation of oxidative radicals alongside SPF 50 + formulation59.
In summary, this study simulated repeated sun exposure and tested whether SPF 30 mitigates ensuing molecular perturbations. SPF 30 not only prevented cumulative erythema but also substantially reduced UV-responsive DEGs and DMPs. Integration of Hallmarks of Health23 mapping with conventional pathway enrichment across transcriptomic and epigenetic layers, along with validation of findings in independent acute20 and chronic exposed‑versus‑protected datasets21, demonstrates SPF 30’s capacity to protect barrier differentiation programs, preserve immune function, attenuate DNA‑damage checkpoint activation, and dampen early collagen and VEGF‑linked vascular remodeling. At the epigenome–transcriptome interface, repeated UV increased DNAm variance and accelerated DNAm age, whereas SPF 30 constrained regulatory variance and prevented age acceleration.
Conclusions
This study underscores the pivotal role of sunscreen in preserving genomic integrity and overall skin health under simulated sun exposure. The Hallmarks of Health are employed as a comprehensive framework, demonstrating that SPF 30 not only prevents erythema but also maintains critical molecular pathways, thereby mitigating UV-induced skin aging at both transcriptomic and epigenomic levels. These results confirm sunscreen as the key skin protective measure, while highlighting opportunities for advancing formulations by adding active ingredients, following rigorous application practices to limit cumulative molecular perturbations. Collectively, these elements underpin a comprehensive approach to photoprotection, delivering immediate defense and sustaining long-term skin health and resilience against UV-driven damage.
Methods
Study design and data collection
The study’s participants encompassed 32 Caucasian women with ages ranging from 40 to 65 years and Fitzpatrick phototypes ranging from I to IV. The study was conducted between October and November. All irradiations were performed using a SOL 500 full spectrum solar simulator (Hönle UV Technology). MED was determined individually for all subjects as the amount and time of UV irradiation to show first signs of reddening or tanning, following the protocols described in DIN EN ISO 24,444. This included irradiation of the skin with 6 different UV dosages (light staircase with a 25% increment), following an assessment of erythema or tanning 24 h after irradiation to determine the MED. After MED determination, treatment was performed. For each participant, three sites on the lower back were assigned to: (i) untreated control (shielded, no UV), (ii) unprotected UV exposure (0.9 MED per day for 3 consecutive days), and (iii) sunscreen protected UV exposure (broad‑spectrum SPF 30 applied at 2 mg/cm2 by a technician 15 min before the same 0.9 MED exposure on each day). The sunscreen used in this study was an oil-in-water emulsion designed for broad-spectrum UV protection. It contained a combination of chemical and physical UV filters, including avobenzone (BMDM), octisalate, octocrylene, homosalate, and titanium dioxide. The recruitment, MED determination, irradiation, and sampling procedures were carried out as previously described60. Photographs were taken of each investigated site 24 h after the final irradiation, immediately before suction-blister sampling. Subsequently, two suction blisters were collected from each site to facilitate the isolation of RNA and DNA for subsequent transcriptomic and DNAm analyses.
Data pre-processing
Transcriptome libraries were prepared using the Illumina TruSeq Library Prep Kit and sequenced on the HiSeq system in single-end mode (1 × 50 bp), reaching a depth of about 100 million reads per sample. Data processing for raw transcriptomic data was performed by using a pipeline including quality control using FastQC v0.12.161, Trimmomatic v0.39 for quality based read trimming62, STAR v2.7.11a for read mapping63, and Salmon v1.10.0 for the quantification64. The human reference genome used was GRCh3865 with Ensembl IDs and the final data included transcript expression in the form of read counts and transcripts per million (TPM) for every gene. The raw counts were filtered to include only protein coding genes, which resulted in a total number of 19,653 genes.
