Abstract
Introduction Approximately 1–5% of pregnant women experience recurrent pregnancy loss (RPL). Early detection and evaluation of high-risk variables allow for the initiation of recommended treatments while also reducing the likelihood of RPL in these couples. Objective The goal of this study was to build a deep learning model to identify an immune-lifestyle pattern in RPL patients based on clinical and laboratory findings. Methods We retrospectively collected the data from 16,818 RPL patients and 19,979 healthy women from across five clinics throughout Iran from December 2014 to April 2024. Fundamental population size and laboratory symptoms were gathered from the all-available data of participants. Data preprocessing involved cleaning and partitioning the data into training and validation groups. We used 22 characteristics to identify patterns via a deep learning model (TabNet). The model performance was evaluated using a confusion matrix, precision-recall curve, calibration plot, and Receiver Operating Characteristics (ROC) curve. Results The model showed robust and practical performance in identifying an immunological pattern based on the selected variables. The model yielded an AUC of 0.985, an accuracy of 0.946, a precision of 0.936, a specificity of 0.921, and a sensitivity of 0.968. Over-fitting was mitigated via repeated 5-fold cross validation (CV); no feature leakage detected. Conclusion Patterns linking immune and lifestyle factors to pregnancy loss were identified with high reliability using a deep learning approach. These findings may support a deeper understanding of the biological mechanisms underlying RPL and help guide the development of targeted treatment strategies.
Similar content being viewed by others
Introduction
According to European Society of Human Reproduction and Embryology (ESHRE) guideline recurrent pregnancy loss (RPL) is characterized as at least two consecutive pregnancy failures before the 24th week of pregnancy1. In 50% of RPL cases, no identifiable etiology has been found, which is known as unexplained RPL (uRPL)2. RPL affects approximately 1–5% of women who are trying to conceive. The intricacy and difficulty of RPL in reproductive medicine have raised concerns and anxiety among couples of reproductive age, and it imposes a significant psychological load on patients3. Studies demonstrate that the probability of re-miscarriage increases with a significant history of prior losses4. As a result, progress toward reliable diagnosis and early prediction of recurrent miscarriage is stalled.
Various risk factors are involved in the development of RPL, including: genetic or acquired abnormalities, immune system disorders, abnormal karyotypes, endocrine system, maternal age, lifestyle and environmental factors5,6. Among the numerous risk factors associated with RPL, several have been extensively studied to have notable effects on the pregnancy outcome. Notably, age, BMI, alcohol use, smoking status, vitamin D3, thyroid stimulating hormone (TSH), T-helper 1 (Th1)/Th2 ratio, T-cell (CD3), B-cells (CD19), helper T-cells (CD3/CD4), cytotoxic T-cells (CD3/CD8), Natural Killer (NK) cells (CD16/56), antiphospholipid antibody (aPL), anti-b2 glycoprotein antibody, anti-transglutaminase (TTG) antibody, antinuclear antibodies (ANAs), anti-ds-DNA, anti‐thyroperoxidase (TPO) antibodies, and anti‐thyroglobulin (TG) antibodies are among most studied risk factors7,8,9,10,11,12.
BMI and age have well known effect on fertility and RPL. It’s commonly understood that increased BMI can cause elevated levels of oxidative stress and systemic inflammation, decreased uterine acceptance, impaired oocyte metabolism and maturation, and metabolic syndrome, all of which are linked to dire reproductive results13. Smoking and alcohol use are powerful indicators of increased risk of pregnancy loss3. Cytotoxic T-cells and Th cells are essential for immune tolerance during pregnancy. Disruptions in the balance between these cell types may lead to immune-related infertility or RPL7. Moreover, In the first trimester, a shift toward Th1-like cells and cytokines creates the aseptic inflammation needed for embryo implantation. Predominance of Th2 responses is necessary after implantation preserving the fetus and balance the Th1 responses7,14. Both peripheral and decidual NK cells frequency and cytotoxicity are associated with RPL15. Thyroid dysfunction is a prevalent endocrine condition that affects 2–3% of women during their reproductive years and has been identified as a possible cause of RPL10. Furthermore, while the exact mechanism of most of autoantibodies such as aPL, anti-thyroid antibodies, ANAs and anti-TTG are not clear, their pathologic role suggest a possible involvement in RPL16.
Risk prediction models are widely used in the study of complicated diseases, particularly to analyze the association between baseline features and future outcomes in patients with specific conditions. Recently, artificial intelligence (AI), more specifically machine learning (ML), has been applied as a consistent model and forecasting tool11,17,18,19. For example, twenty ML algorithms have proven their robustness and adaptability across a variety of biomedical domains by successfully predicting hospitalization risk among COVID-19 patients using large clinical cohorts20, and in another study, two gradient‑boosting algorithms, XGBoost and CatBoost used both clinical and metabolic data for early type 2 diabetes risk prediction21. In reproductive medicine, several ML models have been developed to predict the pregnancy outcome using different characteristics. These models allow for the easy incorporation of vast volumes of data in addition to the clinical information accessible in the electronic health record (EHR) and the prognosis allows doctors to make suitable treatment options for patients. Moreover, ML models can improve over time by recalibrating each time new data is entered. As a result, our goal was to apply the risk probability deep learning model, TabNet, to identify the likelihood of an immune-lifestyle problem in RPL patients.
