Abstract
Macrophage infiltration is critically involved in the pathogene and progression of IgA nephropathy (IgAN). However, the underlying molecular mechanisms remains unclear. This study aimed to identify hub genes associated with macrophage infiltration in IgAN. We further sought to validate a key candidate, VSIG4, and to elucidate its expression pattern and functional role in the disease. Datasets for IgAN were sourced from the GEO database. Differentially expressed genes (DEGs) and Macrophage-related hub genes were identified, and their correlation with clinical characteristics was analyzed for searching biomarkers. The IgAN rat model was established to investigate the expression and distribution of hub genes in renal tissue and serum. An IgAN cohort comprised of 107 patients and 55 normal humans was conducted for validating. From two available datasets, 153 DEGs were identified and mostly enriched in the complement and coagulation cascades pathway. Three macrophage-related hub genes (C1QA, C1QB, and VSIG4) were up-regulated in IgAN, with VSIG4 showing the closest connection with clinical features, which dramatically negatively correlated with GFR (p = 0.004, r=-0.825). The expression of VSIG4, which encoded by VSIG4, was up-regulated in both the renal tissues and serum of IgAN rats, while VSIG4 primarily co-localized in the renal interstitium with the macrophage marker. A total of 107 IgAN patients were included, with the urinary VSIG4 levels elevated in IgAN patients, which showed negatively correlated with eGFR ( r = -0.272, P.adj = 0.013 ), while positively correlated with serum creatinine ( r = 0.292, P.adj = 0.009 ), 24 h urinary protein ( r = 0.273, P.adj = 0.013 ) and Oxford pathological classification T ( r = 0.227, P.adj = 0.019 ). The ROC curve indicated an AUC of 0.796 (95% CI: 0.729–0.864, P < 0.001) for urinary VSIG4 in diagnosing IgAN. VSIG4 is a crucial hub gene that affects macrophage infiltration in IgAN patients. It is highly expressed in IgAN and is closely related to various clinicopathological features. VSIG4 holds certain clinical value for diagnosing IgAN and suggests its potential role to be a sensitive biomarker and a novel intervention target for IgAN.
Similar content being viewed by others
Introduction
IgA nephropathy (IgAN) is the most common primary glomerulonephritis worldwide1, and associates with a higher risk of progressing to chronic kidney disease (CKD) or even end stage kidney disease (ESKD). However, the pathogenesis of IgAN is not yet completely understood2. Previous studies have demonstrated that macrophages play a significant role in IgAN development3. Especially, the intensity of macrophages infiltration in glomeruli is closely related with renal prognosis and predicts the response after receiving immunosuppression therapy in patients with IgAN4. Nevertheless, the mechanism of macrophage involved in the progression of IgAN remains to be investigated.
Macrophages are the core phagocytes of innate immunity5. Studies have shown that macrophages can exhibit different polarization states, i.e., M1 pro-inflammatory and M2 anti-inflammatory phenotypes6, in various diseases or at different stages of the same disease7. Hu et al. reported a close relationship between the M2 macrophage subtypes and pathological classification of IgAN8. Additionally, the infiltration degree of M2 macrophages positively correlated with the levels of serum creatinine and proteinuria in IgAN patients8,9,10,11. However, the key genes and specific mechanisms by which M2 macrophages influence Currently, the diagnosis of IgAN still relies on a kidney biopsy. There is also a significant lack of specific biomarkers for non-invasive diagnosis and prognosis prediction. Achieving a precise diagnosis and treatment for IgAN, along with early prognosis assessment, could provide a useful tool for clinical practices.
In this study, we used bioinformatics analysis to identify key genes and potential functions associated with macrophage infiltration in IgAN. VSIG4 (V-set and immunoglobulin domain-containing 4) was identified as a candidate biomarker. In IgAN animal model and patients, VISG4 was up-regulated and correlated with clinicopathological factors. Furthermore, urinary VSIG4 served as a sensitive biomarker for the diagnosis of IgAN. This finding will provide clues and evidence for further exploring the mechanisms of macrophage-related genes in the pathogenesis and progression of IgAN.
Methods
Screening for macrophage-related hub genes
Source of data
We screened relevant datasets for IgAN from GEO (www.ncbi.nlm.nih.gov/geo/) using the following search terms: ((IgAN) OR (IgA nephropathy) OR (Berger’s disease)) AND (gene expression). Based on the retrieved data, we established inclusion and exclusion criteria. Inclusion criteria included: (1) Human samples; (2) Samples obtained from kidney biopsy tissues; (3) High-throughput transcriptome data. Data from tissues subjected to manipulation, lack of normal controls, presence of a significant number of missing values in the expression matrix, and sample size < 3 per group were excluded.
