Abstract
In breast cancer management, predicting axillary lymph node (ALN) metastasis using whole-slide images (WSIs) of primary tumor biopsies is a challenging and underexplored task for pathologists. We developed METACANS, an multimodal artificial intelligence (AI) model that integrates WSIs with clinicopathological features to predict ALN metastasis. METACANS was trained on 1991 cases and externally validated across five cohorts with a total of 2166 cases. Across all validation cohorts, METACANS achieved an area under the curve (AUC) of 0.733 (95% CI, 0.711–0.755), with an overall negative predictive value of 0.846, sensitivity of 0.820, specificity of 0.504, and balanced accuracy of 0.662. Without additional annotations, METACANS identified pathological imaging patterns linked to metastatic status, such as micropapillary growth, infiltrative patterns, and necrosis. While its predictive performance may not yet support immediate clinical application, METACANS addresses the task of predicting ALN metastasis using WSIs and clinicopathological features, and demonstrates the feasibility of multimodal AI approaches for preoperative axillary staging in breast cancer.
Similar content being viewed by others
Introduction
The global burden of breast cancer necessitates advancements in the precision and efficiency of diagnosis and staging, owing to its high incidence and mortality rates of 46.3 and 16.3 per 100,000 population, respectively1. Axillary lymph node (ALN) metastasis represents the migration of cancer cells from the primary tumor through the lymphatic system and often indicates a higher probability of further metastasis, complicating management and suggesting a poorer prognosis2. Therefore, the early detection and prediction of ALN metastasis can substantially impact patient outcomes.
Traditionally, due to challenges in the preoperative diagnosis of ALN metastasis, it has been diagnosed postoperatively based on ALN dissection (ALND), which is an invasive procedure for patients with breast cancer3,4,5. However, ALND can significantly impede lymphatic circulation, leading to an increased risk of lymphedema and chronic inflammation5. Recent advancements have aimed to predict ALN metastasis using sentinel lymph node biopsy (SLNB) to reduce the potential side effects. Nevertheless, intraoperative assessment of SLNB prolongs the operative time and carries risks such as lymphatic leakage or sensory loss4,6,7. Moreover, SLNB carries the risk of inaccurate predictions and does not utilize the pathological characteristics of the primary tumor8. Furthermore, although modalities like ultrasound and magnetic resonance imaging (MRI) are commonly used to assess ALN metastasis, they primarily provide structural and macro-level morphological information. In contrast, whole slide images (WSIs) enable a detailed examination of histopathological features at the cellular level, such as cancer cell invasion into lymphatic vessels, offering additional insights into lymphatic spread. As such, WSIs may provide a deeper understanding of the metastatic process. Consequently, diagnosing ALN metastasis directly from primary tumor samples could offer immense clinical value and enable preemptive measures before surgery.
WSIs from patients with breast cancer provide a valuable resource for predicting ALN metastasis from primary tumors9. They allow detailed exploration of morphological and proliferation characteristics, such as tubule formation, nuclear pleomorphism, and mitotic rate10,11,12. Recently, deep learning (DL) methods have advanced the analysis of these pathological traits in WSIs. For example, Xu et al.9 proposed a deep learning model that uses manually segmented tumor areas from WSIs as input to predict ALN status and discovered three nuclear characteristics most relevant to the diagnosis, thereby demonstrating that core needle biopsy data could be beneficial for predicting ALN status. Ding et al.13. Developed a multi-modal DL model that integrates clinicopathological data and digital pathological images from core needle biopsy specimens to predict breast cancer lymph node metastasis, achieving high accuracy across multiple validation sets, particularly in the triple-negative breast cancer subtype. One contrasting study, Marmé et al.14, showed that hematoxylin/eosin-stained images did not provide significant benefit in predicting sentinel lymph node status across all three external validation cohorts. While significant progress has been made, most studies have primarily focused on the detection and classification of metastatic WSIs in resected lymph nodes15. In addition, many studies have directly analyzed observable pathological characteristics such as cancer subtyping16. In contrast, the use of WSIs from primary tumor biopsy samples provides a less invasive approach for predicting ALN metastasis. Nevertheless, it is less explored and presents a greater challenge because it does not rely on the direct observation of the axilla.
Therefore, this study aimed to develop and validate an artificial intelligence (AI) model that predicts ALN metastasis based on WSIs from primary tumor biopsies and clinicopathological characteristics. By utilizing biopsy data before performing ALND, the proposed model offers the potential to reduce unnecessary invasive surgery.
