Abstract
With increased use of agrochemicals to protect crops against phytopathogenic diseases, the environment is facing serious hazardous effects. The proven antimicrobial properties of nanomaterials may provide a new level of sustainable development and rapid research for the potential to replace chemical fungicides in crop protection. The objective of this study was to investigate the in vitro and in vivo antifungal activity of chitosan-copper nanocomposite (Ch-Cu NPs) against Rhizoctonia solani and Fusarium oxysporum to control root rot and wilt disease of marjoram plants. The chemically synthesized Ch-Cu NPs nanocomposite was characterized with respect to its morphological features and size, which were obtained by Transmission Electron Microscope (TEM) analysis, and the presence of functional chemical groups responsible for the synthesis of Ch-Cu NPs was determined using Fourier Transform Infrared (FTIR) spectrophotometer. Antifungal activity against the two pathogenic fungi was assessed using three concentrations (25, 50, and 100 mg/l) of Ch-Cu NPs. The laboratory evaluation showed that mycelial radial growth of F. oxysporum and R. solani was inhibited by 80.55% for two fungi at 100 mg/L of Ch-Cu NPs. The morphology of F. oxysporum and R. solani was investigated, and molecular identification was conducted to prove that F. oxysporum and R. solani were the causal agents of root rot and wilt disease of marjoram plants in Egypt. Pathogenicity test showed that both F. oxysporum and R. solani were virulent with the measured disease incidence was 59.14% and the disease severity was 28.47%, respectively. In this work, Ch-Cu NPs at 50 mg/L were found to be the appropriate concentration that could lead to a reduction in disease incidence by 23.67% and disease severity by 5.57% with induction of plant defense. The results of HPLC analysis revealed an increase in 7 out of 9 polyphenols in 50 mg/L Ch-Cu NPs application compared to the untreated control plants. Molecular studies using qRT-PCR technique on marjoram leaf tissue were done to determine the relationship between polyphenols biosynthesis and the expression of responsible genes affected by Ch-Cu NPs application. Genes such as CAD, C4H, TAT, FLS, EGS, HPPR, PAL, and RAS showed an upregulation of their expression in marjoram treated leaves compared to the control leaves. The release profile of Ch-Cu NPs also showed an important insight into the mode of disease control. The conclusion of this work is considered as an important step toward the application of nanomaterials in agriculture, where the biosynthesized and biodegradable Cu-chitosan NPs have better biological control compatibility as it can induce similar effects to natural plant defense elicitors and antioxidant defense to prime the crop for better resistance.
Similar content being viewed by others
Introduction
Plant pathogens lead to 16% of crop yield loss1. Pest attack causes 20–40% loss of agricultural crops annually in the world2. The most important problem that faces global agricultural system is crop diseases, particularly in high-value economic crops, and suffering a lot in substantial losses in both yield and quality3,4. Soil fungi pathogens cause root rot and wilt diseases like Fusarium wilt disease5,6. Chemical control methods may have a difficulty and hazard problem. The process of pathogen spread by climate change is one of the factors that accelerated the transmission, so Fusarium wilt disease created an urgent need for effective and safe management strategies, and the scientific community began to search for non-chemical management ways that can safely replace chemical pesticides and act against the disease7,8,9. Chemical fungicides work as a high performance in plant disease management, but with big problems in environment and human health. That forced scientists to search for a scientific basis to establish a disease-management strategy to replace synthetic chemical fungicides in controlling plant fungal disease with green and unhazardous fungicides to protect economic crops and not pose a risk to human health after their consumption10.
Copper nanoparticles (Cu NPs) have been shown to have antimicrobial and anti-biofilm activity against a wide range of plant pathogens. This has prompted the use of Cu NPs as components of antimicrobial mixtures. They are possible alternative materials that can be used to replace some of the more hazardous chemical fungicides. The proposed use of Cu NPs in the management of fungal diseases of agricultural crops can also qualify them as nanofungicides11. Application of CuO NPs enhanced plant resistance against fungal root rot, wilt and leaf spot diseases12,13,14,15. Cu NPs has been described as nutritional supplements that can promote maize growth and improve plant defense responses16. However, the tendency of pure Cu NPs to aggregate diminishes their antimicrobial activity. This may be improved by using copper nanoparticles conjugated to biodegradable nano carrier with unique antimicrobial features, minimizing copper agglomeration in plant cells. Carrier chitosan was found to be an efficient carrier for loading bioactive agents for the management of plant disease, because of its biodegradable, non-toxic and biocompatible polymer nature appropriate for the usage in agriculture17. This product is known to have many well-documented features, including antimicrobial18, immune-modulating19 and plant growth promoting activities20. Recent studies show that chitosan nanoparticles (Ch NPs) can induce antioxidant and defense enzymes system21,22. A microarray transcription analysis of plants treated with Ch NPs showed that the level of defense response is increased in Ch NP treated plants with upregulation of the expression level of defense genes. This finding clearly elucidated the role of Ch NPs in potentiating plant innate immunity23. Induced resistance in plant disease is considered a good replacement for the genetic resistance strategy for controlling infection caused by soil-borne pathogens that normally treated with the chemical pesticide in agriculture. The widespread application of systemic acquired resistance (SAR) is a new management technique for handling pathogens in a wide variety of crop species24,25.
The current study aimed to identify the ability of Chitosan-Copper nanoparticles (Ch-Cu NPs) to manage wilt and root rot disease pathogens; Fusarium oxysporum and Rhizoctonia solani causing marjoram (Origanum majorana L.) wilt and root rot disease. The study was divided into three main objectives (1) to evaluate the antifungal activity of Ch-Cu NPs nanocomposite as inhibitor of root rot and wilt diseases by laboratory and greenhouse evaluation experiments (2) to assess their ability to induce systemic resistance against these diseases in marjoram in greenhouse conditions and (3) to study the mode of resistance induction by checking the expression patterns of related regulatory and defense genes.
Materials and methods
Ch-Cu NPs source
The Ch-Cu NPs utilized in this study, possessing 99.9% purity, were sourced from the Nanotechnology and Advanced Materials Central Laboratory (NAMCL) at the Regional Center for Food and Feed (RCFF), Agricultural Research Center (ARC), located in Giza 12,619, Egypt.
