Abstract
Pediatric low-grade gliomas (pLGGs) are the most common brain tumors in children with varying degrees of infiltration. Despite having a positive prognosis, if the standard treatment, gross total resection, is impossible due to tumor location or diffuseness, outcomes worsen. Development of targeted therapeutics for diverse subtypes of pLGGs is limited by a lack of genetic models. We generated five fly pLGG models using patient-derived fusion genes to investigate molecular subtype-specific pathology, and found glial overexpression of QKI::RAF1, associated with pilocytic astrocytomas and glioneuronal tumors, induced aberrant glial migration and infiltration. Both repulsive guidance signaling and GPR180/CG9304 mediated glial infiltration, which was suppressed by glial overexpression of Robo2 or PlexA/B, or knockdown of GPR180/CG9304. ROBO2 and GPR180/CG9304 were down and upregulated, respectively, in flies and patients with RAF fusions. Our study provides mechanistic insights into pLGG tumorigenesis and suggests targeting repulsive guidance signaling and GPR180/CG9304 as potential therapeutics for pLGG subtypes.
Similar content being viewed by others
Introduction
Pediatric low-grade gliomas (pLGGs) are the most diagnosed brain tumors in children1,2. While the prognosis of pLGGs is favorable, progression-free survival (PFS) is highly dependent on the feasibility of surgical gross total resection (GTR): the 10-year PFS exceeds 85% in patients where GTR was possible, but falls below 50% in patients where GTR was impossible3 due to the location and the diffuse/infiltrative nature of the tumor1,4. While adjuvant therapies such as radiation and/or chemotherapy have been used in conjunction with surgical resection to treat pLGGs, both can result in significant side effects, leading to poor functional outcomes5. These challenges highlight the importance of bettering our understanding of pLGGs so we can identify new therapeutic strategies and targets to improve outcomes and minimize side effects, particularly in patients where GTR is impossible.
As the paradigm in pLGG treatment has shifted towards the development of personalized medicine, it is critical to understand the diverse tumor biology of pLGGs to identify new targeted molecular therapeutics6. Recognizing the need to more specifically diagnose and treat patients, in 2021, the World Health Organization (WHO) recategorized central nervous system (CNS) tumors, focusing on identifying molecular and histological tumor characteristics7. Pediatric-type diffuse LGGs became a classifier of interest due to their distinct molecular subcategories and diffuse pattern. The diffuse pattern makes GTR more challenging, making it a priority to understand the molecular pathology to develop new and targeted treatment options. However, progress has been limited by a lack of accurate models8.
To fill this knowledge gap, we aimed to generate pLGG models that would inform a better understanding of the molecular underpinnings of tumorigenic mutations, focusing on those that best model tumors with diffuse characteristics. To that end, we created five Drosophila melanogaster (fruit fly) pLGG models by overexpressing patient-derived fusion genes in glia. We chose to model gene fusions due to their frequent role as drivers of gliomagenesis and glioma progression6. Drosophila melanogaster was chosen as the model system due to its genetic and molecular similarity to humans as well as its genetic accessibility, which enables fidelity and throughput. In addition, we specifically wanted an in vivo system to enable visualization of an infiltrative phenotype. This approach builds on our previous success in using fruit flies as a model system to study gliomas9,10,11.
When characterizing our models, we found glial overexpression of the fusion gene, QKI::RAF1, resulted in significant glial cell migration defects in the fly nervous system, mimicking the diffuse/infiltrative tumor pattern that results in challenging GTR in human patients. QKI::RAF1 is a known driver of tumorigenesis associated with pilocytic astrocytomas and glioneuronal tumors. While pilocytic astrocytomas are largely characterized as non-diffuse, there is emerging literature that some have infiltrative characteristics12. Given the small number of QKI::RAF1 cases, it is inconclusive whether they are diffuse or not. In addition, QKI::RAF1 has been shown to act via the mitogen-activated protein kinase (MAPK) pathway, which is known to be altered in pLGGs13. QKI::RAF1 fusions are resistant to first and second-generation RAF inhibitors, highlighting the importance of developing new treatments14. QKI::RAF1 is one of many RAF fusions, a group characterized by having either RAF1 or BRAF as the C-terminal fusion partner. Studying QKI::RAF1-mediated tumorigenesis thus presents an entry point to understand the pathways underlying various subtypes of pLGGs, especially those with RAF1/BRAF fusions. Given that glial QKI::RAF1 overexpression best replicated the diffuse phenotype we aimed to model, we primarily focused on characterizing and understanding the molecular mechanisms that underlie its infiltrative properties, not only to better understand this fusion, but also to elucidate how diffuse pLGGs acquire their invasive properties. We have identified two potential therapeutic avenues: repulsive guidance signaling and a G-protein coupled receptor (GPCR)—GPR180/CG9304 for patients with RAF1/BRAF fusions. We also demonstrate the translatable potential of our pipeline and findings as we move fluidly between fly tumor models, an in vitro mouse system, and human patient tissues/imaging to generate models, identify translatable phenotypes, determine common pathways, and ultimately identify therapeutic targets.
Results
Drosophila melanogaster genetic models overexpressing pLGG-derived gene fusions show glial aberrations
As the most common cause of pLGGs is gene fusions or single gene rearrangements6, we established five distinct fly models via glial overexpression of patient-derived fusion genes: QKI::RAF1, MKRN1::BRAF, FGFR1::TACC1, QKI::NTRK2, and KIAA1549::BRAF to better understand the pathology of each fusion. A fusion gene is defined as two independent genes merging, being transcribed, and translated as a new, hybrid unit. Typically, this fusion results in abnormal activation of downstream signaling pathways, and in the case of pLGGs, often activation of MAPK and mTOR13,15,16. QKI::RAF1 is a RAF1 kinase fusion with the QKI RNA-binding protein associated with pilocytic astrocytomas14 and subgroups of glioneuronal tumors (GNTs)17. MKRN1::BRAF is a BRAF fusion with the E3 ubiquitin ligase MKRN1 and is present in patients with LGGs, thyroid gland papillary carcinoma, and colon adenocarcinoma13,14,18,19. FGFR1::TACC1 is a fusion of fibroblast growth factor receptor (FGFR1) with transforming acidic coiled-coil TACC1 associated with LGGs15. QKI::NTRK2 is a fusion with the QKI RNA-binding protein and neurotrophic tyrosine kinase receptor 2 (NTRK2) associated with LGGs15. KIAA1549::BRAF is the most frequent molecular alteration in pilocytic astrocytomas, resulting from a fusion of the active BRAF kinase domain with the N-terminus of the KIAA1549 protein20. Thus, we modeled the most common fusion, KIAA1549::BRAF, present in 5–10% diffuse gliomas as well as 70–80% pilocytic astrocytomas, and the less common QKI::RAF1, MKRN1::BRAF, and FGFR1::TACC1 fusions with the incidence below 5% across diffuse LGGs, pilocytic astrocytomas, and other mixed glioneuronal tumors21.
To study the pathology of these fusion genes, we conducted in vivo imaging of the ventral nerve cord (VNC) of the fly CNS in third instar larvae (Fig. 1A), a region that is analogous to the spinal cord in humans and subsequently conducted brain and VNC dissections (Fig. 1B). We used the pan-glial repo-Gal4 driver22 to express an mRFP reporter as well as to overexpress each fusion gene and the positive control—the established fly glioblastoma model for constitutively active Drosophila EGFR and PI3K (dEGFRACT; dPI3KACT)23. We found that overexpression of dEGFRACT; dPI3KACT, QKI::NTRK2, and FGFR1::TACC1 resulted in elongation of the VNC, with the two fusion genes having a more pronounced phenotype (Fig. 1A, E). Moreover, we found a significant increase in brain lobe glial volume in both dEGFRACT; dPI3KACT and MKRN::BRAF overexpression, while QKI::NTRK2 showed a reduction (Fig. 1B, F). Next, we conducted immunostaining of brain and VNC samples to determine the number of glia (repo) and the number of mitotic glia (pHistone 3 (pH3)). We quantified the number of repo+; mRFP+ glia and found there was a significant increase in the dEGFRACT; dPI3KACT and KIAA1549::BRAF models and a significant decrease in the QKI::NTRK2 model compared to the wildtype (WT) control (Fig. 1C, G). We quantified the number of pH3+; mRFP+ mitotic glia and found there was a significant increase in MKRN::BRAF and KIAA1549::BRAF (Fig. 1D, D’, H), indicating increased glial proliferation. In addition, we validated KIAA1549::BRAF protein expression and localization by visualizing the N-terminal EGFP tag (Fig. S1A).
