Abstract
Hematoma expansion (HE) is a critical therapeutic target in spontaneous intracerebral hemorrhage (sICH), yet its reliable early identification remains challenging. We developed an automated pipeline for HE prediction using non-contrast computed tomography from 2020 patients across five centers. The modular framework comprised automated segmentation, synthetic data augmentation, and Vision Transformer (ViT)-based classification. High-quality hematoma masks were generated by the full-scale U-Mamba model, identified as the optimal architecture through comprehensive benchmarking. Two augmented training sets were constructed using synthetic HE images from the Diffusion-UKAN model: UKAN-Balanced (HE: NHE = 1:1) and UKAN-Semibalanced (HE: NHE = 1:2). The ViT-1:2 classifier, trained on the UKAN-Semibalanced dataset, achieved a training set AUC of 0.815 and demonstrated robust cross-institutional generalization with external validation AUCs of 0.793 and 0.781 on two independent datasets. These findings suggest that the proposed modular approach provides a promising front-line tool for rapid HE risk stratification in acute care settings, with potentially improving clinical decision-making in sICH management.
Similar content being viewed by others
Introduction
Hematoma expansion (HE) occurs in approximately 20-30% of patients with spontaneous intracerebral hemorrhage (sICH) and is strongly associated with early neurological deterioration, long-term disability, and mortality1,2. Each additional 1 mL increase in hematoma volume raises the risk of death or functional dependency at 90 days by 5-7%3,4. Given its profound impact on outcomes, HE is a critical therapeutic target for acute-phase anti-expansion strategies5,6. Previous studies have shown that ultra-early intensive blood pressure reduction significantly reduces hematoma growth and improves clinical outcomes, underscoring the importance of rapid intervention in sICH patients at high risk of HE7,8,9. However, most expansions occur with 3-6 hours of symptom onset, leaving only a narrow therapeutic window7,10. This temporal constraint highlights the urgent need for reliable, accessible tools that accurately stratify risk and identify patients most likely to benefit from anti-expansion therapies.
Non-contrast computed tomography (NCCT) remains the frontline imaging modality for hematoma evaluation due to its broad availability, rapid acquisition, and suitability for intensive care settings11. Several predictors have been proposed, including NCCT-based radiological features, CT angiography (CTA) spot signs, and various clinical variables12,13,14. Yet these markers have inherent limitations: NCCT feature interpretation suffers from inter-observer variability, CTA is not routinely performed in the hyperacute phase, and clinical variables alone provide limited discriminative power15,16,17. Moreover, acquiring these variables in clinical practice is often time-consuming, which to some extent hinders their utility in acute settings requiring urgent decision-making18. Deep learning has emerged as a promising approach for HE prediction19,20,21,22; however, its clinical translation remains constrained by the scarcity of high-quality medical imaging data, and the relatively low incidence of HE further compounds this challenge23,24,25. Generative artificial intelligence offers a potential solution26. By producing realistic synthetic images, generative models can augment training data, reduce bias from class imbalance, and enhance performance across diverse downstream tasks, including classification, segmentation, and detection27. A recent study suggests that integrating synthetic with real-world data enhances HE prediction20, but the finding remains exploratory and requires rigorous validation across diverse clinical settings. Another practical barrier is that manual or semi-automated hematoma segmentation is labor-intensive and impractical for real-world workflows28. While recent deep learning approaches have automated segmentation and classification, most were evaluated on limited external datasets with relatively few HE cases21,22.
To address these challenges, we developed an end-to-end modular deep learning pipeline for HE prediction in sICH patients. This framework comprises three sequential components: automated segmentation for precise hematoma localization, synthetic data augmentation to alleviate class imbalance, and a classification module for patient-level risk prediction. By leveraging a large multicenter dataset, this approach aims to enable reliable early identification of high-risk patients and facilitate timely, data-driven decision-making in acute care settings.
Results
Study population and dataset construction
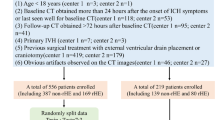
This multicenter retrospective study comprised 2020 patients with sICH recruited from five tertiary medical centers (Fig. 1). The training dataset incorporated 1103 patients from three centers (Centers 1, 2, and 3), including 218 patients (19.8%) with HE and 885 (80.2%) with NHE. For external validation, we utilized two independent cohorts: external validation set 1 included 513 patients from Center 4 (112 HE, 21.8%; 401 NHE, 78.2%), while external validation set 2 consisted of 404 patients from Center 5 (81 HE, 20.0%; 323 NHE, 80.0%). All models were trained on the multicenter training dataset and validated on two independent external validation datasets. Detailed baseline characteristics across all cohorts are presented in Table 1.
sICH Spontaneous intracerebral hemorrhage, NCCT non-contrast CT, HE hematoma expansion, NHE non-hematoma expansion.
