Abstract
Liver fibrosis is a reversible precursor to cirrhosis, and early detection is key to halting disease progression. Tongue diagnosis provides a non-invasive and cost-effective insight into internal health; however, its subjectivity limits clinical reliability. We developed TongVMoe, a multi-task deep learning model trained on 2202 tongue images from 1601 patients, to detect liver fibrosis and simultaneously classify seven key tongue features. The model achieved an area under the curve (AUC) of 0.8061, outperforming State-of-the-Art methods such as DiffMIC-v2 (0.6929), HorNet (0.7018), InceptionNeXt (0.7012), LSNet (0.6971), and TransXNet (0.7062). TongVMoe also demonstrated robust recognition of tongue features, with AUCs of 0.9752 for cracks and 0.9232 for greasy coating. Among these features, petechiae emerged as a significant clinical indicator, showing a strong correlation with liver fibrosis (χ² = 19.516, P < 0.001). We further integrated the model into a WeChat mini-program and simulated remote screening, achieving an accuracy of 77.8% and a sensitivity of 86.2%. These findings suggest that the TongVMoe has the potential to serve as an interpretable and mobile-compatible tool for the early detection and monitoring of liver fibrosis, particularly in resource-limited areas. Trial registration: Chinese Clinical Trial Registry (ChiCTR2100053676, registered 27 November 2021).
Similar content being viewed by others
Introduction
Liver disease is the eleventh-leading cause of death globally, with most fatalities attributed to complications associated with cirrhosis and hepatocellular carcinoma1. As a precursor to cirrhosis, liver fibrosis develops from chronic hepatic injury but can regress with appropriate aetiologic treatment2. Therefore, dynamic monitoring and early detection of liver fibrosis are crucial for controlling disease progression and reducing the overall disease burden3 effectively.
Although liver biopsy remains the reference standard for assessing fibrosis, its clinical utility is limited by its invasive nature and suboptimal intra- and inter-rater reliability4. Non-invasive tests (NITs), including serological tests5 and imaging examinations6, have been developed for the early diagnosis of liver fibrosis. However, these conventional diagnostic methods, with high acquisition and maintenance costs, are unsuitable for routine dynamic monitoring, particularly in primary care settings with limited technical capabilities and resources7.
Tongue inspection, as one of the most essential diagnostic methods in Traditional Chinese Medicine (TCM), provides practitioners with valuable insights into a patient’s overall health8, including the state of internal organs and the severity of illnesses. However, traditional tongue diagnosis heavily relies on practitioner experience and is inherently subjective. Recently, with the advancement of artificial intelligence (AI), tongue diagnosis has undergone significant evolution. Innovative technologies have been developed to enhance the objectivity and accuracy of tongue diagnosis8, enabling it to serve as a screening tool for the early detection of diseases such as breast cancer9 and non-alcoholic fatty liver disease10. Also, specialized diagnostic instruments are now being applied in research and clinical practices10 to expand the scope of tongue diagnosis. The Tongue and Face Diagnosis Analysis-1 (TFDA-1) instrument, developed by Shanghai University of TCM, has been used for diabetes11 and other diseases12. Nevertheless, because they rely on large models and high-performance hardware, TCM diagnostic instruments are expensive and relatively bulky, requiring patients to visit specialized institutions for evaluation.
Our study aims to develop an AI-powered tongue diagnosis system for home-based dynamic monitoring of liver fibrosis via mobile devices. We introduce the TongVMoe model, a multi-task interpretable framework that simultaneously identifies liver fibrosis and key tongue features, and further deploy it within a WeChat mini program to allow patients to upload tongue images and receive real-time diagnostic feedback (Fig. 1). This model was developed and validated on a prospective cohort using ultrasound elastography as the reference standard, and its diagnostic performance was rigorously benchmarked against a range of modern multi-task learning and state-of-the-art architectures. Complete methodological details regarding the study population, image preprocessing, network design, and statistical analysis are delineated in the Methods section. This approach facilitates accessible, continuous monitoring of liver fibrosis, paving the way for a practical follow-up tool suitable for everyday use at home.
The study consists of two stages. The first stage involves developing the TongVMoe model, which is trained using tongue images based on a multi-task model. The second stage involves remote health monitoring simulation by embedding the TongVMoe model into the WeChat mini-program.
Results
The study consists of two stages: the development of the TongVMoe model and the remote health monitoring simulation. The workflow is presented in Fig. 1.