Materials and methods
Clinical records and cohort definition
The data was collected from the Tehran, Shiraz, Ardebil, Mashhad, and Tabriz fertility centers between December 2014 to April 2024. Each clinic provided their various data sets after they internally removed any patient identifiers. The dataset was then consolidated by the research team in prepared for analysis. Overall, the initial data was consisting of 19,167 RPL patients and 22,325 healthy women. Following the application of exclusion criteria, records were also removed if any numeric field (e.g., biomarker result or quantitative variable) contained text, symbols, or other non-numeric entries. After these exclusions, a total of 36,797 data points (16,818 patients and 19,979 healthy women) were included in the analysis. In some cases, the data came from patients with pregnancy problems where both partners underwent a comprehensive workup and the issue was determined to be male-related. Participants in the healthy group were followed up either by visiting the clinic and providing a history, or via phone call.
All methods were carried out in accordance with relevant guidelines and regulations. This retrospective study used fully de-identified datasets provided by participating clinics, each of which removed all patient identifiers prior to data sharing. The consolidated dataset was used solely for research purposes to develop a deep learning model. No human participants or human tissue samples were involved in this study. Due to the retrospective nature of the study, Research Ethics Committee of Tabriz University of Medical Sciences waived the need of obtaining informed consent. The study protocol was reviewed and approved by the Research Ethics Committee of Tabriz University of Medical Sciences, under approval number IR.TBZMED.REC.1403.1084.
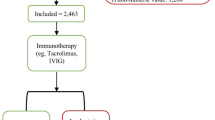
The latest ESHRE guidelines1 on the definition of RPL were used to include patient data in the cohorts. The inclusion criteria were as follows: 1) Women aged between 18 and 45 years who had experienced RPL that meets the diagnostic criteria of the ESHRE1. The exclusion criteria included the following: (1) chromosomal abnormalities, (2) Uterine anomalies, (3) Abnormal pregnancy outcomes such as ectopic pregnancy, and hydatidiform mole, (4) lost to follow up, (5) Endocrine dysfunction such as hyperprolactinemia, (6) Chronic endometritis, (7) Polyps, (8) Intrauterine adhesion, (9) Endometriosis, (10) Hydrosalpinx, 11) Adenomyosis, 12) Uterine Fibroids, 13) Abnormal ovarian reserve, 14) Mullerian abnormalities,, 15) Polycystic ovarian syndrome (PCOS), 16) Inherited/Acquired thrombophilias, 17) antithrombin deficiency, 18) history of prior immunological treatment, 19) infertility, and 20) male-related conditions. Figure 1 displays the patient selection and workflow diagram.
Workflow and patient selection. We used strict criteria to exclude patients with anatomical and gynecological anomalies.
Predictive variables and laboratory protocols
The main outcome of this study was to identify a pattern in RPL patients to determine whether an immune-lifestyle issue was present. The following laboratory tests were performed as part of each patient’s routine diagnostic workup at the fertility clinics, independent of this research study. Candidate variables were classified into 5 categories: (1) Demographic characteristics: age, BMI, (2) Biochemical tests: Vitamin D3, TSH, (3) Immune assays: Th1/Th2 ratio, CD3, CD16, CD19, CD56, CD3/CD4, CD3/CD8, CD16/56, and (4) Measurement of autoantibodies: Antiphospholipid antibody (IgG, IgM), Anticardiolipin antibody (IgG, IgM), Anti b2 glycoprotein antibody (IgG, IgM), Anti TTG antibody (IgG, IgA), ANA, Anti ds-DNA, Anti TPO, Anti TG; and (5) Life style: smoking and alcohol use. These markers have been carefully chosen based on clinical experience and published literature, as detailed in Table S1 (Supplementary File 1). Measurement methods and normal ranges are also provided in the same file.
Clinical diagnostic tests were performed at least 12 weeks after the last pregnancy loss in RPL patients, and 12 weeks prior to a pregnancy in healthy women, both in luteal phase. This timing was selected as part of routine clinical evaluation to support diagnosis and guide treatment planning. To minimize inter-center variability, unified assay kits and thresholds were used in all centers.
Data preprocessing
Missing laboratory values occurred across several immune markers. Rather than filling these gaps using statistical imputation, which can blur genuine biological differences, we used a single numeric placeholder (–1) that lies outside the physiological range of every variable. This allowed the model to treat a missing value as its own state rather than as an estimated number. TabNet is well suited to this strategy because its feature-selection steps and attention masks are able to learn separate internal pathways for observed and absent inputs, preserving the natural structure of the data without forcing assumptions about what a missing measurement “should” have been22. Conventional imputations, even simple mean or median replacement, can introduce bias and reduce the clarity of relationships in clinical datasets, a limitation highlighted in recent reviews of missing-data handling in deep learning23,24. Using a single, consistent rule across all features also avoids discrepancies between training and inference and keeps the preprocessing pipeline reproducible22,23,24. To confirm that this approach did not artificially influence model behavior, we compared the sentinel-based model with a complete-case version and found that the latter produced lower AUC and poorer specificity. We also repeated the analysis after altering half of the sentinel entries to a different placeholder and observed almost no change in performance, indicating that the model was not dependent on the specific value itself. Overall, the sentinel strategy preserved the integrity of the dataset while allowing the model to make use of all available records. Details of model fitting and validation tests are provided in Table S2 (Supplementary File 1).
