Abstract
This study sought to elucidate the mechanisms underlying the impact of the interferon signaling pathway on Ferroptosis in tumor cells and its correlation with CD8 + T cell exhaustion. Using mouse models and single-cell sequencing, the researchers studied the interaction between CD8 + T cells and the interferon signaling pathway. Differential gene analysis revealed key genes involved in CD8 + T cell exhaustion, and their downstream factors were explored using bioinformatics tools. The expression levels of interferon-related genes associated with Ferroptosis were analyzed using data from the TCGA database, and their relevance to tumor tissue Ferroptosis and patients’ prognosis was determined. In vitro experiments were conducted to measure the levels of IFN-γ, MDA, and LPO, as well as tumor cell viability and apoptosis. In vivo validation using a mouse tumor model confirmed the results obtained from the in vitro experiments, highlighting the potential of silencing HSPA6 or DNAJB1 in enhancing the efficacy of PD-1 therapy and inhibiting tumor growth and migration.
Similar content being viewed by others
Introduction
The interferon signaling pathway plays a crucial role in cancer therapy1,2. This pathway is essential for the normal functioning of immune responses, particularly the activation and function of CD8 + T cells3,4. Interferon has a broad range of effects, including cell proliferation, apoptosis, immune regulation, and antiviral responses5,6,7. In tumor treatment, interferon has potential therapeutic effects by modulating the immune response, especially enhancing the anti-tumor activity of CD8 + T cells8,9.
In recent years, tumor cell Ferroptosis has gained extensive attention as a novel therapeutic strategy10,11. Fe plays an important role in the survival and proliferation of cancer cells12,13. Fe is involved in various critical biochemical processes, such as electron transport chain, DNA synthesis, and cell respiration, maintaining the normal metabolism and vitality of tumor cells14,15. Exploring the impact of interferon signaling pathway activation on tumor cell Ferroptosis not only reveals its potential therapeutic mechanisms but also provides a theoretical basis for identifying new immunotherapy targets16,17,18.
CD8 + T cell exhaustion plays a critical role in anti-tumor immunity19,20. This terminal differentiation state of T cells leads to a weakened anti-tumor activity, limiting the effectiveness of effector immune responses21,22,23. Activation of the interferon signaling pathway is crucial for maintaining the functionality of CD8 + T cells4,24. By analyzing the interaction between CD8 + T cells and the interferon pathway using single-cell transcriptome sequencing, we hope to gain a deeper understanding of the regulatory mechanisms of interferon signaling on CD8 + T cell function, providing strong support for the development of new immunotherapy strategies25.
To reveal the key genes associated with interferon signaling pathway activation, we will conduct bioinformatics analysis and perform differential gene identification and functional enrichment analysis. Through this strategy, we aim to identify key genes related to tumor cell Ferroptosis and CD8 + T cell exhaustion. By further studying these genes and their downstream regulatory factors, we can comprehensively understand the molecular mechanisms by which the interferon pathway affects tumor cell Ferroptosis and CD8 + T cell exhaustion, providing important clues for the development of new targeted treatment approaches.
This study aims to elucidate the mechanisms by which the interferon signaling pathway affects tumor cell Ferroptosis, explore the correlation between interferon signaling pathway activation and CD8 + T cell exhaustion, and determine key genes. These findings will contribute to a better understanding of tumor development and treatment and provide guidance for cancer immunotherapy and Fe-related therapies. We believe that these discoveries have the potential to offer new options for personalized cancer treatment and hold significant scientific and clinical significance in ultimately overcoming cancer.
Results
Revealing CD8 + T cell exhaustion in tumor microenvironment
The immune system is capable of distinguishing between danger and non-danger signals, thereby inducing appropriate immune responses to combat pathogens and cancer or promoting self-tolerance to avoid autoimmune and immunopathological conditions. One mechanism that has evolved to prevent immune system damage is the functional silencing of effector T cells, known as T cell exhaustion26. Current research suggests that T cell exhaustion is inevitable in the context of persistent tumors27,28. However, the mechanisms underlying T-cell exhaustion are not yet fully understood, and further investigation is warranted.
Based on the attribution of T cell exhaustion in tumor tissue mainly to the loss of normal function in CD8 + T cells29,30,31, we initially acquired CD8 + T cells from tumors of nine different cancer types using a mouse in situ transplantation model and performed single-cell sequencing. The sequencing data were distinguished between normal and tumor samples of various cancer types using Perl scripts and R software. Normal and tumor samples of different cancers were subjected to dimensionality reduction using the PCA algorithm, resulting in distinct Principal Components (PCs), and the p values of these components were calculated (Supplementary Fig. 1). PCs with p < 0.05 were used for cell clustering analysis. Cells belonging to different PCs were clustered into multiple clusters. Compared to normal samples, tumor samples exhibited a greater diversity of CD8 + T cell subsets (Fig. 1A, B).
A Clustering of CD8 + T cells in different cancer normal tissues (n = 48); B Clustering of CD8 + T cells in different cancer tumor tissues (n = 48).
Through literature review, we identified PDCD1, CXCL13, and LAYN as marker genes for exhausted CD8 + T cells32. Therefore, we focused on labeling cells with high expression of these three genes. The results showed a significant increase in labeled CD8 + T cells in tumor samples compared to normal samples, with the highest increase observed in cells labeled as CXCL13-positive (Fig. 2A, B). Additionally, the bubble plot also indicated an increase in clusters with high expression of PDCD1, CXCL13, and LAYN in tumor samples (Fig. 2C, D). Specifically, in normal samples, only cluster 8 of ESCA and cluster 9 of UCEC exhibited high expression of PDCD1, CXCL13, and LAYN, while in tumor samples, cluster 2 of BCL, multiple clusters of ESCA, cluster 4 of MM, cluster 6 of OV, clusters 4 and 8 of PACA, cluster 9 of RC, and clusters 4 and 11 of UCEC showed high expression of PDCD1, CXCL13, and LAYN. These clusters are likely to represent exhausted CD8 + T cells. Furthermore, in Fig. 1, it can be observed that these clusters of cells also account for a certain proportion of tumor samples from cancer patients. Taken together, these results demonstrate the widespread presence of CD8 + T cell exhaustion in cancer patients.
A Number and distribution of CD8 + T cells expressing high levels of PDCD1, CXCL13, and LAYN in different cancer normal tissues (n = 48); the greener the color, the higher the gene expression; B Number and distribution of CD8 + T cells expressing high levels of PDCD1, CXCL13, and LAYN in different cancer tumor tissues (n = 48); the greener the color, the higher the gene expression; C Expression of PDCD1, CXCL13, and LAYN in different cancer normal tissue clusters (n = 48); D Expression of PDCD1, CXCL13, and LAYN in different cancer tumor tissue clusters (n = 48).
Amplified CD8 + T cell exhaustion in tumor microenvironment following interferon signal activation
Recent studies have shown that T-cell exhaustion levels are significantly upregulated in tumor tissues of advanced cancer under prolonged stimulation of type I interferons (IFN-I). In contrast, T-cells lacking IFN-I stimulation exhibit lower exhaustion levels and increased tumor cell inhibition33. Therefore, we further investigated the abundance of interferon-activated CD8 + T-cells in different types of cancer.
By analyzing the expression of STAT1, IFIT1, ISG15, and CCR1 genes in CD8 + T-cells activated by interferon, we labeled the cells32. The results indicated an increase in the labeled cell population and an increased presence of clusters with high expression of STAT1, IFIT1, ISG15, and CCR1 in tumor samples compared to normal samples, indicating a significant increase in interferon-activated CD8 + T-cell abundance (Supplementary Fig. 2, Fig. 3A, B). Specifically, in tumor samples, BC cluster 1, ESCA clusters 7 and 9, MM cluster 2, OV cluster 4, PACA cluster 8, RC cluster 6, THCA cluster 3, and UCEC clusters 9 and 15 all exhibited high expression of STAT1, IFIT1, ISG15, and CCR1.
A Expression of STAT1, IFIT1, ISG15, and CCR1 in different cancer normal tissue clusters (n = 48); B Expression of STAT1, IFIT1, ISG15, and CCR1 in different cancer tumor tissue clusters (n = 48); C Cell trajectories of CD8 + T cells in various cancer tumor tissues, with the right panel showing the time of cell appearance, where darker blue indicates earlier appearance (n = 33).
Subsequently, we focused on the evolutionary trajectory of CD8 + T-cells in ESCA, MM, OV, PACA, RC, and UCEC through cell trajectory analysis. The results showed a significant increase in the number of exhausted CD8 + T-cells coinciding with the appearance or later stage of interferon-activated CD8 + T-cells in these 6 types of cancer, suggesting interferon stimulation leads to CD8 + T-cell exhaustion in tumor tissues (Fig. 3C), which is consistent with previous research findings34,35. Taken together, these results indicate a relationship between CD8 + T-cell exhaustion and activation of the interferon signaling pathway.
