Abstract
Breath analysis offers a promising noninvasive strategy for early cancer detection by capturing disease-specific volatile organic compound (VOC) signatures in exhaled breath. In this study, we developed a hierarchical deep convolutional neural network (HD-CNN)-based platform for dual-cancer classification using a multimodal gas sensor array. Clinical breath samples were collected from 206 participants, including 67 healthy controls (HC), 78 lung cancer (LC), and 61 gastric cancer (GC) patients. The sensor array, composed of semiconductor metal oxide (SMO), electrochemical (EC), and photoionization detector (PID) sensors, generated time-resolved signals that were converted into 2D response maps for classification. The HD-CNN model employed a two-stage structure: a coarse classifier to distinguish HC from cancer patients (CP), followed by a fine classifier to separate LC and GC. In 5-fold cross-validation, the HD-CNN achieved classification accuracies of 82.1% (HC), 84.0% (LC), and 88.1% (GC), and average AUCs of 0.89, 0.92, and 0.89, respectively. Compared to a 1D CNN, the HD-CNN demonstrated superior class separability, increased prediction confidence, and overall enhanced performance. Additionally, we evaluated multiple coarse–fine configurations and found that isolating HC in the first stage resulted in the highest overall performance. These results support the utility of hierarchical learning and multimodal sensing for robust and scalable breath-based multi-cancer screening.
Similar content being viewed by others
Introduction
Cancer is one of the leading global health threats of the twenty-first century, accounting for over 16% of all deaths and nearly one-third of premature deaths from noncommunicable diseases in adults aged 30–69. It ranks among the top three causes of mortality in the majority of countries worldwide, posing a substantial burden on both public health and healthcare systems. Lung cancer (LC) and gastric cancer (GC) are among the most lethal malignancies worldwide, ranking first and fifth in cancer-related deaths, respectively1,2,3. According to statistics, LC accounted for approximately 18.7% of all cancer deaths, and GC caused an estimated 660,000 deaths in 2022. Both cancers continue to pose a significant global health burden due to frequent late-stage diagnoses and poor 5-year survival rates4,5. It highlights the need for noninvasive early detection technologies to improve clinical outcomes and reduce mortality.
Breath analysis has recently emerged as a promising alternative to conventional cancer diagnostics due to its noninvasive nature, simplicity, and potential for real-time application6,7,8,9,10,11,12,13. This technique captures volatile organic compounds (VOCs) present in exhaled breath, which are byproducts of metabolic processes and can serve as disease-specific biomarkers14,15,16,17,18,19,20,21,22. Because cancer alters cellular metabolism, the composition of exhaled VOCs can differ significantly between healthy individuals and cancer patients. These biochemical alterations provide a valuable diagnostic signature, enabling breath-based sensing platforms to detect pathological states without the need for invasive procedures.
Previous studies have demonstrated the feasibility of breath analysis for diagnosing specific cancers such as LC22,23,24, GC23,25,26, and breast cancer27,28. However, most of these investigations have been focused on single-cancer detection using binary classification models. In clinical practice, multiple cancer types may present with overlapping metabolic features or co-exist in high-risk populations29,30,31,32. Therefore, the development of a diagnostic platform capable of simultaneously identifying multiple cancer types from a single breath sample represents a critical step toward practical, scalable cancer screening. Such a system could not only reduce diagnostic burden but also improve accessibility and cost-efficiency across diverse clinical settings.
Accurate multi-cancer diagnosis based on breath analysis, however, presents several challenges. The biomarkers of different cancers may be partially overlapping or subtly distinct, and their representation in sensor responses can vary across individuals32. Traditional machine learning models often struggle to extract and generalize these complex patterns33,34,35. To overcome these limitations, more advanced deep learning techniques are required—models that go beyond traditional machine learning or shallow architectures in their ability to extract and generalize from complex, high-dimensional input data. Cancer-related breath profiles often contain subtle, nonlinear, and overlapping features that vary across individuals and disease stages. Therefore, a robust classification system must be capable of modeling such intricate patterns with high sensitivity and specificity. Recent advances in deep learning, including attention mechanisms, residual networks, ensemble learning, and hierarchical architectures, offer powerful tools to capture these complex relationships. By enabling deeper representation learning and more flexible decision boundaries, these models can significantly enhance multi-class discrimination performance and are particularly well-suited for tasks such as multi-cancer classification from breath-based VOC patterns.