Performance of benchmarking and full-scale segmentation model
The performance of four advanced 3D segmentation architectures (U-Mamba, nnU-Net, nnFormer, and UNETR + +) was benchmarked using a preliminary dataset of 1000 NCCT images (Fig. 2). As illustrated in Fig. 3A–C, U-Mamba consistently achieved the best segmentation accuracy, with the highest average Dice score of 0.937 and Intersection over Union (IoU) of 0.934. Meanwhile, U-Mamba also displayed faster convergence and more stable loss curves. Additionally, U-Mamba showed superior robustness in handling imaging artifacts, and accurately identified hematomas with complex shapes, ill-defined boundaries, substantial intralesional density variation, and intraventricular hemorrhage, thereby highlighting its strong generalization capability in challenging clinical scenarios (Fig. 3D).
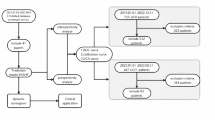
The modular pipeline comprises three sequential stages: automated hematoma segmentation, synthetic minority data augmentation, and automated hematoma classification. Stage 1: Four state-of-the-art 3D segmentation networks (U-Mamba, nnU-Net, nnFormer, and UNETR++) were benchmarked on a preliminary dataset of 1000 NCCT scans (baseline and follow-up) from 500 sICH patients randomly selected from the full training cohort of 1,103 patients. The best-performing model (U-Mamba) was then trained on all 2206 scans from the complete training cohort to generate high-quality hematoma masks. Stage 2: Synthetic minority data augmentation employed Diffusion-UKAN to generate high-fidelity synthetic HE images, yielding two augmented training sets: UKAN-Balanced (HE: NHE = 1:1) and UKAN-Semibalanced (HE: NHE = 1:2). Stage 3: Automated classification used a Vision Transformer trained on three consecutive slices centered on maximum hematoma area (Max−1, Max, Max+1). Patient-level predictions were obtained by averaging slice-level probabilities, with Grad-CAM providing visual interpretation of discriminative regions. ROI, region of interest; HE, hematoma expansion; NHE, non-hematoma expansion.
A Training curves showing pseudo-Dice (dashed lines) and exponential moving average (EMA) pseudo-Dice (solid lines) over 500 epochs for U-Mamba, nnU-Net, nnFormer, and UNETR++. B Training and validation loss curves over epochs for the four architectures. C Comparison of Dice coefficient and Intersection over Union (IoU) values across models on the preliminary benchmarking dataset. D Representative segmentation outputs from each model compared with ground truth annotations.
Based on these comprehensive evaluations, U-Mamba was identified as the optimal segmentation architecture and subsequently retrained on the full training dataset comprising 2206 NCCT scans. The full-scale U-Mamba model exhibited excellent generalization and robustness across all datasets (Fig. 4A, B). Segmentation accuracy remained consistently high with Dice scores of 0.983 (training), 0.980 (external validation set 1), and 0.982 (external validation set 2), with corresponding IoU values of 0.968, 0.963, and 0.965, respectively. Visual analysis revealed accurate delineation across diverse hematoma presentations and anatomical distributions (Fig. 4C).
A Training and validation loss curves alongside pseudo-Dice and exponential moving average (EMA) pseudo-Dice metrics over 500 epochs during full-scale training on 2206 scans. B Dice coefficient and Intersection over Union (IoU) values achieved on the training set and two external validation sets. C Representative segmentation results from the full-scale U-Mamba model, showing input images, ground truth annotations, and model predictions with corresponding Dice and IoU scores for each case.
Quantitative and qualitative evaluation of synthetic data augmentation
To mitigate original training dataset class imbalance, which comprised 654 HE and 2655 NHE images (HE: NHE ≈ 1:4), two augmented datasets were constructed using Diffusion-UKAN synthetic HE images: UKAN-Balanced (1:1 ratio with 654 real HE, 2001 synthetic HE, 2655 real NHE) and UKAN-Semibalanced (1:2 ratio with 654 real HE, 674 synthetic HE, 2655 real NHE). The capability of the Diffusion-UKAN model was comprehensively assessed using both quantitative metrics and qualitative expert visual assessment. As shown in Fig. 5A, the training loss of the model steadily decreased over 1000 epochs, indicating effective convergence and stable learning. The Fréchet Inception Distance (FID), measuring distributional similarity between synthetic and real images, was 36.856 for UKAN-Balanced and 38.744 for UKAN-Semibalanced datasets (Fig. 5B). Similarly, the Inception Score (IS), assessing image quality and diversity, was 1.961 for UKAN-Balanced and 1.998 for UKAN-Semibalanced datasets, closely approximating the benchmark IS of 2.137 computed from the original Real-Imbalanced training set, suggesting adequate diversity in the generated images.
A Training loss curve with smoothed trend (window size = 10) over 1000 epochs for the Diffusion-UKAN model. B Fréchet Inception Distance (FID) and Inception Score (IS) values for UKAN-Balanced (1:1), UKAN-Semibalanced (1:2), and Real-Imbalanced (1:4) datasets. C Representative hemorrhage expansion CT images stratified by expert radiological assessment. Images were evaluated by three independent radiologists using a standardized 5-point Likert scale across two dimensions: overall image quality and hematoma/surrounding structure feature authenticity.