Characterization of Ch-Cu NPs
The synthesized Ch-Cu NPs were verified using UV spectrophotometry (Unico UV-2100 spectrophotometer, Unico Instrument Co. Ltd, China). CH NPs were undergone characterization via transmission electron microscopy (TEM) (JEOL, JEM-1010, Boston, USA). TEM enabled morphological examination of both nanoparticle varieties. Fourier transform infrared (FTIR) spectrophotometry (Bruker tensor 27 IR with KBr discs, Massachusetts, United States) was performed for functional group identification. CH NPs chemical functional groups were identified within 4000–400 cm-1 wavelength ranges.
Isolation and identification of the causal pathogens
Naturally infected marjoram plants (90 days old) showing typical root rot and wilt symptoms were collected from four nurseries in Egypt’s Giza governorate during the spring 2024 growing season and brought to the laboratory for isolation, as described by Ahmed26. According to the formula below, the frequency of the isolated fungus was calculated.
Morphological identification
Cultural and microscopic examinations served as the foundation for preliminary isolate identification. Purification of fungal colonies was achieved through single spore or hyphal tip techniques. Morphological characteristics guided fungal isolation procedures, incorporating colony morphology assessment and cultural attributes such as growth rates, conidial morphology, and pigment production on PDA media. Identification procedures followed established protocols outlined by Booth and et al.27,28,29,30.
Pathogenicity test
Pathogenicity tests were conducted in pots experiment under greenhouse conditions at the Ornamental Medicinal and Aromatic Plant Diseases Research Department, Plant Pathology Research Institute, Agricultural Research Centre, Giza, Egypt. Evaluation of the disease potentials was done for the three pathogens in marjoram plants according to the procedure described by Ahmed26. Loamy clay soil was used and placed in clay pots. The soil was sterilized using commercial formalin solution (5%) and covered with polyethylene sheets. The pots were aerated for two weeks. Fungal inocula were made by incubating each fungus in sterilized sand-sorghum medium in Erlenmeyer flasks for two weeks. The fungal concentration was 2.5% of soil infestation, followed by irrigation and one-week equilibration periods. Experimental design included three replicate pots, planted with 4 seedlings were established per pot (30 cm) in three treatment replicates. Control treatments received equivalent volumes of sterilized, uninoculated sand-sorghum grain medium mixed into sterilized soil. Plant assessment occurred 90 days following planting, with both inoculated and control plants evaluated for root rot and wilt incidence alongside disease severity measurements. Disease incidence percentages for root rot and wilt were determined using the following calculation:
Disease severity (DS) was scored based on El-Mohamedy31 using the following formula:
where: n = number of infected plants, c = category number, N = total number of examined plants, and C = the highest category number of infections.
Molecular Identification of causal pathogen
F. oxysporum and R. solani isolates were inoculated and grown in double-layer (one solid and one liquid) media in 90-mm Petri dishes for 5 days. DNA was extracted as previously described Amer32. PCR was performed using an Applied Biosystems™ MiniAmp Thermal Cycler (ABI, Norwalk, CT, USA).
Molecular identification of F. oxysporum and R. solani isolates was performed using the specific primer. The amplification program and the PCR products were analyzed by electrophoresis in a 1.5% agarose gel33,34.
Effect of Ch-Cu NPs on root rot, wilt diseases of marjoram
Plastic pots measuring 30 cm in diameter containing sterilized soil (un-infested) were utilized for cultivation purposes in this experimental setup. Soil preparations infested with pathogens F. oxysporum and R. solani were prepared using the same protocols used for the pathogenicity test. Seedlings of marjoram were given 30-min of immersion treatment before transplantation into infested soil. Treatment groups included negative controls (seedlings immersed in water and transplanted into un-infested soil), positive controls (seedlings given 30-min root immersion in water before transplantation into infested soil), and additional treatment groups consisting of 30-min root immersion in Ch-Cu NPs at different concentrations (25, 50, 100 mg/L) before establishment in infested soil. Applications of Ch-cu NPs were made three times, the first one after planting (two weeks) and then repeated every two weeks for two times. Fertilization and irrigation were carried out using standard agricultural practices and following Ministry of Agriculture guidelines for ornamental tree production. Root rot and wilt disease incidence and severity were assessed at 90 days post-transplantation using the same criteria as in the pathogenicity test.
Determination of Total Phenolic compounds (TPC) Content
Total phenolic compounds (TPC) in fresh marjoram leaves were quantified following established protocols35. To 100 mg of fresh marjoram leaves, 20 mL of 80% (v/v) methanol was added, and the mixture was macerated for 24 h. 0.2 mL of methanolic extract was mixed with 1 mL Folin–Ciocalteu’s reagent (10%). The mixture was vortexed for 30 s. After three min, 0.8 mL of sodium carbonate solution (7.5% w/v) was added. The resulting solution was shaken and incubated for 30 min at room temperature before reading at 765 nm using a spectrophotometer. Results are expressed as mg gallic acid equivalents per gram fresh weight (mg GAE/g/FW), ethanol was used as control.
Determination of total flavonoid (TFC) Content
Total soluble flavonoid content was determined according to the method of Oriakhi36. 1 mL of the methanolic marjoram leaf extract was added to 1 mL of aluminum chloride solution (2% in methanol). The mixture was shaken vigorously and allowed to stand for 15 min at room temperature. Absorbance was measured at 430 nm. The results are expressed in mg rutin equivalents g/FW (mg RE/g/FW), using ethanol as blank.
Estimation of individual phenolic compounds by HPLC
Liquid nitrogen was employed to pulverize frozen marjoram leaves (1 g), with subsequent extraction using 10 mL methanol-chloroform solution (4:1 ratio) under ambient conditions. Extracts were sonicated for 30 min at 37°C and centrifuged at 6000 rpm for 10 min at room temperature. After phase separation, the upper layer was filtered through 0.22 μm syringe filters, and analyzed for total phenolic compounds and flavonoids. Chromatographic separation of phenolic constituents utilized an Agilent 1100 series HPLC system equipped with a reverse-phase C18 analytical column (3.0 × 250 mm, 5 μm) maintained at 25°C, with automated 20 μL sample injection. Mobile phase composition consisted of methanol (solvent A) and 2% acetic acid (v/v) (solvent B), following this gradient program: 0–2 min, 13% A; 2–7 min, 22% A; 7–15 min, 40% A; 15–25 min, 75% A; 25–35 min, 90% A; 35–40 min, 95% A. Mobile phase flow rate remained constant at 0.5 mL/min. Detection employed multiwavelength diode-array detector (DAD) monitoring with simultaneous chromatogram acquisition and standard compound calibration curves. Data processing utilized Agilent Chemstation Rev. B.02.01-SR1 (260) software. Compound identification relied on retention time comparison and UV absorption spectrum analysis. Phenolic compound concentrations were reported as mg/g plant extract based on fresh weight.