Raw DNAm data were generated from epidermis samples using the Infinium MethylationEPIC BeadChip array (Illumina Inc.). Preprocessing of the data was conducted using the R Bioconductor ‘minfi’ package66,67. Firstly, samples and methylation loci (probes) with poor signal quality were identified and excluded based on detection p-values exceeding a predefined threshold of p > 0.01. Specifically, samples with more than 10% of probes failing this threshold were removed, as were probes with more than 10% of samples failing the detection criteria. Probes located on the sex chromosomes (X and Y) were excluded to mitigate confounding factors associated with sex-specific methylation. Furthermore, probes known to be cross-reactive, along with those overlapping with common single nucleotide polymorphisms (SNPs), were filtered out to minimize technical artefacts and allele-specific bias, resulting in 811,734 CpG sites. The remaining data were normalized using the ‘preprocessFunnorm’ function. Following normalization, β-values were extracted for downstream analyses.
PCA
Overview of high dimensional data structure was generated by PCA. Initially, the raw count data was voom normalized, followed by PCA conducted employing the ‘stats’ R package68.
Differential analysis
Differential analyses of both DNAm and gene expression profiles in epidermal samples were conducted. Linear modelling was conducted using preprocessed β-values (for DNAm) and voom normalized expression values (for gene expression), with the ‘limma’ R package69. MED value and donor age were included as covariates to account for potential confounding factors. DMPs and DEGs were identified through comparisons of control vs. 0.9 MED, and control vs. “SPF 30 + 0.9 MED” groups. Statistical significance was determined using BH adjusted p-values, with an FDR threshold set at < 0.05.
Hallmarks of Health pathway assignment
Hierarchical Reactome pathways70 were conceptually assigned to the Hallmarks of Health based on the framework proposed by López-Otín and Kroemer23. Leaf‑level pathways were propagated to their corresponding hallmark, then curated by removing duplicates and excluding genes assigned to more than three hallmarks to enhance specificity. Subsequently, hallmark definitions were refined by integrating Gene Ontology (GO) terms selected as anchor terms71,72. GOsemantic similarity analysis73 was applied to cross-validate genes to a second database, thereby increasing both coverage and biological specificity. Results of pathway assignment are summarized in Supplementary Table S3.
Enrichment analysis
Pathway enrichment analysis was performed on all DEGs with a significant p-adjusted value p < 0.05 using the ‘fgsea’ R package74. All genes were ranked by log2FC as input for the analysis and Reactome pathways testing70 (minimum gene set size 10, maximum 500). Pathway significance was calculated using BH method with an adjusted p-value threshold of < 0.05. To validate UV-driven pathway regulation, we reanalyzed an independent transcriptomic dataset of five female volunteers in which lower-arm skin received a single 2 MED UV exposure with matched unexposed controls and biopsies at 24 hours20, applying the same UV-versus-control preranked enrichment to confirm differentially regulated pathways.
Pathway enrichment analysis of DMPs was performed in the primary cohort using the ‘methylGSA’ R package75. Reactome pathways70 were used as the reference gene sets, and significance was assessed using a BH adjusted p-value threshold of < 0.05. To validate these findings independently, we applied the same analytical pipeline to epidermal samples from the dataset published by Vandiver, et al.21 which included DNAm profiles from 19 paired outer (sun exposed) and inner (sun protected) forearm samples collected from healthy adult donors using the HumanMethylation450 BeadChip array (Illumina Inc.). Differential methylation analysis in the validation cohort included age and gender as covariates, and pathway enrichment was carried out for the 450 k platform. This enabled direct comparison of the biological pathways associated with chronic UV exposure between the primary and validation datasets.
GSVA
GSVA was performed using the R package ‘GSVA’76. The input consisted of normalized gene expression values, and gene sets were defined from a GSEA or Hallmark of Health analysis. GSVA was run using the Gaussian kernel density estimation. Pathway scores were centered to the control group by subtracting the control mean per pathway. Pairwise differences between conditions were assessed using two-sided paired Wilcoxon signed-rank tests, with p < 0.05 considered significant.
Promoter and enhancer variance analysis
We quantified DNAm variance at promoter- and enhancer-associated CpG sites in epidermal samples from 32 volunteers. Promoter CpGs were defined using the Illumina EPIC array manifest77, while enhancer CpGs were annotated based on ChromHMM segmentation of the keratinocyte (NHEK cell line) from the UCSC database78,79. For each regulatory category, we first calculated the variance in DNAm for each sample across the selected CpGs. Group differences in variance were then assessed using permutational multivariate analysis of variance (PERMANOVA) with the ‘vegan’ R package80, by comparing the distributions of variances between the experimental groups. Statistical significance was determined using a p-value threshold of 0.05.
