Abstract
Breakthrough infections and reinfection are key factors leading to recurrent epidemic waves. However, sustained control strategies can lead to unnecessary resource wastage when tacking these issues. There is an urgent need to establish dynamic intervention systems capable of rapid response and efficient resource utilization. To address the question of how breakthrough infections and reinfections affect the dynamics of the pandemic, this study develops an infectious disease model that incorporates both breakthrough infections and reinfections, and while introduces bang-bang optimal control as an efficient public health intervention strategy to provide a new perspective and solutions. In theoretical analysis, we derive basic reproduction number via next-generation matrix method, prove the global stability of the disease-free equilibrium, and establish sufficient conditions for the existence of multiple endemic equilibria and the occurrence of backward bifurcation. Numerical simulations further confirm the critical role of breakthrough infections and reinfection in disease persistence and recurrent outbreaks. In control strategy research, we prove the existence of bang-bang optimal solutions based on optimal control theory and demonstrate their distinct advantages in rapidly outbreaks while minimizing operational costs. Simulation results show that a combined strategy implemented under the bang-bang control—reducing transmission rates, expanding vaccine coverage, and enhancing vaccine protection—most effectively contains disease spread. This results provide both theoretical foundation and practical guidance for developing efficient control strategies against recurrent infectious disease outbreaks.
Similar content being viewed by others
Introduction
Breakthrough infection refers to the occurrence of infection by the corresponding pathogen after vaccination, indicating that the vaccine fails to confer complete immunity to the recipient1. Breakthrough infection is closely related to various immune response states such as immune failure, immune ineffectiveness, and immune decay2. Its occurrence is affected by multiple factors, including vaccine efficacy, the recipient’s own immunity, viral antigen variation, and individual differences. The occurrence of breakthrough infection makes it difficult for the vaccine protection rate to reach 100%3. Currently, vaccines for SARS-Cov-2, influenza, chickenpox, mumps and other viruses can lead to breakthrough infections4,5,6. Usually, the symptoms of breakthrough infection are milder than those of ordinary infection, and even asymptomatic infection may occur7,8.
Reinfection refers to a subsequent infection with the same virus or after recovery9,10 For example, COVID-19, influenza, HIV/AIDS, malaria, dengue fever, and other diseases all carry a risk of reinfection11,12,13,14,15 . Many people who have experienced multiple COVID-19 infections experience milder symptoms due to immunity from prior infections. Though exceptions exist primarily immunocompromised individuals, the elderly, or those with a history of severe infection16. Many people who had a severe first infection may be hospitalized or need to go to the hospital again because of reinfection17. For example, people living with HIV have a higher rate of reinfection than those without HIV18,19.
Breakthrough infections and reinfection are key factors driving disease recurrence, as evidenced by the large-scale COVID-19 waves observed from late 2021 to 2022. As the virus continues to evolve, the emergence of novel variants capable of evading existing immunity may further increase the risks of breakthrough infection and reinfection20. This study aims to investigate the impact of these factors on disease transmission dynamics and the effectiveness of control measures.
Mathematical models are indispensable for understanding the dynamics of infectious disease transmission and forecasting epidemic trajectories. In particular, models that account for breakthrough infections and reinfection have been the subject of extensive study. In infectious disease modeling, breakthrough infections have been increasingly recognized as a critical factor shaping transmission dynamics and control effectiveness. Perkins et al. developed a stochastic dengue model, demonstrating the substantial impact of breakthrough infections on vaccination effectiveness and underscoring the need to integrate this factor into such models21. Azimaqin et al. proposed an age-structured model to investigate the 2015 mumps resurgence in China, linking the outbreak to vaccine failure, seasonal effects, and shifts in population structure22. Xu et al. established a dynamic transmission model using Chinese hepatitis B data, highlighting that increasing vaccination rates could significantly enhance disease control23. Elbasha et al. introduced a vaccination model with waning immunity, showing that backward bifurcation may occur when vaccine protection is partial and its duration falls below a critical threshold24. Yuliana et al. analyzed an SIVS model under vaccine failure, concluding that breakthrough infections substantially contribute to sustained high transmission25. Finally, Jing et al. developed a COVID-19 model incorporating breakthrough infection and age structure, emphasizing that high vaccine coverage combined with effective antiviral drugs is essential to achieve disease eradication26.
In the study of reinfection in infectious disease modeling, several works have provided critical insights. Montalban et al. developed a herd immunity model incorporating individual heterogeneity and reinfection, demonstrating that epidemic growth can be effectively suppressed once the proportion of immune population exceeds the herd immunity threshold27. Rehman et al. proposed a fractional-order malaria model accounting for memory effects, relapse, and reinfection, identifying vaccination as a key factor in preventing disease recurrence28. Anggriani et al. introduced a multi-strain dengue model, showing that reinfection with the same serotype elevates both primary and secondary case numbers29. Gomes et al. compared SIR and SIS frameworks to evaluate the impact of post-recovery immunity, revealing that below the reinfection threshold, primary infections dominate with low incidence, whereas above it, reinfection drives high infection levels30. Agusto developed an Ebola model incorporating recurrence and reinfection, identifying reinfection as a trigger for backward bifurcation31. Rodrigues et al. assessed tuberculosis risks under partial immunity, indicating that the average reinfection rate may exceed that of primary infections32. In response to recurrent hepatitis B outbreaks, Megala et al. integrated reinfection with Crowley-Martin incidence and ongoing vaccination, underscoring the combined influence of vaccination, reinfection, and preventive measures on HBV transmission dynamics33.