Gene expression analysis
RNA extraction and cDNA synthesis
Marjoram leaf tissue samples (100 mg) were harvested from Ch-Cu NPs-treated and control plants, then pulverized into fine powder utilizing liquid nitrogen before homogenization in 1 mL TRIzol RNA isolation reagent (Invitrogen, USA). The RNA extraction protocol followed procedures established by Ahmad37. cDNA synthesis employed 1 μg RNA using the iScript cDNA synthesis kit (Bio-Rad, Hercules, CA, USA) according to manufacturer’s protocols.
Quantitative real-time PCR (qRT-PCR)
Gene expression analysis employed qRT-PCR primers specific to phenylpropanoid biosynthesis-related genes, utilizing actin gene (GU570135.2) for normalization purposes. Primer sequences followed those reported by Kisa38 (Table 1). Reaction mixtures incorporated SYBR Green Supermix (Bio-Rad, Hercules, CA, USA) prepared according to manufacturer’s specifications. PCR amplification and detection were performed on a real-time analysis system (Rotor-Gene 6000, Qiagen, Hilden, Germany). Expression levels of target genes were calculated from qRT-PCR data employing the 2-ΔΔCt relative quantification approach outlined by Livak and Schmittgen39. Each reaction was performed in triplicate.
Statistical analysis
The collected data was subjected to one-way ANOVA analysis using the Gomez and Gomez40 procedure. The least significant difference (LSD) test was used for pairwise comparisons of treatment means at the 0.05 significance level.
Results
Characterization of Chitosan-Copper nanoparticles
UV–Vis spectra
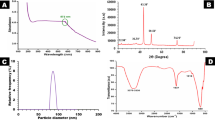
The synthesis of Ch-Cu NPs was confirmed by the formation of an absorbance peak at 585nm under UV–visible spectroscopy, as shown in Fig. 1.
UV–visible spectra indicate the Ch-Cu NPs alone and bulk chitosan showing at 580nm.
FTIR spectroscopy
FTIR analysis of Ch-Cu NPs was elucidated and presented in Fig. 2. Dominant peaks were observed at 3457.12 cm-1, 3169.75 cm-1, 2914.23 cm-1 and followed by smaller peaks in bulk Ch-Cu NPs indicated at 1613.94 cm-1, 1467.51 cm-1, and 1063.48 cm-1. Table 2 provides absorption bands and all primary functional group specifications.
FTIR spectra of Ch-Cu NPs.
TEM
TEM analysis was performed on the Ch-Cu NPs to determine the size and shape of the nanoparticles as shown in Fig. 3. The morphological study of the Ch-Cu NPs revealed that they were spherical with no signs of agglomeration. TEM indicated nanoparticle sizes ranging from 53.8 to 56.2 nm (Fig. 3).
TEM image of Ch-Cu NPs.
Isolation, identification and frequency of fungi isolated from diseased marjoram plants
Fungal isolation procedures yielded six species encompassing 85 total colonies recovered from marjoram plants exhibiting damping-off and wilt symptoms (Table 3). Three genera were represented among the isolated organisms: Alternaria sp., F. oxysporum, F. roseum, F. solani, F. semitectum, and R. solani, all obtained from naturally diseased specimens. Among infected marjoram samples acquired from various nursery facilities throughout Giza Governorate, Egypt, F. oxysporum and R. solani demonstrated the highest isolation frequencies.
Pathogenicity test
Pathogenicity evaluations demonstrated that each tested isolate F. oxysporum, F. roseum, F. semitectum, and R. solani successfully induced root rot and wilt symptom development in marjoram plants with variable severity levels, while simultaneous inoculation with F. oxysporum and R. solani produced comparable disease manifestations. F. oxysporum generated significantly elevated disease incidence (36.88%) and severity indices (21.79%) compared to R. solani, which produced 43.74% and 17.63% values, respectively. Combined inoculation using F. oxysporum and R. solani resulted in maximum disease incidence (59.14%) and severity measurements (28.47%) (Table 4).
Molecular identification
The polymerase chain reaction (PCR) was applied using two sets of primers for the identification of F. oxysporum and R. solani. Conventional PCR studied the specificity of the primers FoF1 and FoR1 for F. oxysporum. A single fragment of 340 bp was amplified when genomic DNA from F. oxysporum was used (Fig. 4). On the other hand, the specific primer pair ARSF1/ARSR1 amplified a 530 bp fragment in R. solani (Fig. 4). No other fragment was observed with PCR product from two isolates.
PCR-amplified products using the specific primers, Lane M: 100 bp DNA ladder, lane 1: F. oxysporum, lane 2: R. solani.
Antifungal Activity of Ch-Cu NPs against mycelial growth of F. oxysporum and R. solani
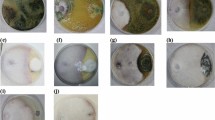
Cultivation of F. oxysporum and R. solani occurred on potato dextrose agar (PDA) medium supplemented with Ch-Cu NPs at concentrations of 25, 50, and 100 mg/L (Fig. 5). Linear growth suppression demonstrated concentration-dependent enhancement for both fungal species with increasing Ch-Cu NPs levels. Radial growth measurements of F. oxysporum revealed significant reduction across all Ch-Cu NPs-amended PDA plates relative to untreated control plates (Fig. 5). Specifically, Ch-Cu NPs at 50 mg/L concentration achieved 72.2% growth suppression of F. oxysporum, while 100 mg/L treatment produced 80.5% inhibition, representing maximum growth suppression relative to controls. Similar concentration-dependent responses emerged when R. solani was exposed to identical Ch-Cu NPs concentrations, with 100 mg/L treatment yielding 80.5% growth suppression (Fig. 5).
Effect of Ch-Cu NPs on the radial growth of F. oxysporum (A) and R. solani (B) with different concentration of Ch-Cu NPs.