a In vivo images depict the VNC of third instar fly larvae, with repo-Gal4 driving mRFP expression in glia. The VNC of the WT is outlined with a dashed line. Scale bar = 100 µm. b Confocal images of dissected third instar larval brain lobes and VNC expressing mRFP in glia (red), staining for repo labeling glia (green), and staining for mitotic cells pH3 (blue). c Confocal images of third instar brain lobes with mRFP and repo. Confocal images of third instar brain lobes with mRFP and pH3 (d) or pH3 alone (d’). Scale bar = 100 µm. e–h Quantifications of VNC length, glial tissue volume, number of total glial cells, and number of mitotic glial cells. For WT, dEGFRACT; dPI3KACT, QKI::RAF1, QKI::NTRK2, MKRN1::BRAF, FGFR1::TACC1, KIAA1549::BRAF, N = 6, 6, 11, 9, 13, 12, 7 for VNC length, N = 23, 17, 21, 20, 38, 17, 17 for glial tissue volume, N = 14, 5, 6, 17, 28, 13, 11 for number of total glial cells, N = 5, 6, 9, 7, 23, 13, 7 for number of mitotic glial cells. All data are means ± SEM. One-way ANOVA followed by Dunnett’s test, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
QKI::RAF1 induces aberrant glial cell migration in flies
When conducting in vivo imaging of QKI::RAF1 glial overexpressing larvae, we noticed unusual glial migration in an area adjacent to the CNS known as leg imaginal discs, which are epithelial structures in the larvae (Fig. 2A). This aberrant glial migration was of particular interest as it is a good model of pediatric-type diffuse LGGs, specifically their diffuse/infiltrative phenotype. Therefore, we focused on QKI::RAF1 phenotypic characterization, molecular mechanisms, and connections to human patients. We also noticed a general brain malformation in the QKI::RAF1 model, but decided to focus on glial migration in the imaginal discs, which are more accessible and tractable.
a Schematic depicting typical glial cell migration patterns in the leg discs of third instar WT and QKI::RAF1 overexpression larvae generated using Biorender.com. In vivo imaging of glia in WT and QKI::RAF1 larvae leg imaginal discs, indicated by arrows. b Confocal images of dissected third instar larval leg discs of larvae expressing mRFP in glia (red) with a DAPI counterstain (blue). c Quantification of the length of migrating glia from the base of the leg disc to the edge of the disc. For WT, QKI::RAF1, QKI.E48G::RAF1, QKI::RAF1.R401H, N = 35, 30, 10, 18. d Quantification of glia bundle area in the “normal pathway” as measured by the area of the disc populated with glial cells normalized to the area of the entire disc. For WT, QKI::RAF1, QKI.E48G::RAF1, QKI::RAF1.R401H, N = 17, 15, 5, 9. e Evaluation of fly survival to eclosion with glial overexpression of QKI::RAF1 and the two mutants. N = 8. All data are means ± SEM. One-way ANOVA followed by Dunnett’s test, *P < 0.05, ****P < 0.0001. f Confocal images of dissected third instar leg discs with repo (green), pH3 (blue, lower left panels), and HRP (blue, lower right panels) staining. Yellow arrows indicate migrating glia. Scale bar = 100 μm.
To observe the migratory trajectory of glial cells in and around the leg discs, we performed leg disc dissections in third instar larvae and examined glial localization. In WT larvae, glial cells run through the middle of the leg disc in a tight nerve bundle (Fig. 2A, B), which we termed the “normal pathway” and do not run along the leg imaginal discs. In contrast, glial overexpression of the fusion gene QKI::RAF1 resulted in migratory defects characterized by glial cells extending onto the edge of the imaginal disc (Fig. 2A, B, arrows). This invasion-like behavior was consistently observed in our QKI::RAF1 model, but was rarely seen in other fusion gene models, except in some MKRN1::BRAF samples (Fig. S1B). It is also worth noting that while the invasive behavior was present in the MKRN1::BRAF samples, the extent of infiltration was not as pronounced as what was observed in the QKI::RAF1 model. Moreover, the area of the glial bundle in the region containing the “normal pathway” was expanded with QKI::RAF1 overexpression (Fig. 2B, D), which might result from increased glial influx, a region-specific increase of glial proliferation, or defasciculation—the separation of glial processes from the bundle. To distinguish between those possibilities, we performed immunostaining of the dissected disks with repo and pH3 antibodies. We found QKI::RAF1 led to glial cells along the aberrantly migrated glial processes around the leg disc edges (Fig. 2F arrows), which are absent in WT. There was also an increase of glial cells in the “normal pathway” without a drastic increase of local glial cell proliferation (Fig. 2F). Worth noting, the axon bundle, labeled by HRP staining, appeared to be normally wrapped by glial processes in the “normal pathway” in both WT and QKI::RAF1 (Fig. 2F).
To examine the contribution of each QKI::RAF1 fusion partner, we generated two QKI::RAF1 point mutation fly lines: QKI.E48G::RAF1 and QKI::RAF1.R401H that have previously been studied in vitro14. In vitro, QKI.E48G::RAF1 was shown to disrupt the QKI-mediated dimerization of QKI::RAF1, resulting in decreased signaling and tumorigenic potential, and QKI::RAF1.R401H was shown to disrupt the RAF1-mediated dimerization, resulting in decreased downstream RAF1 kinase signaling14. In flies, glial overexpression of both mutations resulted in a decrease in glial migration length as measured by the distance glial cells extended from the base to the edge of the leg disc (Fig. 2B, C). To add, the QKI.E48G::RAF1 flies had significantly reduced glial leg disc area as compared to QKI::RAF1 flies (Fig. 2D). We found glial overexpression of both QKI::RAF1 and QKI.E48G::RAF1 is lethal during the pupal stage, but glial QKI::RAF1.R401H overexpression improved overall survival, as some flies survived to adulthood (Fig. 2E). The phenotypic difference observed between QKI.E48G::RAF1 and QKI::RAF1.R401H mutants highlights the distinction in the tumorigenic role of the N-terminal and C-terminal fusion partners, suggesting that the R401 RAF1 site may play a more critical role in mediating QKI::RAF1’s oncogenic function.
Robo2 and PlexA/B rescue glial migration defects in the QKI::RAF1 pLGG model
Abnormal glial cell migration is associated with dysregulation of repulsive guidance signaling24, which includes the roundabout (Robo) and plexin (Plex) transmembrane receptors and their respective ligands, Slit and Semaphorin. Initially discovered as regulators of commissural axon crossing, the Robo/Slit and Plex/Semaphorin pathways help guide glial cells to their final destinations24. Specifically, glial expression of Robo2 and PlexA/B is required to prevent breakaway from the nerve at the larval leg imaginal disc24. While repulsive guidance signaling has also been associated with tumor cell proliferation and invasion in glioblastomas and pancreatic cancer25,26,27, its precise function in tumor development is yet to be determined. We hypothesized that dysregulation of repulsive guidance results in abnormal glial migration in our QKI::RAF1 model.
To investigate if Robo2 or PlexA/B expression impacts QKI::RAF1-induced migration, we overexpressed Robo2 or PlexA/B with QKI::RAF1 in glia. We found glial co-overexpression partially ameliorated the aberrant migration phenotype (Fig. 3A, B). PlexA overexpression also resulted in reduced glial leg disc area (Fig. 3A, C). Additionally, we found that simultaneous glial overexpression of Robo2, PlexB, and QKI::RAF1 did not result in a further reduction in the migration phenotype (Fig. 3A, B), suggesting a partially overlapping function of Robo2 and PlexA/B.
a Confocal images of dissected third instar larval leg discs with mRFP in glia (red) and DAPI (blue). The orange dashed lines outline the glial bundle area. The yellow arrows indicate the furthest point the glia reach; if glia enter and migrate along the leg disc, this is the “migratory path.” b Quantification of the distance traveled by migrating glia from the base of the leg disc to the edge of the disc. For WT, QKI::RAF1, QKI::RAF1+CD4tdTom, QKI::RAF1+robo2, QKI::RAF1+plexinA, QKI::RAF1+plexinB, QKI::RAF1+robo2+plexinB, N = 34, 12, 44, 51, 56, 73, 55. c Quantification of glial bundle area, as measured by the area of the disc populated with glial cells, normalized to the area of the entire disc. For WT, QKI::RAF1, QKI::RAF1+CD4tdTom, QKI::RAF1+robo2, QKI::RAF1+plexinA, QKI::RAF1+plexinB, QKI::RAF1+robo2+plexinB, N = 17, 6, 22, 26, 28, 37, 28. d Confocal images of dissected third instar larval leg discs with mRFP in glia (red), robo2HA (green), and DAPI (blue). Boxed areas are shown to the right of the images. The red outline indicates the normal path. The blue outlined region depicts the edge of the WT leg disc, while the orange outlined region depicts the edge of a QKI::RAF1 leg disc, the location of the “migrating path.” e Quantification of robo2HA expression in glial cells following the normal and migrating pathway, normalized to its expression in regions in the leg disc without glia. For WT, QKI::RAF1 normal path, and QKI::RAF1 migrating path, N = 11, 14, 8. f Confocal images of dissected third instar larval leg discs with mRFP in glia (red), PlexA-myc (green), and DAPI (blue) in WT and QKI::RAF1. g Quantification of PlexA-myc expression in glial cells following the normal and migratory pathway, normalized to its expression in regions in the leg disc without glia. For WT, QKI::RAF1 normal path, and QKI::RAF1 migrating path, N = 8, 12, 12. All data are means ± SEM. One-way ANOVA followed by Dunnett’s test, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Scale bar = 50 µm.
Robo2 is downregulated in aberrantly migrating glial cells in QKI::RAF1 pLGG model
To visualize glial Robo2 expression in our QKI::RAF1 model, we used a robo2HA knock-in28, expressed mRFP in glia, and performed immunostaining. In WT larvae, Robo2 was expressed throughout the leg disc and in glia of the normal pathway (Fig. 3D, E). In the QKI::RAF1 larvae, Robo2 was also expressed throughout the leg disc and at the same level as WT in the glial cells of the normal pathway (Fig. 3D, E, pink dashed outline). However, Robo2 expression in the glia migrating along the leg disc was significantly lower than both the WT and QKI::RAF1 expression in the normal pathway (Fig. 3D, E). Specifically, in QKI::RAF1 samples, glial cells migrated along the edge of the disc, showing a low level of robo2 expression (Fig. 3D orange dashed outline). In WT, no glial cells are present in the same region (Fig. 3D blue dashed outline). This result suggests that the loss of Robo2 induces glial cell infiltration.