Extensive research has confirmed the critical role of breakthrough infections and reinfection in shaping disease dynamics and complicating control initiatives. Notably, several major infectious diseases—such as COVID-19, influenza, and dengue exhibit both phenomena concurrently. Nevertheless, existing modeling frameworks have largely failed to integrate these two mechanisms into a unified structure, leaving a notable gap in the development of effective and feasible intervention strategies. To address this limitation, we develop an epidemic model that simultaneously incorporates breakthrough infections and reinfection, and employ bang-bang optimal control as the central strategy for intervention design. A key advantage of the bang-bang approach lies in its “all-or-nothing” switching nature, which offers public health authorities clear, implementable, and resource efficient phased control policies. Our study aims not only to enhance the realism of disease transmission modeling but also to provide a theoretical foundation and practical strategy for deploying intensity-varying control measures in resource-constrained scenarios.
This paper is organized as follows. Firstly, an infectious disease model with breakthrough infection and reinfection is established, and we derive the basic reproduction number, prove the local stability of the disease-free equilibrium and the existence of backward bifurcations. Secondly, the theoretical results are verified by combining numerical simulations. Bang-Bang optimal control of the minimum-time problem is applied to the model to prove the existence of the optimal solution. Finally, a simulation research of Bang-Bang control is conducted to compare various control strategies to find the optimal strategy to control the disease in the shortest time.
Mathematical modelling
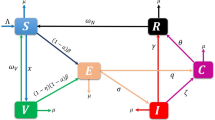
The population is divided into four categories, susceptible, vaccinated, infected, and recovered, with S(t), V(t), I(t), and R(t) representing the number of individuals in each class at time t. The disease transmission flow chart considering breakthrough infection and reinfection is shown in Fig. 1.
Flow chart of disease transmission.
According to Fig. 1, the model can be expressed as
where \(N=S+V+I+R\). The description of parameters in system (1) is given in Table 1 .
System (1) incorporates breakthrough infection and reinfection into the dynamic system by introducing two key parameters. Breakthrough infection is characterized by the parameter \(\sigma \in [0,1]\), which represents the failure rate of the vaccine. When \(\sigma =0\) , the vaccine provides complete protection, indicating no breakthrough infections; when \(\sigma =1\), vaccinated have the same risk of infection as susceptible. Reinfection is described by the parameter \(\epsilon \in [0,1]\), which represents the failure rate of natural immunity of recovered. \(\epsilon > 0\) means that recovered may be reinfected due to incomplete or weakened immunity. Furthermore, the model also includes the process of waning immunity. Vaccinated lose vaccine-induced protection at a rate of \(\theta\), and recovered lose natural immunity at a rate of \(\gamma\), both returning to the susceptible compartment. This system (1) characterizes both breakthrough infections and reinfections through the imperfectness and waning nature of immune protection.
Basic reproduction number
Lemma 1
Assuming that the initial conditions (S(0), V(0), I(0), R(0)) of system (1) are non-negative, then
is a positively invariant set for system (1), where C is a constant.
See Appendix for the proof process of Lemma 1.
Let \(s=\frac{S}{N}\), \(v=\frac{V}{N}\), \(i=\frac{I}{N}\), \(r=\frac{R}{N}\), then \(s+v+i+r=1\). Normalize system (1) and substituting \(r=1-s-v-i\) into system (1), we obtain,
Obviously, the positive invariant set of system (2) is \(\Omega =\left\{ (s,v,i)\in R_3^+|0\le s+v+i\le 1\right\}\). This paper will analyze the system (2). It explains that system (2) is equivalent to system (1).
Let the right side of system (2) equal to 0, and we can get the disease-free equilibrium \(P_0=\left( \frac{\theta }{\alpha +\theta }, \frac{\alpha }{\alpha +\theta }, 0\right)\). Next, we use the next generation matrix method to give the basic reproduction number of system (2)34. First, consider the following two vectors
The Jacobian matrices of vectors f and g at the disease-free equilibrium \(P_0\) are
The inverse of the matrix G is
Then, accoroding to the next generation approach, the basic reproduction number is given as \(R_0=\rho (FG^{-1})=\frac{\beta \left( \sigma \alpha +\theta \right) }{\eta (\alpha +\theta )}\).
Stability of disease-free equilibrium
Theorem 1
When \(R_0<1\), the disease-free equilibrium \(P_0\) of system (2) is locally asymptotically stable. When \(R_0=1\) and \(R_0 \ne R_*\) \(\left( R_*=\frac{[(\alpha \sigma +\theta )(\gamma +\eta )+\gamma \eta \sigma ](\alpha \sigma +\theta )}{\eta (\alpha +\theta )[(\alpha \sigma +\theta )\epsilon +\gamma \sigma ]}\right)\), the disease-free equilibrium \(P_0\) is a saddle node. When \(R_0=1\) and \(R_0 = R_*\), the disease-free equilibrium \(P_0\) is a stable node. When \(R_0>1\), the disease-free equilibrium \(P_0\) of system (2) is unstable.
See Appendix for the proof process of Theorem 1.
Existence of endemic equilibria
First, let the right side of system (2) equal to 0,
Solving Eq. (3) yields
Substituting formula (4) into the third equation of system (3) yields
where
Case \(R_0=1\)
Next, We first discuss case \(R_0=1\) in which
the roots of equation (5) are
-
(a)
If \(b_1<0\), then \(R_0>R_*\) and \(i_{02}<0\). Equation (5) has a unique positive root \(i_{01}\). It is obvious that \(i_{01}\in [0,1]\). Then system (2) has an endemic equilibrium \(P_{3}\) in \(\Omega\), where
$$\begin{aligned} P_{3}=\left( \frac{(\beta i_{01} \sigma +\theta )\gamma (1-i_{01})}{\beta ^2i_{01}^2\sigma +\beta (\alpha \sigma +\gamma \sigma +\theta )i_{01}+\gamma (\alpha +\theta )}, \frac{\alpha \gamma (1-i_{01})}{\beta ^2i_{01}^2\sigma +\beta (\alpha \sigma +\gamma \sigma +\theta )i_{01}+\gamma (\alpha +\theta )}, i_{01}\right) . \end{aligned}$$ -
(b)
If \(b_1=0\), then \(R_0 = R_*\), that is \((\alpha \sigma +\theta )(\gamma +\eta )=\frac{\eta (\alpha +\theta ) (\alpha \sigma +\theta )\epsilon +\alpha \gamma (\sigma -\sigma ^2)}{\alpha \sigma +\theta }\), \(b_2=\frac{(\alpha +\theta )^2\eta ^2}{(\alpha \sigma +\theta )^3} \left[ \frac{\alpha \gamma \sigma (\sigma -\sigma ^2)}{\alpha \sigma +\theta } +(\alpha \sigma +\theta )^2\epsilon \right] .\) Therefore, \(b_2\ge 0\). Furthermore, it can be seen that Eq. (5) does not have any positive roots.
