Abstract
We developed a CancerMaster-targeted next generation sequencing (NGS) panel covering 524 key genes specifically designed for the multidimensional molecular profiling of solid tumors. We aimed to establish CancerMaster as an asynchronous and parallel one-stop automated analysis pipeline with a reporting system to overcome the delay in existing NGS approaches. This panel can detect common genomic alteration types, including single nucleotide variants/indels and copy number variants (CNVs), fusions, Epstein-Barr virus/human papillomavirus infection, microsatellite instability (MSI), tumor mutational burden (TMB) status, and human leukocyte antigen typing. Using reference materials, CancerMaster demonstrated 100% reproducibility, 99% analytical sensitivity, and high accuracy (94%). Analytical performance for other biomarker classes—including CNVs, fusions, MSI, TMB, and viral detection—was evaluated separately. In a cohort of 668 patients, we identified actionable mutations, including in TP53, KRAS, and PIK3CA, and CNVs, e.g., ERBB2 amplification in gastric (n = 412) and colorectal (n = 66) cancers. MSI and TMB were strongly correlated in all patients (n = 668, r = 0.75; p < 10− 15), with consistent results in the gastric (n = 412, r = 0.75; p < 10− 15) and colorectal (n = 66, r = 0.87; p < 10− 15) cancer groups. Furthermore, directly comparing with the TruSight Oncology 500 (TSO500) panel, CancerMaster demonstrated high concordance while uniquely identifying certain clinically relevant alterations, including an ERBB2 missense mutation. Hence, the CancerMaster panel demonstrated high analytical performance and strong clinical potential for supporting clinical decisions regarding personalized cancer treatment.
Similar content being viewed by others
Introduction
The identification of active genetic alterations is crucial for predicting cancer prognosis and making treatment decisions, especially for patients with treatment-refractory solid tumors. These genetic alterations provide potential therapeutic guidance for selecting cancer-specific or pan-cancer-targeted immune checkpoint inhibitor (ICI) therapies. KRAS, NRAS, and BRAF mutations are crucial for predicting therapeutic responses in metastatic colorectal carcinoma1. ICIs are used for treating multiple cancers based on programmed death-ligand 1 expression, microsatellite instability (MSI) or deficiency in mismatch repair (dMMR) genes, and tumor mutation burden (TMB)2,3. Next-generation sequencing (NGS) technologies, such as whole-genome sequencing (WGS) and whole-exome sequencing (WES), are highly sensitive and accurate for genetic variation assessment, promoting precision oncology based on tumor molecular signatures. Although WGS and WES offer comprehensive, sensitive, and accurate genetic variation assessments, they have several problems, such as higher costs, more complex and time-consuming data processing, and lower sequencing depth than targeted sequencing. A lower sequencing depth may lead to reduced resolution and lower ability to detect rare or low-frequency variants. Consequently, targeted NGS panels, such as FoundationOne and MSK-IMPACT, are widely adopted in clinics owing to their cost-effectiveness, operational feasibility, and accuracy4,5. Nonetheless, these panels have notable limitations, including restricted biomarker coverage, lack of integrated viral detection, slower turnaround times, absence of automated reporting, and challenges in analyzing tumor-only samples4,5,6,7.
There are variations in the design and implementation of targeted NGS panels among clinical laboratories owing to differences in target gene selection, reagent kit sources, sequencing depth and coverage, and data analysis software. Developing custom NGS panels requires high-level operational and bioinformatics infrastructure, which can be challenging8,9,10. Despite these obstacles, research institutions prefer to develop in-house NGS panels to add newly discovered actionable targets and to achieve scalability for rapidly changing clinical needs and profound genomic research. By developing in-house NGS panels, institutions can effectively address their clinical needs by reducing turnaround time, adding new targets, and updating analysis systems for new clinical trials. Moreover, they can achieve higher-level research goals, gain greater panel control and flexibility, secure commercial advantages and protect intellectual property, and ultimately justify their investments in infrastructure and resources.
Herein, we present “CancerMaster,” an in-house hybrid capture-based NGS panel developed for the rapid and comprehensive genomic profiling of diverse tumors (Fig. 1). Unlike widely used panels, such as FoundationOne, MSK-IMPACT, and TSO500, which are limited by restricted coverage, lack of viral detection, and face challenges with tumor-only analysis, CancerMaster supports the detection of copy number variants (CNVs), fusions, Epstein-Barr virus (EBV)/human papillomavirus (HPV), MSI, TMB, and human leukocyte antigen (HLA) typing. This study aimed to develop and validate CancerMaster, an asynchronous and parallel one-stop automated NGS analysis pipeline and reporting system capable of detecting diverse clinically relevant genomic alterations, including single nucleotide variants (SNVs)/indels, CNVs, gene fusions, viral infections (EBV/HPV), MSI, TMB, and HLA typing, to address delays and the limitations of existing approaches.
Work-flow of the main NGS analysis steps. This figure illustrates the workflow of the main NGS steps used in the CancerMaster panel. To achieve efficient data analysis, the pipeline analyzes input samples in parallel at each step and performs asynchronous and parallel analyses on specific modules that can be analyzed independently, such as virus modules. After the final analysis of all the modules, the results were integrated into a JSON file and submitted to the reporting system to generate a basket/academic report. NGS, next generation sequencing.