Effect of Ch-Cu NPs on development of root rot and wilt disease
Collected data revealed substantial reductions in marjoram root rot and wilt disease parameters across all Ch-Cu NPs concentrations at 90 days post-treatment relative to untreated infected controls (Table 5 and Fig. 6). Treatment of diseased marjoram plants with 50 mg/L Ch-Cu NPs produced marked decreases in disease incidence and severity values (23.67% and 5.57%, respectively), with 100 mg/L Ch-Cu NPs treatment showing secondary effectiveness, demonstrating statistically significant variation between treatments. The 25 mg/L Ch-Cu NPs treatment resulted in a weak effect against root rot and wilt disease incidence and disease severity compared to higher concentrations.
Effect of Ch-Cu NPs on root rot and wilt disease on marjoram plants treated with 25, 50 and 100 mg/L of Ch-Cu NPs.
Effect of Ch-Cu NPs on TPC and TFC in marjoram leaves
Treatment with each evaluated Ch-Cu NPs concentration promoted substantial TPC accumulation in diseased marjoram plants relative to untreated healthy control specimens (Fig. 7). Maximum TPC levels (13 mg/g FW) were recorded in specimens receiving 50 mg/L Ch-Cu NPs treatment. The 100 mg/L Ch-Cu NPs application demonstrated comparable significant effects to the 50 mg/L concentration.
Effect of Ch-Cu NPs on TPC and TFC in the leaves of marjoram. The numerical values were exhibited as mean value ± standard deviation (SD), and different letters show significant differences at p ≤ 0.05.
Qualitative analysis of phenylpropanoid metabolites of marjoram
The HPLC chromatogram of the contents of major individual polyphenolic compounds in healthy marjoram plants compared with infected marjoram plants treated with 100 mg/L Ch-Cu NPs (Fig. 8). The main identified phenolic compounds were gallic acid, chlorogenic acid, vanillic acid, caffeic acid, coumaric acid, ferulic acid, benzoic acid, rutin and rosmarinic acid. The retention time of the phenylpropanoid metabolites of marjoram leaves was determined using HPLC analysis (Table 6). Also, these table appeared to show that different concentrations of Ch-Cu NPs have increased fold change for these metabolites.
HPLC chromatograms of polyphenolic compounds in control (A) and treated with 100 mg/L Ch-Cu NPs.
Application of Ch-Cu NPs significantly enhanced all phenolic compounds of infected seedlings, but notably, the major phenolic compounds present in the extract were Coumaric acid (15.87%), Ferulic acid (14.87%), rosmarinic acid (10.37%), Caffeic acid (11.76%), Vanillic acid (9.15%), Rutin (7.33%), Chlorogenic acid (6.31%), Gallic acid (5.97%) and Benzoic acid (5.69%).
These nine compounds accounted for a significant proportion, totaling 87.32% of the total identified compounds (96.78%). The application of Ch-Cu NPs at the levels of 25, 50, and 100 mg/L elevated the promotion of major phenolic compound content in infected marjoram plants to a higher level than in healthy plants (Fig. 9). The use of Ch-Cu NPs increased the levels of major phenolic compounds; at 25 mg/L Ch-Cu NPs, the range increased from 1.01 (rutin) to 1.91 (rosmarinic acid); at 50 mg Ch-Cu NPs, the range increased from 1.05 (rutin) to 1.91 (caffeic acid); and at 100 mg/L, the range increased from 1.03 (rutin) to 1.80 (caffeic acid).
Effect of Ch-Cu NPs on the phenolic compound profiles identified in the leaves of marjoram. The numerical values were exhibited as mean value ± standard deviation (SD), and different letters show significant differences at p ≤ 0.05.
Transcript expression of phenylpropanoid biosynthetic pathway genes
Analysis of twelve target genes (4CL, C4H, CAD, COMT, CVOMT, EGS, EOMT, FLS, HPPR, PAL, RAS, and TAT) in marjoram foliar tissue investigated correlations between gene expression profiles and phenolic compound accumulation following treatment with varying Ch-Cu NPs concentrations (Fig. 10). Assessment focused on transcriptional changes in C4H and PAL, which encode initial enzymatic components within the phenylpropanoid biosynthetic pathway. Treatment with 50 and 100 mg/L Ch-Cu NPs induced substantial PAL up-regulation, achieving 3.93-fold and 3.33-fold increases respectively, relative to infected controls. Similarly elevated C4H expression emerged, displaying 4.51-fold and 4.34-fold enhancement at 50 and 100 mg/L Ch-Cu NPs treatments (Fig. 10). C4H and PAL genes belonged to the first group of genes, which also includes four genes (C4H, CVOMT, RAS, and TAT). This group has a highly significant up-regulation with a range from 3 to 4.5-fold change. The second group included four genes (CAD, COMT, EGS, EOMT) with significant up-regulation with a range from 1.8 to 2.5-fold change. For example, EGS was significantly up-regulated in 50 and 100 mg/l Ch-Cu NPs treatment plants, with transcript level 1.89–2.07-fold higher than the control (Fig. 10).
Effect of Ch-Cu NPs on the relative expression levels of phenylpropanoid biosynthesis-related genes in the leaves of marjoram (i) 4CL, (ii) C4H, (iii) CAD, (iv) COMT, (v) CVOMT (vi) EGS, (vii) EOMT (viii) FLS, (ix) HPPR, (x) PAL, (xi) RAS and (xii) TAT. The numerical values were exhibited as mean value ± standard deviation (SD), and different letters above columns show significant differences at p ≤ 0.05.
On the other hand, the obtained data indicated that the FLS and HPPR genes were not induced by the 100 mg/l treatment, with down-regulation with relative expression levels of − 0.83 and − 0.25 for 100 mg/l Ch-Cu NPs. These genes belong to the third group (Fig. 10).
Among the examined genes, 83% (10 genes) demonstrated elevated transcript levels across all treatment conditions relative to control samples, with particularly pronounced increases observed at Ch-Cu NPs concentrations of 50 and 100 mg/l. The PAL gene exhibited the most substantial response, achieving a 4.5-fold increase in relative expression level following treatment with 50 mg/l Ch-Cu NPs.
The twelve genes’ expression of all treatments had different expression patterns in diseased marjoram plants treated with 25, 50, and 100 mg/l Ch-Cu NPs. The highest expressions of studied genes were recorded at 50 mg/l Ch-Cu NPs C4H, CAD, PAL, CVOMT, RAS, and TAT, as evidenced by the heatmap of the selected genes (Fig. 11).