Patient characteristics
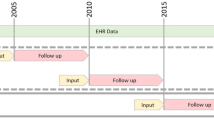
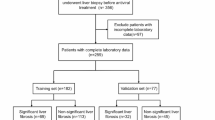
A total of 1601 patients who contributed 2202 tongue images were included in the AI model development phase. The training, validation, and test sets were comprised of 1280 (non-fibrosis: 927 [42.7 ± 10.7 years old], fibrosis: 353 [48.4 ± 12.2 years old]), 160 (non-fibrosis: 111 [42.0 ± 11.5 years old], fibrosis: 49 [47.0 ± 11.9 years old]), 161 (non-fibrosis: 107 [42.6 ± 11.8 years old], fibrosis: 54 [45.1 ± 12.2 years old]) patients, respectively. A total of 108 cases (from 103 individual patients) were enrolled in the remote health monitoring simulation, including 72 non-fibrosis cases and 36 fibrosis cases. Among them, 98 patients underwent the test once, whereas 5 patients completed the test twice at different time points. Table 1 summarizes the clinical characteristics of patients across different datasets.
Correlation between tongue features and liver fibrosis
The analysis of the correlation between various tongue features assessed by TCM expert and hepatic fibrosis status revealed no statistically significant associations between tongue color (χ2 = 4.836, P = 0.089), cracks (χ2 = 0.637, P = 0.425), tongue coating color (χ2 = 01.873, P = 0.171), tooth marks (χ2 = 0.038, P = 0.845), tongue coating thickness (χ2 = 0.155, P = 0.561), or greasy coating (χ2 = 0.536, P = 0.464) with LSM. In contrast, petechiae exhibited a highly significant association with liver stiffness (χ2 = 19.516, P < 0.001), highlighting its potential value as a clinical indicator for liver fibrosis.
Therefore, we conducted a subgroup analysis focusing on tongue petechiae to examine whether their presence affected the models’ predictive accuracy and to compare the performance of different models across these subgroups.
Diagnostic performance of the TongVMoe model
The TongVMoe model demonstrated robust performance across most tasks in the test set. For liver fibrosis prediction, the model achieved an accuracy of 77.98%, with a specificity of 87.42% and an AUC (Area under the curve) of 0.8061. In the task of tongue feature recognition, the model excelled in identifying cracks (Accuracy: 91.74%, AUC: 0.9752) and greasy coating (Accuracy: 90.37%, AUC: 0.9232) while displaying balanced performance in tongue coating color (Accuracy: 86.24%, AUC: 0.9310) and tooth marks (Accuracy: 87.16%, AUC: 0.9257). Additionally, the model achieved high specificity (90.80%) in the petechiae task, with an accuracy of 87.16% and an AUC of 0.8912. More details of the performance measurements of the test set are shown in Table 2. The receiver operating characteristic curves (ROC curves) for binary classification data, such as cracks, tongue coating color, tooth marks, greasy coating, petechiae, and LSM, are shown in Fig. 2a. For three-class classification data such as tongue color and tongue coating thickness, confusion matrices are illustrated in Fig. 2c and d (c: tongue color, d: tongue coating thickness)
a Receiver operating characteristic (ROC) curves of the TongVMoe model for cracks, tongue coating color, tooth marks, greasy coating, petechiae, and liver stiffness measurement (LSM) prediction tasks. b ROC curves of different deep learning models for liver fibrosis diagnosis based on tongue images. c Confusion matrix of the TongVMoe model for tongue color. d Confusion matrix of theTongVMoe model for tongue coating thickness classification.
Comparison of different models for the prediction of liver fibrosis from tongue images
We compared the TongVMoe with three multi-task learning approaches: Hard Parameter Sharing (HPS)13, Customized Gate Control (CGC)14, and DSelect-k15; and five backbone architectures: Diffusion-based Medical Image Classifier v2 (DiffMIC-v2)16, InceptionNeXt17, Large-Small Network (LSNet)18, TransXNet19, and HorNet20. Results showed the TongVMoe model exhibited superior diagnostic performance in diagnosing liver fibrosis from tongue images (Table 3). Specifically, its AUC was significantly higher than those of HPS (Accuracy: 69.72%, AUC: 0.6526) and CGC (Accuracy: 71.56%, AUC: 0.7375), and comparable to DSelect-k (Accuracy: 73.85%, AUC: 0.7643). Among the five backbone models, DiffMIC-v2 (Accuracy: 69.72%, AUC: 0.6929), InceptionNeXt (Accuracy: 71.10%, AUC: 0.7012), LSNet (Accuracy: 74.31%, AUC: 0.6971), TransXNet (Accuracy: 70.18%, AUC: 0.7062) and HorNet (Accuracy: 66.97%, AUC: 0.7018) all showed lower AUCs compared with the TongVMoe (p < 0.05). The ROC curves comparing these models are illustrated in Fig. 2b.
