Abstract
Endophytes are microorganisms that colonize plants, often via commensal or symbiotic associations, and regulate plant growth and metabolism. Such organisms are usually suitable alternatives to therapeuticallly relevant, bioactive, and commercially essential metabolites, particularly under optimized bioprocess conditions. The current study highlights the key research challenge of maximizing the production of Indole-3-Acetic Acid (IAA), a compound difficult to isolate from the plant due to low yield, by elucidating the genetic composition of an isolated endophyte and determining the biosynthetic pathway using the KEGG pathway. Moreover, deciphering the functional genomic and refining production optimization remain significant challenges. The whole-genome sequencing of the endophytic bacterium Bacillus cereus SKAM2 (Strain SKAM2) revealed a genome size of 5.6 Mb, a GC content of 36%, multiple tryptophan-dependent and tryptophan-independent pathways. Furthermore, the IAA biosynthetic pathways pave the way for process-optimization studies. The influence of various abiotic parameters and media supplements on IAA production in both intra- and extracellular media was compared, using a full-factorial design of experiments (DOE). The results showed the highest yield in the extracellular fraction, a 3.81-fold increase, exceeding the intracellular IAA yield. The results highlight the strong potential of strain SKAM2 as a microbial platform for sustainable IAA production.
Similar content being viewed by others
Introduction
The endophytes are microbes that establish symbiotic or commensal relationships with plants, providing a range of benefits to their host plants, such as enhanced defense mechanisms, increased resilience to stress, and improved overall physiological health1. The phytohormones play a significant role in plant growth and metabolism2. One of the most ubiquitous phytohormones, IAA (auxin), stimulates growth processes, tissue development, cell elongation, proliferation, and specialization in response to environmental signals3,4,5,6. The global market indicates a rising demand for IAA, which is expected to reach USD 36 million by 20287. In addition, earlier research has examined the therapeutic applications of IAA, particularly its anti-inflammatory effects8, hepatoprotective9, anticancer10, and the efficacy of chemotherapy in pancreatic cancer11.
The IAA production has been reported in various microorganisms, including actinomycetes, fungi, bacteria, and yeasts12,13,14,15,16. Aligned with sustainable development, IAA-producing microbial strains have attracted considerable attention as potential strains for providing cost-effective, eco-friendly, and large-scale biosynthesis of IAA12,17,18. Despite numerous studies on endophytic bacteria, IAA production19. Understanding functional genomics and optimizing IAA production strategies remains challenging. In addition, whole-genome analysis of endophytic bacteria facilitates the identification of genes underlying IAA production and reveals insights into their molecular and functional mechanism20. In this study, the genome sequence of an isolated endophytic bacterium, Strain Bacillus cereus SKAM2, associated with the medicinal plant Humulus lupulus, was analyzed to identify genes and pathways involved in IAA biosynthesis. Furthermore, the present study investigates the optimization of intra- and extracellular IAA production by the strain SKAM2 using a full-factorial DOE with various abiotic factors. Several studies have aimed to optimize microbial IAA production21,22,23,24; however, only a few have employed a full-factorial DOE to evaluate the influencing factors for intra- and extracellular IAA production. To the best of our knowledge, such studies are scarce in the current literature review to combine genome-level insights with a design-of-experiments approach to maximize IAA yield.
Result
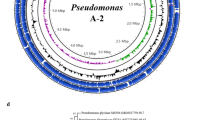
The genome of strain SKAM2 revealed a size of 5.6 Mb, with an average GC content of 36%. The genome contains a circular topology, a chromosome size (bp) of 1 main chromosome (~ 5.5 Mbp), 90 tRNAs, 4 rRNAs, and 5836 protein-coding genes (CDS). The genome sequencing data of endophytic strain SKAM2, deposited under the accession number JBMFCI000000000.1 and the BioProject: PRJNA1237547. Figure 1 describes the genomic properties of the strain SKAM2.
Circular map of the strain SKAM2. Showing the coding sequence gene, transfer RNA (tRNA), ribosomal RNA (rRNA), transfer messenger RNA (tmRNA), guanine-cytosine (GC Content), and GC Skew.
Comparative genomics analysis of the SKAM2 strain
The genomic relatedness of strain SKAM2 was investigated by calculating pairwise Average Nucleotide Identity (ANI) values using the ANIm algorithm using JSpecies v1.5. SKAM2 shared ANI values exceeding 98.45% with five other Bacillus cereus genomes: M72-4 (CP119300.1), QKG-2024 (NZ_CP168976.1), DLOU-Tangshan (CP040340.1), ATCC 14,579 (JBJJSR010000003.1), and CPT56D-587-MTF (CP090081.1). Figure 2 represents the heatmap of ANI between SKAM2 and its closest relatives.
Pairwise ANI heatmap between strain SKAM2 and its closest relatives.
The antiSMASH was employed to investigate and predict secondary metabolite gene clusters. Table 1 describes the prediction and analysis of secondary metabolite BGCs. The findings indicated that the SKAM2 strain encodes multiple gene clusters involved in the synthesis of secondary metabolites. Among them are four non-ribosomal peptide synthases (NRPSs), one betalactone (betalactone-type fengycin), two terpenes, one NI-siderophore (petrobactin), and one RiPP-like (cerecyclin). Compared with known biosynthetic gene clusters related to petrobactin, cerecyclin, and bacillibactin, those in this study exhibited high similarity. In contrast, clusters associated with molybdenum cofactor and fengycin showed low similarity.