Identification of differentially expressed genes and pathways
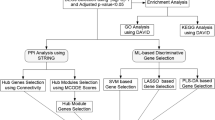
After datasets merging, we used the sva package (version 3.42.0) to remove the batch effects from the combined dataset. Subsequently, we employed the limma package (version 3.50.3), using |log2FC| > 1 and adjusted p-value < 0.05 as filtering criteria, to identify differentially expressed genes (DEGs) between IgAN patients and healthy controls12. The limma algorithm was employed to identify differentially expressed genes (DEGs) between IgAN and non-IgAN groups. The use of adjusted p-values with this method effectively controls for Type I error. Furthermore, Type II error was minimized by applying a threshold of |log2FC| > 1 and a sample size of up to 78. We conducted GO and KEGG analyses on the DEGs using the Metascape website to explore the potential functions involved in IgAN preliminarily13. Additionally, we employed the STRING database (http://string-db.org/)14 to create a protein-protein interaction (PPI) network, and the visualization was achieved using Cytoscape (V3.8.0) software.
Identification of immune infiltration specific biomarkers
In this investigation, the proportions of 22 distinct immune cell types were initially quantified in both normal and IgAN kidney tissues, using the CIBERSORT R package (version 0.1.0) on the integrated dataset15. The WGCNA package (version 1.72.1) was used to construct the weighted gene co-expression network analysis (WGCNA) analysis to identify modules most highly correlated with immune cells in IgAN patients. Then a correlation expression matrix was performed. The definitive soft threshold was selected, allowing the correlation matrix to be converted into an adjacency matrix, from which a topological overlap matrix (TOM) was generated. Genes exhibiting similar expression levels were classified into the same module through WGCNA clustering. After the WGCNA analysis, two modules demonstrating the strongest correlation with immune items were identified as hub modules for further analysis. Finally, the genes from DEGs were intersected with the key modules, and these overlapping genes were described as the differentially-expressed immune-related hub genes.
Correlation analysis between hub genes and clinical features of IgAN
We extracted IgAN-related gene expression matrices and clinical information (including serum creatinine and GFR) from the Ju et al. CKD Glom and Ju et al. CKD TubInt datasets within the Nephroseq v5 database (http://v5.nephroseq.org)16,17. Then the correlation between hub genes and clinical characteristics of IgAN was explored.
IgAN animal model
Twenty SPF male Sprague-Dawley rats (weighing from 150 to 170 g) were provided by the Laboratory Animal Center of Guangxi Medical University. The animal experimental procedures were all conducted in accordance with the ARRIVE guidelines and relevant institutional guidelines and regulations for the care and use of laboratory animals, while approved by the Animal Ethics Committee of The First Affiliated Hospital of Guangxi Medical University (No.2019-KY-079). The rats were randomly divided into IgAN group (n = 10) and normal control group (n = 10). The IgAN models were established as previously described18. Briefly, rats were gavaged with Bovine Serum Albumin (BSA) at 600 mg/kg every other day for 8 consecutive weeks. At the same time, a combined solution of 0.1 ml CCL4 and 0.3 ml castor oil was subcutaneously injected into the rats once a week for 9 consecutive weeks. At week 6 and 8, rats were intravenously injected with 0.05 mg of Lipopolysaccharide (LPS). In normal control group, rats were gavaged or injected with saline. Finally, at the end of week 16, rats were euthanized through cervical dislocation after deep anesthesia with isoflurane to minimize distress, samples of kidney tissue and serum were collected.
Verification of IgAN model
Immunofluorescence of IgA
Frozen fresh kidney tissue was cut into 4 μm sections, then incubated with goat anti-rat IgA (FITC labeled) antibody (ab97184, Abcam, Cambridge, MA) overnight at 4 °C. Images were captured using a fluorescence microscope (Olympus, Japan).
Evaluating kidney histologic lesions
Formalin fixed paraffin embedded sections were stained for hematoxylin and eosin (HE) and Periodic Acid-Schiff (PAS) staining. Proliferations of mesangial cells, mesangial matrix and endothelial cells were recorded.
Expression of VSIG4
qRT-PCR
Extract total RNA from rat kidney tissue was carried out using Trizol RNA isolation reagent (B511311-0100, Sangon Biotech, China). Take 1 ug of total RNA and reverse transcribe it into cDNA using the reverse transcription system kit (RR092A, TaKaRa, Japan) according to the manufacture instructions. RT-qPCR was performed using the SYBR Green method, and GAPDH served as the internal control gene. Primer sequences are as follows: VSIG4 forward, 5’-GGTGAGCAGGATGTATGCCA-3’ and reverse, 5’-TCAGAGCCTGGGAATCTGGT-3’; and GAPDH forward, 5’-TCTCTGCTCCTCCCTGTTCTA-3’ and reverse, 5’-GATACGGCCAAATCCGTTCA-3’.