Results
Sequencing performance and target region coverage
We analyzed 743 samples, including 75 cell lines that met the quality control (QC) criteria. The samples were sequenced using a CancerMaster panel designed for comprehensive genomic profiling in cancer research. The performance metrics across these samples are summarized in Supplementary Table S1. On average, the panel generated 71 million reads per sample, with the read length distribution peaking at 99 bp. More than 98% of the samples achieved at least 50 million total reads, and 65–94% of these reads were unique (i.e., non-duplicate). The mapping rate ranged from 90.4% to 99.7% (mean: 97.7%), indicating high alignment efficiency to the reference genome. For target coverage, the mean sequencing depth ranged from 801.6× to 1981.5×, with an average of 1203.7×, and 96% of targeted regions exceeded 100× coverage (Supplementary Fig. S1). The panel’s uniformity, defined as the proportion of targeted bases with a depth of 0.2× or more of the mean, ranged from 80.2% to 98.5% (mean: 88.7%), consistent with values reported for other hybrid-capture panels (80–90%). This high uniformity highlights the panel’s ability to deliver consistent results across target regions. Furthermore, less than 1% of the targeted regions were not covered, with a mean of only 0.2%, demonstrating the robustness of CancerMaster in capturing the intended genomic regions.
Analytical validation and comparison with a commercial panel
To evaluate the analytical performance of CancerMaster, we validated it using standardized reference materials (NA12878 and OncoSpan HD827) and compared it with a commercial panel. For NA12878, which was assayed 17 times, CancerMaster consistently detected all the known hotspot mutations within the expected variant allele frequency (VAF) range, demonstrating high reproducibility (Fig. 2a). Three replicate assays were performed using OncoSpan HD827, which contains 386 characterized variants across 152 cancer genes. Among the 187 variants located within the CancerMaster target region, 185 were correctly identified, yielding an overall sensitivity of 99% and a precision of 100% based on pre-defined criteria for true positives and coverage depth thresholds (Supplementary Table S2). Two variants were not detected: one owing to insufficient coverage and one large indel beyond the scope of the current variant calling pipeline. To evaluate the agreement between expected and observed allele frequencies, a Bland–Altman analysis was performed. The mean bias between the two measurements was 2.13%, indicating a slight overall overestimation in observed frequencies. The 95% limits of agreement ranged from − 10.5% to + 14.8%, suggesting that the majority of mutations fell within this acceptable error margin. Notably, FLT3 and FANCD2 gene mutations showed relatively lower observed frequencies than expected, whereas NOTCH2, FGF3, and DDR2 gene mutations demonstrated higher observed values. Despite these individual variations, no apparent systematic trend was observed across the frequency range, supporting the analytical accuracy and reproducibility of the assay for detecting allele frequencies across a broad dynamic range (Fig. 2b).
Assessment of reproducibility and analytical sensitivity. This figure shows an assessment of the reproducibility and analytical sensitivity of the CancerMaster panel. (a) The control sample (NA12878) was assayed 17 times to compare the VAF between each experiment. The range of variant frequencies for 32 hotspot variants detected in gold standard samples from the National Institute of Standards and Technologies is shown. (b) Bland-Altman plot comparing expected vs. observed allele frequencies across 187 COSMIC mutations. The x-axis represents the mean allele frequency of expected and observed values, while the y-axis indicates the difference between observed and expected frequencies (Expected − Observed, %). The dashed line represents the mean bias (− 10.5% to + 14.8%).
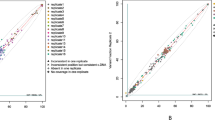
To benchmark against a commercial assay, 23 tumor samples were analyzed in parallel using CancerMaster and TruSight Oncology 500 (TSO500). Most alterations were concordant across both panels (Fig. 3, Supplementary Table S3). Six alterations were discordant: CancerMaster uniquely identified a clinically relevant ERBB2 missense mutation not reported by TSO500, while TSO500 reported one gene fusion and four CNVs not reported by CancerMaster. The gene fusion event was supported by four span reads below the reporting cutoff defined for CancerMaster, and one of the four CNVs (EGFR amplification) underwent orthogonal validation and was negative, consistent with the results of CancerMaster. The agreement between the two panels was defined as concordant variant calls within the same genomic coordinates and comparable VAF ranges (within ~ 10% difference). Together, these results demonstrate that CancerMaster achieves high analytical sensitivity and reproducibility across reference materials and provides results concordant with a widely used commercial panel, while also identifying unique variants of potential clinical significance.
Comparison of CancerMaster and the commercial panel TSO500 using 23 cancer tissue samples. This figure compares the CancerMaster panel with the commercial panel TSO500 using 23 cancer tissue samples. One missense mutation in ERBB2 was reported in CancerMaster but not in TSO500, and one fusion and four CNVs were only reported in TSO500. CNV, copy number variant.
Analytical and clinical validation
Of the 743 cancer tissue samples collected, 288 (including 32 cell lines) were selected for comparison with clinical data. These samples were chosen because they had more than 20% tumor cellularity and known clinical information, such as immunohistochemistry or polymerase chain reaction for genetic alterations (amplification status of ERBB2, MET, EGFR, and FGFR2; EBV/HPV; MSI status). In this subset, the overall accuracy, sensitivity, and specificity of the CancerMaster panel were 94.1%, 76.5%, and 98.0%, respectively.