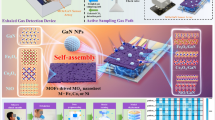
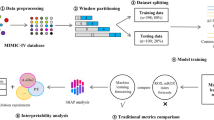
In this study, we developed a breath-based diagnostic platform for dual-cancer classification by integrating a multimodal gas sensor array with a hierarchical deep convolutional neural network (HD-CNN), as illustrated in Fig. 1. Exhaled breath samples from patients with LC, GC, and healthy controls (HCs) were analyzed using an electronic nose system equipped with a multimodal sensor array composed of semiconductor metal oxide (SMO), electrochemical (EC), and photoionization detector (PID) sensors. This heterogeneous sensor configuration enhances the system’s capability to capture complex and subtle biochemical signatures associated with specific diseases. The collected sensor responses were converted into 2D response maps through preprocessing, and subsequently classified using a two-stage HD-CNN framework. The model consists of a coarse classifier that first distinguishes HCs from cancer patients, followed by a fine classifier that differentiates between LC and GC. To validate the performance of the proposed HD-CNN, we compared it with a conventional single one-dimensional convolutional neural network (1D CNN) model. In addition, multiple coarse–fine configurations were evaluated to identify the optimal HD-CNN architecture achieving the highest classification accuracy.
a Exhaled breath from healthy controls (HCs), lung cancer (LC), and gastric cancer (GC) patients is analyzed using a multimodal gas sensor array to generate time-resolved breathprints. b The sensor signals are preprocessed into response maps and classified using a hierarchical deep convolutional neural network (HD-CNN) consisting of a coarse classifier and a fine classifier.
Results and discussion
Development of a breath analyzer with a multimodal sensor array
To enable robust and real-time detection of VOCs in exhaled breath, we developed an upgraded breath analysis system based on our previous prototype22. Figure 2a shows a 3D rendering of the developed system. While the overall chamber geometry and multimodal sensor layout were retained to maintain fluidic uniformity, several key modifications were implemented to improve measurement reliability and facilitate system scalability.
a 3D rendering of the complete breath analysis device. b Internal layout of the system, including gas chamber, sampling module, temperature controller, and embedded PC. c FPGA-based ADC board composed of a signal amplifier block, FPGA block, ADC block, and MCU block. d Main control board containing power management, communication I/O, and PLC interface blocks. e System-level block diagram that illustrates the configuration of 16-bit ADC channels and parallel data acquisition through an FPGA.
First, the thermal control system was significantly improved. In the previous version, prolonged operation led to degradation of the Tenax sorbent tube due to overheating and thermal instability. To address this, the heating unit was redesigned to incorporate closed-loop control with precise thermal feedback, effectively preventing overheating while maintaining consistent desorption efficiency. This modification ensures reproducible VOC release and extends the lifespan of the sorbent tube. Second, to reduce susceptibility to ambient temperature fluctuations, the gas sensor chamber—which was previously exposed to the external environment—was enclosed within a thermally insulated housing. This structural enhancement minimizes baseline drift caused by external thermal perturbations, thereby improving signal stability during extended measurements. Figure 2b illustrates the complete system architecture, including the thermal control design. Third, signal acquisition performance was enhanced through the integration of a custom Field-Programmable Gate Array (FPGA)-based analog-to-digital converter (ADC) board. The structure of the FPGA-based ADC PCB is illustrated in Fig. 2c. Unlike previous microcontroller-based systems limited by sequential processing and constrained sampling rates, the FPGA enables true parallel signal processing. It allows high-speed, independent data acquisition across multiple sensor channels. As a result, the system achieves higher signal fidelity and improved temporal resolution. In addition, it supports integration of heterogeneous sensor types and offers modular scalability for future expansion. A multi-channel power management system with noise isolation was implemented to supply stable and independent power to each sensing module and control unit. This design ensures consistent system performance while effectively minimizing electrical interference. Figure 2d presents the main control PCB, including digital signal processing logic, power converters for optimized multi-sensor operation, and additional computational resources for advanced signal analysis. The complete hardware block diagram is illustrated in Fig. 2e. It shows the parallel acquisition of sensor signals via 16-bit ADCs and real-time processing through an FPGA module. The digitized data are then transmitted to the main controller for classification and system control. The buffer amplifier, as shown in Fig. 2e, is used as a voltage follower after the front-end gain stage. It provides: (1) load isolation and drive—decouples the high-impedance network from the ADC sampling cap; (2) low output impedance—reliably drives multi-channel capture and anti-alias RC; (3) preserved stability/bandwidth—isolates variable loads.