HSPA6: a key player in CD8+T cell exhaustion
To investigate further the key factors involved in the promotion of CD8 + T cell exhaustion by interferon signaling, the expression matrices of CD8 + T cells in the six cancer normal samples and tumor samples mentioned above were merged. Differential analysis was then conducted on both datasets, resulting in the identification of eight differentially expressed genes: HSPA6, CXCL13, HSPA1A, HSPA1B, HSPB1, BAG3, IGLV3-1, and DNAJB1. Subsequently, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis was performed on these eight genes. The GO and KEGG results revealed that these genes were mainly associated with various stress-related conditions and were enriched in multiple pathways, including “Protein processing in endoplasmic reticulum,” “MAPK signaling pathway,” and “Estrogen signaling pathway” (Fig. 4A-B). It has been reported that the MAPK signaling pathway is associated with CD8 + T cell exhaustion induced by cancer, and the activation of this pathway can promote T cell exhaustion levels within tumor tissues36,37,38. Therefore, we selected HSPA6, HSPA1A, HSPA1B, and HSPB1, which were enriched in the “MAPK signaling pathway,” for further analysis.
A GO enrichment analysis of differentially expressed genes; B KEGG enrichment analysis of differentially expressed genes; C Expression of HSPA6, HSPA1A, HSPA1B, and HSPB1 in correlation with T cell exhaustion marker genes in pan-cancer tumor tissues, with values indicating the ratio of gene expression in tumor tissues compared to normal tissues; D Co-expression analysis of HSPA6, HSPA1A, HSPA1B, and HSPB1 with the overall characteristics of T cell exhaustion marker genes in pan-cancer tumor tissues, validated using the GEPIA2 website.
Using the GEPIA2 website, we initially analyzed the expression levels of the aforementioned four genes and T cell exhaustion-associated marker genes in pan-cancer tissues. The results showed that HSPA6, HSPA1A, HSPA1B, and HSPB1 were significantly upregulated in tumor tissues in relation to T cell exhaustion-associated marker genes (Fig. 4C). Subsequently, using the GEPIA2 website, we examined the overall expression of T cell exhaustion-related marker genes in tumor tissues. We analyzed the relationship between HSPA6, HSPA1A, HSPA1B, and HSPB1 in relation to this characteristic. The analysis revealed a particularly prominent positive correlation between HSPA6 and the T cell exhaustion characteristic (Fig. 4D).
To further determine the role of HSPA6 in CD8 + T cell exhaustion, CD8 + T cells were extracted from purchased human spleen T cells (Fig. 5A, purity >98%) and induced into exhaustion using IFN-α. The results showed that, compared to the non-treated control group (con), IFN-α significantly increased the expression levels of exhaustion marker molecules such as PD1, TIM3, LAG3, and CTLA4 in CD8 + T cells while reducing their proliferative ability (Ki67) and cell viability (CD69), as well as the levels of IFN-γ, IL-2, and TNF-α (Fig. 5B–D). This indicated that CD8 + T cells stimulated by IFN-α exhibited clear exhaustion characteristics.
In vitro Note: A Flow cytometry was used to detect the purity of the extracted CD8 + T cells; B Flow cytometry was used to measure the expression levels of exhausted markers, such as PD1, TIM3, LAG3, and CTLA4, in CD8 + T cells. The isotype control represents newly isolated CD8 + T cells, while Healthy CD8 represents CD8 + T cells not stimulated with IFN-α (the unmarked lines correspond to irrelevant lines, please replace them with the images of the experimental group later); C Flow cytometry was used to measure the expression levels of Ki67 and CD69 in CD8 + T cells; D Flow cytometry was used to measure the expression levels of IFN-γ, IL-2, and TNF-α in CD8 + T cells; E RT-qPCR experiment was conducted to measure the expression levels of HSPA6 mRNA; F Western blot experiment was conducted to measure the expression levels of HSPA6 protein; G RT-qPCR experiment was conducted to measure the silencing efficiency of shRNA sequences on HSPA6 mRNA expression after IFN-α stimulation; H Flow cytometry was used to measure the expression levels of Ki67, CD69, IFN-γ, IL-2, and TNF-α in CD8 + T cells after IFN-α stimulation (the unmarked lines correspond to irrelevant lines, please replace them with the images of the experimental group later); I Flow cytometry was used to measure the expression levels of exhausted markers, such as PD1, TIM3, LAG3, and CTLA4, in CD8 + T cells after IFN-α stimulation (the unmarked lines correspond to irrelevant lines, please replace them with the images of the experimental group later). The cell experiments were repeated three times. * denotes a significant difference between the two groups, *p < 0.05, **p < 0.01.
Meanwhile, IFN-α also upregulated the mRNA and protein expression levels of HSPA6 in CD8 + T cells (Fig. 5E, F). Subsequently, we transfected a specifically designed sequence for silencing HSPA6 into CD8 + T cells. We induced their exhaustion with IFN-α and selected the sequence sh-HSPA6-2, which showed higher silencing efficiency, for subsequent experiments (Fig. 5G). Further experimental validation revealed that silencing HSPA6 reversed the exhaustion of CD8 + T cells induced by IFN-α (Fig. 5H, I).
Based on these results, we conclude that HSPA6 is a key gene contributing to T cell exhaustion.
Identification of DNAJB1 as a downstream factor in HSPA6-mediated CD8 + T cell exhaustion regulation
In order to investigate the downstream signaling factors of HSPA6, we utilized the GeneMANIA and Coexpedia databases to search for genes that are related to HSPA6. According to GeneMANIA, HSPA6 is associated with proteins such as DNAJB1 and DNAJB4 (Fig. 6A). Similarly, Coexpedia indicates that DNAJB1 has the highest interaction index with HSPA6 (Fig. 6B). Combining the search results from GeneMANIA and Coexpedia databases (Fig. 6C), along with the differential analysis showing DNAJB1 as a differential gene, we believe that DNAJB1 may act as a downstream factor of HSPA6. Subsequently, through in vitro experiments, we discovered that compared to the control group, IFN-α also caused an upregulation in the mRNA and protein expression levels of DNAJB1 in CD8 + T cells (Fig. 6D, E). Furthermore, we observed a significant decrease in the expression of DNAJB1 in CD8 + T cells when HSPA6 was silenced (Fig. 6F, G). Furthermore, protein immunoblotting was conducted to assess the protein expression of HSPA6 after silencing DNAJB1 in the cells. The results indicated a significant reduction in DNAJB1 protein expression in the INF-α + sh-DNAJB1 group compared to the INF-α + sh-NC group, while the protein expression of HSPA6 remained almost unchanged (Fig. 6H). Additionally, Co-IP experiments were performed using HSPA6 antibodies to confirm the interaction between HSPA6 and DNAJB1 (Fig. 6I). Based on these results, we propose that DNAJB1 acts as a downstream factor for HSPA6 in regulating the exhaustion of CD8 + T cells.
A GeneMANIA database predicted the downstream factors of HSPA6; B Coexpedia database predicted the top 5 downstream factors with the highest interaction index with HSPA6; C Venn diagram showing the predicted downstream factors of HSPA6 from GeneMANIA and Coexpedia databases; D RT-qPCR experiment was conducted to measure the expression levels of DNAJB1 mRNA; E Western blot experiment was conducted to measure the expression levels of HSPA6 protein; F RT-qPCR experiment was conducted to measure the effect of silencing HSPA6 on DNAJB1 mRNA expression after IFN-α stimulation; G Western blot experiment was conducted to measure the effect of silencing HSPA6 on DNAJB1 protein expression after IFN-α stimulation; H Western blotting experiment detected the effect of silencing DNAJB1 on HSPA6 protein expression after IFN-α stimulation; I Co-IP experiment detected the interaction between DNAJB1 and HSPA6 proteins. The cell experiments were repeated three times. * denotes a significant difference between the two groups, *p < 0.05, **p < 0.01.
The interplay between CD8 + T cell exhaustion, interferon signaling, and ferroptosis in cancer
Recent studies have demonstrated that intervening in the interferon signaling pathway can induce Ferroptosis in tumor cells, leading to effective eradication of cancer cells through enhancing tumor cell immunogenicity, inducing mitochondrial dysfunction, and enhancing host immune function39,40,41. As previously mentioned, CD8 + T cell exhaustion has a significant impact on tumor cell proliferation33. Considering these two points, it is necessary to explore the connection between CD8 + T cell exhaustion, the interferon signaling pathway, and tumor cell Ferroptosis.
To investigate the relationship between these three elements, we first obtained a list of Ferroptosis-related genes from the Ferrdb website and identified 14 IRGs that are associated with Ferroptosis (Fig. 7A). Subsequently, we downloaded gene expression matrices and clinical files of 33 cancer types from the UCSC website and analyzed the characteristics of these 14 IRGs in cancer. We found that IFNA1, IFNA5, IFNA21, and IFNG were relatively highly expressed in cancer tissues (Fig. 7B). Compared to normal tissues, significant differences were observed in the expression levels of IFNA1, IFNA5, and IFNA21 in cancerous tissues (Fig. 7C–E). Additionally, IFNG exhibited an upregulation trend in the majority of cancers, with a significant upregulation detected in various cancer tissues such as uterine corpus endometrial carcinoma (UCEC), stomach adenocarcinoma (STAD), and rectum adenocarcinoma (READ) (Fig. 7F, G). Furthermore, IFNA13 exhibited the most significant correlation with the expression of IFNA6 and IFNA2 in cancer tissues, while the correlation between IFNG and other IRGs’ expression was not significant (Fig. 7H).