Subgroups (with/without petechiae) analysis of models in diagnosing liver fibrosis
To further assess the impact of tongue petechiae on fibrosis prediction, we stratified the test set into a petechiae group (44 tongue images) and a non-petechiae group (174 tongue images). In the non-petechiae group, the TongVMoe achieved the highest AUC (0.7902) with an accuracy of 78.16%, significantly outperforming HPS (AUC: 0.6277), CGC (AUC: 0.6711), DiffMIC-v2 (AUC: 0.6098), HorNet (AUC: 0.6105), InceptionNeXt (AUC: 0.6182), LSNet (AUC: 0.6584), and TransXNet (AUC: 0.6265), while demonstrating comparable performance to DSelect-k (AUC: 0.6882). In contrast, in the petechiae group, the TongVMoe reached an AUC of 0.8606, with less pronounced differences compared to other models. Overall, all nine models demonstrated superior diagnostic performance in the petechiae group compared to the non-petechiae group, with AUCs increasing by 7.04–33.50 percentage points. Notably, HorNet (+33.50), InceptionNeXt (+32.95), and TransXNet (+32.34) exhibited the most significant gains, whereas the TongVMoe showed a modest improvement (+7.04). More details are provided in Table 4.
Interpretability of the AI diagnosis
The interpretability of the TongVMoe lies in both its architecture and visualization results. Structurally, the model produced explicit predictions for each tongue feature alongside the fibrosis classification demonstrated previously, thereby revealing the underlying associations between observable features and disease states. For visual explanations of the model’s decision-making process, we applied EigenCAM21 to observe the regions attended by the TongVMoe model during prediction. We evaluated whether these areas corresponded to clinical features commonly examined by physicians. Analysis of the heatmaps revealed that the model’s highlighted regions closely matched the areas clinicians typically focused on. Figure 3a–d illustrate four representative cases where the model accurately diagnosed liver fibrosis by focusing on clinically relevant tongue features. For instance (Fig. 3a), when tongue cracks were present, the model focused on the cracked areas, consistent with the regions clinically assessed by physicians, underscoring the clinical validity of the AI model’s predictions.
a Tongue image exhibiting cracks. b Tongue image with tooth marks. c Tongue image showing petechiae. d Tongue image with thick and greasy coating.
Home monitoring simulation using the TongVMoe model
We integrated the TongVMoe model into a WeChat mini-program that passed the hospital’s review process. Figure 4 presents the interface of the developed mini-program for tongue image acquisition and liver fibrosis assessment. Overall, the mini-program achieved an accuracy of 77.8%, a sensitivity of 86.2%, and a specificity of 73.7%.
The mini-program comprises three sections: a the homepage, b the tongue image acquisition page, and c the result page. The original Chinese text in the interface has been translated into English, with translations provided according to the numbered annotations (①–⑨): ① Tongue Image-Based Liver Fibrosis Intelligent Evaluation System ② Upload tongue image ③ Rapid detection ④ Intelligent Analysis ⑤ Start ⑥ Ensure that the entire tongue is positioned within the shooting frame; Adequate lighting, no backlight, no exposure, no reflection; Make sure the tongue is not stained ⑦ Result ⑧ Tongue Analysis: Damp-heat syndrome, Tongue color: Red; Tongue coating color: Yellow; Cracks: None; Tooth marks: Yes; Tongue coating thickness: Medium; Greasy coating: Yes; Petechiae: None. ⑨ Screening Result: Liver fibrosis (+); Referral to hepatology is recommended.
Notably, among the 36 cases diagnosed with liver fibrosis, 27 were single-time measurements from individual patients, while the remaining nine cases came from five follow-up patients who underwent elastography twice at different time points. Of these five patients, four were consistently identified as having liver fibrosis in both examinations. In contrast, one patient initially tested negative but was later diagnosed with liver fibrosis during a follow-up visit three months later. Remarkably, the mini-program predicted liver fibrosis in this patient during the second visit as the disease progressed.