Expert radiological evaluation further confirmed the clinical authenticity of synthetic images. No significant difference was observed between real and synthetic HE in overall image quality (3.54 ± 0.60 vs. 3.41 ± 0.78, p = 0.156) or hematoma/structural feature authenticity (3.31 ± 0.65 vs. 3.15 ± 0.93, p = 0.207), with 87% and 74% of synthetic images scoring ≥3 in each dimension, respectively. Inter-rater agreement was good to excellent (ICC = 0.775 and 0.814) (Table 2). Fig. 5C illustrates this quality distribution with representative examples. The high-quality synthetic images (Likert scores ≥ 3 in both dimensions) closely resembled real HE images with diverse hematoma morphologies and realistic radiological textures. In contrast, low-quality synthetic images (scores ≤ 2) displayed distinctive artifacts readily identified by radiologists, including abnormal color distortions with yellowish tints inconsistent with standard CT grayscale presentation, unrealistic hematoma morphology and density patterns deviating from clinical expectations, and anatomical distortions in surrounding tissue structures. Despite these identifiable artifacts in a minority of cases, only 24% of synthetic images were correctly identified in a binary classification task (Fleiss’ κ = 0.735). These findings demonstrate that Diffusion-UKAN predominantly generates high-quality, clinically realistic HE images that are largely indistinguishable from real cases by expert observers.
Patient-level classification performance and interpretability of vision transformer (ViT)
The patient-level classification efficacy of the three ViT-based models (ViT-1:1, ViT-1:2, and ViT-1:4) for HE prediction is shown in Fig. 6A–G and Table 3. The ViT-1:2 model, trained on the UKAN-Semibalanced dataset, achieved the best performance across the training and validation sets. In the training set, ViT-1:2 reached an AUC of 0.815 (95% CI: 0.790–0.841). This superior performance was maintained in both external validation sets. In external validation set 1, ViT-1:2 obtained an AUC of 0.793 (95% CI: 0.750–0.837), which was significantly higher than ViT-1:1 (AUC = 0.752, 95% CI: 0.701–0.806; ΔAUC = 0.041, p < 0.001, DeLong test) and ViT-1:4 (AUC = 0.714, 95% CI: 0.661–0.770; ΔAUC = 0.079, p < 0.001). Similar results were observed in external validation set 2, where ViT-1:2 (AUC = 0.781, 95% CI: 0.733–0.827) significantly outperformed ViT-1:1 (AUC = 0.738, 95% CI: 0.681–0.792; ΔAUC = 0.043, p = 0.004) and ViT-1:4 (AUC = 0.728, 95% CI: 0.667–0.783; ΔAUC = 0.053, p < 0.001).
A–F ROC curves and corresponding performance metrics (AUC, accuracy, sensitivity, specificity, PPV, and NPV) for ViT-1:1, ViT-1:2, and ViT-1:4 models on the training set (A, B), external validation set 1 (C, D), and external validation set 2 (E, F). G Confusion matrices of ViT-1:1, ViT-1:2, and ViT-1:4 models on both external validation sets. H Grad-CAM visualizations from the optimal ViT-1:2 model showing representative HE cases and NHE cases, with red regions indicating high importance for classification decisions. ViT-1:1: trained on the UKAN-balanced dataset (1:1); ViT-1:2: trained on the UKAN-semibalanced dataset (1:2); ViT-1:4: trained on the original real-imbalanced dataset (1:4).
Regarding model interpretability, the Gradient-weighted Class Activation Mapping (Grad-CAM) heatmaps from the optimal ViT-1:2 model clearly highlighted attention regions highly relevant to hematoma classification in both HE and NHE cases. Notably, the high-importance regions (shown in red) substantially overlapped with the actual hematoma locations, while activation intensities were relatively low in the perilesional regions. This anatomical precision demonstrates that the model learned to focus on clinically relevant regions for HE prediction, thereby further validating the model’s effectiveness (Fig. 6H).
Discussion
In this study, we developed and validated an end-to-end automated deep learning pipeline for predicting HE in sICH patients using admission NCCT images. A noteworthy feature of the framework is its modular processing structure, including automated hematoma segmentation, diffusion-based synthetic minority data augmentation, and ViT-based classification. Specifically, the benchmark-identified full-scale U-Mamba segmentation network generated high-quality hematoma masks. The Diffusion-UKAN model generated diverse and highly realistic synthetic HE images, which were used to construct two augmented training sets: UKAN-Balanced and UKAN-Semibalanced, thereby enhancing the performance of downstream analysis. The optimal ViT classifier, trained on the UKAN-Semibalanced dataset with an HE: NHE ratio of 1:2, demonstrated robust generalization capability, achieving AUCs of 0.793 and 0.781 on external validation sets 1 and 2, respectively. Overall, this deep learning pipeline represents a promising clinical tool capable of rapidly and accurately identifying sICH patients at high risk of HE, thus potentially facilitating early and precise clinical decision-making in acute intensive care settings.