Age clock prediction and statistical testing
The DNAm age was estimated from normalized β-values using the principal component–based GrimAge (PCGrimAge) clock22,81 implemented in the ‘dnaMethyAge’ R package82. An analysis was conducted on subjects with all three conditions present (untreated control, SPF 30 pre-application followed by UV exposure, and unprotected UV exposure). Predicted ages were compared within donors using two-sided paired Wilcoxon signed-rank tests for each pairwise contrast, with p < 0.05 considered statistically significant.
Data availability
Data supporting the findings of this study are openly available in ArrayExpress under accession numbers E-MTAB-9251 and E-MTAB-9249. The release of supplementary data is restricted due to confidential reasons. However, following the establishment of a data-use agreement with the corresponding author, these additional datasets can be shared upon reasonable request.
References
Mohania, D. et al. in Ultraviolet Light in Human Health, Diseases and Environment (ed Shamim I. Ahmad) 71–87 (Springer International Publishing, 2017).
Lastowiecka-Moras, E., Bugajska, J. & Młynarczyk, B. Occupational exposure to natural UV radiation and premature skin ageing. Int. J. Occup. Saf. Ergon. 20, 639–645. https://doi.org/10.1080/10803548.2014.11077079 (2014).
Young, A. R., Orchard, G. E., Harrison, G. I. & Klock, J. L. The detrimental effects of daily sub-erythemal exposure on human skin in vivo can be prevented by a daily-care broad-spectrum sunscreen. J. Invest. Dermatol. 127, 975–978. https://doi.org/10.1038/sj.jid.5700606 (2007).
Keurentjes, A. J. et al. Tape stripping the stratum corneum for biomarkers of ultraviolet radiation exposure at sub-erythemal dosages: a study in human volunteers. Biomarkers 25, 490–497. https://doi.org/10.1080/1354750X.2020.1792551 (2020).
Harrison, G. I., Young, A. R. & McMahon, S. B. Ultraviolet radiation-induced inflammation as a model for cutaneous hyperalgesia. J. Invest. Dermatol. 122, 183–189. https://doi.org/10.1046/j.0022-202X.2003.22119.x (2004).
Alhasaniah, A., Sherratt, M. J. & O’Neill, C. A. The impact of ultraviolet radiation on barrier function in human skin: molecular mechanisms and topical therapeutics. Curr. Med. Chem. 25, 5503–5511. https://doi.org/10.2174/0929867324666171106164916 (2018).
Orazio, J., Jarrett, S., Amaro-Ortiz, A. & Scott, T. UV Radiation and the Skin. Int. J. Mol. Sci. 14, 12222–12248 (2013).
Halliday, G. M., Byrne, S. N., Kuchel, J. M., Poon, T. S. C. & Barnetson, R. S. C. The suppression of immunity by ultraviolet radiation: UVA, nitric oxide and DNA damage. Photochem. Photobiol. Sci. 3, 736–740. https://doi.org/10.1039/b313199h (2004).
Kullavanijaya, P. & Lim, H. W. Photoprotection. J. Am. Acad. Dermatol. 52, 937–958. https://doi.org/10.1016/j.jaad.2004.07.063 (2005).
Seité, S. et al. A broad-spectrum sunscreen prevents cumulative damage from repeated exposure to sub-erythemal solar ultraviolet radiation representative of temperate latitudes. J. Eur. Acad. Dermatol. Venereol. 24, 219–222. https://doi.org/10.1111/j.1468-3083.2009.03332.x (2010).
Keurentjes, A. J. et al. Stratum corneum biomarkers after in vivo repeated exposure to sub-erythemal dosages of ultraviolet radiation in unprotected and sunscreen (SPF 50+) protected skin. Photodermatol, Photoimmunol. Photomed. 38, 60–68. https://doi.org/10.1111/phpp.12717 (2022).
