Abstract
Reliable lymph node metastasis (LNM) assessment underpins staging and treatment decisions in lung adenocarcinoma (LUAD), yet approximately 10% of patients remain pNx due to sampling limitations. Existing LNM prediction methods mainly rely on imaging or molecular data, while the potential of histopathological whole-slide images (WSIs) remains underexplored. We introduce CTMIL, a customized transformer-based multiple instance learning framework for predicting LNM directly from primary LUAD WSIs. Three CTMIL models were trained on 320 TCGA cases using patch-level features extracted from ResNet50, InceptionResNetV2, and UNI backbones, and internally validated on 80 held-out cases. External evaluation on two independent cohorts (XY cohort, 149 cases, and RM cohort, 164 cases) showed that CTMIL models consistently outperformed attention-based, TransMIL, and baseline approaches under identical inputs. The best-performing model, UNI_CTMIL, achieved AUROCs of 0.8640, 0.8216, and 0.8090 on the validation, XY, and RM cohorts, respectively. Interpretability analyses showed that UNI_CTMIL attends to metastatic hallmarks such as micropapillary and solid patterns, mucin, poor differentiation, and high tumor cell density. These findings highlight CTMIL’s promise for histology-based LNM prediction, potentially in pNx cases.
Similar content being viewed by others
Introduction
Lymph node metastasis (LNM) is an independent predictor of poor prognosis in non-small cell lung cancer (NSCLC)1. Specifically, the five-year survival rate in NSCLC declines drastically from approximately 87% in localized disease to around 36% once regional LNM occurs2,3. Lung adenocarcinoma (LUAD), the most prevalent and notably heterogeneous subtype of NSCLC4,5,6, frequently exhibits occult nodal metastasis7, and around 6–11% of patients remain with undetermined nodal status (pNx) postoperatively8,9,10. This persistent uncertainty makes accurate nodal staging particularly challenging in LUAD. Improving LNM assessment and prediction in this subtype is therefore critical for optimal clinical management.
To date, efforts to predict LNM in LUAD have primarily focused on radiological imaging and molecular profiling approaches. Numerous studies have applied deep learning models to PET/CT imaging for the preoperative prediction of lymph node involvement11,12,13,14,15,16. For instance, Zhong et al. developed a PET/CT-based deep learning model (DLNMS) demonstrating substantial predictive capability in detecting occult nodal metastasis preoperatively11. Concurrently, molecular profiling studies have identified biomarkers associated with LNM, including differential gene expression17,18 and genomic alterations such as ALK rearrangements19 and EGFR exon 19 deletions20. Although these imaging and molecular approaches have revealed important aspects of metastatic progression, they often fail to capture key morphological and microenvironmental features that are preserved in histopathological tissue samples. This gap presents an opportunity for computational pathology to uncover morphological correlates of LNM by analyzing spatial and cellular features preserved in routine H&E-stained WSIs.
Histopathological whole-slide images (WSIs) from primary tumors constitute an abundant yet underexploited resource for metastasis prediction21,22. WSIs preserve rich spatial and cytomorphological information reflective of tumor biology, potentially harboring biomarkers predictive of LNM. Pathological studies corroborate this notion, as aggressive histologic subtypes, such as micropapillary or solid patterns in LUAD, have been strongly linked to increased LNM risk23. Computational pathology enables the identification of histological markers of metastasis that are invisible to radiological imaging, highlighting its potential to expand the clinical understanding of tumor biology beyond what imaging alone can provide. However, research exploring direct LNM prediction from primary LUAD histopathology WSIs remains limited. Recent studies have started to apply deep learning approaches for nodal metastasis prediction using WSIs in cancers such as lung squamous cell carcinoma (LUSC)24, colorectal cancer25,26,27, cervical cancer28, and bladder cancer29, but no established approaches currently exist specifically for LUAD.
Multiple instance learning (MIL) has emerged as a promising computational paradigm for weakly supervised analysis of WSIs, wherein a slide is represented as a collection (“bag”) of image patches (“instances”), enabling predictions from slide-level labels without detailed region-level annotations30,31. Among MIL methods, attention-based MIL (AMIL) introduces trainable attention mechanisms, effectively identifying diagnostically significant patches within WSIs32. Recently, transformer-based MIL architectures (for example, TransMIL) have advanced this methodology by capturing complex contextual relationships among patches through self-attention mechanisms33. Such transformer-based approaches have achieved notable success in various WSI tasks, including diagnosis, tumor subtyping, and survival prediction34,35,36,37,38,39, underscoring their potential in computational pathology tasks. Nevertheless, the application of MIL, particularly transformer-based models, remains largely unexplored for the intricate task of LNM prediction from primary LUAD WSIs.
In this study, we propose CTMIL, a customized transformer-based MIL framework specifically designed to predict LNM from WSIs of primary LUAD. Building upon the foundational TransMIL architecture, CTMIL incorporates deeper self-attention transformer layers, positional encoding, and an auxiliary patch-level supervision strategy to better capture morphological and spatial features. Through training, validation, and external testing across multiple independent LUAD cohorts, we demonstrate the strong performance of CTMIL-based models in LNM prediction and highlight the potential of transformer-based MIL frameworks for addressing complex, biologically structured tasks in computational pathology beyond conventional classification.
Results
Overview of the study
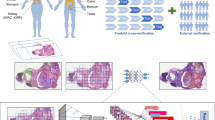
This study presents CTMIL, a customized transformer-based MIL framework designed to predict LNM from primary WSIs in LUAD (Fig. 1). A tumor patch detection model was first developed to isolate tumor-positive regions, ensuring that downstream analyses focused on diagnostically relevant areas. Feature embeddings extracted from these tumor patches using three backbone networks (ResNet50, InceptionResNetV2, and UNI) were separately input into the CTMIL architecture, which incorporates spatial coordinates via positional embeddings and employs a deeper transformer encoder along with auxiliary patch-level supervision to enhance discriminative capacity. The TCGA cohort was used for model training and internal validation, while two independent cohorts (RM and XY) served as external test sets to evaluate generalization performance (Fig. 2). Comparative evaluation was conducted between CTMIL-based models and AMIL-, original TransMIL-based models under multiple feature extraction settings, as well as other baseline models. To enhance interpretability, we further conducted in-depth analyses on the best-performing CTMIL model (UNI_CTMIL). Specifically, we visualized the contribution of each patch to the final prediction using attention scores from the last transformer layer and assessed its directional influence on positive or negative outcomes via patch-level classification scores. We also investigated the histopathological features associated with model predictions.
WSIs whole slide images, MIL multiple instance learning, CTMIL customized transformer-based multiple instance learning, AMIL attention-based multiple instance learning, IRV2 InceptionResNetV2, RN50 ResNet50.