Discussion
In this study, we developed a deep-learning AI model, the TongVMoe, designed to screen for liver fibrosis and identify associated tongue features. Furthermore, we integrated the model into a WeChat mini-program and simulated a remote health monitoring scenario for liver fibrosis screening. Our study demonstrates that, with the assistance of AI, tongue images can be used not only for the preliminary screening of liver fibrosis but also as a tool for monitoring disease progression in patients with chronic liver disease during routine follow-ups.
Previously, we investigated the use of tongue images to develop an AI model capable of detecting liver fibrosis22. However, the model lacked interpretability regarding relevant tongue features, which undermined the trust of both physicians and patients in the diagnostic results and limited its clinical applicability. In addition, the training dataset was relatively small, resulting in suboptimal model performance. In the current study, we addressed both issues by enhancing the model’s ability to recognize tongue features and expanding the training dataset to boost overall performance and generalizability.
According to TCM theory, physicians can assess the condition of the liver by observing specific tongue features23. However, tongue diagnosis is highly subjective, and the diversity of tongue features further complicates its assessment. In this context, petechiae serve as a crucial diagnostic indicator. The subtle reddish-purple dots are considered in TCM to correspond to impaired circulation and stagnation of liver Qi and blood. From a modern perspective, this may parallel the microcirculatory disturbances that occur during the progression of hepatic fibrosis. Although visually less prominent than features such as cracks or greasy coating, petechiae provided stronger pathophysiological specificity for fibrosis. This may explain why all models exhibited better performance in the petechiae subgroup. In contrast, the non-petechiae subgroup, which presented more heterogeneous and nonspecific features, posed greater challenges for fibrosis detection. Notably, the TongVMoe model maintained stable performance across both subgroups, underscoring its generalizability in real-world clinical settings where tongue presentations are diverse. Overall, these findings reveal a gap between visual prominence and clinical relevance, suggesting that future model development should strike a balance between visually apparent features and clinically meaningful yet subtle ones.
Deep learning is often considered a “black box,” but Explainable Artificial Intelligence (XAI)24 offers solutions by providing objective and interpretable insights. Heatmaps generated by EigenCAM showed that the TongVMoe model consistently focused on clinically relevant tongue regions when predicting different features, supporting its visual interpretability. In addition to such visual evidence, the multi-task framework allows the model to reason about tongue features in parallel with disease diagnosis, thereby partially simulating the holistic and pattern-based diagnostic thinking of TCM physicians. This not only enhances transparency but also bridges the gap between AI decision-making and clinical reasoning, providing a more intuitive and trustworthy interpretation for practitioners.
We simulated a remote health scenario in which patients captured tongue images using their smartphones. Despite the potential decline in image quality, the AI model maintained robust performance and high sensitivity, enabling early detection of liver abnormalities. The follow-up results further support the model’s potential for early warning and longitudinal monitoring of liver fibrosis, highlighting its practical value in dynamic disease management, particularly in primary care and rural settings. Importantly, patient privacy is strictly protected, as no personal information is collected.
Our study has several limitations. First, we selected ultrasound elastography as the diagnostic reference standard due to its feasibility and reliability. Although ultrasound elastography has demonstrated promising results for the non-invasive assessment of liver fibrosis25, its accuracy may be influenced by factors such as the patient’s body mass index (BMI), the degree of liver inflammation, and the type of equipment used. These potential variabilities may introduce noise into the reference labels and influence the model’s performance evaluation. Second, the tongue images for the mini-program were all captured during clinic visits. Although the pictures were taken using patients’ smartphones, the lighting conditions were relatively uniform. This setting does not fully reproduce the complex lighting and positional variations that are likely to occur in real home environments, which may affect the model’s performance in broader real-world applications. Third, the dataset for simulating home monitoring was relatively small, mainly consisting of single-time measurements, which limited the validation of the model’s longitudinal predictive ability and dynamic monitoring performance. Moreover, the study was conducted in a single clinical center, and external validation across different populations and imaging conditions is still needed to ensure generalizability. In future studies, we plan to expand the dataset and further refine the tongue diagnosis model to improve its robustness and clinical applicability.
In conclusion, this study provides preliminary evidence supporting the use of deep learning-based tongue image analysis as a non-invasive approach to liver fibrosis screening. While further validation in larger and more diverse populations is required, our findings suggest that the AI model could potentially serve as an assistive tool for remote monitoring and follow-up management in chronic liver disease.