Green, A. C., Williams, G. M., Logan, V. & Strutton, G. M. Reduced melanoma after regular sunscreen use: randomized trial follow-up. J. Clin. Oncol. 29, 257–263. https://doi.org/10.1200/jco.2010.28.7078 (2011).
van der Pols, J. C., Williams, G. M., Pandeya, N., Logan, V. & Green, A. C. Prolonged prevention of squamous cell carcinoma of the skin by regular sunscreen use. Cancer Epidemiol. Biomarkers Prev. 15, 2546–2548. https://doi.org/10.1158/1055-9965.Epi-06-0352 (2006).
Marionnet, C., Pierrard, C., Lejeune, F. & Bernerd, F. Modulations of gene expression induced by daily ultraviolet light can be prevented by a broad spectrum sunscreen. J. Photochem. Photobiol. B: Biol. 116, 37–47. https://doi.org/10.1016/j.jphotobiol.2012.08.001 (2012).
Young, A. R., Claveau, J. & Rossi, A. B. Ultraviolet radiation and the skin: Photobiology and sunscreen photoprotection. J. Am. Acad. Dermatol. 76, S100-s109. https://doi.org/10.1016/j.jaad.2016.09.038 (2017).
Passeron, T. et al. Sunscreen photoprotection and vitamin D status. Br. J. Dermatol. 181, 916–931. https://doi.org/10.1111/bjd.17992 (2019).
Syring, F. et al. Investigation of model sunscreen formulations comparing the sun protection factor, the universal sun protection factor and the radical formation ratio. Skin Pharmacol. Physiol. 29, 18–23. https://doi.org/10.1159/000441037 (2016).
Sun, X., Kim, A., Nakatani, M., Shen, Y. & Liu, L. Distinctive molecular responses to ultraviolet radiation between keratinocytes and melanocytes. Exp. Dermatol. 25, 708–713. https://doi.org/10.1111/exd.13057 (2016).
Köhler, F. & Rodríguez-Paredes, M. DNA methylation in epidermal differentiation, aging, and cancer. J. Invest. Dermatol. 140, 38–47. https://doi.org/10.1016/j.jid.2019.05.011 (2020).
Skopelja-Gardner, S. et al. The early local and systemic Type I interferon responses to ultraviolet B light exposure are cGAS dependent. Sci. Rep. 10, 7908. https://doi.org/10.1038/s41598-020-64865-w (2020).
Vandiver, A. R. et al. Age and sun exposure-related widespread genomic blocks of hypomethylation in nonmalignant skin. Genome. Biol. 16, 80. https://doi.org/10.1186/s13059-015-0644-y (2015).
Lu, A. T. et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (Albany NY) 11, 303–327. https://doi.org/10.18632/aging.101684 (2019).
López-Otín, C. & Kroemer, G. Hallmarks of health. Cell 184, 33–63. https://doi.org/10.1016/j.cell.2020.11.034 (2021).
Brenner, M. & Hearing, V. J. The protective role of melanin against UV damage in human skin. Photochem. Photobiol. 84, 539–549. https://doi.org/10.1111/j.1751-1097.2007.00226.x (2008).
Tadokoro, T. et al. UV-induced DNA damage and melanin content in human skin differing in racial/ethnic origin. Faseb J. 17, 1177–1179. https://doi.org/10.1096/fj.02-0865fje (2003).
Maresca, V., Flori, E. & Picardo, M. Skin phototype: a new perspective. Pigment Cell Melanoma. Res. 28, 378–389. https://doi.org/10.1111/pcmr.12365 (2015).
Gorlov, I. P. et al. Human genes differ by their UV sensitivity estimated through analysis of UV-induced silent mutations in melanoma. Hum. Mutat. 41, 1751–1760. https://doi.org/10.1002/humu.24078 (2020).
Jang, S., Jung, Y., Kang, N. & Kim, E. LB805 a study of the human skin changes according to UV shock. J. Invest. Dermatol. 141, B24. https://doi.org/10.1016/j.jid.2021.07.063 (2021).