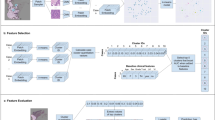
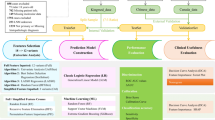
Process of the case enrollment.
Selection of the optimal CNN model for tumor patch detection
We systematically compared five pre-trained CNN architectures (ResNet50, InceptionV3, InceptionResNetV2, UNI, and Virchow2) for automated tumor patch detection. Performance was evaluated across training, validation, and independent test sets using standard classification metrics (accuracy, AUC, precision, recall, specificity, and F1-score) (Supplementary Table 3). InceptionResNetV2 consistently outperformed the other models across all datasets. On the test set, it achieved the highest accuracy (0.9892), AUC (0.9997), and F1-score (0.9893), demonstrating excellent balance between sensitivity (0.9967) and specificity (0.9817). ResNet50 also performed well but slightly lagged behind, while InceptionV3, UNI, and Virchow2 showed comparatively lower accuracy and AUC. Based on these results, InceptionResNetV2 was selected as the optimal patch classifier for downstream analyses. Optimal classification thresholds for each model were empirically selected based on validation performance and are reported in Supplementary Table 3.
Comparative evaluation of CTMIL-based and baseline MIL models for LNM prediction
We trained LNM prediction models using patch-level features extracted by three backbone networks (ResNet50, InceptionResNetV2, and UNI) and two MIL architectures (CTMIL and AMIL), resulting in six paired variants. An ensemble approach (Ensemble_CTMIL) was constructed by averaging predictions from the three CTMIL models. For baseline comparison, non-attention-based MIL models were also included, each using different aggregation strategies for slide-level inference. All models were evaluated by AUROC and AUPRC (Fig. 3A–L, Supplementary Fig. 1A–D). Subsequent evaluations prioritized the validation and external test cohorts (RM and XY) to assess the generalizability of these models.
Performance (AUROC) of CTMIL models and baseline methods for LNM prediction on the training set (A, E, I), validation set (B, F, J), and two independent test cohorts (C, D, G, H, K, L). Patch_baseline, AUROC computed from raw patch-level predictions. AvgPool_baseline, per-slide AUROC obtained by averaging patch-level predicted probabilities. Prop_baseline, per-slide AUROC obtained by calculating the proportion of positively predicted patches per slide. AMIL, attention-based multiple instance learning. CTMIL, customized transformer-based multiple instance learning. IRV2_*, models using features extracted by the InceptionResNetV2 backbone. RN50_*, models using features extracted by the ResNet50 backbone. UNI_*, models using features extracted by the UNI backbone. Ensemble_CTMIL: Averaged slide-level predictions from the three CTMIL variants.
On the validation set, all CTMIL-based models (Fig. 3J) surpassed their AMIL counterparts (Fig. 3F) and non-attention-based baseline models (Fig. 3B). For instance, UNI_CTMIL achieved the highest AUROC (0.8640) among all CTMIL variants, outperforming UNI_AMIL (0.7514); likewise, ResNet50_CTMIL (0.7023) and IRV2_CTMIL (0.7187) exceeded their AMIL counterparts (0.6944 and 0.6318, respectively). Baseline models showed markedly lower performance, with AUROCs of 0.5964 (Patch_baseline), 0.6834 (AvgPool_baseline), and 0.6890 (Prop_baseline). In terms of AUPRC, UNI_CTMIL again led among CTMIL variants (0.7539, Supplementary Fig. 1B), while Ensemble_CTMIL achieved the highest overall performance (AUROC = 0.8768, AUPRC = 0.7824).
On the XY test set, CTMIL-based models maintained superior performance compared to their AMIL counterparts and baseline models (Fig. 3K, G, C). Among all CTMIL variants, UNI_CTMIL again achieved the highest AUROC (0.8216), exceeding UNI_AMIL (0.7830); IRV2_CTMIL and ResNet50_CTMIL achieved AUROCs of 0.8069 and 0.7886, both surpassing their AMIL counterparts (0.5970 and 0.7049). Among baseline models, the best AUROC (0.7206) was observed in the Prop_baseline model. AUPRC values followed a similar pattern, with UNI_CTMIL reaching 0.7969, the highest among the CTMIL variants. The ensemble model further improved results (AUROC = 0.8407, AUPRC = 0.8400), in line with validation set trends.
On the RM cohort, CTMIL-based models again outperformed all corresponding AMIL counterparts and baseline models (Fig. 3L, H, D). UNI_CTMIL attained the best individual performance among CTMIL variants, with an AUROC of 0.8090, exceeding UNI_AMIL (0.6697). ResNet50_CTMIL and IRV2_CTMIL reached AUROCs of 0.7349 and 0.6934, compared to 0.6096 and 0.6000 for their AMIL counterparts. Baseline models performed consistently worse, with the highest AUROC at 0.6553 (AvgPool_baseline). Ensemble_CTMIL slightly improved the results (AUROC = 0.8208, AUPRC = 0.6994), maintaining the performance trend observed across datasets.
In summary, CTMIL-based models consistently outperformed their AMIL counterparts and non-attention-based baseline models. Among all evaluated variants, UNI_CTMIL demonstrated the strongest and most consistent predictive performance, underscoring the advantage of integrating a semantically enriched, self-supervised feature extractor (UNI) with a transformer-based MIL framework for effective and generalizable LNM prediction.
Ablation analysis of CTMIL architectural components
To elucidate the individual contributions of key architectural components within CTMIL, we conducted a stepwise ablation analysis using the best-performing variant, the UNI_CTMIL model. Starting from the full ensemble configuration, we sequentially ablated ensemble aggregation, auxiliary patch-level supervision, and transformer depth (from six to two layers), with the fully reduced model corresponding to the original TransMIL (Supplementary Fig. 2E–H). Performance comparisons were conducted across the validation set and two external test cohorts (Supplementary Fig. 2).
