Abstract
The circadian clock is an internal timekeeping system that generates ~ 24-h cycles in physiology and behavior, maintaining a remarkably consistent period across physiological conditions and temperature1. Sodium chloride (NaCl) is a key physiological ion whose concentration varies across species, yet its influence on circadian rhythms remains poorly understood. Using a reconstituted cyanobacterial oscillator composed of KaiA, KaiB, and KaiC, we investigated how NaCl modulates circadian period and temperature compensation. Increasing NaCl concentrations progressively shortened the circadian period without substantially affecting oscillation amplitude, mimicking the effects of KaiB titration. While temperature compensation was maintained across varying KaiB concentrations, changes in NaCl partially disrupted temperature compensation, as Q₁₀ values correlated positively with salt concentration. We propose that NaCl perturbs the equilibrium of KaiB conformations and oligomerization, normally stabilized across physiological temperatures, thereby modulating circadian period and temperature compensation. These findings provide insight into how physiological salt levels influence circadian timing and drive the diversification of clock proteins across species.
Similar content being viewed by others
Introduction
Dietary salt, particularly NaCl, is an essential component of the human body, playing a vital role in maintaining proper physiological function. Adequate sodium levels support the transport of nutrients across cell membranes and influence enzyme activity, which in turn affects energy metabolism1. Although numerous clinical studies have reported a direct impact of sodium on circadian rhythms2,3,4, the molecular mechanisms underlying this effect within individual cellular clocks remain largely unexplored.
The circadian clock is an internal timekeeping system that generates rhythmic signals with an approximately 24-h period5. An interesting feature of the circadian rhythm is its ability to maintain a consistent period across the normal physiological temperature range, a property known as temperature compensation6. The circadian clock governs sleep–wake patterns, hormone secretion, body temperature, metabolism, and various cellular functions. Circadian clocks are conserved across nearly all light-perceiving organisms, from bacteria to humans.
Cyanobacteria possess one of the simplest known circadian clocks, making them an ideal model for studying the molecular mechanisms of biological timekeeping7.
Their core clock machinery consists of three proteins—KaiA, KaiB, and KaiC—which together form a circadian oscillator. Remarkably, this oscillator can be fully reconstituted in vitro by simply mixing the purified Kai proteins with ATP and magnesium ions, enabling to observe and analyze circadian oscillations outside the cellular environment8,9.
In halotolerant cyanobacteria, salt stress affects the molecular regulation of clock-related proteins and can attenuate circadian rhythmicity10. Building on these observations, the potential correlation between NaCl concentration and the circadian period can be examined using an in vitro system, which allows separation of these effects from other biological processes. In this study, we used in vitro circadian oscillators reconstituted from cyanobacteria to test the influence of salt on both the circadian period and temperature compensation.
Results
NaCl modulates the circadian period
We previously demonstrated that magnesium ions (Mg2+) play a crucial role in regulating the cyanobacterial circadian clock. Mg2⁺ is indispensable for the generation of circadian oscillations, and its absence abolishes rhythmicity11. Moreover, fluctuations in Mg2⁺ concentration modulate the circadian period, underscoring the importance of ionic conditions in clock regulation. Building on these findings, we investigated the influence of NaCl—an essential component of the cellular environment—on circadian period.
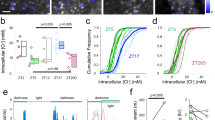
A cyanobacterial circadian oscillator is typically reconstituted with 150 mM NaCl, which supports a robust oscillation of KaiC phosphorylation12. To investigate how salt concentration affects the circadian period, we reconstituted cyanobacterial circadian oscillators under four different NaCl conditions (100, 150, 200, and 250 mM). Across this concentration range, the oscillations remained robust, although the periods displayed slight variations (Fig. 1a). Notably, we observed that increasing NaCl concentration progressively shortened the circadian period, indicating that ionic strength directly modulates the period of the oscillator (Fig. 1b).