Methods
This prospective study was approved by the Institutional Committee on Ethics (ICE) for Clinical Research and Animal Trials at the First Affiliated Hospital of Sun Yat-sen University (Approval No. 2021464), and informed consent was obtained before collecting the tongue images from each participant. This study was registered at the Chinese Clinical Trial Registry (ChiCTR2100053676; registered on 27 November 2021) in accordance with the World Health Organization International Clinical Trials Registry Platform (WHO ICTRP) requirements.
Participants
Our study was conducted in two stages. Stage 1 involved the development of the AI diagnostic model. During this phase, patients who met the following criteria were enrolled between April 2021 and March 2024: (1) age ≥ 18 years; (2) a prior diagnosis of liver diseases, including chronic hepatitis, non-alcoholic fatty liver disease (NAFLD), abnormal transaminase levels and other related conditions, was confirmed in the Department of Gastroenterology at our hospital; (3) consented to tongue image collection. In Stage 2, participants were invited to test the mini-program during outpatient visits before undergoing liver elastography in the Department of Medical Ultrasonics between August 2024 and March 2025.
Tongue images collection and annotation
Participants were required to fast for 4–6 h before undergoing the ultrasound examination, ensuring that their tongues’ appearances were not affected by food or beverages. After receiving ultrasound elastography, patients were instructed to extend their tongues naturally for the collection of tongue images. In the first phase, photos were captured in no-flash mode using a Sony DSC-RX100 camera, and multiple images were taken for some patients to ensure data diversity. In the second phase, patients scanned a QR code to access the mini-program, then used their smartphones to capture a tongue image or upload a pre-existing one. Adhering to data privacy principles, our tongue diagnosis WeChat mini-program did not collect patient data, and diagnostic results were recorded on-site. There were no restrictions on the type of smartphone or camera resolution.
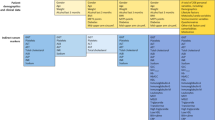
We invited a senior TCM physician with 20 years of clinical experience to annotate tongue images by identifying and labeling their tongue features. To standardize the process, we categorized the features into seven main types, each further divided into two or three subcategories based on tongue features commonly observed in clinical practice. Table 5 details these tongue features and their corresponding indications in TCM theory. The physician referred to the categories outlined in this table during the annotation process to ensure consistency.
Reference standard: ultrasound elastography examination
Ultrasound elastography was selected as the reference standard for assessing the degree of fibrosis in this study, with radiologists having at least 10 years of experience performing real-time two-dimensional shear wave elastography (2D SWE). The patient lay supine, and the transducer was positioned in the intercostal space to obtain clear images of the liver. The propagation of shear waves was measured, and the stiffness value, often expressed in kilopascals (kPa), was calculated. The procedure lasted for 10–20 min. The cutoff value of Liver stiffness measurement (LSM) was set to 726, dividing the patients into “fibrosis (LSM ≥ 7 kPa)” and “non-fibrosis (LSM < 7 kPa)”.
Image pre-processing
Tongue images were divided at an 8:1:1 ratio into training, validation, and test sets. Stratified sampling was conducted based on sub-attributes, ensuring that the class distribution within each attribute remained approximately consistent across all sets (Table 5). There were no overlapping patients between data sets. We employed the TongueSAM27 to segment the tongue images and minimize interference from irrelevant content. For the input tongue images, we performed a series of data augmentation and normalization operations to simulate various real-world scenarios, thereby enhancing the model’s generalizability and robustness.
The data pre-processing pipeline included the following steps: The input images were resized to 256 × 256 pixels using bilinear interpolation; Random padding was performed with a padding of 20 pixels and following with a random crop to 224 × 224 pixels; Horizontal flipping was applied with a probability of 0.5; Random affine transformations was applied including rotations (±10°) and translations (up to 5% of image dimensions); Sharpness adjustment was randomly applied with a sharpening factor of 2.0 (p = 0.3); Random transformations was performed with a randomized sequence including rotation (±10°, p = 0.5), perspective distortion (scale = 0.2, p = 0.5), sharpness adjustment (factor=2.0, p = 0.3), and Gaussian blur (kernel size 5–9, sigma 0.1–5.0, p = 0.4); Then images were converted to tensor format and normalized using dataset-specific statistics (mean = [0.765, 0.73, 0.746], standard deviation = [0.28, 0.315, 0.309]).