Additionally, the COG function analysis investigation detected 5,312 proteins in the genome that matched entries in the eggNOG database. Notably, 304 genes, accounting for 5.72% of the total, were associated with cell wall and membrane functions. A total of 493 genes, representing 9.28% of the genome, were linked to amino acid transport and metabolism, and 5.91% (314) of the genes were grouped into ion transport and metabolism. In addition, 5.69% (302) of carbohydrate metabolism genes were found in strain SKAM2. Figure 3 represents the COG function analysis of the genome. Furthermore, the strain SKAM2 reveals that the IAA biosynthesis and tryptophan metabolism genes were detected through the KEGG pathway (Figure S1). Table 2 describes the identified genes involved in IAA biosynthesis and tryptophan metabolism.
The COG classification of the genome strain SKAM2.
Optimization of IAA yield from intra- and extracellular sources via full factorial DOE
The investigation of the influence of different factors and culture conditions on IAA production was examined. Table 3: Describe the yield of IAA achieved in various factors. In addition, MINITAB® statistical tools results reveal that the main effects plot (Fig. 4), interaction plot (Fig. 5), Pareto chart (Fig. 6), and contour diagram (Fig. 7) were generated using MINITAB® to assess the relationships between variable effects, both intra- and extracellular IAA yield. Additionally, kinetic models for IAA yield in both intra- and extracellular IAA production processes were illustrated in Eqs. 1 and 2. Furthermore, analysis of variance (ANOVA) was used to determine the relative contribution and significance of each experimental factor governing IAA yield. The results, summarized in Table 4, include the sum of squares, mean square values, and significance levels (p-values) for both extracellular and intracellular IAA production.
Show the main effect plot, (A) extracellular and (B) intracellular IAA yield.
Shows an interaction plot for studying factors (A) extracellular and (B) intracellular IAA yield.
Illustrates the relative significance of various factors using a Pareto chart: (A) extracellular and (B) intracellular IAA yield.
Show the contour plots (A) of extracellular and (B) intracellular IAA yield.
Validation of the DOE model
The validation of the DOE model was conducted under optimized conditions predicted by the model. The validation run was performed using cultures grown under tryptophan-supplemented conditions, with static conditions, and incubated for 5 days (Table 3, run number 3), then compared with cultures grown under the same model conditions using the predicted model values. The calculated yield of the model was 0.67 mg/mL for extracellular and 0.25 mg/mL for intracellular. The experimental yield was achieved extracellular at 0.70 mg/mL and intracellular at 0.25 mg/mL. The experimental observation and predicted yields demonstrated close correspondence, confirming the model’s predictive accuracy under the optimized conditions. Figure 8 demonstrates the UPLC chromatograms corresponding to the validation runs for IAA optimization. The kinetic models for the validated model for extracellular and intracellular IAA production were represented by Eqs. 3 and 4.
UPLC chromatogram of the validation run of (A) extracellular and (B) intracellular IAA.
Discussion
The endophytic microorganisms isolated from plants are well known for conferring benefits to the host plant, ranging from growth to metabolite production25. Bacillus cereus, a scarcely identified endophyte, has garnered increasing attention for its genomic features and potential applications in sustainable agriculture and therapeutic development26. The genomic exploration of Bacillus strains demonstrated considerable biotechnological and pharmaceutical potential, offering a valuable avenue for elucidating the molecular mechanisms underlying secondary metabolite biosynthesis27. Microbial genome size and GC content act as crucial factors in shaping their adaptability to diverse environmental conditions28. Microbial strains possessing larger genome sizes are often better adapted to complex environments, potentially enabled by their capacity to encode a broader array of metabolic functions and stress-response mechanisms29. However, numerous studies have proposed that smaller bacterial genomes might offer a competitive edge, offering benefits in terms of energy efficiency and reproductive rate30,31. The current study highlighted the genome size of strain SKAM2 to be 5.6 Mb, with a GC content of 36%. The variations in the genome size of Bacillus cereus strains have been reported, ranging from 5.39 to 6.40, with similar GC content, exhibiting possible conservation of genetic composition among the genera26. Furthermore, the ANI quantifies the average level of nucleotide identity between matching genomic fragments from microorganisms. The ANI serves as a standard tool for distinguishing closely related species, owing to its high discriminatory power, with values ≥ 95% indicating the same species32,33. The ANI result of strain SKAM2 showed the highest ANI with Bacillus cereus strain M72-4 (98.59%) and Bacillus cereus strain QKG-2024 (98.58%).
In addition, genome mining was performed using the antiSMASH tools that uncovered a diverse array of biosynthetic gene clusters (BGCs)34,35 that revealed the secondary metabolic richness of strain SKAM2. The AntiSMASH analysis highlighted thirteen BGC-associated pathways. Among them, six gene clusters encoding unknown secondary metabolites were identified, including three RiPP-like clusters, NI Siderophores, and one NRPS-type cluster, indicating that the secondary metabolite profile of this strain is distinct from other reported Bacillus cereus genomes36,37. The applications of these metabolites, such as siderophores, indicate a high recurrence of plant-associated components38. The detection of RiPP-like clusters reveals the capacity of strain SKAM2 for producing ribosomally-synthesized and post-translationally-modified peptides, a group of metabolites broadly acknowledged for their potent antimicrobial activity and ecological relevance39,40. Together, the genomic insights paved a strong platform for deciphering and validating novel bioactive metabolites.
