Abstract
Multidisciplinary tumor boards (MDTs) are central to cancer care but remain constrained by scarce experts and variable decision quality. EvoMDT employs a self-evolution loop that updates prompts, consensus weights, and retrieval scope based on expert feedback and outcome signals, improving robustness without sacrificing traceability. This matters clinically because MDT workloads and evidence shift over time, requiring adaptive yet auditable decision support. Agents perform domain-specific inference over lesion-level clinical data with structured knowledge retrieval; a consensus protocol resolves conflicts and generates traceable, evidence-linked recommendations. Evaluation spanned six public oncology QA benchmarks and four real-world datasets (breast, liver, lung, lymphoma), followed by single-blind physician assessment. Quantitative metrics (ROUGE, BERTScore) and automated safety checks assessed factuality and guideline concordance, while clinicians rated clinical appropriateness and usability. EvoMDT outperformed frontier Large Language Models (LLMs) baselines (e.g., Llama-3-70B, Claude-3, Med-PaLM 2), improving guideline concordance and semantic alignment with expert plans (BERTScore 0.62–0.68) and reducing safety violations. In physician review, EvoMDT achieved decision quality comparable to human MDTs while shortening response time by 30–40%. These results position EvoMDT as an interpretable, evidence-traceable framework that operationalizes AI reasoning for multidisciplinary oncology practice and offers a scalable foundation for trustworthy, lesion-level precision cancer care.
Similar content being viewed by others
Introduction
Precision oncology now requires index-lesion decisions that integrate heterogeneous evidence under operational constraints. Lesion-level management must reconcile genomics, histopathology, and imaging with RECIST/iRECIST anchors, together with organ function and patient context, within narrow clinical timelines1,2. In routine MDT practice, these lesion-level decisions are multi-constraint trade-offs made under strict time limits: clinicians must balance therapeutic intent and sequencing with patient-specific organ reserve, comorbidities, prior therapies, and feasibility constraints. As case volume rises, complex longitudinal histories and competing objectives are often compressed into short discussion windows, and the most consequential reasoning steps may remain implicit—for example, why one guideline-concordant option was prioritized over another, which safety gates were checked, and how disagreements were resolved. When such trade-offs and safety checks are not captured in a structured form, it becomes difficult to audit decisions retrospectively, communicate rationale across specialties, or ensure consistent quality outside high-resource tumor boards. Multidisciplinary Team (MDT) deliberations remain the reference standard and are associated with guideline adherence and improved outcomes across tumor types3,4,5. For example, a meta-analysis that aggregated 37 multivariable-adjusted studies covering over 56,000 cancer patients found that the MDT consultation group had superior overall survival compared to the non-MDT group, particularly in subgroup analyses of liver and breast cancer, with adjusted hazard ratios (HR) of 0.82 and 0.86, respectively6. Additionally, in a cohort study of colorectal cancer, patients managed with MDT had a 3-year disease-free survival rate and overall survival rate of 78% and 83%, respectively, which were significantly higher than the 65% and 69% observed in the non-MDT group7.
However, rising case volumes and workforce pressures impose throughput limits, variable depth of discussion, and delays that are increasingly difficult to absorb1,8,9. Early clinical decision support systems (CDSS) offered rule-based standardization, but in many deployments they function primarily as stage-to-regimen lookup tables or alert engines rather than end-to-end decision workflows: they can list eligible options and flag contraindications, yet they rarely integrate interacting guideline families and longitudinal lines of therapy within a single patient, and they seldom document why one reasonable option was selected over alternatives in the face of competing objectives. More recent LLM-style assistants can produce fluent narrative recommendations, but without explicit role accountability, enforced safety gating, and auditable conflict resolution, a single opaque answer remains difficult to verify and operationalize in MDT settings10,11. The field lacks scalable tools that preserve MDT-grade, lesion-specific reasoning while improving efficiency, transparency, and auditability.
Across oncology practice, four representative malignancies were selected to encompass the full spectrum of lesion-level clinical complexity. Hepatocellular carcinoma (HCC) couples tumor burden with hepatic reserve, requiring BCLC staging with Child–Pugh/ALBI gating, portal-vein tumor thrombus strategies, transplant/bridging logistics, and ICI caveats in transplant contexts8. Lung adenocarcinoma demands actionable driver integration and PD-L1 thresholds within TNM-8 anatomy, with peri-operative IO/TKIs and lesion-directed therapy for oligometastatic brain disease3,8. Lymphoma decisions hinge on histology and risk (Lugano/Ann Arbor; IPI/CNS-IPI), marrow/CNS work-ups, and lines of therapy spanning chemo-immunotherapy to cellular therapies4. Breast cancer requires biologic subtype fidelity (ER/PR/HER2, Ki-67) to steer (neo)adjuvant vs metastatic pathways, alongside fertility, menopause, and survivorship planning4. Together these settings span anatomic staging, organ-function constraints, targeted/immune therapeutics, and hematologic complexity—a representative testbed for index-lesion decision systems.
LLMs have advanced biomedical synthesis, but monolithic models often misalign with MDT’s role-based reasoning and clinical governance. Although recent work demonstrates competence on standardized knowledge and patient-facing queries12,13,14,15, a single-model architecture typically collapses diagnosis, treatment selection, safety checks, and follow-up into one response, making it difficult to enforce role accountability, to surface trade-offs, or to audit how safety constraints were applied in complex cases11,12,13,14. In parallel, conventional CDSS implementations provide valuable standardization, yet their rule-centric designs commonly operate as stage-to-regimen lookup tables and alert engines; they can enumerate options or flag contraindications, but they rarely integrate interacting guideline families and longitudinal treatment lines into an explicit, end-to-end plan, and they seldom retain the rationale for choosing one reasonable option over another in the face of competing objectives8,9. Retrieval-augmented and agentic patterns improve grounding and task fidelity, yet lesion-centric, oncology-focused multi-agent systems that jointly provide auditable evidence provenance, explicit safety gates, consensus mechanisms, and structured trade-off handling remain scarce16,17,18,19.
Clinically, a useful system must pre-structure MDT thinking at the lesion level and output implementable, traceable plans. Recommendations should (i) respect staging thresholds (e.g., TNM-8, BCLC, Lugano), (ii) enforce organ-function gates (e.g., Child–Pugh/ALBI; performance status), (iii) surface contraindications and drug–drug/organ constraints, and (iv) specify monitoring cadence with RECIST/iRECIST or hematologic anchors—each citation-backed with native guideline grades and years. Such structured outputs enable pre-MDT triage, reduce variability for junior teams, accelerate time-critical decisions, and create provenance logs aligned with CONSORT-AI/SPIRIT-AI and DECIDE-AI for governance and equitable care20,21.
We present EvoMDT, a self-evolving multi-agent system that mirrors MDT practice at the index-lesion level. Instead of a single oracle model, EvoMDT coordinates specialized Diagnostic, Treatment, Safety, and Monitoring agents with a Coordinator that resolves conflicts by safety-first logic and consensus strength. Each agent operates on a shared, pan-cancer, lesion-centric knowledge base with retrieval-augmented reasoning and explicit evidence provenance, so that every recommendation can be traced back to guideline- or literature-derived atomic rules. Operating on this substrate, EvoMDT produces structured, evidence-traceable recommendations with explicit role accountability, making its behavior closer to a digital tumor board than to a conventional rule-based CDSS or monolithic LLM. As illustrated in Fig. 1, EvoMDT organizes the end-to-end MDT-style workflow from lesion-level inputs to evidence-grounded recommendations via explicit agent roles, coordination, and traceable rationale. We validate EvoMDT using a multi-stage design: an expert-curated cohort of synthetic cases across lung adenocarcinoma, breast cancer, HCC, and lymphoma to stress lesion-level diversity22,23,24, and a single-blind, multi-physician head-to-head evaluation on six predefined dimensions. EvoMDT attains competitive decision quality with pronounced gains in evidence use and plan completeness, and materially shorter end-to-end times, supporting a shift from single-model “oracle” AI to collaborative agent systems that better reflect real-world oncology decision-making13,15,20,21,24.
a Evidence overload versus clinician bandwidth across guidelines, literature, and patient data; safety-critical, multi-objective decisions must balance guideline concordance, accuracy, safety, and implementability. b What has been tried (rule-based CDSS, knowledge graphs, ML on structured EHR, single-agent LLM, naïve multi-agent LLM) and what is missing. c Gap-to-solution mapping: EvoMDT integrates ontology-grounded RAG, a dedicated Safety Agent, coordinator-mediated consensus, longitudinal Monitoring Agent, self-evolution, and three-tier validation. RAG retrieval-augmented generation, EHR electronic health record.