This diverse range of augmented samples improved the model’s generalization ability. In contrast, the validation and test phases employed only basic preprocessing (resizing, tensor conversion, and normalization) to ensure consistent evaluation conditions.
Network architecture
Our preliminary research22 demonstrated the effectiveness of the deep learning model for tongue diagnosis in liver fibrosis. However, the model lacks interpretability, limiting its ability to provide clinically meaningful explanations. This time, we employed the state-of-the-art visual VMamba28 for feature extraction and Multi-gate Mixture-of-Experts (MMoE)29 as the network backbone.
Feature Extraction Network: VMamba utilizes a 2D Selective Scan (SS2D) module to scan the image along four directions, efficiently capturing both local and global characteristics. It learns hierarchical visual representations by stacking Visual State Space (VSS) blocks, effectively serving diverse tasks such as tongue feature classification and liver fibrosis diagnosis.
Multi-task Learning with MMoE: The features extracted by VMamba are fed into the MMoE module. MMoE generates independent gates for each task to control the contributions of shared experts, modeling inter-task relationships and learning task-specific feature combinations. Given input features \(x\), the shared bottom layers learn a common representation \(z=f\left(x;\theta \right)\). Then, \(K\) expert networks \({\phi }_{k},k=1,2,...,K\) extract representations softly shared across tasks. The output for the\(\,i\)-th task is:
where\({g}_{i}^{k}\left(z\right)\) is the gating network for the \(k\)-th expert and the \(i\)-th task. The final prediction is:
Gradient Balancing with Conflict-Averse Gradient(CAGrad)30: CAGrad was employed to balance the gradients across the eight tasks and mitigate task imbalance. Let \({{\mathscr{L}}}_{i}\left(\theta \right)\) be the loss for task \(i\). The average loss is:
CAGrad finds an update vector \({\bf{d}}\) by solving:
where \({{\bf{g}}}_{0}=\frac{1}{K}{\sum }_{i=1}^{K}{{\bf{g}}}_{i}\) is the average gradient, and \(c\in [0,1)\) is a hyperparameter.
Finally, the task-specific features are input to the corresponding prediction heads for attribute classification and liver fibrosis diagnosis tasks. Figure 1 shows the schematic diagram of the model architecture.
Training process
The model was trained on a single NVIDIA H100 GPU with a batch size of 16 for 100 epochs. The optimizer was Adam with a learning rate of 1 × 10⁻⁵ and weight decay of 1 × 10⁻⁷. The learning rate scheduler employed a cosine annealing strategy (CosineAnnealingLR) with Tmax = 100 and ηmin = 1e-6. An early stopping mechanism was applied, halting training if the change in validation AUC over the past 10 epochs was less than 0.01. Other key hyperparameters included: Dropout Rate (0.5), Drop Path Rate (0.5), MTL CAGrad \({c}_{\alpha }\) (0.5), and specific initialization for MLP layers.
For binary classification tasks, a probability threshold of 0.5 was applied, where predictions with probabilities greater than or equal to 0.5 were classified as positive.
For multi-class classification tasks, the model produced a probability distribution over all categories through a softmax layer, and the class with the highest predicted probability was assigned as the final label.
Comparison with other multi-task learning models and modern architectures
To comprehensively assess the robustness and generalizability of our model, we compare it with both widely used multi-task learning methods and recently developed backbone designs. For multi-task learning, we include three representative models: HPS, CGC, and DSelect-k, each adopting different strategies for feature sharing across tasks. In addition, we benchmark our approach against five state-of-the-art backbone architectures: DiffMIC-v2, InceptionNeXt, LSNet, TransXNet, and HorNet. These models encompass diverse design philosophies, including diffusion-based learning (DiffMIC-v2), modernized convolutional networks (InceptionNeXt), integrated self-attention module (LSNet), hybrid token mixing (TransXNet), and high-order interaction modeling (HorNet). The performance of each model was evaluated using the same dataset, where we analyzed their effectiveness in diagnosing liver fibrosis.
Statistical analysis
Continuous variables were described as mean ± standard deviations (SD) and were compared by the t-test or Mann–Whitney U test. The association between individual tongue characteristics (e.g., presence of cracks or tooth marks) and liver fibrosis status was analyzed using the Pearson χ² test. The performances of all models were evaluated in terms of Area Under Curve (AUC), accuracy (ACC), precision (Pre), sensitivity (Sen), specificity (Spe), and F1-score(F1). AUC values were compared using the DeLong method. Acc, Pre, Rec, Spe, and F1 were compared using a paired samples t-test after verifying the normality of differences. Results with two-sided P-values of less than 0.05 indicated a statistically significant difference. Confidence intervals (CIs) were computed at a level of 95% using 1000 bootstrap samples. The analyses were conducted using Python 3.8 (Python Software Foundation).