-
(c)
If \(b_1>0\), then \(R_0 < R_*\), \((\alpha \sigma +\theta )(\gamma +\eta )>\frac{\eta (\alpha +\theta ) (\alpha \sigma +\theta )\epsilon +\alpha \gamma (\sigma -\sigma ^2)}{\alpha \sigma +\theta }\). So
$$\begin{aligned} b_2>\frac{(\alpha +\theta )^2\eta ^2}{(\alpha \sigma +\theta )^3}\left[ \frac{\alpha \gamma \sigma (\sigma -\sigma ^2)}{\alpha \sigma +\theta } +(\alpha \sigma +\theta )^2\epsilon \right] \ge 0. \end{aligned}$$Therefore, Eq. (5) does not have any positive roots.
Case \(R_0<1\)
When \(R_0<1\), \(b_0>0\). The necessary and sufficient conditions for \(b_1=0\) is \(R_0=R_*\).
-
(a)
If \(R_*>1\), then \(0<R_0<1<R_*\), \(b_1>0\). Furthermore \(\beta <\frac{(\alpha \sigma +\theta ) (\gamma +\eta )+\gamma \eta \sigma }{(\alpha \sigma +\theta )\epsilon +\gamma \sigma }\),
$$\begin{aligned} \begin{aligned}&\quad (-\beta \epsilon +\alpha \epsilon +\eta +\gamma )\sigma +\epsilon \theta \\&>(\frac{1}{(\alpha \sigma +\theta )\epsilon +\gamma \sigma }\left\{ (\alpha \sigma +\theta )^2\epsilon ^2+\gamma \sigma (\alpha \sigma +\theta )\epsilon +\gamma ^2\sigma ^2 +\eta \gamma \sigma ^2(1-\epsilon )\right\} \ge 0. \end{aligned} \end{aligned}$$Therefore, \(b_2\ge 0\). It can be seen that Eq. (5) does not have any positive roots.
-
(b)
If \(R_0<R_*=1\), then \(b_1=0\), further \(b_2\ge 0\), so Eq. (5) does not have positive roots.
-
(c)
If \(R_*<1\).
When \(0<R_0<R_{*}<1\), \(b_1>0\), it has \(b_2\ge 0\). So Eq. (5) does not have positive roots.
When \(0<R_*<R_{0}<1\), \(b_1<0\), taking partial derivatives of both sides of Eq. (5) with respect to i, we can obtain
The two real roots of Eq. (6) are
Then the necessary and sufficient conditions for the existence of positive roots of Eq. (5) is \(b_3i_{11}^3+b_2i_{11}^2+b_1i_{11}+b_0\le 0\), that is, \(R_0\ge R_{**}\), where
If \(R_0<R_{**}\), \(b_3i_{11}^3+b_2i_{11}^2+b_1i_{11}+b_0>0\). Thus, Eq. (5) does not have positive roots.
When \(0<R_*<R_{0}<1\) and \(R_0=R_{**}\), it has \(b_1<0\) and \(b_3i_{11}^3+b_2i_{11}^2+b_1i_{11}+b_0=0\). Therefore, Eq. (5) has a positive root \(i_1\), as shown in Fig. 2a, and system (2) has an endemic equilibrium \(P_1\) in \(\Omega\), where
When \(0<R_*<R_{0}<1\) and \(R_0>R_{**}\), it has \(b_1<0\) and \(b_3i_1^3+b_2i_1^2+b_1i_1+b_0<0\). Then, Eq. (5) has three different real roots, as shown in Fig. 2b. One of them is negative, and the other two are positive real roots, namely \(i_2\) and \(i_3\).
When \(R_*<R_0<1\), the existence of the root of Eq. (5) under different conditions. (a) \(R_0=R_{**}\), (b) \(R_0>R_{**}\).
Therefore, system (2) has two endemic equilibrium \(P_2\) and \(P_3\) respectively
When \(i=1\), \(F(1)=\eta \left[ \beta ^2\sigma +(\alpha \sigma +\gamma \sigma +\theta )\beta +\gamma (\alpha +\theta )\right] >0\), so \(i_2,i_3\in [0,1]\).
Case \(R_0>1\)
In this section, we will discuss the existence of endemic equilibria in system (2) when \(R_0>1\). If \(R_0>1\), then \(b_0<0\). It is clear that Eq. (5) has at least one positive root.
-
(i)
If \(R_*\ge R_0>1\), then \(b_1>0\), \(b_2>0\). As shown in Fig. 3b, c, in the positive semi-axis region of the x-axis, F(i) will monotonically increase from \(b_0\) and cross the x-axis into the first quadrant, so Eq. (5) has a positive root.
-
(ii)
If \(R_*<R_0\), then \(b_1<0\). As shown in Fig. 3a, in the positive half-axis region of the x-axis, F(i) is a concave function. It initially decreases monotonically from \(b_0\)(\(b_0<0\)) to a limit, and then increases to a positive value. It is easy to see that Eq. (5) has only one positive root \(i_1^*\).
When \(i=1\), since \(F(1)>0\). In summary, when \(R_0>1\), system (2) has a endemic equilibrium \(P_3\) in \(\Omega\).
When \(R_0>1\), the existence of the root of Eq. (5) under different conditions. (a) \(R_*<R_0\), (b) \(R_*>R_0\), (c) \(R_*=R_0\).