| Circadian oscillation of KaiC phosphorylation under varying NaCl and KaiB concentrations. (a), KaiC phosphorylation at four NaCl concentrations (100 mM, green; 150 mM, orange; 200 mM, blue; 250 mM, red). Open circles indicate measured values; lines are for visualization. Two additional replicates are shown in Supplementary Information (Fig. S1). (b), Circadian periods at each NaCl concentration (mean of three independent measurements ± s.e.m.). The linear correlation is described by y = 23.78 – 0.018x (R2 = 0.982, p = 0.01). (c), Oscillation amplitudes at each NaCl concentration (n = 3). The linear correlation is described by y = 28.76 – 0.019x (R2 = 0.586, p = 0.23). (d), KaiC phosphorylation at four KaiB concentrations (1.7 µM, green; 3.4 µM, orange; 6.8 µM, blue; 10.2 µM, red). (e), Circadian periods at different KaiB concentrations (n = 3). The linear correlation is described by y = 21.32 – 0.019x (R2 = 0.946, p = 0.03). (f), Oscillation amplitudes at different KaiB concentrations (n = 3). The linear correlation is described by y = 29.26 – 0.318x (R2 = 0.777, p = 0.12).
To determine which Kai protein contributes to the period changes observed at different NaCl concentrations, we compared the rate constants for phosphorylation (KaiC + KaiA) and dephosphorylation (KaiC alone). Because each experiment is completed in less than 24 h, the differences are small and difficult to visualize on a standard graph (Fig. S2). Therefore, we performed nonlinear regression by fitting the data to the simplified KaiC phosphorylation model previously described by Nakajima et al.8.
The correlations between NaCl concentration and the fitted rate constants were not statistically significant for either phosphorylation (p = 0.61) or dephosphorylation (p = 0.56) (Fig. S2). These results indicate that the NaCl-dependent changes in circadian period are not mediated through KaiA or KaiC directly.
The circadian period of the in vitro oscillator can also be modulated by altering the concentration of Kai proteins, with increases or decreases in their levels leading to corresponding shortening or lengthening of the period12,13. These effects are also reflected in the oscillation amplitude, which shows strong dependence on KaiA and KaiC concentrations. In contrast, KaiB concentration has minimal impact on amplitude. To further explore this relationship, we measured oscillation amplitudes under varying NaCl concentrations and found no significant changes, remaining within a narrow range of 23% to 28% (Fig. 1c).
Since the oscillation pattern under varying NaCl concentrations resembled the behavior observed when altering KaiB levels13, we next examined the circadian period and amplitude by systematically changing KaiB concentrations in the in vitro oscillator. Oscillations remained robust across the tested range of KaiB concentrations (Fig. 1d). Similar to the salt concentration experiments, increasing KaiB concentration progressively shortened the circadian period (Fig. 1e), while oscillation amplitudes remained relatively stable, ranging from 25 to 29% (Fig. 1f). Thus, both NaCl and KaiB exert similar effects on the circadian oscillation, primarily modulating period length without substantially affecting amplitude.
Temperature compensation remained stable across the different KaiB concentrations
KaiB plays a central role in regulating KaiC phosphorylation, enabling the generation of circadian oscillations. Its functional dynamics depend on oligomeric state transitions, shifting between tetramers and monomers. The monomeric KaiB exists in two conformations: the ground-state form (gs-KaiB), which remains unbound, and the fold-switched form (fs-KaiB), which binds to KaiC14. The equilibrium between these oligomeric and conformational states is critical for defining the physical properties of the circadian oscillation. Recently, KaiB has also been implicated in the mechanism of temperature compensation15. To investigate how KaiB concentration influences temperature compensation, we reconstituted in vitro circadian oscillators across a range of KaiB concentrations and measured their periods at two different temperatures (25 °C and 35 °C).