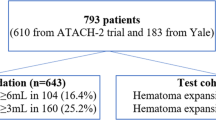
As artificial intelligence gains momentum in hemorrhagic stroke research, an increasing number of studies have shown that deep learning-based methods using NCCT imaging yield encouraging results in hematoma segmentation, medical image synthesis, and risk stratification for HE20,21,22,28,29,30. Yu et al.29 attained expert-level accuracy in hematoma segmentation and volume estimation using a 2D CT slice-based DR-UNet model. Similarly, Ironside et al.28 developed an automated CNN model that reached high segmentation accuracy (Dice ≈ 0.90), strong volumetric correlation (R² ≈ 0.98), and markedly improved efficiency compared to manual and semiautomated approaches. Despite these advances, the clinical utility of these studies remains limited, as they solely focus on volumetric analysis without extending these approaches to downstream tasks such as hematoma risk stratification and outcome prediction. Tran et al.21 proposed a fully automated, uncertainty-aware deep learning model to predict supratentorial HE from admission NCCT images in a multicenter cohort of 793 patients, achieving an AUC of 0.81 for HE ≥ 6 mL with high-confidence predictions. However, the limited number of positive cases in the test set (n = 26) may compromise statistical power and generalizability. A similar limitation was observed in another multicenter study, where despite prospective validation strengthening clinical translatability, only 65 of 500 sICH patients in the test set experienced HE22. Furthermore, in Tran et al.’s study, a pronounced class imbalance (104 HE versus 530 NHE cases) was observed in the training/cross-validation set, potentially biasing the model toward non-expansion predictions and limiting real-world performance31. To explore potential mitigation strategies, Yalcin et al.20 investigated the utility of synthetic image augmentation by incorporating five generated variants per original NCCT image during training. This approach significantly improved HE prediction performance, increasing sensitivity from 0.51 to 0.77 and F1-score from 0.53 to 0.82. Nevertheless, the synthetic images were generated from a limited single-center cohort of 122 patients with only 35 HE cases, raising concerns about potential overfitting and limited diversity32.
This study presents a modular deep learning pipeline for predicting HE in sICH patients using baseline NCCT, comprising automated segmentation, synthetic data augmentation, and classification components. We validated its robustness and generalizability across multicenter cohorts. The modular design enables the use of task-specific networks suited to each component’s characteristics33. For automated hematoma segmentation, U-Mamba emerged as the optimal model after benchmarking against three advanced 3D architectures (nnU-Net, nnFormer, and UNETR + +), achieving superior boundary accuracy and regional consistency. This performance is primarily attributed to its hybrid architecture, which integrates convolutional neural networks with structured state space models to enable effective multi-scale local feature extraction while efficiently modeling long-range dependencies34. To alleviate the scarcity of HE samples in the training set, we implemented a generative augmentation strategy based on Diffusion-U-KAN, which is capable of synthesizing strong anatomical realism and visual plausibility under data-constrained conditions35. By incorporating synthetic HE samples, we constructed augmented datasets with optimized class distributions, including both fully balanced and semi-balanced training cohorts. These synthetic images exhibited robust diversity while maintaining high fidelity to real images, as reflected in favorable FID and IS metrics. Moreover, expert radiological assessment provided independent validation of their clinical authenticity, with experienced radiologists unable to reliably distinguish synthetic from real images. Importantly, the augmentation strategy yielded significant downstream benefits. The ViT-1:2 model, trained on a semi-balanced training set (HE: NHE = 1:2) augmented by Diffusion-UKAN, outperformed ViT classifiers trained on both fully balanced and original real-imbalanced datasets. Grad-CAM interpretability analyses further validated the effectiveness of the ViT-1:2 model. The spatial distribution of attention weights demonstrated precise anatomical correspondence with hematoma regions, indicating the model’s capability to prioritize intrinsic hematoma heterogeneity. Notably, despite being trained with fewer synthetic HE samples than the fully balanced ViT-1:1 model, ViT-1:2 achieved superior performance across external validation cohorts. Similar observations have been reported in previous studies20,26. These results suggest that a semi-balanced training strategy may offer a more favorable trade-off between class representation and data realism.
There are several limitations to our study. First, although the model demonstrated strong generalizability across two external validation cohorts, prospective evaluation in real-world emergency settings is warranted to support broader clinical adoption. Second, the expert visual evaluation has inherent limitations. Reviewers’ awareness of participating in a synthetic image evaluation task may have induced heightened scrutiny beyond routine clinical practice. Additionally, images were assessed in isolation without accompanying clinical context, which diverges from real-world diagnostic conditions. Nevertheless, the consistently high-quality scores and low correct identification rate of synthetic images provide compelling evidence for their clinical authenticity. Third, the retrospective design of this study limited the ability to systematically evaluate the relationship between HE prediction and clinical outcomes, such as 90-day modified Rankin Scale (mRS) scores and mortality. Future prospective studies incorporating longitudinal follow-up data are needed to rigorously assess the prognostic value of the deep learning model and further enhance its clinical applicability. Finally, while this analysis intentionally focused on baseline NCCT images alone to ensure practical applicability in acute settings, future studies could explore whether combining our automated approach with readily available clinical parameters might offer additional prognostic value without compromising efficiency.