On the validation set, performance declined progressively with the removal of key components: Ensemble_CTMIL yielded the highest AUROC (0.8768), followed by UNI_CTMIL (0.8640) (Fig. 3J), the model without auxiliary supervision (0.7172) (Supplementary Fig. 2B), and the model based on the original TransMIL framework (0.7258) (Supplementary Fig. 2F). On the XY test cohort, the Ensemble_CTMIL achieved the best AUROC (0.8407), followed by the UNI_CTMIL (0.8216). The variant without auxiliary supervision slightly improved AUROC (0.8371) (Supplementary Fig. 2C). The TransMIL-based model again showed the weakest performance, with an AUROC of 0.7850 (Supplementary Fig. 2G). On the RM cohort, a partial reversal of this trend occurred: the model without auxiliary supervision achieved the highest AUROC (0.8338) (Supplementary Fig. 2D), slightly outperforming both UNI_CTMIL (0.8090) and Ensemble_CTMIL (0.8208). The original TransMIL remained the weakest performer (0.7353) (Supplementary Fig. 2H). Notably, similar trends were observed in IRV2_CTMIL and ResNet50_CTMIL, both of which achieved higher AUROC values than their TransMIL-based counterparts in the XY and RM cohorts (0.6934 vs. 0.4966 for IRV2_CTMIL; 0.7349 vs. 0.6929 for ResNet50_CTMIL).
Taken together, these results confirm the additive value of auxiliary supervision and deeper transformer depth in enhancing predictive performance. UNI_CTMIL consistently outperformed the original TransMIL architecture across all evaluation settings.
Patch-level interpretability and morphological assessment
To understand how the model distributes predictive focus across tumor regions, we visualized four representative WSIs (from two LNM-positive and two LNM-negative cases) based on prediction results of the UNI_CTMIL model, the top-performing individual model among the three CTMIL variants, and the primary contributor to the ensemble prediction (Fig. 4). Each row displays the original WSI, the tumor region delineated by the tumor patch detection model, an attention heatmap derived from the last-layer transformer attention scores, and the classification score heatmap obtained by feeding each patch independently through the trained UNI_CTMIL model. Since both model training and inference were confined to the automatically annotated tumor regions, attention and score maps were generated exclusively within these areas. Classification score maps identified patches that, when assessed individually, contributed strongly to a positive LNM prediction, reflecting regions with high local discriminative confidence. Notably, regions with high attention scores frequently overlapped with those showing high classification scores in LNM cases (Fig. 4A, B), indicating alignment between the model’s global attention focus and local predictive confidence. However, some patches exhibited high classification scores but received low attention, and vice versa. These discrepancies underscore the complementary roles of attention scores, which reflect the model’s internal decision focus, and classification scores, which capture patch-level predictive strength in isolation. In LNM-negative cases (Fig. 4C, D), classification scores across tumor regions were generally low, supporting the model’s negative slide-level predictions. While a small number of patches showed mismatches, such as concurrently high attention and high classification scores, the majority of high-attention regions corresponded to low classification scores.
A, B LNM-positive cases, C, D LNM-negative cases. From left to right, each column shows the original WSI, tumor regions detected by the tumor patch detection model, the attention heatmap (final-layer transformer attention scores), and the classification heatmap (per-patch prediction probabilities from the trained model). Attention scores are normalized within each WSI. Classification scores are output probabilities ranging from 0 to 1, with higher values indicating greater likelihood of LNM.
To explore the histopathological correlates of the model-derived predictions, we analyzed 300 patches from LNM-positive and LNM-negative patients, selected based on attention and classification scores as detailed in the Methods section (Supplementary Data 2). Examples from both groups are shown in Fig. 5A. Chi-square test revealed that patches from the LNM-positive group more frequently exhibited high tumor cell content (>60%) and poorly differentiated morphology, with statistically significant differences compared to the LNM-negative group (P < 0.001) (Fig. 5B). Regarding growth patterns, acinar, micropapillary, and solid morphologies were significantly enriched in the LNM-positive group (P < 0.001). A significantly higher proportion of mucinous-containing patches was observed in the LNM-positive group compared to the LNM-negative group (P < 0.001; Fig. 5C). In addition, granulocyte- or plasma cell-containing patches were more frequently observed in the LNM-positive group (P < 0.05), suggesting a potential involvement of these immune cells in metastasis progression, consistent with previous findings40,41,42. To further explore co-occurring histological traits, we constructed a Sankey diagram illustrating the joint distribution of the most informative features, with flow paths color-coded by group (LNM-positive vs. LNM-negative) (Fig. 5D). Patches from LNM-positive patients predominantly exhibited a feature path characterized by solid/acinar/micropapillary growth patterns, poorly differentiated morphology, mucin positivity, and high tumor cell content. In contrast, patches from LNM-negative patients more often showed papillary or lepidic patterns, moderate or well-differentiated morphology, mucin negativity, and lower tumor cell ratios, and were more likely to contain macrophages.
A representative tumor patches from LNM-positive (left) and LNM-negative (right) groups, selected based on combinations of high attention with high or low classification scores. B bar plots showing the number of patches exhibiting each histopathological feature. C radar plot showing the proportion of patches positive for each microenvironmental component. D Sankey diagram illustrating co-occurrence patterns of key histopathological features in the two groups. nLNM, LNM-negative. None (not evaluable), insufficient tumor cells or representative architecture within a patch to assess growth pattern or differentiation. *, P < 0.05; **, P < 0.01; ***, P < 0.001.
Discussion
The study introduces CTMIL, a customized transformer-based multiple-instance learning framework, designed for predicting LNM status from primary WSIs of LUAD. By incorporating enhanced spatial encoding, deeper transformer layers, and patch-level auxiliary supervision, CTMIL-based models, especially UNI_CTMIL, demonstrate superior performance and robust generalizability across independent multicenter cohorts, representing a notable advancement over TransMIL and AMIL frameworks.
Most existing efforts for LNM prediction have focused on radiological or molecular modalities, with limited attention to the histopathological evaluation of the primary tumor. Our study fills this gap by directly leveraging routine H&E-stained WSIs to predict LNM in LUAD, representing a shift toward histopathology-based inference of LNM status. Until recently, few studies had attempted to infer LNM status solely from primary tumor morphology. One proof-of-concept study in LUSC demonstrated that weakly supervised deep learning applied to H&E-stained WSIs could achieve modest predictive performance using a MIL framework24. Similar efforts in cervical cancer, bladder cancer, and colorectal cancer have also shown that primary tumor histology encodes predictive signals relevant to nodal involvement. However, prior pathology-based models predominantly relied on CNN-based MIL architectures and conventional attention pooling mechanisms, which are limited in their ability to capture the spatial and morphological heterogeneity inherent in solid tumors. Our approach introduces two key innovations. First, to our knowledge, it is the first multicenter study to investigate WSI-driven prediction of LNM in LUAD. Second, CTMIL replaces standard attention pooling with multi-head self-attention, allowing the model to capture long-range spatial dependencies and complex intratumoral patterns more effectively. Compared to earlier AMIL frameworks, CTMIL enables more expressive aggregation of patch-level information across entire WSIs. Through this architecture, our study advances beyond previous proof-of-concept efforts and provides a potentially generalizable framework for LNM prediction grounded in histopathology.