Despite these modifications, the gas sensor chamber retains the same internal dimensions (30 × 10 × 4 cm) and edge-chamfered structure as in our previous study22, preserving the uniform gas distribution confirmed through prior flow simulations. The multimodal sensor array was reconfigured by incorporating previously validated sensors, along with several improvements to enhance performance. Supplementary Fig. 1 presents the output values for each sensor in the implemented system upon exposure to standard gases. The test gases (ethanol and NO2) were each evaluated at concentrations of 150, 300, 600, and 1200 ppb. Sensor responses varied depending on both gas concentration and type, demonstrating the potential to distinguish different gases and their levels through pattern analysis. These improvements over previous work result in a more stable and scalable hardware platform. The system acquires high-resolution breath profiles with high reproducibility and provides a solid foundation for accurate disease classification through deep learning models.
Analysis of demographic information and clinical breath samples
A total of 206 participants were included in this study, comprising 67 HC, 78 LC, and 61 GC. The sex and age distribution for each group is presented in Fig. 3a. Participants were well-balanced across age groups and included both male and female individuals. The cancer cohorts covered various clinical stages, with the majority diagnosed at early stages (I and II), as shown in Fig. 3b. To validate the feasibility of early detection using our platform, we collected breath samples from a larger number of patients diagnosed at early stages.
a Distribution of participants in the HC, LC, and GC groups by sex and age. A total of 206 participants were included: 67 HC, 78 LC, and 61 GC. b Cancer stage distribution of LC and GC patients. Most patients were diagnosed in early stages (I and II), with a smaller number in advanced stages (III and IV). c Representative sensor responses of the multimodal gas sensor array for each group. Transient responses are shown for semiconductor metal oxide (SMO), electrochemical (EC), and photoionization detector (PID) sensors.
Figure 3c shows that the multimodal sensor array generated distinct response patterns across HC, LC, and GC. The breath sample desorbed from the Tenax tube enters the sensor chamber around 900 s into the measurement, and most sensors show peak responses near 1100 s. Variations in response value and time across the multimodal sensor array were attributed to their differing sensitivity to specific VOCs present in exhaled breath. Each sensor detects a distinct subset of chemical compounds, resulting in unique response patterns depending on the breath composition of each subject. As a result, some sensors responded more strongly or more rapidly in certain patient groups. Specifically, the sensors showed differences in response, peak time, and recovery time. SMO#2 (blue), SMO#4 (cyan), and SMO#5 (green) showed larger response changes in LC and GC compared to HC. In addition, EC#2 (sky blue) and EC#9 (brown) showed stronger responses in cancer patients. The variability among sensors contributes to diverse response patterns, enabling more detailed characterization of exhaled breath. Such diversity enhances the diagnostic accuracy of the system, particularly in distinguishing between disease states like LC and GC.
Compared to the previous breath analyzer22, the overall sensor responses have increased in magnitude, and the signals show smoother transitions with reduced noise. Notably, the peak response time has significantly decreased, suggesting that the system can now capture more informative and dynamic changes in VOC profiles within a shorter measurement period.