In conclusion, our modular pipeline demonstrates robust performance in automated hematoma segmentation, synthetic HE image generation, and risk stratification for sICH patients. The results underscore the advantages of task-specific module optimization within an integrated framework. This approach offers a practical solution for HE prediction in acute care settings, with the potential to improve triage decisions and personalize treatment strategies.
Methods
Patient cohort and image acquisition
This retrospective study was approved by the institutional review boards of all participating centers, with waiver of informed consent. Inclusion criteria were: (1) spontaneous intracerebral hemorrhage; (2) availability of complete clinical and imaging data; (3) age ≥18 years; and (4) baseline NCCT obtained within 24 hours of symptom onset with follow-up NCCT performed within 72 hours. Exclusion criteria were: (1) poor image quality; (2) surgical intervention before follow-up imaging; (3) primary intraventricular hemorrhage; (4) baseline or follow-up NCCT with slice thickness >5 mm; and (5) baseline hematoma involving fewer than 3 contiguous axial slices. HE was defined as absolute hematoma volume growth >6 mL and/or relative hematoma volume growth >33% between baseline and follow-up CT scans12,15,36,37.
Baseline and follow-up NCCT scans were acquired at participating institutions using local acquisition protocols, detailed in Supplementary Table S1. All NCCT images were resampled to uniform voxel spacing (0.5 × 0.5 × 5 mm³) using linear interpolation to ensure standardized spatial resolution for subsequent analysis.
Automated hematoma segmentation
Automated hematoma segmentation was performed through two sequential steps. (1) Step 1: Segmentation model benchmarking. We conducted benchmarking experiments to compare four state-of-the-art 3D segmentation architectures: U-Mamba, nnU-Net, nnFormer, and UNETR + +. This benchmarking utilized a preliminary dataset comprising 1000 NCCT scans (baseline and follow-up) from 500 sICH patients randomly selected from the full training cohort of 1103 patients. The dataset was partitioned into training and validation subsets (4:1 ratio) exclusively for benchmarking purposes. Manual segmentation masks were delineated by a neuroradiologist with 6 years of experience using ITK-SNAP 3.8.0. Segmentation efficacy was evaluated using Dice score and IoU metrics to identify the optimal architecture. (2) Step 2: Full-scale model training and validation. The best-performing architecture from Step 1 was subsequently trained on the complete dataset, incorporating 2206 NCCT images (baseline and follow-up) from all 1103 patients in the training cohort. Initial segmentation masks were generated through automatic inference using the Step 1 model, then manually refined to ensure accurate 3D hematoma annotations for full-scale training. Following model training, segmentation accuracy was validated on three independent datasets to assess accuracy and robustness. The high-quality segmentation masks generated by the full-scale model were utilized to support downstream tasks, including synthetic data augmentation and patient-level classification.
All segmentation models were trained using identical hyperparameters. We used Stochastic Gradient Descent (SGD) as the optimizer with an initial learning rate of 0.001, Nesterov momentum of 0.99, and weight decay of 3 × 10⁻⁵. The loss function combined Dice loss and cross-entropy loss. Models were trained for 500 epochs with a batch size of 2. For intensity normalization, CT voxel intensities were clipped to the 0.5–99.5 percentile range to remove outliers, followed by z-score normalization using global mean and standard deviation.
Segmentation-based hematoma volume calculation
Volumetric analysis was performed using 3D NCCT images and their corresponding binary masks, which were manually refined from the initial outputs of the full-scale segmentation model to ensure accurate hematoma delineation. Final masks were converted to closed triangular surfaces using the Marching Cubes algorithm implemented in PyRadiomics (https://pyradiomics.readthedocs.io/en/latest), and mesh volumes were calculated using the following formula38:
where p₁, p₂, and p₃ are the three vertex coordinates of each triangular face T, and the determinant represents the signed volume of the tetrahedron formed by the triangle and the origin.
Diffusion-based synthetic minority data augmentation
The initial dataset was constructed by cropping 2D rectangular regions of interest (ROI) from three consecutive axial NCCT slices for each hematoma, guided by automatically generated 3D segmentation masks. The selected slices comprised the slice with maximum hematoma area (Max) and its immediately adjacent superior (Max+1) and inferior (Max-1) slices. During cropping, images were standardized using a fixed window level of 40 HU and window width of 80 HU with linear intensity transformation. This process yielded an imbalanced training dataset of 654 HE and 2,655 NHE images (HE: NHE ≈ 1:4). To mitigate this class imbalance, we utilized Diffusion-UKAN, a generative model that learns the distribution of HE cases through iterative denoising of Gaussian noise via forward and reverse diffusion processes35, to generate synthetic HE images. The model was trained exclusively on HE samples from the training set to ensure high-fidelity synthetic generation of the minority class only. Two augmented training cohorts were subsequently constructed by combining the original real images with synthetic HE images: the UKAN-Balanced (HE: NHE = 1:1) and UKAN-Semibalanced (HE: NHE = 1:2) training sets. These augmented datasets were used to train downstream classifiers, aiming to improve model robustness and reduce bias from class imbalance.
