Abstract
Prostate cancer (PCa) ranks among the most prevalent cancers in men worldwide. Biochemical recurrence (BCR) presents a major clinical challenge in PCa management, with significant prognostic heterogeneity observed among patients post-recurrence. This study aimed to develop machine learning models for predicting both the diagnosis and prognosis of PCa patients. Using WGCNA, we initially identified 16 BCR-related target genes. Cluster analysis revealed these genes were significantly associated with PCa prognosis, drug sensitivity, and immune infiltration. We constructed a robust diagnostic model integrating multiple machine learning algorithms, demonstrating strong predictive capability for PCa. Furthermore, a BCR-related prognostic model built using the LASSO algorithm also yielded satisfactory performance. Among the differentially expressed BCR-associated prognostic genes, COMP emerged as a critical regulatory factor. Both in vitro and in vivo experiments confirmed COMP’s role in influencing PCa progression. Additionally, COMP demonstrates significant potential as a dual biomarker for both the diagnosis and recurrence prediction of PCa.
Similar content being viewed by others
Introduction
Prostate cancer (PCa) ranks as the most common malignant tumor in men and represents a major contributor to cancer-related deaths1,2. The primary treatments recommended for localized PCa include radical prostatectomy (RP) and radical radiotherapy (RT)3. Recent technological improvements have increased the effectiveness of both RP and RT; however, approximately 20–60% of individuals who undergo these radical treatments experience biochemical recurrence (BCR) within ten years4. BCR is generally characterized by a prostate-specific antigen (PSA) level greater than 0.2 ng/mL following radical surgery or exceeding 2 ng/mL above the lowest point after receiving radiotherapy5,6. The occurrence of BCR is linked to negative outcomes in PCa, such as local recurrence, distant metastasis, and increased mortality7,8. Consequently, even with progress in the early detection and management of PCa, BCR continues to pose a considerable challenge for healthcare providers9,10. Prompt prediction of BCR in PCa patients can help clinicians formulate individualized treatment strategies and adjust follow-up frequency based on the risk of recurrence. This method will aid in the early identification of disease progression or recurrence and encourage timely therapeutic interventions.
PSA testing is integral to the surveillance of PCa, especially for the early identification of BCR11,12. Studies suggest that changes in PSA levels can serve as preliminary signals of BCR, making consistent PSA testing a vital component for tracking the recurrence of PCa13,14. Nonetheless, it is important to recognize the limitations inherent in PSA testing. To begin with, the elevated false-positive rates associated with PSA screenings can lead to unnecessary additional tests and procedures, which in turn may exacerbate patients’ psychological stress and increase healthcare costs15. Furthermore, variations in PSA levels do not always have a direct correlation with cancer recurrence. For instance, after specific therapies—such as high-intensity focused ultrasound—PSA levels might increase sharply; this increase, however, does not invariably indicate a recurrence of cancer16. This occurrence, referred to as PSA bounce, can result in misinterpretation and excessive treatment17. Therefore, while PSA testing significantly contributes to monitoring PCa recurrence, its limitations also restrict its overall value.
In recent years, the rapid advancement of artificial intelligence has emerged as a pivotal factor in the biomedical field18,19,20,21,22. Gene expression-based diagnostic and therapeutic models have become essential tools in guiding personalized treatment for cancers, including PCa23. This study aims to systematically identify key genes and regulatory networks associated with BCR in PCa by integrating multi-omics data and machine learning algorithms. It seeks to construct a recurrence risk model with high predictive performance, further evaluate its clinical application value, and contribute to the advancement of precision medicine. By integrating various machine learning methods, we anticipate identifying key biomarkers associated with PCa diagnosis, prognosis, and their influence on the immune microenvironment. This research could provide potential targets for the development of personalized treatment strategies and precise management of PCa. Ultimately, this work not only seeks to enhance the prognostic assessment of PCa patients but may also offer a theoretical foundation for the development of novel treatment options.
Results
Identification of BCR related genes based on Weighted gene co-expression network analysis (WGCNA)
WGCNA is a systems biology method that describes gene association patterns across different samples. This approach can identify gene sets that change collaboratively and pinpoint candidate organisms based on the interconnectivity of these gene sets, as well as their association with phenotypes, marker genes, or therapeutic targets. In this study, we employed the WGCNA method to identify regulatory genes associated with BCR of PRAD, utilizing 248 samples from the GSE116918 data set. Initially, we screened and determined the optimal soft threshold, followed by the clustering analysis of the 248 samples within the GSE116918 data set (Fig. 1A–C). The GSE116918 sample was ultimately categorized into 11 stable modules, with the pink module exhibiting the highest correlation with BCR of PRAD. Furthermore, the genes within the pink module also demonstrated a significant correlation with BCR (Fig. 1D–F). This pink module comprises 162 genes, of which only 16 were found to be highly expressed in PRAD and associated with patient progression-free interval (PFI) (Fig. 1G, H). Notably, these 16 genes exhibited a positive correlation with one another in the TCGA-PRAD dataset (Fig. 1I). Finally, we validated the correlation of these genes with BCR in PRAD patients using the GSE116918 dataset. Our analysis revealed that the expression of these genes was significantly higher in the group with BCR compared to the group of patients without BCR (Fig. 1J).