Next, we focused on identifying genes for IAA biosynthesis and tryptophan metabolism in the genome of strain SKAM2. Earlier investigations have highlighted the catalogues of endophytic microorganisms for diverse IAA biosynthesis pathways41, encompassing both tryptophan-dependent and tryptophan-independent mechanisms42,43. The tryptophan-dependent routes, including the IPyA pathway, tryptamine (TAM) pathway, indole-3-acetamide (IAM) pathway, and the indole-3-acetonitrile (IAN) pathway, were evaluated5,44. The strain SKAM2 harbors multiple genes associated with IAA biosynthesis, including ipdC, which encodes indole-3-pyruvate decarboxylase, a key gene involved in the IPyA pathway. Earlier studies have shown that the transcriptional activity of the ipdC gene correlates with the upregulation or downregulation of IAA production45,46. Further analysis of tryptophan metabolism revealed the presence of a cas operon that encompasses essential biosynthetic and metabolic genes, including trpA, trpB, trpC, trpD2, trpE, and trpF, which are crucial for IAA biosynthesis via tryptophan metabolism47. Previous results reveal that tryptophan serves as the primary precursor for the IAA biosynthetic pathway47. In addition, a tryptophan decarboxylase gene was identified in the genome of strain SKAM2, suggesting potential involvement of the TAM pathway, in which tryptophan is decarboxylated to tryptamine, an intermediate that can be further converted to IAA48. Earlier findings demonstrated that the bacteria possess multiple pathways for IAA biosynthesis49. Collectively, our observations support that the presence of a tryptophan-mediated biosynthetic pathway; the potential existence of tryptophan-independent routes in the endophytic strain SKAM2 cannot be ruled out.
Based on the genomic insights, strain SKAM2 reveals multiple tryptophan-dependent IAA pathways and tryptophan metabolism genes. To validate the genomic predictions and confirm the functional expression of these pathways, further analysis was done using full factorial DOE, which showed both intra- and extracellular IAA production due to the cascades of biosynthetic pathways. The secretion of IAA into the culture broth is rather unique and necessitates the exploration of the detailed mechanisms underlying this secretion process. The main effect (Fig. 4) analysis highlighted the significant effect of IAA yield from different factors. In extracellular IAA production, a distinct negative influence of factor rotation speed on IAA yield. In contrast, factors such as time, tryptophan, and glucose exhibited positive main effects, each contributing to a progressive enhancement in IAA yield. In addition, the main impact of intracellular IAA production factors, such as time and rotation speed, showed less impact on IAA yield. Similar observations have been reported that over-agitation has been shown to impair IAA yield in a wide range of microbial systems50,51. The tryptophan and glucose showed a significant impact on the intracellular production of IAA yield. Based on the main effect of both intra- and extracellular IAA yield, tryptophan was found to have the major impact on IAA yield. These results are consistent with previous studies that reveal tryptophan is a precursor of IAA42,52. Earlier investigations have highlighted that supplementing the medium with glucose enhances the IAA yield53. The Pareto analysis was employed to evaluate statistically significant factors influencing IAA yield. The Pareto chart (Fig. 6) reveals that, in both intra- and extracellular IAA, tryptophan was observed to be higher in both fractions. The assessment of the Pareto graph indicates significant factors required for regulating bacterial growth and IAA production. The Pareto charts deliver a clear visual ranking of effects, and ANOVA offers a more sensitive, model-based evaluation54. This complementary model enhances the accuracy of the experimental conclusions, consistent with previously reported DOE methodologies54. Furthermore, to identify the significant factor, ANOVA was conducted (Table 4), results highlighted that tryptophan had a statistically significant impact (p < 0.05) on both intra- and extracellular IAA yield. Our finding is consistent with earlier reports describing that supplements of tryptophan enhanced the IAA yield7,42,51,52. Furthermore, to validate these observations and the optimised studies, the validation of the DOE model was conducted. The optimization using DOE and validation are crucial steps to verify the predicted model values and the practical applicability of the developed model53,55. The validation result revealed a 5% deviation between the model-predicted (Table 3, run number 3) value and the experimentally validated data (Fig. 8). The validation results reveal a robust concordance between the experimental and predicted models, attesting to the model’s robustness and predictive accuracy.
Our findings suggest that whole-genome sequencing confirmed the presence of genes associated with the tryptophan-dependent IAA biosynthetic pathway, while also indicating the potential involvement of tryptophan-independent routes, forming the basis for design-of-experiments-guided optimization. Guided by this genome-level insight, statistical optimization identified conditions that maximized IAA production in extracellular production. As demonstrated in earlier reports, extracellular IAA production from microbes confers multiple ecological and biotechnological benefits, such as enhancing plant–microbe interactions44, enabling direct root uptake of auxin56, and minimizing feedback inhibition5. Furthermore, extracellular IAA production also enhanced the efficiency of metabolite extraction in downstream processes, aiding the application of agriculture and pharmaceuticals57. To the best of our knowledge, this is one of the few studies where an association with the elucidated metabolic pathways and the IAA biosynthesis has been deciphered. The study not only provides insight into the role of abiotic factors in modulating IAA biosynthesis in both intracellular and extracellular media, but also paves the way for elucidating the detailed mechanisms quantifying IAA in both intracellular and extracellular fractions. The findings demonstrated that quantifying IAA in both extracellular and intracellular fractions resulted in a higher total IAA yield by the strain SKAM2.