Results
Experimental setup
All experiments were conducted on a high-performance server running the Ubuntu operating system, equipped with eight NVIDIA RTX 4090 GPUs (24 GB memory each) that enabled large-scale model training and parallel inference. The software environment was based on Python 3.10 and PyTorch 2.1.0, with GPU acceleration provided by CUDA 12.1 and cuDNN. Additional deep learning libraries such as Hugging Face Transformers, scikit-learn, and timm were employed to support model construction, optimization, and evaluation. To ensure reproducibility, random seeds were fixed across all experiments, and strict version control was applied to data loading, distributed training, and inference pipelines.
Benchmark verification results
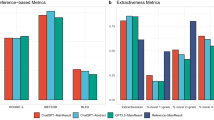
We evaluated the generalization capability of EvoMDT across six medical question-answering datasets, including three standardized multiple-choice benchmarks (MedQA, MedQA-Cancer, MedMCQA-Cancer) and three free-response datasets (MedQuAD, MedQARo, and ROND). Each dataset contains physician-verified ground truth, and we assume that a model output more semantically aligned with the reference indicates higher decision accuracy. All metrics were normalized to ensure comparability across different scoring systems, allowing the radar chart in Fig. 2a to jointly represent both accuracy and BERTScore. As shown in Fig. 2a, EvoMDT achieved the highest performance across all six datasets. For multiple-choice benchmarks (Fig. 2b, left), EvoMDT reached 0.86 on MedQA, 0.89 on MedQA-Cancer, and 0.75 on MedMCQA-Cancer, outperforming Llama3-70B (0.76/0.76/0.63) and Claude-3 (0.79/0.75/0.69). These results demonstrate strong robustness and consistency in oncology-related reasoning and structured clinical decision tasks. For free-response datasets (Fig. 2b, right), EvoMDT achieved the best BERTScores on MedQARo (0.66), MedQuAD (0.63), and ROND (0.68), surpassing Claude-3 (0.61/0.59/0.65) and Med-PaLM 2 (0.62/0.63/0.63). While BERTScore measures only linguistic similarity rather than clinical logic, the consistent advantage indicates that EvoMDT produces responses semantically closer to physician-level treatment plans. Figure 2c presents a representative example from MedQA, where the model correctly identified Cyclophosphamide as the causative agent of hemorrhagic cystitis in a patient with non-Hodgkin lymphoma following chemotherapy. This highlights EvoMDT’s capability for pharmacologic reasoning and clinically consistent inference.
a Comprehensive comparison of model performance across six medical question–answering benchmarks, including MedQA, MedQA-Cancer, MedMCQA-Cancer, MedQuAD, MedQARo, and ROND. The radar chart shows that EvoMDT consistently outperforms baseline large language models (Llama3-70B, Claude-3, and Med-PaLM 2) in both accuracy-based and semantic similarity metrics. b Quantitative performance comparison on standardized medical benchmarks (left) and custom oncology datasets (right). EvoMDT achieved the highest accuracy (0.86–0.89) on multiple-choice datasets and superior BERTScores (0.63–0.68) on generative question–answering tasks, indicating improved semantic fidelity to expert responses. c Representative multiple-choice example from the MedQA dataset. The model correctly identifies cyclophosphamide (d) as the causative agent of hemorrhagic cystitis in a patient with non-Hodgkin lymphoma following chemotherapy, demonstrating clinically consistent pharmacologic reasoning.
Multi-cancer validation on real-world datasets
All performance metrics in this study were computed with respect to expert-defined reference answers. Each dataset contained physician-verified diagnostic decisions and treatment plans, allowing us to assume that model-generated outputs closer to these reference plans represent higher clinical accuracy and appropriateness. Based on this assumption, we employed multiple evaluation metrics, including lexical overlap (ROUGE), semantic similarity, and AI-based composite scores (Fig. 3). Notably, the relatively low ROUGE scores (approximately 0.04–0.06) are expected, as ROUGE measures only surface-level word overlap rather than underlying clinical reasoning or semantic equivalence. In contrast, the semantic similarity values ranging from 0.62 to 0.68 indicate that EvoMDT captures expert-level treatment logic even when lexical alignment is limited.
a Overall performance by cancer type across evaluation metrics including ROUGE-1, ROUGE-L, semantic similarity, and AI composite score; b AI-based evaluation scores using DeepSeek, comparing model predictions with two baselines (simple baseline and random baseline); c Recall distributions for ROUGE-1 and ROUGE-L metrics; d Semantic similarity of treatment plans measured by cosine similarity.
To provide meaningful reference points, we introduced two baselines. The first, a random baseline, was generated by shuffling treatment plans among patients with the same cancer type. This tests whether the model can differentiate between individual cases that share the same disease label but require distinct therapeutic strategies. The second, a simple baseline, consisted of deliberately incorrect instructions such as “take cold medicine,” serving as a lower-bound reference for obviously invalid outputs. As shown in Fig. 3b–d, EvoMDT consistently outperformed both baselines across all cancer types and evaluation metrics. For example, in AI-based scoring, the model achieved 6–9 points across breast, liver, lung, and lymphoma datasets, whereas the random and simple baselines scored around 2–3 points. Similarly, higher ROUGE recall and cosine similarity further confirm that EvoMDT is capable of generating individualized treatment recommendations that align closely with expert clinical reasoning (Fig. 3a). Collectively, these findings support the model’s robustness and reliability across diverse cancer types.
Single-blind physician assessment of clinical decision quality
In the single-blind evaluation of 405 standardized case dossiers (n = 405) derived across four cancer types (lung adenocarcinoma, breast cancer, hepatocellular carcinoma, and lymphoma), EvoMDT demonstrated competitive performance across all six evaluation dimensions. The system achieved a composite mean score of 3.44 (95% CI, 3.29–3.59), placing it within the mid-to-upper range of the five participating physicians (Fig. 4a–c). At the dimension level, EvoMDT showed differentiated strengths (Fig. 4a, d). Response Completeness and Safety & Ethics were highest (both 3.68, 95% CI, 3.58–3.78; high-score rate ≥4 points: 63.7% for each). Clinical Appropriateness, Response Efficiency, Evidence Utilization, and Patient-Centeredness were comparable (each 3.32, 95% CI, 3.23–3.42; high-score rate 43.5%). These patterns indicate that the system consistently preserved core clinical content and safety safeguards while maintaining balanced performance in communication efficiency and patient-centric elements.
a Six-dimensional score distribution (Likert 1–5) pooled across four tumor types. b Response time comparison between five physicians and EvoMDT (minutes; median with IQR; pairwise Mann–Whitney U with FDR adjustment). c Overall performance ranking: composite mean score (95% CI) by rater and cancer type. d Multi-dimensional radar profile showing mean scores on six dimensions. e Inter-rater reliability by dimension: ICC (2,1) with 95% CI (two-way random, absolute agreement). f Inter-dimension correlation matrix (Spearman’s ρ; FDR-adjusted significance). ICC intraclass correlation coefficient, IQR interquartile range, FDR false discovery rate.