Stability of endemic equilibria
Theorem 2
When \(R_{**}<R_{0}<1\) and \(R_*<R_0\), the endemic equilibrium \(P_2\) of system (2) is unstable.
See Appendix for the proof process of Theorem 2.
Theorem 3
If \(\sigma =0\), when \(R_*\ge 1\) and \(R_{0}>1\), or \(R_{**}<R_{0}\), \(R_*<R_0\) and \(R_*<1\), the endemic equilibrium \(P_3\) of system (2) is locally asymptotically stable.
See Appendix for the proof process of Theorem 3.
Next, the existence and stability conditions of equilibria of system (2)under different conditions are summarized in Table 2.
Numerical simulation
In this section, we will verify the theoretical results in “Stability of endemic equilibria” through numerical simulations. From Fig. 4, we can see that when \(R_0=1\) and \(R_*\ge 1\), the disease-free equilibrium \(P_0\) is stable. When \(R_0=1\) and \(R_*<1\), the disease-free equilibrium \(P_0\) is unstable, and the endemic equilibrium \(P_{3}\) is stable. From Fig. 5, we can see that when \(R_0<1\), \(R_*\ge 1\) or \(R_*<1\) and \(R_{**}>R_0\), the disease-free equilibrium \(P_0\) is stable. When \(R_*<1\) and \(R_{**}=R_0<1\), the disease-free equilibrium \(P_0\) and the endemic equilibrium \(P_{1}\) are bistable. When \(R_*<1\) and \(R_{**}<R_0<1\), the endemic equilibrium \(P_2\) is unstable, \(P_0\) and \(P_{3}\) are bistable.
When \(R_0=1\), the phase diagram of system (2) under different conditions. (a) \(R_*=1\). (b) \(R_*<1\). (c) \(R_*>1\). The red dots represent the disease-free equilibrium \(P_0\), and the green dots represent the endemic equilibrium \(P_{3}\). The values of the parameters are given in the text, as well as in other figures.
Next, we use the Matcont package in Matlab to draw the bifurcations diagram of system (2). We set the parameters as \(\beta =0.25\), \(\epsilon =0.8\), \(\sigma =0.1\), \(\theta =0.001\), \(\gamma =0.05\), \(\eta =\frac{1}{7}\), \(\alpha =0.01\). Then, \(R_0=0.9625<1\), \(R_*=0.7911<1\). The corresponding backward bifurcations diagram is shown in Fig. 6b. Taking parameters \(\epsilon =0.3\), \(R_0=0.9625<1\), \(R_*=1.3154>1\), the corresponding forward bifurcations diagram is shown in Fig. 6a. From Fig. 7, we can see that when \(\sigma\) and \(\epsilon\) are very small, that is, when the breakthrough infection and reinfection rates are very low, the disease will be extinct and will not become endemic. When the initial value is set to (0.5, 0.4, 0.08), Fig. 8 shows the change of infected people over time under different values of \(\sigma\) and \(\epsilon\). When the parameter value exceeds the red interface or the value of the red dotted line, the disease will break out on a large scale, otherwise the disease will be a small epidemic or extinction. When the disease is prevalent, improving the immunity of the vaccine and reducing breakthrough infection play a vital role in epidemic prevention and control. Strengthening the natural immunity of the recovered and reducing the reinfection rate can reduce the final epidemic scale of the disease, but the disease cannot be eradicated.
When \(R_0<1\), the phase diagram of system (2) under different conditions. (a) \(R_*\ge 1\). (b) \(R_*<1\) and \(R_{**}<R_0\). (c) \(R_*<1\) and \(R_{**}=R_0\). (d) \(R_*>1\) and \(R_{**}=R_0\). The red point represents the disease-free equilibrium \(P_0\), and the yellow, purple and green points represent the endemic equilibria \(P_{1}\), \(P_{2}\) and \(P_{3}\), respectively.
Bifurcations diagrams under different conditions. (a) \(R_*\ge 1\). (b) \(R_*< 1\). The dotted line indicates an unstable equilibrium, and the solid line indicates a stable equilibrium.
The change of equilibrium i under different \(\sigma\) and \(\epsilon\). The equilibrium in the red area is equal to 0.
The change of infected i(t) over time under different \(\sigma\) and \(\epsilon\).
When \(\sigma =0\), \(R_*=\frac{\theta (\gamma +\eta )}{\eta (\alpha +\theta )\epsilon }\). When \(\epsilon =0\), \(R_*=\frac{[(\alpha \sigma +\theta ) (\gamma +\eta )+\gamma \eta \sigma ] (\alpha \sigma +\theta )}{\gamma \eta \sigma (\alpha +\theta )}\). Take \(\theta =0.005\), then \(R_0=2.56\). From Fig. 9, we can see that when at least one of \(\sigma\) and \(\epsilon\) is not 0, system (2) may have a backward bifurcations. However, when \(\sigma\) and \(\epsilon\) are both 0, system (2) will have no backward bifurcations.
Relationship between \(\sigma\), \(\epsilon\) and \(R_*\) under different conditions. (a) \(\gamma =0.05\). (b) \(\gamma =0.0333\). (c) \(\gamma =0.0083\).
Optimal control
In this section, we will investigate a formalized time control problem. First, we propose a definition of disease eradication time for an infectious disease system.
Definition 1
The disease eradication time T of an infectious disease system is the moment when the number of infected first reaches a threshold k (\(0<k<\frac{1}{N}\)), where N is the total population.
In order to make Definition 1 meaningful, we choose the initial value of the number of infected in system (2) to be strictly greater than k. T is the time when the number of infected i(t) drops to k for the first time, that is, \(T=\text {inf}\left\{ t>0|i(t)=k\right\}\). From a biological perspective, setting the condition \(k<\frac{1}{N}\) to determine the disease eradication time means that the disease will become extinct when the number of infected is less than 1.