Conclusion
The isolated endophytic bacterial strain SKAM2 has been identified as a potential source of IAA, and, like many other well-explored endophytes, exhibits evidence of tryptophan-dependent pathways. The culture may also be adapted for tryptophan-independent IAA production, as analyzed in the DOE, where the absence of tryptophan was shown to affect IAA production. The results further suggest that interactions among abiotic factors modulate IAA yield. However, the role of interaction in the transcriptional regulation of IAA biosynthesis requires an in-depth analysis. The cumulative production of IAA in both intracellular and extracellular environments synergistically improves the overall yield. However, an interesting aspect of exploration will be the underlying mechanism responsible for the regulated release of IAA into the extracellular media. The study may be further extended to understand the molecular regulation of IAA biosynthesis and optimized production for designing a scale-up process.
Materials and methods
Collection of microorganisms
In our previous studies, our lab isolated a endophytic bacterial strain, Bacillus cereus SKAM2, from Humulus lupulus. The isolation and characterization have been done in our previously published work58. The biochemical and molecular techniques were employed for the identification of bacteria58. and their 16 S rRNA sequence has been submitted to GenBank under the accession number PQ305264. The isolated strain was stored at 4 °C for future use.
Whole genome sequencing of endophytic strain SKAM2
Library preparation and quality assessment
The whole genome sequencing of Bacillus cereus SKAM2 was performed at BioKart India Pvt. Ltd, Bengaluru, India. The Genomic DNA was quantified using the Qubit 3 Fluorometer with the dsDNA High Sensitivity (HS) assay. A total of 100 ng of intact DNA was enzymatically fragmented to generate 200–300 bp fragments. These fragments underwent end repair to convert overhangs into blunt ends. The 3′ to 5′ exonuclease activity removed 3′ overhangs, while polymerase activity filled in 5′ overhangs. Following end repair, a single adenine (A) nucleotide was added to the 3′ ends (adenylation), facilitating the ligation of loop adapters. The adapter-ligated DNA was treated with the uracil-specific excision reagent (USER) enzyme to cleave uracil-containing sites and purified using AMPure XP beads. A paired-end sequencing library was constructed26. The resulting library was then PCR-amplified for 6 cycles using the NEBNext Ultra II Q5 Master Mix with Illumina universal primers and sample-specific indexed (octamer) primers. Post-PCR cleanup was performed again with AMPure XP beads, and the final DNA library was eluted in 15 µL of 0.1× TE buffer. Library quantification was performed using a Qubit 3 Fluorometer with the dsDNA HS reagent. Fragment size distribution and quality were assessed by loading 1 µL of the library onto an Agilent DNA 7500 chip and analyzing it using the Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, California, United States)59. The quality control of raw reads was performed using the “FastQC quality control” tool of Galaxy v.0.7260. The genome of Bacillus cereus ATCC 10,987 was utilized as the reference61. Furthermore, A phylogenetic tree (Figure S2) was constructed using the Molecular Evolutionary Genetics Analysis software (MEGA, version 11)62 using the Neighbor-Joining algorithm with the Maximum Composite Likelihood model. To examine the robustness of each node, 1,000 bootstrap samples were performed63.
Genome component analysis, comparative genomics analysis, and functional annotation of strain SKAM2
The Genomic component analysis was done using PROKKA version 1.14.6 (rapid prokaryotic genome annotation)64. The JSpecies is one of the most popular, utilized, and recognized tool in microbial genomics65. The closely related genomes were obtained from the NCBI database. Genomic ANI was calculated using the NUCmer algorithm via JSpecies v1.5 with default settings47. The prediction and analysis of secondary metabolite biosynthetic gene clusters (BGCs) in the bacterial genome sequences were carried out using the antiSMASH software version 8.0.166. Gene function was studied through the cluster of orthologous groups (COG)67 from the EggNOG mapper Galaxy with default parameters68. Furthermore, the sequencing results were compared with the Kyoto Encyclopedia of Genes and Genomes (KEGG)47. Using the predicted genomic data, a genome map of strain SKAM2 was created using GC Viewer69.
Optimization of the production of IAA by the full factorial DOE
A full factorial DOE was employed to evaluate factor interactions. The analysis was performed using the MINITAB® statistical software to optimize different factors: incubation time, rotation speed, tryptophan, and glucose concentration. A total of 32 replicated experiments were conducted using a two-level factorial design (−1, + 1), defining the minimum and maximum levels of different factors, as presented in Table 5. The experimental run was carried out to assess changes in IAA production in both extracellular and intracellular fractions.
Preparation of the extract for IAA production
The extracellular IAA production was carried out using the strain SKAM2, which was grown in M9 minimal salt medium (1X) (HiMedia, Mumbai, India)58, incubated at 37 °C74, with different culture conditions mentioned in Table 5. The biomass of the culture was extracted by centrifugation, and the resulting supernatant was dried at room temperature. The intracellular IAA production was conducted using the bacterial biomass, which was subjected to sonication using the Sonics Vibra-Cell ultrasonic processor VCX 130 (Sonics & Materials, Inc., CT, USA) at an amplitude of 80 Å for 2 min, employing 10-second on-and-off cycles, using the protocol with modification established by Singh75. The lysate was centrifuged for 10 min at 4 °C to obtain the intracellular supernatant.