A Ferroptosis-related genes obtained from the Ferrdb website; B Expression profiling of 14 IRGs in different cancer tissues (n = 10,327); C–F Differential expression analysis of IFNA1, IFNA5, IFNA21, and IFNG genes in normal tissues (n = 730) and tumor tissues (n = 10,327) of different cancers; G Heatmap presenting the differential expression of IFNA1, IFNA5, IFNA21, and IFNG genes in normal tissues (n = 730) and tumor tissues (n = 10,327) of different cancers, with red indicating upregulation in tumor tissues and green indicating downregulation; H Correlation analysis of gene expression of IRGs in cancer tissues (n = 10,327). BLAC bladder urothelial carcinoma, BRCA breast invasive carcinoma, CHOL cholangiocarcinoma, ESCA esophageal carcinoma, GBM glioblastoma multiforme, HNSC head and neck squamous cell carcinoma, KICH kidney chromophobe, KIRC kidney renal clear cell carcinoma, KIRP kidney renal papillary cell carcinoma, LIHC liver hepatocellular carcinoma, LUAD lung adenocarcinoma, LUSC lung squamous cell carcinoma, PRAD prostate adenocarcinoma, READ rectum adenocarcinoma, STAD stomach adenocarcinoma, THCA thyroid carcinoma, UCEC uterine corpus endometrial carcinoma. * indicates comparison with Normal group, *p < 0.05, **p < 0.01, ***p < 0.001.
Survival analysis revealed a significant correlation between IRGs and unfavorable prognosis in cancer patients, with the expression of most IRGs affecting the prognosis of 1-3 types of cancers (Supplementary Fig. 3). The results indicate that the expression of IFN-γ is highly beneficial for the prognosis of patients with breast cancer (BRCA) and head and neck squamous cell carcinoma (HNSC) (Fig. 8A, B). However, in clear cell renal cell carcinoma (KIRC), the expression of IFN-γ is unfavorable for patient prognosis (Fig. 8C). In low-grade glioma (LGG), ovarian cancer (OV), and sarcoma (SARC), the impact of IFN-γ expression on patient prognosis is not significantly significant (Fig. 8D–F). In skin cutaneous melanoma (SKCM), the expression of IFN-γ is advantageous for patient prognosis (Fig. 8G); conversely, in thymoma (THYM), IFN-γ expression is detrimental to patient prognosis (Fig. 8H). Nonetheless, in uterine corpus endometrial carcinoma (UCEC), the expression of IFN-γ is beneficial for patient prognosis (Fig. 8I, J). Furthermore, we further validated the relationship between IFNG and prognosis in different cancer patients through the GEPIA website, with the results aligning closely with the aforementioned analysis, indicating a negative correlation between IFNG expression and the survival time of THYM patients (Fig. 8K).
A–J Analysis of the overall survival time in different cancer patients (n = 10327) based on the expression level of IFNG; K Validation of the correlation between the expression level of IFNG and prognosis in THYM patients (n = 115) using the GEPIA website. LGG brain lower grade glioma, OV ovarian serous cystadenocarcinoma, SARC sarcoma, SKCM skin cutaneous melanoma, THYM thymoma, UVM uveal melanoma.
In conclusion, the expression levels of Ferroptosis-related IRGs in tumor tissues of cancer patients are associated with their prognosis, with IFNG showing the closest relationship to patient prognosis.
Impact of IFN-α and IFN-γ on CD8 + T cell-mediated ferroptosis in cancer cells
It is well known that IFN-γ is mainly produced by CD8 + T cells39,42. We first examined the changes in IFN-γ levels in CD8 + T cells after treatment with IFN-α and silencing of HSPA6 or DNAJB1. The results showed that, compared to normal CD8 + T cells (con), IFN-α inhibited the production of IFN-γ in CD8 + T cells, while silencing of HSPA6 or DNAJB1 reversed this phenomenon (Fig. 9A).
A ELISA experiment examining the influence of IFN-α on the IFN-γ content of CD8 + T cells when HSPA6 or DNAJB1 are silenced; B MTT experiment measuring cell viability of tumor cells after treatment with IFN-α; C Flow cytometry analysis to determine the level of apoptosis in tumor cells after treatment with IFN-α; D RT-qPCR experiment analyzing the expression levels of COX2, ACSL4, NOX1, and GPX4 mRNA in tumor cells after treatment with IFN-α; E ELISA experiment measuring the levels of MDA and LPO in tumor cells after treatment with IFN-α; F MTT experiment measuring cell viability of tumor cells after treatment with IFN-γ; G Flow cytometry analysis to determine the level of apoptosis in tumor cells after treatment with IFN-γ; H RT-qPCR experiment analyzing the expression levels of COX2, ACSL4, NOX1, and GPX4 mRNA in tumor cells after treatment with IFN-γ; I ELISA experiment measuring the levels of MDA and LPO in tumor cells after treatment with IFN-γ; J RT-qPCR experiments detected the expression levels of COX2, ACSL4, NOX1, and GPX4 mRNA in tumor cells after treatment with IFN-γ and anti-IFN-γ antibodies; K ELISA experiments measured the levels of MDA and LPO in tumor cells after treatment with IFN-γ and anti-IFN-γ antibodies. Cell experiments were repeated three times. * indicates comparison with the control group, *p < 0.05.
To investigate whether IFN-α induction affects Ferroptosis in tumor cells, we co-cultured CD8 + T cells with tumor cells. Nine different cancer cell lines were chosen, and CD8 + T cells were stimulated with IFN-α in vitro. After continuous stimulation with IFN-α, we found that the cell viability of all tumor cells significantly increased while the apoptosis level decreased (Fig. 9B, C). Additionally, as shown in Fig. 9D, E, compared to the con group, the expression levels of COX2, ACSL4, and NOX1 mRNA in tumor cells from the IFN-α group significantly decreased, while GPX4 increased significantly. Furthermore, the levels of lipid peroxidation products MDA and LPO were significantly reduced, suggesting that CD8 + T cells attenuated Ferroptosis in co-cultured tumor cells when stimulated by IFN-α.
To further validate the impact of IFN-γ secreted by CD8 + T cells on Ferroptosis in tumor cells, we selected nine different cancer cell lines and performed cultivation and stimulation with IFN-γ in vitro. After continuous stimulation with IFN-γ, we observed significant inhibition of cell viability in all tumor cells, along with an increase in apoptosis levels (Fig. 9F, G). Moreover, as depicted in Fig. 9H, I, compared to the con group, the expression levels of COX2, ACSL4, and NOX1 mRNA in tumor cells from the IFN-γ group significantly increased, while GPX4 decreased significantly. Additionally, the levels of lipid peroxidation products MDA and LPO were significantly elevated, indicating that Ferroptosis occurred in tumor cells following stimulation with IFN-γ. Upon stimulation with IFN-γ, we treated the cells with IgG and anti-IFN-γ antibodies separately. The results demonstrated that compared to the IFN-γ + IgG group, the expression levels of COX2, ACSL4, and NOX1 mRNA in tumor cells was significantly decreased in the IFN-γ+anti-IFN-γ group, while GPX4 expression was significantly increased (Fig. 9J). Additionally, the levels of lipid peroxidation products MDA and LPO were also significantly reduced (Fig. 9K) in the IFN-γ+anti-IFN-γ group. Considering the results of the bioinformatics analysis in Fig. 8, we believe that the main reason for the favorable prognosis of cancer patients with high IFN-γ expression is likely due to its induction of Ferroptosis in tumor cells.
Enhanced antitumor effect of PD-1 blockade by silencing HSPA6 in CD8 + T cells
Tumor immunotherapy is an emerging treatment for drug-resistant tumor cells and involves the use of the human immune system to combat diseases43. Research has shown that tumor-specific CD8 + T cells play a crucial role in anti-tumor immunity, making them a key target for immunotherapy, especially when combined with anti-PD-1 therapy for enhanced efficacy44,45,46. To investigate the impact of HSPA6 or DNAJB1-treated CD8 + T cells on the efficacy of anti-PD-1 therapy, we established a subcutaneous xenograft model of melanoma using the SK-MEL-1 cell line in mice (immunotherapy-tolerant model). Subsequently, all mice were intraperitoneally injected with anti-PD-1.