We compared the CNV detection results between CancerMaster and the clinical data using 193 patient samples (Table 1). The accuracy of FGFR2 was 100%, whereas the specificity and sensitivity of MET were 100% and 94.1%, respectively. For ERBB2, 143 samples (40 positive and 103 negative) showed an accuracy of 91.6%, sensitivity of 70.0%, and specificity of 100%, with 12 false-negative samples. EGFR amplification was compared in 96 samples (11 positive and 85 negative), with an observed accuracy of 92.7%, sensitivity of 45.5%, and specificity of 98.8%, with one false-positive and six false-negative samples. The sensitivity of ERBB2 and EGFR genetic alterations was relatively low in the patient tumor samples, likely because of the high intra-tumoral heterogeneity of gastric cancer. In contrast, all the cell lines (Supplementary Table S4) demonstrated 100% sensitivity. Although the number of cell line samples was limited, these findings suggest that the developed panel reliably detected CNVs in 100% of the samples with high tumor purity. However, detecting such genetic alterations in patients with real-world cancers with lower tumor purity may be more challenging, particularly when only tumor samples are available.
In a cohort of 110 samples assessed for EBV infection, the CancerMaster panel demonstrated an accuracy of 96.4%, sensitivity of 87.5%, and specificity of 97.1% (Table 1). Three false-positives and one false-negative case were identified. The false-negative sample detected a small number of EBV sequences which did not meet the pre-defined threshold for sample contamination; hence, it was categorized as EBV-negative. After excluding the possibility of contamination in the three false-positive samples, we suspected that the false-negative EBER-ISH results may have been caused by RNA degradation. Our observations using the Integrative Genomics Viewer (IGV) (Supplementary Fig. S2), which revealed a significant number of EBV sequences, led us to consider these samples as genuine EBV-infected samples8.
Nine samples were evaluated for HPV infection, including seven clinically positive and two clinically negative cases. Two clinically positive cases were not detected by the CancerMaster panel, resulting in a sensitivity of 71.4% and specificity of 100%. No HPV-aligned reads were observed in the two discordant samples using IGV.
After excluding the initially validated training data (29 MSI-H and 88 MSS cases), we evaluated the MSI detection performance of the CancerMaster panel using an independent cohort of 125 subsequently collected samples consisting of 11 MSI-H and 114 MSS cases. The analysis yielded an accuracy of 95.2%, sensitivity of 90.9%, and specificity of 95.6%. CancerMaster successfully identified 10 of 11 clinically positive samples as positive. One MSI-H sample was misclassified as MSS although it was close to the pre-defined threshold. Among the clinically negative samples, CancerMaster falsely identified five as positive samples. These findings suggest that CancerMaster demonstrates high accuracy and sensitivity in detecting MSI. Nonetheless, caution should be exercised when interpreting the results for samples near the decision threshold. Furthermore, discrepancies between the results may be attributed to differences in the MSI markers used for NGS and clinical MSI testing. Continued research and refinement of CancerMaster algorithms may help improve MSI detection and classification.
Molecular alteration landscape in the patient cohort
The CancerMaster panel was used to identify actionable alterations in the collected patient samples, which were predominantly gastric cancer samples, followed by colorectal and other cancer samples with fewer than 50 cases each (Supplementary Fig. S3 and Supplementary Table S5). The distribution of alteration types, high-frequency actionable gene alterations of SNVs/indels, and CNVs in each cancer type are shown in Supplementary Figs. S4–S6. In colorectal cancer, there was a low frequency of CNVs and a high proportion of clinically actionable mutations, particularly in TP53, KRAS, PIK3CA, and APC mutations, which is consistent with previous studies8,9,10. Among the major cancer types in our cohort, TP53, KRAS, and PIK3CA mutations, and ERBB2 amplification were the most frequent (Supplementary Figs. S6).
MSI and TMB status
Given the similar incidence of MSI-High (MSI-H) and TMB-High (TMB-H) across cancer types, we compared the CancerMaster-derived MSI status and TMB scores with clinical data (Fig. 3). Of the 668 samples (r = 0.75; p < \(\:{10}^{-15}\)), TMB-H (n = 45) and MSI-H (n = 60) were analyzed, and 42 (93.3%) of the TMB-H samples were MSI-H. Of the 412 stomach cancer samples (r = 0.75; p < \(\:{10}^{-15}\)), TMB-H (n = 21, 5.1%) and MSI-H (n = 35, 8.5%) were analyzed; 20 of the TMB-H samples were MSI-H. Similarly, of the 67 colorectal cancer samples (r = 0.87; p < \(\:{10}^{-15}\)), TMB-H (n = 16) and MSI-H (n = 18) were analyzed, and all the TMB-H (100%) samples were MSI-H (Supplementary Fig. S7).
Across all cancer types, TMB-H cases were more likely to be MSI-H but not necessarily vice versa. Notably, colorectal cancer showed particularly high concordance between TMB-H and MSI-H. A comparison of TMB-H with clinical MSI results yielded similar patterns, likely reflecting the fact that MSI assesses local variation in short tandem repeats, whereas TMB-H estimates global genetic variation and encompasses MSI (Fig. 4).
Distribution of MSI and TMB. (a) This figure presents the distribution of MSI and TMB in various cancer patient samples analyzed with the CancerMaster panel. (b) Venn diagrams of NGS MSI-H and TMB-H for patients with all cancers, gastric cancer, and colorectal cancer. MSI, microsatellite instability; TMB, and tumor mutational burden; NGS, next generation sequencing; H, high.