TransMIL is a transformer-based architecture specifically adapted for WSI analysis33, addressing the limitations of conventional instance-level MIL by modeling inter-patch dependencies and long-range contextual relationships. In recent years, TransMIL and its derivatives have shown promising performance across various computational pathology tasks, including classification37,43,44,45 and biomarker prediction46. However, transformer-based methods have been scarcely applied to the LNM prediction task. To address this gap, we proposed CTMIL, an enhanced variant of the TransMIL framework, incorporating two key architectural innovations. First, we increase the depth of the transformer encoder from two to six layers, enhancing its ability to integrate global contextual features across spatially distant tumor regions, a key factor in detecting subtle morphological cues associated with metastatic potential. Second, we incorporate auxiliary patch-level supervision, enabling the model to jointly learn from both global slide-level and localized patch-level signals. This dual-objective design facilitates fine-grained discriminative learning and encourages attention to histologically informative regions. These enhancements lead to consistent gains in predictive performance across validation and external test cohorts. When paired with the UNI feature extractor, a self-supervised backbone trained to capture high-level semantic representations47, the UNI_CTMIL model achieved the highest performance, indicating strong synergy between expressive feature encoding and transformer-based spatial modeling. Collectively, our findings highlight the promise of transformer-based MIL frameworks in advancing histopathology-informed prediction tasks in LUAD.
Histopathology offers a complementary perspective, providing direct visualization of tumor architecture and morphological features associated with metastatic potential. For example, micropapillary and solid growth patterns in LUAD have been associated with increased aggressiveness and a higher likelihood of lymph node involvement48, yet these features can be subtle and are often undetectable through imaging alone. By applying deep learning to high-resolution whole-slide histology, our approach systematically captures these complex morphologic signals. The attention mechanism in CTMIL enhances interpretability by highlighting regions most influential to model predictions. Qualitative review revealed that the model consistently attended to regions aligned with known aggressive phenotypes, such as acinar, micropapillary, and solid growth patterns23, mucinous components49, poorly differentiated morphology, and high tumor cell density. These findings underscore the model’s ability to extract morphologic correlates of nodal spread. Notably, this capability may be especially useful in clinical settings where lymph node dissection is incomplete or assessment yields equivocal results. In such cases, histology-based inference of metastatic risk could serve as a valuable adjunct to standard staging. This may facilitate earlier identification of high-risk patients and support treatment decision-making.
Despite its promise, our study has several limitations that must be acknowledged. First, although CTMIL-based models demonstrated strong performance on internal and external cohorts, particularly the UNI_CTMIL model, the small numbers in some subcategories, especially N3, limit the stability of per-class (N1/N2/N3) prediction. Because all cohorts had complete nodal evaluation, generalizability to pNx remains uncertain; thus, the present study should be considered proof of concept. Further validation in larger, prospective, stage-stratified, and multi-institutional datasets, including pNx and biopsy-only settings, is warranted. Second, the retrospective nature of our study constrains its direct clinical applicability. Although the model predicted LNM from primary tumor histology, the ground truth was derived from postoperative pathological nodal evaluation. As such, the current setup offers proof-of-concept validation but does not fully emulate a preoperative diagnostic scenario, where only a limited biopsy or partial resection may be available. The applicability of CTMIL to small biopsy specimens remains untested, and its ability to infer LNM risk from such limited tissue is an open question. Third, although high-attention and high-score regions provide visual cues, establishing completely consistent correspondence with specific, human-interpretable histopathological or molecular features remains challenging. Fourth, each case in our cohorts was represented by one pathologist-selected, quality-controlled WSI, which may not capture intra-case variability. Future work will evaluate multi-WSI aggregation strategies for case-level inference and assess robustness when multiple slides per case are available. Finally, as the model operates exclusively on histopathological images, it does not account for relevant factors such as tumor size, radiologic findings, and genomic alterations. While this single-modality design enhances interpretability and scalability, it may limit predictive power and underscores the value of future multimodal integration. Nonetheless, our findings demonstrate the potential of CTMIL as a pathology-based framework for LNM prediction in LUAD, and more broadly highlight the promise of transformer-based MIL architectures in computational pathology. Future research may extend this approach to more complex endpoints, including immunotherapy response, recurrence risk, and long-term prognosis.
In conclusion, we present CTMIL, a customized transformer-based multiple-instance learning framework that achieves robust performance in predicting LNM from LUAD histopathology. This multicenter study demonstrates that CTMIL consistently outperforms conventional TransMIL and attention-based methods through the integration of deeper transformer encoders, spatial encoding, and auxiliary patch-level supervision. Visual and quantitative analyses confirm the interpretability of the best-performing UNI_CTMIL model, which reliably attends to histological regions associated with metastatic potential. Future research should consider integrating CTMIL with complementary data modalities, such as radiologic imaging and genomic profiling, which may further enhance predictive precision and clinical applicability for metastasis assessment and treatment response prediction.
Methods
Study cohorts
In this study, three LUAD cohorts were included: the Cancer Genome Atlas (TCGA) cohort, accessed via the Genomic Data Commons (GDC; https://portal.gdc.cancer.gov/); the RM cohort from Renmin Hospital of Wuhan University, China; and the XY cohort from the Third Xiangya Hospital of Central South University, China. Institutional review board approvals were obtained for both the RM and XY cohorts, and the requirement for informed consent was waived due to the retrospective nature of the study. All selected cases had histologically confirmed LUAD with clearly documented LNM status, categorized as pathological N0 (no metastasis) or N1-N3 (confirmed metastasis). Corresponding hematoxylin and eosin (H&E)-stained WSIs were available for all cases. WSIs exhibiting substantial artifacts, such as staining defects, air bubbles, tissue folds, or extraneous markings, were excluded to ensure image quality consistency across cohorts. WSIs from both flash-frozen and formalin-fixed paraffin-embedded (FFPE) slides were used to develop the tumor patch detection model, as most normal samples were only available in flash-frozen format. In contrast, only FFPE-derived WSIs were included for LNM prediction task due to their superior morphological preservation.
In the TCGA cohort, patients who had received neoadjuvant therapy were excluded to avoid therapy-induced alterations in tumor histology. When multiple WSIs were available for a patient, three board-certified pathologists independently reviewed the slides and reached consensus on the most representative image, prioritizing abundant tumor regions, broad tissue coverage, and minimal necrosis. Transcriptomic data associated with the selected slides were also downloaded from the GDC.