The circadian oscillations remained robust across the tested KaiB concentration range at 35 °C (Fig. 2a). At 25 °C, the oscillation amplitude decreased somewhat but still exhibited clear circadian rhythmicity (Fig. 2b). The circadian period shortened with increasing KaiB concentrations at 25 °C, whereas at 35 °C this trend showed slight perturbation when excess KaiB was added, though the changes remained within the experimental error range (Fig. 2c). Overall, variation in temperature had only minimal impact on the circadian period over the physiological temperature range. To evaluate temperature compensation, we calculated the Q₁₀ values of circadian frequency, which were essentially conserved within experimental error range (Fig. 2d). Therefore, temperature compensation was maintained across the different KaiB concentrations.
Temperature compensation of circadian oscillations under varying KaiB concentrations. (a), KaiC phosphorylation at 35 °C with four KaiB concentrations (1.7 µM, green; 3.4 µM, orange; 6.8 µM, blue; 10.2 µM, red). Closed circles represent measured values, and dotted lines are drawn for visualization. Two additional replicates are provided in the Supplementary Information (Fig. S3). (b), KaiC phosphorylation at 25 °C with the same four KaiB concentrations. Open circles represent measured values, and solid lines are drawn for visualization. Two additional replicates are provided in the Supplementary Information. (c), Circadian periods at each KaiB concentration and temperature (25 °C, open circles; 35 °C, closed circles). The linear correlation at 25 °C is described by y = 21.30 – 0.129x (R2 = 0.871, p = 0.07) and at 35 °C by y = 21.11 – 0.101x (R2 = 0.551, p = 0.26). Data represent the mean of three independent measurements ± s.e.m. (d), Temperature compensation of circadian oscillations expressed as Q₁₀ values of circadian frequency (n = 3). The linear correlation is described by y = 1.0088 – 0.0016x (R2 = 0.136, p = 0.63).
NaCl modulates the temperature compensation.
To investigate the effect of NaCl on temperature compensation, we reconstituted four in vitro circadian oscillators with different NaCl concentrations. These oscillators exhibited robust circadian rhythms across the tested NaCl range at 35 °C (Fig. 3a). At 25 °C, the oscillations showed the expected reduction in amplitude but remained rhythmic (Fig. 3b). Analysis of the periods revealed a nearly linear correlation with NaCl concentration at both temperatures (Fig. 3c). Interestingly, the slope of this correlation was steeper at 35 °C than at 25 °C. Unlike the case of varying KaiB concentrations, period variation was more pronounced when NaCl deviated from 150 mM, a condition under which the period became temperature-independent. The calculated Q10 values displayed a clear linear correlation with NaCl concentration, although the average values were not significantly changed (Fig. 3d). Taken together, these results suggest that the in vitro oscillator maintains temperature compensation across a range of NaCl concentrations, but the Q10 values can be perturbed, showing a positive linear dependence on NaCl concentration.
Temperature compensation of circadian oscillations under varying NaCl concentrations. (a), KaiC phosphorylation at 25 °C with four NaCl concentrations (100 mM, green; 150 mM, orange; 200 mM, blue; 250 mM, red). Open circles represent measured values, with solid lines drawn for visualization. Two additional replicates are provided in the Supplementary Information (Fig. S4). (b), KaiC phosphorylation at 35 °C with the same four NaCl concentrations. Closed circles represent measured values, with dotted lines drawn for visualization. Two additional replicates are provided in the Supplementary Information. (c), Circadian periods at each NaCl concentration and temperature (25 °C, open circles; 35 °C, closed circles). The linear correlation at 25 °C is described by y = 21.88–0.0074x (R2 = 0.984, p = 0.01) and at 35 °C by y = 25.24 – 0.0256x (R2 = 0.995, p = 0.002). Data represent the mean of three independent measurements ± s.e.m. (d), Temperature compensation of circadian oscillations, expressed as Q₁₀ values of circadian frequency (n = 3). The linear correlation is described by y = 1.158 – 0.00083x (R2 = 0.992, p = 0.004).