Model architecture & training
We used a compact TabNet configuration (n_d = n_a = 16) to reduce the risk of overfitting on our ~ 36,000-patient dataset while maintaining clinical interpretability. Five attention steps (n_steps = 5) with a moderate feature reuse penalty (gamma = 1.5) promoted diversity in feature selection, and a light sparsity regularizer (lambda_sparse = 1e-4) encouraged meaningful feature attribution without compromising AUC. Training was stabilized using Ghost Batch Normalization (2048 batch size with virtual batches of 256), and early stopping (patience = 20, max_epochs = 300) was applied to the validation set only. The model was trained using fixed random seeds (seed = 42) with stratified 90/10 splits to ensure reproducibility and class balance across the train, and validation sets. The decision threshold for classification was fixed at 0.5. All confusion-matrix values correspond to this cutoff. Furthermore, the hyperparameters of the model were systematically constrained and predetermined in accordance with established guidelines for the application of TabNet to clinical datasets of similar dimensions, as opposed to being optimized for the purpose of maximizing performance metrics. This approach was used to mitigate the risk of overfitting, improve stability across resampled folds, and maintain the interpretability of feature attributions. Techniques such as early stopping and repeated cross-validation were employed to assess generalization; however, comprehensive hyperparameter tuning or automated search methodologies were purposefully eschewed to minimize post-hoc optimization bias. Consequently, the performance reported is indicative of reproducible model behavior under biologically relevant constraints rather than representing the peak of benchmark performance.
Because of its attention-based architecture, which permits transparent feature attribution and interaction analysis, and its suitability for structured tabular clinical data, TabNet was chosen as the modelling framework in advance25. Crucially, the goal of this study was not to compare and benchmark various machine learning algorithms. Rather, to limit analytic flexibility and concentrate on biological pattern discovery, a single, pre-specified architecture was employed. This strategy supports methodological continuity rather than post-hoc model selection and is in line with the applied TabNet model to immunological risk stratification25.
Statistical analysis
We randomly divided the dataset into training and validation cohorts at a 9:1 ratio, where the former was utilized for model development and the latter for internal validation. Continuous data with a normal distribution are presented as mean ± standard deviation (SD), while categorical variables are described using counts and percentages. To compare the means across three groups, we used one-way ANOVA (analysis of variance) when the assumptions of normality and homogeneity of variance were met. If these assumptions were violated, we used a Kruskal-Wallis H test for a non-parametric alternative. For categorical data, chi-square test was used. Moreover, logistic regression algorithm and multivariable regression analysis were employed with the training group to generate odds ratios. P-value < 0.5 was considered significant. The evaluation metrics for the model comprised the AUC, the receiver operating characteristic curve (ROC), calibration plots, accuracy, precision, recall, F1-score, and specificity. The model’s performance was assessed by comparing predicted and true values using a confusion matrix.
Results
Baseline characteristics of patients
Overall, 19,979 healthy controls and 16,818 RPL patients were included in the study (Fig. 1). Table 1 presents the pre-pregnancy characteristics of these individuals, including blood tests indicators and lifestyle factors of RPL patients and healthy controls. To better understand the patient population, we divided RPL patients into two groups of abnormal and normal tests, based on standard reference ranges. Abnormal tests were defined as values falling outside the normal range. Significant differences were observed among these characteristics between healthy controls, RPL patients with normal tests, and RPL patients with abnormal tests, except for Vitamin D3 (p = 0.06). (Table 1). To ensure balanced representation across cohorts, the data were randomly shuffled and split using stratified sampling, maintaining class proportions across the training, and validation sets.
Univariable and multivariable logistic regression analysis
Following univariable and multivariable logistic regression, most of the variables examined were found to be significant independent predictors (p < 0.05). Only the alcohol use (OR = 0.87, 95% CI: 0.63–1.21), smoking (OR = 0.98, 95% CI: 0.81–1.18), vitamin D3 (OR = 1.00, 95% CI: 1.00–1.00), and TSH (OR = 0.98, 95% CI: 0.96–1.00) were not significant risk factors. Odds ratios (95% CI) for all variables are reported in Table 2.
Performance of the TabNet model
According to the confusion matrix (Fig. 2A), the model produced high true-positive (TP = 1,549) and true-negative (TN = 1,934) counts relative to false positives (FP = 133) and false negatives (FN = 64), indicating strong classification accuracy and balanced discrimination across both outcome classes. As shown in Table 3, the TabNet classifier achieved an accuracy of 0.946, specificity 0.921, and sensitivity 0.968 on the validation set (Fig. 2A).
TabNet Model performance.(A)The confusion matrix, (B)Precision-Recall curve,(C) ROC curve for training set, and (D)ROC curve for validation set. TP: True positive; FP: False positive; FN: False Negative; TN: True Negative.
The precision–recall curve (Fig. 2B) remained consistently high, underscoring reliability in imbalanced-class contexts and minimizing false-negative outcomes. An AUC of 0.985, which indicates excellent class separation, was obtained from ROC analysis. ROC curves of both the training and validation datasets (Fig. 2C and D) showed strong generalizability and minimal overfitting, with both curves closely aligned in the upper-left quadrant. The training curve displayed tight calibration, and the validation plot’s small mid-range deviations most likely reflect sparse bin counts rather than misspecification. Calibration plots (Fig. 3) affirmed that predicted probabilities were closely aligned with observed event frequencies. In addition to the initial 90/10 train–validation split, repeated 5 × 5 stratified cross-validation was performed to verify robustness. This procedure produced a mean AUROC = 0.984 ± 0.007, confirming model stability and excluding overfitting across resampled folds. The classification threshold was fixed at 0.5, and all confusion-matrix statistics correspond to this cutoff. Minor rounding differences between the numerical metrics and the matrix counts arise from averaging across cross-validation iterations.
Calibration plots for the training set (A) and validation set (B). The training plot demonstrates near-perfect calibration, while the validation plot reveals only minor deviation around mid-range probabilities.