Assessment of IAA by ultra-performance liquid chromatography (UPLC)
The quantitative assessment of IAA was conducted using the method with modification described by Szkop and Bielawski76. The extract was dissolved in HPLC-grade methanol and subjected to chromatographic separation at 37 °C. UPLC was performed using a Waters Acquity UPLC (Waters Corporation, Milford, MA, USA) system equipped with a BEH C18 column (100 mm × 2.1 mm, 1.7 μm particle size). The mobile phase consisted of solvent A: B in a 40:60 ratio (0.01% glacial acetic acid in water: acetonitrile). The column temperature was maintained at 35 °C, with a detection wavelength of 265 nm using a PDA eλ detector. The total run time was 5 min, with a 1 µL injection volume and a flow rate of the mobile phase of 01 mL/min. Furthermore, five different solutions of IAA (Central Drug House (P) Ltd) with concentrations from 50 µg/ml to 250 µg/ml were prepared from UPLC to establish the calibration curve. The representative chromatograms and the standard calibration curve are provided in the supplementary file (Figure S3). The resulting chromatographic peak was analyzed and compared with the retention time of the standard IAA obtained from Central Drug House (P) Ltd.
Statistical analysis
All experiments were conducted in duplicate, and the reported data represent the average of the observations.
Data availability
The whole-genome sequencing data of strain *Bacillus cereus* SKAM2 generated in this study have been deposited in the National Center for Biotechnology Information (NCBI) GenBank database under accession number JBMFCI000000000.1 and the BioProject accession number PRJNA1237547.
References
Abo Elsoud, M. M., Hasan, S. F. & Elhateir, M. M. Optimization of Indole-3-acetic acid production by Bacillus velezensis isolated from Pyrus rhizosphere and its effect on plant growth. Biocatal. Agric. Biotechnol. 50, 102714. https://doi.org/10.1016/j.bcab.2023.102714 (2023).
Adeleke, B., Ayangbenro, A. & Babalola, O. Genomic Analysis of Endophytic Bacillus cereus T4S and Its Plant Growth-Promoting Traits. Plants 10 (9), 1776. https://doi.org/10.3390/plants10091776 (2021).
Almirón, C. et al. Functional and genomic analyses of plant growth promoting traits in Priestia aryabhattai and Paenibacillus sp. isolates from tomato rhizosphere. Sci. Rep. 15 (1), 3498. https://doi.org/10.1038/s41598-025-87390-0 (2025).
Arora, P., Tabssum, R., Gupta, A. P., Kumar, S. & Gupta, S. Optimization of indole acetic acid produced by plant growth promoting fungus, aided by response surface methodology. Heliyon 10 (14), e34356. https://doi.org/10.1016/j.heliyon.2024.e34356 (2024).
Benadjila, A., Zamoum, M., Aouar, L., Zitouni, A. & Goudjal, Y. Optimization of cultural conditions using response surface methodology and modeling of indole-3-acetic acid production by Saccharothrix texasensis MB15. Biocatal. Agric. Biotechnol. 39, 102271. https://doi.org/10.1016/j.bcab.2021.102271 (2022).
Blin, K. et al. antiSMASH 8.0: extended gene cluster detection capabilities and analyses of chemistry, enzymology, and regulation. Nucleic Acids Res. 53 (W1), W32–W38. https://doi.org/10.1093/nar/gkaf334 (2025).
Boondaeng, A. et al. Biological Conversion of Agricultural Wastes into Indole-3-acetic Acid by Streptomyces lavenduligriseus BS50-1 Using a Response Surface Methodology (RSM). ACS Omega. 8 (43), 40433–40441. https://doi.org/10.1021/acsomega.3c05004 (2023).
Bunsangiam, S., Thongpae, N., Limtong, S. & Srisuk, N. Large scale production of indole-3-acetic acid and evaluation of the inhibitory effect of indole-3-acetic acid on weed growth. Sci. Rep. 11 (1), 13094. https://doi.org/10.1038/s41598-021-92305-w (2021).
Cen, X., Li, H., Zhang, Y., Huang, L. & Luo, Y. Isolation and Plant Growth Promotion Effect of Endophytic Siderophore-Producing Bacteria: A Study on Halophyte Sesuvium portulacastrum. Plants 13 (19), 2703. https://doi.org/10.3390/plants13192703 (2024).
Chandra, S., Askari, K. & Kumari, M. Optimization of indole acetic acid production by isolated bacteria from Stevia rebaudiana rhizosphere and its effects on plant growth. J. Genetic Eng. Biotechnol. 16 (2), 581–586. https://doi.org/10.1016/j.jgeb.2018.09.001 (2018).
Chaudhary, T., Yadav, D., Chhabra, D., Gera, R. & Shukla, P. Low-cost media engineering for phosphate and IAA production by Kosakonia pseudosacchari TCPS-4 using Multi-objective Genetic Algorithm (MOGA) statistical tool. 3 Biotech. 11 (4), 158. https://doi.org/10.1007/s13205-021-02690-2 (2021).
Cheynier, V., Comte, G., Davies, K. M., Lattanzio, V. & Martens, S. Plant phenolics: Recent advances on their biosynthesis, genetics, and ecophysiology. Plant Physiol. Biochem. 72, 1–20. https://doi.org/10.1016/j.plaphy.2013.05.009 (2013).
Cimermancic, P. et al. Insights into Secondary Metabolism from a Global Analysis of Prokaryotic Biosynthetic Gene Clusters. Cell 158 (2), 412–421. https://doi.org/10.1016/j.cell.2014.06.034 (2014).
Duca, D., Lorv, J., Patten, C. L., Rose, D. & Glick, B. R. Indole-3-acetic acid in plant–microbe interactions. Antonie van Leeuwenhoek. 106 (1), 85–125. https://doi.org/10.1007/s10482-013-0095-y (2014).