EvoMDT’s principal operational advantage was response time (Fig. 4b). The system generated comprehensive multidisciplinary recommendations with a median of 10.6 min (IQR, 9.5–12.0; mean ± SD, 10.8 ± 1.7), significantly faster than all five physicians across seniority levels (medians 15.8–18.9 min). The relative reduction was computed against physician medians. Mann–Whitney U tests confirmed large and statistically robust differences between EvoMDT and each physician (all P < 10⁻⁹⁸; rank-biserial r = 0.74–0.85). Notably, despite this speed advantage, EvoMDT retained high Response Completeness (mean 3.68), suggesting efficient integration of multi-source evidence without compromising thoroughness. These times were measured under the standardized case-dossier setting used throughout this study; richer raw clinical inputs would require additional upstream structuring/triage, which may increase end-to-end latency and is not benchmarked here. Inter-rater reliability, assessed as ICC(2,1) (two-way random effects, absolute agreement, single rater), showed good-to-excellent agreement across all dimensions (Fig. 4e). Estimates ranged 0.79–0.89, with all lower 95% CI bounds >0.75: Safety & Ethics 0.89 (95% CI, 0.85–0.93), Response Completeness 0.86 (0.82–0.90), Clinical Appropriateness 0.85 (0.81–0.89), Response Efficiency 0.84 (0.80–0.88), Evidence Utilization 0.83 (0.79–0.87), and Patient-Centeredness 0.79 (0.75–0.83). These findings indicate the rubric was well-calibrated and reproducible across clinicians with diverse backgrounds.
Spearman correlations revealed predominantly weak associations (|ρ| < 0.10 for most pairs; Fig. 4f), supporting the multi-dimensional independence of the framework. The strongest observed association was Evidence Utilization × Patient-Centeredness (ρ = 0.095, P < 0.05), suggesting modest convergence between evidence-based reasoning and individualized care. Response Completeness × Clinical Appropriateness showed a weak negative correlation (ρ = −0.068), hinting that maximal comprehensiveness may occasionally trade off with strictly focused relevance. Response Efficiency displayed near-zero correlations with other dimensions (ρ = −0.054 to 0.060), confirming it captures a distinct “speed vs depth” facet. Performance was stable across the four cancer types (The specific results are shown in Table 1). Composite means clustered between 3.3 and 3.5 without statistically significant between-cancer differences (Kruskal–Wallis P > 0.10), underscoring the generalizability of EvoMDT’s multi-agent architecture across heterogeneous oncologic contexts and guideline ecosystems.
Representative case demonstrations of clinical reasoning
To further illustrate the reasoning capability and clinical decision quality of EvoMDT in oncology, Fig. 5 showcases representative real-world cases drawn from the four validated cancer cohorts. Each example pairs the model’s structured recommendation with the corresponding expert consensus treatment plan, serving as a gold-standard reference for comparison. We clarify how this “model-generated reasoning” is produced by role-separated intermediate outputs and coordinator synthesis. In the representative HCC case shown in Fig. 5a, the Diagnostic Agent organizes the lesion-centric description (tumor context and background liver disease) into a structured problem representation; the Treatment Agent proposes the primary management strategy and key adjunct components consistent with the retrieved guidelines; the Safety Agent acts as a gating layer that checks feasibility and major contraindication risks before the plan is finalized; and the Monitoring Agent specifies phase-adapted follow-up and surveillance logic. The Coordinator Agent then reconciles these components into a single coherent MDT-style recommendation, resolving any disagreements by prioritizing safety and implementability when trade-offs emerge. This division of responsibilities is intended to mirror real MDT workflows, making the reasoning pathway inspectable rather than collapsing all decisions into a single “oracle” answer. In Fig. 5a (hepatocellular carcinoma), the case involves a patient with HBV-related cirrhosis and a small lesion located in liver segment S6. EvoMDT accurately recommends laparoscopic liver resection combined with Entecavir antiviral therapy, fully consistent with the expert decision. This concordance highlights the model’s ability to integrate imaging-derived anatomical information with antiviral management principles. Overall, EvoMDT not only reproduces clinically appropriate treatment components but also generates transparent, human-interpretable reasoning that mirrors expert decision logic. These representative cases underscore the system’s potential as a trustworthy AI collaborator in multidisciplinary oncology decision-making.
Each panel presents a representative patient case with structured clinical data, model-generated treatment reasoning, and the corresponding expert reference plan used for evaluation. a Hepatocellular carcinoma (HCC). b Breast cancer. c Lung adenocarcinoma. d Lymphoma.
Discussion
This study presents EvoMDT, a self-evolving multi-agent system designed to emulate multidisciplinary oncology decision-making through transparent, evidence-based reasoning. By integrating structured knowledge retrieval, lesion-level clinical data, and multi-agent consensus mechanisms, EvoMDT extends beyond conventional large language models that operate primarily at the text-response level. In practical terms, EvoMDT’s contribution is not simply “a stronger model,” but a different operating contract for clinical decision support. Rule-based CDSS are dependable for standard pathways and contraindication checks, yet they typically cannot represent the negotiation MDTs actually perform when multiple guideline-supported options compete under patient-specific constraints; importantly, they also struggle to preserve a decision rationale that can be reviewed across specialties. By structuring deliberation into role-specialized agents and forcing evidence provenance at each step, EvoMDT makes the trade-offs and safety checks explicit and reviewable. Likewise, compared with monolithic LLM assistants, EvoMDT separates responsibilities (e.g., diagnosis vs. therapy vs. safety vs. monitoring), surfaces disagreements rather than hiding them inside a single narrative, and resolves them through a coordinator with safety-first logic—closely matching how real tumor boards protect against individual blind spots. Our multi-layer validation—spanning public medical QA benchmarks, real-world multi-cancer datasets, and single-blind physician evaluations—demonstrated that EvoMDT consistently outperformed state-of-the-art foundation models such as Llama3-70B, Claude-3, and Med-PaLM 2 across both semantic similarity and expert-aligned decision quality. Importantly, its outputs not only achieved high quantitative agreement with expert consensus plans but also exhibited interpretable, stepwise reasoning reflective of real-world clinical logic. While ROUGE/BERTScore quantify textual/semantic alignment, for unstructured, dynamic clinical narratives, EvoMDT pre-structures inputs with lesion-level templates and ontology-grounded RAG, and maintains an auditable evidence trail; future deployments will integrate full-text notes and imaging reports via controlled parsers with human-in-the-loop verification25. Clinical adequacy ultimately requires human judgment and safety verification; our single-blind physician ratings and low safety-violation rates complement benchmark gains, aligning with CONSORT-AI/DECIDE-AI emphases on human-aligned evaluation26. Together, these findings establish EvoMDT as a promising framework for safe, explainable, and generalizable AI-assisted decision-making in oncology.
Compared with traditional single-agent or monolithic large language model (LLM) approaches, the multi-agent collaborative framework proposed in this study offers several distinct advantages for clinical decision-making. First, by decomposing the decision process into specialized yet interdependent agents—covering diagnostic reasoning, treatment planning, safety surveillance, evidence retrieval, and consensus coordination—EvoMDT achieves a modular, interpretable, and auditable reasoning architecture. This design mirrors the workflow of real multidisciplinary tumor boards, where different specialties contribute complementary perspectives under a unified coordination protocol. Second, the consensus-based conflict resolution mechanism enables the system to dynamically balance safety, efficacy, and evidence strength, reducing the risk of overconfident or hallucinated recommendations that often arise in single-agent LLMs27. Third, through its retrieval-augmented and evidence-traceable knowledge layer, EvoMDT ensures that every treatment suggestion is grounded in verifiable clinical guidelines or peer-reviewed literature, enhancing transparency and trustworthiness. Finally, the system’s self-evolving learning loop, informed by real-world feedback and physician evaluations, allows adaptive refinement without compromising regulatory traceability—an essential prerequisite for clinical translation.