Results
ALN metastasis prediction using deep learning and machine learning
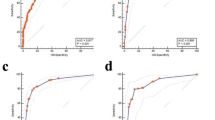
In this study, we proposed METACANS, a model combining the clinicopathological data-based model, ClinicML, and the WSI-based model, PathDL. The overall performance of each model for ALN metastasis prediction is summarized in Table 1. METACANS achieved an area under the receiver operating characteristics (ROC) curve (AUC) of 0.743 (95% confidence interval [CI], 0.694–0.792) in the internal validation cohort. For the five external validation cohorts A–E, AUCs of 0.715 (95% CI: 0.683–0.747), 0.785 (95% CI: 0.742–0.828), 0.681 (95% CI: 0.612–0.751), 0.743 (95% CI: 0.672–0.814), and 0.801 (95% CI: 0.728–0.873) were obtained, respectively (see Fig. 1a, b). When evaluating METACANS on the combined external validation datasets (cohorts A–E), the model achieved an AUC of 0.733 (95% CI, 0.711–0.755), which was significantly better than PathDL (AUC = 0.650 [95% CI, 0.626–0.674], p < 0.0001) and ClinicML (AUC = 0.713 [95% CI, 0.690–0.735], p = 0.0030), respectively.
a Receiver operating characteristics (ROC) curves for each validation cohort based on the METACANS. b Violin plots representing the probabilities of METACANS, with the black dashed line at 0.434 indicating the cut-off threshold for decision-making. AUC area under the ROC curve.
The maximum Youden index is used to select the cutoff on a cross-validation fold within the eight-fold cross-validation of the training set, as it maximizes the sum of sensitivity and specificity. As a result, the cutoff probabilities for METACANS, PathDL, and ClinicML were 0.434, 0.428, and 0.530, respectively.
In the internal validation cohort, the sensitivity and negative predictive value (NPV) were 0.795 (95% CI: 0.717–0.861) and 0.886 (95% CI: 0.839–0.924), respectively. The sensitivity and NPV for the external validation cohorts were as follows: 0.806 (95% CI: 0.763–0.845) and 0.824 (95% CI: 0.783–0.859) in validation cohort A, 0.839 (95% CI: 0.771–0.893) and 0.893 (95% CI: 0.846–0.930) in validation cohort B, 0.832 (95% CI: 0.741–0.901) and 0.750 (95% CI: 0.626–0.850) in validation cohort C, 0.746 (95% CI: 0.616–0.850) and 0.845 (95% CI: 0.758–0.911) in validation cohort D, and 0.935 (95% CI: 0.821–0.986) and 0.935 (95% CI: 0.821–0.986) in validation cohort E. Performing ALND when there was no actual ALN metastasis resulted in unnecessary surgical intervention. An example of visual illustration of the prognosis results is presented in Fig. 2.
Each icon, representing one of the patients with breast cancer, is color-coded as follows: green indicates true positives, red indicates false negatives, blue indicates true negatives, and yellow indicates false positives. ALN axillary lymph node, ALND ALN dissection.
In the internal validation cohort, our model correctly identified 105 out of 132 patients with ALN metastasis and 210 out of 368 patients without ALN metastasis. When we combined the prediction results from the five external validation cohorts, our model correctly identified 600 out of 732 patients with ALN metastasis as true positives (82.0% sensitivity) and 723 out of 1434 patients without ALN metastasis as true negatives (50.4% specificity). In our retrospective validation using a multi-institutional dataset of 2166 patients, METACANS achieved an overall NPV of 0.846. The evaluation of the clinical utility using decision curve analysis is shown in Supplementary Fig. 1.
Explainable AI for the deep learning model
We conducted further analyzes to identify the key information PathDL relies on for ALN metastasis prediction. PathDL applies attention-based multiple instance learning (ABMIL) to assign attention values to patches within WSIs, highlighting those most relevant to the model’s final decision17,18. Patches with higher attention values contributed more significantly to the slide-level predictions. We investigated the regions of interest in cases where PathDL correctly identified patients with ALN metastasis as true positives, as shown in Fig. 3a–c (see also Supplementary Figs. 2, 3, and 4 for more examples). Areas associated with micropapillary growth, infiltration, and necrosis are predominantly observed.
a true positive cases, b false positive cases, c false negative cases.
Feature importance analysis and the impact of clinicopathological characteristics on machine learning model performance
ClinicML was trained using patient tumor size, number of cancerous lesions, and age for ALN metastasis prediction. Subsequent feature importance analysis revealed that tumor size was the most influential factor, followed by the number of cancerous lesions and patient age. The feature importances are shown in Supplementary Fig. 5. Additionally, we trained several random forest (RF) models19, each integrating various characteristics from clinical reports. These characteristics include tubule formation, nuclear pleomorphism, mitotic count, histological grade, estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2), and Ki-67 labeling index (LI [%]), all of which are crucial for breast cancer diagnosis20,21,22,23. Nevertheless, the inclusion of these additional features did not improve the performance of any model (Supplementary Tables 1 and 2). To analyze the results, we conducted a comprehensive analysis using all available data to examine the relationship between the clinicopathological characteristics and ALN metastasis in a large patient cohort. From the total dataset, 6599 cases were considered, with 4471 (67.8%) being ALN metastasis-negative and 2128 (32.2%) being ALN metastasis-positive.
As detailed in Table 2, factors such as nuclear grade, histological grade, tubule formation, nuclear pleomorphism, ductal carcinoma in situ (DCIS), lobular carcinoma in situ (LCIS), and tumor size were found to significantly correlate with ALN metastasis. Clinically, this means that patients presenting more aggressive forms of these characteristics may have a higher risk of cancer spreading to the lymph nodes, indicating the need for careful monitoring and potential further intervention. Despite the statistical significance of these factors, the difference in the ALN metastatic rate, excluding tumor size, ranged between 8.5% and 14.5%. For instance, 24.8% of patients with histologic grade 1 exhibited ALN metastasis, compared to 33.3% of those with grade 3 tumors. However, when the histological grade variable was incorporated into the predictive model, there was no notable difference in the performance (Supplementary Table 2). Furthermore, while factors such as ER, PR, and HER2 status were statistically significant (p < 0.05), the differences in metastatic rates were only 3.2%–4.9%. This implies that while receptor status is crucial for guiding treatment decisions (e.g., hormonal or HER2-targeted therapies), its role in predicting immediate lymphatic spread may be limited.