Discussion
The mechanism by which the circadian period is generated in the in vitro oscillator, a simplified model system of the cyanobacterial clock, has been studied extensively11,14,16,17,18. KaiB plays a central role in regulating KaiC phosphorylation and thereby sustaining circadian rhythm generation. A key feature of this process is the conformational transition between gs-KaiB and fs-KaiB. While gs-KaiB predominantly assembles into tetramers and is generally regarded as a non-functional conformation, fs-KaiB adopts a structurally distinct, functional state that specifically recognizes and binds fully phosphorylated KaiC14. This interaction suppresses KaiC’s autokinase activity, which is stimulated by KaiA16, and enhances its autophosphatase activity19, a pivotal reaction necessary to reset the dephosphorylation cycle and sustain circadian rhythm generation. The equilibrium between these two states is primarily governed by proline isomerization20 and the balance between tetrameric and monomeric forms of KaiB21,22.
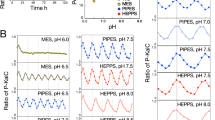
Because free fs-KaiB is scarce in the in vitro oscillator14, it is technically challenging to measure equilibrium shifts with such a small amount using current methods. To investigate the influence of NaCl on the equilibrium of different KaiB forms, we cross-linked KaiB under varying NaCl concentrations. Although this approach does not fully resolve the tetramer–monomer equilibrium23, our results showed that increasing NaCl concentration promoted tetramer formation (Fig. S5). Salt concentration can also affect the equilibrium between fold-switch states24, although this has not been studied in the case of KaiB. Thus, altering NaCl levels perturbed the equilibrium and ultimately led to changes in the circadian period. Surprisingly, NaCl also influenced temperature compensation. Because the KaiB equilibrium is tightly regulated by temperature25, it likely buffers the abundance of fs-KaiB across physiological temperature ranges, thereby stabilizing the circadian period (Fig. 4). When this equilibrium was perturbed by a non-thermal factor such as salt, we observed partial disruption of temperature compensation, although the effect was relatively modest. Identifying additional cofactors that exert a stronger influence on this equilibrium may provide deeper insight into the underlying mechanism of temperature compensation.
Hypothetical model of KaiB equilibrium. At low temperature, fold-switched (fs-KaiB, PDBID: 5jwo) and tetrameric KaiB (PDBID: 4kso) are favored, whereas at high temperature, ground-state (gs-KaiB, PDBID: 4kso) predominates. This equilibrium may regulate the circadian period and contribute to temperature compensation.
Variations in physiological conditions—particularly intracellular salt concentration—can shift the circadian period, producing non-resonant rhythms that entrain to the Earth’s day/night cycle with suboptimal phase angles. Previous studies have shown that cyanobacteria with such non-resonant rhythms are at a selective disadvantage in competitive environments due to temporal misalignment, whereas those possessing resonant rhythms persist more effectively26,27,28. This evolutionary pressure may drive diversification of circadian clock components, particularly protein sequences, as organisms adapt their rhythms to fluctuating environmental conditions29. Consequently, the distinct oscillatory properties observed under a single buffer condition in an in vitro oscillator may not faithfully represent the true circadian dynamics across different species. Taken together, the results demonstrate that accurate interpretation of circadian rhythm properties requires careful consideration of species-specific physiological conditions.
Methods
Purification of KaiA and KaiB proteins
KaiA or KaiB sequence were cloned into the pET28a( +) vector. A small ubiquitin-like modifier (SUMO) tag and ubiquitin-like protease 1 (Ulp1) cleavage site were introduced at the N-terminus of the Kai protein sequences30. E. coli BL21(DE3) cells (Sigma-Aldrich) were inoculated into 1 L of LB medium and grown at 37 °C. Protein expression was induced by adding 1 mL of 1 M Isopropyl β-D-1-thiogalactopyranoside (IPTG) stock solution to the culture at an OD₆₀₀ of ~ 0.7. After 6 h of expression, cells were harvested by centrifugation at 3,200 × g for 20 min.