The three-layer validation collectively demonstrates the robustness, clinical reliability, and interpretability of EvoMDT, while highlighting opportunities for refinement. Layer-1 confirmed strong factual accuracy and semantic reasoning across benchmarks, yet underscored that high benchmark scores alone cannot guarantee bedside safety—necessitating further clinical validation. Layer-2, based on real-world multi-cancer data, verified that EvoMDT achieved high guideline concordance, plan-level agreement, and evidence traceability while maintaining low safety-violation rates, proving its capacity to generate clinically coherent and traceable decisions. Layer-3, involving single-blind physician ratings, validated that EvoMDT’s recommendations were comparable to expert outputs across six dimensions, with notable strengths in Safety & Ethics and Completeness, though raters observed occasional over-detailed plans in complex cases. The three layers form a dialectical evaluation system—balancing quantitative rigor with qualitative insight. Benchmark tests validate the model’s foundational reasoning, real-world datasets test contextual generalization, and physician assessments ensure human-aligned interpretability.
The clinical implications of EvoMDT extend beyond algorithmic performance to the redefinition of how multidisciplinary oncology decisions can be supported by artificial intelligence. By aligning with the workflow and evidence hierarchy of real-world tumor boards, EvoMDT offers a scalable digital counterpart to the human MDT, capable of providing structured reasoning, traceable evidence chains, and reproducible recommendations even in resource-limited settings28,29. In regions or hospitals lacking full multidisciplinary expertise, the system can function as a virtual MDT facilitator, ensuring that treatment decisions remain consistent with up-to-date clinical guidelines and real-world practice patterns30. Moreover, its transparent reasoning architecture enables human–AI collaboration, allowing clinicians to audit, refine, and customize recommendations according to patient-specific needs, rather than replacing professional judgment. From a translational standpoint, EvoMDT establishes a prototype for clinically compliant, explainable AI systems that can evolve with accumulating medical knowledge and feedback loops. Its multi-agent modular design facilitates regulatory traceability and selective module validation—features essential for future integration into hospital information systems or clinical trial workflows.
While EvoMDT demonstrates promising performance across benchmarks and real-world validations, several limitations remain. The retrospective datasets, though multi-center and multi-cancer, are still limited in scale and geographic diversity, necessitating broader external validation to ensure generalizability. Although records were clinician-reviewed and standardized for consistency, residual site-specific practice patterns and documentation differences are unavoidable in retrospective multi-center cohorts; therefore, we interpret real-world performance as reflecting both clinical complexity and pragmatic heterogeneity rather than a fully harmonized trial-like setting. Our focus on lung adenocarcinoma, breast cancer, HCC, and lymphoma was intentional. Together, these disease groups occupy a central position in real-world MDT practice, combining high case volumes with guideline-intensive, multi-constraint decisions that are difficult to standardize and audit at scale. At the same time, they expose EvoMDT to four distinct decision grammars and MDT structures, from organ-based boards to hematology-focused conferences, which makes them more than a convenience sample. Nevertheless, we did not evaluate EvoMDT in other important domains such as non-HCC gastrointestinal malignancies, gynecologic cancers, sarcomas, or rare tumors. The present results should therefore be interpreted as evidence that the EvoMDT framework can operate across several contrasting and representative MDT archetypes, rather than as proof of coverage for the entire oncology spectrum. Extending and prospectively validating EvoMDT in additional disease groups is an important direction for future work. The current knowledge base relies primarily on published guidelines and literature; future work should incorporate dynamic real-world evidence from clinical registries and prospective trials to enhance adaptability. Although single-blind physician evaluations confirmed clinical reliability, prospective deployment and patient-outcome studies are required to assess real-world impact and trust calibration. Moreover, the observed response-time advantage was evaluated on structured MDT-style case dossiers; evaluating end-to-end latency on raw, high-complexity clinical records (including unstructured notes and richer longitudinal context) remains an important direction for future prospective deployments. Technically, current interpretability mechanisms—while improving transparency—may not fully capture complex clinical reasoning or ethical nuances. Future research should emphasize multi-modal data fusion, human-in-the-loop continual learning, and regulatory-grade auditability, advancing EvoMDT toward a trustworthy, adaptive, and ethically aligned clinical decision-support system.
This study introduces EvoMDT, a self-evolving multi-agent framework that emulates the collaborative reasoning of multidisciplinary oncology teams through transparent, evidence-grounded decision-making. By integrating structured knowledge retrieval, lesion-level clinical data, and cross-agent consensus mechanisms, EvoMDT bridges the gap between general-purpose language models and clinically accountable decision-support systems. Multi-layer validation—spanning public medical benchmarks, real-world multi-cancer datasets, and single-blind physician assessments—demonstrated the system’s robustness, interpretability, and alignment with expert clinical logic. Beyond achieving high quantitative concordance, EvoMDT provides human-auditable reasoning and verifiable evidence traces, offering a viable path toward safe, explainable, and adaptive AI collaboration in precision oncology. As future iterations incorporate prospective validation and multi-modal clinical data, EvoMDT holds the potential to serve as a foundational infrastructure for trustworthy, next-generation clinical decision support in cancer care.
Methods
Knowledge base construction
We developed a single, pan-cancer, lesion-centric knowledge base (KB) that acts as the evidence substrate for EvoMDT’s retrieval-augmented, role-based reasoning. Rather than a static document repository, the KB stores atomic recommendation units—diagnostic and staging rules, therapeutic algorithm steps with dose and schedule, contraindications and drug–drug/organ constraints, and surveillance intervals—each linked to human-readable provenance (source organization, version/year, section/page/table anchors) and governed by immutable version hashes, timestamps, and diffs. Native certainty and consensus schemes (e.g., NCCN Category, ESMO level/grade, ASCO strength/certainty, and corresponding Chinese systems) are preserved verbatim and consistently mapped to an internal ordinal scaffold for machine reasoning without loss of interpretability. Routine quarterly sweeps incorporate scheduled updates, while hot-fix ingestion captures safety alerts or major regulatory approvals.
To enable consistent computation across diseases at the index-lesion level, guideline texts (PDF/HTML) are segmented by a deterministic parser into atomic units and normalized to canonical ontologies and data models (ICD-10/11 for diagnoses; contemporary solid-tumor staging frameworks such as TNM-8; standardized drug vocabularies such as RxNorm/ATC; toxicity and response anchors via CTCAE v5.0 and RECIST/iRECIST). All fields are cast into machine-readable schemas covering indication/context, line of therapy, biomarker thresholds, organ-function gates, and explicit exceptions, while retaining the original wording for audit. The KB contains no identifiable patient information; it represents structured guidance, constraints, and follow-up templates that are later bound to case inputs elsewhere in the pipeline.
Evidence retrieval combines a hybrid retriever (BM25 + domain embeddings) with a lightweight medical knowledge graph capturing high-value relations (e.g., disease–stage–biomarker–therapy, and drug–drug/organ constraints). Queries return ranked, deduplicated snippets accompanied by click-through provenance triplets (source ID, anchor, version) to ensure traceable reproducibility. When sources diverge, conflicts are arbitrated by an explicit policy—safety first, then higher-certainty/higher-strength guidance, jurisdiction-matched documents preferred, and recency as a final tie-breaker—with the KB logging rationale and considered alternatives for governance and auditability. Supplementary Tables 1 and 2 lists the international and China local guideline catalog that populated the KB; entries are used strictly for knowledge extraction and traceable retrieval, not for redistribution.