To visualize glial PlexA expression in our QKI::RAF1 model, we used a Myc-tagged PlexA BAC transgenic strain29. In both WT and QKI::RAF1 larvae, PlexA was expressed throughout the leg disc and in the glia of the normal pathway at the same level (Fig. 3F, G). While not statistically significant, PlexA expression in the main glial bundle in QKI::RAF1 larvae appeared to be downregulated as opposed to WT. In QKI::RAF1 larvae, the PlexA glial expression level was similar between the migrating glia and the glia of the normal pathway (Fig. 3F, G, orange dashed outline).
QKI::RAF1 overexpressing mouse astrocytes and pLGGs with RAF gene fusions show reduced expression of Robo2
To determine if repulsive guidance signaling is relevant across species and types of pLGGs, we performed gene expression profiling of primary mouse astrocytes (PMAs) overexpressing QKI::RAF114 and looked for differential mRNA expression of axon guidance receptors. Robo2, Plxna1, Plxna2, and Plxnb3 were significantly downregulated in the QKI::RAF1 PMAs (Fig. 4A). Other axon guidance receptors were unaffected, indicating QKI::RAF1 overexpression specifically dysregulates Robo2 and Plexin (Fig. 4A).
a Relative mRNA levels of Tp53-null primary mouse astrocytes (PMAs) overexpressing QKI::RAF1. N = 3–5 biological replicates. b Bar plots showing mean values of ROBO2 expression values (TPM) derived from RNA sequencing of low-grade gliomas within the pediatric brain tumor atlas. LGGs are classified based on cases with no gene fusions (N = 125) and those with the presence of BRAF/RAF1 gene fusions (N = 148), QKI gene fusions (N = 8), RAF1 gene fusions (N = 5). c Volcano plot of DEGs in the QKI::RAF1 pLGG fly model comparing tumor and control CNS tissues. d Volcano plot of DEGs in OLGGs comparing BRAF/RAF1 fusion versus no-fusion samples. e Heatmap of the shared DEGs between fly and human samples. f Pathway enrichment analyses of fly, human and shared DEGs. g Venn diagrams showing the shared genes between fly and human samples. All data are means ± SEM. The data were analyzed by the Kruskal–Wallis test followed by Dunn’s multiple comparisons test, where the control was compared to the QKI::RAF1 condition for each gene (a). The data were analyzed by one-way ANOVA followed by Dunnett’s multiple comparisons test (b). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
We examined ROBO2 expression in human pLGG patient tumors and found ROBO2 was downregulated in tumors containing either RAF1 or BRAF fusions compared to tumors containing no fusions or QKI fusions without RAF1 as its fusion partner (Fig. 4B). We included both RAF1 and BRAF fusions due to limited patient samples for RAF1 fusions (Table S1), and the presence of some glial infiltration in the MKRN::BRAF fly model. The downregulation of ROBO2 in RAF1/BRAF tumors indicates that the C-terminal fusion partner (RAF1/BRAF) plays an essential role in the dysregulation of the repulsive guidance signaling pathway in pLGGs. To note, ROBO2 expression is unchanged in pLGGs resulting from the BRAF-V600E mutation, highlighting the diversity of molecular profiles induced by different mechanisms and underscoring the importance of studying each mutation and fusion independently.
Integrative transcriptomic profiling and a candidate genetic screen reveal GPR180/CG9304 as a regulator of glioma infiltration
To better understand the molecular pathways in pLGG fusions and identify possible therapeutic targets, we looked for overlapping differentially expressed genes (DEGs) in our fly QKI::RAF1 model and human patients (Table S2). We performed bulk mRNA transcriptomic profiling of fly CNS tissue and compared it to the pLGG expression dataset from the Pediatric Brain Tumor Atlas (PBTA). We found 635 DEGs in flies with glial QKI::RAF1 overexpression vs. WT (Fig. 4C) and 4531 DEGs in human patients with BRAF/RAF1 fusions vs. no BRAF/RAF1 fusions (Fig. 4D). We next intersected the fly and the human DEGs (Fig. 4E) and found 31 and 56 overlapping upregulated or downregulated genes, respectively (Fig. 4G). We found both fly and human DEG enrichment in pathways related to neurexins and neuroligins, as well as protein-protein interactions at synapses (Fig. 4F). This may be related to glioma neuron signaling during tumorigenesis30,31,32.
To determine which DEGs played a role in the migration phenotype in the fly tumor model, we performed a candidate-based screen. Ten genes were selected based on fold change, homology, and functional relevance. We used our QKI::RAF1 model to perform glial knockdown or overexpression of upregulated or downregulated genes, respectively. We next performed in vivo imaging to assess the glial phenotypes (Fig. 5A, B) and quantified the length of aberrant migration (Fig. 5C). We found knockdown of GPR180/CG9304 via two independent RNAi lines significantly reduced migration length (Fig. 5B, C). To ensure our findings were the result of a true reduction in migration length and not due to the leg disc being obstructed during in vivo imaging, we repeated our data analysis and excluded all data points with zero migration (Fig. 5D). We found GPR180/CG9304 knockdown still significantly reduced migration length of the two RNAi lines (Fig. 5D), validating that our imaging method does not masking the phenotype. To further characterize the effects of GPR180/CG9304 on glial invasion, we dissected the imaginal leg discs from QKI::RAF1 larvae with GPR180/CG9304 glial knockdown and analyzed migration (Fig. 6A). Consistent with in vivo imaging, there was reduced aberrant glial migration in dissected QKI::RAF1 larvae following GPR180/CG8304 knockdown (Fig. 6C). However, GPR180/CG9304 glial knockdown did not reduce the glial bundle area, suggesting glial GPR180/CG9304 has a specific role in regulating glial infiltration along the disc (Fig. 6D). As a control, glial knockdown of CG9304 in the WT background did not cause an obvious defect in glial migration, proliferation, or axon pathfinding (Figs. 6B–D and S3). To confirm the phenotypes associated with CG9304 knockdown, we employed an insertional mutant of CG9304–CG9304MI09328. The insertion in the 3′UTR disrupts CG9304 splicing and reduces the overall transcript levels (Fig. S2) and is thus at least a loss-of-function hypomorphic allele. We found that the leg disc glial infiltration induced by QKI::RAF1 expression was suppressed by the CG9304MI09328 mutation as well as a modest reduction in the glial bundle area (Fig. 6B–D), the latter may be caused by CG9304 function in cell types other than glia. However, no obvious defects in glial migration, proliferation, or axon pathfinding were detected in the mutants (Figs. 6B–D and S3). These data indicate a pro-infiltration function of GPR180/CG9304 in the context of QKI::RAF1 fusion.
a Schematic illustrating an in vivo screen of QKI::RAF1 flies to identify tumor-suppressing genes and oncogenes generated using Biorender.com. b In vivo images of the VNC and leg discs in third-instar larvae with mRFP in glia (red). The white outlines indicate the area of the leg discs, and the yellow arrows indicate the furthest point glia reach. If glia enter and migrate along the leg disc, this is the “migratory path.” Scale bar = 100 µm. c, d Quantification of glial migration in QKI::RAF1 flies with either an overexpression or knockdown. The only difference between (c) and (d) quantifications is that (c) includes all values, while d does not include any values of zero, which may indicate that the leg disc was not in position or obscured. For CD4-tdTomato, Nuf, Slit-D, Cap RNAi#1, Cap RNAi#2, Gol RNAi, Vrpl RNAi, Godzilla RNAi, Nrg RNAi, Prip RNAi, CG9304 RNAi#1, CG9304 RNAi#2, CG9304MI09328, N = 63, 26, 31, 32, 29, 32, 38, 19, 20, 68, 52, 66, 54 (c), and N = 37, 18, 21, 21, 20, 25, 12, 7, 12, 34, 8, 8,19 (d). All data are means ± SEM. The data were analyzed by one-way ANOVA followed by Dunn’s multiple comparisons test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Scale bar = 100 µm.
a Confocal images of dissected third instar larval leg discs with mRFP in glia (red) and DAPI (blue). The orange dashed line outlines the glial bundle area in the center of the disc. The yellow arrows mark the furthest point glia reach. If glia enter and migrate along the leg disc, this is the “migratory path.” b Confocal images of dissected third instar leg discs with mRFP in glia (red), as well as repo (green) and pH3 (blue) immunostaining. The yellow arrows mark the location of the furthest point glia reach. c Quantification of glial migration in dissected leg discs. N = 57, 42, 25, 12, 20, 11, 11, 16. d Quantification of glial bundle area in dissected leg discs. N = 17, 22, 14, 4, 11, 6, 4, 7. All data are means ± SEM. The data were analyzed by one-way ANOVA followed by Tukey’s multiple comparisons test. **P < 0.01, ****P < 0.0001. Scale bar = 100 µm.