A Plot scale independence. B Plot average connectivity. C Draw sample clusters. D Dividing the GSE116918 sample into 11 modules. E Module-phenotype correlation heat map. F Scatter plot of correlation between gene significance and module membership. G Heatmap of BCR-related gene expression in TCGA-PRAD. H Prognostic forest plot of BCR-related genes in TCGA-PRAD. I Heatmap of correlation of biochemical relapse-related genes in TCGA-PRAD. J Differential expression of BCR-related genes in BCR and non-BCR samples in GSE116918.
Functional analysis of genes associated with BCR
The potential functions of these 16 genes associated with BCR were analyzed using KEGG and GO databases. The KEGG analysis revealed that these genes are significantly linked to several pathways, including ECM-receptor interaction, phagosome formation, focal adhesion, and the PI3K-Akt signaling pathway. Meanwhile, the GO analysis indicated that these genes predominantly contribute to the regulation of B cell differentiation, autophagic cell death, and the regulation of macrophage differentiation (Fig. 2A). Furthermore, we conducted an analysis of these genes using the GSCA database, which indicated that they are associated with the activation of the cell cycle, epithelial-mesenchymal transition (EMT), activation of estrogen receptors, and inhibition of the RTK pathway (Fig. 2B). Subsequently, we analyzed the correlation between these genes and the frequency of copy number variation. Our findings indicated that CTHRC1, ITGBL1, FAP, THBS2, MSR1, and OLR1 exhibited significant changes in copy number variation (CNV) frequency (Fig. 2C). Finally, we examined the expression differences of these genes across various pathological stages within the TCGA-PRAD dataset, revealing that all genes were highly expressed in higher pathological T stages, N stages, and Gleason scores (Fig. 2D–F).
A KEGG and GO analysis of BCR-related genes in PRAD. B GSCA database-based analysis of BCR-associated gene function in PRAD. C Analysis of the correlation between BCR related genes and the frequency of copy number variants. D–F Differential expression of BCR-related genes in different pathological stages of TCGA-PRAD. ***p < 0.001.
Cluster analysis based on BCR-related genes
The TCGA-PRAD samples were clustered using the Negative matrix factorization (NMF) clustering algorithm. To identify the most suitable method for categorizing TCGA-PRAD sample subgroups for our subsequent research, we employed a judgment standard based on co-expression curves, which is currently regarded as the clearest method. The optimal grouping is defined by the vertex corresponding to the largest drop in the co-expression curve. Our study demonstrates that dividing TCGA-PRAD samples into two clusters based on co-expression curves is the most appropriate approach. Additionally, the clustering heat map indicates that color distribution is more concentrated when the samples are divided into two clusters (Fig. 3A, B). When samples were categorized into two groups, individuals in cluster 1 exhibited a significantly better prognosis compared to those in cluster 2 (Fig. 3C). Furthermore, we analyzed the expression of genes associated with BCR across different groups, revealing significant differences in gene expression between the groups (Fig. 3D). Additionally, we incorporated various pathological parameters to compare the distribution of patient numbers across different clusters. Our results consistently demonstrate significant differences in patient counts among the clusters, regardless of the pathological T stage, N stage, or variations in PSA and Gleason scores (Fig. 3E–H).
A, B Consensus map of NMF clustering. C Survival differences between clusters. D Differences in the expression of BCR related genes between different clusters. E–H Differences in the distribution of different subgroups in different pathological stages of PRAD. ****p < 0.0001.
Analysis of BCR-related genes and immune infiltration and chemosensitivity in PCa patients
The level of immune cell infiltration in each sample from the TCGA-PRAD dataset was evaluated using the XCELL algorithm. Significant differences (p < 0.005) were observed in the levels of myeloid dendritic cell activation, T cell CD4+ memory, T cell CD4+ effector memory, common lymphoid progenitor, common myeloid progenitor, myeloid dendritic cell, granulocyte-monocyte progenitor, hematopoietic stem cell, macrophage, macrophage M1, monocyte, T cell CD4 + Th1, T cell CD4 + Th2, and regulatory T cells (Tregs) between the two clusters (Fig. 4A, B). A heat map illustrating the levels of immune cell infiltration was also generated (Fig. 4C). Furthermore, we analyzed differences in the IC50 scores of various compounds across the two clusters. Notably, several compounds, including commonly used therapeutic drugs for PCa patients, exhibited significant differences between the clusters, particularly bicalutamide (Fig. 4D). To further investigate the mechanisms underlying these findings, we conducted gene enrichment analysis on the two clusters. Given that patients in cluster 2 have a worse prognosis, we focused our analysis on pathways enriched in this cluster. The results indicated that WNT, PI3K-AKT, NOTCH1, VEGF, EGFR, and immune-related pathways were significantly enriched in cluster 2 (Fig. 4E).