The results revealed that compared to the control group, the growth of tumors in the sh-NC group showed no significant changes. However, the growth of tumors in the sh-HSPA6 or sh-DNAJB1 groups was suppressed, resulting in reduced tumor size and weight. Additionally, the use of anti-PD-1 alone had no significant effect on tumor growth compared to the control group. In contrast, treatment with sh-HSPA6+anti-PD-1 or sh-DNAJB1+anti-PD-1 was beneficial in suppressing the growth of implanted SK-MEL-1 tumors, leading to decreased tumor size and weight (Fig. 10A, B). Additionally, sh-HSPA6 or sh-DNAJB1 downregulated the expression of PD1, TIM3, LAG3, and CTLA4 in tumor tissues, upregulated the expression of Ki67 and CD69, and enhanced the secretion of effector factors in CD8 + T cells. Similarly, treatment with sh-HSPA6+anti-PD-1 or sh-DNAJB1+anti-PD-1 downregulated the expression of PD1, TIM3, LAG3, and CTLA4 in tumor tissues, upregulated the expression of Ki67 and CD69, and enhanced the secretion of effector factors in CD8 + T cells, when compared to the sh-NC+anti-PD-1 group (Fig. 10C–E). Moreover, sh-HSPA6 or sh-DNAJB1 upregulated the levels of IFN-γ and Ferroptosis in tumor tissues. In comparison to the sh-NC+anti-PD-1 group, treatment with sh-HSPA6+anti-PD-1 or sh-DNAJB1+anti-PD-1 increased the levels of IFN-γ and Ferroptosis in tumor tissues (Fig. 10F–H). Based on the findings of previous studies33, we propose that the enhanced anti-tumor effect of PD-1 is attributed to the improvement of CD8 + T cell immune response through silencing HSPA6.
A Changes in tumor volume of subcutaneous transplanted tumors in mice within 30 days (n = 6); B Representative images and weights of subcutaneous transplanted tumors in mice after 30 days of the experiment (n = 6); C Expression levels of exhausted markers PD1, TIM3, LAG3, CTLA4, etc. in CD8 + T cells detected by flow cytometry (n = 6); D Expression levels of Ki67 and CD69 in CD8 + T cells detected by flow cytometry (n = 6); E Expression levels of IFN-γ, IL-2, and TNF-α in CD8 + T cells detected by flow cytometry (n = 6); F Detection of IFN-γ content in tumor tissue by ELISA (n = 6); G Expression levels of COX2, ACSL4, NOX1, and GPX4 mRNA in tumor tissue detected by RT-qPCR (n = 6); H Levels of MDA and LPO in tumor tissue detected by ELISA (n = 6). * indicates a comparison between the two groups, *p < 0.05.
Discussion
The interferon signaling pathway plays a critical role in tumor immunotherapy47,48,49. The production of interferons activates the immune system and hinders the growth and spread of tumor cells, making its association with tumor immunotherapy a subject of extensive research and focus4,50,51. In this study, the impact mechanism of the interferon signaling pathway on Ferroptosis in tumor cells was revealed, providing important insights into the underlying mechanisms of tumor immunotherapy.
Compared to previous studies, this research employed single-cell sequencing technology, enabling more accurate analysis of the functional state and gene expression changes in CD8 + T cell subpopulations25. This application significantly reduced human error and enhanced data quality, particularly in uncovering the role of CD8 + T cells in the interferon signaling pathway52,53,54. The employment of single-cell sequencing technology in this study yielded more precise conclusions and offered new perspectives and ideas for further research.
A key gene, HSPA6, was found by this study to be highly expressed in CD8 + T cells and played a vital role in regulating CD8 + T cell exhaustion. HSPA6 enhanced the immune-killing effect of CD8 + T cells on tumors by modulating the cell cycle and mitochondrial function, thereby improving the effectiveness of tumor immunotherapy55. This finding provides crucial clues for a deeper exploration of the mechanisms of CD8 + T cell exhaustion and the identification of new therapeutic targets for exhaustion. However, the heat shock protein family is induced when encountering pathogens. Most cancer patients have compromised immune responses to some extent, making them susceptible to pathogenic attacks. Therefore, the expression of HSPA6, a biomarker for depleted T cells in cancer patients, is likely related to the patient’s potential infection status rather than directly associated with the tumor condition. To control for this potential confounding factor, we used bioinformatics to analyze whether the expression level of HSPA6 directly correlates with tumor staging. Our study found a correlation in colorectal and esophageal cancers but not in pancreatic, endometrial, renal, or thyroid cancers. To delve deeper, we retrieved data on HSPA6 expression in tumor cells from databases; however, individual tumor cells may not adequately represent the overall tumor status, especially considering the considerable inconsistency in HSPA6 expression among different tumor cells of the same cancer type. Furthermore, the tumor microenvironment likely plays a pivotal role in this process, warranting further in-depth investigation.
The key gene IFN-γ, revealed by this study, plays an important role in interferon-induced Ferroptosis of tumor cells. IFN-γ promotes tumor cell iron starvation and Ferroptosis by modulating the antioxidant enzyme system and iron metabolism pathways in tumor cells9,56,57. In comparison to previous studies, this research delved deeper into the potential role of IFN-γ in tumor treatment, offering new perspectives and directions for the development of tumor immunotherapy.
The influence of IFN-α on CD8 + T cells and tumor cells was experimentally validated in vitro, supporting the regulatory role of the interferon signaling pathway in tumor cell Ferroptosis at the cellular level. IFN-α significantly increased the activity and cytotoxicity of CD8 + T cells while inhibiting the growth and spread of tumor cells. This finding further emphasizes the importance and potential role of the interferon signaling pathway in tumor immunotherapy.
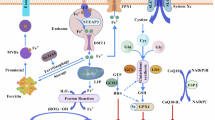
The in vivo study using a mouse model validated the in vitro results and evaluated changes in tumor growth and migration, highlighting the potential enhanced therapeutic effect of PD-1 treatment. Continued stimulation with IFN-α led to exhaustion of CD8 + T cells by activating the HSPA6-DNAJB1 axis, which consequently decreased IFN-γ secretion and inhibited IFN-γ-induced ferroptosis in tumor cells. Simultaneously, the silence of HSPA6 and anti-PD-1 treatment effectively suppressed tumor cell proliferation in the host (Fig. 11). Currently, PD-1 (PDL-1) antibodies have gradually gained FDA approval for the treatment of various cancers, including non-small cell lung cancer, renal cell carcinoma, bladder cancer, Hodgkin lymphoma, head and neck cancer, Merkel cell carcinoma, liver cancer, and gastroesophageal junction cancer58,59. However, it must be acknowledged that only a small subset of cancer patients can ultimately benefit from these treatment regimens. Further analysis reveals that only patients with CD8 + T cell infiltration in the tumor exhibit a more favorable clinical response to immunotherapy60. HSPA6 is a member of the HSP70 protein family and shows high expression in various cancers61,62,63,64,65,66. Studies suggest that the HSP70 family promotes malignant tumor progression by inhibiting apoptosis, enabling tumor cells to evade senescence mechanisms, interfering with immunity, and promoting angiogenesis67,68. Additionally, HSP70 (HSPA6 or HSPA7) and TLR-2 are positively correlated with dendritic cells, Tregs, or CD4 + T cells, and negatively correlated with CD3+ or CD8 + T cells, indicating that activation of the HSP70/TLR-2 axis mediates immune suppression by antagonizing CD8 + T cells69. In recent years, HSPA6 has also been identified as an immune-related gene and biomarker across different cancer types70,71. These research findings provide a solid theoretical basis and practical support for the enhancing effect of the HSPA6-DNAJB1 axis on the potential efficacy of PD-1 therapy.
Molecular mechanism diagram of IFN-α inducing CD8 + T cell exhaustion and reducing tumor cell Ferroptosis through the HSPA6-DNAJB1 axis.
The significant findings of this study underscore the scientific and clinical value of the interferon signaling pathway in tumor immunotherapy. The research provides important clues for investigating the mechanisms underlying the interferon signaling pathway in CD8 + T cell exhaustion and tumor cell Ferroptosis and offers a theoretical basis for optimizing the application strategies of the interferon signaling pathway. However, this study has some limitations. Firstly, it heavily relies on mouse models and in vitro experiments, needing more clinical sample verification and clinical data support. Secondly, although the study reveals the important role of the interferon signaling pathway in CD8 + T cell exhaustion and tumor cell Ferroptosis, the detailed molecular mechanisms and signal transduction pathways remain to be fully elucidated. Future research can address these limitations through comprehensive and in-depth experimental designs and bioinformatics analysis.
Looking ahead, further exploration of the mechanisms linking the interferon signaling pathway with CD8 + T cell exhaustion and tumor cell Ferroptosis is warranted. Firstly, we can further investigate other factors regulating CD8 + T cell activity and exhaustion and explore their interaction with the interferon signaling pathway to identify additional targets for therapy. Secondly, a more thorough understanding of the molecular mechanisms underlying tumor cell Ferroptosis would help identify key genes and pathways regulating tumor cell death. Lastly, the application strategies of the interferon signaling pathway in tumor immunotherapy, including the rational design of interferon dosage and timing, as well as its combination with other tumor treatment approaches, should be optimized. Through these endeavors, we can make progress and better utilize the interferon signaling pathway for treating tumors, ultimately bringing improved therapeutic outcomes for patients.