Genetic alteration landscape in gastric cancer
CancerMaster identified several cancer-specific alterations in the gastric cancer cohort (n = 412). Frequent copy number alterations included ERBB2 (11.7%), EGFR (3.2%), FGFR2 (2.7%), and MET (1.2%). ICI–related biomarkers were also identified, including MSI-H (8.5%), TMB-H (5.1%), and EBV positivity (5.1%) (Fig. 5).
Actionable variations in the landscape of gastric cancer. This figure illustrates the actionable variations in the landscape of gastric cancer identified using the CancerMaster panel.
When compared with public datasets, including The Cancer Genome Atlas (TCGA), Memorial Sloan Kettering - Metastatic Events and Tropisms, and China Pan-Cancer (cBioPortal), CancerMaster showed broadly consistent detection of major alterations, although mutation frequencies in TCGA were generally higher, likely reflecting methodological differences (whole-exome versus deep-targeted sequencing) (Supplementary Fig. S8). These cross-cohort comparisons highlight the robustness of CancerMaster, while providing a refined view of genetic alterations specific to gastric cancer.
Discussion
In this study, we developed and validated CancerMaster, a hybrid capture-based NGS panel comprising 524 cancer-associated genes and an automated one-stop analysis pipeline. CancerMaster demonstrated high reproducibility and analytical sensitivity using reference standards and showed strong concordance with a widely used commercial panel. Clinical validation confirmed the high specificity and accuracy of CNVs, EBV/HPV detection, and MSI assessment. Notably, the panel identified actionable alterations across multiple tumor types, including frequent TP53, KRAS, and PIK3CA mutations, and ERBB2 amplification in gastric cancer11,12,13,14. The MSI status and TMB scores were highly correlated, particularly in colorectal cancer. These findings highlight the clinical utility of a comprehensive tumor-only sequencing strategy, which is critical given the unavoidable presence of non-cancerous components in cancer tissues15.
In the absence of paired normal samples, comprehensive tumor-only sequencing has become a practical and essential strategy to guide individualized treatment decisions for patients with advanced cancer. Tumors harboring actionable alterations can be matched to Food and Drug Administration (FDA)-approved targeted therapies or included in clinical trials evaluating novel agents. As the number of gene variants, molecular signatures, and pan-cancer biomarkers with approved therapies continues to expand, genomic profiling of tumor tissues has become widely recognized as a cornerstone for selecting targeted ICI therapies16,17,18. Nevertheless, most FDA-approved companions or complementary diagnostics are designed for univariate analysis, focusing on a single biomarker or drug rather than providing a comprehensive molecular profile19. In contrast, current clinical practice guidelines, such as those of the NCCN, recommend a broad panel-based approach that integrates multiple biomarker assessments within a single assay to preserve limited tumor tissue and maximize clinical utility1.
For clinically useful and reliable NGS using only tumor tissues, we designed and validated a CancerMaster panel. This hybrid capture-based assay, which covers 524 cancer-associated genes and is equipped with an asynchronous one-stop automated pipeline, was developed to provide rapid and comprehensive genomic profiling to support timely clinical decision-making. CancerMaster integrates the detection of diverse alteration types, including SNVs/indels, CNVs, gene fusions, EBV/HPV infection, MSI, TMB, and HLA typing within a single platform. Validation with reference standards confirmed its reproducibility and analytical sensitivity, and comparison with a widely used commercial panel demonstrated overall concordance, reinforcing its reliability for clinical application. Although minor discrepancies were observed, they were consistent with known variability across sequencing platforms and analysis pipelines. Notably, in the OncoSpan (HD827) reference material, the automated CancerMaster workflow did not detect a low-VAF (~ 2%) EGFR exon 19 deletion, a large indel exceeding 10 bp. This event, while clinically important in non-small cell lung cancer, falls outside the primary clinical scope of CancerMaster, which was developed for solid tumors other than lung cancer. Manual inspection confirmed that the underlying sequencing data captured the indel signal, indicating that the limitation lies in the current automated caller rather than in sequencing performance. We recognize this as a methodological constraint of the present version and plan to incorporate improved algorithms for large indel detection in future updates. Taken together, these findings suggest that CancerMaster is a practical and accurate tool for detecting clinically relevant alterations and can be flexibly adapted to meet the evolving clinical needs in precision oncology20.
In our head-to-head comparison with the TSO500 assay, CancerMaster demonstrated high overall concordance across SNVs, indels, fusions, and CNVs, with only a small number of discordant findings. Most discrepancies were attributable to differences in reporting thresholds or assay-specific design features, such as the lower spanning-read support for the TSO500-only fusion event and the orthogonally negative EGFR amplification call. These observations are consistent with expected variability between targeted sequencing platforms and underscore how algorithmic parameters and target-region structure affect CNV detection. Among the CNV discrepancies, MYC amplification detected by TSO500 was not reported by the automated CancerMaster workflow. Because MYC contains only three targeted regions in our panel, and the current CancerMaster CNV algorithm requires more than three consecutive altered regions to call a gain, amplifications in small genes may be underreported. Manual inspection confirmed elevated signal across all MYC target regions, indicating that the underlying sequencing performance was adequate. Future updates to the CancerMaster algorithm will incorporate modified criteria to improve CNV detection in compact genes with limited probe coverage.