For the RM and XY cohorts, 164 and 149 high-quality WSIs were retrospectively and randomly collected from patients who underwent surgical resection between January 2021 and March 2025. Inclusion criteria were: (1) availability of diagnostic H&E slides with sufficient tumor area; and (2) complete clinical documentation of pathological LNM status retrievable from electronic medical records. Patients who had received neoadjuvant therapy were not included. All slides were scanned at 20× magnification and saved in SVS format.
Overview of workflow
Figure 1 presents a computational workflow for LNM prediction in LUAD using primary WSIs. The analysis pipeline consisted of three main stages: (1) development of a tumor patch detection model to identify tumor patches within WSIs (Module 1); (2) extraction of patch-level features from the detected tumor patches using three pre-trained networks (ResNet50, InceptionResNetV2, and UNI), each yielding representations of different dimensionality (Module 2); (3) training of multiple LNM prediction models within the established CTMIL framework, with each model trained on features from a specific backbone network (Modules 3 and LNM prediction part). An AMIL model was also implemented as a comparative baseline. Notably, in contrast to the methodology of Wagner et al.46, which utilized all available patches within each WSI, our approach restricted analysis to tumor-derived patches to mitigate potential confounding effects from non-tumor regions.
To ensure that downstream analysis focused on diagnostically meaningful regions, a tumor patch detection model was developed to automatically delineate tumor areas while excluding irrelevant regions such as normal tissue, background, and necrosis. This automated approach effectively reduces reliance on manual annotation, improves preprocessing consistency across samples, and enhances computational efficiency and scalability by focusing analysis on tumor regions.
WSIs processing
WSI reprocessing followed protocols consistent with our previous work50, with one key modification: all patches were standardized to 512 × 512 pixels. Each WSI, scanned at 20× magnification, was segmented into non-overlapping RGB image patches using the OpenSlide Python library. To exclude background and ensure sufficient tissue content, patches with a mean RGB intensity above 230 and a standard deviation below 15 were discarded. Remaining patches were subjected to stain normalization using Macenko’s method51 to reduce inter-slide staining variability.
Dataset preparation for tumor patch detection
Patch-level annotation was performed following the criteria described by Chen et al.52, in which each 512 × 512-pixel patch was labeled as tumor-positive if it contained at least five histologically identifiable tumor cells; otherwise, it was labeled tumor-negative. To construct a balanced dataset of approximately 400,000 annotated patches, three experienced pathologists randomly selected 460 high-quality WSIs, comprising 144 tumor and 316 normal slides, from the full TCGA LUAD cohort (n = 1133; 541 tumor and 592 normal slides; Supplementary Table 1). The numbers of tumor and normal slides were not pre-specified; instead, they were iteratively adjusted based on post-quality control (QC) patch yields to achieve approximately 1:1 patch-level class balance and to maximize the histologic diversity of negative patches. These selected WSIs yielded a total of 591,333 patches, from which 377,790 artifact-free patches with clear cellular morphology were curated (Supplementary Table 1 and Data 1). The final balanced dataset consisted of 189,038 tumor-positive and 188,872 tumor-negative patches, randomly divided into training, validation, and test sets at an 8:1:1 ratio. Importantly, all patches from a single sample were exclusively assigned to the same set to prevent data leakage.
Development, evaluation, and selection of the tumor patch detection model
Five pre-trained deep neural networks (ResNet5053, InceptionV354, InceptionResNetV255, UNI47, Virchow256) were trained for binary patch-level tumor classification and systematically evaluated. All models were initialized with publicly available pre-trained weights and adapted by appending a two-class output layer. To facilitate transfer learning, the feature extraction layers were either frozen (UNI and Virchow2) or fine-tuned (ResNet50, InceptionV3, and InceptionResNetV2), depending on the model. Input patches were 512 × 512 pixels by default, and only resized to 518 × 518 for compatibility with the Virchow2 model. Training patches were augmented through random horizontal flips, rotations, and color jitter to enhance model generalizability. Model optimization employed the Adam optimizer with a cross-entropy loss function, along with adaptive learning rate schedules: one-cycle learning rate policy for UNI and Virchow2, and plateau-based reduction for the other models. Early stopping based on validation loss was used to prevent overfitting. The optimal classification threshold was determined by maximizing the Youden index on the validation set. Final model performance was assessed on the test set using standard metrics: accuracy, precision, recall, specificity, F1-score, and the area under the receiver operating characteristic curve (AUC). The best-performing model was selected based on a comprehensive comparison of all five models across these metrics.
Tumor patch feature extraction
The best-performing classification model (InceptionResNetV2) was first applied to identify tumor-positive patches from WSIs. From these patches, three deep neural network backbones (InceptionResNetV2, ResNet50, and UNI) were independently used to extract feature embeddings, producing patch-level representations of 1536, 2048, and 1024 dimensions, respectively. All networks were initialized with publicly available pre-trained weights, and their classification heads were replaced with identity layers to enable feature extraction from the penultimate layer. Each tumor patch was processed in inference mode through the modified networks, generating three distinct sets of feature vectors. This strategy enabled systematic comparison of patch-level representations derived from different architectures, each capturing unique morphological and contextual features. The resulting backbone-specific embeddings were subsequently used as inputs for downstream LNM prediction models.
Customized transformer-based MIL architecture
Building on the original TransMIL framework33, we adapted our CTMIL architecture with two key modifications aimed at enhancing spatial representation and improving instance-level discriminability. First, we expanded the transformer encoder depth from two to six layers to enhance modeling of global contextual relationships. Second, we introduced auxiliary supervision at the patch level to reinforce fine-grained learning and improve the discriminative quality of patch representations.
For each WSI, the extracted patch-level feature vectors were first projected into a shared 512-dimensional latent space via a linear transformation. In parallel, the corresponding two-dimensional spatial coordinates (x, y) of each patch were normalized and mapped into 512-dimensional positional embeddings using a separate linear layer. These embeddings were then fused through element-wise addition to jointly encode morphological and spatial context. To form the input sequence for the transformer module, a learnable 512-dimensional classification token (CLS token) was prepended to the fused patch embeddings. As a result, the model input for each slide consisted of (n + 1) tokens, each of dimensionality 512, where n is the number of tumor patches per WSI (Module 3 in Fig. 1).