The cell pellet was washed and resuspended in Ni column binding buffer (500 mM NaCl, 20 mM Tris, 5 mM imidazole, pH 7.0). Cells were lysed using a high-pressure homogenizer, and the lysate was clarified by centrifugation at 20,000 × g for 1 h. The supernatant was loaded onto a Ni column at a flow rate of 2 mL/min and washed with 4 column volumes of binding buffer. Bound proteins were eluted using a linear gradient of elution buffer (500 mM NaCl, 20 mM Tris, 500 mM imidazole, pH 7.0).
Eluted Kai protein fractions were desalted into anion exchange binding buffer (50 mM NaCl, 20 mM Tris, pH 7.0) and applied to an anion exchange column for further purification. Proteins were eluted with a linear gradient of elution buffer (1 M NaCl, 20 mM Tris, pH 7.0). Fractions containing Kai proteins were incubated overnight at 4 °C with Ulp1 protease to remove the His-SUMO tag. The cleaved tag was separated by passing the sample through a Ni column, and remaining impurities were removed with an additional round of anion exchange chromatography. The buffer of purified Kai proteins was exchanged into reaction buffer (150 mM NaCl, 20 mM Tris, 5 mM MgCl₂, 0.5 mM EDTA, 1 mM ATP, pH 8.0). Protein concentration was determined by Bradford assay, and samples were stored at –80 °C for later use.
Purification of KaiC proteins
The KaiC sequence was cloned into the pET41a( +) protein expression vector. An human rhinovirus 3C (HRV 3C) protease cleavage site was introduced at the N-terminus of KaiC to allow removal of the glutathione S-transferase (GST) tag from the purified protein. Expression of KaiC was carried out similarly to KaiA, except that the temperature was reduced to 25 °C upon induction with IPTG (0.2 mL of 1 M IPTG stock)31. After overnight expression at 25 °C, cells were harvested under the same conditions as KaiA. The cell pellet was resuspended in GST binding buffer (150 mM NaCl, 50 mM Tris, 5 mM MgCl₂, 0.5 mM EDTA, 5 mM ATP, pH 7.3) and lysed using the same method as for KaiA.
For purification, the clarified supernatant was loaded onto a GST column at a flow rate of 1 mL/min at 30 °C to minimize non-specific binding of endogenous DnaK. The column was washed with 20 column volumes of GST binding buffer to remove endogenous proteases and ATPases from E. coli. KaiC was then eluted with GST elution buffer (150 mM NaCl, 50 mM Tris, 5 mM MgCl₂, 0.5 mM EDTA, 1 mM ATP, 10 mM glutathione, pH 7.3). The eluted fraction was incubated overnight at 4 °C with HRV 3C protease to cleave the GST tag.
The next day, buffer exchange was performed to remove glutathione, and the sample was reapplied to the GST column to separate the free GST tag. The eluted KaiC fraction was buffer-exchanged into reaction buffer. Protein concentration was determined using the Bradford assay, and samples were stored at –80 °C for later use.
Preparation of in vitro oscillation reaction mixtures
In vitro oscillation assays were performed using purified KaiA, KaiB, and KaiC proteins. Oscillation reactions contained KaiC (3.4 µM), KaiA (1.2 µM), KaiB (3.4 µM) in the reaction buffer. Reaction mixtures were incubated at 30 °C in sealed glass vials without agitation. NaCl and KaiB concentrations were varied (NaCl: 100, 150, 200, and 250 mM, KaiB: 1.7, 3.4, 6.8, and 10,2 µM). For temperature-compensation experiments, reactions were incubated at two different temperatures (25 °C and 35 °C) using the same stock of reaction mixture.