A, B Analysis of BCR-related genes correlating with PRAD immune infiltration. C Heat map of different immune cell infiltration levels. D Analysis of biochemical relapse-related genes and chemotherapy sensitivity in PRAD patients. E Gene enrichment analysis of two clusters. ns på 0.05, *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
Construction of a diagnostic model based on the expression of biologic recurrence-related genes
To thoroughly investigate the role of biologic recurrence-related genes in PRAD, we assessed the predictive capability of these genes for diagnosing PRAD patients using ROC curves. Our analysis revealed that certain genes exhibit significant predictive ability for the diagnosis of PRAD, while COL1A1, INHBA, RCN3, and THBS2 demonstrate limited predictive capacity for PRAD diagnosis (Fig. 5A). Consequently, we endeavored to develop diagnostic models for PRAD based on these genes, utilizing six datasets from PRAD: the TCGA-PRAD dataset for training and GSE32571, GSE62872, GSE16120, GSE14206, and GSE38241 for validation. Among the 108 algorithm combinations tested, the LASSO + LDA algorithm emerged as the most effective for model development. The area under curve (AUC) value for the training set TCGA-PRAD was 0.911, while the AUC values for the validation cohorts GSE32571, GSE62872, GSE16120, GSE14206, and GSE38241 were 0.764, 0.616, 0.824, 0.860, and 0.897, respectively (Fig. 5B). Notably, 13 genes associated with BCR were incorporated into the diagnostic model constructed using the LASSO + LDA algorithm. These genes include ASPN, BGN, COMP, CTHRC1, FAP, INHBA, ITGBL1, MSR1, NOX4, OLR1, RCN3, SPP1, and TREM2 (Fig. 5C).
A An examination of how biologic recurrence-related genes predict diagnosis in PCa patients. B AUC values for diagnostic models created using various algorithm combinations. C The count of genes included in diagnostic models that were formed with different algorithm combinations.
Constructing a prognostic model for BCR
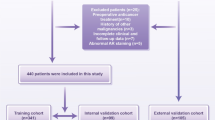
To construct a prognostic model related to BCR, we initially collected clinical data from patients with BCR in the TCGA-PRAD dataset. We defined the time node as the number of days prior to the first BCR, with the occurrence of BCR serving as the outcome measure. A total of 70 samples with comprehensive information from TCGA-PRAD were included in the analysis. Additionally, we incorporated data from 248 patients from the GSE116918 dataset into our study. Subsequently, samples from both the GSE116918 and TCGA-PRAD datasets were merged and randomly divided into a training set (50%), a validation set (50%), and a complete dataset using a random seed for randomization. Prognosis-related single genes were screened using univariate Cox analysis with a p-value cutoff of less than 0.05. This study identified a total of six genes: OLR1, COMP, INHBA, COL1A1, RCN3, and ASPN (Fig. 6A). Subsequently, genes were compressed using LASSO and stepwise regression, leading to the construction of a model through multivariate Cox analysis (Fig. 6B). In the entire dataset, our prognostic model demonstrated a robust predictive capability for BCR-related prognosis in patients. Notably, patients categorized in the high-risk group exhibited significantly poorer BCR-free prognosis compared to those in the low-risk group (Fig. 6C, D). Additionally, we presented risk factor plots for COMP and INHBA, which are included in the model, in both the test set and train set (Fig. 6E, F). The results derived from the ROC curves and Kaplan-Meier curves indicated that our BCR-related prognostic model exhibited strong predictive value in both the test set and train set (Fig. 6G–J).
A Screening of prognosis-related genes by univariate Cox analysis. B Genes were screened through lasso/stepwise regression, and a model was constructed using multivariate Cox analysis. C ROC curve analysis of the model’s predictive ability for BCR-related prognosis in the entire dataset. D The Kaplan-Meier survival curves for the high-risk and low-risk groups in the entire dataset. E Risk factor plot of the test dataset. F Risk factor plot of the train dataset. G ROC curve analysis of the model’s predictive ability for BCR-related prognosis in patients within the test dataset. H The Kaplan-Meier survival curves for the high-risk and low-risk groups in the test dataset. I ROC curve analysis of the model’s predictive ability for BCR-related prognosis in patients within the train dataset. J The Kaplan-Meier survival curves for the high-risk and low-risk groups in the train dataset.
Machine learning algorithm identifies key BCR regulatory genes
To further identify the key regulatory genes associated with BCR in PCa, we employed the XGBoost algorithm and interpreted the results through SHAP, using BCR as the outcome variable to analyze the correlation between the genes included in the model and the recurrence. We displayed the top 15 genes associated with BCR in the TCGA-PRAD and GSE116918 datasets (Fig. 7A, B). Subsequently, we analyzed the importance of these genes through Friends analysis (Fig. 7C). Among the genes incorporated into our diagnostic and therapeutic model, COMP and INHBA consistently emerged as pivotal regulatory genes. Based on the results obtained from the combination of the XGBoost algorithm and Friends analysis, we found that the importance of COMP and its correlation with BCR were both higher than those of INHBA. Therefore, we selected COMP as the key gene for subsequent analysis. We divided the samples in the TCGA-PRAD dataset into two groups based on the median expression level of COMP: samples with expression levels above the median were designated as the COMP high-expression group, while those with expression levels below the median were designated as the COMP low-expression group. Our findings confirm a significant relationship between COMP expression and the level of immune cell infiltration (Fig. 7D, E). To further investigate the binding affinity of COMP with commonly used clinical drugs for PCa, we conducted molecular docking analysis. The three selected PRAD-related drugs demonstrated strong binding capabilities with COMP (Fig. 7F). We also performed gene enrichment analysis on COMP, revealing its significant association with immunotherapy pathways in tumors (Fig. 7G, H). The results of the TIDE algorithm indicated that patients with high COMP expression had poorer responses to immunotherapy (Fig. 7I).