Heatmap for selected genes’ expression linked to diseased marjoram plants treated with 25, 50, and 100 mg/l Ch-Cu NPs. The scale color bar in the picture represents low (blue) to high (red) gene expression.
Discussion
This investigation validated Ch-Cu NPs synthesis from a commercial source through UV–visible spectrophotometry, FTIR spectroscopy, and TEM analysis. UV–visible spectrophotometric study confirmed the synthesis of Ch-Cu NPs as well. A peak was observed at 585 nm. Absorption bands in the wavelength range of 570–600 nm for chitosan-coupled copper NPs were reported earlier41,42. FTIR analysis identified multiple functional groups at wavelengths 3457.12, 3169.75, 2914.83, 1613.94, 1467.51 and 1063.48 cm-1. These spectral patterns correspond with previously reported observations23,42, corroborating the present results. TEM analysis demonstrated exceptional capability in characterizing nanoparticle dimensions, distribution patterns, and morphology through its superior resolution capabilities. TEM measurements revealed Ch-Cu particle dimensions between 53.8 and 56.2 nm. These measurements align with previously published data16,43. Egypt cultivates marjoram (Origanum majoranum L.) as a significant aromatic crop species across numerous geographical locations within various governorates. Primary cultivation regions encompass Giza, Beni Suef, and Minya governorates. Marjoram cultivation areas have experienced root rot and wilt disease outbreaks, causing diminished vegetative development and reduced plant populations, thereby decreasing essential oil production from harvested material44,45. Our result findings, three genera comprising six fungal species were recovered from naturally diseased plants: Alternaria sp., F. oxysporum, F. roseum, F. solani, F. semitectum, and R. solani. Diseased marjoram specimens obtained from various nurseries throughout Giza Governorate, Egypt yielded F. oxysporum and R. solani as the predominant isolates. These observations align with previous research44,46, which documented soil-borne marjoram diseases in Egypt and.
their fungal pathogens. Frequency analysis and pathogenicity testing in this investigation established F. oxysporum and R. solani as causative organisms for marjoram root rot and wilt diseases within Egypt. Literature records indicate Ali et al.47 first documented R. solani and F. oxysporum on Egyptian marjoram plants, with subsequent isolations by48,49. Italian researchers isolated this fungus from wild marjoram in 200750. F. oxysporum and F. solani were recovered by51 in Argentina and49 in Italy. Italian isolation of R. solani occurred in 201352. F. oxysporum and R. solani are plant pathogenic fungi that have caused large scale disease epidemics leading to significant biological and economical losses in many important agricultural crops worldwide. Early detection of pathogens in planting material is one of the key steps in the management of plant diseases. The proposed method is very useful and efficient for the identification of F. oxysporum. In addition to that, this method practically can be used for diagnosis, epidemiology studies, and management strategies of root rot and wilt of marjoram33,34. Fungal ecological research has incorporated molecular identification via DNA barcoding as a fundamental and indispensable component over the past two decades53. The internal transcribed spacer (ITS) region of nuclear DNA is the universal and best DNA barcoding marker to identify individual taxa and environmental (complex) samples, and it is now accepted as the official primary fungal barcoding marker54. Fungal ribosomal DNA’s ITS region exhibits substantial variability crucial for PCR-based species differentiation, including Fusarium spp.55, Aspergillus spp.56, R. solani anastomosis groups57, and Alternaria spp58.
Precise and expeditious pathogen identification represents a fundamental component in disease epidemiology research and remains critical for developing effective management strategies. Our investigation developed conventional PCR methodology for molecular identification, employing a PCR approach utilizing previously established internal transcribed spacers (ITS1 and ITS2)59,60,61. Earlier research demonstrated successful antifungal properties of CH@CuO NPs tested at varying concentrations (1, 25, and 50 mg/L) against pathogenic F. oxysporum wild isolates affecting tomatoes. The antifungal activity was shown to increase with increasing CH@CuO NPs concentrations in a concentration dependent manner. The CH@CuO NPs at 50 mg/L concentration were shown to have better antifungal activity than that of a commercial chemical fungicide “Kocide 2000” at 2.5 g/L concentration with percentage inhibitions of 96.9% and 77.3%13. Another study also showed a maximum inhibition of Botrytis cinerea from tomato at 250 mg/L concentration of CH@CuO NPs (92%)62. These observations align with existing research demonstrating antimicrobial pathogen suppression by nanocarrier systems and CuO nanoparticles (NPs)18, establishing their potential as synthetic fungicide alternatives for crop fungal pathogen management. Such approaches could minimize dependence on hazardous fungicides near food production areas63,64. Evidence indicates CH@CuO NPs represent viable substitutes for conventional fungicides in crop protection applications.
Research analysis revealed Ch-Cu NPs enhancement of mRNA transcript levels for target genes associated with phenylpropanoid biosynthesis and metabolism within Ch-Cu NPs-treated marjoram leaves, corresponding to polyphenol accumulation patterns. Plant polyphenolic compounds functioning as secondary metabolites contribute significantly to defensive mechanisms against microbial pathogens. Polyphenol biosynthetic compound production occurs primarily through the phenylpropanoid pathway. Co-relations among the patterns of accumulation of these compounds and the levels of resistance suggest that the phenylpropanoids perform critical protective roles against various microbial pathogens. Expression of twelve genes which code for the key enzymes which regulate the polyphenol biosynthetic pathway were compared. Unlike conventional pesticides targeting pathogens directly, NPs demonstrate plant immunity induction through defense-related gene transcriptional activation, phytohormone signaling regulation, and enhancement of induced systemic resistance (ISR) and systemic acquired resistance (SAR)65. Combining the above-mentioned mechanisms has a more profound and long-lasting defensive effect on plant resistance to pathogens. Therefore, NP-based disease management strategies are potential integral components in the maintenance of sustainable agricultural disease management66. Transcriptomics studies showed that metal NPs such as Ag-NPs, Cu-NPs, and ZnO-NPs induce accumulation of phenolic compounds, PR proteins, and antioxidant enzymes. This is done by modulating genetic expressions to improve plant defense. Genetic expression alterations in plants were through secondary metabolite biosynthesis and ROS production to boost plant resistance to pathogens9,67. Research identifies additional pathogen invasion deterrent mechanisms of silica nanoparticles through chemical barrier formation, including enhanced defense-related gene expression and phenylpropanoid biosynthesis pathway support, representing fundamental plant disease resistance pathways68. F. solani-challenged plants treated with Si NPs at 150 mg/L concentration exhibited 93.2% PAL gene expression elevation compared to controls. The 4CL gene expression increased 96.8%, while CHS gene expression levels rose 98.7% [68]. Selenium nanoparticles stimulate phenylpropanoid metabolism directing antimicrobial phenolic acid synthesis against Fusarium graminearum. This nanoparticle-mediated strategy integrates seed vigor enhancement with innate immunity strengthening, presenting sustainable maize protection methods69.