Based on system (7), the following three parameters are considered as control variables
-
(i)
\(u_1(t)\in U\) indicates the percentage of infection rate reduction due to reduced detection by wearing masks, maintaining social distance, et al.
-
(ii)
\(u_2(t)\in U\) indicates increasing vaccination rates among susceptible;
-
(iii)
\(u_3(t)\in U\) indicates that the protection rate of the vaccine is improved.
Where \(U=\left\{ u_1, u_2, u_3|0\le u_1, u_2, u_3 \le 1\right\}\) is control set. Our goal is to eradicate the disease in the shortest possible time. The corresponding state equation is
Let \(\vec {x}(t)=(s(t), v(t), i(t))\), \(\vec {u}(t)=(u_1(t), u_2(t), u_3(t))^{T}\),
then, system (7) can be rewritten as \(\vec {x}'=\vec {f}(t,\vec {x},\vec {u}), \vec {x}(0)=x^0\). For system (7), we set the initial conditions to \(i(0)>k\), \(s(0)>0\) or \(v(0)>0.\) Under this initial condition, the solution of system (7) is represented by \(\vec {x}(t)\), and its set is marked as X.
Eradicating the disease in the shortest possible time is of great significance. To achieve this goal, we study the following time optimal control problem: \(J(u)={\bf {min}}\int _{0}^{T} 1dt, i(T)=k\).
Existence of the optimal solution
Next, according to Theorem 4.1 and the corollary method in reference35, we prove that system (7) has sufficient conditions for optimal control. This is shown in Lemma 2 below.
Lemma 2
For the time optimal control problem, there exists an optimal control \(\vec {u}^*(t)\).
See Appendix for the proof process of Lemma 2.
Theorem 4
If \(\vec {u}^*(t)\) is the minimum value of the time-optimal control problem, then \(\vec {u}^*(t)\) is a Bang-Bang control.
See Appendix for the proof process of Theorem 3.
Simulation of optimal control
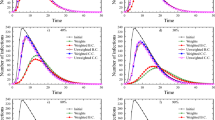
In this summary, we use the fmincon function to simulate Bang-Bang control. Here, the initial value and parameter values of system (2) in the simulation research are set as \(s(0)=0.4\), \(v(0)=0.5\), \(i(0)=0.1\), \(\beta =0.9\), \(\epsilon =0.4\), \(\sigma =0.1\), \(\theta =0.02\), \(\gamma =0.001\), \(\eta =0.1667\), \(\alpha =0.01\) \(k=1\times 10^{-3}\). Then, \(R_0=3.7792>1\). In real life, every control measure cannot reach 100%, so it is assumed that \(u_{i\text {min}}=0\), \(u_{i\text {max}}=0.98\), where \(i=1,2,3\). The control goal is to control the spread of the disease as much as possible in the shortest time. Next, the disease prevalence under various control measures is compared to find the best prevention and control measure. Figures 11, 12, 13, 14, 15, 16 and 17 show the dynamic changes of infected under no control measures and different control measures. Table3 shows the disease control time and cumulative infection proportion corresponding to each control measure.
Without any control measures, the changes of state variables s, v, and i over time.
Considering only the control of the infection rate, the changes of the control variable \(u_1\) and the state variables s, v, i over time.
Considering only the vaccination rate, the change of control variable \(u_2\) and state variables s, v, i over time.
Considering only the control of vaccine protection rate, the change of control variable \(u_3\) and state variables s, v, i over time.
Considering the control of infection rate and vaccination rate, the change of control variables \(u_1\), \(u_2\) and state variables s, v, i over time.
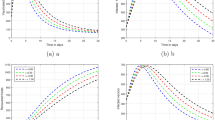
From Fig. 10, we can see that if no control measures are taken, the disease will become endemic with a high prevalence trend. When only the vaccination rate is increased without implementing other control strategies, as shown in Fig. 12, the disease will be controlled in a short period of time, but the vaccine protection rate will gradually weaken over time, and the negligence of protective measures will cause the recurrence of the disease. However, even if the vaccine protection rate is increased for a long time without strengthening the vaccination rate through publicity and education, as shown in Fig. 13, the disease will not be controlled, it will continue to exist and become endemic. As shown in Fig. 11, if social distance, mask wearing, and reduced outdoor activities are taken in the short term, the epidemic can be effectively controlled at time \(t=3.15\). From Figs. 14, 15 and 16, we can see that the disease can be quickly controlled when two control strategies are considered simultaneously. Among them, controlling the infection rate and increasing the vaccination rate take the shortest time to reach the control target, while controlling the infection rate and increasing the vaccine protection rate take the longest time to reach the control target, and it takes a period of time in the early stage to be controlled, which may be related to the relatively low vaccination rate.
Considering the control of infection rate and vaccine protection rate, the change of control variables \(u_1\), \(u_3\) and state variables s, v, i over time.
Considering the control of vaccination rate and protection rate, the change of control variables \(u_2\), \(u_3\) and state variables s, v, i over time.
Considering all control measures, the changes of control variables \(u_1\), \(u_2\), \(u_3\) and state variables s, v, i over time.
If the three control measures are considered at the same time, as shown in Fig. 17, it can be found that the control target is achieved at time \(t=2.835\), and the control time is often shorter. Table 3 shows that by implementing all control strategies, the disease containment target can be achieved in the shortest possible time while minimizing the cumulative infection scale. Therefore, in order to control the spread of the disease in the short term, when the epidemic comes, it must strengthen the propaganda of infectious disease prevention and control, and reduce contact with infected people through non-drug intervention measures such as wearing masks, maintain social distance, and reduce going out, especially for patients in the recovery period to prevent secondary outbreaks. With the continuous emergence and threat of new and recurrent infectious diseases, in order to prevent and control the spread of diseases in the long term, it is necessary to develop infectious disease vaccines with high immunity, so as to effectively prevent and control the continuous occurrence of diseases. Although the long-term implementation of prevention and control measures can curb the continuous spread of diseases, it will also cause waste of resources and even seriously interfere with the normal operation of the economy and society. However, Bang-Bang control provides an effective strategy, which can achieve the effect of disease prevention and control by implementing prevention and control strategies in a very short time.