A, B Xgboost algorithm identifies key regulatory genes for BCR. C Friend analysis of dominant genes in BCR-regulated genes. D, E Correlation analysis of COMP with the level of immune cell infiltration. F Molecular docking of COMP with drugs. G, H Functional analysis of COMP in PRAD. I Correlation Analysis Between COMP and Immunotherapy. *p < 0.05; **p < 0.01; ***p < 0.001.
The expression analysis of COMP in PRAD
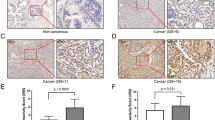
Our study highlights the critical role of COMP as a gene associated with BCR in PCa. We collected a total of 60 PRAD samples along with their corresponding normal prostate tissue samples. To investigate the differences in COMP expression and its relationship with recurrence and diagnosis in PRAD patients, we employed immunohistochemical staining techniques. Our results indicate that COMP expression is significantly higher in PRAD tissues compared to normal prostate tissues, with even more pronounced expression in recurrent samples than in non-recurrent samples (Fig. 8A–I). Furthermore, we utilized violin plots to visually illustrate the differences in COMP expression between PRAD and normal tissues (Fig. 8J). We evaluated the predictive potential of COMP for diagnosing PRAD patients through ROC curve analysis, which indicated that COMP possesses significant predictive value for this diagnosis (Fig. 8K). Finally, we analyzed the expression differences of COMP in recurrent versus non-recurrent samples, confirming that COMP is expressed at higher levels in recurrent samples and exhibits strong predictive ability for recurrence in PRAD patients (Fig. 8L, M).
A–C The expression of COMP in normal prostate tissue. D–F Expression of COMP in non-recurrence PRAD tissues. G–I Expression of COMP in recurrence PRAD tissues. J Differences in COMP expression between normal prostate tissue and PRAD. K The predictive value of COMP in the diagnosis of PRAD patients. L The expression difference of COMP in recurrence and non-recurrence PRAD samples. M The predictive value of COMP for recurrence in PRAD patients. *p < 0.05; ***p < 0.001.
Inhibiting COMP expression can suppress tumor progression
We first confirmed the knockdown efficiency of COMP siRNA in PCa cells using qRT-PCR (Fig. 9A). Our results demonstrated that COMP knockdown significantly impaired PCa cell proliferation (Fig. 9B, C). Furthermore, Transwell assays revealed that COMP knockdown markedly inhibited the migration and invasion capabilities of PCa cells (Fig. 9D, E). To investigate the in vivo role of COMP in tumor proliferation and metastasis, we established subcutaneous xenograft and lung metastasis models using male BALB/c nude mice (Fig. 9F). COMP knockdown significantly inhibited prostate tumor growth, evidenced by a marked reduction in both tumor volume and weight compared to controls (Fig. 9G–J). Immunohistochemical analysis indicated that COMP knockdown resulted in a notable decrease in Ki67 staining within tumors (Fig. 9K, L). Subsequently, we established a tail vein lung metastasis model and quantified metastatic burden using in vivo imaging. COMP knockdown effectively suppressed lung metastasis of prostate tumors, with nude mice exhibiting reduced lung fluorescent signals compared to the control group (Fig. 9M, N).
A Verification of COMP knockdown efficiency in PCa cells. B, C Colony formation assay used to evaluate the effect of COMP on PCa cell proliferation. D, E Transwell assays to assess the effect of COMP on the metastatic ability of PCa cells. F Tumor modeling in male BALB/c nude mice. G, H Subcutaneous tumor volume change curve. I Final subcutaneous tumor weight. J Final subcutaneous tumor size. K, L IHC staining images of COMP and Ki-67 in subcutaneous tumors. M, N Measurement of lung metastasis luciferase activity via in vivo imaging system. **p < 0.01; ***p < 0.001.
Discussion
PCa is the most prevalent tumor in the male genitourinary system24. According to statistics from the American Cancer Society in 2023, there were 288,300 new cases of PCa and 34,700 deaths among male cancer patients, making it the leading cancer in incidence and the third in mortality25. Treatment options for patients with low-risk localized PCa primarily include active surveillance, RP, radiotherapy, and androgen deprivation therapy. However, relevant statistics indicate that 30% to 50% of patients experience BCR following RP treatment26. Consequently, investigating the factors associated with BCR in PCa is both urgent and challenging.
WGCNA is a technique used to uncover relationships of co-expression among genes, and it is widely applied in the fields of bioinformatics and genomic studies. This approach allows for the organization of genes into modules of co-expression, which are often linked with particular biological functions or phenotypic characteristics, thus enhancing our comprehension of intricate biological mechanisms. By examining the correlations between gene modules and phenotypic information, WGCNA can identify essential genes involved in specific biological processes27,28. These significant genes could act as biomarkers or serve as potential targets for therapy. Therefore, our aim was to identify critical regulatory genes related to BCR in PCa through the application of the WGCNA technique. By incorporating clinical data from the TCGA-PARD dataset, we identified 16 differential prognostic genes associated with BCR. Due to the significant disparity in the number of death samples compared to non-death samples in the TCGA-PRAD dataset, we focused on PFI as the prognostic measure. In the GSE116918 dataset, the expression levels of these 16 genes were all upregulated in samples with BCR. Notably, CTHRC1 has been validated as a predictor of PCa recurrence, while ITGBL1 has been implicated in promoting the invasion and migration of PCa cells by activating the NF-κB signaling pathway in PCa29,30. Furthermore, the macrophage cytokine Spp1 has been shown to enhance the growth of prostate intraepithelial neoplasia, thereby facilitating prostate tumor progression31. TREM2 serves as an independent predictor of poor prognosis in PCa patients and promotes PCa cell migration via the PI3K/AKT axis32. The results of the enrichment analysis indicate that the interaction between the ECM and receptors is associated with genes related to BCR. The ECM functions as a barrier to the progression of cancer cells at both primary and metastatic sites. To navigate this ECM barrier, cancer cells modify their production of specific proteases that degrade ECM components33. Consequently, BCR in patients with PCa may be linked to this mechanism. Additionally, GO results demonstrate that genes associated with BCR are involved in the regulation of B cell differentiation and the Toll-like receptor signaling pathway, underscoring the significant role of these genes in the immune microenvironment of PCa.