Conclusion
In summary, the current study is the first systematic study to identify phenylpropanoid biosynthesis‑related genes for qRT-PCR studies in marjoram plants against root rot and wilt diseases treated with Ch-Cu NPs. C4H, CAD, PAL, CVOMT, RAS, and TAT genes belonged to the first group, which had a highly significant up-regulation with a range from 3 to 4.5-fold change indices in diseased marjoram plants treated with 50 and 100 mg/l Ch-Cu NPs. The second group included four genes (CAD, COMT, EGS, and EOMT) with significant up-regulation with a range from 1.8 to 2.5-fold change. On the other hand, the obtained data indicated that the FLS and HPPR genes were not induced by the 100 mg/l treatment, with down-regulation with relative expression levels of − 0.83 and − 0.25 for 100 mg/l Ch-Cu NPs. These genes belong to the third group. In conclusion, the examined genes, 83% (10 genes), demonstrated elevated transcript levels across all Ch-Cu NPs treatment conditions relative to control samples, with particularly pronounced increases observed at Ch-Cu NPs concentrations of 50 and 100 mg/l. We need new research to shed light on the expression of numerous genes linked in the biosynthesis pathways of plant secondary metabolites having a regulation function in plants.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Ficke, A., Cowger, C., Bergstrom, G. & Brodal, G. Understanding yield loss and pathogen biology to improve disease management: Septoria nodorum blotch-a case study in wheat. Plant Dis. 102, 696–707. https://doi.org/10.1094/PDIS-09-17-1375-FE (2018).
Gullino, M. L., Albajes, R., Al-Jboory, I., Angelotti, F., Chakraborty, S. & Garrett, K. A. et al. Scientific Review of the Impact of Climate Change on Plant Pests. Scientific review of the impact of climate change on plant pests-A global challenge to prevent and mitigate plant pest risks in agriculture, forestry and ecosystems. Presented at Secretariat of the International Plant Protection Convention, hosted by FAO (2021).
Sarrocco, S., Mauro, A. & Battilani, P. Use of competitive filamentous fungi as an alternative approach for mycotoxin risk reduction in staple cereals: State of art and future perspectives. Toxins 11, 701 (2019).
Granzow, S. et al. The effects of cropping regimes on fungal and bacterial communities of wheat and faba bean in a greenhouse pot experiment differ between plant species and compartment. Front. Microbiol. 8, 902 (2017).
Stoddard, F., Nicholas, A. H., Rubiales, D., Thomas, J. & Villegas-Fernández, A. Integrated pest management in faba bean. Field Crops Res. 115, 308–318 (2010).
Levenfors, J. Soil-borne pathogens in intensive legume cropping-aphanomyces spp. and root rots, Doctoral thesis. Swedish University of Agricultural Sciences, Uppsala (2003).
Rosenzweig, A. Iglesius, X.-B. & Yang, P. R. Epstein, Chivian E. Climate change and extreme weather events-Implications for food production, plant diseases, and pests 2, 90–104 (2001).
Abdelaziz, A. M. et al. Inhibition of Aspergillus flavus growth and aflatoxin production in Zea mays L. using endophytic Aspergillus fumigatus. J. Fungi 8, 482 (2002).
Abdelaziz, A. M. et al. Protective role of zinc oxide nanoparticles based hydrogel against wilt disease of pepper plant. Biocatal. Agric. Biotechnol. 35, 1020833.1 (2021).
Saleem, H., Akbar, M., Anar, M., Chaudhary, H. J. & Munis, M. F. H. Biogenic iron oxide nanoparticles (Fe2O3 NPs) adeptly diminish leaf blight of Rosa chinensis by exhibiting higher expression of RcWRKY29 and increasing antioxidant enzymatic activities. S. Afr. J. Bot. 171, 337–346 (2024).
Abdelfattah, N. A. et al. Influence of biosynthesized magnesium oxide nanoparticles on growth and physiological aspects of cowpea (Vigna unguiculata L.) plant, cowpea beetle, and cytotoxicity. Biotechnol. J. https://doi.org/10.1002/biot.202300301 (2023).
El-Abeid, S. E. et al. Antifungal activity of copper oxide nanoparticles derived from Zizyphus spina leaf extract against Fusarium root rot disease in tomato plants. J. Nanobiotechnology 12(1), 28. https://doi.org/10.1186/s12951-023-02281-8 (2024).
Mosa, M. A. & El-Abeid, S. E. Chitosan-loaded copper oxide nanoparticles: A promising antifungal nanocomposite against Fusarium wilt disease of tomato plants. Sustainability 15, 14295. https://doi.org/10.3390/su151914295 (2023).
Gaba, S., Rai, A. K., Varma, A., Prasad, R. & Goel, A. Biocontrol potential of mycogenic copper oxide nanoparticles against Alternaria brassicae. Front. Chem. 10, 966396. https://doi.org/10.3389/fchem.2022.966396 (2022).
Consolo, V. F., Torres-Nicolini, A. & Alvarez, V. A. Mycosinthetized Ag, CuO and ZnO nanoparticles from a promising Trichoderma harzianum strain and their antifungal potential against important phytopathogens. Sci. Rep. 24, 20499. https://doi.org/10.1038/s41598-020-77294-6 (2020).
Choudhary, R. C. et al. Cu-chitosan nanoparticle boost defense responses and plant growth in maize (Zea mays L.). Sci. Rep. 7, 9754. https://doi.org/10.1038/s41598-017-08571-0 (2017).
Katiyar, D., Hemantaranjan, A. & Singh, B. Chitosan as a promising natural compound to enhance potential physiological responses in plant: A review. Indian J. Plant Physiol. 20, 1–9 (2015).