Conclusion
In order to investigate the impact of breakthrough infection and reinfection on the continuous outbreak of infectious diseases, a disease transmission model with breakthrough infection and reinfection is established. First, a dynamic analysis of the model is conducted. The basic reproduction number \(R_0\) is obtained by using the next generation matrix method. The local stability of disease-free equilibrium when \(R_0<1\) is proved. By analyzing the stability of disease-free equilibrium when \(R_0=1\) and the existence of multiple endemic equilibria, we find that when \(R_*<1\), system (1) exhibits a backward bifurcation, which means that \(R_0\) is no longer the threshold for judging whether the disease will persist or not. Secondly, the local stability of the endemic equilibrium \(P_2\) and the instability of \(P_3\) under certain conditions are proved. Numerical simulations show that backward bifurcation occurs in system (1) when at least one of the parameters \(\sigma\) (breakthrough infection) or \(\epsilon\) (reinfection) is non-zero. Obviously, breakthrough infection and reinfection play a vital role in the multiple outbreaks of the disease. When \(\sigma\) and \(\epsilon\) approach 0, the disease will be effectively controlled and will not evolve into an endemic disease. On the contrary, the disease will spread extensively and eventually become an endemic disease when at least one of \(\sigma\) or \(\epsilon\) approaches to 1. In the context of pandemic control, real-world conditions such as resource constraints and socioeconomic factors often limit the long-term continuous implementation of intervention measures. Therefore, this study attempts to explore feasible pathways for dynamically switching different prevention and control methods within a limited period, based on the Bang-Bang control strategy under a discrete control framework. Finally, Bang-Bang control is used to conduct an optimization control, and the existence of an optimal solution is established. Simulations are conducted for single, dual and triple control measures. The research shows that the time to reach the control target is the shortest when the three control measures are considered comprehensively. If only the vaccination rate or the vaccine protection level is increased separately, it will not control the spread of the disease effectively, even will cause a secondary outbreak of the disease or an endemic state.
The model established in this paper is based on simple assumptions of homogeneous population mixing and spatiotemporal invariance, without considering realistic factors such as population heterogeneity and spatial distribution. Additionally, we construct Bang-Bang control is a piecewise continuous strategy under unconstrained conditions, without accounting for practical considerations such as intervention costs, resource limitations, and random disturbances.In the future, we will conduct research from aspects such as age and spatial heterogeneity, and perform simulation studies using actual case data. Additionally, we will construct an optimal control framework with resource constraints, cost-effectiveness, and random disturbances to quantify the return on investment of prevention and control measures, providing more specific and reliable theoretical support for infectious disease prevention and control decisions.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Centers for Disease Control and Prevention. COVID-19 Vaccination. https://www.cdc.gov/coronavirus/2019-ncov/vaccines/effectiveness/why-measure-effectiveness/breakthrough-cases.html (2022).
Li, Y., Qin, S., Dong, L. et al. Long-term effects of Omicron BA. 2 breakthrough infection on immunity-metabolism balance: A 6-month prospective study. Nat. Commun. 15(1), 2444 (2024).
Osterholm, M. T. et al. Efficacy and effectiveness of influenza vaccines: A systematic review and meta-analysis. Lancet Infect. Dis. 12(1), 36–44 (2012).
Gupta, R. K. & Topol, E. J. COVID-19 vaccine breakthrough infections. Science 374(6575), 1561–1562 (2021).
Centers for Disease Control and Prevention. Seasonal Influenza (Flu). https://www.cdc.gov/flu/professionals/antivirals/antiviral-use-influenza.htm (2021).
Centers for Disease Control and Prevention. Chickenpox (Varicella). https://web.archive.org/web/20170225050958/https://www.cdc.gov/chickenpox/hcp/clinical-overview.html (2017).
Bergwerk, M. et al. COVID-19 breakthrough infections in vaccinated health care workers. N. Engl. J. Med. 385(16), 1474–1484 (2021).
Alcendor, D. J. et al. Breakthrough COVID-19 infections in the US: implications for prolonging the pandemic. Vaccines 10(5), 755 (2022).
Abu-Raddad, L. J. et al. Relative infectiousness of SARS-CoV-2 vaccine breakthrough infections, reinfections, and primary infections. Nat. Commun. 13(1), 532 (2022).
Chemaitelly, H. et al. Addressing bias in the definition of SARS-CoV-2 reinfection: Implications for underestimation. Front. Med. 11, 1363045 (2024).
Pulliam, J. R. C. et al. Increased risk of SARS-CoV-2 reinfection associated with emergence of Omicron in South Africa. Science 376(6593), 4947 (2022).
Fonville, J. M. et al. Antibody landscapes after influenza virus infection or vaccination. Science 346(6212), 996–1000 (2014).
Van der Kuyl, A. C. & Cornelissen, M. Identifying HIV-1 dual infections. Retrovirology 4, 1–12 (2007).
Ghosh, M., Olaniyi, S. & Obabiyi, O. S. Mathematical analysis of reinfection and relapse in malaria dynamics. Appl. Math. Comput. 373, 125044 (2020).
Anggriani, N. et al. The effect of reinfection with the same serotype on dengue transmission dynamics. Appl. Math. Comput. 349, 62–80 (2019).
Wang, J. et al. COVID-19 reinfection: A rapid systematic review of case reports and case series. J. Invest. Med. 69(6), 1253–1255 (2021).
Glynn, J. R. et al. High rates of recurrence in HIV-infected and HIV-uninfected patients with tuberculosis. J. Infect. Dis. 201(5), 704–711 (2010).
Costenaro, P. et al. SARS-CoV-2 infection in people living with HIV: A systematic review. Rev. Med. Virol. 31(1), 1–12 (2021).