Western Blot analysis
Rat kidney tissues were lysed and proteins were extracted, and the protein concentration was measured by BCA kits (P0012S, Beyotime Biotechnology, China). The SDS-PAGE gel electrophoresis was performed to separate the protein, followed by transferring the protein to a PVDF membrane. The membrane was blocked at room temperature for 2 h with a solution containing 5% skimmed milk powder. After blocking, the membrane was washed and incubated overnight at 4 °C with anti-VSIG4 (ER1918-60, HUABio, China) diluted 1:1000 and anti-GAPDH (10494-1-AP, Proteintech, China) diluted 1:2000. In the next day, the membrane was rewashed and incubated at room temperature for 1 h with HRP-labeled secondary antibodies. Eventually, the bands were visualized by chemiluminescent imaging system (Servicebio, China). The grayscale values were analyzed using ImageJ.
Immunohistochemistry staining
After deparaffinized and rehydrated, sections received proteinase K induced antigen retrieval and blocking, then incubated with the primary antibody VSIG4 (ER1918-60, HUABio, China) for 60 min at 37℃. A secondary antibody (G1303, Servicebio, China) was added and incubated at 37℃ for 30 min. All sections were visualized by HRP-DAB system kit (G1212, Servicebio, China).
ELISA analysis
The levels of serum VSIG4 in IgAN rats were measured using a commercial ELISA kit (MM-72609R1, MEIMIAN, China) according to the manufacture instructions.
Study population
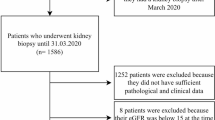
Patients with biopsy proved primary IgAN admitted at The First Affiliated Hospital of Guangxi Medical University from July 2020 to July 2025 were screened. Exclusion criteria included: age < 18 years, secondary IgAN (such as lupus nephritis, hepatitis B virus associated-glomerulonephritis, and IgA vasculitis) and the patients received glucocorticoid and immunosuppressant before kidney biopsy. This study was approved by the ethics committee of the First Affiliated Hospital of Guangxi Medical University (2016(036)). All participants provided written informed consent.
Urine sample collection and measurement
Samples of spot urine were collected on the day of kidney biopsy, we also took samples from age- and sex-matched healthy people as normal controls. The levels of urinary VSIG4 were measured using a commercial ELISA kit as above described. Urine albumin and creatinine was measured using a automated biochemical analyzer (SYP-H4308, UpingBio, China). The levels of urinary VSIG4 were normalized to urine creatinine concentration, and expressed as ng/mg UCr.
Statistical analysis
Continuous variables were expressed as mean ± SD or median (IQR). Categorical data were expressed as counts and percentages (%). A Student’s t test or a Mann-Whitney U test was used for comparison between-group differences for continuous variables, and the χ2 test was used for categorical variables. Correlation analysis was conducted to explore the association between hub genes and clinical pathological indicators. ROC curves were plotted to evaluate the value of urinary hub genes in diagnosing IgAN.
All statical analyses were performed using R version 4.3.3. Two-tailed P < 0.05 were considered a statistically significant.
Results
Identification of candidate biomarkers for IgAN
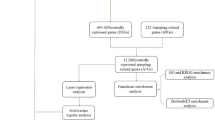
According to the inclusion and exclusion criteria, we identified two available high-throughput datasets for IgAN (GSE37460 and GSE93798), comprising a total of 49 IgAN patients and 29 healthy controls. After merging the two datasets and removing batch effect, we conducted differential analysis and identified 109 down-regulated genes and 44 up-regulated genes in IgAN (Fig. 1A,B). Subsequently, the GO analysis and KEGG pathway analysis were performed by using these DEGs, and the results revealed that the top three biological functions were: inflammatory response, collagen-containing extracellular matrix, and response to growth factor (Fig. 1C), and the top three pathways were: protein digestion and absorption, the Complement and coagulation cascades, and the IL-17 signaling pathway (Fig. 1D). To elucidate the complex interactions among the DEGs, a protein-protein interaction (PPI) network was constructed using the STRING database and visualized through Cytoscape software—the resulting network, as depicted in Fig. 1E.
Identified differentially expressed genes(DEGs) and functional enrichment analyses of DEGs. (A) A heatmap of DEGs. (B) A volcano plot of DEGs. (C) Bubble diagrams of the GO functional analysis of DEGs. (D) Bubble diagrams of the KEGG pathway enrichment analysis of DEGs. (E) The PPI network of DEGs was established using string and Cytoscape, comprised 55 nodes and 166 edges.