Discussion
In this study, we developed a model called METACANS using DL and ML techniques to enable the preoperative prediction of ALN metastasis in patients with breast cancer. Recent protocols have used sentinel lymph node frozen biopsy during surgery to identify patients suspected of having ALN metastasis, and selective ALND was subsequently performed for those identified24,25,26. However, this approach leads to an extended duration of the intraoperative assessment. In contrast, METACANS offers the potential to predict the presence or absence of ALN metastasis before surgery, which may help avoid unnecessary ALND, though its clinical utility remains constrained by limited specificity. METACANS leverages imaging features extracted from preoperative primary tumor biopsy samples and clinicopathological characteristics. Across five independent validation cohorts with a total of 2166 patients with breast cancer, METACANS demonstrated the potential to avert invasive procedures in approximately 50% of patients without ALN metastasis while maintaining a sensitivity of 0.820 and an NPV of 0.846. However, these results should be interpreted with caution, as clinical practice decision-making often integrates a broader array of patient-specific data, such as radiological findings, genomic information, and physician expertise, which were not incorporated into our model. Therefore, METACANS should be regarded as a valuable but adjunctive diagnostic tool rather than a standalone solution.
Most previous studies using WSIs have primarily focused on analyzing metastatic cancer in ALN samples from breast cancer15,16. Although such studies are essential for understanding the metastatic nature of cancer and determining appropriate personalized treatments, they do not necessarily reduce the need for invasive procedures. This is because pathologists can diagnose the presence or absence of metastatic cancer in resected samples with considerable accuracy through microscopic inspection, given sufficient time15. In contrast, our study emphasizes the prediction of ALN metastasis using primary tumor biopsy samples, a more complex task that nonetheless holds substantial clinical value. Traditionally, pathologists determine the presence of ALN metastasis by examining ALN samples directly after ALND. Currently, SLNB is used as a less invasive alternative to ALND27. However, despite being less invasive, SLNB still involves surgery and lacks full standardization, with ongoing technical controversies24,26. Therefore, predicting ALN metastasis from primary tumor biopsy samples could further reduce the need for invasive procedures and provide earlier indications for metastasis. Despite its potential advantages, this approach has been largely unexplored because of the absence of robust methods.
In this study, we collected a training set from a single institution with 2491 patients and multiple external cohorts from independent institutions with 1090, 486, 246, 197, and 147 patients, respectively. Across these cohorts, our model demonstrated AUCs ranging from 0.681 to 0.801. Predicting distant metastases, such as ALN metastasis, from primary tumor biopsy samples is inherently challenging, making these outcomes promising. Predicting distant metastases, such as ALN metastasis, from primary tumor biopsy samples is inherently challenging, which makes these outcomes promising. A potential clinical advantage of our model lies in its ability to identify patients who may not require ALND based on the predicted absence of ALN metastasis. If used in conjunction with existing staging methods, such predictions could inform decisions aimed at reducing overtreatment. Therefore, our model may support the pursuit of less invasive approaches in managing breast cancer and potentially contribute to improved patient outcomes, reduced morbidity, and fewer postoperative complications, such as edema and inflammation. Nevertheless, given the model’s low specificity, there is a risk of recommending unnecessary ALND; its predictions must therefore be interpreted cautiously and validated against current clinical standards.
We analyzed important metadata for ALN metastasis prediction using the feature importance of ClinicML. Tumor size had the most significant effect, followed by the number of cancerous lesions and age. Larger tumors are more likely to be surrounded by enlarged and hyperplastic peritumoral lymphatic vessels, facilitating cancer cell metastasis through these vessels. This phenomenon is attributed to the tumor microenvironment, where tumor and stromal cells release factors that promote lymphangiogenesis, driving the growth of lymphatic endothelial cells and the formation of lymphatic capillaries28. Commonly used factors for breast cancer diagnosis, such as the ER, PR, and HER2, as well as traditional pathological imaging features like tubule formation, nuclear pleomorphism, and mitotic count, did not enhance predictive accuracy in this study. While these factors are critical for breast cancer diagnosis, their direct pathobiological impact on ALN metastasis remains uncertain20,21. Given the limited predictive contribution of traditional clinicopathological features, we next examined the potential of pathology images to reveal additional patterns associated with ALN metastasis.