The complete (n + 1) × 512 sequence was passed through a stack of six transformer encoder layers. Each layer followed a standard composition of operations designed to capture complex intra-slide relationships: (1) pre-layer normalization was applied to the input sequence; (2) multi-head self-attention was performed using eight attention heads with a dropout rate of 0.1. This mechanism enabled each token to attend to spatially and semantically relevant regions across the WSI. Formally, for the normalized input XER(n+1)×512, the attention output was computed as:
where LayerNorm(.) denotes layer normalization and MultiHeadSelfAttention(.) follows the formulation proposed by Vaswani et al.57 and was implemented via torch.nn.MultiheadAttention in Python. The attention output Z was added element-wise to the input X as a residual connection, followed by dropout (rate = 0.1), yielding:
(3) The updated sequence X′ was passed through a second layer normalization and then a position-wise feed-forward network (FFN) composed of two fully connected layers. The first layer expanded the hidden dimension from 512 to 2048, followed by a ReLU activation, and the second projected it back to 512 dimensions (implemented via torch.nn.Linear and torch.nn.ReLU in Python). This transformation can be formulated as:
The FFN output was added to the input via a residual connection and regularized with dropout (rate = 0.05), yielding:
This sequence of operations was repeated across six transformer encoder layers, with each layer maintaining the shape of the sequence at (\(n\)+1) × 512. Stacking multiple such layers enabled the model to progressively capture both local interactions between patches and global slide-level dependencies, with the CLS token aggregating the overall contextual representation of the WSI.
Unlike the original TransMIL framework, CTMIL incorporated an auxiliary supervision mechanism inspired by the CLAM methodology58. During training, the model was optimized using a dual-objective scheme that combined global and patch-level predictive signals. The global slide-level prediction was obtained by feeding the CLS token embedding through a dedicated classification head. Simultaneously, each patch token embedding was passed through a separate classifier to produce patch-level logits, which were aggregated into an auxiliary slide-level prediction using attention weights corresponding to the CLS token’s focus on each patch. This auxiliary path encouraged the model to focus more effectively on discriminative regions, enhancing both feature expressiveness and classification accuracy. Accordingly, model optimization was guided by a joint loss function that integrated both the global and attention-weighted patch-level predictions, structured as follows:
where BCE represents binary cross-entropy, ZCLS is the logit derived from the CLS token, Zi is the logit corresponding to the ith patch, the attention weight αi represents the normalized attention weight assigned to the ith patch by the CLS token, derived from the final transformer layer and constrained such that \({\sum }_{i=1}^{N}{\alpha }_{i}=1\). N is the number of patches, and Y is the binary ground truth for the slide.
Model parameters were optimized using the Adam algorithm, with early stopping based on validation AUC to prevent overfitting. The learning rate was adaptively adjusted using a plateau-based scheduler, and class imbalance was addressed by applying class weights to the loss function. The model checkpoint with the highest validation AUC was selected for final evaluation. During inference, only the global slide-level prediction from the CLS token was used. The predicted probability of a positive class was obtained by applying a sigmoid function to the CLS logit ZCLS:
where σ (⋅) denotes the sigmoid function.
Attention-based MIL Architecture
An AMIL architecture32 was implemented as a comparative baseline to evaluate against the performance of CTMIL (Module 3 in Fig. 1). AMIL employs a simple attention mechanism to aggregate patch-level features into a single slide-level representation for classification. Given a set of patch-level feature vectors hi for i = 1,…, N, where N is the number of patches in a slide, the attention score for each patch is computed using a two-layer fully connected network with a hyperbolic tangent activation:
These scores are normalized via a softmax function with a learnable temperature parameter t, which controls the sharpness of the attention distribution:
The final slide-level feature representation is obtained by an attention-weighted sum of patch features. This aggregated feature is passed through a fully connected layer and a sigmoid activation to yield the final prediction probability P for each slide:
where W and b denote the weights and bias of the final fully connected layer, αi denotes the normalized attention score assigned to the ith patch, and σ (⋅) denotes the sigmoid function.
LNM prediction models
Three primary LUAD cohorts (TCGA, XY, and RM cohorts) were employed to develop and evaluate models for predicting LNM. All WSIs underwent a standardized processing pipeline consisting of tumor patch detection and subsequent extraction of patch-level features (Module 2 in Fig. 1). To ensure sufficient tumor patch content for model training, TCGA cases with fewer than 100 detected tumor patches were excluded from further analysis. The remaining 400 TCGA cases were randomly allocated to training (80%) and validation (20%) sets, while the XY (n = 149) and RM (n = 164) cohorts were reserved as independent test sets (Fig. 2). Detailed case counts by nodal stage for each cohort, stratified by LNM status, are summarized in Supplementary Table 2. For each WSI, three distinct types of patch-level feature vectors were extracted as previously described, based on ResNet50, InceptionResNetV2, and UNI. Models based on the CTMIL architecture (as illustrated in Modules 3 and 4 of Fig. 1) were independently trained on each of the three feature vector types, resulting in three model variants: (1) RN50_CTMIL model, (2) IRV2_CTMIL model, and (3) UNI_CTMIL model. A fourth model (Ensemble_CTMIL) was also evaluated, in which slide-level prediction probabilities from the three CTMIL variants were averaged. For comparison, three additional models based on the AMIL architecture were trained using the same feature sets: (1) RN50_AMIL model, (2) IRV2_AMIL model, and (3) UNI_AMIL model. Model performance was primarily evaluated using the area under the receiver operating characteristic curve (AUROC), with the area under the precision-recall curve (AUPRC) included as a complementary metric to account for class imbalance. In addition, the performance of the CTMIL framework was also compared with the original TransMIL architecture based on these two metrics.
As an additional baseline, we trained a patch-level classification model using InceptionResNetV2, which was selected based on its superior performance in the tumor patch detection task. In this approach, all tumor-positive patches from LNM-positive slides were labeled as “LNM”, and those from LNM-negative slides as “non-LNM”, thereby forming a binary patch-level dataset. The TCGA cohort was split at an 8:2 ratio, with all patches from each WSI assigned exclusively to either the training or validation set to prevent data leakage. The optimal decision threshold was determined by maximizing the Youden index (TPR-FPR) on the validation set. Model performance was subsequently evaluated from three complementary perspectives: (1) raw patch-level prediction probabilities (Patch_baseline), (2) slide-level scores obtained by averaging patch-level predictions within each slide (AvgPool_baseline), and (3) the proportion of patches classified as LNM within each slide (Prop_baseline).
Visualization and explainability
We applied the attention mechanism to quantify the contribution of individual patches to slide-level prediction by focusing on the attention weights from the CLS token to each patch token. Specifically, we extracted the last-layer attention matrix and interpreted the CLS-to-patch weights as indicators of each patch’s relative contribution to the final prediction. In parallel, patch-level classification scores were obtained by feeding each patch individually through the trained model and recording the resulting slide-level probabilities. These scores represent the model’s estimated likelihood of LNM for each patch. Although not used for supervision during training, they provided post hoc insights into the spatial distribution of predictive relevance. Both attention scores and classification scores were mapped back to the spatial coordinates of the corresponding patches and visualized as heatmaps, highlighting regions that were either strongly attended to or highly predictive of LNM.