Methods
Ethical statement
The Animal Ethics Committee of Zibo Central Hospital Affiliated to Binzhou Medical University has approved this experimental program and animal usage scheme (20190305).
Tumor sample acquisition and single-cell sequencing
The human cell lines TE354.T (Bio-132776) derived from basal cell carcinoma, BJAB (Bio-133059) representing B-cell lymphoma, TE-1 (Bio-73172) for esophageal cancer, KMS-11 (Bio-129971) for multiple myeloma, OVCAR-3 (Bio-54032) originating from ovarian cancer, PANC-1 (Bio-73125) from pancreatic cancer, A-498 (Bio-73195) from renal carcinoma, ACT-1 (Bio-54288) from thyroid carcinoma and KLE (Bio-73074) from endometrial carcinoma were procured from the Biobw database in China. These cell lines were cultured in DMEM medium (11965092, Gibco) supplemented with 20% fetal bovine serum (FBS, 12483020, Gibco), 2 mM L-glutamine (A2916801, Gibco), 100 units/mL penicillin, and 100 µg/mL streptomycin (15140148, Gibco). The cultures were maintained at 37 °C in a 5% carbon dioxide atmosphere72. All materials were purchased from Thermofisher.
Subsequently, NOG mice (408, NOD.Cg-PrkdcscidIL2rgtm1Sug/JicCrl) were procured from Beijing Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China). After intratumoral injection of 4 × 106 various cancer cells and intravenous injection of 2 × 106 normal human CD8 + T cells, the mice were euthanized 30 days post-injection. Tumor tissue, adjacent tissue (defined as “tumor tissue” and “normal tissue”), and CD8 + T cells were extracted for single-cell sequencing. Normal human T cells were obtained from human spleen sourced from Mingzhou Biotech (Zhejiang, China) and purified using a CD8 + T Cell Isolation Kit (Invitrogen 11348D). CD8 + T cells were cultured in RPMI 1640 medium (Gibco 11875101) containing 30 U/mL IL-2 (Gibco PHC0021). Cell purity, assessed by flow cytometry using anti-CD3 + T cell antibody (Biolegend 317308) and anti-CD8 + T cell antibodies (Biolegend 344722, 344706), exceeded 95%, meeting experimental standards73. All materials were purchased from Sigma and Thermofisher.
Single-cell cDNA library preparation and quality control analysis of scRNA-seq datasets
CD8 + T cells were loaded onto the 10x Genomics Chromium chip following the manufacturer’s protocol. Gel bead-in-emulsion (GEM) formation was achieved by mixing single-cell suspension, gel beads, and oil in the 10x Genomics Chromium controller. After droplet formation, samples were transferred to PCR tubes and reverse transcribed on a T100 Thermal Cycler (Bio-Rad) at 53 °C for 45 min, followed by 85 °C for 5 min and then held at 4 °C. The generated cDNA was amplified, and its quality was assessed using an Agilent Bioanalyzer 2100. Libraries were constructed by adding P5 and P7 primers, Read 2 (reads 2 sequencing primer site), and Sample Index, followed by quality control checks before sequencing on an Illumina HiSeq4000 PE125 platform. The obtained sequencing data were further analyzed using 10x Cell Ranger (version 2.2.0) to convert the base call files (BCLs) from the Illumina sequencer to FASTQ files for alignment, filtering, barcode, and UMI counting. Human reference transcriptome read alignment was carried out with STAR (Spliced Transcripts Alignment to a Reference). Cell Ranger was employed for primary quality control (QC) to generate high-quality data.
Data processing and analysis of single-cell sequencing
The R software was used to classify normal and cancer cells in the dataset. The data was subjected to quality control, standardization, and filtering using Seurat, with the following criteria: min. cell = 3 & min. feature = 50, nFeature_RNA > 500, nCount_RNA > 1000, nCount_RNA < 20000, and percent.mt <25. The correlation between the number of genes and sequencing depth was analyzed using the Pearson correlation coefficient. The dimensionality-reduced single-cell RNA sequencing data was further analyzed using PCA and t-SNE algorithms, including clustering and nonlinear dimensionality reduction74. The R package “limma” was utilized to normalize and merge the expression matrices of normal and tumor samples in various cancers. Differential analysis was performed using the criteria: |log2FC | > 1 and p < 0.05. Subsequently, the R language package “ClusterProfiler” was employed for GO and KEGG enrichment analysis of differentially expressed genes, with p < 0.05 as the selection criterion. GeneMANIA (http://genemania.org/) and Coexpedia (https://www.coexpedia.org/) databases were used to identify downstream factors of key differentially expressed genes in order to validate their co-expression relationship with T cell exhaustion marker genes.
TCGA data download and organization
To download TCGA pan-cancer transcriptome data and clinical data, please visit the UCSC database (http://xena.ucsc.edu/). Use Perl scripting to organize these transcriptome data. To obtain a gene set associated with Ferroptosis, visit the Ferrdb website. Within this gene set, you can also find interferon-related genes (IRGs). Subsequently, extract the expression levels of IRGs from the pan-cancer dataset using the R package “limma” and analyze the differential expression of these genes in tumor and normal tissues across different cancer cases, as well as their correlation with gene expression. Finally, merge the IRG gene matrix from the pan-cancer dataset with survival information using the R software and analyze the association between IRGs and adverse prognosis using the “survival” package33.
Culturing CD8 + T cells
To induce T cell exhaustion, CD8 + T cells were cultured continuously for 8 days. Every 48 h, RPMI 1640 medium supplemented with 30 U/mL of IL-2 and Dynabeads™ CD3/CD28 (Thermofisher 11161D) was used for passaging. Throughout the entire process, recombinant human IFN-α (Sigma 1000 U/mL) was added. Subsequently, cells were incubated in the medium for 48 h and stimulated with Dynabeads™ CD3/CD28 for 4 h. Cells were collected for the detection of various parameters33.
Co-culture systems for cells
In order to assess the impact of transfected CD8 + T cells on tumor cells, we employed a cell Transwell co-culture system. A 0.4 µm Transwell chamber (3412, Corning, USA) was utilized for the co-culture of CD8 + T cells and tumor cells. Tumor cells were placed in the upper chamber with a density of 2 × 105/mL, while CD8 + T cells were placed in the lower chamber with a density of 1 × 106/mL. The co-culture was conducted for 48 h, after which tumor cells were extracted for further analysis75. In our experiments involving antibodies, we co-incubated an IgG antibody (BioXCell, BE0297) and a neutralizing antibody against IFN-γ (BioXCell, BE0235) in tumor cells following IFN-γ stimulation. Subsequently, the tumor cells were subjected to further analysis according to the methods stated in a previous publication76.
Transfection of cells
To generate the following cell lines (Supplementary Table 1), we employed a lentiviral-mediated transduction method: the silencing of HSPA6 in CD8 + T cells (sh-HSPA6), the silencing of DNAJB1 in CD8 + T cells (sh-DNAJB1), and a control group (sh-NC). Plasmids for the silencing of HSPA6 and DNAJB1, as well as their corresponding derivative lentiviruses, were obtained from Hanheng Biotechnology Co., Ltd. in Shanghai, China. The plasmids and helper plasmids were co-transfected into 293 T cells (bio-73410) to generate packaged lentiviruses, which were subsequently validated, amplified, and purified. For lentiviral-mediated cell transduction, 5 × 105 cells were seeded in a 6-well plate. When the cell confluence reached 70–90%, an appropriate amount of packaged lentivirus (MOI = 10, working titer approximately 5 × 106 TU/mL) and 5 μg/mL polybrene (Merck, TR-1003, USA) were added to the medium for transduction. Four hours after transduction, an equal volume of medium was added to dilute the polybrene, and the fresh medium was replaced after 24 h. To select stably transfected cell lines, 1 μg/mL puromycin (Thermo Fisher, A1113803, USA) was added 48 h post-transduction. All lentiviruses were transduced into extracted CD8 + T cells at a concentration of 5 µg/mL puromycin (A1113803, Gibco) with an MOI of 1072.
Flow cytometry
Cell counting technology was employed to calculate the quantity and concentration of exhausted T cells. For cell surface staining, cells were resuspended in 2% PBS and then incubated with the following antibodies for flow cytometry detection: anti-CD279 (PD-1, 557946, BD), anti-TIM3 (565558, BD), anti-CD223 (LAG3, 369307, BioLegend), anti-CTLA4 (369606, BioLegend), anti-Ki67 (558616, BD), and anti-CD69 antibodies (310904, BioLegend). After incubation on ice for 30 min, the cells were washed twice with 2% PBS and then subjected to flow cytometry analysis. For intracellular staining, cells were fixed and permeabilized, followed by incubation with fluorochrome-conjugated anti-IL-2 (500310, BioLegend), anti-IFN-γ (502515, BioLegend), and anti-TNF-α antibodies (559321, BD) at 4 °C for 30 min before flow cytometry analysis. Cell viability and apoptosis levels were assessed using the Beyotime Cell Proliferation Assay Kit (C0009S) and Apoptosis Assay Kit (C1062L), respectively. For cells requiring cytokine detection, T cells were stimulated with a cell stimulation mixture (phorbol 12-myristate 13-acetate, ionomycin, Brefeldin A, and monensin in the presence of a protein transport inhibitor, 00-4975-93, ThermoFisher) at 37 °C for 6 h, followed by staining as needed. Cell flow cytometry was conducted using the FACS Aria II Cell Sorter from BD Biosciences, and data analysis was performed using FlowJo software (TreeStar)77. All materials were purchased from BD, BioLegend, and ThermoFisher.