Given that the model focusses on a proximal immune-lifestyle signature rather than a distal clinical endpoint, the high AUROC observed makes biological sense. Numerous well-known, high-signal immunological axes in RPL are represented by many of the top-ranked predictors. Furthermore, performance is less likely to be influenced by narrow cohort artefacts due to the large sample size and multi-center heterogeneity.
Over-fitting, feature-leakage assessment, and missingness
To ensure the robustness and generalizability of the model, several validation strategies were implemented. A repeated 5 × 5 stratified cross-validation demonstrated high internal consistency, with a mean AUROC of 0.984 ± 0.007 (range: 0.961–0.992), only 0.009 below the single hold-out score (0.993), closely matching the single hold-out validation performance, and also indicating low variance across folds and no evidence of overfitting.
To rule out feature-to-label leakage, a permuted-label sanity check was conducted, which yielded an AUROC of 0.516 - near random performance - confirming no inadvertent use of identifying or time-based information. The model was trained with early stopping (patience = 20 epochs, max = 300 epochs), further reducing overfitting risk.
A fixed random seed (42) ensured reproducibility across NumPy, Python, and PyTorch environments, with a 90/10 patient-level stratified split to avoid data leakage across sets.
Finally, a sentinel-flip stress test confirmed that the model’s performance was not artificially inflated by the use of “−1” as the missing-value indicator. Replacing half of these sentinel values with “−2” led to an imperceptible AUROC drop (Δ = 0.001), supporting the conclusion that the model relies on physiological signal rather than exploiting encoding artifacts. Calibration was assessed using Brier score (0.036) and a calibration curve, both of which confirmed clinically acceptable probability estimation. The information regarding the tests is provided in Table S2 (supplementary file 1).
Out of the 22 input features used in the TabNet model, 10 had missing values, with the most affected being CD3, CD3/CD4, and CD3/CD8 - each missing in approximately 13.5% of records (refer to Table S3). Rather than relying on statistical imputation, missing entries were replaced with “−1”, a value that falls clearly outside the normal biological range for all quantitative markers. This choice ensures that the model can distinguish between true measurements and absent values without introducing noise or artificial bias. TabNet, which is well-suited to learning from structured inputs with missingness, treats such placeholder values as meaningful signals during training. Despite moderate gaps in a few immunological features, the model’s overall performance remained strong, suggesting it is not overly sensitive to missing data. Because the gaps were spread across multiple inputs and didn’t affect the target variable (R-LiveBirth), there’s little evidence of training bias or leakage. Moreover, several features - including TSH, Vitamin D, and lifestyle-related inputs - were complete and offered consistent support for stable split learning throughout the model.
To assess whether missing values influenced model performance, we compared two approaches: one trained on the full dataset with missing entries replaced by a sentinel (“−1”), and another restricted to rows with no missing data. The model using all records performed slightly better overall, with an accuracy of 94.6% and an AUC of 0.985. By comparison, the complete-case model reached 93.3% accuracy and an AUC of 0.971 (Table S4). While recall was marginally higher in the complete-case version (0.980 versus 0.968), it came with a notable drop in specificity (0.838 vs. 0.921). This suggests that removing incomplete rows may have reduced the model’s ability to generalize, possibly due to the smaller training set. Keeping all rows and marking missingness explicitly gave the model access to more information, which seems to have helped without introducing noise. Based on these results, handling missing data through sentinel substitution appears to be both practical and effective in this setting.
Importance of features
When all input variables were considered, the model’s feature importance analysis (Fig. 4) highlighted CD3/CD4 ratio and Th1/Th2 ratio as the most influential predictors. Other key contributors included CD19, BMI, age, CD3, and several autoantibodies, suggesting that both immune markers and lifestyle factors played a central role in the model’s predictions.
Lambda Plots of TabNet model showing feature importance. Overall importance of variables in identifying immune pattern between healthy women and RPL patients. The x-axis generally reflects the importance score of each feature in the model’s decision-making process, with higher values indicating greater importance.
Feature importance in deep learning models like TabNet is inherently less intuitive compared to traditional models. Unlike simpler models such as Random Forests that use metrics like Gini importance, TabNet learns which features to attend to at each decision step using its attention mechanism. This allows it to capture complex, context-dependent interactions between features, making the contribution of each variable dynamic and sometimes only meaningful in combination with others.
While some variables such as CD16, Vitamin D, Smoking, Alcohol, TSH, Anti-dsDNA, Anti-cardiolipin, Anti-TPO, and CD3/CD8 appear less important at first glance, they still serve critical roles in the model. For instance, in scenarios with missing data, these “low importance” features can compensate and allow the model to maintain high predictive performance. In fact, our experiments showed that the model remained effective even when using only a subset of 6–8 key features. This adaptability highlights one of the key strengths of deep learning approaches in clinical settings, which is robustness to incomplete data and the ability to dynamically adjust to the context of available features.
By retaining all variables and utilizing a model capable of handling missing data directly, we can apply the same architecture to retrospective clinical datasets where data may be incomplete or inconsistently recorded. This approach allows us to maximize validation opportunities using real-world patient records and supports ongoing investigation into the clinical relevance of each marker, even in the presence of sparsity or data gaps.
Feature correlation analysis
To investigate the relationships between all potential predictors, including immunological, biochemical, and lifestyle factors, a correlation analysis was performed. All continuous variables had their Pearson’s correlation coefficients calculated, and Fig. 5 shows the correlation matrix that was produced.