After temperature equilibration (5–10 min), aliquots (20 µL) were withdrawn every 2 h for 48 h using a sample changer (Gilson). Samples were quenched with sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) loading buffer and stored at –20 °C to preserve KaiC phosphorylation states. Protein phosphorylation dynamics were analyzed by SDS–PAGE followed by Coomassie staining, and band intensities were quantified by densitometry using ImageJ (NIH)32.
Estimation of circadian periods and amplitudes
Circadian oscillation parameters were quantified from KaiC phosphorylation data using the Fast Fourier Transform–Nonlinear Least Squares (FFT–NLLS) method implemented in Bodare233. Densitometry values obtained from SDS–PAGE were normalized to total KaiC intensity to correct for gel loading variation. The normalized phosphorylation percentage was then used as the input dataset for analysis. The FFT component was first applied to identify dominant frequencies in the dataset and provide initial estimates of the circadian period. These estimates were subsequently refined by nonlinear least-squares curve fitting to a cosine wave model, allowing more accurate determination of period length and amplitude. Amplitude was defined as half the peak-to-trough difference of the fitted oscillatory curve. Three replicate datasets were analyzed independently, and the mean ± standard error of mean for period and amplitude were calculated with the equations.
Estimation of the apparent rate constants of KaiC phosphorylation and dephosphorylation
The apparent rate constants of KaiC phosphorylation and dephosphorylation were determined by fitting the data to previously published rate Eqs. 8. For the dephosphorylation assay, the “KaiC alone” sample was incubated at 30 °C without magnesium chloride to reach maximal phosphorylation11. For the phosphorylation assay, the “KaiC + KaiA” sample was dephosphorylated by incubating at 30 °C in the absence of KaiA to reach minimal phosphorylation. Prior to collecting the time 0 sample, magnesium chloride was added to the “KaiC alone” sample, and KaiA was added to the “KaiC + KaiA” sample to initiate dephosphorylation and phosphorylation, respectively.
Phosphorylation and dephosphorylation data collected in the presence or absence of KaiA, respectively, were analyzed using nonlinear least-squares regression to compare reaction rate constants across different NaCl concentrations. Phosphorylation levels of KaiC (dependent variable) were fit to a first-order exponential model of the form
In this Eq. 8, the fitted parameters correspond to key physical characteristics: Pequi represents the final asymptotic value (the saturation level approached at long times), P(0) is the initial phosphorylation level (the value at t = 0), and k1 + k2 is the first-order rate constant (h⁻1) governing the transition between phosphorylation and dephosphorylation.
Curve fitting was performed using a computational programming environment (e.g., Python’s scipy.optimize.curve_fit via Gemini), which minimizes the sum of squared residuals (χ2) between the observed data and model-predicted values, yielding optimal estimates for Pequi, P(0), and k1 + k2.
Zero-length cross-linking with 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC)
KaiB cross-linking experiments were performed using EDC as a zero-length cross-linker. Purified KaiB was exchanged into phosphate buffer (25 mM sodium phosphate, 20–2000 mM NaCl, pH 7.2) to remove residual Tris that could interfere with cross-linking. Reactions were initiated by mixing KaiB (10 µM) with freshly prepared EDC (2 mM). Samples were incubated at room temperature for 10 min with gentle mixing. The reaction was quenched by addition of 50 mM Tris–HCl (pH 7.5). Cross-linking was analyzed by SDS–PAGE under denaturing conditions. Protein bands corresponding to KaiB monomer, dimer, and higher-order oligomers were visualized by Coomassie staining and quantified using densitometry (ImageJ, NIH)32.
Calculation of Q10 values
Temperature compensation of the in vitro oscillator was evaluated by calculating Q₁₀ values from the frequency of KaiC phosphorylation rhythms measured at 25 °C and 35 °C. Q₁₀ was then calculated using the standard equation:
where f1 and f2 represent the oscillation frequencies at 25 °C (T1) and 35 °C (T2), respectively.