The Diffusion-UKAN model used 1000 denoising steps with a linear noise schedule (β₁ = 1 × 10⁻⁴, β₂ = 0.02). Training images were resized to 256 × 256 pixels for unconditional generation. The model was trained for 1000 epochs using the Adam optimizer with a learning rate of 1 × 10⁻⁴, batch size of 32, and cosine annealing schedule. Synthetic image quality was evaluated using FID and IS39,40. The FID measures the similarity between feature distributions of real and synthetic images, with lower values indicating higher fidelity. Let \({\mu }_{r}\), \({\varSigma }_{r}\) and \({\mu }_{s}\), \({\varSigma }_{s}\) denote the empirical mean vectors and covariance matrices of real and synthetic image features, respectively, extracted by a pretrained Inception network.
where \(\text{Tr}\) denotes matrix trace and \({({\varSigma }_{r}{\varSigma }_{s})}^{1/2}\) represents the principal matrix square root.
The IS assesses both image quality and diversity through classification confidence, with higher scores indicating better quality. A pretrained Inception network classifies generated images to obtain a conditional label distribution \(p(y|x)\). High-quality images yield confident (low entropy) predictions, while a diverse set produces a high entropy marginal distribution \(p(y)\).
where \({D}_{{\rm{KL}}}\) is the Kullback-Leibler divergence measuring the difference between \(p(y|x)\) and \(p(y)\), and \(p(y)\) is the marginal class distribution of the generated dataset, defined as: \(p(y)=\frac{1}{N}\mathop{\sum }\limits_{i=1}^{N}p(y|{x}_{i})\).
To further validate the clinical authenticity of synthetic images, we conducted a comprehensive expert evaluation study. We randomly selected 100 real HE maximum-slice CT images from the original training set and 100 synthetic HE images from the UKAN-Semibalanced augmented training set (HE: NHE = 1: 2). All images were anonymized and randomly coded to ensure unbiased evaluation. Three senior radiologists from different medical centers, each with over 5 years of neuroimaging diagnostic experience, served as independent reviewers. The evaluation employed a standardized 5-point Likert scale assessing two critical dimensions: (1) overall image quality, evaluating image clarity, noise, and artifacts; (2) hematoma/surrounding structure feature authenticity, assessing the clinical plausibility of morphological and density features of both hematoma and surrounding structures. Detailed scoring criteria are provided in Supplementary Table S2. Additionally, reviewers performed binary classification (real or synthetic) for each image. Correctly identified (%) is based on majority vote (≥2/3), meaning that for each image, the final label was determined as the category selected by at least two out of three independent reviewers. To ensure consistency in evaluation criteria, all reviewers underwent standardized training before the formal assessment.
ViT-based patient-level hematoma classification
A patient-level automated classification framework was developed using Vision Transformer (ViT) to predict HE. For each patient, three consecutive axial 2D rectangular ROI were extracted from NCCT slices based on automated segmentation masks: the slice with maximum hematoma area (Max) and its immediately adjacent superior (Max+1) and inferior (Max-1) slices. Each slice, treated as an independent training sample, was divided into non-overlapping patches that were subsequently flattened and linearly projected into embedding vectors. These embeddings were augmented with positional and aggregation embeddings to preserve spatial context and facilitate global feature extraction41. All embedded tokens were processed through a standard transformer encoder, enabling the model to capture complex spatial and semantic relationships across the input slice. The resulting representations were passed through a multilayer perceptron head to generate a classification probability for each input slice. The final patient-level prediction was obtained by averaging the predicted probabilities of the three slices. In addition, to further identify the optimal classification model and validate the effectiveness of diffusion-based synthetic augmentation, we compared the predictive performance of three ViT models trained on datasets with different class ratios: ViT-1:1 trained on the UKAN-balanced dataset (1:1), ViT-1:2 trained on the UKAN-semibalanced dataset (1:2), and ViT-1:4 trained on the original real-imbalanced dataset (1:4).
The classification models were trained for 100 epochs using the Adam optimizer with a cosine annealing learning rate schedule that decayed from 2 × 10⁻⁴ to 1 × 10⁻⁴, and batch size of 256. Input images were resized to 224 × 224 pixels and models were initialized with ImageNet pretrained weights. Performance was evaluated on independent validation cohorts using AUC, accuracy, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV). To enhance model interpretability, Grad-CAM was applied to visualize discriminative regions contributing to model predictions42. The LayerNorm layer from the final transformer block served as the target layer, with token embeddings reshaped into spatial feature maps for Grad-CAM compatibility. The resulting heatmaps highlighted regions most relevant to HE classification, facilitating clinical interpretation and validation.