We further validated CancerMaster against clinical data for CNV, EBV/HPV, and MSI status. While the panel showed strong overall performance, ERBB2 and EGFR amplifications were more difficult to detect in real-world gastric cancer samples, reflecting the impact of tumor heterogeneity and variable tumor purity in tumor-only sequencing21. This challenge is consistent with the known limitations of CNV detection in heterogeneous solid tumors. For SNV and TMB interpretation, we referenced the ACMG variant classification guidelines and integrated population-level germline resources such as gnomAD and other curated germline databases. Although tumor-only sequencing cannot completely eliminate the risk of germline–somatic misclassification, applying these filters substantially reduces false-positive somatic calls and increases the reliability of downstream biomarker assessment. For CNV analysis, we constructed gene-specific normalization baselines by identifying presumed diploid regions across a large cohort of clinical tumor samples. This reference framework enhances the discrimination of true copy-number alterations from technical noise and compensates for variability in tumor purity. Together, these approaches help overcome key challenges associated with tumor-only assays and support robust multi-layer biomarker profiling in diverse clinical specimens. In contrast, the panel reliably detected EBV/HPV and MSI, supporting its clinical applicability in identifying patients who may benefit from ICI therapy. These findings emphasize both the potential and current technical boundaries of tumor-only sequencing approaches and highlight areas for further refinement to maximize their clinical utility.
In our patient profiling study, CancerMaster captured clinically actionable alterations across multiple cancer types, which is consistent with previously reported mutational landscapes. For gastric cancer, the frequent detection of ERBB2 amplification underscores its relevance as a therapeutic target for HER2-directed therapies, such as trastuzumab. In addition to targeted therapy, the ability of the panel to assess MSI and TMB provides important insights for immunotherapy. We observed a strong correlation and frequent co-occurrence of MSI-H and TMB-H, particularly in colorectal cancer, which is in line with previous studies21,22,23. This reflects overlapping yet distinct biological mechanisms: TMB represents the global mutational burden, whereas MSI indicates defects in mismatch repair at tandem repeats. As a result, most TMB-H tumors also exhibit TMB-H, although the relationship is not absolute; some TMB-H tumors may arise independently of MSI, and conversely, a subset of MSI-H tumors may present with lower TMB. These findings highlight that MSI and TMB, while often co-occurring, provide complementary information and should be evaluated independently in clinical practice to optimize patient selection for ICIs.
These findings highlight the clinical value of comprehensive panel testing in informing biomarker-driven therapy but also underscore the limitations inherent to molecular diagnostics. False positives and negatives may arise because of sample quality, assay performance, or tumor heterogeneity, requiring careful interpretation in the context of clinical and pathological findings. Furthermore, emerging evidence suggests that the host immune background contributes to treatment response; for example, HLA class I supertypes have been linked to differential outcomes, with HLA-B44 associated with extended survival and HLA-B62 or loss of heterozygosity at HLA-I linked to poor prognosis24. Because the CancerMaster panel includes HLA-related genomic regions, these findings suggest that HLA typing and immunogenomic stratification may become feasible using CancerMaster in future studies as additional validation and analytical development are completed. Collectively, these insights demonstrate how integrated genomic and immunogenomic profiling can refine patient stratification for targeted and immune-based therapies.
In conclusion, CancerMaster represents a valuable tool for the comprehensive genomic profiling of solid tumors. Our study demonstrates its accuracy and sensitivity in detecting diverse genetic alterations, CNVs, EBV/HPV infections, and MSI status, thus highlighting its clinical utility. Beyond analytical performance, the asynchronous and parallel one-stop pipeline enables rapid and integrated reporting of clinically actionable variants while conserving a limited amount of tumor tissue. Together, these features highlight the potential of CancerMaster to improve patient outcomes through personalized cancer treatment and serve as a resource for the discovery of novel biomarkers and therapeutic targets25. Continuous refinement of the panel and pipeline, guided by clinical feedback, will further enhance its impact on treatment-refractory patients and support advancements in drug development.
Methods
Panel design and capture protocol
“CancerMaster” uses a customized RNA probe designed for targeted enrichment sequencing. It encompasses all reported exons (7,811 regions) from 524 tumor-related genes. This panel covered 2,456,492 bases of hg19 (2.5 Mb). All gene transcripts reported by the UCSC hg19 were included to thoroughly detect SNVs, small indels, and structural variants. The CNVs of all 524 genes were detected using a normalized baseline comparison; we identified 143 genes of interest. In addition, 214 fusion breakpoint candidate regions were assessed, enabling the detection of genomic rearrangements in 18 genes (Supplementary Tables S6 and S7). The target panel size was designed to be sufficient for calculating the TMB. Furthermore, the panel contains specific RNA probes for detecting EBV and HPV. The CancerMaster panel was designed to capture 25 HPV subtypes and two EBV subtypes. All regions sharing similarities with the human genome were removed from the design before probe synthesis26,27. To capture all the target regions, libraries and capture probes were hybridized using a Celemics-Targeted Sequencing Kit (Celemics, Seoul, Korea), strictly following the manufacturer’s recommended protocol for custom panel designs. The captured DNA fragments were further amplified through post-PCR to enrich the sample. Pooled libraries containing the captured DNA fragments were subsequently sequenced on an Illumina NextSeq 500 Sequencing System as 2 × 100 bp paired-end reads.