Statistics
Error bars shown in the figures represent the standard error of the mean (SEM), which was calculated using the standard error equation:
where σ is the standard deviation and n is the sample size.
The linear regression parameters—the slope (m) and the y-intercept (b)—were estimated using the Ordinary Least Squares (OLS) method, fitting the data to the linear equation y = mx + b. To assess the statistical significance of the linear relationship, a two-tailed t-test was performed on the slope coefficient using Gemini. The t-statistic was calculated as the ratio of the estimated slope to its standard error (t = m / SEm). The corresponding p-value was then derived from the t-distribution with N-2 degrees of freedom (N = 4). Statistical significance for the overall model was defined as a p-value less than 0.05.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Wu, Q., Burley, G., Li, L.-C., Lin, S. & Shi, Y.-C. The role of dietary salt in metabolism and energy balance: Insights beyond cardiovascular disease. Diabetes Obes. Metab. 25, 1147–1161 (2023).
Gizowski, C. & Bourque, C. W. Sodium regulates clock time and output via an excitatory GABAergic pathway. Nature 583, 421–424 (2020).
Kimura, G., Dohi, Y. & Fukuda, M. Salt sensitivity and circadian rhythm of blood pressure: The keys to connect CKD with cardiovasucular events. Hypertens. Res. 33, 515–520 (2010).
Nikolaeva, S. et al. The circadian clock modulates renal sodium handling. J. Am. Soc. Nephrol. JASN 23, 1019–1026 (2012).
Patke, A., Young, M. W. & Axelrod, S. Molecular mechanisms and physiological importance of circadian rhythms. Nat. Rev. Mol. Cell Biol. 21, 67–84 (2020).
Sorek, M. & Levy, O. The effect of temperature compensation on the circadian rhythmicity of photosynthesis in Symbiodinium, coral-symbiotic alga. Sci. Rep. 2, 536 (2012).
Fang, M., LiWang, A., Golden, S. S. & Partch, C. L. The inner workings of an ancient biological clock. Trends Biochem. Sci. 49, 236–246 (2024).
Nakajima, M. et al. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415 (2005).
Chavan, A. G. et al. Reconstitution of an intact clock reveals mechanisms of circadian timekeeping. Science 374, eabd4453 (2021).
Waditee-Sirisattha, R. & Kageyama, H. Halotolerance, stress mechanisms, and circadian clock of salt-tolerant cyanobacteria. Appl. Microbiol. Biotechnol. 107, 1129–1141 (2023).
Jeong, Y. M. et al. Magnesium regulates the circadian oscillator in Cyanobacteria. J. Biol. Rhythms 34, 380–390 (2019).
Kageyama, H. et al. Cyanobacterial circadian pacemaker: Kai protein complex dynamics in the KaiC phosphorylation cycle in vitro. Mol. Cell 23, 161–171 (2006).
Nakajima, M., Ito, H. & Kondo, T. In vitro regulation of circadian phosphorylation rhythm of cyanobacterial clock protein KaiC by KaiA and KaiB. FEBS Lett. 584, 898–902 (2010).
Chang, Y.-G. et al. A protein fold switch joins the circadian oscillator to clock output in cyanobacteria. Science 349, 324–328 (2015).
Zhang, N. et al. Temperature-dependent fold-switching mechanism of the circadian clock protein KaiB. Proc. Natl. Acad. Sci. 121, e2412327121 (2024).
Kim, Y.-I., Dong, G., Carruthers, C. W. J., Golden, S. S. & LiWang, A. The day/night switch in KaiC, a central oscillator component of the circadian clock of cyanobacteria. Proc. Natl. Acad. Sci. U. S. A. 105, 12825–12830 (2008).
Nishiwaki, T. et al. A sequential program of dual phosphorylation of KaiC as a basis for circadian rhythm in cyanobacteria. EMBO J. 26, 4029–4037 (2007).