Egamberdieva, D., Wirth, S. J., Alqarawi, A. A., Abd_Allah, E. F. & Hashem, A. Phytohormones and Beneficial Microbes: Essential Components for Plants to Balance Stress and Fitness. Front. Microbiol., 8. (2017). https://doi.org/10.3389/fmicb.2017.02104
Elsayed, H. H., Abdallah, N. A. & Amer, S. K. Sustainable bioconversion of rice straw into indole-3-acetic acid by Streptomyces coelicoflavus using response surface methodology. Biomass Convers. Biorefinery. https://doi.org/10.1007/s13399-024-06252-3 (2024).
Etesami, H. & Glick, B. R. Bacterial indole-3-acetic acid: A key regulator for plant growth, plant-microbe interactions, and agricultural adaptive resilience. Microbiol. Res. 281, 127602. https://doi.org/10.1016/j.micres.2024.127602 (2024a).
Etesami, H. & Glick, B. R. Bacterial indole-3-acetic acid: A key regulator for plant growth, plant-microbe interactions, and agricultural adaptive resilience. Microbiol. Res. 281, 127602. https://doi.org/10.1016/j.micres.2024.127602 (2024b).
Fang, Z., Zhang, J., Liu, B., Du, G. & Chen, J. Enhancement of the catalytic efficiency and thermostability of < scp> S tenotrophomonas sp. Microb. Biotechnol. 9 (1), 35–46. https://doi.org/10.1111/1751-7915.12300 (2016). keratinase < scp>KerSMD by domain exchange with < scp>KerSMF
Felsenstein, J. CONFIDENCE LIMITS ON PHYLOGENIES: AN APPROACH USING THE BOOTSTRAP. Evolution 39 (4), 783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x (1985).
Folkes, L. K. & Wardman, P. Oxidative activation of indole-3-acetic acids to cytotoxic species— a potential new role for plant auxins in cancer therapy. Biochem. Pharmacol. 61 (2), 129–136. https://doi.org/10.1016/S0006-2952(00)00498-6 (2001).
Goris, J. et al. DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. MicroBiol. 57 (1), 81–91. https://doi.org/10.1099/ijs.0.64483-0 (2007).
Guo, D., Kong, S., Chu, X., Li, X. & Pan, H. De Novo Biosynthesis of Indole-3-acetic Acid in Engineered Escherichia coli. J. Agric. Food Chem. 67 (29), 8186–8190. https://doi.org/10.1021/acs.jafc.9b02048 (2019).
Hu, J. et al. Genomic and metabolic features of Bacillus cereus, inhibiting the growth of Sclerotinia sclerotiorum by synthesizing secondary metabolites. Arch. Microbiol. 205 (1), 8. https://doi.org/10.1007/s00203-022-03351-5 (2023).
Imada, E. L., Rolla dos Santos, A. A., de Oliveira, P., de Hungria, A. L. M., Rodrigues, E. P. & M., & Indole-3-acetic acid production via the indole-3-pyruvate pathway by plant growth promoter Rhizobium tropici CIAT 899 is strongly inhibited by ammonium. Res. Microbiol. 168 (3), 283–292. https://doi.org/10.1016/j.resmic.2016.10.010 (2017).
Jiang, L. et al. Genome insights into the plant growth-promoting bacterium Saccharibacillus brassicae ATSA2T. AMB Express. 13 (1), 9. https://doi.org/10.1186/s13568-023-01514-1 (2023).
Kautsar, S. A. et al. MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res. (2019) https://doi.org/10.1093/nar/gkz882.
Khalifa, A. & Alsowayeh, N. Whole-Genome Sequence Insight into the Plant-Growth-Promoting Bacterium Priestia filamentosa Strain AZC66 Obtained from Zygophyllum coccineum Rhizosphere. Plants, 12(10), 1944. (2023). https://doi.org/10.3390/plants12101944
Khan, S. & Mathur, A. LC–MS and GC-MS-based bioactive metabolites profiling of endophytic bacterium from Humulus lupulus and production of Indole-3-acetic acid. Curr. Trends Biotechnol. Pharm. 19 (3), 2422–2432. https://doi.org/10.5530/ctbp.2025.3.27 (2025).
Khan, S., Mathur, A. & Khan, F. Endophytic fungi-bioinspired nanoparticles potential to control infectious disease. Crit. Rev. Microbiol. 1–23. https://doi.org/10.1080/1040841X.2025.2497795 (2025).
Koskiniemi, S., Sun, S., Berg, O. G. & Andersson, D. I. Selection-Driven Gene Loss in Bacteria. PLoS Genet. 8 (6), e1002787. https://doi.org/10.1371/journal.pgen.1002787 (2012).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 35 (6), 1547–1549. https://doi.org/10.1093/molbev/msy096 (2018).
Law, D. M. Gibberellin-enhanced indole‐3‐acetic acid biosynthesis: D‐Tryptophan as the precursor of indole‐3‐acetic acid. Physiol. Plant. 70 (4), 626–632. https://doi.org/10.1111/j.1399-3054.1987.tb04316.x (1987).
Leontidou, K. et al. Plant growth promoting rhizobacteria isolated from halophytes and drought-tolerant plants: genomic characterisation and exploration of phyto-beneficial traits. Sci. Rep. 10 (1), 14857. https://doi.org/10.1038/s41598-020-71652-0 (2020).
Lin, L. & Xu, X. Indole-3-Acetic Acid Production by Endophytic Streptomyces sp. En-1 Isolated from Medicinal Plants. Curr. Microbiol. 67 (2), 209–217. https://doi.org/10.1007/s00284-013-0348-z (2013).
Mano, Y. & Nemoto, K. The pathway of auxin biosynthesis in plants. J. Exp. Bot. 63 (8), 2853–2872. https://doi.org/10.1093/jxb/ers091 (2012).