Correlation matrix among candidate predictors. Red and blue shades denote positive and negative Pearson correlations, respectively. Strong intercorrelations were observed within lymphocyte subsets, while biochemical and lifestyle variables showed weak or no correlation with immunological features.
Different patterns of association within immunological markers were revealed by the matrix. A high degree of co-regulation across lymphocyte populations was indicated by the strong positive correlations found between the T-cell and natural killer (NK)-cell subsets, including CD3, CD3/CD4, CD3/CD8, CD16, CD56, and CD19 (r > 0.70). In line with the functional balance between T-helper subsets and cytotoxic immune activity, the Th1:Th2 ratio showed moderate inverse correlations with a number of these CD markers (r ≈ − 0.40 to − 0.60).
The presence of weak to moderate positive correlations (r = 0.30–0.55) between autoantibody markers, including Anti-Phospholipid (Anti-Phospho), Anti-Cardiolipin (Anti-Cardio), Anti-β2-Glycoprotein (Anti-B2 Glyco), Anti-TTG, Anti-TPO, and Anti-TG, suggests that there may be overlap in autoimmune activity. Antibodies against ANA and dsDNA, on the other hand, showed weaker associations with other autoimmune markers, indicating a somewhat different immunopathological profile.
TSH and vitamin D were among the biochemical variables that had a weak correlation (|r| < 0.25) with immune indices. Similarly, there was little correlation between lifestyle factors like age, BMI, smoking, and alcohol and immunological or serological markers, suggesting that these factors have mostly independent effects on the model (Table S5-S13). Overall, the correlation structure shows that, aside from the expected clustering among lymphocyte subsets, the majority of predictors provide distinct information to the model with little redundancy. Since multicollinearity was not found to be significant (no pairwise correlation was found to exceed |r| > 0.90), these results support the inclusion of all listed features in subsequent multivariable modeling.
Discussion
Overview
In our study, the presence of an immune-lifestyle pattern in patients suffering from RPL was predicted using TabNet model based on several key parameters with high accuracy. To the best of our knowledge, this is the first study to discover an immune-lifestyle pattern in a large group of patients and a healthy population using 22 specific parameters. This method offers a new multivariable approach to evaluating this condition, combining the 22 parameters to find an immunological problem, rather than the traditional clinical focus on individual parameters. This approach may assist physicians in identifying patients who are likely to benefit from immune-modulating treatments. The model targets a proximal immune-lifestyle signature rather than a distal clinical endpoint, making the high performance observed biologically plausible.
Comparison with previous machine learning models
The domain of ML has seen significant improvements in deep learning, particularly in representing complex clinical outcomes from EHR data. Shickel et al. investigated the role of deep learning in clinical activities (information extraction, representation learning, and outcome prediction) using EHR data26. ML models can continually improve by recalibrating when new data is input. Studies have identified numerous contributing factors to RPL and attempted to develop several models to predict RPL. Bashiri et al., using a multivariable logistic regression model, proposed prognostic indicators including as material age, previous pregnancy loss, positive RPL, and primary versus secondary RPL27. In a study in 2020, six classification algorithms were created and tested to accurately predict early pregnancy loss (EPL) following the development of fetal cardiac activity during in vitro fertilization-embryo transfer (IVF-ET)28. When including fetal heart rate as a feature, Random Forest was the most accurate of the six predictive models, with a recall ratio and F1 of up to 97% and an AUC of up to 0.97.
Previous models using immunological and clinical parameters
Regarding the prediction of RPL, two studies used pre-conception immunological parameters11,18, which is similar to our study. Li et al. highlighted the significance of maternal peripheral blood immunological characteristics as indicators of future pregnancy outcomes in individuals with unexplained RPL. They developed and evaluated four models based on immunological profiles and discovered five predictors in the following order: 1, 2) complement system (e.g., C3 and C4), 3) dyslipidemia (e.g., triglycerides levels), 4, 5) number and percentage of peripheral NK cells (pNK)11. Their second model achieved the AUC of 0.96 with only using seven risk factors in predicting clinical pregnancy (> 28 weeks of gestational age). Similarly, Yang et al.18. employed pre-conception RPL patients’ information and clinical laboratory biomarkers to predict EPL (< 10 weeks of gestational age). Their GBM model achieved AUC of 0.805 using 16 variables. Their findings revealed that age, BMI, previous induced abortion, number of previous miscarriages, anticardiolipin antibody (ACA), homocysteine, IgM, low-density lipoprotein to high-density lipoprotein ratio (LHR), and platelet to neutrophilic ratio (PNR) were key risk factors for EPL18.
Recently, Li et al.17 proposed a nomogram to predict subsequent PL in RPL patients. Their model was based on nine features and the AUC was 0.795. Similarly, Mu et al.10 established a nomogram using thyroid and thromboelastograph (TEG) parameters to predict the risk of subsequent PL in RPL patients. Their variables included: maternal age, BMI, T4, TSH, TGAb, TPOAb, and TEG examination (e.g., reaction time, kinetic time, alpha angle, etc.). Their nomogram achieved AUC of 0.767.
Feature selection and model construction
To date, most proposed models for pregnancy outcome have been constructed based on features selected through multivariable regression or the least absolute shrinkage and selection operator (LASSO)10,11,17,18,19. However, this approach has introduced significant variability in identifying the independent or key risk factors as previously discussed. Such variability may stem from heterogeneity in populations or different pregnancy outcomes including EPL, clinical pregnancy, and live birth. To address this issue, we selected the features based on the comprehensive review of the current literature regarding RPL, combined with our own experience and expertise. This approach allowed us to prioritize the most relevant risk factors without relying on data-driven selection techniques which may overlook critical features due to their statistical constraints.