GPR180 expression correlates with pLGG molecular subtypes and adult glioma survival
After identifying GPR180/CG9304’s pro-invasion role in our fly pLGG model, we next investigated its clinical significance in human gliomas by examining three human glioma datasets: PBTA, TCGA (The Cancer Genome Atlas), and CGGA (Chinese Glioma Genome Atlas). We found that the GPR180 mRNA level is significantly more enriched in BRAF-fusion pLGGs than in other molecular subtypes (Fig. 7A). BRAF-fusion induced pLGGs are a unique proteomic and phosphoproteomic cluster identified via tissue composition analyses33. GPR180 expression was both highest in and highly correlated with the proteomic (Fig. 7B) and phosphoproteomic (Fig. 7C) BRAF-fusion clusters. We also performed pathway enrichment analyses on the TCGA dataset comparing tumor patients with high vs. low GPR180 expression and found that GPCR, KRAS, and receptor tyrosine kinase signaling are highly enriched in pLGGs induced by RAF1/BRAF fusions (Fig. S3A, B). Given that RAF fusions (both RAF1 and BRAF) are known to be associated with increased MAPK signaling14, the correlation between GPR180 expression level and MAPK-related pathways further supports both that GPR180 expression is correlated with RAF1/BRAF fusions and that GPR180 may be a potential co-target for treating RAF1/BRAF-fusion driven pLGGs.
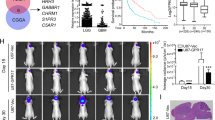
a Transcriptomic profiling of GPR180 mRNA expression in pLGG molecular subtypes. b GPR180 mRNA expression compared to the proteomic composition of pLGG molecular subtypes. c GPR180 mRNA expression compared to the phosphoproteomic composition of pLGG molecular subtypes. A Wilcoxon test was conducted in a one-vs-rest approach; asterisks indicate that an individual box is significantly different from the mean of the entire dataset. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. d–f The scatter plots with fitted regression lines indicate correlations between three of the highly correlated ADC-derived features with GPR180 expression levels. The box plots show the differences among the three imaging features in patients with low versus high GPR180 levels. N = 71 subjects that had ET. Unpaired Student’s t-test. *P < 0.05, ****P < 0.0001. g–i Correlation between GPR180 and survival/progression-free survival of LGG patients from the TCGA dataset. g is the overall survival of all LGG patients, h is the overall survival of astrocytoma patients, and i is the progression-free survival of astrocytoma patients. P value: log-rank test.
To assess the clinical significance of GPR180 expression levels, we queried the PBTA dataset to determine survival differences in patients with high vs. low GPR180 expression. We saw no significant correlation in overall survival (OS) (Fig. S5A, B) or in PFS (Fig. S5C) in either fusion-pLGGs or all pLGGs. Our inability to detect a difference in either OS or PFS could be related to the relatively long survival of pLGG patients, which may obscure small differences34.
We next sought to determine if there were detectable differences in MRI imaging of high vs. low GPR180-expressing pLGG patients. To investigate the relationship between GPR180 expression and radiographic imaging characteristics, we conducted a multiparametric MRI analysis of pLGG patients. Diffusion-weighted imaging (DWI) and its derived quantitative map, apparent diffusion coefficient (ADC), are MRI techniques that measure the proton mobility within the tissue. The information generated with these techniques reflects changes in tissue microstructure and cellular density that are markers of tumor progression35. In adult glioblastomas, lower ADC values in enhancing solid tumor and peritumoral edema regions within a tumor (definitions provided in Fig. S6) have been associated with elevated cell density and tumor infiltration36,37, making them a potential marker for tumor progression38. ADC values are not only affected by the cell density within the tumor but also by other factors, including tissue microstructure, such as tumor cell size, and microvascular perfusion37. As tumor tissues often exhibit heterogeneity, histogram analysis (described in Methods and Fig. S6), which measures the distribution of ADC values within a region of a given tumor, has been utilized to aid in the characterization and assessing treatment response in various tumors39. After analyzing the values of ADC maps within the tumorous subregions (detailed in Methods and Fig. S6) collected from 77 pLGG patients, we found significant differences in 10 ADC-derived features (Table S3). Three histogram-based features in the enhancing solid tumor region were particularly noteworthy: mean ADC (with correlation coefficient, R = 0.49, P = 0.000016), ADC root mean square (R = 0.49, P = 0.000017), and ADC histogram skewness (i.e., the measure of asymmetry of the probability distribution of a variable about its mean, where a negatively skewed intensity distribution might suggest a predominance of high-intensity areas within the tumor; R = -0.42, P = 0.000241). The former two features were positively correlated with GPR180 levels (Fig. 7D, E), while the latter was negatively correlated (Fig. 7F), suggesting that patients with higher GPR180 levels have higher ADC magnitude and predominance of high ADC values within the enhancing solid tumor.
To take it a step further and test whether ADC values within tumor regions could predict GPR180 expression levels, we applied a radiogenomic analysis approach that utilized a machine learning approach, multivariate logistic regression, along with ADC features to classify the patients into either GPR180 high or low groups. The performance of the resulting classification model was assessed using the AUC–ROC metric, which returned a value of 0.80 for differentiation of GPR180 low vs. high expression. The AUC–ROC (ranging between values of 0 and 1) is a measure of how well a model can distinguish between two classes, with higher values indicating better performance. Our classification result shows that the model, generated using ADC features, has a strong ability to differentiate high vs low GPR180 expression levels. This suggests the potential of ADC values to predict GPR180 expression levels36,37. Correlation of the volumes of the tumor subregions (detailed in “Methods”) was not statistically significant (P > 0.05).
We next examined whether the GPR180 expression level is correlated with survival in adult LGGs and all gliomas. In the TCGA dataset, we found that lower GPR180 expression is associated with better OS in LGG patients (Fig. 7G), and stratification to include only astrocytoma patients revealed not only better OS (Fig. 7H), but also better PFS (Fig. 7I). In the CGGA datasets, lower GPR180 expression is associated with better OS in patients with primary and recurrent gliomas (Fig. S5F, G). To add, GPR180 expression level is negatively correlated with the outcome of the first-course treatment in TCGA LGG patients; there are significantly more cases of “complete remission” and fewer cases of “progressive disease” in patients with low vs. high GPR180 expression (Fig. S5D, E). Overall, these data show that GPR180 expression level may serve as a biomarker in all pLGGs and a possible novel therapeutic target in pLGGs with RAF1/BRAF fusions.
Lastly, we decided to broaden the scope of our work to determine if GPR180/CG9304 may regulate tumorigenesis beyond gliomas. We used an established fly neuroblast-derived tumor model that overexpresses the bHLH transcriptional repressor dpn in the neuroblast lineage, which results in over-proliferation of both type I and type II neuroblasts and brain tumor formation40,41. Consistent with previous results, dpn overexpression in neuroblasts driven by insc-Gal4 drastically increased brain size. We focused on the type-ID neuroblasts in the VNC (Fig. S7A), the most accessible to in vivo live imaging, and quantified the neuroblast area. Overexpression of dpn doubled the neuroblast area, while knockdown of CG9304 in the model did not lead to an obvious reduction (Fig. S7B, C). This result suggests that GPR180/CG9304 may be more involved in gliomas, supporting its specificity and the importance of precision targeting in tumor types and molecular subtypes.
Discussion
The goal of this work was to focus on the phenotypic and molecular subset of pLGGs with the lowest PFS, where GTR is impossible due to the diffuse nature of the tumor. The recent shift towards targeted therapeutics requires developing an understanding of the molecular pathways underlying the diverse subtypes of pLGGs, which has been limited by a lack of genetic models6. To fill this gap, we developed the first five fly models of pLGGs, a system we felt best served our purpose to identify and examine aberrant glial migration, as an in vivo model can effectively replicate the infiltrative nature of pediatric-type diffuse LGGs, where GTR is more challenging42.
We selected the fruit fly for our purpose because it is a genetically tractable in vivo system that has been previously utilized to model brain tumors23,40,41, enabling us to screen a variety of gene fusions and possible modifiers for their impact on an infiltrative phenotype. We developed the fly models by identifying pLGG patient fusion genes of interest (QKI::RAF1, QKI::NTRK2, MKRN1::BRAF, FGFR1::TACC1, and KIAA1549::BRAF), overexpressed them in fly glia, and found that the aberrant glial migration induced by QKI::RAF1 overexpression best modeled the diffuse tumor pattern in human patient pLGGs. Due to our interest in this diffuse phenotype, we chose to focus on the QKI::RAF1 fusion to determine the causes of aberrant migration, with the goal of identifying novel targets not only for this fusion, but also for other diffuse/infiltrative pLGGs.
To determine how to ameliorate abnormal migration, we first used the fly to explore known and unknown regulators of glial migration and tumorigenesis. We found that overexpression of repulsive guidance signaling molecules Robo2 or PlexA/B, known regulators of glial migration and tumorigenesis24,43, was sufficient to suppress aberrant glial cell migration and observed that reduced Robo2, but not PlexA, expression in aberrantly migrating glia in the fly QKI::RAF1 model. The lack of differential expression of PlexA in contrast with Robo2 suggests different roles in regulating glial migration; Robo2 may function to prevent glia from migrating onto the leg disc periphery, while PlexA may function to restrict tumor cells from leaving the center glial bundle. The idea that PlexA may normally serve to restrict glia could explain why PlexA expression appears lower in the main glial bundle in the QKI::RAF1 model and why glial overexpression of PlexA in that model was able to reduce the overall glial bundle area.