With the advent of high-throughput sequencing technology, the size and complexity of genomic data have increased significantly. Machine learning algorithms are adept at processing and analyzing these large-scale datasets, allowing for the extraction of valuable information and patterns. These models can predict gene functions based on characteristics such as gene sequences and expression data, which is critical for understanding the roles of genes in organisms and their potential implications in disease34,35,36. NMF cluster analysis, a form of machine learning, was employed to categorize PCa patients into two distinct clusters. These clusters displayed differing clinical characteristics, with patients in cluster 2 generally experiencing poorer outcomes in terms of PFI. Gene enrichment analysis revealed that well-known signaling pathways, including WNT, PI3K-AKT, NOTCH1, VEGF, and EGFR, were significantly enriched in cluster 2, potentially explaining the unfavorable PFI observed in these patients. To facilitate the early identification of PCa, we developed a diagnostic model utilizing multiple machine learning algorithms, which was subsequently validated across various datasets. Consequently, we assert that the diagnostic model we constructed possesses exceptionally high predictive value. Additionally, we established a prognostic model related to BCR using the LASSO algorithm, which was also validated. The receiver operating characteristic (ROC) curve further corroborates the robust predictive power of the model we developed.
Using the Xgboost algorithm, we identified COMP as the most significant regulatory gene. Concurrently, we analyzed the correlation between COMP and the immune microenvironment in PCa. Molecular docking and correlation analyzes of drug IC50 scores demonstrated the strong binding affinity of COMP to bicalutamide and other therapeutic agents for PCa. Furthermore, results from immunohistochemical experiments, as well as in vivo and in vitro studies, indicate that COMP can serve as a biomarker for the diagnosis and recurrence of PCa, in addition to being a potential therapeutic target.
Our study acknowledges certain limitations. Although we have preliminarily demonstrated the stability and predictive performance of the diagnostic and therapeutic model constructed based on BCR-related genes, the sample size remains insufficient. Validation with larger and more diverse independent cohorts will further enhance the model’s generalizability. In the future, we plan to collect and analyze additional independent clinical cohort data to continuously optimize and validate the model.
Methods
Datasets and patient samples
The current research mainly made use of the TCGA-PRAD and GSE116918 datasets to pinpoint genes linked to BCR. Various diagnostic models were created and assessed through different datasets, such as TCGA-PRAD, GSE16120, GSE32571, GSE38241, GSE62872, and GSE14206. A total of sixty samples of PCa and adjacent non-cancerous tissues were obtained from Shanghai Aoduo Biotechnology Company. The tissue microarray analysis incorporated patients who had surgical procedures performed between January 2011 and December 2014. Follow-up assessments are set to persist until November 2021, with the ultimate follow-up duration varying between 6 to 10 years. The study received approval from the Shanghai Outdo Ethical Committee (SHYJS-CP-2207001) and the Nantong Tumor Hospital Ethical Committee (2024-118).
WGCNA identified BCR-related genes
Initially, we utilized gene expression profiles to calculate the median absolute deviation (MAD) for each gene individually. Subsequently, we eliminated the top 50% of genes exhibiting the smallest MAD and employed the R package WGCNA to construct a scale-free co-expression network. Following the establishment of the scale-free network, we constructed a clustering dendrogram with a minimum gene module size of 30 and a threshold of 0.25. We then calculated module membership (MM) and gene significance (GS) to assess the correlation between key modules and clinical features37.
Cluster Analysis Using NMF
The NMF methodology reveals coefficients within the gene expression matrix that possess biological significance, effectively arranging genes and samples to uncover the fundamental structural elements of the data, which facilitates the grouping of samples [9]. For the analysis of differential expression, a comparison was made between clusters A and B utilizing the ‘Limma’ R package, with thresholds established at |logFC | > 0.5 and an adjusted p-value of less than 0.05. Subsequent to this, all samples were organized into clusters based on the differentially expressed genes (DEGs) found within the subclusters, employing the ‘NMF’ R package to unveil possible molecular subtypes. The ‘brunet’ algorithm was utilized with 100 iterations for each assigned value, examining a range of clusters from 2 to 10. The identification of the most suitable cluster counts took into account factors such as cophenetic correlation, dispersion, and silhouette width38.