Our findings demonstrated discrepancies in the selection of key risk factors when comparing multivariable analysis with the deep learning model. For example, although multivariable analysis revealed significant differences in factors such as CD16 + cells, anti-dsDNA, anti-TPO antibodies, and anticardiolipin antibody (IgG, IgM) between the two groups (Table 2), these same variables were among the lowest in importance according to the model’s feature importance plot (Fig. 4).
Immune–metabolic and lifestyle interactions
Feature attribution in our model indicates that metabolic inflammation, influenced mainly by high BMI and cigarette use, intensifies Th1-dominant immune responses. This relationship provides a plausible biological link between lifestyle and immune imbalance in RPL. In several patient subgroups, markers such as the Th1:Th2 ratio, CD3/CD4, and thyroid autoantibodies contributed most strongly to prediction, suggesting that altered immune polarity is central to outcome differences.
Patients with elevated BMI showed clearer immune separation, consistent with reports that obesity promotes low-grade inflammation, enhances Th1 cytokine release, and shifts T-cell activation thresholds29,30. Smoking appears to act in a similar direction by increasing oxidative stress and endothelial reactivity, both of which disturb tolerance at the maternal–fetal interface31,32. These mechanisms help explain why the model was particularly sensitive in overweight and smoking groups, while specificity fell slightly due to overlapping inflammatory signals in subclinical cases.
Women who reported moderate alcohol intake showed a somewhat different pattern, with improved calibration and higher specificity. This may reflect the transient anti-inflammatory effect of light ethanol exposure, previously linked to better HDL function and reduced cytokine activation33,34,35. Overall, the data suggest that everyday behaviors leave measurable fingerprints on immune–metabolic balance, influencing implantation success and pregnancy maintenance.
Our model therefore extends beyond prediction. It highlights a reproducible immune–metabolic signature through which modifiable lifestyle factors shape immune tolerance. This understanding may guide future interventional studies combining lifestyle adjustment with targeted immune therapy to restore reproductive balance.
For instance, these results highlight the significance of including lifestyle change into RPL management procedures from a clinical standpoint. In addition to smoking cessation programs, weight optimization by diet and physical exercise interventions may help lower systemic inflammation and restore Th1/Th2 balance prior to conception. Clinicians may be able to personalize preconception counseling and identify patients who might benefit from supplementary immunomodulatory medications by routinely evaluating metabolic and immunological indicators found by the model. These integrated strategies, which target immunological and behavioral mechanisms, may improve pregnancy maintenance in women with RPL.
Model strengths, interpretability, and practical implications
Our study’s key strengths include the substantial patient cohort and high AUC achieved by our model using 22 variables. Importantly, the model focuses on identifying a biological/behavioral signature, not predicting a complex pregnancy outcome directly. That is more realistic given our dataset and could be the basis for a powerful risk stratification or decision-support tool in the clinic. Leveraging our experience and extensive research, we identified and selected most relevant risk factors instead of relying on automated selection methods to determine features from a large pool. While our model generated feature importance plots (Lambda plots), we recognized that even the least important feature in the plot could prove essential in cases of missing data.
A degree of interpretive tension exists between clinical reasoning and AI-based modeling. Clinicians often seek bottom-up, pathophysiological explanations for model predictions, whereas deep learning methods such as TabNet identify high-dimensional interaction patterns that are not easily reduced to linear cause–effect relationships. The model therefore captures emergent immune–lifestyle signatures that are physiologically consistent but exceed the resolution of traditional statistical approaches. Furthermore, a key component of contemporary biomedical AI is the use of neural architectures that remain robust in the face of noisy or incomplete clinical data. For example, recent research in oncology has shown how specialised neural network techniques, like Self-organising Maps with block processing, can greatly improve diagnostic accuracy even when there is a lot of noise in the dataset36. In a similar vein, we use TabNet’s attention masks to ensure that the overall integrity of the patient’s immune-lifestyle profile is not compromised by missing measurements. This framework enhances clinical judgment by revealing latent biological relationships that need more investigation, rather than replacing it.
Furthermore, the cost of these laboratory tests is considerably less than that of advanced therapies such as Lymphocyte immunotherapy (LIT), intravenous immunoglobulin (IVIG), and Immunosuppressives (e.g., Tacrolimus). Reducing the number of features could potentially decrease the AUC of the model, leading to less accurate predictions and imposing significant economic and psychological burden on both patients and healthcare system. Therefore, our aim was to construct the model with the highest possible accuracy.
Additionally, the model user interface (UI) is very simple and easy to use. Since it only requires users to download an excel file (available at the website), then fill the excel with the 22 parameters, and upload it to the web-based model. Finally, after analysis by the model, the model will provide a downloadable report that consists information regarding the immune status of the patient, probability of live birth, etc. Clinicians can use the report to conceive better decisions and guide their approach such as the need and type of intervention for each patient (Supplementary file 2).
Translational value and novel contributions
The novelty of our work is the quantitative integration of these immune parameters with modifiable lifestyle variables within a single, high-performance deep learning framework, even though the immunologic mechanisms linked to RPL, such as Th1/Th2 imbalance and autoantibody activation, have been previously described. This model uses routine laboratory data to create a composite immune-metabolic fingerprint that can identify high-risk women prior to conception, rather than just confirming known risk factors. Clinically, this makes it possible to incorporate targeted interventions like weight loss, quitting smoking, or selective immunomodulation before conception or assisted reproduction, as well as early, individualized counseling. Additionally, by converting complex immune profiles into useful insights, the model’s explainable outputs (feature attributions and risk probabilities) can aid in clinical decision-making and patient education. In this way, the model acts as a link between bedside risk assessment and laboratory immunology, opening the door for future validation and the creation of AI-assisted reproductive immunology decision-support tools.