Pathology images provide a microscopic view that reveals detailed structural information about cancer cells, including their aggregation, morphology, and size. This level of detail may not be as apparent in certain radiological images due to their lower resolution. In our study, we observed that patients with breast cancer exhibiting micropapillary growth, infiltrative patterns, and necrosis had a high incidence of ALN metastasis. In particular, the micropapillary growth pattern has been associated with lymphovascular invasion and an increased incidence of ALN metastasis29,30,31. Recognizing this pattern is important, as it is often linked to advanced disease stages32. Another study reported the presence of infiltrative patterns in patients with ALN metastasis33. These patterns may facilitate metastasis to lymph nodes by allowing tumor cells to access lymphatic channels more easily34. Therefore, careful assessment of such patterns can assist in stratifying metastasis risk. Notably, PathDL was trained exclusively on ALN metastasis status, which was not directly evident from the WSI of the primary tumor biopsy samples. Even without explicit annotations of the tumor area in WSI or consideration of other clinicopathological features, PathDL recognized the significance of these patterns.
In this manner, PathDL utilizes the micro-level information of patients, whereas ClinicML incorporates macro-level information, such as tumor size. However, the extent of performance improvement when ensembling these two models varied among cohorts, and in some cohorts, it was not statistically significant. Despite these variations, METACANS showed significantly better performance than both PathDL and ClinicML, with p values of <0.0001 and 0.0030, respectively, when aggregating all the data from the external validation cohorts.
Nevertheless, this study has several limitations. First, although its performance was validated using multi-institutional data, this was a retrospective study and has not yet been prospectively validated. Second, the prediction performance varied across institutions. These variations could be attributed to differences in regional factors, institutional practices, imaging devices, and periods of data acquisition. Such disparities highlight the need for more advanced data standardization methods and generalized models. Third, although PathDL demonstrated promising results in terms of pathological translation, its performance was generally lower compared to ClinicML. While METACANS outperforms both PathDL and ClinicML in most validation cohorts, the relatively lower performance of PathDL may have limited its contribution to the overall performance gains when combined with ClinicML. Moreover, several methods are currently under development to predict ALN metastasis preoperatively using data sources other than primary tumor biopsy samples35,36,37. Although axillary staging with primary tumor biopsy samples is innovative and can aid in uncovering the pathological basis, its performance is still not as effective as that of other techniques. With recent shifts in clinical practice away from routine ALND, the low specificity in this study could also pose a significant concern. The results presented demonstrate the potential of METACANS for reducing unnecessary ALNDs in breast cancer patients, as evidenced by its high NPV. However, the model’s relatively low specificity (50.4%) warrants careful consideration. This lower specificity, while not negating the potential benefit of reduced ALNDs in negative cases, highlights the risk of false positives, potentially leading to unnecessary surgical interventions. This limitation underscores the need for a nuanced interpretation of the model’s output and the importance of integrating METACANS into a broader clinical workflow.
Current clinical practice frequently incorporates preoperative ultrasound and intraoperative SLNB to refine patient selection for ALND. While these methods offer valuable information, they are subject to limitations in both sensitivity and specificity, and their effectiveness is dependent on the experience and expertise of the clinicians involved. Preoperative ultrasound, though non-invasive and rapid, can yield false negatives. SLNB, while highly sensitive in detecting metastasis, exhibits lower specificity and adds complexity and time to the surgical procedure.
METACANS offers a unique approach by providing an objective, experience-independent assessment of ALNM risk based on the analysis of primary tumor biopsy WSIs. This approach aims to complement existing clinical methods, mitigating their individual limitations. The model’s high NPV suggests its potential to reduce the number of patients undergoing unnecessary ALNDs. However, the lower specificity necessitates a strategy that incorporates METACANS within a broader clinical decision-making framework, potentially using it in conjunction with preoperative ultrasound or SLNB to improve overall accuracy and minimize the risk of false positives. Future research will focus on improving the model’s specificity through various strategies, including model refinement, data augmentation, and feature selection, to enhance its clinical utility and reliability. Ultimately, the goal is to optimize the balance between sensitivity and specificity to provide a more robust and clinically valuable tool for breast cancer management.
In summary, our study introduced the METACANS model, an approach for predicting ALN metastasis from primary tumor biopsy samples in patients with breast cancer. Although efficient diagnostic methods such as SLNB are being developed to prevent invasive surgeries in patients, the significance of our research lies in diagnosing images from primary tumor biopsy samples and analyzing them in relation to metastasis using pathobiological knowledge. This study aligns with the growing trend toward patient-friendly diagnostic methods, potentially preventing unnecessary ALND and reducing invasive procedures. Although the current performance may not be optimal for immediate clinical application, the insights gained from this research set the stage for further innovation in this field.
Methods
Study design and participant
This retrospective, multicenter diagnostic study was conducted in South Korea using data from six independent breast cancer cohorts. The institutional review boards (IRBs) of the participating institutions waived the requirement for written informed consent: Sinchon Severance Hospital (SS; IRB no. 4-2021-0029), Keimyung University Dongsan Medical Center (KUDMC; IRB no. 2021-08-112), Gangnam Severance Hospital (GS; IRB no. 3-2021-0071), Ewha Womans University Mokdong Hospital (EWUMH; IRB no. 2021-08-013-007), Cha Bundang Medical Center (CBMC; IRB no. 2021-09-021), and Dankook University Hospital (DKUH; IRB no. 2022-03-041). The study was conducted in accordance with the Declaration of Helsinki. An overview of the data collection is shown in Fig. 4, and the details of the patient characteristics are provided in Table 3 and Supplementary Table 1.
a Data flow diagram of patients. In this retrospective study, patients were collected from six independent institutions and were assigned to a training and internal validation cohort, as well as to external validation cohorts A, B, C, D, and E. b Geographical distribution of the multi-institutions in this study. ALNM− axillary lymph node metastasis-negative, ALNM+ axillary lymph node metastasis-positive.