Statistical analysis
Statistical analyses were conducted using MedCalc 20.009 and Python 3.9. Normally distributed continuous variables were expressed as mean ± standard deviation and compared across cohorts using one-way ANOVA. Non-normally distributed continuous variables were presented as median (interquartile range) and analyzed using the Kruskal-Wallis test. Categorical variables were reported as frequencies (percentages) and compared using the chi-square test. Differences in AUC values between models were quantified as ΔAUC and tested for statistical significance using the DeLong test. In the expert radiological evaluation, differences in Likert scale scores between real and synthetic images were assessed using the Mann–Whitney U test. Inter-rater reliability of Likert ratings was quantified using the intraclass correlation coefficient [ICC (2,1), two-way random-effects model, absolute agreement]. Binary classification results were determined according to the majority principle (≥2/3 reviewers in agreement), and inter-rater consistency for classification was evaluated using Fleiss’ Kappa. A two-sided p-value < 0.05 was considered statistically significant. All deep learning models were implemented in Python using the PyTorch framework and trained on an NVIDIA A100 GPU (40 GB) running Ubuntu.
Data availability
The datasets analyzed in this study are available from the corresponding author upon reasonable request.
Code availability
The source code for this model is openly accessible via GitHub (https://github.com/DeepMedX/sICH_HE) under a non-commercial research license, permitting use exclusively for academic and scientific purposes.
References
Dowlatshahi, D. et al. Defining hematoma expansion in intracerebral hemorrhage: relationship with patient outcomes. Neurology 76, 1238–1244 (2011).
Morotti, A. et al. Intracerebral haemorrhage expansion: definitions, predictors, and prevention. Lancet Neurol. 22, 159–171 (2023).
Delcourt, C. et al. Hematoma growth and outcomes in intracerebral hemorrhage: the INTERACT1 study. Neurology 79, 314–319 (2012).
Davis, S. M. et al. Hematoma growth is a determinant of mortality and poor outcome after intracerebral hemorrhage. Neurology 66, 1175–1181 (2006).
Puy, L. et al. Intracerebral haemorrhage. Nat. Rev. Dis. Prim. 9, 14 (2023).
Ruff, I. M. et al. 2024 AHA/ASA Performance and Quality Measures for Spontaneous Intracerebral Hemorrhage: A Report From the American Heart Association/American Stroke Association. Stroke 55, e199–e230 (2024).
Wang, X. et al. Effects of blood pressure lowering in relation to time in acute intracerebral haemorrhage: a pooled analysis of the four INTERACT trials. Lancet Neurol. 24, 571–579 (2025).
Li, Q. et al. Ultra-Early Blood Pressure Reduction Attenuates Hematoma Growth and Improves Outcome in Intracerebral Hemorrhage. Ann. Neurol. 88, 388–395 (2020).
Rodriguez-Luna, D. et al. Effects of Achieving Rapid, Intensive, and Sustained Blood Pressure Reduction in Intracerebral Hemorrhage Expansion and Functional Outcome. Neurology 102, e209244 (2024).
Al-Shahi Salman, R. et al. Absolute risk and predictors of the growth of acute spontaneous intracerebral haemorrhage: a systematic review and meta-analysis of individual patient data. Lancet Neurol. 17, 885–894 (2018).
Winstein, C. J. et al. Guidelines for Adult Stroke Rehabilitation and Recovery: A Guideline for Healthcare Professionals From the American Heart Association/American Stroke Association. Stroke 47, e98–e169 (2016).
Demchuk, A. M. et al. Prediction of haematoma growth and outcome in patients with intracerebral haemorrhage using the CT-angiography spot sign (PREDICT): a prospective observational study. Lancet Neurol. 11, 307–314 (2012).
Law, Z. K. et al. Noncontrast Computed Tomography Signs as Predictors of Hematoma Expansion, Clinical Outcome, and Response to Tranexamic Acid in Acute Intracerebral Hemorrhage. Stroke 51, 121–128 (2020).
Morotti, A. et al. Standards for Detecting, Interpreting, and Reporting Noncontrast Computed Tomographic Markers of Intracerebral Hemorrhage Expansion. Ann. Neurol. 86, 480–492 (2019).
Morotti, A. et al. Using Noncontrast Computed Tomography to Improve Prediction of Intracerebral Hemorrhage Expansion. Stroke 54, 567–574 (2023).
Pszczolkowski, S. et al. Quantitative CT radiomics-based models for prediction of haematoma expansion and poor functional outcome in primary intracerebral haemorrhage. Eur. Radiol. 31, 7945–7959 (2021).
Arba, F. et al. Noncontrast Computed Tomography Markers of Cerebral Hemorrhage Expansion: Diagnostic Accuracy Meta-Analysis. Int J. Stroke 17, 835–847 (2022).
Liu, J. et al. Prediction of hematoma expansion in spontaneous intracerebral hemorrhage using support vector machine. EBioMedicine 43, 454–459 (2019).
Zhao, X. et al. Explainable CT-based deep learning model for predicting hematoma expansion including intraventricular hemorrhage growth. iScience 28, 112888 (2025).
Yalcin, C. et al. Hematoma expansion prediction in intracerebral hemorrhage patients by using synthesized CT images in an end-to-end deep learning framework. Comput. Med. Imaging Graph. 117, 102430 (2024).