Identification of macrophages-associated hub genes
The CIBERSORT algorithm revealed immune infiltration characteristics of IgAN patients, as shown in Fig. 2A, we used Batch variance analysis to further identify six immune cell types that exhibited comparable differences between IgAN patients and normal controls, including activated NK cells (p.adj = 0.0004), neutrophils (p.adj = 0.0017), resting CD4 T cells (p.adj = 0.0030), naive B cells (p.adj = 0.0149), M2 macrophages (p.adj = 0.0149), and activated dendritic cells (p.adj = 0.0149). Following WGCNA analysis utilizing all samples, the soft threshold power was calibrated to 7 (scale-free R^2 = 0.90). Subsequently, a total of 40 modules were obtained, each with its corresponding expression profiles. Notably, the ‘orange’ module exhibited a strong positive correlation with M2 macrophages (r = 0.74, p < 0.0001), while the ‘violet’ module showed a negative correlation (r = 0.66, p < 0.0001) (Fig. 2B). By intersecting DEGs and genes selected from the ‘orange’ and ‘violet’ modules, we identified three genes that considered as hub genes associated with M2 macrophage in IgAN: VSIG4 (AveExpr:6.373029413, adj.P.Val = 0.00021, log2FC = 1.164519), C1QA (AveExpr: 6.32086141, adj.P.Val = 0.000738, log2FC = 1.08447119), and C1QB (AveExpr: 6.170875945, adj.P.Val = 0.001307, log2FC = 1.2089183) (Fig. 3). All three hub genes were found to be upregulated in the IgAN group compared to the normal group, with VSIG4 showing the significant statistical difference.
Immune infiltration analysis and construction of weighted gene co-expression networks. (A) 22 immune cells in samples with IgAN and normal renal tissues of integrated dataset. (B) 40 modules revealed by the WGCNA.
The hub genes are obtained by taking the intersection of DEGs and immune-related genes(IRGs). Three hub genes were identified: VSIG4, C1QA, and C1QB.
The correlation between hub genes with clinical features
Additionally, we explored the relationship between hub genes and clinical features of IgAN using the Nephroseq V5 database. As shown in Fig. 4, all three key hub genes were positively correlated with serum creatinine levels in IgAN patients ( VSIG4 r = 0.768, C1QA r = 0.513, C1QB r = 0.537), while exhibiting negative correlations with glomerular filtration rate (VSIG4 r=-0.825, C1QA r=-0.787, C1QB r=-0.832). Analysis of the Nephroseq V5 database revealed a statistically significant correlation between VSIG4 expression and clinical indicators, with notably high statistical values. Based on these findings, VSIG4 was selected for further experimental validation and clinical analysis.
Correlation between hub genes and clinical parameters. Correlation of hub genes with glomerular filtration rate and serum creatinine in IgAN patients.
Validation of VSIG4 expression in IgAN rat
The successful establishment of IgAN rat model was supported by proliferation of mesangial cells and deposition of IgA in mesangial area (Fig. 5A-B). The mRNA and protein expression levels of VSIG4 in renal tissues were significantly elevated in the IgAN rats compared to the normal control group (Fig. 6A,B), and the mRNA and protein expression levels of VSIG4 in renal tissues increased over time with the progression of the modeling (Fig. 7A,B). Additionally, ELISA results also indicated an increase in serum VSIG4 levels in IgAN rats (Fig. 6C), which were consistent with the earlier bioinformatics findings. Furthermore, we examined the localization of VSIG4 in renal tissues using IHC. The results revealed positive staining for VSIG4 in IgAN, localized in the glomeruli and renal tubulointerstitial. The IgAN rats exhibited a higher positive cell rate and staining intensity than the normal control group (Fig. 6D), indicating an upward trend of VSIG4 in this condition.
Validation of the IgAN rat model. (A) HE staining of renal tissue; (B) Immunofluorescence of renal tissue. The deposition of electron-dense substances in the mesangial area observed under electron microscopy and the green fluorescence suggests IgA immunecomplex.
Validation of VSIG4 expression levels in the IgAN rats. (A) The mRNA expression in renal tissue; (B) The protein expression in renal tissue; (C) Serum VSIG4 levels; (D) The localization of VSIG4 in renal tissue. Samples were from rats at week 16. Data were expressed as means ± SD, *P < 0.05, indicated the significance between the two group.
Changes in expression of VSIG4 in renal tissue of IgAN rats with modeling time. (A) Changes in the mRNA in renal tissue with modeling time; (B) Changes in the protein in renal tissue with modeling time.
Studies have shown that VSIG4 is a protein primarily expressed on the surface of macrophages, and our bioinformatics analysis suggests its strong association with macrophages. We conducted co-localization staining on serial sections to examine the co-localization between VSIG4 and the macrophage marker F4/80 in IgAN rat kidney tissue. The results showed that VSIG4 and the macrophage marker F4/80 primarily co-localized in the renal tubulointerstitial, with a minor portion co-localizing in the glomeruli (Fig. 8).
Co-localization of VSIG4 and the macrophage marker via immunohistochemical serial sections. Samples were from rats at week 16.