For model training and internal validation, 5921 patients treated between July 2005 and June 2020 at the Sinchon Severance Hospital were initially included. The exclusion criteria were as follows: (1) absence of biopsy specimens (n = 2621), (2) non-invasive areas (n = 664), and (3) insufficient tissue (n = 145). After applying these criteria, 2491 patients remained. They were randomly divided into two groups: 80% for training (n = 1991) and 20% for internal validation (n = 500). The training set included 1457 (73%, ALN metastasis-negative) and 534 (27%, positive) patients. The internal validation set comprised 368 (74%, negative) and 132 (26%, positive) patients. Within the training set (n = 1991), an eight-fold cross-validation strategy was implemented to optimize the model during training. Seven folds (n = 1743) were used as the training folds, while one fold (n = 248) was used as the cross-validation fold. The cross-validation fold served to monitor model training, optimize hyperparameters, and determine cut-off thresholds.
Data from five additional independent cohorts were collected for external validation using the same exclusion criteria. Cohort A comprised 1090 patients treated at the KUDMC between November 2001 and December 2020. Cohort B comprised 486 patients from the GS who were treated between January 2007 and January 2021. Cohort C comprised 246 patients from the EWUMH, treated between January 2005 and June 2010. Cohort D comprised 197 patients from the CBMC who were treated between June 2011 and September 2017. Cohort E comprised 147 patients from the DKUH treated between January 2004 and September 2016. Further information is shown in Supplementary Figs. 6 and 7.
Patch generation
Given the large size of WSIs, patch-level analysis is often employed to mitigate computational demands. Moreover, because of the presence of abundant non-informative areas in WSIs, selecting the relevant tissue areas is necessary for computational efficiency. In this study, we extracted 224 × 224 × 3 red-green-blue (RGB) patches with 10× resolution (each pixel represents ~1.0 × 1.0 μm2) from WSIs. For each patch, we selected only those with a T-value greater than 50, which were considered informative patches, as described in Eqs. 1 and 2. Constant C was empirically set to 8. The notations R, G, and B represent RGB channels.
Additionally, to ensure a more accurate patch selection, we utilized the hue-saturation-value (HSV) domain. We converted the RGB domain image into the HSV domain and selected patches for which the average hue (H) domain value was empirically larger than 70. This process resulted in 2,330,202 patches from the training and internal validation cohorts. For external validation cohorts A–E, 888,294; 392,356; 213,301; 157,845; and 142,287 patches were extracted, respectively.
Stain normalization
Histopathological slide preparation involves a staining process to enhance the contrast and detail. However, this process can introduce variability in the color and intensity of the stain, potentially affecting the performance of computer-aided diagnosis systems. Hence, stain normalization is a crucial preprocessing step to mitigate these inconsistencies and ensure reliable image analysis.
To address variations in WSI scanners, staining methods, and tissue processing across institutions, we applied the Macenko method for stain normalization38,39. This method is a widely adopted technique in digital pathology that reduces the variance in color representation stemming from differences in the staining procedures. This method extracts the color deconvolution matrix from a reference image and then applies this matrix to the target images. This process effectively standardizes the color distribution across all images, thus mitigating the effects of staining variability and facilitating a more accurate and consistent downstream image analysis. The results of the stain normalization in this study are shown in Supplementary Fig. 8.
Deep learning model for feature extraction
In our study, we adopted the CTransPath40 model, implemented in the PyTorch framework, to extract image features from each patch using an NVIDIA RTX A6000 graphics processing unit (GPU) with a batch size of 1000. The CTransPath model combines convolutional neural networks (CNNs) with multi-scale Swin Transformers41, effectively capturing both local and global information. The underlying CNN layers focus on extracting detailed spatial features, whereas the Swin Transformer handles long-range dependencies, making it particularly powerful for histopathological image analysis, where both fine-grained and global contextual understanding are critical. The model was pretrained on datasets from The Cancer Genome Atlas (TCGA) and the Pathology AI Platform (PAIP), including approximately 15 million patches taken from over 30,000 WSIs. TCGA and PAIP together cover multiple organs and a variety of cancers, with over 25 anatomical locations and 32 different cancer subtypes, ensuring a diverse sample range that aids in training universal feature representations suitable for various histopathological images. The integrated design allows CTransPath to act as an effective local-global feature extractor, generating universal feature representations that are highly suitable for histopathological image analysis tasks. We chose CTransPath because of its hybrid architecture, which has demonstrated significant capability in capturing both local and contextual features effectively, making it ideal for complex medical image analysis tasks. Using CTransPath, each patch was transformed into a 768-dimensional vector, which is termed a patch-level representation. Consequently, a WSI with N patches was represented as an N × 768 feature matrix.