Importantly, the current model does not define current clinical subtypes RPL or suggest a new biological phenotype. Instead, it combines well-established lifestyle and immunological variables into a single, data-driven risk signature that represents the non-linear interactions between these variables at the individual level. The TabNet framework emphasizes conditional interaction patterns, such as the joint contribution of Th1/Th2 imbalance, CD3/CD4 ratios, and metabolic indicators such as BMI, whose combined influence is not easily apparent from univariate or multivariable statistics alone, in contrast to conventional regression-based approaches that estimate independent effects. These patterns of interaction show a spectrum of immune-metabolic dysregulation rather than distinct clusters. Therefore, the model’s main contribution in the AI-based approach to an immune-lifestyle condition, which suggests rather than focusing on immune markers in isolation, future studies should emphasis on how combination of several factors including metabolic inflammation, life style, and immune profile disrupts maternal-fetal tolerance. Without creating new disease categories, this integrative viewpoint offers a biologically sound framework for planning targeted experimental and interventional research.
Conclusion
In conclusion, we built a deep learning model, TabNet, based on 22 variables and data from over 36,000 patients to identify a specific immune-lifestyle risk profile, which could help stratify RPL patients for targeted preventive care or immunomodulation.
Limitations
The primary limitation of this study is that it is based on a single-country dataset with internal validation only. Additionally, the model was developed using 22 predefined variables, some of which - particularly immunological markers - are not yet incorporated into international guidelines. Nonetheless, these factors have been extensively studied for their role in feto-maternal tolerance, especially in cases of unexplained RPL. As the model is still in an early developmental stage, it does not yet differentiate between humoral, cellular, or mixed immunological profiles. Moreover, as we acknowledge that further validation through external, multi-center studies and clinical trials is needed to assess the model’s performance and generalizability, which we aim to undertake in future work. We recognize that temporal hold-out validation and leave-one-center-out (LOCO) would provide more information about the model’s practicality, particularly with regard to site-specific and time-dependent variability. However, due to lack of date stamps and center identifiers in the current dataset, these analyses were not yet possible. Moreover, we found the model to be stable across the checks we were able to perform, but we also recognize that a simple 90/10 split does not capture differences that may exist between clinics. Although this restricts formal evaluation of of site-specific effects, participating clinics used harmonized diagnostic protocols and assay platforms, reducing systematic inter-center bias. Future external validation studies will explicitly incorporate center-aware and temporal validation frameworks.
Future direction
The next stage of our study will concentrate on thorough external validation to improve generalizability. To acquire diverse patient cohorts, we intend to form data-sharing partnerships with fertility and reproductive immunology facilities in several nations. Temporal hold-out validation and LOCO will be performed using newly collected data. To facilitate these analyses, we are now standardizing variable definitions, timestamps, and data formats amongst institutions. This methodical approach will guarantee a thorough evaluation of the model’s repeatability and practicality in a range of clinical contexts.
Data availability
The deep learning model, and the datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
RPL, E. G. G. et al. ESHRE guideline: recurrent pregnancy loss. Hum. Reprod. open. 2018 (2), hoy004 (2018).
Daya, S. & Stephenson, M. D. Frequency of factors associated with habitual abortion in 197 couples. Fertil. Steril. 66 (1), 24–29 (1996).
Tong, F. et al. The epidemiology of pregnancy loss: global burden, variable risk factors, and predictions. Hum. Reprod. 39 (4), 834–848 (2024).
Brigham, S. A., Conlon, C. & Farquharson, R. G. A longitudinal study of pregnancy outcome following idiopathic recurrent miscarriage. Hum. Reprod. 14 (11), 2868–2871 (1999).
Larsen, E. C. et al. New insights into mechanisms behind miscarriage. BMC Med. 11, 1–10 (2013).
Aslanian-Kalkhoran, L. et al. Understanding main pregnancy complications through animal models. J. Reprod. Immunol. 153, 103676 (2022).
Esparvarinha, M. et al. Dominant immune cells in pregnancy and pregnancy complications: T helper cells (TH1/TH2, TH17/Treg cells), NK cells, MDSCs, and the immune checkpoints. Cell. Biol. Int. 47 (3), 507–519 (2023).
Aslanian-Kalkhoran, L. et al. The effect of lymphocyte immunotherapy (LIT) in modulating immune responses in patients with recurrent pregnancy loss (RPL). Int. Immunopharmacol. 121, 110326 (2023).
Sarkesh, A. et al. The Immunomodulatory effect of intradermal allogeneic PBMC therapy in patients with recurrent spontaneous abortion. J. Reprod. Immunol. 156, 103818 (2023).
Mu, F. et al. A new prognostic model for recurrent pregnancy loss: assessment of thyroid and thromboelastograph parameters. Front. Endocrinol. 15, 1415786 (2024).
Li, Y. et al. Immunological parameters of maternal peripheral blood as predictors of future pregnancy outcomes in patients with unexplained recurrent pregnancy loss 1444–1456 (Acta Obstetricia et Gynecologica Scandinavica, 2024).