One-stop automated analysis bioinformatics pipelines
Stringent QC criteria were established to ensure the reliability and accuracy of data generation. QC criteria focused on several key metrics, including uniformity (≥ 80% of 0.2× the mean coverage), GC content (approximately 50%), Q30 score (≥ 75%), mapping ratio (≥ 90%), duplication rate (≤ 35%), and mean coverage depth (≥ 800×). These QC criteria allowed us to effectively assess the quality of the NGS data and make informed decisions regarding their suitability for further analysis and interpretation.
Data processing was performed on a high-performance computing cluster using an asynchronous and parallel one-stop automated analysis pipeline developed using open-source software tools (Fig. 1)28,29,30. SNVs/indels were identified using VarScan 2.4.0, and CNVs were identified by comparing the depth of coverage over targeted regions in a tumor-only sample relative to a reconstructed baseline31,32,33. Fusion genes were identified using TopHat-Fusion 2.0.13 and reported with fusion scores higher than 0 and fusions with more than five spanning reads34. Low-quality reads were removed using an in-house script that filtered out read pairs that met at least two of the following criteria: a low mean base quality of less than 20; more than 5% bases with a base quality of less than 2; R1 and R2 having the same sequence. Among reads that passed the FASTQ filtration criteria, adaptor sequences overlapping more than three bases were trimmed from each read with Trimmomatic 0.33 and reads shorter than 36 bases were removed after trimming8. Reads were aligned to hg19 using BWA-aln version 0.7.10. Duplicate reads were marked using Picard Mark Duplicates version 1.115. The following processes recognized these marks to avoid bias. Called indels were realigned with GATK version 2.3.9 IndelRealigner, and base quality values were recalibrated with GATK BaseRecalibrator using a read group, reported quality score, cycle, and local sequence context covariates.
We constructed a viral reference genome set that included the EBV and 25 HPV sequence subtypes. We used mSINGS and a set of microsatellite markers to detect MSI (Supplementary Fig. S9)35. TMB (mutations [mut]/megabase [Mb]) was estimated as the total number of detected mutations (SNV VAF > 5% and indel VAF > 10%); synonymous and putative germline mutations were removed after correcting the misannotation of multiple nucleotide variants with BCFtools csq and WhatsHap36,37,38,39,40. We defined TMB-High as 20 or more, TMB-Low as less than 10, and TMB-Intermediate as between 20 and 10 mutations/Mb. We utilized SOAP-HLA version 2.2 to describe the HLA type and reported alleles with a type score higher than 5041.
Alteration calling
SNV and indels were called using VarScan 2.4.0 with a strand-filter and a minimum VAF of 1%. A coverage depth of > 10 was used as the cutoff for somatic variants with low allele frequencies. Annovar 2020 Jun07 was used to annotate the variants.
CNV were identified by comparing the depth of coverage over the targeted regions in a tumor-only sample relative to a reconstructed baseline. For the internal normalization of sequencing depth variation, the regional depth was divided by the median depth of each sample. Extreme values were removed from the normalized exonic depths and pooled values. This process was repeated until 30% of the value was retained. The mean value was extracted for each target region to construct the reference “normal” sample. For evaluating CNVs, we also used the segmental copy number calculated using the circular binary segmentation algorithm with “DNAcopy” R package version 1.60.0. Copy gain or loss was determined based on whether the segmental relative value was higher than 2 or lower than 0.5, genic-fold changes were greater than 3 or lower than 0.5, and if at least two regions had the same events. Genomic fusion breakpoints were detected using TopHat2 (v2.0.13)42 in split-read structural variant mode. All RNA-specific splice-junction inference features were disabled (--no-coverage-search) to ensure that TopHat2 operates as a general-purpose DNA structural variant mapper. Candidate fusion events were summarized using TopHat-Fusion-Post and reported only when the fusion score was greater than 0 and more than five split reads supported the breakpoint.
These criteria ensure that no splice-site motifs (GT–AG, GC–AG, AT–AC) or transcript-based predictions are required. The reported fusions represent high-confidence DNA-level rearrangements captured within the 214 targeted breakpoint-candidate regions across 18 clinically actionable genes in the panel.
EBV & HPV
Viral detection was performed using a two-step hybrid-mapping workflow. First, sequencing reads were aligned to the human reference genome (hg19) using BWA-MEM (v0.7.17). Reads unmapped or improperly mapped to the human genome were subsequently realigned to a curated viral reference genome set consisting of EBV (including EBV-WT) and 27 HPV subtypes (HPV9, 16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 68, 92, 96, 136, 140, 154, 156, 178, 179, 201, and 379).
To minimize false-positive viral signals, viral reads with low mapping quality (MAPQ < 20), multimapping reads, and alignments shorter than 30 bp were filtered out.
For HPV detection, samples with > 300 bp of uniquely aligned viral sequence were classified as HPV-positive. For EBV, the viral read depth was normalized to one million total aligned bases, and samples with normalized depth > 50 were considered EBV-positive.
Microsatellite instability
We employed mSINGS and a set of microsatellites to detect microsatellite instability17. First, we selected MSI-H and MSS samples that were previously clinically validated as the training set. Second, we identified microsatellites using MSIA and constructed a baseline by pooling the MSS samples. Third, we removed microsatellites with the same status from all MSS and MSI-H samples, as they were not helpful for classifying the MSI status. Fourth, we compared the MSS and MSI-H samples to select the set of microsatellites most beneficial for detecting MSI (Supplementary Fig. S9A). Fifth, microsatellite instability was detected using mSINGS with a selected set of microsatellites (Supplementary Fig. S9B).