Immunoprecipitation experiment (Co-IP)
Initially, cells were cultured to appropriate density, the supernatant was removed, and cells were washed with 2 mL of chilled PBS solution. After the complete removal of PBS, 400 μL of IP lysis buffer (Beyotime P0013B) was added, and the cells were lysed on ice for 15 min. The proteins were scraped off using a cell scraper, transferred to a 1.5 mL Eppendorf tube, and sonicated at 4 °C using an ultrasonic homogenizer. Subsequently, the protein suspension was centrifuged at 4 °C for 15 min, and the supernatant was collected as the desired protein. Next, the 400 μL protein solution was divided into three parts: the first part for whole-cell protein preparation, the second part for the IgG group, and the third group for the IP group. The first group was directly mixed with 5× loading buffer to prepare the whole-cell lysate (WCL). The second group had 5 μL of IgG antibody added (Thermofisher #MA5-42729), and the third group had HSPA6 antibody added (Thermofisher #PA5-87019, 1:100) for overnight incubation. The following day, magnetic beads (Thermofisher #80106G) were added and incubated for 3 h to form a protein-antibody-magnetic bead complex. The complex was centrifuged in a 4 °C cold centrifuge, washed three times with RIPA lysis buffer (Beyotime P0013B), and finally, the magnetic bead pellet was retained and mixed with 2× loading buffer to prepare protein loading samples. Ultimately, the samples were subjected to standard protein electrophoresis procedures, incubated with the corresponding antibodies, developed in a gel imaging system, and the experimental results were saved for subsequent analysis78.
ELISA assay
CD8 + T cells or tumor tissues were collected and dispersed, followed by lysis. After centrifugation, the supernatant of the lysate was collected, and the levels of IFN-γ (H025-1), MDA (S0131M, Beyotime), and LPO (A106-1, Nanjing Jiancheng) were determined using an enzyme-linked immunosorbent assay (ELISA) method provided by the reagent supplier, Nanjing Jiancheng79.
RT-qPCR assay
First, total RNA was isolated from cells and tissues. Tissues (100 mg each) were homogenized in 1 mL TRIzol reagent (10296010, Thermo Fisher), followed by the addition of 200 μL chloroform and vigorous mixing. The mixture was then centrifuged at 4 °C, 12,000 g, for 10 min. The supernatant was collected, and RNA was precipitated by mixing with 500 μL isopropanol. The RNA was dissolved in RNase-free water (10-30 μL) and quantified using a Nanodrop spectrophotometer (Nanodrop 3300, Thermo Fisher). Reverse transcription was performed on 1 μg of total RNA using TaqMan Reverse Transcription reagents (N8080234, Thermo Fisher). PCR analysis was performed on the samples using the PowerUp SYBR Green PCR Master Mix (A25741, Thermo Fisher). The mRNA levels of COX2, ACSL4, NOX1, and GPX4, as Fe-induced cell death markers, were determined using GAPDH as the internal control, and the relative expression levels of each gene were analyzed using the 2−ΔΔCT method80,81. The primer sequences are provided in Supplementary Table 2.
Western blot
Total protein from cells and tissues was extracted using RIPA lysis buffer (Beyotime P0013B) containing 4% protease inhibitor, following the manufacturer’s instructions (1 mL per 100 mg of tissue). The protein concentration was determined using the BCA protein assay kit (Beyotime P0010S). Equal amounts of protein samples were loaded onto a sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and subsequently transferred to a PVDF membrane (Beyotime FFP24) using a wet transfer method. The membrane was blocked in TBST (5 mL) containing 5% skimmed milk (Beyotime P0216-300 g) at room temperature for 1 h and then incubated overnight at 4 °C with the following primary antibodies: HSPA6-70 kDa (Thermo Fisher #PA5-87019, 1:500), DNAJB1-40 kDa (Thermo Fisher #PA5-17382, 1:500), and GAPDH-36 kDa (Thermo Fisher #PA1-987, 1:500). The aforementioned primary antibodies were purchased from Thermo Fisher. After washing with TBST, the membranes were incubated with a horseradish peroxidase-conjugated goat anti-rabbit secondary antibody (Beyotime A0208, 1:1000) for 1 h. Then, the membranes were developed using the ECL chemiluminescence detection system (Beyotime P0018FS) and visualized with the ChemiDoc XRS+ system (Bio-Rad). The protein quantification analysis was performed using ImageJ software by determining the ratio of the intensity of each protein to the intensity of the reference protein, GAPDH82. Three independent experiments were conducted for reproducibility. Please refer to Supplementary Fig. 4 for full, un-cropped images of all blots
Culturing tumor cell lines
Human colon cancer cells HT-29 (Bio-73059), human liver cancer cells HuH-7 (Bio-73061), human lung cancer cells A-549 (Bio-72958), human cervical cancer cells HeLa (Bio-73042), human breast cancer cells MCF-7 (Bio-73090), human gastric cancer cells MKN-45 (Bio-73233), and human skin melanoma cells SK-MEL-1 (Bio-73381) were obtained from the Chinese Microorganism Culture Collection Center (Biobw, China). All cells were cultured in DMEM medium (11965092, Gibco) supplemented with 20% fetal bovine serum (FBS, 12483020, Gibco), 2 mM L-glutamine (A2916801, Gibco), 100 units/mL penicillin, and 100 µg/mL streptomycin (15140148, Gibco) at 37 °C in a 5% carbon dioxide incubator. These materials were purchased from Thermo Fisher72.
Generation of animal models
After subcutaneous injection of 4 × 106 SK-MEL-1 cells into NOG mice, the mice were randomly divided into four groups: con group (intravenous injection of normal CD8 + T cells not transfected with lentivirus + intraperitoneal injection of IgG1 isotype antibody), anti-PD-1 group (intravenous injection of normal CD8 + T cells not transfected with lentivirus + intraperitoneal injection of anti-PD-1 antibody), sh-NC+anti-PD-1 group (intravenous injection of control lentivirus-transfected CD8 + T cells + intraperitoneal injection of anti-PD-1 antibody), and sh-HSPA6+anti-PD-1 group (intravenous injection of sh-HSPA6 lentivirus-transfected CD8 + T cells + intraperitoneal injection of anti-PD-1 antibody), with 6 mice per group. The con group and anti-PD-1 group were injected with normal CD8 + T cells not transfected with lentivirus via tail vein injection at a dose of 2 × 106 cells/100 µLPBS83. The sh-NC+anti-PD-1 group was injected with control lentivirus-transfected CD8 + T cells at a dose of 2 × 106 cells/100 µLPBS83. The sh-HSPA6+anti-PD-1 group was injected with sh-HSPA6-lentivirus transfected CD8 + T cells at a dose of 2 × 106 cells/100 µLPBS83. Except for the con group, the other three groups of mice were given a dose of 2 mg anti-PD-1 antibody (329902, BioLegend) per 1 kg of body weight. The anti-PD-1 antibody was dissolved in 100 µL PBS and administered by intraperitoneal injection for PD-1 blockade therapy twice a week. The con group was injected with an equal dose of IgG1 isotype antibody (401402, BioLegend)84. Mice were observed every 5 days, and after 30 days, approved animal experiments were performed to investigate tumor tissue. The functionality of CD8 + T cells in tumor tissue was evaluated by flow cytometry83.
Acquisition of mouse CD8 + T cells
The method for obtaining CD8 + T cells from tumor tissue is as follows29. First, the tumor tissue is cut into 1–2 mm pieces and incubated with a final concentration of 30 μg/ml DNase I (D7076, Beyotime), 0.1 mg/ml hyaluronidase (ST1384, Beyotime), and 1 mg/ml collagenase (ST2303, Beyotime) at room temperature with gentle shaking for 2 h. Subsequently, a single-cell suspension is prepared by passing through a 70 μm filter and layered onto 10 ml of Ficoll-Paque (10771, Sigma). Centrifugation at 400 g and 20 °C for 30 min separates the lymphocyte layer as a milky interphase. Next, the isolated cells are further purified using a CD8 + T cell isolation kit (11348D, Invitrogen). To ensure purity meeting experimental standards, the isolated cells are subjected to flow cytometry analysis using anti-CD3 + T cell antibodies (317308, Biolegend) and anti-CD8 + T cell antibodies (344722, 344706, Biolegend), aiming for purity exceeding 95%.