Tran, A. T. et al. Uncertainty-aware deep-learning model for prediction of supratentorial hematoma expansion from admission non-contrast head computed tomography scan. NPJ Digit. Med. 7, 26 (2024).
Li, N. et al. A Deep Learning-Based Framework for Predicting Intracerebral Hematoma Expansion Using Head Non-contrast CT Scan. Acad. Radiol. 32, 347–358 (2025).
Li, Y. et al. Hematoma expansion prediction based on SMOTE and XGBoost algorithm. BMC Med. Inf. Decis. Mak. 24, 172 (2024).
Khazrak, I. et al. Addressing small and imbalanced medical image datasets using generative models: a comparative study of DDPM and PGGANs with random and greedy k sampling. arXiv https://arxiv.org/abs/2412.12532 (2024).
Fan, W. et al. The performance of artificial intelligence in image-based prediction of hematoma enlargement: a systematic review and meta-analysis. Ann. Med. 57, 2515473 (2025).
Wang, J. et al. Self-improving generative foundation model for synthetic medical image generation and clinical applications. Nat. Med. 31, 609–617 (2025).
Ktena, I. et al. Generative models improve fairness of medical classifiers under distribution shifts. Nat. Med. 30, 1166–1173 (2024).
Ironside, N. et al. Fully Automated Segmentation Algorithm for Hematoma Volumetric Analysis in Spontaneous Intracerebral Hemorrhage. Stroke 50, 3416–3423 (2019).
Yu, N. et al. A Robust Deep Learning Segmentation Method for Hematoma Volumetric Detection in Intracerebral Hemorrhage. Stroke 53, 167–176 (2022).
Zhong, J. W. et al. Deep learning for automatically predicting early haematoma expansion in Chinese patients. Stroke Vasc. Neurol. 6, 610–614 (2021).
Carvalho, M., Pinho, A. J. & Brás, S. Resampling approaches to handle class imbalance: a review from a data perspective. J. Big Data. 12, 71 (2025).
Khosravi, B. et al. Synthetically enhanced: unveiling synthetic data’s potential in medical imaging research. EBioMedicine 104, 105174 (2024).
Wang, G. et al. A deep-learning pipeline for the diagnosis and discrimination of viral, non-viral and COVID-19 pneumonia from chest X-ray images. Nat. Biomed. Eng. 5, 509–521 (2021).
Ma, J., Li, F. & Wang, B. U-Mamba: Enhancing Long-range Dependency for Biomedical Image Segmentation. arXiv https://arxiv.org/abs/2401.04722 (2024).
Li, C. et al. U-KAN makes strong backbone for medical image segmentation and generation. Proc. AAAI Conf. Artif. Intell. 39, 4652–4660 (2025).
Li, Q. et al. Island sign: an imaging predictor for early hematoma expansion and poor outcome in patients with intracerebral hemorrhage. Stroke 48, 3019–3025 (2017).
Rodriguez-Luna, D. et al. Multiphase CT angiography improves prediction of intracerebral hemorrhage expansion. Radiology 285, 932–940 (2017).
Lorensen, W. E. & Cline, H. E. Marching cubes: A high resolution 3D surface construction algorithm. ACM SIGGRAPH Computer Graphics, 163-169 (1987).
Heusel, M., Ramsauer, H., Unterthiner, T., Nessler, B. & Hochreiter, S. GANs Trained by a Two Time-Scale Update Rule Converge to a Local Nash Equilibrium. arXiv https://arxiv.org/abs/1706.08500 (2017).
Salimans, T. et al. Improved Techniques for Training GANs. arXiv https://arxiv.org/abs/1606.03498 (2016).
Dosovitskiy, A., Beyer, L., Kolesnikov, A., Weissenborn, D. & Houlsby, N. An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. arXiv https://arxiv.org/abs/2010.11929v1 (2020).
Selvaraju, R. R. et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. in Proceedings of the IEEE International Conference on Computer Vision. 618-626 (2017).
Acknowledgements
This study was supported by the Sichuan Foundation for Distinguished Young Scholars (2022JDJQ0049); 1.3.5 project of Center for High Altitude Medicine, West China Hospital, Sichuan University (Project No. GYYX24004). We gratefully acknowledge all participants and patients involved in this study.
Author information
Authors and Affiliations
Contributions
X.X., Z.W., J.P., and M.W. conceived the study design and supervised the project as co-corresponding authors. Q.Y., X.F., and J.L. contributed equally as co-first authors, conducting data analysis, model development, and drafting the manuscript. Y.N., S.L., W.J., F.L., X.Y., and Q.L. were responsible for data collection and quality control. Q.H., Q.L., Z.W., and J.P. assisted with data analysis. X.X., Z.W., J.P., and M.W. revised the manuscript critically. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yu, Q., Fan, X., Li, J. et al. An end-to-end deep learning pipeline for hematoma expansion prediction in spontaneous intracerebral hemorrhage based on non-contrast computed tomography. npj Digit. Med. 9, 39 (2026). https://doi.org/10.1038/s41746-025-02213-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02213-w