Validation of VSIG4 expression in IgAN patients
A total of 107 patients with IgA nephropathy (IgAN) were included in this study. The cohort had a male-to-female ratio of 1.20:1 and a mean age of 37.62 ± 12.46 years. The mean estimated glomerular filtration rate (eGFR) for the entire IgAN cohort was 82.75 ± 31.37 mL/min/1.73 m2. Among these, 32 patients (29.9%) presented with impaired renal function, defined as a baseline eGFR < 60 mL/min/1.73 m2, with a mean eGFR of 43.75 ± 10.46 mL/min/1.73 m2. Compared to patients with preserved renal function, those with impaired renal function were significantly older and had a higher prevalence of hypertension, along with elevated mean arterial pressure (MAP), triglyceride (TG) levels, 24-hour urinary total protein (24h_UTP), and a greater proportion of Oxford pathological classification T lesions (all P < 0.05). These data are summarized in Table S1.
As shown in Fig. 9A, the results revealed a significantly elevated level of urinary VSIG4 in the IgAN group compared to healthy controls group (P < 0.001). Furthermore, IgAN patients with decreased renal function exhibited higher urinary VSIG4 levels than those with normal renal function (P = 0.039) (Fig. 9B).
Comparison of urinary VSIG4 levels among different group. (A) Comparison between IgAN and normal control group; (B) Comparison between normal renal function and decreased renal function group in IgAN. Data were expressed as means ± SD, *P < 0.05, ****P < 0.0001, indicated the significance between the two group.
Correlation between urinary VSIG4 and clinicopathological indicators
As summarized in Table 1, urinary VSIG4 levels in the 107 patients demonstrated a significant negative correlation with eGFR (r = -0.272, P = 0.0046), and significant positive correlations with serum creatinine (Scr) (r = 0.292, P = 0.0092), 24-hour urinary protein (r = 0.272, P = 0.0045), and the Oxford Pathology Score T (r = 0.227, P = 0.0186). These results indicate a potential association of urinary VSIG4 with baseline renal function impairment, the degree of proteinuria, and pathological features related to mesangial proliferation and tubulointerstitial fibrosis in IgAN.
The value of urinary VSIG4 levels in distinguishing IgAN patients from healthy pepole
To evaluate the diagnostic performance of urinary VSIG4 levels in IgAN patients, a receiver operating characteristic (ROC) curve was constructed. The analysis revealed an area under the curve (AUC) of 0.796 (95% CI: 0.729–0.864, P < 0.001). Using an optimal cut-off value of 12.12 ng/mg urinary creatinine (UCr), the sensitivity and specificity were 70.09% and 76.67%, respectively. as shown in Fig. 10, suggesting its potential clinical value in distinguishing IgAN patients from healthy pepole.
ROC curve of urinary VSIG4 levels for the diagnosis of IgAN. The AUC was 0.796, P < 0.001.
Discussion
Previous research has shown that the disease process of IgAN is characterized as a “four-hit” process. First, the body produces excessive poorly O-glycosylated galactose-deficient IgA1 (Gd-IgA1). Subsequently, antiglycan autoantibodies recognize the Gd-IgA1, leading to the aggregation of Gd-IgA1 into polymeric IgA1 immune complexes. These immune complexes then deposit in the glomerular mesangium, activating the complement system, and ultimately leading to renal tubulointerstitial inflammation and kidney injury19. Various immune cells play a role in the fourth hit of IgAN pathogenesis. Single-cell sequencing analyses have shown that macrophages are the most prevalent type of immune cell in the renal tissues of IgAN patients20. Recent studies have reported that macrophages are closely associated with clinical indicators such as renal function and proteinuria, which may serve as independent risk factors and markers for predicting adverse outcomes in IgAN21,22. However, the key genes involved and the specific mechanisms still require further investigation.
Our study preliminarily explored the characteristics of immune cell infiltration in the renal tissues of IgAN patients using CIBERSORT analysis. The results showed that the abundance of M2 macrophages in the renal tissues of the IgAN group was significantly higher than that of the normal control group. Bin Li et al., who utilized CIBERSORT analysis, observed a significant increase in the ratio of M1 and M2 macrophages within the glomerular and interstitial compartments in IgAN, with a higher proportion of resting CD4 + memory T cells and M2 macrophages in the tubulointerstitial area23. Wenxue Hu et al. reported that M2 subpopulations were involved in the progression of IgAN8. An elevated presence of macrophages in the glomerular and tubulointerstitial regions may contribute to oxidative stress and the activation of the complement system24. The previous research findings are consistent with our results. M2 macrophages could regulate the immune response of inflammation and the type I adaptive immunity, facilitate the clearance of debris, boost angiogenesis, mediate anti-inflammatory effects, inhibit cell apoptosis, enhance cell proliferation, and support tissue regeneration25. In recent years, numerous studies have investigated the role of M2 macrophages in IgAN. An increased number of M2 macrophages has been observed in patients with IgAN. These macrophages are positively correlated with the serum creatinine and 24-hour proteinuria, yet are negatively correlated with eGFR in IgAN. Additionally, M2 macrophages play a crucial role in the progression of IgAN-associated fibrosis11, which indicates that the innate immune response was closely associated with IgAN26. Our pathway analysis also revealed the pivotal role of immune-related pathways, particularly the IL-17 signaling pathway. A study based on a limited sample size has reported increased plasma and urinary concentration of IL-17 A in IgAN patients27. One study also reported that genes overexpressed in IgAN patients are mainly enriched in the IL-17 signaling pathway28. In conclusion, immune-related biological mechanisms, such as M2 macrophages and the IL-17 signaling pathway, are the hub processes in IgAN. Therefore, it is necessary to further explore the specific mechanisms of M2 macrophages and their related key genes in the progression of IgAN.