MartÃnez-Cano, D. J. et al. Evolution of small prokaryotic genomes. Front. Microbiol. 5 https://doi.org/10.3389/fmicb.2014.00742 (2015).
Melini, F. et al. Optimization of the growth conditions through response surface methodology and metabolomics for maximizing the auxin production by Pantoea agglomerans C1. Front. Microbiol. 14 https://doi.org/10.3389/fmicb.2023.1022248 (2023).
Myo, E. M. et al. Indole-3-acetic acid production by Streptomyces fradiae NKZ-259 and its formulation to enhance plant growth. BMC Microbiol. 19 (1), 155. https://doi.org/10.1186/s12866-019-1528-1 (2019).
Numponsak, T., Kumla, J., Suwannarach, N., Matsui, K. & Lumyong, S. Biosynthetic pathway and optimal conditions for the production of indole-3-acetic acid by an endophytic fungus, Colletotrichum fructicola CMU-A109. PLOS ONE. 13 (10), e0205070. https://doi.org/10.1371/journal.pone.0205070 (2018).
Nutaratat, P., Monprasit, A. & Srisuk, N. High-yield production of indole-3-acetic acid by Enterobacter sp. DMKU-RP206, a rice phyllosphere bacterium that possesses plant growth-promoting traits. 3 Biotech. 7 (5), 305. https://doi.org/10.1007/s13205-017-0937-9 (2017).
Nutaratat, P., Srisuk, N., Arunrattiyakorn, P. & Limtong, S. Plant growth-promoting traits of epiphytic and endophytic yeasts isolated from rice and sugar cane leaves in Thailand. Fungal Biology. 118 (8), 683–694. https://doi.org/10.1016/j.funbio.2014.04.010 (2014).
Özdal, M., Gür Özdal, Ö., Sezen, A. & Algur, Ö. F. Biosynthesis Of Indole-3-Acetic Acid By Bacillus cereus Immobilized Cells. Cumhuriyet Sci. J. 37(3), 212. https://doi.org/10.17776/csj.34085 (2016).
Patten, C. L., Blakney, A. J. C. & Coulson, T. J. D. Activity, distribution and function of indole-3-acetic acid biosynthetic pathways in bacteria. Crit. Rev. Microbiol. 39 (4), 395–415. https://doi.org/10.3109/1040841X.2012.716819 (2013).
Patten, C. L. & Glick, B. R. Role of Pseudomonas putida Indoleacetic Acid in Development of the Host Plant Root System. Appl. Environ. Microbiol. 68 (8), 3795–3801. https://doi.org/10.1128/AEM.68.8.3795-3801.2002 (2002).
Peng, Y., He, Y., Wu, Z., Lu, J. & Li, C. Screening and optimization of low-cost medium for Pseudomonas putida Rs-198 culture using RSM. Brazilian J. Microbiol. 45 (4), 1229–1237. https://doi.org/10.1590/S1517-83822014000400013 (2014).
Put, H. et al. Bacillus subtilis as a host for natural product discovery and engineering of biosynthetic gene clusters. Nat. Prod. Rep. 41 (7), 1113–1151. https://doi.org/10.1039/D3NP00065F (2024).
Ranea, J. A. G., Buchan, D. W. A., Thornton, J. M. & Orengo, C. A. Evolution of Protein Superfamilies and Bacterial Genome Size. J. Mol. Biol. 336 (4), 871–887. https://doi.org/10.1016/j.jmb.2003.12.044 (2004).
Rey, M. W. et al. Complete genome sequence of the industrial bacterium Bacillus licheniformis and comparisons with closely related Bacillusspecies. Genome Biol., 5(10), r77. (2004). https://doi.org/10.1186/gb-2004-5-10-r77.
Richter, M., Rosselló-Móra, R., Glöckner, O., Peplies, J. & F., & JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32 (6), 929–931. https://doi.org/10.1093/bioinformatics/btv681 (2016).
Sam-on, M. F. S. et al. Mining the genome of Bacillus velezensis FS26 for probiotic markers and secondary metabolites with antimicrobial properties against aquaculture pathogens. Microb. Pathog. 181, 106161. https://doi.org/10.1016/j.micpath.2023.106161 (2023).
Sarmiento-López, L. G. et al. Production of indole-3-acetic acid by Bacillus circulans E9 in a low-cost medium in a bioreactor. J. Biosci. Bioeng. 134 (1), 21–28. https://doi.org/10.1016/j.jbiosc.2022.03.007 (2022).
Sasirekha, B. (ed, S.) Statistical optimization for improved indole-3-acetic acid (iaa) production by Pseudomonas aeruginosa and demonstration of enhanced plant growth promotion. J. Soil. Sci. Plant. Nutr. ahead 0-0 https://doi.org/10.4067/S0718-95162012005000038 (2012).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30 (14), 2068–2069. https://doi.org/10.1093/bioinformatics/btu153 (2014).
Seo, Y. D. & Wargo, J. A. From bugs to drugs: Bacterial 3-IAA enhances efficacy of chemotherapy in pancreatic cancer. Cell. Rep. Med. 4 (5), 101039. https://doi.org/10.1016/j.xcrm.2023.101039 (2023).
Shahzad, R. et al. Indoleacetic acid production and plant growth promoting potential of bacterial endophytes isolated from rice (Oryza sativa L.) seeds. Acta Biol. Hung. 68 (2), 175–186. https://doi.org/10.1556/018.68.2017.2.5 (2017).