Data availability
The tongue images and corresponding clinical metadata generated and used in this study are not publicly available due to patient privacy considerations. However, de-identified data can be made available from the corresponding author upon reasonable request and with approval from the Ethics Committee of the First Affiliated Hospital of Sun Yat-sen University.
Code availability
The code used for model development, training, and evaluation in this study is openly available at https://github.com/MedAI-UAIX/TongVMoe. Additional scripts related to data preprocessing and deployment can be obtained from the corresponding author upon reasonable request.
References
Devarbhavi, H. et al. Global burden of liver disease: 2023 update. J. Hepatol. 79, 516–537 (2023).
Taru, V., Szabo, G., Mehal, W. & Reiberger, T. Inflammasomes in chronic liver disease: hepatic injury, fibrosis progression and systemic inflammation. J. Hepatol. 81, 895–910 (2024).
Rinella, M. E. et al. AASLD practice guidance on the clinical assessment and management of nonalcoholic fatty liver disease. Hepatology 77, 1797–1835 (2023).
Soon, G. S. T. et al. Artificial intelligence improves pathologist agreement for fibrosis scores in nonalcoholic steatohepatitis patients. Clin. Gastroenterol. Hepatol. 21, 1940–1949.e1943 (2023).
Oh, J. H. et al. Diagnostic performance of non-invasive tests in patients with MetALD in a health check-up cohort. J. Hepatol. 81 https://doi.org/10.1016/j.jhep.2024.05.042 (2024).
Jung, K. S. & Kim, S. U. Clinical applications of transient elastography. Clin. Mol. Hepatol. 18, 163–173 (2012).
Chang, M. et al. Degree of discordance between FIB-4 and transient elastography: an application of current guidelines on general population cohort. Clin. Gastroenterol. Hepatol. 22, 1453–1461.e1452 (2024).
Jia, L. Y. et al. Modernizing tongue diagnosis: AI integration with traditional Chinese medicine for precise health evaluation. IEEE Access 12, 161670–161678 (2024).
Lo, L. C., Cheng, T. L., Chen, Y. J., Natsagdorj, S. & Chiang, J. Y. TCM tongue diagnosis index of early-stage breast cancer. Complement. Ther. Med. 23, 705–713 (2015).
Jiang, T. et al. Application of computer tongue image analysis technology in the diagnosis of NAFLD. Comput. Biol. Med. 135, 104622 (2021).
Zhang, J. F. et al. Diagnostic method of diabetes based on support vector machine and tongue images. Biomed. Res. Int. 2017, 7961494 (2017).
Shi, Y. L. et al. A new approach of fatigue classification based on data of tongue and pulse with machine learning. Front. Physiol. 12, 708742 (2022).
Caruana, R. Multitask learning: a knowledge-based source of inductive bias. Proc. Int. Conf. Mach. Learn. 41–48 (ICML, 1993).
Tang, H., Liu, J., Zhao, M. & Gong, X. Progressive layered extraction (PLE): a novel multi-task learning model for personalized recommendations. In Proc. ACM Conf. Recomm. Syst. (ACM, 2020).
Hazimeh, H. et al. Dselect-k: differentiable selection in the mixture of experts with applications to multi-task learning. Proc. NeurIPS 34, 29335–29347 (2021).
Yang, Y. J., Fu, H. Z., Aviles-Rivero, A. I., Xing, Z. H. & Zhu, L. DiffMIC-v2: medical image classification via improved diffusion network. IEEE Trans. Med. Imaging 44, 2244–2255 (2025).
Yu, W. H., Zhou, P., Yan, S. C., Wang, X. C. & IEEE Computer Society. InceptionNeXt: when Inception meets ConvNeXt. In Proc. IEEE/CVF Conf. Comput. Vis. Pattern Recognit. (CVPR), 5672–5683 (IEEE, 2024).
Wang, A., Chen, H., Lin, Z., Han, J. & Ding, G. LSNet: see large, focus small. In Proc. IEEE/CVF Conf. Comput. Vis. Pattern Recognit. (CVPR), 9718–9729 (IEEE, 2025).