It was critical to validate our fly findings in vitro and in humans as the fly does not sufficiently recapitulate the complexity of human pLGGs. Primary mouse astrocytes overexpressing QKI::RAF1 had reduced Robo2 expression, as did human patients with RAF1/BRAF fusions, suggesting the broader translatability of these findings and making repulsive guidance signaling molecules interesting candidates for the development of targeted therapeutics. Future studies into how different RAF1/BRAF fusions in pLGGs drive misexpression of repulsive guidance receptors as well as the signaling pathways downstream of ROBO2 and PlexA/B receptors, will be informative to better understand erroneous glial migration and tumorigenesis as well as identify additional targets. Additionally, while we know ROBO2 has a known role in metastasis in various tumors25,26,27 and our results demonstrate the importance of ROBO2 dysregulation in pLGG patients with RAF1/BRAF fusions, we have yet to explore ROBO2 in diffuse tumors with other genetic causes. Further exploration is critical to determining if repulsive guidance signaling is a common pathway in diffuse pLGGs.
Previously unknown regulators of glial migration and tumorigenesis were identified by conducting transcriptomic profiling of the fly QKI::RAF1 model and human pLGG patients with RAF1/BRAF fusions. The QKI::RAF1 fly model was utilized to screen the resulting candidates, and we found a promising target, GPR180/CG9304, a very understudied gene not previously linked to gliomas. In flies, glial GPR180/CG9304 loss-of-function reduced aberrant glial migration, linking GPR180/CG9304 to pLGG tumor cell infiltration. Follow-up clinical investigation utilizing transcriptomic, proteomic, and phosphoproteomic profiling identified GPR180 as a potential biomarker for RAF1/BRAF fusion-driven pLGGs.
Multiparametric imaging analyses from pLGG tumors showed that high GPR180 expression resulted in increased magnitude of ADC values and negative ADC histogram skewness, suggesting predominance of high ADC values in the enhancing solid tumor region. This may suggest increased interstitial water content, a phenomenon that can result from increased inflammation44. The radiogenomic analysis, employing machine learning along with ADC-derived features, demonstrated the potential of these imaging characteristics in predicting GPR180-high and GPR180-low tumor groups. Given that many of the pLGGs are not completely resectable, particularly when they are situated in deeply seated brain regions or exhibit high infiltration21, the predictive radiogenomics model suggested in this study offers a noninvasive approach for assessing GPR180 levels. This could help in guiding targeted treatments tailored to the molecular status of each patient. To note, we observed no significant association between gross tumor subregion volumes (enhancing, non-enhancing, cystic) and GPR180 expression. This is not entirely surprising, as volume is a relatively coarse morphologic metric that may not capture the microenvironmental remodeling driven by GPR180. Whereas volumetry reflects the macroscopic extent of tumor mass, GPR180 may predominantly influence tissue architecture at the cellular and interstitial levels. Consequently, diffusion metrics appear more sensitive to these subtle microstructural changes than bulk volume measurements. Thus, the absence of a volume correlation does not undermine our ADC-derived radiogenomic associations; instead, it underscores that ADC features and volumetry provide complementary, orthogonal information. Future studies incorporating voxel-wise mapping of GPR180 expression against both diffusion and volumetric maps will be critical to fully disentangle these relationships. In addition, we acknowledge that ADC is an imperfect surrogate for tumor invasion. Future studies should integrate more advanced diffusion techniques, such as diffusion kurtosis imaging and direct histologic quantification of cellularity and extracellular volume fraction, to disentangle these factors. Finally, further mechanistic research is needed to elucidate the relationship between ADC characteristics and GPR180 expression levels and to clarify the association between ADC and cell density as well as infiltration in pLGGs.
Additionally, through analysis of tumor databases, we saw that lower GPR180 expression is associated with better survival in adult patients with all LGGs, astrocytoma, and gliomas, particularly in recurrent cases. These findings highlight the therapeutic potential of targeting GPR180 in RAF1/BRAF fusion-induced pLGGs and in gliomas in general.
Our findings indicate that both repulsive guidance signaling and GPR180/CG9304 expression are dysregulated in the pLGG QKI::RAF1 model, contributing to the invasion-like phenotype we propose in Fig. S8. It is likely that QKI::RAF1 fusion induces glioma cell invasion by hijacking pathways involved in normal glial cell migration, which is reminiscent of recent reports suggesting gliomas can take control of pathways involved in neurodevelopment and plasticity30,45,46. We show that overexpression of the repulsive guidance signaling receptors Robo2 and PlexA/B or knockdown of GPR180 can mitigate fly aberrant glial migration, suggesting they are promising therapeutic targets. While we have demonstrated that repulsive guidance signaling and GPR180 are critical to glial infiltration phenotypes in the QKI::RAF1 fly model, we do not know how QKI::RAF1 downregulates Robo2 or upregulates GPR180. There are a variety of mechanisms that could be involved, including, but not limited to, methylation, transcription factor binding, and chromatin level regulations. We would encourage further exploration into the causes of infiltration to aid in discovering new therapeutic targets. Specifically, we believe GPR180 is an ideal target, yet very understudied and novel to the field of gliomas; additionally, there is currently no small molecule that directly targets GPR180; we would highlight this avenue as particularly important for future work. Our clinical findings: aberrant ROBO2 and GPR180/CG9304 expression in RAF1/BRAF fusion patients and ADC values that may be associated with increased inflammation in high GPR180 expressing pLGG patients underscore the promise of ROBO2 and GPR180 as therapeutic targets not only in RAF1/BRAF fusions, but also in other types of pLGGs. Further investigation is needed to determine if GPR180 is, in fact, a more general oncogenic driver in multiple types of pLGGs. In addition, while we found evidence to suggest GPR180 may be specific to gliomas, as we saw no effect of knockdown in a fly neuroblast tumor model, we still saw a general role of Robo2, PlexA/B, and GPR180 in glial infiltration. Therefore, more exploration is warranted to identify possible dysregulation in metastatic tumors. Overall, this study demonstrated a translatable pipeline for the development of novel tumor models to identify targeted therapeutics for pLGG patients that can be utilized to drive innovation in molecular-targeted medicine.
Methods
Fly stocks
Repo-Gal422, UAS-mRFP47, UAS-CD4-tdTomato48, UAS-dEGFRACT; dPI3KACT 23, UAS-robo249, UAS-PlexA50, UAS-PlexB51, robo2HA 28, PlexA-myc29, UAS-Nuf52, UAS-Slit-D53, insc-Gal4, UAS-dpn, UAS-mCD8-GFP40,54 have been previously described. UAS-Cap RNAiBL30506 (RNAi1), UAS-Cap RNAiBL36663 (RNAi2), UAS-Gol RNAiBL28785, UAS-Vrpl RNAiBL36704, UAS-Prip RNAiBL50695, UAS-CG9304 RNAiBL42544 (RNAi1) and CG9304MI09328 were from the Bloomington stock center. UAS-Godzilla RNAiv7518, UAS-Nrg RNAiv27201, and CG9304 RNAiv11142 (RNAi2) were from VDRC.
To create the UAS-QKI::RAF1, UAS-MKRN1::BRAF, UAS-FGFR1::TACC1, UAS-QKI::NTRK2, UAS-KIAA1549::BRAF, UAS-QKI.E48G::RAF1, and UAS-QKI::RAF1.R401H stocks, the fusion coding sequences were cloned into the pACU2 fly expression vector. The mEGFP-KIAA1549::BRAF plasmid55 was acquired from Addgene. The plasmids were then injected into fly embryos via φC31-mediated site-specific insertion (Rainbow Transgenic Flies Inc). All fly crosses, including larvae, were kept at 25 °C.
Live imaging in flies
Live imaging was conducted as previously described56,57. Randomly selected female and male third instar larvae were collected in vials containing standard fly food and screened for the correct genotype. Larvae were anesthetized using diethyl ether and individually mounted in halocarbon oil. The Zeiss LSM 880 microscope was used for live imaging. Consistency across imaging sessions was ensured by maintaining identical microscope settings, including laser intensity, format, scanning speed, and zoom factor.
Migration analysis
To quantify the invasion-like phenotype of the QKI::RAF1 flies, the length that glial cells traveled along the edges of the leg disc (in μm) was measured using the FIJI software.
Dissection and fixation
Third-instar larval brains and leg imaginal discs were dissected in PBS. The dissected tissues were fixed in 4% paraformaldehyde for 20 min, followed by three rinses in PBS. For samples that did not require immunostaining, the tissues were mounted and sealed for imaging. Brain samples were mounted with Vectashield mounting medium, while leg imaginal disc samples were mounted with Vectashield mounting medium with DAPI to facilitate visualization of disc cell morphology. Glial processes were visualized with repo-Gal4 > mRFP. Imaging was performed using a Leica SP8 confocal microscope. Consistency across imaging sessions was ensured by maintaining identical microscope settings, including laser intensity, format, scanning speed, and zoom factor.
Immunohistochemistry and quantifications
Standard fixation and preparation of tissue were performed and repeated at least three times (N > 3 each time). Following fixation, tissues were blocked at room temperature for 1 h in a blocking buffer containing 10% Normal Donkey Serum and 90% 1 × PBS + 0.3% triton (PBST). After blocking, the buffer was removed, and the samples were incubated with the primary antibody solution overnight at 4 °C. The primary antibody solution was then removed, and the samples were rinsed three times with blocking buffer. Next, the secondary antibody solution was added, and the samples were incubated at room temperature for 2 h. Following incubation, tissues were rinsed three times with 1 × PBST. Brain samples were subsequently mounted for imaging. For disc samples, an additional preparation step was conducted. After rinsing, the discs were incubated in a 50:50 solution of 1 × PBST and glycerol for 20 minutes. The samples were then incubated with Vectashield mounting medium with DAPI at 4 °C overnight before final mounting.