Multi-agent EvoMDT
EvoMDT is a five-agent, coordinator-mediated framework that mirrors MDT roles and synthesizes auditable, lesion-level recommendations. The architecture orchestrates five specialized agents—each embodying distinct clinical subspecialty expertise—that engage in structured collaborative decision-making, synthesizing diagnostic assessments, therapeutic strategies, safety evaluations, and surveillance protocols into cohesive clinical recommendations. Unlike conventional single-model approaches, EvoMDT implements distributed reasoning with consensus mechanisms and autonomous prompt optimization, enabling adaptive performance refinement without human intervention. The overall architecture and data flow are shown in Fig. 6, spanning a multi-modal, ontology-normalized knowledge base; a five-agent collaborative reasoning framework; structured MDT outputs with evidence provenance; and multi-level performance validation.
a Multi-modal clinical knowledge base integrating guidelines, medical literature, and clinical cases with ontology normalization (SNOMED CT, ICD-10/11, RxNorm) and retrieval. b Five-agent collaborative reasoning: Diagnostic (Bayesian inference; TNM/BCLC/Lugano staging), Treatment (multi-criteria decision analysis; sequencing; pharmacogenomics), Safety (risk scoring; DDI/organ constraints; contraindication screening), Monitoring (three-phase protocols; RECIST/iRECIST anchors), and Coordinator (weighted consensus; conflict resolution). c Integrated MDT recommendation with primary diagnosis/staging, treatment plan, safety profile, monitoring protocol, and evidence citations plus confidence. d Multi-level performance validation: benchmarks, real-patient lesion-level validation, and single-blind expert ratings. DDI drug–drug interaction, MDT multidisciplinary team.
The EvoMDT architecture implements a hierarchical multi-agent system with five specialized reasoning modules, each instantiated from foundation large language models but configured with domain-specific prompts and knowledge retrieval mechanisms. EvoMDT’s core modules are open-sourced on GitHub to facilitate collaborative research and benchmarking (https://github.com/KesselZ/EvoMDT).
Diagnostic Agent: executes comprehensive tumor characterization through Bayesian diagnostic inference, computing posterior probabilities P(D|S) = P(S|D)·P(D)/P(S) for candidate diagnoses D given clinical presentation S, incorporating TNM/BCLC staging algorithms, molecular subtyping logic, and prognostic score calculations.
Treatment Agent: formulates personalized therapeutic recommendations via multi-criteria decision analysis, optimizing the utility function U(T) = Σwᵢ·vᵢ(T) across efficacy, safety, cost-effectiveness, and patient-specific factors, while integrating pharmacogenomic variants (CYP450, HLA alleles) and treatment sequencing through Markov decision processes.
Safety Agent: implements computational pharmacovigilance, calculating composite risk scores R = ΣαiRi + ΣβjIj incorporating individual risk factors Riand interaction terms Ij, querying drug interaction databases (DrugBank, Lexicomp, FAERS), and applying signal detection algorithms (proportional reporting ratios, Bayesian confidence propagation).
Monitoring Agent: generates phase-adapted surveillance protocols, optimizing monitoring utility M(t) = Σ pᵢ(t)·sᵢ·cᵢ⁻¹ balancing detection probability pᵢ(t), clinical significance sᵢ, and monitoring cost cᵢ across acute (0–3 months), intermediate (3–12 months), and long-term (>12 months) phases.
Coordinator Agent: synthesizes multi-specialist inputs through weighted consensus D_final = Σwᵢ·Dᵢ·Cᵢ where Dᵢ represents specialist recommendations, wᵢ denotes domain-relevance weights, and Cᵢ indicates agent confidence scores.
Inter-agent collaboration employs a Byzantine fault-tolerant consensus protocol, ensuring robust decision synthesis even when up to f agents (f < n/3, where n = 5) provide erroneous or conflicting outputs. Communication utilizes structured JSON message passing with semantic grounding in shared clinical ontologies (SNOMED CT, RxNorm), enabling precise cross-agent information exchange. In addition to role separation, EvoMDT is designed to expose a concrete, auditable reasoning trace rather than a single opaque narrative. Concretely, each agent emits a structured JSON output that contains: (i) a role-specific intermediate conclusion (e.g., diagnostic characterization/staging summary; candidate treatment plan; explicit safety constraints and contraindication checks; phase-adapted monitoring plan); (ii) a confidence estimate used in the weighted consensus; and (iii) an evidence list returned by retrieval, where each cited item is attached to provenance triplets (source ID, anchor, version) from the knowledge base. The Coordinator then compiles these intermediate outputs into the final MDT-style recommendation while logging (a) which options were accepted or rejected, (b) which conflicts were detected (semantic disagreement vs risk/implementability), and (c) the safety-first rationale used for tie-breaking when trade-offs arise. This trace structure makes EvoMDT’s collaboration process inspectable at the level of agent responsibilities and evidence provenance, enabling clinicians to audit not only “what the system recommends,” but also “which role concluded what, grounded on which sources, and why the final plan was selected.” The consensus algorithm implements dynamic weight adjustment based on case-specific factors: diagnostic complexity modulates Diagnostic Agent influence, treatment-refractory disease elevates Treatment Agent weighting, high-risk scenarios amplify Safety Agent priority, and surveillance-intensive cases enhance Monitoring Agent contribution. Conflict detection quantifies disagreement severity S_conflict = α·D_semantic + β·R_risk + γ·I_implementation triggering hierarchical resolution that prioritizes safety over efficacy when trade-offs emerge.
EvoMDT works through specialized agents (diagnostic, treatment, safety, monitoring, and coordination) collaborating to simulate real multidisciplinary team discussions. Each agent focuses on a specific task, complementing each other and coordinating decisions. When disagreements arise between agents, the coordinator agent plays a key role. The coordinator adjusts the influence of each agent dynamically, based on their recommendations and confidence levels, and resolves conflicts through a weighted consensus mechanism. For example, in the selection of a treatment plan, if the diagnostic agent recommends surgery, but the safety agent believes the patient has a high surgical risk, the coordinator will consider both agents’ recommendations and make a final decision based on risk assessments and the patient’s clinical situation.
For instance, in the case of a hepatocellular carcinoma patient, the diagnostic agent assesses that the tumor is suitable for surgical resection. However, the safety agent evaluates that the patient has severe liver dysfunction, making the surgical risk high, and therefore recommends alternative treatment (such as TACE). In this case, the coordinator will integrate the opinions of both the diagnostic and safety agents, considering the patient’s overall condition (such as liver function score, tumor biological characteristics, etc.), and balance safety and treatment effectiveness to provide the final treatment recommendation.
Within the EvoMDT framework, the foundation large language model (LLM) is designed to be interchangeable, allowing flexible replacement depending on task requirements and computational resources. During the development phase, we benchmarked several mainstream models. Ultimately, we selected DeepSeek V3 and DeepSeek R1 as the primary foundation models, based on a balance of efficiency and cost-effectiveness. Both models have demonstrated strong performance on authoritative medical benchmarks such as MedQA, confirming their medical reasoning capabilities and domain knowledge coverage. Furthermore, their inference speed and affordability substantially outperform comparable LLMs, making them a scalable and economically viable choice for supporting lesion-level clinical decision-making in EvoMDT.