Kim, M. A., Jung, Y. C., Suh, B.-F., Lee, H. N. & Kim, E. Skin biophysical properties including impaired skin barrier function determine ultraviolet sensitivity. J. Cosmetic Dermatol. 21, 5066–5072. https://doi.org/10.1111/jocd.14964 (2022).
Laughery, M. F., Wilson, H. E., Sewell, A., Stevison, S. & Wyrick, J. J. The surprising diversity of UV-induced mutations. Adv. Gene. 5, 2300205. https://doi.org/10.1002/ggn2.202300205 (2024).
Ding, S. et al. Chronic sun exposure is associated with histone acetylation changes. Br. J. Dermatol. 179, e61–e61. https://doi.org/10.1111/bjd.16852 (2018).
Pruim, B. & Green, A. Photobiological aspects of sunscreen re-application. Austr. J. Dermatol. 40, 14–18. https://doi.org/10.1046/j.1440-0960.1999.00309.x (1999).
Petersen, B. & Wulf, H. C. Application of sunscreen − theory and reality. Photodermatol., Photoimmunol. Photomed. 30, 96–101. https://doi.org/10.1111/phpp.12099 (2014).
Chouinard, N. et al. Repeated exposures of human skin equivalent to low doses of ultraviolet-B radiation lead to changes in cellular functions and accumulation of cyclobutane pyrimidine dimers. Biochem. Cell Biol. 79, 507–515 (2001).
Schäfer, M. & Werner, S. The Cornified Envelope: A First Line of Defense against Reactive Oxygen Species. J. Invest. Dermatol. 131, 1409–1411. https://doi.org/10.1038/jid.2011.119 (2011).
Biniek, K., Levi, K. & Dauskardt, R. H. Solar UV radiation reduces the barrier function of human skin. Proc. Natl. Acad. Sci. 109, 17111–17116. https://doi.org/10.1073/pnas.1206851109 (2012).
Jacques, C. et al. Multi-omics analysis to evaluate the effects of solar exposure and a broad-spectrum SPF50+ sunscreen on markers of skin barrier function in a skin ecosystem model. Photochem. Photobiol. 101, 373–385. https://doi.org/10.1111/php.14001 (2025).
Berkey, C., Biniek, K. & Dauskardt, R. H. Screening sunscreens: protecting the biomechanical barrier function of skin from solar ultraviolet radiation damage. Int. J. Cosmetic Sci. 39, 269–274. https://doi.org/10.1111/ics.12370 (2017).
Halliday, G. M., Damian, D. L., Rana, S. & Byrne, S. N. The suppressive effects of ultraviolet radiation on immunity in the skin and internal organs: Implications for autoimmunity. J. Dermatol. Sci. 66, 176–182. https://doi.org/10.1016/j.jdermsci.2011.12.009 (2012).
Poon, T. S. C., Barnetson, R. S. C. & Halliday, G. M. Sunlight-induced immunosuppression in humans is initially because of UVB, then UVA, followed by interactive effects. J. Invest. Dermatol. 125, 840–846. https://doi.org/10.1111/j.0022-202X.2005.23894.x (2005).
Narbutt, J. et al. Sunscreen applied at ≥ 2 mg cm−2 during a sunny holiday prevents erythema, a biomarker of ultraviolet radiation-induced DNA damage and suppression of acquired immunity. Br. J. Dermatol. 180, 604–614. https://doi.org/10.1111/bjd.17277 (2019).
Dong, K. K. et al. UV-induced DNA damage initiates release of MMP-1 in human skin. Exp. Dermatol. 17, 1037–1044. https://doi.org/10.1111/j.1600-0625.2008.00747.x (2008).
Liu, J., Liu, L., He, J., Xu, Y. & Wang, Y. Multi-omic analysis of altered transcriptome and epigenetic signatures in the UV-induced DNA damage response. DNA Repair 106, 103172. https://doi.org/10.1016/j.dnarep.2021.103172 (2021).
Young, A. R. et al. Sub-optimal application of a high SPF sunscreen prevents epidermal DNA damage in Vivo. Acta Derm. Venereol. 98, 880–887. https://doi.org/10.2340/00015555-2992 (2018).