Dashti, M. et al. Impact of lymphocyte immunotherapy (LIT) on fertility rates in recurrent pregnancy loss (RPL) women with antinuclear antibodies: a randomized clinical trial. J. Reprod. Immunol. 168, 104432 (2025).
Cavalcante, M. B. et al. Obesity and recurrent miscarriage: A systematic review and meta-analysis. J. Obstet. Gynecol. Res. 45 (1), 30–38 (2019).
Ozkan, Z. S. et al. What is the impact of Th1/Th2 ratio, SOCS3, IL17, and IL35 levels in unexplained infertility? J. Reprod. Immunol. 103, 53–58 (2014).
Ahmadi, M. et al. NK cell frequency and cytotoxicity in correlation to pregnancy outcome and response to IVIG therapy among women with recurrent pregnancy loss. J. Cell. Physiol. 234 (6), 9428–9437 (2019).
D’Ippolito, S. et al. The pathogenic role of autoantibodies in recurrent pregnancy loss. Am. J. Reprod. Immunol. 83 (1), e13200 (2020).
Li, M. et al. A nomogram and risk stratification to predict subsequent pregnancy loss in patients with recurrent pregnancy loss. Hum. Reprod. 39 (10), 2221–2232 (2024).
Yang, X. et al. Predicting risk of the subsequent early pregnancy loss in women with recurrent pregnancy loss based on preconception data. BMC Women’s Health. 24 (1), 381 (2024).
Wei, C. et al. Identification and verification of diagnostic biomarkers in recurrent pregnancy loss via machine learning algorithm and WGCNA. Front. Immunol. 14, 1241816 (2023).
Salehnasab, Z. et al. Predictive modeling of COVID-19 hospitalization using Twenty machine learning classification algorithms on cohort data. Front. Health Inf., 12. (2023).
Rafie, Z. et al. Leveraging XGBoost and explainable AI for accurate prediction of type 2 diabetes. BMC Public. Health. 25 (1), 3688 (2025).
Arik, S. Ö. & Pfister, T. Tabnet: Attentive interpretable tabular learning. in Proceedings of the AAAI conference on artificial intelligence. (2021).
Liu, M. et al. Handling missing values in healthcare data: A systematic review of deep learning-based imputation techniques. Artif. Intell. Med. 142, 102587 (2023).
Che, Z. et al. Recurrent neural networks for multivariate time series with missing values. Sci. Rep. 8 (1), 6085 (2018).
Dashti, M. et al. Immunological risk factors for recurrent implantation failure using a deep learning model: a multicenter retrospective cohort study. Sci. Rep. 15 (1), 42822 (2025).
Shickel, B. et al. Deep EHR: a survey of recent advances in deep learning techniques for electronic health record (EHR) analysis. IEEE J. Biomedical Health Inf. 22 (5), 1589–1604 (2017).
Bashiri, A. et al. A proposed prognostic prediction tool for a live birth among women with recurrent pregnancy loss. J. Maternal-Fetal Neonatal Med. 35 (19), 3736–3742 (2022).
Liu, L. et al. Machine learning algorithms to predict early pregnancy loss after in vitro fertilization-embryo transfer with fetal heart rate as a strong predictor Vol. 196, 105624 (Computer Methods and Programs in Biomedicine, 2020).
Bae, H. R. et al. IFNγ is a key link between obesity and Th1-mediated autoimmune diseases. Int. J. Mol. Sci. 22 (1), 208 (2020).
Schmidt, F. M. et al. Inflammatory cytokines in general and central obesity and modulating effects of physical activity. PloS One. 10 (3), e0121971 (2015).
Arnson, Y., Shoenfeld, Y. & Amital, H. Effects of tobacco smoke on immunity, inflammation and autoimmunity. J. Autoimmun. 34 (3), J258–J265 (2010).
Pereira, R. D. et al. Angiogenesis in the placenta: the role of reactive oxygen species signaling. Biomed. Res. Int. 2015 (1), 814543 (2015).
Romeo, J. et al. Moderate alcohol consumption and the immune system: a review. Br. J. Nutr. 98 (S1), S111–S115 (2007).
Imhof, A. et al. Effect of alcohol consumption on systemic markers of inflammation. Lancet 357 (9258), 763–767 (2001).
Brien, S. E. et al. Effect of alcohol consumption on biological markers associated with risk of coronary heart disease: systematic review and meta-analysis of interventional studies. Bmj, 342. (2011).
Porkar, P. et al. Enhancing cancer zone diagnosis in MRI images: A novel SOM neural network approach with block processing in the presence of noise. Iran. J. Blood Cancer. 17 (2), 34–45 (2025).
Funding
This research was funded by iFertility Technologies.
Author information
Authors and Affiliations
Contributions
M.Y. and A.A. designed the study. Ma.Da., S.B., F.C., S.R., N.N., K.B., S.K., J.Z., P.K., and S.Da. contributed to the data collection and extraction. S.D. and J.A.H selected the patients. A.A. carried out statistical analysis and constructed the model. S.D., and M.D. interpreted the data. M.D. drafted the manuscript. A.G. prepared the figures. J.A.H., M.Y., and A.A. revised the manuscript. All authors approved the final manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
M.Y. is the Chief Scientist, and A.A. is the CEO/Chief Engineer of iFertility Technologies.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dashti, M., Aslanian-Kalkhoran, L., Doustfateme, S. et al. Multivariable AI-based analysis of immune–lifestyle patterns associated with recurrent pregnancy loss: an exploratory retrospective study. Sci Rep 16, 8250 (2026). https://doi.org/10.1038/s41598-026-38941-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-38941-6