Rust, M. J., Markson, J. S., Lane, W. S., Fisher, D. S. & O’Shea, E. K. Ordered phosphorylation governs oscillation of a three-protein circadian clock. Science 318, 809–812 (2007).
Kitayama, Y., Iwasaki, H., Nishiwaki, T. & Kondo, T. KaiB functions as an attenuator of KaiC phosphorylation in the cyanobacterial circadian clock system. EMBO J. 22, 2127–2134 (2003).
Wayment-Steele, H. K. et al. The conformational landscape of fold-switcher KaiB is tuned to the circadian rhythm timescale. Proc. Natl. Acad. Sci. U. S. A. 121, e2412293121 (2024).
Hitomi, K., Oyama, T., Han, S., Arvai, A. S. & Getzoff, E. D. Tetrameric architecture of the circadian clock protein KaiB: A novel interface for intermolecular interactions and its impact on the circadian rhythm. J. Biol. Chem. 280, 19127–19135 (2005).
Murakami, R. et al. The roles of the dimeric and tetrameric structures of the clock protein KaiB in the generation of circadian oscillations in cyanobacteria. J. Biol. Chem. 287, 29506–29515 (2012).
Iwase, R. et al. Functionally important substructures of circadian clock protein KaiB in a unique tetramer complex. J. Biol. Chem. 280, 43141–43149 (2005).
Tuinstra, R. L. et al. Interconversion between two unrelated protein folds in the lymphotactin native state. Proc. Natl. Acad. Sci. 105, 5057–5062 (2008).
LiWang, A. & Orban, J. Unveiling the cold reality of metamorphic proteins. Proc. Natl. Acad. Sci. 122, e2422725122 (2025).
Ouyang, Y., Andersson, C. R., Kondo, T., Golden, S. S. & Johnson, C. H. Resonating circadian clocks enhance fitness in cyanobacteria. Proc. Natl. Acad. Sci. U. S. A. 95, 8660–8664 (1998).
Woelfle, M. A., Ouyang, Y., Phanvijhitsiri, K. & Johnson, C. H. The adaptive value of circadian clocks: An experimental assessment in cyanobacteria. Curr. Biol. 14, 1481–1486 (2004).
Kawamoto, N., Nakanishi, S. & Shimakawa, G. Light wavelength as a contributory factor of environmental fitness in the cyanobacterial circadian clock. Plant Cell Physiol. 65, 798–808 (2024).
Mukaiyama, A. et al. Evolutionary origins of self-sustained Kai protein circadian oscillators in cyanobacteria. Nat. Commun. 16, 4541 (2025).
Kim, Y.-I., Boyd, J. S., Espinosa, J. & Golden, S. S. Detecting KaiC phosphorylation rhythms of the cyanobacterial circadian oscillator in vitro and in vivo. Methods Enzymol. 551, 153–173 (2015).
Kim, P., Kaszuba, A., Jang, H.-I. & Kim, Y.-I. Purification of GST-fused cyanobacterial central oscillator protein KaiC. Appl. Biochem. Microbiol. 56, 395–399 (2020).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Zieliński, T., Hay, J. & Millar, A. J. Period estimation and rhythm detection in timeseries data using BioDare2, the free, online, community resource. Methods Mol. Biol. Clifton N. J. 2398, 15–32 (2022).
Acknowledgements
This research was made possible by the NASA Established Program to Stimulate Competitive Research, Grant # 80NSSC22M0027.
Funding
NASA Established Program to Stimulate Competitive Research, 80NSSC22M0027
Author information
Authors and Affiliations
Contributions
E.K. and Y.K. designed the research; All authors performed the research and analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, E., Adams, M., Tyree, S. et al. Effects of sodium chloride on circadian period and temperature compensation of KaiC phosphorylation. Sci Rep 16, 10319 (2026). https://doi.org/10.1038/s41598-026-40224-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-40224-z