Through differential expression analysis and WGCNA, three genes—C1QA, C1QB, and VSIG4—were identified as potentially key players in IgA nephropathy (IgAN), all of which are linked to M2 macrophage-related pathways29. In the context of IgAN, studies by Siqing Wang, Hong-Joo Lee, and colleagues have demonstrated that patients with positive C1q deposition are at higher risk of progressing to end-stage renal disease30,31. Furthermore, research by Amir Horowitz et al. has established that C1QA is associated with macrophages and serves as a reliable marker for macrophage identification32. A high-throughput analysis of peripheral blood mononuclear cells (PBMCs) also suggests that C1QB may act as a potential diagnostic biomarker for IgAN33. In contrast, the role of VSIG4 in IgAN remains largely unexplored. However, in our analysis of the Nephroseq V5 database, VSIG4 exhibited stronger statistical associations with clinical indicators compared to C1QA and C1QB. Consequently, VSIG4 was selected for further experimental and clinical validation. Our results confirmed that urinary VSIG4 levels are closely associated with clinicopathological indices in IgAN, and demonstrated its ability to distinguish IgAN from non-IgAN cases, as well as to differentiate between patients with and without impaired baseline renal function.
The V-set Ig domain containing 4 (VSIG4)34 is a B7 family-related protein that is specifically expressed in macrophages, capable of binding to receptor molecules on the cell surface35, also known as the complement receptor of the immunoglobulin Ig superfamily molecule (CRIg). Studies suggest that VSIG4 is closely associated with the phenotype of macrophages24. Our study calculated the relationship between VSIG4 and the immune microenvironment via the CIBERSORT algorithm. Additionally, through WGCNA analysis and protein-protein interaction network screening, we found that VSIG4 is a core gene closely related to M2 macrophages, which is consistent with previous research. In this study, a total of 49 samples were included in the WGCNA. According to established methodological standards, a sample size exceeding 15 is sufficient to control Type II error in this algorithm36. Our results demonstrated that the two key modules identified had p-values of less than 0.0001, indicating a low probability of Type I error. LI Y et al. indicated that the expression value of VSIG4 was up-regulated in mice with acute kidney injury (AKI), and the highly expressed VSIG4 alleviated the lesions of kidney tissue in AKI mice by inhibiting the secretion of inflammatory factors in M1 macrophages, playing a protective role37. Sang Youb Han et al. pointed out that VSIG4 was up-regulated in diabetic nephropathy, which was associated with proteinuria and profibrotic markers38. Additionally, a previous study has demonstrated that the expression of VSIG4 is up-regulated in both animal and cell CKD models39. However, there are few studies indicating a relationship between VSIG4 and IgAN currently. In our study, we utilized the external dataset from the Nephroseq V5 database to confirm that the VSIG4 levels were elevated in IgAN. Meanwhile, we constructed an IgAN rat model and collected urine samples from IgAN patients to further verify the expression of VSIG4 in this condition. The results were consistent with the bioinformatics analysis findings: both mRNA and protein expression levels of VSIG4 in the renal tissues and serum of IgAN rats were significantly higher than those of the normal control group. Likewise, we observed that urinary VSIG4 levels in IgAN patients were also significantly elevated compared to those in the normal population. Additionally, our study also observed the distribution of VSIG4 in the renal tissues of IgAN rats using IHC, and the results showed that positive staining of VSIG4 could be observed in both the glomeruli and tubulointerstitium. Tang et al. demonstrated that VSIG4 was mainly elevated in the glomeruli and renal tubules of LN patients34, and VSIG4 could also be observed in the damaged renal tubules of another CKD mouse model39. We also detected whether VSIG4 and the macrophage marker F4/80 were co-expressed by continuous section co-localization staining. And the results showed that VSIG4 and F4/80 were mainly co-localized in the renal tubulointerstitium, with a small portion co-localized in the glomerulus. Yu Sasaki et al. utilized serial tissue section co-localization techniques to assess the spatial relationship between SNX gene expression and podocin localization in IgA nephropathy40. Therefore, we suggested that VSIG4 overexpression may play an important role in the progression of IgAN by affecting the activation of M2 macrophages.