Deep learning model for ALN metastasis classification
To aggregate the patch-level representations (N × 768 feature matrix) into a slide-level representation (1 × 768 feature vector), we employed an ABMIL model, which has been applied to various digital pathology image analyzes, including breast cancer detection, cancer subtyping, and survival prediction. ABMIL uses a weighted average of patch representations, with weights determined by an attention mechanism using a neural network. Let \(H={\{h}_{1},\ldots ,{h}_{N}\}\) be a bag of N instances; ABMIL uses weighted averaging of each representation to obtain a bag-level (WSI-level) representation z:
where:
where \({\rm{w}}\in {{\mathbb{R}}}^{L\times 1}\), \({\rm{V}}\in {{\mathbb{R}}}^{L\times M}\), and \({\rm{U}}\in {{\mathbb{R}}}^{L\times M}\) are parameters. \(\odot\) is an element-wise multiplication. The ABMIL method utilizes nonlinearity with a sigmoid activation function. In this study, L and M were set as 768 and 192, respectively. The bag-level representation was then passed to a fully connected layer to obtain the final prediction probability. We refer to this process as the DL model, which uses pathological images and is termed PathDL.
The model was trained using the following key parameters: binary cross-entropy loss function, Adam optimizer42 (β1 = 0.9, β2 = 0.999), weight decay of 0.0005, learning rate of 0.001 with a cosine annealing learning rate scheduler, and batch size of 1. Dropout layers with a probability of 0.10 were added before both the attention gating and the last fully connected layer to improve the robustness of the model.
Machine learning model for ALN metastasis classification
We developed an ML model based on an RF to predict ALN metastasis using clinicopathological characteristics. The RF model integrates multiple decision trees to enhance the precision and stability of predictions. This approach reduces overfitting and improves generalization performance. We selected the RF model due to its ability to handle complex interactions between features, which is crucial given the heterogeneity of clinicopathological data. In this study, we trained the RF model using tumor size, number of cancerous lesions, and age. Tumor size and number of cancerous lesions were determined using ultrasound image or magnetic resonance imaging, while age was derived from clinical reports. To prioritize practical applications for clinicians, our model was trained using only the clinicopathological characteristics obtained before WSI analysis. We refer to this ML-based model as ClinicML.
We used the RandomForestClassifier module from Python’s sklearn.ensemble package to train the RF model. The parameters we employed were n_estimators = 2000, max_depth = 3, min_samples_split = 50, class_weight = “balanced,” and random_state = 42.
To prevent model overfitting and simplify the model, age was categorized into two classes: patients aged 55 years and above, and those aged under 55 years. The cutoff of 55 years was selected based on the average age of 53.7 years in the training set. In cases of multiple cancers, the longest diameter of the largest tumor was used as the tumor size. The number of cancerous lesions was classified as single or multiple. To make our model more convenient for clinicians, we opted not to use any additional information that could be obtained from further analysis of the WSIs.
Ensemble of two ALN metastasis classification models
Given that each prediction model has a different predictive power, simply adding the prediction results of the two models with equal weights could potentially degrade the performance. To effectively integrate the continuous probabilities obtained from both models for the final prediction, we calculated the weights proportional to the predictive power of each individual model. This was achieved by applying a weighted ensemble of probabilities, as outlined in Eq. 5. Here, wPathDL and wClinicML were calculated based on the performance gain, which signifies the actual predictive power of the model, as shown in Eq. 7. As the theoretical AUC was 0.500 when performing a random guess, we defined the actual predictive power, or performance gain, G, as AUC = 0.500 in this study. The AUC value used to calculate the performance gain and weights was based on the AUC value for a single validation fold from the 8-fold cross-validation with the training set.
where:
where:
In this study, the AUCPathDL and GPathDL were 0.639 and 0.139, respectively. Similarly, the AUCClinicML and GClinicML were 0.729 and 0.229, respectively. At this point, wPathDL and wClinicML were obtained as 0.378 and 0.622, respectively, according to Eq. 6 (see “Materials and Methods”). Subsequently, we performed min-max normalization using a minimum probability of 0.235 and a maximum probability of 0.771 from the validation fold to adjust the overall probability between 0 and 1. This operation, which intuitively adjusts the range of values, does not affect the performance of the model. We then clipped the final calculated probability to the range [0, 1] for all cohorts. These weights and processes were used to calculate the final ALN metastasis prediction probabilities for internal and external validation cohorts.
METACANS
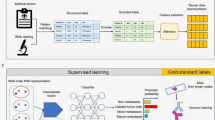
In this study, we ensembled the probabilities from the DL-based model (PathDL) and ML-based model (ClinicML) using a weighted summation, where the weights were determined based on the performance of each model to optimize the final prediction by assigning more importance to the model with higher performance. The result was the final probability of predicting ALN metastasis. In this paper, this ensemble model is referred to as METACANS, which encapsulates the concept of METAstasis CANcer Scope. Overall process of the METACANS is shown in Fig. 5.
PathDL analyzes whole slide images from primary tumor biopsies. ClinicML focuses on clinicopathological data. METACANS combines predictions from both models through a weighted ensemble.
Statistical analysis
Statistical analysis was conducted using the R software (R Core Team, 2020). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. URL: https://www.R-project.org/). The DeLong’s method43 was used to compare the two AUC values. Statistical significance was set at P < 0.05. We set the cut-off threshold for model decisions using maximum Youden’s index on a cross-validation fold within the eight-fold cross-validation of the training set.