Analysis of BCR-related genes with PCa immune infiltration and drug sensitivity
To assess the immune score for each sample within the TCGA-PRAD dataset, we utilized the immunedeconv tool, an R package that incorporates six sophisticated algorithms: xCell, TIMER, MCP-counter, CIBERSORT, EPIC, and quanTIseq39. For our investigation, we chose to focus on the xCell algorithm because of its capability to evaluate numerous immune cell types, which aligns well with our research aims. Furthermore, we accessed the Genomics Drug Sensitivity Database of Cancer to forecast the chemotherapy response for each sample. This forecasting was performed using the pRRophetic R software package.
Functional analysis of genes associated with BCR
In our gene set enrichment analysis (GSEA), we employed the GSEA software, version 3.040. The samples were divided into two categories according to elevated and reduced expression levels of important genes. Furthermore, we retrieved the c2.cp.reactome.v7.4.symbols.gmt sub-collection from the Molecular Signatures Database to evaluate pertinent pathways and molecular mechanisms41. To perform functional enrichment analysis of gene sets, we employed the Gene Ontology (GO) as well as the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway annotations derived from the R software package (version 3.1.0) to establish a background for mapping the genes. The enrichment analysis was carried out with the aid of the R software package clusterProfiler (version 3.14.3) to derive the results of gene set enrichment. We defined the minimum size of a gene set as 5 and the maximum as 5000.
Diagnostic and prognostic modeling of genes associated with BCR
To develop predictive models for PCa diagnosis, we tested multiple configurations of machine learning algorithms. The training cohort was obtained from TCGA-PRAD, while validation was performed using the GSE16120, GSE32571, GSE38241, GSE62872, and GSE14206 datasets. Each algorithm configuration was evaluated based on the AUC metric. The configuration achieving the highest average AUC was selected as the optimal diagnostic model. Subsequently, to build a prognostic model for BCR, we pooled samples with associated BCR clinical data from both the TCGA-PRAD and GSE116918 datasets. This combined cohort was analyzed using the LASSO regression algorithm.
Immunohistochemical validation
Initially, the tissue microarray was heated at 85 °C for 10 min, followed by immersion in xylene for 15 min and hydration through a graded ethanol series (100%, 95%, 80%, and 70%). The microarray was then processed in an autoclave using a citric acid solution for antigen retrieval. After cooling, slides were washed with phosphate-buffered saline (PBS) and treated with hydrogen peroxide for 20 min. Subsequently, the COMP antibody (No. 28369-1-AP, 1:100) was applied and incubated at room temperature for 2 h. Following three PBS washes, the microarray was incubated with secondary immunohistochemistry antibodies for 20 min at room temperature. After three additional PBS rinses, staining was performed using 3,3’-diaminobenzidine followed by hematoxylin counterstaining. Dehydration proceeded through a graded ethanol series (70%, 80%, 90%, and 100%), followed by an 8-minute immersion in xylene. Blocking procedures were then performed. Immunostaining intensity was scored as: 0 (negative), 1 (weak), 2 (moderate), or 3 (strong). The percentage of positively stained cells was scored as: 1 (0–25%), 2 (26–50%), 3 (51–75%), or 4 (76–100%). The final score was calculated as the product of the intensity score and the percentage score.
Cell culture and transfection
Human PCa cell lines (DU145 and PC3) were obtained from the American Type Culture Collection (ATCC) and authenticated by short tandem repeat (STR) profiling. DU145 cells were cultured in DMEM medium (Gibco, USA), and PC3 cells were cultured in F-12K medium (MeilunBio, China). Both media were supplemented with 10% fetal bovine serum (FBS; VivaCell Biosciences, China) and 1% penicillin-streptomycin (Beyotime, China). Cells were maintained at 37 °C in a humidified 5% CO2 atmosphere. Transient transfection was performed using SuperKine™ Lipo3.0 (Abbkine, China) according to the manufacturer’s protocol. GenePharma designed and synthesized small interfering RNAs (siRNAs) targeting COMP with the following sequences: Sense: 5’-GGUCACGGAUGACGACUAUTT-3’, Antisense: 5’-AUAGUCGUCAUCCGUGACCTT-3’.
Proliferation ability analysis
The assessment of cell viability was conducted using a colony formation assay. PCa cells were grown in 6-well plates, with each well initially seeded with 2000 cells. These cells were incubated at 37 °C for two weeks in an incubator. Upon completion of the incubation period, the visible cell colonies in each well were treated with a 0.1% crystal violet solution for 15 min, which facilitated better visualization and quantification of the colonies.
Transwell assay
A total of 600 µL of medium with 10% FBS was placed in the lower chamber, whereas 200 µL of a PCa cell suspension (4 × 104 cells) in serum-free medium was added to the upper chamber of the Transwell apparatus. The distinction between migration and invasion assays is based on whether the upper chamber is pre-coated with Matrigel. Following a 48-hour incubation period, gently remove the cells adhered to the upper membrane using a cotton swab. The cells that migrated to the lower membrane were subsequently fixed, stained, captured in images, and quantified using ImageJ software.
qRT-PCR
Total RNA was extracted from fresh tissues or cells using TRIzol reagent (Takara, T9108, Dalian, China). Complementary DNA (cDNA) was synthesized from total RNA using the Hiscript II Q RT SuperMix for qPCR (+ gDNA wiper) kit (Vazyme R223, Nanjing, China). qRT-PCR was performed on an ABI Prism 7500 Fast Real-Time PCR System (Applied Biosystems, StepOnePlus, USA) using ChamQ Universal SYBR qPCR Master Mix (Vazyme, Q711) with specific PCR primers (Sangon Biotech Co., Ltd., Shanghai, China). GAPDH was used as the normalization control, and the fold changes were calculated using the 2-ΔΔCT method. Below are the primer pairs’ sequences for the target gene COMP (Forward: ACACAGGGTCAAGGAGATCAC, Reverse: AGACTACGCCAGGGAAGCA) and GAPDH (Forward: GGAGCGAGATCCCTCCAAAAT, Reverse: GGCTGTTGTCATACTTCTCATGG).