Goyal, P., Chakraborty, S. & Misra, S. K. Multifunctional Fe3O4-ZnO nanocomposites for environmental remediation applications. Environ. Nanotechnol. Monit. Manag. 10, 28–35 (2018).
Popova, E., Domnina, N., Kovalenko, N., Sokornova, S. & Tyuterev, S. Effect of chitosan and vanillin-modified chitosan on wheat resistance to spot blotch. Appl. Biochem. Microbiol. 52, 537–540 (2016).
Sathiyabama, M., Bernstein, N. & Anusuya, S. Chitosan elicitation for increased curcumin production and stimulation of defence response in turmeric (Curcuma longa L.). Ind. Crops Prod. 89, 87–94 (2016).
Sathiyabama, M. & Parthasarathy, R. Biological preparation of chitosan nanoparticles and its in vitro antifungal efficacy against some phytopathogenic fungi. Carbo. Polymers 151, 321–325 (2016).
Chandra, S. et al. Chitosan nanoparticles: A positive modulator of innate immune responses in plants. Sci. Rep. 5, 15195 (2015).
Saharan, V. et al. Synthesis and in vitro antifungal efficacy of Cu-chitosan nanoparticles against pathogenic fungi of tomato. Int. J. Biol. Macromol. 75, 346–353 (2015).
Stoimenov, P. K., Klinger, R. L., Marchin, G. L. & Klabunde, K. J. Metal oxide nanoparticles as bactericidal agents. Langmuir 18, 6679–6686 (2002).
Nelson, E. B. & Boehm, M. J. Compost-induced suppression of turf grass diseases. Bio. Cycle 43, 51 (2002).
Ahmed, H. Z., Armanious, H. A. H., Abdel-Moneim, M. L. & Gala, A. A. Control of root rot and wilt diseases of marjoram by using some mineral compounds and antioxidants. Minia J. Agric. Res. Dev. 37, 141–156 (2017).
Booth, C. The Genus Fusarium 1–237 (Commonwealth mycological Institute, 1971).
Barnett, H. & Hunter, B. Illustrated Genera of Imperfect Fungi 4th edn. (The American Phytopathological Society (1998).
Summerell, B. A., Salleh, B. & Leslie, J. F. A utilitarian approach to Fusarium identification. Plant Dis. 87, 117–128 (2003).
Gilman, J. C. A Manual of Soil Fugi 450 (Iowa State university, 1957).
El-Mohamedy, R. S. R., Abdalla, A. M. & Adam, S. M. Preliminary studies on response of Moringa oleifera plants to infection with some soil-borne plant pathogenic fungi. Int. J. Curr. Microbiol. Appl. Sci. 3, 389–397 (2014).
Amer, O. E., Mahmoud, M. A., El-Samawaty, A. M. A. & Sayed, S. R. M. Non liquid nitrogen-based-method for isolation of DNA from filamentous fungi. Afr. J. Biotechnol. 10, 14337–14341 (2011).
Duan, C. et al. Identification of pathogenic Fusarium spp. causing maize ear rot and potential mycotoxin production in China. Toxins 8, 186. https://doi.org/10.3390/toxins8060186 (2016).
Dubey, S. C., Tripathi, A., Upadhyay, B. K. & Kumar, A. Development of conventional and real time PCR assay for detection and quantification of Rhizoctonia solani infecting pulse crops. Biologia 71, 133–138. https://doi.org/10.1515/biolog-2016-0027 (2016).
Hossain, M. A. & Shah, M. D. A study on the total phenols content and antioxidant activity of essential oil and different solvent extracts of endemic plant Merremia borneensis. Arab. J. Chem. 8, 66–71 (2015).
Oriakhi, K. et al. Comparative antioxidant activities of extracts of Vernonia amygdalina and Ocimum gratissimum leaves. J. Agric. Sci. 6, 13–20 (2014).
Ahmad, J. et al. Comparative assessment of four RNA extraction methods and modification to obtain high-quality RNA from Parthenium hysterophorus leaf. 3 Biotech 7, 373–386 (2017).
Kisa, D. et al. The interactive effect of aromatic amino acid composition on the accumulation of phenolic compounds and the expression of biosynthesis-related genes in Ocimum basilicum. Physiol. Mol. Biol. Plants 27(9), 2057–2069. https://doi.org/10.1007/s12298-021-01068-1 (2021).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCt method. Methods 25, 02–408 (2001).
Gomez A. K. & Gomez. Statistical Procedures for Agriculture Research 2nd Ed. John Wiley & sons New Youk, 680 PP (1980).
Mallick, K., Witcomb, M. J. & Scurrell, M. S. In situ synthesis of copper nanoparticles and poly (o-toluidine): A metal-polymer composite material. Eur. Polym. J. 42, 670–675 (2006).
Vanti, G. L. et al. Synthesis and application of chitosan-copper nanoparticles on damping off causing plant pathogenic fungi. Int. J. Biol. Macromol. https://doi.org/10.1016/j.ijbiomac.2019.11.179 (2018).
Hidangmayum, A. & Dwivedi, P. Chitosan based nanoformulation for sustainable agriculture with special reference to abiotic stress: A review. J. Polym. Environ. 30, 1264–1283 (2022).
Hilal, A. A., Heweidy, M. A., El-Deeb, A. A. & El-Ghareeb, L. A. Root–rot and wilt diseases affecting marjoram in Egypt. Proceedings of 6th Congress Phytopathology, Cairo, Egypt, 3, 402–417 (1990).
Hassanin, M. Pathological studies on some important fungal diseases of marjoram and their management. M.Sc. Thesis, Faculty of Agriculture, Al-Azhar University, Cairo, Egypt, 137pp (2007).
El-Gebaly, M. M. Studies on root-rot and wilt disease of marjoram Ph.D. Thesis, Fac.Agric., Al-AzharUniversity, Cairo, Egypt, 153pp (1988).
Ali, M. D. H., Morsi, A. A. & El-Sherif, M. A. Identification of agricultural research carried out in Egypt (1900–1970). I. Plant pathology. N.I.D.C.,Nat. Acad. Sci.Tech , 489 pp. (in Arabic) (1972).
Eisa, E. A., Meligy, M. M. & Ziedan, H. E. Application of composts and potassium sulphate on root rot incidence, morphological growth, yield components, oil content and constitutes of marjoram plants (Majorana hortensis L.). Biocatal. Agric. Biotechnol. 42, 102334 (2022). https://doi.org/10.1016/j.bcab.2022.102334.