Data availability
The data generated in this study are not publicly available because of privacy and security concerns, but are available upon reasonable request from the corresponding author.
Code availability
The underlying code for this study can be accessed via the following link: https://github.com/DoohyunPark/METACANS.
Change history
19 June 2025
In the Acknowledgements section of this article the grant number relating to Korea Health Industry Development Institute funded by the Korea government (Ministry of Health and Welfare) was incorrectly given as RS-2021-KH113638 and should have been RS-2021-KH113638.The original article has been corrected.
References
Huang, J. et al. Global incidence and mortality of breast cancer: a trend analysis. Aging 13, 5748–5803 (2021).
Rao, R., Euhus, D., Mayo, H. G. & Balch, C. Axillary node interventions in breast cancer. JAMA 310, 1385 (2013).
Nielsen Moody, A. et al. Preoperative sentinel lymph node identification, biopsy and localisation using contrast enhanced ultrasound (CEUS) in patients with breast cancer: a systematic review and meta-analysis. Clin. Radiol. 72, 959–971 (2017).
Krag, D. N. et al. Technical outcomes of sentinel-lymph-node resection and conventional axillary-lymph-node dissection in patients with clinically node-negative breast cancer: results from the NSABP B-32 randomised phase III trial. Lancet Oncol. 8, 881–888 (2007).
Rahman, M. & Mohammed, S. Breast cancer metastasis and the lymphatic system. Oncol. Lett. 10, 1233–1239 (2015).
Langer, I. et al. Morbidity of Sentinel Lymph Node Biopsy (SLN) alone versus SLN and completion axillary lymph node dissection after breast cancer surgery. Ann. Surg. 245, 452–461 (2007).
Kootstra, J. J. et al. A longitudinal comparison of arm morbidity in stage I–II breast cancer patients treated with sentinel lymph node biopsy, sentinel lymph node biopsy followed by completion lymph node dissection, or axillary lymph node dissection. Ann. Surg. Oncol. 17, 2384–2394 (2010).
van de Vrande, S., Meijer, J., Rijnders, A. & Klinkenbijl, J. H. G. The value of intraoperative frozen section examination of sentinel lymph nodes in breast cancer. Eur. J. Surg. Oncol. 35, 276–280 (2009).
Xu, F. et al. Predicting axillary lymph node metastasis in early breast cancer using deep learning on primary tumor biopsy slides. Front. Oncol. 11, 759007 (2021).
Das, A., Nair, M. S. & Peter, S. D. Computer-aided histopathological image analysis techniques for automated nuclear atypia scoring of breast cancer: a review. J. Digit. Imaging 33, 1091–1121 (2020).
Romo-Bucheli, D., Janowczyk, A., Gilmore, H., Romero, E. & Madabhushi, A. Automated tubule nuclei quantification and correlation with oncotype DX risk categories in ER+ breast cancer whole slide images. Sci. Rep. 6, 32706 (2016).
Veta, M. et al. Assessment of algorithms for mitosis detection in breast cancer histopathology images. Med. Image Anal. 20, 237–248 (2015).
Ding, Y. et al. Multi-center study on predicting breast cancer lymph node status from core needle biopsy specimens using multi-modal and multi-instance deep learning. NPJ Breast Cancer 9, 58 (2023).
Marmé, F. et al. Deep learning to predict breast cancer sentinel lymph node status on INSEMA histological images. Eur. J. Cancer 195, 113390 (2023).
Ehteshami Bejnordi, B. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318, 2199 (2017).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Maron, O. & Lozano-Pérez, T. A framework for multiple-instance learning. Adv. Neural Inf. Process. Syst. 10, 570–576 (1997).
Ilse, M., Tomczak, J. & Welling, M. Attention-based deep multiple instance learning. Proc. Int. Conf. Mach. Learn. PMLR 80, 2127–2136 (2018).
Breiman, L. Random forests. Mach. Learn 45, 5–32 (2001).
Amin, M. B. AJCC Cancer Staging Manual https://doi.org/10.1007/978-3-319-40618-3. (Springer International Publishing, 2017).
Elston, C. W. & Ellis, I. O. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long‐term follow‐up. Histopathology 19, 403–410 (1991).
Gradishar, W. J. et al. Breast cancer, version 3.2024, NCCN clinical practice guidelines in oncology. J. Natl. Compr. Cancer Netw. 22, 331–357 (2024).
Davey, M. G., Hynes, S. O., Kerin, M. J., Miller, N. & Lowery, A. J. Ki-67 as a prognostic biomarker in invasive breast cancer. Cancers 13, 4455 (2021).
Lyman, G. H. et al. Sentinel lymph node biopsy for patients with early-stage breast cancer: American Society of Clinical Oncology clinical practice guideline update. J. Clin. Oncol. 35, 561–564 (2017).
Hindié, E. et al. The sentinel node procedure in breast cancer: nuclear medicine as the starting point. J. Nucl. Med. 52, 405–414 (2011).
Manca, G. et al. Sentinel lymph node biopsy in breast cancer. Clin. Nucl. Med. 41, 126–133 (2016).
Chang, J. M., Leung, J. W. T., Moy, L., Ha, S. M. & Moon, W. K. Axillary nodal evaluation in breast cancer: state of the art. Radiology 295, 500–515 (2020).