To explore the morphological basis underlying the model’s predictions of LNM, we conducted a focused patch-level analysis on 60 patients (30 LNM-positive and 30 LNM-negative, with 10 cases from each of the three cohorts) whose slide-level predictions matched the ground truth. For each case, we selected five patches from two informative categories: (1) high-score/high-attention patches in LNM-positive cases, those with the top-5 attention scores among the top 100 classification-scoring patches, and (2) low-score/high-attention patches in LNM-negative cases, those with the top-5 attention scores among the bottom 100 classification-scoring patches. All selected patches were blindly and independently reviewed by two pathologists to assess key histological features, including tumor cell proportion, degree of differentiation, growth pattern, and stromal ratio, as well as the presence of mucin (intracytoplasmic and extracellular mucin) and non-neoplastic cellular components. Differences in feature distributions between the two groups were assessed using the chi-square test.
All computational analyses were performed using Python (version 3.9.3) and R (version 4.4.2) and run on the NVIDIA A40 GPU.
Data availability
WSIs, clinical and transcriptomics data of the TCGA-LUAD cohort can be downloaded via the GDC (https://portal.gdc.cancer.gov/). Data from the XY and RM cohorts are available from the corresponding authors upon reasonable request.
Code availability
All codes related to the training of tumor patch detection and CTMIL-based models are available via the GitHub repository: https://github.com/huibozh/CTMIL.
References
Lao, S. et al. Prognostic patterns in invasion lymph nodes of lung adenocarcinoma reveal distinct tumor microenvironments. NPJ Precis. Oncol. 8, 164 (2024).
Asamura, H. et al. IASLC lung cancer staging project: the new database to inform revisions in the Ninth Edition of the TNM classification of lung cancer. J. Thorac. Oncol. 18, 564–575 (2023).
Son, J. W. et al. Validation of IASLC 9th edition TNM classification for lung cancer: focus on N descriptor. BMC Cancer 24, 1460 (2024).
Denisenko, T. V., Budkevich, I. N. & Zhivotovsky, B. Cell death-based treatment of lung adenocarcinoma. Cell Death Dis. 9, 117 (2018).
Hua, X. et al. Genetic and epigenetic intratumor heterogeneity impacts prognosis of lung adenocarcinoma. Nat. Commun. 11, 2459 (2020).
Tan, K. S. et al. Novel insights into the international association for the study of lung cancer grading system for lung adenocarcinoma. Mod. Pathol. 37, 100520 (2024).
Bille, A., Woo, K. M., Ahmad, U., Rizk, N. P. & Jones, D. R. Incidence of occult pN2 disease following resection and mediastinal lymph node dissection in clinical stage I lung cancer patients. Eur. J. Cardiothorac. Surg. 51, 674–679 (2017).
Ray, M. A. et al. Comparative effectiveness of a lymph node collection kit versus heightened awareness on lung cancer surgery quality and outcomes. J. Thorac. Oncol. 16, 774–783 (2021).
David, E. A. et al. Does lymph node count influence survival in surgically resected non-small cell lung cancer. Ann. Thorac. Surg. 103, 226–235 (2017).
Cackowski, M. M. et al. The absence of lymph nodes removed (pNx status) impacts survival in patients with lung cancer treated surgically. Surg. Oncol. 48, 101941 (2023).
Zhong, Y. et al. PET/CT based cross-modal deep learning signature to predict occult nodal metastasis in lung cancer. Nat. Commun. 14, 7513 (2023).
Zhao, X. et al. A cross-modal 3D deep learning for accurate lymph node metastasis prediction in clinical stage T1 lung adenocarcinoma. Lung Cancer 145, 10–17 (2020).
Tian, W. et al. Predicting occult lymph node metastasis in solid-predominantly invasive lung adenocarcinoma across multiple centers using radiomics-deep learning fusion model. Cancer Imaging 24, 8 (2024).
Ouyang, M. L. et al. Prediction of occult lymph node metastasis using tumor-to-blood standardized uptake ratio and metabolic parameters in clinical N0 lung adenocarcinoma. Clin. Nucl. Med. 43, 715–720 (2018).
Ma, X., Xia, L., Chen, J., Wan, W. & Zhou, W. Development and validation of a deep learning signature for predicting lymph node metastasis in lung adenocarcinoma: comparison with radiomics signature and clinical-semantic model. Eur. Radio. 33, 1949–1962 (2023).
Li, F., Zhai, S., Fu, L., Yang, L. & Mao, Y. Nomograms for intraoperative prediction of lymph node metastasis in clinical stage IA lung adenocarcinoma. Cancer Med. 12, 14360–14374 (2023).
Zhu, X., Luo, H. & Xu, Y. Transcriptome analysis reveals an important candidate gene involved in both nodal metastasis and prognosis in lung adenocarcinoma. Cell Biosci. 9, 92 (2019).
Dong, B., Wu, C., Huang, L. & Qi, Y. Macrophage-Related SPP1 as a potential biomarker for early lymph node metastasis in lung adenocarcinoma. Front. Cell Dev. Biol. 9, 739358 (2021).
Gallina, F. T. et al. ALK rearrangement is an independent predictive factor of unexpected nodal metastasis after surgery in early stage, clinical node negative lung adenocarcinoma. Lung Cancer 180, 107215 (2023).
Wang, K. et al. Genomics analysis and nomogram risk prediction of occult lymph node metastasis in non-predominant micropapillary component of lung adenocarcinoma measuring </= 3 cm. Front. Oncol. 12, 945997 (2022).
Park, D. et al. Multimodal AI model for preoperative prediction of axillary lymph node metastasis in breast cancer using whole slide images. NPJ Precis. Oncol. 9, 131 (2025).
Gao, F. et al. Deep learning-based pathological prediction of lymph node metastasis for patient with renal cell carcinoma from primary whole slide images. J. Transl. Med. 22, 568 (2024).
Dai, C. et al. Relationship of lymph node micrometastasis and micropapillary component and their joint influence on prognosis of patients with stage i lung adenocarcinoma. Am. J. Surg. Pathol. 41, 1212–1220 (2017).
Xia, L. et al. Lymph node metastasis prediction from in situ lung squamous cell carcinoma histopathology images using deep learning. Lab Investig. 105, 102187 (2025).
Song, J. H. et al. Prediction of lymph node metastasis in T1 colorectal cancer using artificial intelligence with hematoxylin and eosin-stained whole-slide-images of endoscopic and surgical resection specimens. Cancers 16, https://doi.org/10.3390/cancers16101900 (2024).