Abdel-Wahed, G. A., Abd El-Aziz, A. R. M., Ahmed, N. E., Gad, M. A. & Shehata, A.S.F. Biological Control of marjoram root-rot using kombucha and natural antagonists with first report of Fusarium equiseti in Egypt. Egypt. J. Phytopathol. 53, 178–193 (2025). https://doi.org/10.21608/EJP.2025.431979
Garibaldi, A., Minuto, A. & Gullino, M. L. First report of white mold caused by Sclerotini asclerotiorum on Origanum vulgare and Taraxacum officinale in Italy. Plant Dis. 91, 1360 (2007).
Gaetan, S. A., Madia, M. S. & Perez, A. Recent outbreak of Fusarium crown and root rot caused by Fusarium solanion marjoram in Argentina. Australas. Plant Dis. Notes 2, 15–16 (2007).
Garibaldi, A., Bertetti, D., Pensa, P., Poli, A. & Gullino, M. L. First Report of Web Blight on Oregano (Origanum vulgare L.) caused by Rhizoctonia solani AG-1-IB in Italy. Plant Dis. 97, 1119 (2013).
Anderson, I. & Cairney, J. Diversity and ecology of soil fungal communities: Increased understanding through the application of molecular techniques. Environ. Microbiol. 6, 769–779. https://doi.org/10.1111/j.1462-2920.2004.00675.x (2004).
Nilsson, R., Ryberg, M., Abarenkov, K., Sjökvist, E. & Kristiansson, E. The ITS region as a target for characterization of fungal communities using emerging sequencing technologies. FEMS Microbiol. Lett. 296, 97–101. https://doi.org/10.1111/j.1574-6968.2009.01618.x (2009).
Campos, M. D., Patanita, M., Campos, C., Materatski, P., Varanda, C. M. R., Brito, I. & Félix, M. D. R. Detection and quantification of Fusarium spp. (F. oxysporum, F. verticillioides, F. graminearum) and magnaporthiopsis maydis in maize using real-time PCR targeting the ITS region. Agronomy 9, 45 (2019). https://doi.org/10.3390/agronomy9020045.
Rozaliyani, A. et al. Unravelling the molecular identification and antifungal susceptibility profiles of Aspergillus spp. isolated from chronic pulmonary aspergillosis patients in Jakarta, Indonesia: The emergence of cryptic species. J. Fungi 8, 411. https://doi.org/10.3390/jof8040411 (2022).
Amancio, L. C. S. et al. PCR with specific primers for detection of Rhizoctonia solani anastomosis groups reveals lack of specificity. J. Phytopathol. 173, e70162 (2025).
Baramidze, V. et al. A barcoded ITS primer-based nanopore sequencing protocol for detection of Alternaria species and other fungal pathogens in diverse plant hosts. J. Fungi 11, 249. https://doi.org/10.3390/jof11040249 (2025).
Sneh, B., Jabaji-Hare, S., Neate, S. & Dijst, G. Rhizoctonia Species: Taxonomy, Molecular Biology, Ecology (Kluwer Academic Publishers, Dordrecht, 1996).
Liu, B. et al. Advances in crop wilt disease. J. Xiamen Univ. 43, 47–58 (2004).
Fang, Y. & Ramasamy, R. P. Current and prospective methods for plant disease detection. Biosensors 5, 537–561 (2015).
Ismail, A. M., Mosa, M. A. & El-Ganainy, S. M. Chitosan-decorated copper oxide nanocomposite: Investigation of its antifungal activity against tomato gray mold caused by Botrytis cinerea. Polymers 15, 1099. https://doi.org/10.3390/polym15051099 (2023).
Karthik, R. et al. A facile graphene oxide based sensor for eEelectrochemical detection of prostate anti-cancer (anti-testosterone) drug Flutamide in biological samples. RSC Adv. 7, 25702–25709 (2017).
Dreyer, D. R., Park, S., Bielawski, C. W. & Ruoff, R. S. The chemistry of graphene oxide. Chem. Soc. Rev. 39, 228–240 (2010).
Kumari, M. et al. Omics-based mechanistic insight into the role of bioengineered NPs for biotic stress amelioration by modulating plant metabolic pathways. Front. Bioeng. Biotechnol. 8, 242. https://doi.org/10.3389/fbioe.2020.00242 (2020).
Guo, Z. et al. Multiple pathways revealing the CeO2 nanoparticle-biostimulant-Based “stress training” strategy for enhanced Medicago sativa L antiviral capability. ACS Nano https://doi.org/10.1021/ACSNANO.4C10637 (2025).
Elmer, W. H. et al. Foliar application of copper oxide NPs suppresses Fusarium wilt development on Chrysanthemum. Environ. Sci. Technol. 55, 10805–10810. https://doi.org/10.1021/acs.est.1c02323 (2021).
Zhou, J. et al. Silica nanoparticles enhance the disease resistance of ginger to rhizome rot during postharvest storage. Nanomaterials (Basel) 12, 1418. https://doi.org/10.3390/nano12091418 (2022).
Dong, Q. et al. Nano-priming with selenium nanoparticles reprograms seed germination, antioxidant defense, and phenylpropanoid metabolism to enhance Fusarium graminearum resistance in maize seedlings. J. Adv. Res. https://doi.org/10.1016/j.jare.2025.05.057 (2025).
Acknowledgements
The authors extend their appreciation to the Researchers supporting project number (ORF-2026-224)King Saud University, Riyadh, Saudi Arabia.
Funding
This research project was funded by the Researchers supporting Project number (ORF-2026-224) of King SaudUniversity, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
Mohamed A. Mahmoud: Writing – review & editing, Writing – original draft, Visualization, Methodology, Investigation, Funding acquisition, Data curation, Conceptualization. Abeer R. M. Abd El-Aziz: Writing – review & editing, Methodology, Investigation, Data curation. Eman W. R. Ghebrial: Writing – review & editing, Methodology, Investigation. Fatimah S. Al-khattaf: Project administration, Supervision and Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Al-khattaf, F.S., Mahmoud, M.A., Ghebrial, E.W.R. et al. Eco-friendly managing of root rot and wilt diseases in marjoram plants by chitosan-copper nanoparticles: enhancing plant immunity and stimulation of resistance genes. Sci Rep 16, 9232 (2026). https://doi.org/10.1038/s41598-026-44092-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-44092-5