Mary, K. People with HIV at higher risk of COVID reinfection: CDC. https://abcnews.go.com/Health/people-hiv-higher-risk-covid-reinfection-cdc/story?id=104035803 (2023).
Centers for Disease Control and Prevention. What is COVID-19 Reinfection? https://www.cdc.gov/coronavirus/2019-ncov/your-health/reinfection.html (2019).
Perkins, T. A. et al. An agent-based model of dengue virus transmission shows how uncertainty about breakthrough infections influences vaccination impact projections. Plos Comput. Biol. 15(3), e1006710 (2019).
Azimaqin, N. et al. Vaccine failure, seasonality and demographic changes associate with mumps outbreaks in Jiangsu Province, China: Age-structured mathematical modelling study. J. Theor. Biol. 544, 111125 (2022).
Xu, C. et al. A mathematical model to study the potential hepatitis B virus infections and effects of vaccination strategies in China. Vaccines 11(10), 1530 (2023).
Elbasha, E. H., Podder, C. N. & Gumel, A. B. Analyzing the dynamics of an SIRS vaccination model with waning natural and vaccine-induced immunity. Nonlinear Anal. Real World Appl. 12(5), 2692–2705 (2011).
Yuliana, R., Alfiniyah, C. & Windarto, W. Stability analysis of SIVS epidemic model with vaccine ineffectiveness. In AIP Conference Proceedings. Vol. 2329(1) (2021).
Jing, S. et al. Age-structured modeling of COVID-19 dynamics: The role of treatment and vaccination in controlling the pandemic. J. Math. Biol. 90(1), 1–49 (2025).
Montalbán, A., Corder, R. M. & Gomes, M. G. M. Herd immunity under individual variation and reinfection. J. Math. Biol. 85(1), 2 (2022).
Rehman, A., Singh, R. & Singh, J. Mathematical analysis of multi-compartmental malaria transmission model with reinfection. Chaos Solitons Fract. 163, 112527 (2022).
Anggriani, N. et al. The effect of reinfection with the same serotype on dengue transmission dynamics. Appl. Math. Comput. 349, 62–80 (2019).
Gomes, M. G. M., White, L. J. & Medley, G. F. Infection, reinfection, and vaccination under suboptimal immune protection: Epidemiological perspectives. J. Theor. Biol. 228(4), 539–549 (2004).
Agusto, F. B. Mathematical model of Ebola transmission dynamics with relapse and reinfection. Math. Biosci. 283, 48–59 (2017).
Rodrigues, P., Margheri, A., Rebelo, C. & Gomes, M. G. Heterogeneity in susceptibility to infection can explain high reinfection rates. J. Theor. Biol. 259(2), 280–290 (2009).
Megala, T., Nandha Gopal, T., Siva Pradeep, M. et al. Dynamics of re-infection in a hepatitis B virus epidemic model with constant vaccination and preventive measures. J. Appl. Math. Comput. 1–27 (2025).
Van den Driessche, P. Reproduction numbers of infectious disease models. Infect. Dis. Model. 2(3), 288–303 (2017).
Fleming, W. H. & Rishel, R. W. Deterministic and Stochastic Optimal Control (Springer, 1975).
Pontryagin, L. S. Mathematical Theory of Optimal Processes. (CRC Press, 1987).
Acknowledgements
We would like to thank the reviewers for their valuable comments and suggestions which helped us to improve the presentation of this paper significantly.
Funding
This work was supported by the National Natural Science Foundation of China (No. 12471462, No. 12101373, No. 12171291), Fundamental Research Program of Shanxi Province (No. 202403021211154).
Author information
Authors and Affiliations
Contributions
Y. C.: Data curation, Formal analysis, Validation,Writing – original draft, Writing – review & editing. W. J.: Data curation, Formal analysis, Visualization, Writing – original draft, Writing – review & editing. J. Z.: Conceptualization, Investigation, Methodology, Supervision, Writing – review & editing P. Q.: Conceptualization, Data curation, Software, Supervision, Writing – review & editing
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
The proof of Lemma 1
Proof
Summing the four equations of system (1) up yields \(\frac{dN}{dt}=0\). Thus the total population N(t) is a constant. For convenience, let \(N(t)=K\). Next, we prove that the solution of system (1) is non-negative.
Solving to the third equation of system (1) yields
Plugging it into the fourth equation of system (1), we can get
Besides, sloving the first equation of system (1), we obtion
Let \(t_1>0\) be the first time that \(\text {min}\left\{ S(t_1),V(t_1)\right\} =0\). Assuming that \(S(t_1)=0\), then S(t) and \(V(t)>0\), for\(\forall t\in [0,t_1)\). Furthermore we can get
this contradicts the assumption \(S(t_1)=0\). Similarly, let us assume that \(V(t_1)=0\), then according to the second equation of system (1), we can get
which contradicts the assumption of \(V(t_1)=0\). This mean that all solution of system (1) are non-negative and \(\Gamma\) is the positively invariant set of system (1). This completes the proof. \(\square\)
The proof of Theorem 1
Proof
The Jacobian matrix of system (2) at the disease-free equilibrium \(P_0\) is
The characteristic polynomial corresponding to matrix \(K(P_0)\) is
where
The roots of the characteristic polynomial (8) are \(\lambda _1=-\gamma\), \(\lambda _2=-\alpha -\theta , \lambda _3=\frac{\beta (\sigma \alpha +\theta )-\eta (\alpha +\theta )}{\alpha +\theta }\). Obviously, when \(R_0<1\), \(\lambda _3<0\). Therefore, the disease-free equilibrium \(P_0\) of system (2) is locally asymptotically stable, if \(R_0<1\).