The following antibodies were used: mouse anti-HA (BioLegend, 901502, 1:1000), mouse anti-repo (DSHB, 8D12, 0.1 ml concentrate, 1:200), rat anti-histone H3 (Abcam, ab10543, 1:400), rabbit anti-Myc (Cell Signaling, 2272, 1:400), goat anti-HRP (Jackson ImmunoResearch, 123-605-021, 1:1000) and fluorescence-conjugated Alexa Fluor secondary antibodies (Jackson ImmunoResearch; 1:1000). Tissues were imaged using a Leica SP8 confocal microscope and analyzed in FIJI software. Fluorescence intensity was measured in a single plane of Z-stacks.
To quantify glial volume, brains were first dissected, fixed, and imaged on a Leica SP8 confocal. The Imaris software was used for volume quantification. Using Imaris’ segmentation tools, the outline of the fly brains was manually segmented. The software then automatically calculated the volume of each segmented fly brain. To determine the number of glia or mitotic cells in repo-Gal4 > mRFP areas, the following steps were taken. Brains were dissected and stained with anti-pH3 and anti-repo. Samples were then imaged on a Leica SP8 confocal. Using the “channel mask” function in Imaris, the repo channel was selected as the reference channel for creating the mask. This allowed us to isolate the repo+ areas. The software was then used to identify and quantify the number of pH3+ signals within the repo+ mask.
Survival assay
Repo-Gal4, UAS-mRFP flies were crossed with UAS-QKI::RAF1, UAS-QKI.E48G::RAF1, or UAS-QKI::RAF1.R401H. The crosses were kept in fly vials containing standard fly food at 25 °C. Once pupae were observed, the crosses were moved to room temperature and the vials were checked three times a day over the period of a week. To calculate normalized survival, the number of adult flies found to express the tumor fusion gene was first divided by the total number of adult flies collected each day. This value was then divided by the percentage of flies expected to express the tumor fusion gene, based on Mendelian inheritance.
Quantitative PCR (qPCR) and RT-PCR
qPCR was performed using PMAs according to the manufacturer’s protocols. The primers used were the following. For mouse experiments. Robo2: GCTGGACGCTAGATCTCTCC and GGAAGGAAAAGGGGAAGGCA. Plxna1: TTCTGGAAGGCCGAGTCAAC and TGGAAAAGGCACTCGTACCC. Plxna2: CAGACCTTTCCCCATGACACA and ACAAATTGTGTCCTTGGCTTGC. Plxna3: CTTCACCATAGGAGGGTGCC and CTCAATGGGCCCTGTGACAT. Plxna4: GGTGGCCGAGAAGATGTTGA and CTCACTTAGCGAGTAGCGGG. Plxnb1: TGTCTCCTTTGTTCCCAGCG and GCATGACCTGAGCAGGAGTC. Plxnb2: AACGCAAGGAAACGGCTCTA and CCCAGAACTCAAGGCTCCAG. Plxnb3: TGACTGACTTCCTCCAGGCT and GAGGTGATTCACTGCACCCA. Robo1: AAAGATGACGGAGAGCTGCC and AATGTGGCGGCTCTTGAACT. Robo3: CCAAGAGGAGCAAGGATGGG and TGCACAAAACAAGCAGGGAC. DCC: TGATGATTTCCCCACCTCGG and AACGGTGTACAGTCGCACAT. Epha1: ATGTGGACCTCCAGGCCTAT and ACCAGTAGCCATCTGGGGAT. For fly experiments. rp49: CAGTCGGATCGATATGCTAAGCTG and TAACCGATGTTGGGCATCAGATAC. CG9304: CCGCAACACCATGAAGTACG and TGCATAAACTACCGCAACGA, CTGCAGCGATGAAGATGAACC and AGCGTGTTGAGGAGATTGCT.
RNA-sequencing
For the human patient sample ROBO2 analysis, the methods utilized for expression and fusion data generation are as follows. We used STAR to align paired-end RNA-seq reads, and used the associated alignment for all subsequent RNA analyses. We used Ensembl GENCODE 27 “Comprehensive gene annotation” as a reference. We used RSEM for both FPKM and TPM transcript- and gene-level quantification. For the generation of ROBO2 boxplots, we used the TPM data (Table S1).
We set up Arriba and STAR-Fusion fusion detection tools using CWL on CAVATICA. For both tools, we used aligned BAM and chimeric SAM files from STAR as inputs and GRCh38_gencode_v27 GTF for gene annotation. We ran STAR-Fusion with default parameters and annotated all fusion calls with the GRCh38_v27_CTAT_lib_Feb092018.plug-n-play.tar.gz file from the STAR-Fusion release. For Arriba, we used a blacklist file, blacklist_hg38_GRCh38_2018-11-04.tsv.gz from the Arriba release to remove recurrent fusion artifacts and transcripts present in healthy tissue. We provided Arriba with strandedness information for stranded samples, or we set it to auto-detection for poly-A samples. We used FusionAnnotator on Arriba fusion calls to harmonize annotations with those of STAR-Fusion.
For flies, WT and QKI::RAF1 larval brains and VNCs were dissected. 3 WT and 3 QKI::RAF1 samples were prepared, each containing about 50 brains and VNCs. RNA extraction and cDNA library were prepared and sequenced according to the manufacturer’s protocols.
The QKI::RAF1 fly vs normal comparison was performed on a total of 24723 genes (Table S2, Sheet 1). Out of these, 6730 genes were mapped to 9157 human orthologs (Table S2, Sheet 2). These 9157 genes were used as input for the volcano plot. A total of 635 genes were differentially expressed out of the 9157 genes, with 303 genes being down-regulated and 332 being up-regulated at log2FC > 0 and FDR-adjusted P-value < 0.05. We annotated our gene of interest, GPR180, which is upregulated in the tumor samples with a log2 fold change value of 0.837 and FDR P-value of 0.005. The vertical cut-off line is at log2FC of 0, and the horizontal cut-off line is at FDR-adjusted P-value of 0.05 (Fig. 4C).
The pLGG BRAF1/RAF1 fusion vs no-fusion comparison was performed on a total of 26449 genes (Table S2, Sheet 3). A total of 4531 genes were differentially expressed out of the 26449 genes, with 3177 genes being downregulated and 1354 genes being upregulated at log2FC > 0 and FDR-adjusted P-value < 0.05 (Fig. 4D). We annotated GPR180, which is upregulated in the BRAF1/RAF1 fusion samples with a log2 fold change value of 0.44 and FDR P-value of 0.016. The vertical cut-off line is at log2FC of 0, and the horizontal cut-off line is at FDR-adjusted P-value of 0.05 (Fig. 4D). The R package EnhancedVolcano was used to generate the volcano plots.
A total of 87 genes were identified to be differentially expressed (Table S2, Sheet 4) in both comparisons in the same direction: (1) QKI::RAF1 vs normal and (2) pediatric LGG BRAF1/RAF1 fusion vs no-fusion samples. We used these genes to plot a heatmap of the QKI::RAF1 vs normal samples (Fig. 4E). The R package pheatmap was used to generate the heatmap. The R package VennDiagram was used to generate the Venn diagrams.
Pathway enrichment analyses
We used the Compute Overlaps tool by Molecular Signature DB available at http://www.gsea-msigdb.org/gsea/msigdb/annotate.jsp to identify significantly deregulated pathways corresponding to our genes of interest.
Proteomics and phosphoproteomics analyses
Proteomic and phosphoproteomic clustering data from 218 tumors across 7 histologies published previously33 and mRNA gene expression data from OpenPBTA project58 were utilized to establish GPR180 gene expression correlation. Briefly, log2 expression counts of GPR180 were obtained from OpenPBTA release v22 in order to correlate with proteomics and phospho-proteomics clusters. The significance was calculated using the Wilcoxon test by comparing each cluster to the rest of the clusters.
pLGG molecular subtype analysis
Plots were generated using the latest release v22 of OpenPBTA58.
Multiparametric imaging analyses in pLGG patients
Our retrospective study was compliant with HIPAA and obtained approval from the IRB of the Children’s Hospital of Philadelphia (CHOP) through the Children’s Brain Tumor Network (CBTN) Protocol. MRI exams were collected from the subjects enrolled in the CBTN, which is a biorepository (cbtn.org 21) that allows for the collection of specimens, longitudinal clinical and imaging data, and sharing of de-identified samples and data for future research. For a subset of N = 77 subjects, pre-surgical and pre-treatment MRI scans acquired as part of standard of care practice were retrospectively collected for this study. Multiparametric MRI (mpMRI) scans (T1-weighted pre- (T1) and post-contrast (T1-Gd), T2-weighted (T2), and T2-FLAIR imaging, diffusion-weighted imaging (DWI) sequences and apparent diffusion coefficient (ADC) maps derived from DWI on the scanner) were included. DWI scans were obtained with an echo planar imaging (EPI) pulse sequence with a matrix size of 128 × 128, in-plane voxel size of 2 × 2 mm2, and diffusion weightings (b-values) of 0 and 1000 s/mm2.
Cancer Imaging Phenomics Toolkit open-source software (CaPTk, https://www.cbica.upenn.edu/captk) was used to pre-process the images59. The images were co-registered to the SRI atlas60 and resampled to a 1 × 1 × 1 mm3 resolution using a co-registration method named the Greedy tool (https://github.com/pyushkevich/greedy)61. ADC maps were co-registered with their corresponding T1-Gd scans using the Greedy tool. Skull stripping was then performed using our in-house pediatric skull-stripping method.