Song, J. H., Hong, Y., Kim, E. R., Kim, S. H. & Sohn, I. Utility of artificial intelligence with deep learning of hematoxylin and eosin-stained whole slide images to predict lymph node metastasis in T1 colorectal cancer using endoscopically resected specimens; prediction of lymph node metastasis in T1 colorectal cancer. J. Gastroenterol. 57, 654–666 (2022).
Kiehl, L. et al. Deep learning can predict lymph node status directly from histology in colorectal cancer. Eur. J. Cancer 157, 464–473 (2021).
Guo, Q. et al. Predicting lymph node metastasis from primary cervical squamous cell carcinoma based on deep learning in histopathologic images. Mod. Pathol. 36, 100316 (2023).
Zheng, Q. et al. Predicting lymph node metastasis status from primary muscle-invasive bladder cancer histology slides using deep learning: a retrospective multicenter study. Cancers 15, https://doi.org/10.3390/cancers15113000 (2023).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Ilse, M., Tomczak, J. & Welling, M. Attention-based deep multiple instance learning. in International conference on machine learning. 2127–2136 (PMLR).
Shao, Z. et al. Transmil: transformer based correlated multiple instance learning for whole slide image classification. Adv. Neural Inf. Process. Syst. 34, 2136–2147 (2021).
Zhao, R. et al. MIST: multi-instance selective transformer for histopathological subtype prediction. Med. Image Anal. 97, 103251 (2024).
Wang, Z., Yu, L., Ding, X., Liao, X. & Wang, L. Lymph node metastasis prediction from whole slide images with transformer-guided multiinstance learning and knowledge transfer. IEEE Trans. Med. Imaging 41, 2777–2787 (2022).
Javed, S., Mahmood, A., Qaiser, T., Werghi, N. & Rajpoot, N. Unsupervised mutual transformer learning for multi-gigapixel Whole Slide Image classification. Med. Image Anal. 96, 103203 (2024).
Chikontwe, P. et al. FR-MIL: distribution re-calibration-based multiple instance learning with transformer for whole slide image classification. IEEE Trans. Med. Imaging 44, 409–421 (2025).
Jiang, R. et al. A transformer-based weakly supervised computational pathology method for clinical-grade diagnosis and molecular marker discovery of gliomas. Nat. Mach. Intell. 6, 876–891 (2024).
Li, Z., Jiang, Y., Lu, M., Li, R. & Xia, Y. Survival prediction via hierarchical multimodal co-attention transformer: a computational histology-radiology solution. IEEE Trans. Med. Imaging 42, 2678–2689 (2023).
Wang, S. S. et al. Tumor-infiltrating B cells: their role and application in anti-tumor immunity in lung cancer. Cell Mol. Immunol. 16, 6–18 (2019).
Shalapour, S. et al. Immunosuppressive plasma cells impede T-cell-dependent immunogenic chemotherapy. Nature 521, 94–98 (2015).
Huang, C. et al. Downregulation of tropomyosin 2 promotes the progression of lung adenocarcinoma by regulating neutrophil infiltration through neutrophil elastase. Cell Death Dis. 16, 264 (2025).
Wang, C. W. et al. Ensemble transformer-based multiple instance learning to predict pathological subtypes and tumor mutational burden from histopathological whole slide images of endometrial and colorectal cancer. Med. Image Anal. 99, 103372 (2025).
Chen, Y. et al. dMIL-transformer: multiple instance learning via integrating morphological and spatial information for lymph node metastasis classification. IEEE J. Biomed. Health Inf. 27, 4433–4443 (2023).
Cai, H. et al. MIST: multiple instance learning network based on Swin Transformer for whole slide image classification of colorectal adenomas. J. Pathol. 259, 125–135 (2023).
Wagner, S. J. et al. Transformer-based biomarker prediction from colorectal cancer histology: a large-scale multicentric study. Cancer Cell 41(1650–1661), e1654 (2023).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Xue, M. et al. The role of adenocarcinoma subtypes and immunohistochemistry in predicting lymph node metastasis in early invasive lung adenocarcinoma. BMC Cancer 24, 139 (2024).
Li, X. et al. Transmembrane mucins in lung adenocarcinoma: understanding of current molecular mechanisms and clinical applications. Cell Death Discov. 11, 163 (2025).
Zhang, H. et al. Prediction and analysis of tumor infiltrating lymphocytes across 28 cancers by TILScout using deep learning. NPJ Precis Oncol. 9, 76 (2025).
Macenko, M. et al. A method for normalizing histology slides for quantitative analysis. in IEEE International Symposium on Biomedical Imaging: From Nano to Macro. 1107–1110. (IEEE, 2009).
Chen, R. J. et al. Pan-cancer integrative histology-genomic analysis via multimodal deep learning. Cancer Cell 40, 865–878.e866 (2022).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. in IEEE Conference on Computer Vision and Pattern Recognition (CVPR). 770–778. (IEEE, 2016).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the Inception Architecture for Computer Vision. in IEEE Conference on Computer Vision and Pattern Recognition (CVPR). 2818–2826 (IEEE, 2016)
Szegedy, C., Ioffe, S., Vanhoucke, V. & Alemi, A. Inception-v4, inception--resnet and the impact of residual connections on learning. in Proc. AAAI conference on artificial intelligence. 4278–4284 (AAAI Press, 2017).
Vorontsov, E. et al. A foundation model for clinical-grade computational pathology and rare cancers detection. Nat. Med. 30, 2924–2935 (2024).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inform. Process. Syst. 30, 1–11 (2017).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Acknowledgements
This work was funded by the National Natural Science Foundation of China (grant No.82403850 and No.82203502) and the Cross-Innovation Talent project of Renmin Hospital of Wuhan University (JCRCGW-2022-002). We would like to express our sincere gratitude to the Departments of Pathology at Renmin Hospital of Wuhan University and the Third Xiangya Hospital of Central South University for their substantial assistance and valuable contributions to this study.
Author information
Authors and Affiliations
Contributions
H.Z. contributed to the conceptualization of the research, carried out data curation, performed formal analysis, developed the methodology, created the software, conducted visualization, and wrote the original draft of the manuscript. T.W. and J.F. participated in the investigation, contributed significantly to developing the methodology, and performed validation analyses. J.H. assisted with methodology development and data analysis. Y.L. and Q.S. provided resources, participated in conceptualizing the project, and supervised the research activities. B.X. oversaw the conceptualization and methodology development, managed project administration, secured funding, supervised the project, and reviewed and revised the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, H., Wang, T., Feng, J. et al. Customized transformer for lymph node metastasis prediction from lung adenocarcinoma histology in a multicentric study. npj Precis. Onc. 10, 16 (2026). https://doi.org/10.1038/s41698-025-01199-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01199-8