When \(R_0=1\), \(\lambda _3=0\). At this time, the central manifold theorem is used to prove the stability of the disease-free equilibrium \(P_0\) . The following affine transformation is defined
Furthermore,
the linear part here is the Jordan canonical form of the matrix K, and
Let \(R_*=\frac{[(\alpha \sigma +\theta ) (\gamma +\eta )+\gamma \eta \sigma ](\alpha \sigma +\theta )}{\eta (\alpha +\theta ) [(\alpha \sigma +\theta )\epsilon +\gamma \sigma ]}\). Then, if \(R_0\ne R_*\),there exists a one-dimensional central manifold
Therefore, system (2) is simplified to
It is to see that disease-free equilibrium \(P_0\) is a saddle node. When \(R_0=R_*\), according to the center manifold theorem, the center manifold of system (2) at the far point can be expressed as
Since \(\sigma , \epsilon \in [0,1]\), the disease-free equilibrium \(P_0\) is a stable node, when \(R_0=1\) and \(R_0=R_*\). \(\square\)
The proof of Theorem 2
Proof
The Jacobian matrix of system (2) at the equilibrium \(P_2\) is
The characteristic equation of \(K(P_2)\) is
where
Obviously, \(d_2>0\), \(f(i_2)=0\). According to \(R_0<1\) and \(R_*<R_0\), we have that \(g(i_2)<0\), \(d_0<0\). Let the three roots of Eq. (14) be \(\Lambda _1\), \(\Lambda _2\) and \(\Lambda _3\), respectively. According to Vieta’s theorem, it has
Therefore, according to expression (15), the roots of Eq. (14) include the following three cases: three positive roots, one positive root and a pair of conjugate complex roots, one positive root and two negative roots. According to expression (16), Eq. (14) has a pair of conjugate complex roots or negative roots with negative real parts. In summary, there are two cases of the roots of Eq. (14): one positive root and a pair of conjugate complex roots, one positive root and two negative roots. Therefore, the endemic equilibrium \(P_2\) is unstable. \(\square\)
The proof of Theorem 3
Proof
The Jacobian matrix of system (2) at the endemic equilibrium \(P_3\) is
The characteristic equation of \(K(P_3)\) is
where
Obviously, \(e_2>0\), \(e_1>0\), \(F(i_3)=0\). According to \(R_{**}<R_{0}\), \(R_*<R_0\) and \(R_*<1\), we obtain that \(G(i_3)>0\), and then \(e_0>0\). Therefore,
According to the Hurwitz criterion, all the roots of Eq. (17) have negative real parts. Therefore, the endemic equilibrium \(P_3\) is locally asymptotically stable. \(\square\)
The proof of Lemma 2
Proof
Let
Then, system (7) can be rewritten as \(\vec {x}'(t)=\vec {f_*}(t,\vec {x})+\vec {g_*}(t,\vec {x},\vec {u})\vec {u}\). We prove that the following four conditions are satisfied:
-
(i)
\(\vec {f_*}(t,\vec {x})+\vec {g_*}(t,\vec {x},\vec {u})u\) is first-order continuously differentiable, and there exists a constant C such that
$$\begin{aligned} |\vec {f_*}(t,0,0)|\le C, |\vec {f_*}(\vec {x})+\vec {g_*}(\vec {x},\vec {u})\vec {u}|\le C(1+|\vec {u}|), |\vec {g_*}(\vec {x},\vec {u})|\le C. \end{aligned}$$ -
(ii)
The solution set of system (7) corresponding to the control parameters in the control set U is non-empty.
-
(iii)
The dominating set U is a closed convex compact set.
-
(iv)
The integrand in the objective function J is convex in U.
Firstly, it is easy to see that \(\vec {f_*}(t,\vec {x})+\vec {g_*}(t,\vec {x},\vec {u})u\) is first-order continuously differentiable, and \(|\vec {f_*}(t,0,0)|=0\). Since s, v, i are non-negative and bounded, there exists a constant C such that \(|\vec {f_*}(t,0,0)|\le C, |\vec {f_*}(\vec {x})+\vec {g_*}(\vec {x},\vec {u})\vec {u}|\le C(1+|\vec {u}|), |\vec {g_*}(\vec {x},\vec {u})|\le C.\) This shows that condition (i) is true, It can be seen that the system (7) has a unique solution, which means that condition (ii) is established.
The control set U is a closed convex compact set and the integrand of the objective function is a constant function. So conditions (iii) and (iv) are valid. In summary, the time optimal control problem has an optimal solution. \(\square\)
The proof of Theorem 4
Proof
In order to find the minimum disease eradication time and the time-dependent control variable \(\vec {u}^*(t)\), it is equivalent to the problem of minimizing the Hamiltonian system. Let the Hamiltonian system be
where \(\Lambda _1, \Lambda _2, \Lambda _3\) are the adjoint variables associated with the state variables s, v, i. Using the Pontryagin maximum principle36, we can see that the relationship between adjoint control and optimal control is as follows
and the transversality condition is \(\Lambda _1(T)=\Lambda _2(T)=\Lambda _3(T)=0\).
Therefore, the adjoint system is
The corresponding switching functions and their time derivatives are
where
Next, we prove that the switching function \(\Phi\) disappears only at isolated points. Assuming that \(\Phi (\vec {x},\Lambda _1,\Lambda _2,\Lambda _3,t)\) disappears in an open set L, then in the neighborhood L, \(\Phi =\frac{d\Phi }{dt}=0\), so there is \(\Lambda _1=\Lambda _2=\Lambda _3=0\). Then, the Hamiltonian system calculated along the optimal solution can be \(H(\vec {x},\Lambda _1,\Lambda _2,\Lambda _3,t)=1\ne 0\), which creates a contradiction. This implies that \(\vec {u}^*(t)\) is a Bang-Bang control, that is a piecewise function. \(\square\)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, Y., Jing, W., Zhang, J. et al. Bang-bang control optimization in infectious disease model with incorporating breakthrough and reinfection. Sci Rep 16, 15272 (2026). https://doi.org/10.1038/s41598-026-44921-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-44921-7