In medical large language models, retrieval-augmented generation (RAG) is typically implemented through three major approaches: dense retrieval, sparse retrieval, and graph-enhanced retrieval. Dense retrieval leverages semantic embeddings to capture nuanced relationships, but it may underperform when handling rare medical terminology or low-resource languages. Sparse retrieval relies on keyword matching, which is highly effective for precise entity capture such as drug names or staging codes, though its semantic generalization is limited. Graph-enhanced retrieval employs knowledge graphs to represent structured relationships among medical concepts, but this method is complex to construct and maintain, and it depends heavily on graph quality and update frequency. In EvoMDT, we adopted a hybrid retrieval strategy that assigns a dominant weight of 0.8 to semantic retrieval while retaining a complementary weight of 0.2 for keyword-based retrieval. This design ensures robustness in multilingual contexts by combining semantic depth with entity-level precision, thereby enhancing both accuracy and interpretability in clinical decision-making.
We designed specialized prompts for each agent encoding domain-specific clinical reasoning frameworks, evidence-based medicine principles, and structured output requirements. Each prompt incorporates explicit instructions for algorithmic reasoning (Bayesian inference for diagnosis, multi-criteria decision analysis for treatment, risk stratification for safety), evidence integration requirements (citation of guidelines and literature with evidence grading), and quality assurance mechanisms (confidence calibration, uncertainty quantification).
The Diagnostic Agent prompt instructs systematic differential diagnosis generation using dual-process reasoning, TNM/BCLC staging with molecular integration, and investigation planning optimized by diagnostic yield. The Treatment Agent prompt encodes precision medicine principles including pharmacogenomic optimization, treatment sequencing via dynamic programming, and ensemble outcome prediction. The Safety Agent prompt emphasizes conservative risk assessment with comprehensive pharmacovigilance database integration and hierarchical contraindication classification. The Monitoring Agent prompt specifies multi-phase surveillance strategies with Bayesian adaptive protocols and machine learning-based early warning systems. The Coordinator Agent prompt directs weighted consensus computation, multi-layered conflict resolution prioritizing safety, and metacognitive quality assessment. Prompt optimization was performed through iterative refinement based on clinical expert feedback and quantitative performance metrics on development cases. Each prompt underwent validation for clinical appropriateness, completeness of reasoning coverage, and clarity of output specifications by board-certified oncologists with expertise in each respective domain (Details are as shown in Table 2).
The “self-evolution” mechanism in EvoMDT optimizes system efficiency through continuous feedback and automatic adjustments. After each task, the system evaluates its performance, identifying strengths and weaknesses. This process resembles a team manager using regular trial-and-error and review to improve workflows. Specifically, the system optimizes in two areas: First, it refines the instructions given to agents based on task feedback. When an agent encounters issues, the system adjusts its operating manual (i.e., prompts) to make the instructions clearer or more specific. For instance, it may instruct the agent to list a plan before executing or check the details before completing a task. This ensures more precise execution and improves decision quality. Second, the system optimizes the overall workflow by adjusting the team structure. If a single agent cannot complete a task, the system automatically adds another agent for collaboration. Alternatively, it may rearrange the task sequence or switch from a serial to a parallel workflow, similar to reorganizing a team to enhance efficiency. Through this iterative cycle, the system continuously fine-tunes both instructions and structure after each task, gradually evolving from a simple, rough initial design into a well-organized, efficient system tailored to solve specific complex problems.
Cross-domain evaluation on oncology-oriented medical QA benchmarks
To systematically assess EvoMDT’s cross-domain generalization in oncology-related reasoning, we established a two-tier evaluation framework encompassing both multiple-choice and generative medical QA tasks. All datasets were selected from publicly available, high-quality medical QA benchmarks and manually filtered to ensure thematic relevance to oncology or cancer.
The first category, multiple-choice medical QA, includes MedQA31, MedQA-Cancer, and MedMCQA-Cancer. MedQA serves as a general medical examination dataset used to evaluate clinical knowledge and diagnostic reasoning. MedQA-Cancer is a curated subset extracted from MedQA containing only cancer-related questions, while MedMCQA-Cancer consists of oncology questions drawn from the Indian medical entrance dataset MedMCQA32. These datasets present structured multiple-choice questions in which the model must identify the most clinically appropriate answer among several options. Given the unambiguous, fact-based nature of these tasks, Accuracy was used as the primary evaluation metric to quantify correctness in factual recall and reasoning.
The second category, generative medical QA, comprises MedQuAD33, MedQARo34, and ROND35. MedQuAD contains approximately 47,000 question–answer pairs sourced from NIH-affiliated websites such as cancer.gov, covering a range of general medical and cancer-specific topics. MedQARo is a Romanian-language QA benchmark centered on cancer patient case summaries, including over 100,000 QA pairs with both extractive and reasoning-based question types. ROND (Radiation Oncology NLP Database) focuses on the radiation oncology domain, supporting multiple NLP tasks including QA, summarization, and entity recognition, enabling evaluation of model performance in radiotherapy-specific contexts.
Since these datasets involve open-ended answer generation where correctness cannot be evaluated by discrete labels, we employed BERTScore36 as the principal evaluation metric. To enhance multilingual semantic alignment, we implemented BERTScore using BGE (BAAI General Embedding)37 and Qwen3-Embedding38—two of the most recent multilingual embedding models demonstrating state-of-the-art cross-lingual semantic similarity. This hybrid embedding setup ensures stable cross-language comparison across English, Chinese, and mixed biomedical corpora, improving the robustness of semantic evaluation. BERTScore measures the semantic similarity between model-generated responses and expert references, providing a more faithful estimate of clinical reasoning alignment than purely lexical metrics such as ROUGE. We used BERTScore with multilingual encoders (BGE-m3; Qwen3-Embedding) at default pooling; F1 aggregation was reported with resampling over 5 random seeds for stability. This dual evaluation framework enables a comprehensive assessment of EvoMDT’s performance across both structured examination-style tasks and free-form generation tasks, thereby validating its capability to balance factual precision with semantic reasoning in oncology decision-making contexts.
In this study, MedQA and MedQA-Cancer were primarily used as standardized benchmarks to quantify the model’s performance in medical knowledge acquisition and clinical reasoning abilities. Despite the structured nature of these datasets, the excellent results achieved on them demonstrate the model’s deep understanding of medical knowledge. In real-world clinical settings, when faced with unstructured, complex, and dynamic inputs, EvoMDT utilizes knowledge-enhanced reasoning and multi-agent collaborative mechanisms to complete the decision-making process, effectively addressing the diversity and complexity of real-world clinical cases.
Clinical validation on real-patient cases using EvoMDT
To assess the transferability of EvoMDT to lesion-level clinical decision-making, we conducted a multi-center retrospective study using real-world patient data collected from four tertiary hospitals: Peking University Third Hospital (Ethics Approval No. 2025472-01); Shanxi Cancer Hospital (Ethics Approval No. KY2025066); Henan Cancer Hospital (Ethics Approval No. 2024-472-001), and the First Affiliated Hospital of Fujian Medical University (Ethics Approval No. [2015]084-3). All participating centers obtained institutional ethics approval for secondary data analysis of de-identified medical records, and the study was conducted in full accordance with the principles of the Declaration of Helsinki. Because the investigation involved retrospective review of existing data without direct patient contact or intervention, the requirement for informed consent was formally waived by all institutional review boards. The study cohort comprised patients with four representative malignancies—lung adenocarcinoma (n = 100), breast cancer (n = 100), hepatocellular carcinoma (HCC; n = 105), and malignant lymphoma (n = 100)—yielding a total of 405 cases diagnosed and treated between October 2023 and March 2025. Each record was reviewed and standardized by experienced oncologists to ensure internal consistency and completeness across demographic, pathological, and treatment variables. We treated standardization as a clinician-led consistency check rather than an algorithmic “equalization” of centers. Specifically, case variables were recorded under a unified schema (demographics, core pathology context, stage context, lesion descriptors, and initial treatment plan) and reviewed for internal contradictions across fields. When a variable was missing, under-specified, or documented inconsistently in the source record, we retained it as “not available” instead of imputing or inferring values, thereby avoiding artificial uniformity across sites. This approach preserves real-world differences in case mix and documentation style while ensuring that the evaluation is not driven by inadvertent formatting artifacts or hidden assumptions. The statistical unit of analysis was defined as the index lesion at first diagnosis; for multifocal, bilateral, or recurrent disease, only the primary index lesion was included in the main analysis, while accompanying lesions were retained as metadata. For each cancer type, we compiled and statistically summarized real patient data to produce a structured descriptive overview, including age–stage distribution, gender composition, stage-stratified lesion diameters, risk and clinical context combinations, overall stage composition, and initial-treatment patterns. The corresponding six-panel visual summaries for all four malignancies are presented in the Supplementary Figs. 1–4 (Panels A–F for each cancer).