Zwaans, B. M. M. & Bielenberg, D. R. Potential therapeutic strategies for lymphatic metastasis. Microvasc. Res. 74, 145–158 (2007).
Yoo, S. H. et al. A histomorphologic predictive model for axillary lymph node metastasis in preoperative breast cancer core needle biopsy according to intrinsic subtypes. Hum. Pathol. 46, 246–254 (2015).
Tavassoli, F. A. Pathology and Genetics. Tumours of the Breast and Female Genital Organs. (World Health Organization Classification of Tumours, 2003).
Acs, G., Paragh, G., Chuang, S.-T., Laronga, C. & Zhang, P. J. The presence of micropapillary features and retraction artifact in core needle biopsy material predicts lymph node metastasis in breast carcinoma. Am. J. Surg. Pathol. 33, 202–210 (2009).
Pettinato, G., Manivel, C. J., Panico, L., Sparano, L. & Petrella, G. Invasive micropapillary carcinoma of the breast. Am. J. Clin. Pathol. 121, 857–866 (2004).
Makki, J. Diversity of breast carcinoma: histological subtypes and clinical relevance. Clin. Med. Insights Pathol. 8, CPath.S31563 (2015).
Verras, G.-I. et al. Micropapillary breast carcinoma: from molecular pathogenesis to prognosis. Breast Cancer. Targets Ther. 14, 41–61 (2022).
Dihge, L. et al. Prediction of lymph node metastasis in breast cancer by gene expression and clinicopathological models: development and validation within a population-based cohort. Clin. Cancer Res. 25, 6368–6381 (2019).
Zheng, X. et al. Deep learning radiomics can predict axillary lymph node status in early-stage breast cancer. Nat. Commun. 11, 1236 (2020).
Cools‐Lartigue, J. & Meterissian, S. Accuracy of axillary ultrasound in the diagnosis of nodal metastasis in invasive breast cancer: a review. World J. Surg. 36, 46–54 (2012).
Macenko, M. et al. A method for normalizing histology slides for quantitative analysis. In Proc. IEEE International Symposium on Biomedical Imaging: From Nano to Macro 1107–1110 https://doi.org/10.1109/ISBI.2009.5193250 (IEEE, 2009).
Barbano, C. A. et al. Unitopatho, a labeled histopathological dataset for colorectal polyps classification and adenoma dysplasia grading. In Proc. IEEE International Conference on Image Processing (ICIP) 76–80 https://doi.org/10.1109/ICIP42928.2021.9506198 (IEEE, 2021).
Wang, X. et al. Transformer-based unsupervised contrastive learning for histopathological image classification. Med. Image Anal. 81, 102559 (2022).
Liu, Z. et al. Swin Transformer: Hierarchical Vision Transformer Using Shifted Windows. In Proc. IEEE/CVF International Conference on Computer Vision (ICCV) 10012–10022 (IEEE, 2021).
Kingma, D. P. & Ba, J. L. Adam: A Method for Stochastic Optimization. In Proc. 3rd International Conference on Learning Representations (ICLR), San Diego. https://doi.org/10.48550/arXiv.1412.6980 (2015).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44, 837 (1988).
Acknowledgements
This study was supported by a grant from the Korea Health Industry Development Institute funded by the Korea government (Ministry of Health and Welfare) (RS-2021-KH113638); the Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Science and ICT (2021R1C1C2008773, 2022R1A2C2008983); Artificial Intelligence Graduate School Program, Yonsei University (RS-2020-II201361), the KIST Institutional Program (Project No. 2E33801, 2E33800); Yonsei Signature Research Cluster Program of 2024 (2024-22-0161). The funder played no role in study design, data collection, analysis, and interpretation of data, or the writing of this manuscript.
Author information
Authors and Affiliations
Contributions
D.P.: methodology, validation, investigation, formal analysis, writing—original draft, writing—review and editing, and visualization. Y-M.L.: data curation, validation, formal analysis, writing—original draft, writing—review and editing, visualization, and resources. T.E.: validation, formal analysis, and writing—review and editing. H.J.A., H.K., E.P., and Y.J.C.: data curation, validation, writing—review and editing, visualization, and resources. Heejung Park, D.K., S.Y.K., H-R.J., S.-J.S., Hyunjin Park, Y.L., S.P., J.M.K., and S-E.C.: data curation, resources, and writing—review and editing. N.H.C.: methodology, validation, investigation, formal analysis, data curation, writing—original draft, writing—review and editing, resources, supervision, project administration, and funding acquisition. D.H.: methodology, validation, investigation, formal analysis, writing—original draft, writing—review and editing, resources, supervision, project administration, and funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
Doohyun Park was employed by VUNO Inc. after the completion of the project. All other authors declare no financial or non-financial competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, D., Lee, YM., Eo, T. et al. Multimodal AI model for preoperative prediction of axillary lymph node metastasis in breast cancer using whole slide images. npj Precis. Onc. 9, 131 (2025). https://doi.org/10.1038/s41698-025-00914-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-00914-9
This article is cited by
-
Customized transformer for lymph node metastasis prediction from lung adenocarcinoma histology in a multicentric study
npj Precision Oncology (2025)