Tumor segmentation was performed semi-automatically. Pre-processed standard MRI sequences (T1, T1-Gd, T2, and T2-FLAIR) were initially segmented automatically using a pediatric-specific deep learning segmentation method freely available at https://github.com/d3b-center/peds-brain-auto-seg-public, to identify RAPNO-recommended tumor subregions (definitions can be found in refs. 62,63), including enhancing solid tumor (ET), non-enhancing solid tumor (NET), peri-tumoral edema (ED), and cystic components (CC). Manual revisions were performed when necessary using ITK-SNAP (http://www.itksnap.org/)64. Enhancing solid tumors are described by areas with enhancement (brightness) on T1-Gd images as compared to pre-contrast T1. The cystic component appears hyperintense (very bright) on T2 and hypointense (dark) on T1-Gd. Non-enhancing solid tumor represents any abnormal signal intensity within the tumoral region that cannot be defined as enhancing or cystic. Edema is defined by the abnormal hyperintense signal (very bright) on FLAIR scans. Figure S3 shows some of these subregions overlaid on an MRI scan.
Volumes of each tumor subregion, i.e., ET, NET, CC, and ED, the union of all subregions presenting the whole tumor, as well as volumetric ratios, including ET/whole tumor, NET/whole tumor, CC/whole tumor, and ED/whole tumor, were computed. ADC-maps were rescaled to the range of 0 and 255, after removing the outlier pixels with intensities beyond the 99.99th percentile. Histogram analysis was performed to investigate ADC heterogeneity within subregions. Histogram analysis is a statistical method applied to the image for the quantification and characterization of tissue properties within a specified region of interest (here, tumor subregions). The histogram is a graphical representation of the distribution of image intensity values. This analysis method provides valuable insights into the heterogeneity and textural characteristics of tissue. By assessing the spread (range), central tendency (mean or median), and distribution shape (skewness, kurtosis) of the intensity values, histogram analysis helps in differentiating tumors with different characteristics. For each patient, all pixel values on ADC maps within the four tumor subregions, i.e., ET and NET, were divided into 10 bins. 44 histogram features, including mean, standard deviation, root mean square, skewness, kurtosis, entropy, energy, and percentiles (25th, 50th, 75th, and 90th), were calculated from four subregions on the ADC-maps across the tumor subregions, using CaPTk software.
Pearson’s and Spearman’s rank correlation was performed to explore associations between the extracted ADC features and GPR180 expression levels. A level of P = 0.05 was considered for statistical significance. To account for Type I error, correction for multiple comparisons was performed using Bonferroni’s method. Furthermore, to evaluate the potential of the combinations of ADC-derived variables in predicting high and low levels of GPR180, multivariate machine learning classification using logistic regression was performed by splitting the data into 70% training and 30% test. Area under the receiver operating characteristic curve (AUC–ROC), as a metric for evaluation of the performance of the machine learning classification model, was calculated. The patients were stratified into GPR180-low and GPR180-high classes using the median of GPR180 levels calculated in the whole PBTA study.
Publicly available datasets collection
Clinical information and data of molecular biomarkers were generated from PBTA58, TCGA, and CGGA publications65,66,67,68. The clinical information and gene expression datasets of PBTA were accessed from the Pediatric Brain Tumor Atlas via (https://github.com/AlexsLemonade/OpenPBTA-analysis). The clinical information and gene expression datasets of TCGA were accessed from the R2 platform (https://hgserver1.amc.nl). The clinical information and gene expression datasets of CGGA (mRNAseq_693 and mRNAseq_325) were accessed from the Chinese Glioma Genome Atlas (http://www.cgga.org.cn/index.jsp).
Statistics and reproducibility
No statistical methods were used to pre-determine sample sizes, but our sample sizes are similar to those reported in previous publications9,10, and the statistical analyses were done afterward without interim data analysis. Data distribution was assumed to be norma,l but this was not formally tested. Each experiment was repeated independently at least three times. The values of “N” (sample size) are provided in the Fig. legends. Data are expressed as mean in bar graphs for percentage, mean ± SEM in bar graphs and scatter plots. No data points were excluded except where mentioned otherwise. A two-tailed unpaired Student’s t-test was performed for comparison between two groups of samples. One-way ANOVA followed by multiple comparison tests was performed for comparisons among three or more groups of samples. Two-sided Fisher’s exact test was used to compare the percentage. Statistical significance was assigned, *P < 0.05, **P < 0.01, ***P < 0.001. Statistical analyses were performed using GraphPad Prism 8.
Data availability
All data supporting the findings of this study are provided within the paper and its supplementary information. All additional information will be made available upon reasonable request to the corresponding author. Raw data is available on request and can be accessed at https://cavatica.sbgenomics.com/u/d3b-bixu-ops/sd-x11yenk7-rnaseq.
References
Sievert, A. J. & Fisher, M. J. Pediatric low-grade gliomas. J. Child Neurol. 24, 1397–1408 (2009).
Ostrom, Q. T. et al. Alex’s Lemonade Stand Foundation Infant and Childhood Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2007-2011. Neuro Oncol. 16, x1–x36 (2015).
Wisoff, J. H. et al. Primary neurosurgery for pediatric low-grade gliomas: a prospective multi-institutional study from the Children’s Oncology Group. Neurosurgery 68, 1548–1554 (2011).
Jooma, R., Waqas, M. & Khan, I. Diffuse low-grade glioma - changing concepts in diagnosis and management: a review. Asian J. Neurosurg. 14, 356–363 (2019).
Aarsen, F. K. et al. Functional outcome after low-grade astrocytoma treatment in childhood. Cancer 106, 396–402 (2006).
Manoharan, N., Liu, K. X., Mueller, S., Haas-Kogan, D. A. & Bandopadhayay, P. Pediatric low-grade glioma: targeted therapeutics and clinical trials in the molecular era. Neoplasia 36, 100857 (2023).
Louis, D. N. et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol. 23, 1231–1251 (2021).
Aldape, K. et al. Challenges to curing primary brain tumours. Nat. Rev. Clin. Oncol. 16, 509–520 (2019).
Huang, X. et al. EAG2 potassium channel with evolutionarily conserved function as a brain tumor target. Nat. Neurosci. 18, 1236–1246 (2015).
Chen, X. et al. A feedforward mechanism mediated by mechanosensitive ion channel PIEZO1 and tissue mechanics promotes glioma aggression. Neuron 100, 799–815.e797 (2018).
Francisco, M. A. et al. Chloride intracellular channel 1 cooperates with potassium channel EAG2 to promote medulloblastoma growth. J. Exp. Med. 217, e20190971 (2020).
Alturkustani, M. Infiltration in pilocytic astrocytoma: a diagnostic pitfall. Cureus 14, e27940 (2022).
Zhang, J. et al. Whole-genome sequencing identifies genetic alterations in pediatric low-grade gliomas. Nat. Genet. 45, 602–612 (2013).
Jain, P. et al. CRAF gene fusions in pediatric low-grade gliomas define a distinct drug response based on dimerization profiles. Oncogene 36, 6348–6358 (2017).
Jones, D. T. et al. Recurrent somatic alterations of FGFR1 and NTRK2 in pilocytic astrocytoma. Nat. Genet. 45, 927–932 (2013).
Bandopadhayay, P. et al. MYB-QKI rearrangements in angiocentric glioma drive tumorigenicity through a tripartite mechanism. Nat. Genet. 48, 273–282 (2016).
Sievers, P. et al. Epigenetic profiling reveals a subset of pediatric-type glioneuronal tumors characterized by oncogenic gene fusions involving several targetable kinases. Acta Neuropathol. 144, 1049–1052 (2022).
Forshew, T. et al. Activation of the ERK/MAPK pathway: a signature genetic defect in posterior fossa pilocytic astrocytomas. J. Pathol. 218, 172–181 (2009).
Ross, J. S. et al. The distribution of BRAF gene fusions in solid tumors and response to targeted therapy. Int J. Cancer 138, 881–890 (2016).
Jones, D. T. et al. Tandem duplication producing a novel oncogenic BRAF fusion gene defines the majority of pilocytic astrocytomas. Cancer Res. 68, 8673–8677 (2008).
Ryall, S., Tabori, U. & Hawkins, C. Pediatric low-grade glioma in the era of molecular diagnostics. Acta Neuropathol. Commun. 8, 30 (2020).
Sepp, K. J., Schulte, J. & Auld, V. J. Peripheral glia direct axon guidance across the CNS/PNS transition zone. Dev. Biol. 238, 47–63 (2001).
Read, R. D., Cavenee, W. K., Furnari, F. B. & Thomas, J. B. A drosophila model for EGFR-Ras and PI3K-dependent human glioma. PLoS Genet. 5, e1000374 (2009).
Sasse, S. & Klambt, C. Repulsive epithelial cues direct glial migration along the nerve. Dev. Cell 39, 696–707 (2016).
Harburg, G. C. & Hinck, L. Navigating breast cancer: axon guidance molecules as breast cancer tumor suppressors and oncogenes. J. Mammary Gland Biol. Neoplasia 16, 257–270 (2011).
Prasad, A., Fernandis, A. Z., Rao, Y. & Ganju, R. K. Slit protein-mediated inhibition of CXCR4-induced chemotactic and chemoinvasive signaling pathways in breast cancer cells. J. Biol. Chem. 279, 9115–9124 (2004).