We designed a validation strategy based on four representative real-world cancer datasets—breast cancer, hepatocellular carcinoma, lung adenocarcinoma, and lymphoma. We deliberately chose lung adenocarcinoma, breast cancer, hepatocellular carcinoma (HCC), and lymphoma rather than a single tumor type. These diseases are high-burden, guideline-driven indications that account for a substantial share of MDT activity in tertiary centers, and they jointly span both solid and hematologic oncology. Lung adenocarcinoma and breast cancer are prototypical common solid tumors at the core of thoracic and breast tumor boards; HCC generates complex lesion-level decisions in liver tumor boards because transplant eligibility, local therapies, and systemic options must be balanced under liver function constraints; lymphoma represents the regimen- and line-of-therapy-centric decision patterns of hematologic oncology. As a result, the four-disease cohort stresses the ability of a single agentic framework to support lesion-level, MDT-style decision-making across several contrasting but highly representative oncology archetypes. Each dataset contained clinically validated patient records with expert treatment plans, which served as the ground truth for evaluation. Model outputs were compared against these references using three complementary metrics: BERTScore, ROUGE, and an AI-based ensemble score. BERTScore measured semantic similarity through contextual embeddings, capturing how well model-generated recommendations preserved expert intent beyond surface-level wording. ROUGE-1 recall quantified lexical overlap to ensure faithful reproduction of key clinical terminology and decision cues. The AI-based ensemble scoring integrated multiple independent evaluators with different reasoning heuristics, and the averaged results provided a robust estimate of alignment with expert reasoning, mitigating single-model bias. To contextualize performance, two baselines were introduced. The first was a rule-based baseline, producing generic and clinically irrelevant outputs such as “take medicine,” serving as a lower-bound reference for trivial errors. The second was a shuffled baseline, in which expert treatment plans were randomly reassigned within each cancer type, preserving medical language but removing case-specific reasoning. These baselines respectively tested robustness against nonsensical responses and sensitivity to individualized clinical variation. This multi-layered validation framework—integrating real-world datasets, diverse evaluation metrics, and carefully designed baselines—enabled a comprehensive assessment of EvoMDT’s ability to generate treatment recommendations that are semantically consistent with expert reasoning and lexically faithful to clinical practice.
Single-blind physician evaluation of clinical usefulness
To evaluate clinical usefulness under standardized, privacy-safe conditions, we applied a uniform operational and statistical workflow. Five board-certified oncologists were recruited, spanning experience levels of 3, 7, 12, 18, and 25 years (resident, attending, associate chief, chief, and discipline lead), with coverage across lung, breast, liver, and lymphoma subspecialties. Before formal rating, all physicians completed a 60-min rubric training and five pilot cases for anchoring; pilots were used solely for alignment and interface familiarization (target ICC > 0.70).
A single-blind design was used: raters were unaware of method provenance. Each case presented two MDT plans (EvoMDT versus a human comparator) in a paired fashion. A computer-generated stratified block randomization by cancer type and case complexity; both case order and A/B order were randomized, with block size concealed from raters. Ratings were performed independently without discussion; major protocol deviations (e.g., premature unblinding or incomplete records) were excluded prior to analysis. Using a standardized case dossier (demographics; lesion-level staging/biomarkers; organ function; comorbidities), raters scored six predefined dimensions on a 1–5 Likert scale: Safety & Ethics, Efficiency, Completeness, Clinical Appropriateness, Evidence Utilization, and Patient-Centeredness. Data capture used an electronic data-capture (EDC) system with range/logic checks (values constrained to 1–5; missing values disallowed) and an audit trail. We define response time as wall-clock time from when the standardized case dossier is made available to the evaluator/system to when a complete MDT-style plan is submitted. The dossier format used in this study is intentionally structured and bounded to reflect typical MDT discussion packets. Accordingly, the reported timing results quantify decision-generation latency conditioned on this dossier, and do not include additional upstream preprocessing that would be required for substantially richer raw clinical records (e.g., lengthy unstructured notes or dense longitudinal data). Response times <0.5 min triggered review and correction. Single-dimension missingness was handled by listwise deletion at the case–dimension level; whole-case missingness prompted re-rating. Outliers were screened using Tukey fences (±3 IQR) and retained if clinically plausible after blinded adjudication. Post-lock missingness was controlled at <2%. Table 3 shows the evaluation rules for various representative tumor types.
Statistical analysis
Analyses followed a three-layer validation framework aligned with the study objectives. Layer 1 (pan-cancer benchmarks) assessed component-level performance across established metrics to compare alternative LLM settings and retrieval configurations. Layer 2 (lesion-level validation on real-world data) evaluated clinical decision quality in four representative cancers using hospital-derived case dossiers, focusing on (i) guideline concordance, (ii) weighted plan concordance (WPC) with reference MDT plans, (iii) evidence traceability, and (iv) safety-violation rate. Layer 3 (single-blind physician rating) compared EvoMDT with human experts across six dimensions—Safety & Ethics, Efficiency, Completeness, Clinical Appropriateness, Evidence Utilization, and Patient-Centeredness—using a 1–5 Likert scale and recorded response time.
Unless otherwise specified, statistical tests were two-sided with α = 0.05. For multiple comparisons, both raw P values and Benjamini–Hochberg FDR-adjusted q values were reported (q < 0.05 considered significant). Given the non-normal distribution of ordinal scores, Mann–Whitney U and Kruskal–Wallis tests were applied, followed by FDR-adjusted pairwise contrasts. Categorical outcomes (e.g., ≥4-score proportion, guideline concordance) were analyzed using χ² or Fisher’s exact tests as appropriate. Effect sizes were reported as rank-biserial correlations (r) for Mann–Whitney tests and derived analogously for post-hoc comparisons. Inter-dimension correlations used Spearman’s ρ, and inter-rater reliability was quantified via ICC(2,1) (two-way random effects, absolute agreement). Data integrity was ensured through automated range and logic checks; missing values were not permitted, and any incomplete cases were re-rated. Outliers were screened using Tukey fences (±3 IQR) and retained only if deemed clinically plausible after blinded adjudication. Sensitivity analyses included site-fixed effects and covariate adjustment (age, stage) to assess robustness to inter-center heterogeneity.
Data availability
The data supporting this study’s findings are not publicly available due to privacy and confidentiality restrictions. However, the datasets can be obtained from the corresponding author upon reasonable request, subject to approval and compliance with relevant data protection regulations.
Code availability
Core modules and prompts of EvoMDT are available at GitHub: https://github.com/KesselZ/EvoMDT.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Topol, E. J. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 25, 44–56 (2019).
Patkar, V. et al. Cancer multidisciplinary team meetings: evidence, challenges, and the role of clinical decision support technology. Int. J. Breast Cancer 2011, 831605 (2011).
Horlait, M., Baes, S., Dhaene, S., Van Belle, S. & Leys, M. How multidisciplinary are multidisciplinary team meetings in cancer care? An observational study in oncology departments in Flanders, Belgium. J. Multidiscip. Healthc. 12, 159–167 (2019).