Statistical analysis
All data were analyzed using the SPSS 21.0 statistical software produced by IBM. Numerical data were reported as mean ± standard deviation. Comparisons between two groups were conducted using an independent t test, while comparisons among multiple groups were performed using one-way analysis of variance (ANOVA). For data comparisons at different time points, we utilized repeated measures ANOVA. Statistical significance was considered when p < 0.05, p < 0.01, or p < 0.001.
Data availability
All sequencing data generated and analyzed during this study have been deposited in a community-endorsed public repository as required by the Nature Portfolio data availability policy. The data are available in the Sequence Read Archive (SRA), which is part of the International Nucleotide Sequence Database Collaboration (INSDC), under BioProject accession number PRJNA1140668. The submission ID for this project is SUB14628138.
The following accession codes have been assigned:
Disease group:
SRR30009925
SRR30009924
SRR30009915
SRR30009914
SRR30009913
SRR30009912
SRR30009911
SRR30009910
SRR30009909
Control group:
SRR30009908
SRR30009923
SRR30009922
SRR30009921
SRR30009920
SRR30009919
SRR30009918
SRR30009917
SRR30009916
The data are accessible at the National Center for Biotechnology Information (NCBI) SRA repository.
For further information or queries regarding the dataset, please contact Peiqing Zhao at bzjzzpq@sdu.edu.cn.
Change history
11 January 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41698-025-00807-x
References
Jiang, M. et al. cGAS-STING, an important pathway in cancer immunotherapy. J. Hematol. Oncol. 13, 81 (2020).
Bagheri, A. et al. Interferon-inducer antivirals: potential candidates to combat COVID-19. Int. Immunopharmacol. 91, 107245 (2021).
Snell, L. M., McGaha, T. L. & Brooks, D. G. Type I interferon in chronic virus infection and cancer. Trends Immunol. 38, 542–557 (2017).
Benson, L. N., Liu, Y., Deck, K., Mora, C. & Mu, S. IFN-γ contributes to the immune mechanisms of hypertension. Kidney 3, 2164–2173 (2022).
Iuliano, M. et al. Virus-induced tumorigenesis and IFN system. Biology 10, 994 (2021).
Pujantell, M. & Altfeld, M. Consequences of sex differences in Type I IFN responses for the regulation of antiviral immunity. Front. Immunol. 13, 986840 (2022).
El-Asmi, F., McManus, F. P., Thibault, P. & Chelbi-Alix, M. K. Interferon, restriction factors and SUMO pathways. Cytokine Growth Factor Rev. 55, 37–47 (2020).
Zhou, L. et al. A dual role of type I interferons in antitumor immunity. Adv. Biosyst. 4, e1900237 (2020).
Cai, H., Ren, Y., Chen, S., Wang, Y. & Chu, L. Ferroptosis and tumor immunotherapy: a promising combination therapy for tumors. Front. Oncol. 13, 1119369 (2023).
Fei, W. et al. Bioactive metal-containing nanomaterials for ferroptotic cancer therapy. J. Mater. Chem. B 8, 10461–10473 (2020).
Huang, Y., Wang, S., Ke, A. & Guo, K. Ferroptosis and its interaction with tumor immune microenvironment in liver cancer. Biochim. Biophys. Acta Rev. Cancer 1878, 188848 (2023).
Bystrom, L. M. & Rivella, S. Cancer cells with irons in the fire. Free Radic. Biol. Med. 79, 337–342 (2015).
Elliott, R. L., Elliott, M. C., Wang, F. & Head, J. F. Breast carcinoma and the role of iron metabolism. A cytochemical, tissue culture, and ultrastructural study. Ann. N.Y. Acad. Sci. 698, 159–166 (1993).
Morales, M. & Xue, X. Targeting iron metabolism in cancer therapy. Theranostics 11, 8412–8429 (2021).
Chen, Y., Li, X., Wang, S., Miao, R. & Zhong, J. Targeting Iron metabolism and ferroptosis as novel therapeutic approaches in cardiovascular diseases. Nutrients 15, 591 (2023).
Liao, P. et al. CD8+ T cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell 40, 365–378.e6 (2022).
Ke, S., Wang, C., Su, Z., Lin, S. & Wu, G. Integrated analysis reveals critical ferroptosis regulators and FTL contribute to cancer progression in hepatocellular carcinoma. Front. Genet. 13, 897683 (2022).
Ng, C. Y. et al. Targeting the elevated IFN-γ in vitiligo patients by human anti- IFN-γ monoclonal antibody hampers direct cytotoxicity in melanocyte. J. Dermatol. Sci. 110, 78–88 (2023).
Huang, Y., Jia, A., Wang, Y. & Liu, G. CD8+ T cell exhaustion in anti-tumour immunity: the new insights for cancer immunotherapy. Immunology 168, 30–48 (2023).
Rahim, M. K. et al. Dynamic CD8+ T cell responses to cancer immunotherapy in human regional lymph nodes are disrupted in metastatic lymph nodes. Cell 186, 1127–1143.e18 (2023).
Hsieh, C. H. et al. An innovative NRF2 nano-modulator induces lung cancer ferroptosis and elicits an immunostimulatory tumor microenvironment. Theranostics 11, 7072–7091 (2021).
Gocher-Demske, A. M. et al. IFNγ-induction of TH1-like regulatory T cells controls antiviral responses. Nat. Immunol. 24, 841–854 (2023).
Saadey, A. A. et al. Rebalancing TGFβ1/BMP signals in exhausted T cells unlocks responsiveness to immune checkpoint blockade therapy. Nat. Immunol. 24, 280–294 (2023).
Zhang, Q. et al. Immunosuppressive lncRNA LINC00624 promotes tumor progression and therapy resistance through ADAR1 stabilization. J. Immunother. Cancer 10, e004666 (2022).
Jovic, D. et al. Single-cell RNA sequencing technologies and applications: a brief overview. Clin. Transl. Med. 12, e694 (2022).
Catakovic, K., Klieser, E., Neureiter, D. & Geisberger, R. T cell exhaustion: from pathophysiological basics to tumor immunotherapy. Cell Commun. Signal. 15, 1 (2017).
Shao, M. et al. Inhibition of calcium signaling prevents exhaustion and enhances anti-leukemia efficacy of CAR-T cells via SOCE-calcineurin-NFAT and glycolysis pathways. Adv. Sci. 9, e2103508 (2022).
Wasiuk, A. et al. CD27-mediated regulatory T cell depletion and effector T cell costimulation both contribute to antitumor efficacy. J. Immunol. 199, 4110–4123 (2017).
Liu, Z. et al. Epigenetic reprogramming of Runx3 reinforces CD8 + T-cell function and improves the clinical response to immunotherapy. Mol. Cancer 22, 84 (2023).
Chakiryan, N. H. et al. Geospatial characterization of immune cell distributions and dynamics across the microenvironment in clear cell renal cell carcinoma. J. Immunother. Cancer 11, e006195 (2023).
Desai, P. N., et al. Single-cell profiling of CD8+ T cells in acute myeloid leukemia reveals a continuous spectrum of differentiation and clonal hyperexpansion. Cancer Immunol. Res. CIR-22-0961. https://doi.org/10.1158/2326-6066.CIR-22-0961 (2023).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, abe6474 (2021).
Chen, W. et al. Chronic type I interferon signaling promotes lipid-peroxidation-driven terminal CD8+ T cell exhaustion and curtails anti-PD-1 efficacy. Cell Rep. 41, 111647 (2022).
Lukhele, S. et al. The transcription factor IRF2 drives interferon-mediated CD8+ T cell exhaustion to restrict anti-tumor immunity. Immunity 55, 2369–2385.e10 (2022).
Zhu, Y. et al. The combination of PD-1 blockade with interferon-α has a synergistic effect on hepatocellular carcinoma. Cell. Mol. Immunol. 19, 726–737 (2022).
Liu, S. et al. Multi-organ landscape of therapy-resistant melanoma. Nat. Med. 29, 1123–1134 (2023).
He, W. et al. STRA6 regulates tumor immune microenvironment and is a prognostic marker in BRAF-mutant papillary thyroid carcinoma. Front. Endocrinol. 14, 1076640 (2023).
Melese, E. S. et al. CCL5 production in lung cancer cells leads to an altered immune microenvironment and promotes tumor development. Oncoimmunology 11, 2010905 (2021).
Liu, Y. et al. Multi-enzyme co-expressed dual-atom nanozymes induce cascade immunogenic ferroptosis via activating interferon-γ and targeting arachidonic acid metabolism. J. Am. Chem. Soc. 145, 8965–8978 (2023).
Qiu, S. et al. Mitochondria-localized cGAS suppresses ferroptosis to promote cancer progression. Cell Res. 33, 299–311 (2023).
Shen, D. et al. PARPi treatment enhances radiotherapy-induced ferroptosis and antitumor immune responses via the cGAS signaling pathway in colorectal cancer. Cancer Lett. 550, 215919 (2022).