Biskanaki, F. et al. Impact of solar ultraviolet radiation in the expression of Type I collagen in the dermis. Cosmetics https://doi.org/10.3390/cosmetics8020046 (2021).
Sherratt, M. J. et al. Low-dose ultraviolet radiation selectively degrades chromophore-rich extracellular matrix components. The J. Pathol. 222, 32–40. https://doi.org/10.1002/path.2730 (2010).
McCabe, M. C. et al. Alterations in extracellular matrix composition during aging and photoaging of the skin. Matrix Biol. Plus 8, 100041. https://doi.org/10.1016/j.mbplus.2020.100041 (2020).
El-Domyati, M. et al. Intrinsic aging vs. photoaging: a comparative histopathological, immunohistochemical, and ultrastructural study of skin. Exp. Dermatol. 11, 398–405. https://doi.org/10.1034/j.1600-0625.2002.110502.x (2002).
Hartono, S. P. et al. Vascular Endothelial Growth Factor as an Immediate-Early Activator of Ultraviolet-Induced Skin Injury. Mayo Clin. Proceed. 97, 154–164. https://doi.org/10.1016/j.mayocp.2021.08.018 (2022).
Ziller, M. J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481. https://doi.org/10.1038/nature12433 (2013).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495. https://doi.org/10.1038/nature10716 (2011).
Grönniger, E., Max, H. & Lyko, F. Skin Rejuvenation by Modulation of DNA Methylation. Exp Dermatol. 33, e70005. https://doi.org/10.1111/exd.70005 (2024).
Holzscheck, N. et al. Multi-omics network analysis reveals distinct stages in the human aging progression in epidermal tissue. Aging (Albany NY) 12, 12393–12409. https://doi.org/10.18632/aging.103499 (2020).
Schalka, S., dos Reis, V. M. & Cucé, L. C. The influence of the amount of sunscreen applied and its sun protection factor (SPF): evaluation of two sunscreens including the same ingredients at different concentrations. Photodermatol. Photoimmunol. Photomed. 25, 175–180. https://doi.org/10.1111/j.1600-0781.2009.00408.x (2009).
Lourenco, E. A. J. et al. Application of SPF moisturisers is inferior to sunscreens in coverage of facial and eyelid regions. PLoS ONE 14, e0212548. https://doi.org/10.1371/journal.pone.0212548 (2019).
De Villa, D., Nagatomi, A. R., Paese, K., Guterres, S. & Cestari, T. F. Reapplication improves the amount of sunscreen, not its regularity, under real life conditions. Photochem. Photobiol. 87, 457–460. https://doi.org/10.1111/j.1751-1097.2010.00856.x (2011).
van Bodegraven, M. et al. Redefine photoprotection: Sun protection beyond sunburn. Exp. Dermatol. 33, 15002. https://doi.org/10.1111/exd.15002 (2024).
Kohli, I. et al. Greater efficacy of SPF 100+ sunscreen compared with SPF 50+ in sunburn prevention during 5 consecutive days of sunlight exposure: A randomized, double-blind clinical trial. J. Am. Acad. Dermatol. 82, 869–877. https://doi.org/10.1016/j.jaad.2019.09.018 (2020).
Mann, T. et al. High-energy visible light at ambient doses and intensities induces oxidative stress of skin—Protective effects of the antioxidant and Nrf2 inducer Licochalcone A in vitro and in vivo. Photodermatology, Photoimmunology & Photomedicine 36, 135–144 (2020). https://doi.org/10.1111/phpp.12523
Holzscheck, N. et al. Concomitant DNA methylation and transcriptome signatures define epidermal responses to acute solar UV radiation. Sci. Rep. 10, 12918. https://doi.org/10.1038/s41598-020-69683-8 (2020).
Andrews, S. FastQC: A quality control tool for high throughput sequence data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Dobin, A. & Gingeras, T. R. Mapping RNA-seq Reads with STAR. Curr Protoc Bioinform. https://doi.org/10.1002/0471250953.bi1114s51 (2015).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14, 417–419. https://doi.org/10.1038/nmeth.4197 (2017).