Tumor mutation burden
TMB (mutations [mut]/megabase [Mb]) was estimated as the total number of detected mutations (SNV VAF > 5% and indel VAF > 10%), and synonymous and putative germline mutations were removed after correcting the misannotation of multiple nucleotide variants with BCFtools csq and WhatsHap. Putative germline mutations were defined as those reported in the Korean Variant Archive, Korean Reference Genome Database, and gnomAD_exome_EAS. Pathogenic or likely pathogenic mutations were assessed using the American College of Medical Genetics criteria and were divided by the length of the covered coding regions. We defined TMB-High as ≥ 20, TMB-Low as < 10, and TMB-Intermediate as between 20 and 10 mutations/Mb.
HLA type & HLA class I supertype
We utilized SOAP-HLA version 2.2 for HLA type and reported alleles with a type score higher than 50. The HLA allele identified through the SOAP-HLA analysis was reclassified as a supertype and used for subsequent analyses.
Clinically actionable alterations
We called variants within the target regions that resulted in changes to the protein’s primary sequence (e.g., nonsynonymous SNV, stop gain, stop loss, frameshift indel, and in-frame indel). Two separate types of reports were generated: a “Basket report” and an “Academic Report.” The Basket Report lists only the clinically actionable mutations (Supplementary Table S7). If the variants only matched the transcript ID, they were listed in the academic report.
Clinical comparison
This retrospective study included 668 patients with solid tumors who were treated at Yonsei Cancer Center (Seoul, Republic of Korea) between July 2014 and December 2021. All methods were carried out in accordance with relevant guidelines and regulations. The study protocol was reviewed and approved by the Institutional Review Board of Severance Hospital (IRB No: 4-2014-0349). Written informed consent was obtained from all participants and their legal guardians prior to sample collection. Clinical and pathological information was retrieved from electronic medical records and pathology reports. The following variables were collected for analysis: age at diagnosis; sex; tumor differentiation status. Biomarker results, including HER2, MET, EGFR expression, MSI/dMMR status, and EBV/HPV status, were obtained from the clinical diagnostic assays routinely used at Yonsei Cancer Center43.
Statistical analysis and visualization
Continuous variables were compared using the Wilcoxon rank-sum test to evaluate inter-group differences. All statistical analyses were performed in R version 4.0.4 (R Foundation for Statistical Computing, Vienna, Austria) using RStudio Desktop version 1.4.1103. Somatic variant data were processed and visualized using the maftools package. Mutation Annotation Format files generated from variant calling were imported into maftools, and standard functions were applied to summarize and display the mutational landscape. The visualization outputs included Oncoplots (gene-level mutation frequency across samples), variant classification plots (distribution of SNVs and indels by functional consequence), and transition–transversion plots. Bland–Altman analysis was performed to evaluate the agreement between expected and observed allele frequencies. The difference (Expected – Observed) was plotted against the average of the two values for each variant. Bias and 95% limits of agreement (mean difference ± 1.96 × SD) were calculated to assess systematic deviation and variability. Bland-Altman analysis was conducted using GraphPad Prism software (version 10.0, GraphPad software, La Jolla, CA). Co-occurrence and mutual exclusivity analyses among frequently mutated genes were also performed using maftools functions, and the results are displayed as pairwise association plots. All figures related to mutational profiling were generated directly from maftools outputs without additional manual modifications, ensuring reproducibility32.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request, due to restrictions related to patient confidentiality. Targeted panel sequencing data generated in this study have been deposited in the NCBI database under SRA accession codes PRJNA1395781. The remaining data are available within the Article or Supplementary Information.
References
Benson, A. B. et al. Colon Cancer, version 2.2021, NCCN clinical practice guidelines in oncology. J. Natl. Compr. Canc Netw. 19, 329–359 (2021).
Boumber, Y. Tumor mutational burden (TMB) as a biomarker of response to immunotherapy in small cell lung cancer. J. Thorac. Dis. 10, 4689–4693 (2018).
Marcus, L., Lemery, S. J., Keegan, P. & Pazdur, R. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors. Clin. Cancer Res. 25, 3753–3758 (2019).
Cheng, D. T. et al. Memorial sloan Kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Milbury, C. A. et al. Clinical and analytical validation of FoundationOne® CDx, a comprehensive genomic profiling assay for solid tumors. PloS One. 17, e0264138 (2022).
Frampton, G. M. et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat. Biotechnol. 31, 1023–1031 (2013).
Kim, E. et al. Identifying anti-cancer effects and exploring the mechanism of an MPS1/TTK inhibitor in gastric cancer. Cancer Res. Treat. 57, 803–820 (2025).
Akkari, Y. et al. Implementation of cancer next-generation sequencing testing in a community hospital. Mol. Case Stud. 5, a003707 (2019).
Robinson, J. T. et al. Integr. Genomics Viewer Nat Biotechnol 29, 24–26 (2011).
Muzny, D. M. et al. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Bass, A. J. et al. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Ma, F., Laster, K. & Dong, Z. The comparison of cancer gene mutation frequencies in Chinese and U.S. Patient populations. Nat. Commun. 13, 5651 (2022).
Jung, K. W. et al. Prediction of cancer incidence and mortality in Korea, 2025. Cancer Res. Treat. 57, 331–338 (2025).
Park, E. H. et al. Cancer statistics in korea: incidence, mortality, survival, and prevalence in 2022. Cancer Res. Treat. 57, 312–330 (2025).