Koo, Y., Shafiq, J., Yanga, J., Avery, S. & Vinod, S. Quality of decision-making at oncology multidisciplinary team meetings: a structured observational study. Clin. Oncol. 47, 103942 (2025).
Williams, G. J. & Thompson, J. F. Management changes and survival outcomes for cancer patients after multidisciplinary team discussion; a systematic review and meta-analysis. Cancer Treat. Rev. 139, 102997 (2025).
Mangone, L. et al. Impact of multidisciplinary team management on survival and recurrence in stage I–III colorectal cancer: a population-based study in Northern Italy. Biology 13, 928 (2024).
Gouliaev, A. et al. Discrepancies in regional lung cancer multidisciplinary team decisions can be reduced through national consensus meetings. Acta Oncol. 64, 43314 (2025).
Reig, M. et al. BCLC strategy for prognosis prediction and treatment recommendation Barcelona Clinic Liver Cancer (BCLC) staging system. The 2022 update. J. Hepatol. 76, 681–693 (2021).
Sutton, R. T. et al. An overview of clinical decision support systems: benefits, risks, and strategies for success. NPJ Digit. Med. 3, 17 (2020).
Shortliffe, E. H. & Sepúlveda, M. J. Clinical decision support in the era of artificial intelligence. JAMA 320, 2199–2200 (2018).
Kung, T. H. et al. Performance of ChatGPT on USMLE: potential for AI-assisted medical education using large language models. PLoS Digit. Health 2, e0000198 (2023).
Ayers, J. W. et al. Comparing physician and artificial intelligence chatbot responses to patient questions posted to a public social media forum. JAMA Intern. Med. 183, 589–596 (2023).
Bedi, S. et al. Testing and evaluation of health care applications of large language models: a systematic review. JAMA 333, 319–328 (2025).
Lin, C. & Kuo, C. F. Roles and potential of large language models in healthcare: a comprehensive review. Biomed. J. 48, 100868 (2025).
Li, G., Hammoud, H., Itani, H., Khizbullin, D. & Ghanem, B. Camel: communicative agents for “mind” exploration of large language model society. Adv. Neural Inf. Process. Syst. 36, 51991–52008 (2023).
Amugongo, L. M., Mascheroni, P., Brooks, S., Doering, S. & Seidel, J. Retrieval augmented generation for large language models in healthcare: a systematic review. PLoS Digit. Health 4, e0000877 (2025).
Woo, J. J. et al. Custom large language models improve accuracy: comparing retrieval augmented generation and artificial intelligence agents to noncustom models for evidence-based medicine. Arthroscopy 41, 565–573 (2025).
Asif, S. et al. Advancements and prospects of machine learning in medical diagnostics: unveiling the future of diagnostic precision. Arch. Comput. Methods Eng. 32, 853–883 (2025).
Liu, X. et al. Reporting guidelines for clinical trial reports for interventions involving artificial intelligence: the CONSORT-AI extension. Lancet Digit. Health 2, e537–e548 (2020).
Vasey, B. et al. Reporting guideline for the early stage clinical evaluation of decision support systems driven by artificial intelligence: DECIDE-AI. BMJ 377, e070904 (2022).
Walonoski, J. et al. Synthea: an approach, method, and software mechanism for generating synthetic patients and the synthetic electronic health care record. J. Am. Med. Inform. Assoc. 25, 230–238 (2018).
Hussain, A., Ali, S., Farwa, U. E., Mozumder, M. A. I. & Kim, H. C. Foundation models: from current developments, challenges, and risks to future opportunities. In 2025 27th International Conference on Advanced Communications Technology (ICACT) 51–58 (IEEE, 2025).
Sethi, G. K., Sharma, S. & Kumar, R. Explainable AI in Medical Imaging, Personalized Medicine, and Bias Reduction (2025).
Salimparsa, M. et al. Explainable AI for clinical decision support systems: literature review, key gaps, and research synthesis. Informatics 12, 119 (2025).
Wang, L. et al. Artificial intelligence in clinical decision support systems for oncology. Int. J. Med. Sci. 20, 79 (2023).
Oehring, R. et al. Use and accuracy of decision support systems using artificial intelligence for tumor diseases: a systematic review and meta-analysis. Front. Oncol. 13, 1224347 (2023).
Farooq, M. U. B. et al. MDT-based intelligent route selection for 5G-enabled connected ambulances. In 2022 IEEE International Conference on E-health Networking, Application & Services (HealthCom) 81–87 (IEEE, 2022).
Brown, G. T., Bekker, H. L. & Young, A. L. Quality and efficacy of multidisciplinary team (MDT) quality assessment tools and discussion checklists: a systematic review. BMC Cancer 22, 286 (2022).
Jing, X. M. et al. Effect of multidisciplinary treatment (MDT) on survival outcomes of lung cancer patients: experiences from China. Asia Pac. J. Clin. Oncol. 20, 634–642 (2024).
Jin, D. et al. What disease does this patient have? A large-scale open domain question answering dataset from medical exams. Appl. Sci. 11, 6421 (2021).
Pal, A., Umapathi, L. K. & Sankarasubbu, M. Medmcqa: a large-scale multi-subject multi-choice dataset for medical domain question answering. In Conference on Health, Inference, and Learning 248–260 (PMLR, 2022).
Ben Abacha, A. & Demner-Fushman, D. A question-entailment approach to question answering. BMC Bioinforma. 20, 511 (2019).
Rogoz, A. C. et al. MedQARo: A Large-Scale Benchmark for Medical Question Answering in Romanian. arXiv preprint https://doi.org/10.48550/arXiv.2508.16390 (2025).
Liu, Z. et al. The radiation oncology NLP database. arXiv preprint https://doi.org/10.48550/arXiv.2401.10995 (2024).
Zhang, T., Kishore, V., Wu, F., Weinberger, K. Q. & Artzi, Y. BERTScore: Evaluating text generation with BERT. In International Conference on Learning Representations (2020).
Chen, J. et al. M3-Embedding: Multi-Linguality, Multi-Functionality, Multi-Granularity Text Embeddings Through Self-Knowledge Distillation. In Findings of the Association for Computational Linguistics: ACL, 2318–2335 (2024).
Zhang, Y. et al. Qwen3 Embedding: Advancing Text Embedding and Reranking Through Foundation Models. arXiv preprint arXiv:2506.05176 (2025).
Acknowledgements
We acknowledge the guidance provided by the participating institutions during the implementation of this study. This work was supported by National Natural Science Foundation of Fujian (2023Y9052, 2024Y9158); National Key Clinical Specialty Construction Project Fund (No.2022YBL-ZD-02); the project of the China Association for Promotion of Health Science and Technology (Grant No. JKHY2023003), the Capital Health Development Scientific Research Special Project of Beijing Municipal Health Commission (Grant No. 2024-1-1023), and the Clinical Key Project of Peking University Third Hospital (Grant No. BYSYRCYJ2023001).
Author information
Authors and Affiliations
Contributions
Q.L., Z.H., and T.H. conceived of the presented idea. T.H. and Y.N. designed the computational framework and analyzed the data. Q.L., Y.N., X.Z., and S.M. conducted the clinical evaluation. Z.H. and C.L. supervised the findings of this work. F.G., G.K.H., and H.E.K. provided valuable intellectual input during the software refinement process. Q.L. and Z.H. took the lead in writing the manuscript and supplementary information. X.S., Z.Y., and G.Q. provided funding support for this research. All authors discussed the results and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Q., Hu, Z., Huang, T. et al. EvoMDT: a self-evolving multi-agent system for structured clinical decision-making in multi-cancer. npj Digit. Med. 9, 124 (2026). https://doi.org/10.1038/s41746-025-02304-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02304-8