Church, D. et al. Extending reference assembly models. Genome Biology, In Press (2015). https://doi.org/10.1186/s13059-015-0587-3
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369. https://doi.org/10.1093/bioinformatics/btu049 (2014).
Fortin, J. P. et al. Functional normalization of 450k methylation array data improves replication in large cancer studies. Genome Biol. 15, 503. https://doi.org/10.1186/s13059-014-0503-2 (2014).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing (2024).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47. https://doi.org/10.1093/nar/gkv007 (2015).
Ligtenberg, W. reactome.db: A set of annotation maps for Reactome (R package version 1.86.2). Bioconductor. https://doi.org/10.18129/B9.bioc.reactome.db (2023).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat. Genet. 25, 25–29. https://doi.org/10.1038/75556 (2000).
Aleksander, S. A. et al. The Gene Ontology knowledgebase in 2023. Genetics https://doi.org/10.1093/genetics/iyad031 (2023).
Yu, G. Gene ontology semantic similarity analysis using GOSemSim. Methods Mol .Biol. 2117, 207–215. https://doi.org/10.1007/978-1-0716-0301-7_11 (2020).
Korotkevich, G. et al. Fast gene set enrichment analysis. bioRxiv, 060012 (2021). https://doi.org/10.1101/060012
Ren, X. & Kuan, P. F. methylGSA: a Bioconductor package and Shiny app for DNA methylation data length bias adjustment in gene set testing. Bioinformatics 35, 1958–1959. https://doi.org/10.1093/bioinformatics/bty892 (2019).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinform. 14, 7. https://doi.org/10.1186/1471-2105-14-7 (2013).
IlluminaHumanMethylationEPICanno.ilm10b4.hg19: An annotation package for Illumina’s EPIC methylation arrays (R package version 0.6.0). Bioconductor. https://doi.org/10.18129/B9.bioc.IlluminaHumanMethylationEPICanno.ilm10b4.hg19 (2017).
Ernst, J. & Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol. 28, 817–825. https://doi.org/10.1038/nbt.1662 (2010).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49. https://doi.org/10.1038/nature09906 (2011).
Oksanen, J. et al. vegan: Community Ecology Package (R package version 2.7-2). https://CRAN.R-project.org/package=vegan (2025).
Higgins-Chen, A. T. et al. A computational solution for bolstering reliability of epigenetic clocks: Implications for clinical trials and longitudinal tracking. Nat Aging 2, 644–661. https://doi.org/10.1038/s43587-022-00248-2 (2022).
Wang, Y., Grant, O. A., Zhai, X., McDonald-Maier, K. D. & Schalkwyk, L. C. Insights into ageing rates comparison across tissues from recalibrating cerebellum DNA methylation clock. Geroscience 46, 39–56. https://doi.org/10.1007/s11357-023-00871-w (2024).
Acknowledgements
We would like to express our sincere gratitude to Cassandra Falckenhayn, Joern Soehle, Elke Groenniger, Hermann Jan Huette, Emma Luebchow, Dominik Goeddertz, and Ute Breitenbach for their generous support of this project. Their engagement and feedback substantially strengthened the study’s findings.
Funding
This work was funded by Beiersdorf AG.
Author information
Authors and Affiliations
Contributions
AB and JB contributed equally to this work. The curation of the data was conducted by AB and JB. The bioinformatics analysis and visualization were performed by AB, JB, and LE. The administration of the project was overseen by LK, MW, and SG. The initial draft of the manuscript was prepared collaboratively by JB and AB, who both contributed equally to conceptualization and writing. All authors participated in the revision process and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval and consent to participate
The study was conducted in accordance with the ethical principles of the Declaration of Helsinki, and all participants provided written informed consent prior to enrollment. The protocol was approved by the Ethics Committee of the University of Freiburg (study code 016/1672).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bienkowska, A., Boedewadt, J., Elsbroek, L. et al. Sunscreen application substantially mitigates molecular perturbations induced by repetitive UV exposure and maintains healthy skin. Sci Rep 16, 4326 (2026). https://doi.org/10.1038/s41598-026-37232-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37232-4