Shani, E. et al. Plant Stress Tolerance Requires Auxin-Sensitive Aux/IAA Transcriptional Repressors. Curr. Biol. 27 (3), 437–444. https://doi.org/10.1016/j.cub.2016.12.016 (2017).
Shen, J. et al. Indole-3-Acetic Acid Alters Intestinal Microbiota and Alleviates Ankylosing Spondylitis in Mice. Front. Immunol. 13 https://doi.org/10.3389/fimmu.2022.762580 (2022).
Shokri, D. & Emtiazi, G. Indole-3-Acetic Acid (IAA) Production in Symbiotic and Non-Symbiotic Nitrogen-Fixing Bacteria and its Optimization by Taguchi Design. Curr. Microbiol. 61 (3), 217–225. https://doi.org/10.1007/s00284-010-9600-y (2010).
Singh, R. et al. Comparison of freeze-thaw and sonication cycle-based methods for extracting AMR-associated metabolites from Staphylococcus aureus. Front. Microbiol. 14 https://doi.org/10.3389/fmicb.2023.1152162 (2023).
Šmarda, P. et al. Ecological and evolutionary significance of genomic GC content diversity in monocots. Proc. Natl. Acad. Sci., 111(39), (2014). https://doi.org/10.1073/pnas.1321152111
Spaepen, S., Vanderleyden, J. & Remans, R. Indole-3-acetic acid in microbial and microorganism-plant signaling. FEMS Microbiol. Rev. 31 (4), 425–448. https://doi.org/10.1111/j.1574-6976.2007.00072.x (2007).
Stofan, M. & Guo, G. L. Bile Acids and FXR: Novel Targets for Liver Diseases. Front. Med., 7. (2020). https://doi.org/10.3389/fmed.2020.00544
Stothard, P. & Wishart, D. S. Circular genome visualization and exploration using CGView. Bioinformatics 21 (4), 537–539. https://doi.org/10.1093/bioinformatics/bti054 (2005).
Szkop, M. & Bielawski, W. A simple method for simultaneous RP-HPLC determination of indolic compounds related to bacterial biosynthesis of indole-3-acetic acid. Antonie van Leeuwenhoek. 103 (3), 683–691. https://doi.org/10.1007/s10482-012-9838-4 (2013).
Tang, J. et al. Biosynthetic Pathways and Functions of Indole-3-Acetic Acid in Microorganisms. Microorganisms 11 (8), 2077. https://doi.org/10.3390/microorganisms11082077 (2023).
Teale, W. D., Paponov, I. A. & Palme, K. Auxin in action: signalling, transport and the control of plant growth and development. Nat. Rev. Mol. Cell Biol. 7 (11), 847–859. https://doi.org/10.1038/nrm2020 (2006).
Viet Cuong, P., Hoa, N. P., Optimization of culture condition for IAA roduction by Bacillus sp. isolated from cassava field of Vietnam. Vietnam J. Sci. Technol. 59(3), 312. https://doi.org/10.15625/2525-2518/59/3/15600 (2021).
Wagi, S. & Ahmed, A. Bacillus spp.: potent microfactories of bacterial IAA. PeerJ 7, e7258. https://doi.org/10.7717/peerj.7258 (2019).
Wang, Y. et al. Whole-genome analysis revealed the growth-promoting mechanism of endophytic bacterial strain Q2H1 in potato plants. Front. Microbiol. 13 https://doi.org/10.3389/fmicb.2022.1035901 (2022).
Wu, C. F. J. & Hamada, M. Experiments (Wiley, 2021). https://doi.org/10.1002/9781119470007
Yoon, S. H., Ha, S., Lim, J., Kwon, S. & Chun, J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek. 110 (10), 1281–1286. https://doi.org/10.1007/s10482-017-0844-4 (2017).
Zakharova, E. A. et al. Biosynthesis of indole-3‐acetic acid in Azospirillum brasilense. Eur. J. Biochem. 259 (3), 572–576. https://doi.org/10.1046/j.1432-1327.1999.00033.x (1999).
Zeng, Q. et al. Comparative genomic and functional analyses of four sequenced Bacillus cereus genomes reveal conservation of genes relevant to plant-growth-promoting traits. Sci. Rep. 8 (1), 17009. https://doi.org/10.1038/s41598-018-35300-y (2018a).
Zeng, Q. et al. Comparative genomic and functional analyses of four sequenced Bacillus cereus genomes reveal conservation of genes relevant to plant-growth-promoting traits. Sci. Rep. 8 (1), 17009. https://doi.org/10.1038/s41598-018-35300-y (2018b).
Zeng, Q. et al. Comparative genomic and functional analyses of four sequenced Bacillus cereus genomes reveal conservation of genes relevant to plant-growth-promoting traits. Sci. Rep. 8 (1), 17009. https://doi.org/10.1038/s41598-018-35300-y (2018c).
Acknowledgements
The authors acknowledge the Department of Biotechnology, Jaypee Institute of Information Technology, Noida, for providing all necessary support for the successful execution of the study.
Author information
Authors and Affiliations
Contributions
S.K: Writing – original draft, Experiment Validation, Software, Methodology, Investigation, Formal analysis, Data curation, Conceptualization. A.M: Conceptualization and Supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Khan, S., Mathur, A. Genome Insight and factorial design to elucidate the regulation of the tryptophan-mediated IAA biosynthetic pathway in an endophyte. Sci Rep 16, 10376 (2026). https://doi.org/10.1038/s41598-026-40546-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-40546-y