Conroy, J. M. et al. A scalable high-throughput targeted next-generation sequencing assay for comprehensive genomic profiling of solid tumors. PLoS One. 16, e0260089 (2021).
Endris, V. et al. Measurement of tumor mutational burden (TMB) in routine molecular diagnostics: in Silico and real-life analysis of three larger gene panels. Int. J. Cancer. 144, 2303–2312 (2019).
O’Kane, G. M., Connor, A. A. & Gallinger, S. Characterization, detection, and treatment approaches for homologous recombination deficiency in cancer. Trends Mol. Med. 23, 1121–1137 (2017).
Pabla, S. et al. Development and analytical validation of a next-generation sequencing based microsatellite instability (MSI) assay. Oncotarget 10, 5181–5193 (2019).
Ratner, M. First multi-gene NGS diagnostic kit approved. Nat. Biotechnol. 35, 699 (2017).
Jung, M. et al. First-in-human phase 1 study of a B cell- and monocyte-based immunotherapeutic vaccine against HER2-positive advanced gastric cancer. Cancer Res. Treat. 56, 208–218 (2024).
Antoniotti, C. et al. Tumour mutational burden, microsatellite instability, and actionable alterations in metastatic colorectal cancer: next-generation sequencing results of TRIBE2 study. Eur. J. Cancer. 155, 73–84 (2021).
Salem, M. E. et al. Comparative molecular analysis between microsatellite instability-high (MSI-H) tumors with high tumor mutational burden (TMB-H) versus MSI-H tumors with TMB-intermediate/low. Ann. Oncol. 29, viii650–viii651 (2018).
Zhao, Z., Li, W., Zhang, X., Ge, M. & Song, C. Correlation between TMB and MSI in patients with solid tumors. J. Clin. Oncol. 38, e15169–e15169 (2020).
Chowell, D. et al. Patient HLA class I genotype influences cancer response to checkpoint Blockade immunotherapy. Science 359, 582–587 (2018).
Lee, C. K. et al. A single arm phase Ib/II trial of first-line pembrolizumab, trastuzumab and chemotherapy for advanced HER2-positive gastric cancer. Nat. Commun. 13, 6002 (2022).
Hassan, R. et al. Epstein-Barr virus (EBV) detection and typing by PCR: a contribution to diagnostic screening of EBV-positive burkitt’s lymphoma. Diagn. Pathol. 1, 17 (2006).
Monsonego, J. Emerging Issues on HPV Infections: from Science To Practice (S. Karger, 2006).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with bwa-mem. ArXiv 3, 13033997 (2013).
McKenna, A. et al. The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Kim, D. & Salzberg, S. L. TopHat-Fusion: an algorithm for discovery of novel fusion transcripts. Genome Biol. 12, R72 (2011).
Salipante, S. J., Scroggins, S. M., Hampel, H. L., Turner, E. H. & Pritchard, C. C. Microsatellite instability detection by next generation sequencing. Clin. Chem. 60, 1192–1199 (2014).
Danecek, P. & McCarthy, S. A. BCFtools/csq: haplotype-aware variant consequences. Bioinformatics 33, 2037–2039 (2017).
Jung, K. S. et al. KRGDB: the large-scale variant database of 1722 Koreans based on whole genome sequencing. Database (Oxford) baz146 (2020). (2020).
Lee, S. et al. Korean variant archive (KOVA): a reference database of genetic variations in the Korean population. Sci. Rep. 7, 4287 (2017).
Martin, M. Whatshap: fast and accurate read-based phasing. BioRxiv https://doi.org/10.1101/085050 (2016).
Xavier, A., Scott, R. J. & Talseth-Palmer, B. A. TAPES: A tool for assessment and prioritisation in exome studies. PLoS Comput. Biol. 15, e1007453 (2019).
Cao, H. et al. An integrated tool to study MHC region: accurate SNV detection and HLA genes typing in human MHC region using targeted high-throughput sequencing. PLoS One. 8, e69388 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Kim, H. S. et al. Comprehensive expression profiles of gastric cancer molecular subtypes by immunohistochemistry: implications for individualized therapy. Oncotarget 7, 44608–44620 (2016).
Funding
This study was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (2020R1A2B5B02001452) and also supported by a grant from the National R&D Program for Cancer Control, Ministry of Health and Welfare, Republic of Korea (HA16C0018).
Author information
Authors and Affiliations
Contributions
Conceptualization: H. C. Chung and S. Y. Rha; data curation: J. Che, W. S. Kwon, J. Kim, E. Jadamba, H. J. Han, Y. Kim, C. Lee, Y. J. Moon and H. B. Mun; formal analysis: J. Che and W. S. Kwon; investigation: J. Che, W. S. Kwon, J. Kim, E. Jadamba, H. J. Han, Y. Kim and H. C. Chung; methodology: J. Che, W. S. Kwon, J. Kim, E. Jadamba, H. J. Han, Y. Kim; project administration: H. C. Chung and S. Y. Rha; visualization: J. Che and W. S. Kwon; software: J. Che; funding acquisition: S. Y. Rha; writing—original draft: J. Che and W. S. Kwon; writing—review & and editing: all authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.








Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Che, J., Kwon, W.S., Kim, J. et al. Analytical and clinical validation of CancerMaster, an automated targeted NGS panel, for tumor-only precision oncology. Sci Rep 16, 8048 (2026). https://doi.org/10.1038/s41598-026-37991-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37991-0