Subcutaneous tumor and lung metastasis model
Five-week-old male BALB/c nude mice were reared under specific pathogen free (SPF) conditions. The subcutaneous tumor model was established as follows: 2 × 106 PC-3 or DU145 cells (control group and siCOMP group) were inoculated in 100 μL DMEM basal medium, respectively, and subcutaneously injected into the right back of mice, with 5 cells in each group. Tumor size was measured with digital caliper every 3 days, and volume V = 1/2 × (length × width²) was calculated. After 4 weeks, mice were sacrificed by dislocation. Excised tumors were weighed, photographed, and fixed in 10% neutral buffered formalin for 24 h before embedding in paraffin. It was used for subsequent HE and IHC staining. Caudal vein pulmonary metastasis was modeled as follows: 5 nude mice in each group were injected with 2 × 106 tumor cells (PC-3 or DU145, control group and siCOMP group) with stable luciferase. In vivo imaging was performed after 6 weeks. The nude mice were anesthetized with isoflurane. Imaging detection was performed 10 min after injection of sodium fluorescein using an IVIS Small animal in vivo imager. All procedures were approved by the Animal Care and Use Committee of Nantong Tumor Hospital and followed the ARRIVE guidelines (2024-118).
Statistical analysis
The expression of BCR-related genes in both normal tissues and PRAD was evaluated using the Wilcoxon rank-sum test. The log-rank test was employed to assess prognosis. To investigate the relationship between gene expression and stemness scores, Pearson correlation analysis was performed. A p-value threshold of less than 0.05 was established to indicate statistical significance.
Data availability
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
The underlying code for this study is not publicly available but may be made available on reasonable request from the corresponding author.
References
James, N. D. et al. The Lancet Commission on prostate cancer: planning for the surge in cases. Lancet 403, 1683–1722 (2024).
Zhong, J. et al. Combining MRI radiomics, hypoxia gene signature score and clinical variables for prediction of biochemical recurrence-free survival after radiotherapy in prostate cancer. Radiol. Med. (2025).
Cornford, P. et al. EAU-EANM-ESTRO-ESUR-ISUP-SIOG guidelines on prostate cancer-2024 update. Part I: screening, diagnosis, and local treatment with curative intent. Eur. Urol. 86, 148–163 (2024).
Van den Broeck, T. et al. Prognostic value of biochemical recurrence following treatment with curative intent for prostate cancer: a systematic review. Eur. Urol. 75, 967–987 (2019).
Basourakos, S. P. et al. Understanding the impact of salvage radiation on the long-term natural history of biochemically recurrent prostate cancer after radical prostatectomy. Cancer Med. 14, e70988 (2025).
Lin, X. et al. Assessment of biochemical recurrence of prostate cancer (Review). Int J. Oncol. 55, 1194–1212 (2019).
Huang, M. M. et al. Prostate magnetic resonance imaging to predict grade concordance, extra prostatic extension, and biochemical recurrence after radical prostatectomy. Urol. Oncol. 43, 445.e11–e19 (2025).
Paller, C. J. & Antonarakis, E. S. Management of biochemically recurrent prostate cancer after local therapy: evolving standards of care and new directions. Clin. Adv. Hematol. Oncol. 11, 14–23 (2013).
Falagario, U. G. et al. Epidemiology of high-risk biochemical recurrence after primary prostate cancer treatment. Eur. Urol. Oncol. (2025).
Margolese, N. et al. Prognostic factors of biochemical recurrence and impact of pre-treatment MRI for prostate cancer radiotherapy. Br. J. Radiol. (2025).
Hoey, C. et al. Circulating miRNAs as non-invasive biomarkers to predict aggressive prostate cancer after radical prostatectomy. J. Transl. Med. 17, 173 (2019).
Silagy, A. W. et al. Percentage of free to total PSA as a biomarker of survival in metastatic castration-resistant prostate cancer. BJU Int. 136, 329–335 (2025).
Inagaki, T. et al. PSA at postoperative three months can predict biochemical recurrence in patients with pathological T3 prostate cancer following radical prostatectomy. Int J. Urol. 16, 941–946 (2009).
Planas, J. & Morote, J. [Biochemical recurrence criteria after radical prostatectomy. Natural history of the disease]. Arch. Esp. Urol. 65, 4–11 (2012).
Xia, S. J. & Jing, Y. F. Biochemical recurrence of prostate cancer: the controversial recognition and management. Chin. Med J. (Engl.). 124, 1283–1285 (2011).
Inamoto, T., Komura, K., Watsuji, T. & Azuma, H. Rapid increase of the serum PSA level in response to high-intensity focused ultrasound therapy may be a potential indicator of biochemical recurrence of low- and intermediate-risk prostate cancer. Clin. Med. Insights Oncol. 5, 101–106 (2011).