Our analysis revealed that urinary VSIG4 levels were inversely correlated with eGFR and positively correlated with serum creatinine, 24-hour urinary protein, and the Oxford Pathology Score T. These findings suggest that VSIG4 expression is closely associated with baseline renal function impairment, the severity of proteinuria, and tubulointerstitial fibrosis lesions in patients with IgAN. Our study also showed the mRNA and protein expression levels of VSIG4 in the renal tissue of IgAN rats show an increasing trend with the extension of the modeling time, indicating that VSIG4 may be related to the course of IgAN. Furthermore, a ROC curve was constructed to assess the diagnostic potential of urinary VSIG4 in identifying IgAN patients. The results indicated that urinary VSIG4 exhibited favorable diagnostic performance in distinguishing IgAN patients from healthy individuals, with an AUC exceeding 0.75 and a sensitivity above 0.7. These findings suggest that VSIG4 may represent a potential therapeutic target for preserving or improving renal function in IgA nephropathy (IgAN). Furthermore, it holds promise as a non-invasive diagnostic biomarker for the disease.
However, several limitations of this study should be acknowledged. First, dual immunofluorescence staining was not employed to investigate the spatial relationship between VSIG4 and the macrophage marker F4/80; this will be addressed in subsequent experiments. Furthermore, intervention studies and in vitro functional assays to examine the effect of VSIG4 on macrophage polarization or cytokine secretion were not performed. The specific molecular mechanisms through which VSIG4 contributes to IgAN pathogenesis require further comprehensive investigation and elucidation, supported by multidimensional validation in future studies.
Conclusion
In conclusion, our study demonstrates the significantly elevated expression of VSIG4 in both the serum and renal tissues of IgAN rats, which served as a hub gene co-associated with macrophage infiltration, suggesting its potential role in the progression of IgAN through immune modulation. Furthermore, urinary VSIG4 demonstrates a strong correlation with the clinicopathological conditions of these IgAN patients, indicating its potential clinical value in diagnosing IgAN. It is anticipated that VSIG4 may be a sensitive biomarker and a novel target in IgAN.
Data availability
The data presented in this study is available from the corresponding author on request.
References
Pattrapornpisut, P., Avila-Casado, C. & Reich, H. N. IgA Nephropathy: Core Curriculum 2021. Am. J. Kidney Dis. 78(3), 429–441 (2021).
Lai, K. N. et al. IgA nephropathy. Nat. Rev. Dis. Primers. 2, 16001 (2016).
Takahata, A. et al. Crucial Role of AIM/CD5L in the Development of Glomerular Inflammation in IgA Nephropathy. J. Am. Soc. Nephrol. 31(9), 2013–2024 (2020).
Xie, D. et al. Intensity of Macrophage Infiltration in Glomeruli Predicts Response to Immunosuppressive Therapy in Patients with IgA Nephropathy. J. Am. Soc. Nephrol. 32(12), 3187–3196 (2021).
Rogers, N. M., Ferenbach, D. A., Isenberg, J. S., Thomson, A. W. & Hughes, J. Dendritic cells and macrophages in the kidney: a spectrum of good and evil. Nat. Rev. Nephrol. 10(11), 625–643 (2014).
Gordon, S. Alternative activation of macrophages. Nat. Rev. Immunol. 3(1), 23–35 (2003).
Mantovani, A. et al. The chemokine system in diverse forms of macrophage activation and polarization. Trends Immunol. 25(12), 677–686 (2004).
Hu, W. et al. M2a and M2b macrophages predominate in kidney tissues and M2 subpopulations were associated with the severity of disease of IgAN patients. Clin. Immunol. 205, 8–15 (2019).
Lu, J. et al. Discrete functions of M2a and M2c macrophage subsets determine their relative efficacy in treating chronic kidney disease. Kidney Int. 84(4), 745–755 (2013).
Olmes, G. et al. CD163 + M2c-like macrophages predominate in renal biopsies from patients with lupus nephritis. Arthritis Res. Ther. 18, 90 (2016).
Yang, M., Liu, J. W., Zhang, Y. T. & Wu, G. The Role of Renal Macrophage, AIM, and TGF-beta1 Expression in Renal Fibrosis Progression in IgAN Patients. Front. Immunol. 12, 646650 (2021).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43(7), e47 (2015).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53(D1), D672–D7 (2025).
Szklarczyk, D. et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 39(Database issue), D561–D568 (2011).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods. 12(5), 453–457 (2015).
Ju, W. et al. Tissue transcriptome-driven identification of epidermal growth factor as a chronic kidney disease biomarker. Sci. Transl Med. 7(316), 316ra193 (2015).
Liu, J. et al. Klotho exerts protection in chronic kidney disease associated with regulating inflammatory response and lipid metabolism. Cell. Biosci. 14(1), 46 (2024).
Xu, R. et al. Yi-shen-hua-shi granules modulate immune and inflammatory damage via the ALG3/PPARgamma/NF-kappaB pathway in the treatment of immunoglobulin a nephropathy. J. Ethnopharmacol. 319(Pt 2), 117204 (2024).