Latil, A. et al. Quantification of expression of netrins, slits and their receptors in human prostate tumors. Int J. Cancer 103, 306–315 (2003).
Spitzweck, B., Brankatschk, M. & Dickson, B. J. Distinct protein domains and expression patterns confer divergent axon guidance functions for Drosophila Robo receptors. Cell 140, 409–420 (2010).
Pecot, M. Y. et al. Multiple interactions control synaptic layer specificity in the Drosophila visual system. Neuron 77, 299–310 (2013).
Venkatesh, H. S. et al. Electrical and synaptic integration of glioma into neural circuits. Nature 573, 539–545 (2019).
Venkatesh, H. S. et al. Targeting neuronal activity-regulated neuroligin-3 dependency in high-grade glioma. Nature 549, 533–537 (2017).
Venkatesh, H. S. et al. Neuronal activity promotes glioma growth through neuroligin-3 secretion. Cell 161, 803–816 (2015).
Petralia, F. et al. Integrated proteogenomic characterization across major histological types of pediatric brain cancer. Cell 183, 1962–1985.e1931 (2020).
Bandopadhayay, P. et al. Long-term outcome of 4,040 children diagnosed with pediatric low-grade gliomas: an analysis of the Surveillance Epidemiology and End Results (SEER) database. Pediatr. Blood Cancer 61, 1173–1179 (2014).
Padhani, A. R. et al. Diffusion-weighted magnetic resonance imaging as a cancer biomarker: consensus and recommendations. Neoplasia 11, 102–125 (2009).
Lee, C.-Y., Bennett, K. M., Debbins, J. P., Choi, I.-Y. & Lee, P. The relationship between diffusion heterogeneity and microstructural changes in high-grade gliomas using Monte Carlo simulations. Magn. Reson. Imaging 85, 108–120 (2022).
Fathi Kazerooni, A. et al. Characterization of active and infiltrative tumorous subregions from normal tissue in brain gliomas using multiparametric MRI. J. Magn. Reson. Imaging 48, 938–950 (2018).
Song, Y. S. et al. True progression versus pseudoprogression in the treatment of glioblastomas: a comparison study of normalized cerebral blood volume and apparent diffusion coefficient by histogram analysis. Korean J. Radiol. 14, 662–672 (2013).
Just, N. Improving tumour heterogeneity MRI assessment with histograms. Br. J. Cancer 111, 2205–2213 (2014).
Zhu, S. et al. The bHLH repressor Deadpan regulates the self-renewal and specification of Drosophila larval neural stem cells independently of Notch. PLoS ONE 7, e46724 (2012).
Huang, X. et al. Voltage-gated potassium channel EAG2 controls mitotic entry and tumor growth in medulloblastoma via regulating cell volume dynamics. Genes Dev. 26, 1780–1796 (2012).
Thon, N., Tonn, J. C. & Kreth, F. W. The surgical perspective in precision treatment of diffuse gliomas. Onco Targets Ther. 12, 1497–1508 (2019).
Chedotal, A., Kerjan, G. & Moreau-Fauvarque, C. The brain within the tumor: new roles for axon guidance molecules in cancers. Cell Death Differ. 12, 1044–1056 (2005).
Galban, C. J. et al. Evaluation of treatment-associated inflammatory response on diffusion-weighted magnetic resonance imaging and 2-[18F]-fluoro-2-deoxy-D-glucose-positron emission tomography imaging biomarkers. Clin. Cancer Res. 16, 1542–1552 (2010).
Zeng, Q. et al. Synaptic proximity enables NMDAR signalling to promote brain metastasis. Nature 573, 526–531 (2019).
Venkataramani, V. et al. Glutamatergic synaptic input to glioma cells drives brain tumour progression. Nature 573, 532–538 (2019).
Leiss, F. et al. Characterization of dendritic spines in the Drosophila central nervous system. Dev. Neurobiol. 69, 221–234 (2009).
Han, C., Jan, L. Y. & Jan, Y. N. Enhancer-driven membrane markers for analysis of nonautonomous mechanisms reveal neuron-glia interactions in Drosophila. Proc. Natl Acad. Sci. USA 108, 9673–9678 (2011).
Rajagopalan, S., Vivancos, V., Nicolas, E. & Dickson, B. J. Selecting a longitudinal pathway: Robo receptors specify the lateral position of axons in the Drosophila CNS. Cell 103, 1033–1045 (2000).
Winberg, M. L. et al. Plexin A is a neuronal semaphorin receptor that controls axon guidance. Cell 95, 903–916 (1998).
Ayoob, J. C., Terman, J. R. & Kolodkin, A. L. Drosophila Plexin B is a Sema-2a receptor required for axon guidance. Development 133, 2125–2135 (2006).
Bellen, H. J. et al. The BDGP gene disruption project: single transposon insertions associated with 40% of Drosophila genes. Genetics 167, 761–781 (2004).
von Philipsborn, A. C. et al. Neuronal control of Drosophila courtship song. Neuron 69, 509–522 (2011).
Wallace, K., Liu, T. H. & Vaessin, H. The pan-neural bHLH proteins DEADPAN and ASENSE regulate mitotic activity and cdk inhibitor dacapo expression in the Drosophila larval optic lobes. Genesis 26, 77–85 (2000).
Tripathi, S. et al. Defining the condensate landscape of fusion oncoproteins. Nat. Commun. 14, 6008 (2023).
Parrish, J. Z., Emoto, K., Kim, M. D. & Jan, Y. N. Mechanisms that regulate establishment, maintenance, and remodeling of dendritic fields. Annu Rev. Neurosci. 30, 399–423 (2007).
Emoto, K., Parrish, J. Z., Jan, L. Y. & Jan, Y. N. The tumour suppressor Hippo acts with the NDR kinases in dendritic tiling and maintenance. Nature 443, 210–213 (2006).
Shapiro, J. A. et al. OpenPBTA: An Open Pediatric Brain Tumor Atlas. bioRxiv, https://doi.org/10.1101/2022.09.13.507832 (2022).
Davatzikos, C. et al. Cancer imaging phenomics toolkit: quantitative imaging analytics for precision diagnostics and predictive modeling of clinical outcome. J. Med. Imaging 5, 011018 (2018).
Rohlfing, T., Zahr, N. M., Sullivan, E. V. & Pfefferbaum, A. The SRI24 multichannel atlas of normal adult human brain structure. Hum. Brain Mapp. 31, 798–819 (2010).
Yushkevich, P. A. et al. IC-P-174: Fast automatic segmentation of hippocampal subfields and medial temporal lobe subregions in 3 tesla and 7 tesla T2-weighted MRI. Alzheimer’s Dement. 12, P126–P127 (2016).
Erker, C. et al. Response assessment in paediatric high-grade glioma: recommendations from the Response Assessment in Pediatric Neuro-Oncology (RAPNO) working group. Lancet Oncol. 21, e317–e329 (2020).
Fangusaro, J. et al. Response assessment in paediatric low-grade glioma: recommendations from the Response Assessment in Pediatric Neuro-Oncology (RAPNO) working group. Lancet Oncol. 21, e305–e316 (2020).
Yushkevich, P. A., Gao, Y. & Gerig, G. ITK-SNAP: an interactive tool for semi-automatic segmentation of multi-modality biomedical images. In: 2016 38th annual international conference of the IEEE engineering in medicine and biology society (EMBC)). (IEEE, 2016).
Ceccarelli, M. et al. Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164, 550–563 (2016).
Zhao, Z. et al. Comprehensive RNA-seq transcriptomic profiling in the malignant progression of gliomas. Sci. Data 4, 170024 (2017).
Liu, X. et al. A radiomic signature as a non-invasive predictor of progression-free survival in patients with lower-grade gliomas. Neuroimage Clin. 20, 1070–1077 (2018).
Zhao, Z. et al. Chinese Glioma Genome Atlas (CGGA): a Comprehensive Resource with Functional Genomic Data from Chinese Glioma Patients. Genomics Proteom. Bioinforma. 19, 1–12 (2021).
Acknowledgements
We thank G. Bashaw and S. Zhu for sharing fly lines; Bloomington Stock Center and VDRC for fly stocks; GenScript for plasmids; and members of the Song lab for helpful discussions. This work was supported by an NIH grant (1R01NS107392) to Y.S. and NIH grants (U2CHL138346, R01NS085336) to A.R. G.J. is a recipient of the Goldfeder Family Undergraduate Research Grant and the Eliot Stellar Prize.
Author information
Authors and Affiliations
Contributions
Experimental design, G.J., Z.L, P.J., F.L., and Y.S.; methodology, G.J., P.J., Z.L., F.L., A.F., M.P.K., A.J.W., A.N., A.C.R., and Y.S.; data collection and analysis, G.J., Z.L., P.J., F.L., K.S.R., Z. S., X.C., J.S., A.F., Y.S., M.P.K., S.S., A.F.K., A.N., A.N., X.H., A.C.R., and Y.S.; writing—original draft, G.J., Z.L., H.M.D., and Y.S.; writing—review & editing, P.J., X.C., A.F., M.P.K., A.J.W., A.N., and X.H.; funding acquisition, A.C.R., and Y.S.; supervision, A.N., X.H., A.C.R., and Y.S. All authors read, edited, and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jung, G., Liang, Z., Jain, P. et al. Role of repulsive guidance signaling and GPR180 in pediatric low-grade glioma infiltration. npj Precis. Onc. 9, 344 (2025). https://doi.org/10.1038/s41698-025-01121-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01121-2