Lou, M. et al. TransXNet: learning both global and local dynamics with a dual dynamic token mixer for visual recognition. IEEE Trans. Neural Netw. Learn. Syst. 36, 11534–11547 (2025).
Liu, Z., Rao, Y., Zhao, W., Zhou, J. & Lu, J. Efficient high-order spatial interactions for visual perception. In IEEE Trans. Pattern Anal. Mach. Intell. https://doi.org/10.1109/tpami.2025.3603181 (IEEE, 2025).
Rahman, A. N., Andriana, D. & Machbub, C. Comparison between Grad-CAM and EigenCAM on YOLOv5 detection model. In Proc. IEEE Int. Symp. Electron. Smart Devices (ISESD), 1–5 (IEEE, 2022).
Lu, X. Z. et al. Exploring hepatic fibrosis screening via deep learning analysis of tongue images. J. Tradit. Complement. Med. 14, 544–549 (2024).
Wang, R. R. et al. Non-invasive diagnostic technique for nonalcoholic fatty liver disease based on features of tongue images. Chin. J. Integr. Med. 30, 203–212 (2024).
Arrieta, A. B. et al. Explainable artificial intelligence (XAI): concepts, taxonomies, opportunities and challenges toward responsible AI. Inf. Fusion 58, 82–115 (2020).
Archer, A. J., Belfield, K. J., Orr, J. G., Gordon, F. H. & Abeysekera, K. W. M. EASL clinical practice guidelines: non-invasive liver tests for evaluation of liver disease severity and prognosis. Frontline Gastroenterol. 13, 436–439 (2022).
Herrmann, E. et al. Assessment of biopsy-proven liver fibrosis by two-dimensional shear wave elastography: an individual patient data-based meta-analysis. Hepatology 67, 260–272 (2018).
Cao, S., Wu, Q. & Ma, L. TongueSAM: a universal tongue segmentation model based on SAM with zero-shot. In Proc. IEEE Int. Conf. Bioinformatics Biomed. (BIBM), 4520–4526 (IEEE, 2023).
Liu, Y. et al. Vmamba: visual state space model. Neural Comput. Appl. 37, 103031–103063 (2024).
Ma, J. Q. et al. Modeling task relationships in multi-task learning with multi-gate mixture-of-experts. In Proc. ACM SIGKDD Int. Conf. Knowl. Discov. Data Min. (KDD), 1930–1939 (ACM, 2018).
Liu, B., Liu, X. C., Jin, X. J., Stone, P. & Liu, Q. Conflict-averse gradient descent for multi-task learning. Proc. NeurIPS. 34, 18878–18890 (2021).
Acknowledgements
This study was supported by the National Nature Science Foundation of China (NO: 82371983 and NO: 82272076), the Guangxi Key Research and Development Project (No. GuikeAB25069464), and the Guangxi Science and Technology Major Special Project (No. GuikeAA23073013).
Author information
Authors and Affiliations
Contributions
X.Z.L.: Conceptualization, Formal Analysis, Methodology, Data analysis and interpretation, Writing-Original Draft Preparation, Writing-Review & Editing; S. L.: Data Curation, Model development, Data analysis and interpretation, Writing-Original Draft Preparation; X.X.L.: Literature review, Model implementation and validation, Statistical analysis, Figure preparation, Writing-Manuscript Revision; Y.Z.: Data Curation, Methodology, Software, Writing-Original Draft Preparation; J.H.C.: Data Curation, Formal Analysis, Model Development; W.P.K.: Software, Validation, Model Development, Supervision; J.F.D.: Data Curation, Formal Analysis, Investigation, Methodology; M.Q.C.: Literature review, Statistical analysis, Writing-Manuscript Revision; W.L.: Data Curation, Methodology, Investigation; L.D.C.: Project Administration, Validation; Z.K.L.: Project Administration, Supervision, Model development, Writing-Manuscript Revision; B.G.S.: Project Administration, Supervision, Data Curation, Writing-Review & Editing; H.T.H.: Conceptualization, Model Development, Methodology, Supervision, Writing-Review & Editing; W.W.: Conceptualization, Project Administration, Supervision, Writing-Review & Editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lu, XZ., Liu, S., Lin, XX. et al. An AI-powered tongue image model for home-based monitoring of liver fibrosis. npj Digit. Med. 9, 67 (2026). https://doi.org/10.1038/s41746-025-02246-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02246-1