Stamellou, E. et al. IgA nephropathy. Nat. Rev. Dis. Primers. 9(1), 67 (2023).
Zheng, Y. et al. Single-Cell Transcriptomics Reveal Immune Mechanisms of the Onset and Progression of IgA Nephropathy. Cell. Rep. 33(12), 108525 (2020).
Soares, M. F. et al. Relationship between renal CD68(+) infiltrates and the Oxford Classification of IgA nephropathy. Histopathology 74(4), 629–637 (2019).
Gong, S. et al. Urinary Soluble CD163 Levels Predict IgA Nephropathy Remission Status. Front. Immunol. 12, 769802 (2021).
Zeng, H. et al. Single-cell RNA-sequencing reveals distinct immune cell subsets and signaling pathways in IgA nephropathy. Cell. Biosci. 11(1), 203 (2021).
Liu, B. et al. The biology of VSIG4: Implications for the treatment of immune-mediated inflammatory diseases and cancer. Cancer Lett. 553, 215996 (2023).
Murray, P. J. et al. Macrophage activation and polarization: nomenclature and experimental guidelines. Immunity 41(1), 14–20 (2014).
Floege, J. & Feehally, J. The mucosa-kidney axis in IgA nephropathy. Nat. Rev. Nephrol. 12(3), 147–156 (2016).
Watorek, E. & Klinger, M. IL-17A as a potential biomarker of IgA nephropathy. Pol. Arch. Med. Wewn. 125(3), 204–206 (2015).
Tang, R. et al. A Partial Picture of the Single-Cell Transcriptomics of Human IgA Nephropathy. Front. Immunol. 12, 645988 (2021).
Liang, Z., Pan, L., Shi, J., Zhang, L. & C C1QB, and GZMB are novel prognostic biomarkers of skin cutaneous melanoma relating tumor microenvironment. Sci. Rep. 1QA(1), 20460 (2022).
Wang, S. et al. Roles of mesangial C3 and C1q deposition in the clinical manifestations and prognosis of IgAN. Int. Immunopharmacol. 120, 110354 (2023).
Lee, H. J. et al. Association of C1q deposition with renal outcomes in IgA nephropathy. Clin. Nephrol. 80(2), 98–104 (2013).
Horowitz, A., Yu, H., Pandey, S., Mishra, B. & Sahoo, D. C1QA is an invariant biomarker for tissue macrophages. bioRxiv (2024).
Gan, T. et al. Unveiling biomarkers and therapeutic targets in IgA nephropathy through large-scale blood transcriptome analysis. Int. Immunopharmacol. 132, 111905 (2024).
Tang, C. et al. V-Set Immunoglobulin Domain-Containing Protein 4 as a Novel Serum Biomarker of Lupus Nephritis and Renal Pathology Activity. Arthritis Rheumatol. 75(9), 1573–1585 (2023).
Vogt, L. et al. VSIG4, a B7 family-related protein, is a negative regulator of T cell activation. J. Clin. Invest. 116(10), 2817–2826 (2006).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Li, Y. et al. VSIG4 overexpression alleviates acute kidney injury of mice via inhibition of M1-macrophages activation. Ann. Transl Med. 10(10), 559 (2022).
Han, S. Y. et al. Upregulation of VSIG4 in type 2 diabetic kidney disease. Life (Basel) 12(7) (2022).
Han, S. Y. et al. The role of V-Set Ig domain-containing 4 in chronic kidney disease models. Life (Basel) 13(2) (2023).
Sasaki, Y. et al. Sorting Nexin 9 facilitates podocin endocytosis in the injured podocyte. Sci. Rep. 7, 43921 (2017).
Funding
This research was founded by Natural Science Foundation of Guangxi Zhuang Autonomous Region (serial number: 2022GXNSFAA035458).
Author information
Authors and Affiliations
Contributions
Pan L and Tang LQ designed research; Tang LQ and Xu YS analyzed data; Xu YS, Nong ZQ, Li FJ, and Pan L performed rats experimental research; Xu YS, Pan L, and Tang LQ wrote the paper. The first draft of the manuscript was written by Tang LQ, Xu YS. Nong ZQ, Li FJ, and Pan L commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
In our research, the animal experimental procedures were approved by the Animal Ethics Committee of The First Affiliated Hospital of Guangxi Medical University (No.2019-KY-079). Research protocols involving human participants were reviewed and formally approved by the ethics committee of the First Affiliated Hospital of Guangxi Medical University (2016(036)).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tang, L., Xu, Y., Nong, Z. et al. Identifying the hub genes in macrophage infiltration and verifying of the role of VSIG4 in IgA nephropathy. Sci Rep 16, 10211 (2026). https://doi.org/10.1038/s41598-026-40679-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-40679-0