Toledano, A. et al. [Elevation of PSA after prostate radiotherapy: rebound or biochemical recurrence?]. Prog. Urol. 18, 557–561 (2008).
Park, S. Y. et al. Osteoarthritis progression pattern based on patient specific characteristics using machine learning. NPJ Digit Med. 8, 464 (2025).
Phillips, E. et al. Hybrid machine learning for real-time prediction of edema trajectory in large middle cerebral artery stroke. NPJ Digit Med. 8, 288 (2025).
Ishizu, K. et al. Establishment of a machine learning model for predicting splenic hilar lymph node metastasis. NPJ Digit Med. 8, 93 (2025).
Lin, M. et al. Exploring biomarkers of MAPK pathway co-expression in lung adenocarcinoma and their functions based on machine learning algorithms and single-cell analysis. Genes Dis. 12, 101222 (2025).
Mou, L. et al. Phosphorylation-related genes in lupus nephritis: single-cell and machine learning insights. Genes Dis. 12, 101385 (2025).
Wang, Y. et al. Identification of metastasis-related genes for predicting prostate cancer diagnosis, metastasis and immunotherapy drug candidates using machine learning approaches. Biol. Direct 19, 50 (2024).
Culp, M. B., Soerjomataram, I., Efstathiou, J. A., Bray, F. & Jemal, A. Recent Global Patterns in Prostate Cancer Incidence and Mortality Rates. Eur. Urol. 77, 38–52 (2020).
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 73, 17–48 (2023).
Zhang, X. et al. Development and validation of a circulating tumor cells-related signature focusing on biochemical recurrence and immunotherapy response in prostate cancer. Heliyon 9, e22648 (2023).
Zhou, Q. et al. Identification of methylated differentially expressed hub genes in systemic lupus erythematosus patients by WGCNA-based epigenome-wide and transcriptome-wide analysis. Genes Dis. 10, 1771–1774 (2023).
Di, Z. et al. Single-cell and WGCNA uncover a prognostic model and potential oncogenes in colorectal cancer. Biol. Proced. Online 24, 13 (2022).
Zhou, Q. et al. CTHRC1 and PD‑1/PD‑L1 expression predicts tumor recurrence in prostate cancer. Mol. Med Rep. 20, 4244–4252 (2019).
Li, W. et al. ITGBL1 promotes EMT, invasion and migration by activating NF-κB signaling pathway in prostate cancer. Onco Targets Ther. 12, 3753–3763 (2019).
Messex, J. K., Byrd, C. J., Thomas, M. U. & Liou, G. Y. Macrophages Cytokine Spp1 Increases Growth of Prostate Intraepithelial Neoplasia to Promote Prostate Tumor Progression. Int J Mol Sci.;23, (2022).
Gao, H. T. et al. TREM2 as an independent predictor of poor prognosis promotes the migration via the PI3K/AKT axis in prostate cancer. Am. J. Transl. Res. 15, 779–798 (2023).
Stewart, D. A., Cooper, C. R. & Sikes, R. A. Changes in extracellular matrix (ECM) and ECM-associated proteins in the metastatic progression of prostate cancer. Reprod. Biol. Endocrinol. 2, 2 (2004).
Liu, T. et al. Machine learning-directed conversion of glioblastoma cells to dendritic cell-like antigen-presenting cells as cancer immunotherapy. Cancer Immunol. Res. 12, 1340–1360 (2024).
Jayakrishnan, T. T. et al. Multi-omics machine learning to study host-microbiome interactions in early-onset colorectal cancer. NPJ Precis. Oncol. 8, 146 (2024).
Guan, S. et al. Identifying potential targets for preventing cancer progression through the PLA2G1B recombinant protein using bioinformatics and machine learning methods. Int J. Biol. Macromol. 276, 133918 (2024).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 9, 559 (2008).
Wang, X., Rao, J., Zhang, L., Liu, X. & Zhang, Y. Identification of circadian rhythm-related gene classification patterns and immune infiltration analysis in heart failure based on machine learning. Heliyon 10, e27049 (2024).
Sturm, G. et al. Comprehensive evaluation of transcriptome-based cell-type quantification methods for immuno-oncology. Bioinformatics 35, i436–i445 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Acknowledgements
This research was supported by the Natural Science Foundation of Heilongjiang Province (PL2024H044).
Author information
Authors and Affiliations
Contributions
Y.W. and H.Z. spearheaded the methodological framework design and manuscript drafting. J.R. supplemented new experiments during the revision phase and analyzed the data. M.R. developed the study concept and design, secured funding, and revised the manuscript for important intellectual content. All authors have accepted responsibility for the entire content of this manuscript and approved its submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Zhu, H., Ren, J. et al. Integrative machine learning models predict prostate cancer diagnosis and biochemical recurrence risk: Advancing precision oncology. npj Digit. Med. 8, 524 (2025). https://doi.org/10.1038/s41746-025-01930-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01930-6
This article is cited by
-
Talin1 is downregulated in testicular germ cell tumors according to combined bioinformatics and experimental approaches
Scientific Reports (2026)
-
Downregulation of ankyrin 3 (ANK3) promotes malignant behaviors associated with altered adhesion dynamics and actin cytoskeleton remodeling in renal cell carcinoma
Cell and Tissue Research (2025)