Cui, K. et al. Restraint of IFN-γ expression through a distal silencer CNS-28 for tissue homeostasis. Immunity 56, 944–958.e6 (2023).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Koyama-Nasu, R. et al. CD69 imposes tumor-specific CD8+ T-cell fate in tumor-draining lymph nodes. Cancer Immunol. Res. 11, 1085–1099 (2023).
Naito, Y. et al. Tumor-derived semaphorin 4A improves PD-1-blocking antibody efficacy by enhancing CD8+ T cell cytotoxicity and proliferation. Sci. Adv. 9, eade0718 (2023).
Deng, G. et al. Ibrutinib Inhibits BTK signaling in tumor-infiltrated B cells and amplifies antitumor immunity by PD-1 checkpoint blockade for metastatic prostate cancer. Cancers 15, 2356 (2023).
Madani, A. Y. et al. Signal transducer and activator of transcription 3 (STAT3) suppresses STAT1/interferon signaling pathway and inflammation in senescent preadipocytes. Antioxidants 10, 334 (2021).
Han, J., Wu, M. & Liu, Z. Dysregulation in IFN-γ signaling and response: the barricade to tumor immunotherapy. Front. Immunol. 14, 1190333 (2023).
Bhat, A. A. et al. Uncovering the complex role of interferon-gamma in suppressing type 2 immunity to cancer. Cytokine 171, 156376 (2023).
Liu, N., Pang, X., Zhang, H. & Ji, P. The cGAS-STING pathway in bacterial infection and bacterial immunity. Front. Immunol. 12, 814709 (2022).
Agarwala, S. Improving survival in patients with high-risk and metastatic melanoma: immunotherapy leads the way. Am. J. Clin. Dermatol. 4, 333–346 (2003).
Friedensohn, S., Khan, T. A. & Reddy, S. T. Advanced methodologies in high-throughput sequencing of immune repertoires. Trends Biotechnol. 35, 203–214 (2017).
Phetsouphanh, C., Zaunders, J. J. & Kelleher, A. D. Detecting antigen-specific T cell responses: from bulk populations to single cells. Int. J. Mol. Sci. 16, 18878–18893 (2015).
Clauze, A., Enose-Akahata, Y. & Jacobson, S. T cell receptor repertoire analysis in HTLV-1-associated diseases. Front. Immunol. 13, 984274 (2022).
Song, B., Shen, S., Fu, S. & Fu, J. HSPA6 and its role in cancers and other diseases. Mol. Biol. Rep. 49, 10565–10577 (2022).
Jorgovanovic, D., Song, M., Wang, L. & Zhang, Y. Roles of IFN-γ in tumor progression and regression: a review. Biomark. Res. 8, 49 (2020).
Burke, J. D. & Young, H. A. IFN-γ: a cytokine at the right time, is in the right place. Semin. Immunol. 43, 101280 (2019).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Zhang, L. et al. HSPA6, a novel prognostic and therapeutic biomarker, associated with Ming classification in gastric cancer. J. Clin. Lab. Anal. 37, e24763 (2023).
Guan, Y. et al. Upregulation of HSPA1A/HSPA1B/HSPA7 and downregulation of HSPA9 were related to poor survival in colon cancer. Front. Oncol. 11, 749673 (2021).
Shen, S., Wei, C. & Fu, J. RNA-sequencing reveals heat shock 70-kDa Protein 6 (HSPA6) as a novel thymoquinone-upregulated gene that inhibits growth, migration, and invasion of triple-negative breast cancer cells. Front. Oncol. 11, 667995 (2021).
Guo, X. et al. Identification of the prognostic value of immune-related genes in esophageal cancer. Front. Genet. 11, 989 (2020).
Wang, L. et al. Identification of the prognostic value of immune gene signature and infiltrating immune cells for esophageal cancer patients. Int. Immunopharmacol. 87, 106795 (2020).
Chen, H., Luo, J. & Guo, J. Construction and validation of a 7-immune gene model for prognostic assessment of esophageal carcinoma. Med. Sci. Monit.: Int. Med. J. Exp. Clin. Res. 26, e927392 (2020).
Hu, C. et al. Heat shock proteins: Biological functions, pathological roles, and therapeutic opportunities. MedComm 3, e161 (2022).
Richter, K., Haslbeck, M. & Buchner, J. The heat shock response: life on the verge of death. Mol. Cell 40, 253–266 (2010).
Feng, H. et al. Excessive HSP70/TLR2 activation leads to remodeling of the tumor immune microenvironment to resist chemotherapy sensitivity of mFOLFOX in colorectal cancer. Clin. Immunol. 245, 109157 (2022).
Sun, X. & Zhang, T. Identification of immune-related signature for the prognosis and benefit of immunotherapy in triple-negative breast cancer. Front. Genet. 13, 1067254 (2022).
Zhang, Z. et al. Development of a prognostic signature for esophageal cancer based on nine immune related genes. BMC Cancer 21, 113 (2021).
Liu, Z. et al. CircRNA-5692 inhibits the progression of hepatocellular carcinoma by sponging miR-328-5p to enhance DAB2IP expression. Cell Death Dis. 10, 900 (2019).
Jiang, G. et al. Androgen receptor affects the response to immune checkpoint therapy by suppressing PD-L1 in hepatocellular carcinoma. Aging 12, 11466–11484 (2020).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Wu, M. et al. Gene expression profiling of CD8+ T cells induced by ovarian cancer cells suggests a possible mechanism for CD8+ Treg cell production. Cell Prolif. 49, 669–677 (2016).
Clever, D. et al. Oxygen sensing by T cells establishes an immunologically tolerant metastatic niche. Cell 166, 1117–1131.e14 (2016).
Wang, X. et al. TOX promotes the exhaustion of antitumor CD8+ T cells by preventing PD1 degradation in hepatocellular carcinoma. J. Hepatol. 71, 731–741 (2019).
Cooper, J. M. et al. TBK1 provides context-selective support of the activated AKT/mTOR pathway in lung cancer. Cancer Res. 77, 5077–5094 (2017).
Guo, L., Wang, D., Alexander, H. Y., Ren, X. & Ma, H. Long non-coding RNA H19 contributes to spinal cord ischemia/reperfusion injury through increasing neuronal pyroptosis by miR-181a-5p/HMGB1 axis. Aging 14, 5449–5463 (2022).
Franke, M. et al. The NLRP3 inflammasome drives inflammation in ischemia/reperfusion injury after transient middle cerebral artery occlusion in mice. Brain Behav. Immun. 92, 223–233 (2021).
Peng, Z. et al. miR-211-5p alleviates focal cerebral ischemia-reperfusion injury in rats by down-regulating the expression of COX2. Biochem. Pharmacol. 177, 113983 (2020).
Gong, L. et al. RTN1-C mediates cerebral ischemia/reperfusion injury via ER stress and mitochondria-associated apoptosis pathways. Cell Death Dis. 8, e3080 (2017).
Kwon, H. J., Lee, S., Lee, H. H., Cho, H. & Jung, J. Korean red ginseng enhances immunotherapeutic effects of NK cells via eosinophils in metastatic liver cancer model. Nutrients 14, 134 (2021).
Burkard, M. et al. High-dose ascorbate in combination with Anti-PD1 checkpoint inhibition as treatment option for malignant melanoma. Cells 12, 254 (2023).
Acknowledgements
This study was supported by National Natural Science Foundation of China (81972002), Natural Science Foundation of Shandong Province (ZR2023QC168, ZR2021MC165, ZR2021MC083, ZR2023MC136), Xinjiang Uygur Autonomous Region Training Program of Innovation and Entrepreneurship for College Students (S202310760060).
Author information
Authors and Affiliations
Contributions
Wei Hu, Ziqian Zhao, Jianxin Du, Jie Jiang, Minghao Yang, Maojin Tian, Peiqing Zhao contributed to the conceptualization and design of the study. Wei Hu, Ziqian Zhao, Jianxin Du, and Jie Jiang performed the experiments and analyzed the data. Wei Hu, Ziqian Zhao, Jianxin Du, and Jie Jiang contributed to the interpretation of the results. Wei Hu, Ziqian Zhao, and Jianxin Du wrote the manuscript. Minghao Yang, Maojin Tian, and Peiqing Zhao provided critical revisions of the manuscript. Minghao Yang, Maojin Tian, and Peiqing Zhao supervised the study. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hu, W., Zhao, Z., Du, J. et al. Interferon signaling and ferroptosis in tumor immunology and therapy. npj Precis. Onc. 8, 177 (2024). https://doi.org/10.1038/s41698-024-00668-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-024-00668-w
This article is cited by
-
Metal-modulated T cell antitumor immunity and emerging metalloimmunotherapy
Cancer and Metastasis Reviews (2025)
-
METTL5-mediated m6A modification of UBE3C promotes osteosarcoma progression by suppressing ferroptosis via inducing AHNAK ubiquitination
Journal of Molecular Histology (2025)
-
Exome sequencing reveals new insights into the germline landscape of inflammatory breast cancer among Tunisian patients
Journal of Translational Medicine (2025)