Each sensor in the multimodal array is designed to respond to specific volatile compounds such as ethanol, isobutanol, formaldehyde, carbon monoxide, and hydrogen sulfide22. However, the exhaled breath of cancer patients generally contains a complex mixture of numerous VOCs originating from diverse metabolic pathways. As a result, the variations observed across sensors represent composite chemical interactions. Therefore, we focus on the overall “breathprint” pattern that reflects integrated metabolic signatures, rather than targeting individual VOCs.
Construction of an HD-CNN model of dual-cancer screening
To classify breath samples into HC, LC, and GC groups, we developed an HD-CNN model designed to process time-series gas sensor data. As illustrated in Fig. 4, the overall pipeline begins with the collection of clinical breath samples using a multimodal gas sensor array composed of 19 individual channels, including SMO, EC, and PID sensors. The sensor array generates a variety of time-series responses depending on the VOC profile of each breath sample. The raw signals obtained from the multimodal sensor array were first normalized and temporally cropped to a fixed window, producing 2D input response maps of size 19 × 1800 (sensor × time). The cropped data corresponded to the time segment from 900 to 2700 s, starting from the point when the breath sample entered the sensor chamber. Supplementary Fig. 2 shows examples of the input response maps for each group. Heatmaps visualize the normalized sensor responses over time for HC, LC, and GC. Each class exhibits distinct signal intensity patterns, which indicate temporal variations in VOC interactions among groups. In particular, a clear contrast in response characteristics is observed between the SMO sensor array and the PID + EC sensor array.
Clinical breath samples were collected using a multimodal gas sensor array. Raw sensor signals were normalized and cropped to form 2D input response maps (19 sensors × 1800 time points). Input data were split using a 5-fold cross-validation scheme for robust model training and validation. The HD-CNN consists of a coarse classifier that distinguishes HC from cancer (LC + GC), followed by a fine classifier that differentiates between LC and GC. A probabilistic averaging layer integrates the outputs from both classifiers to generate the final three-class prediction.
To ensure generalizable model performance and minimize overfitting, we employed 5-fold validation. In each fold, 80% of the data were used for training and 20% for validation, with participant-level stratification based on disease class (HC, LC, and GC) to maintain class balance across folds. Other demographic variables such as age, sex, and cancer stage were not explicitly used as stratification criteria due to the limited sample size, but their distributions were confirmed to be comparable across folds. The core of the classification architecture is an HD-CNN model composed of two independently trained classifiers: a coarse classifier and a fine classifier. First, the coarse classifier performs binary classification to determine whether a sample belongs to the HC class or cancer patient class (LC + GC). Samples predicted as cancer are subsequently passed to the fine classifier, which further discriminates between LC and GC. Both classifiers employ identical feature extraction layers consisting of two convolutional layers with batch normalization and LeakyReLU activation, followed by fully connected layers and dropout regularization (p = 0.65 for coarse and p = 0.5 for fine). The dropout values were empirically optimized based on model stability and validation performance. Through comparative experiments with multiple dropout combinations, the selected configuration provided the best balance between sensitivity and specificity. To produce the final classification result, a probabilistic averaging layer combines the output probabilities from both classifiers. Specifically, the probability of a sample belonging to the HC class is directly determined by the output of the coarse classifier. In contrast, the probabilities for LC and GC are calculated by multiplying the cancer probability from the coarse classifier by the class probabilities generated by the fine classifier. This probability integration method was adapted from a previously proposed hierarchical classification approach36.
To evaluate the effectiveness of the proposed HD-CNN architecture, we compared its performance with a conventional 1D CNN model using 5-fold validation. Figure 5 summarizes the classification results of both models across various performance metrics. The scatter plots in Fig. 5a, d show the predicted class probabilities generated by the 1D CNN and HD-CNN models, respectively. The predicted probability for each individual sample is illustrated in Supplementary Fig. 3. In the 1D CNN model, predicted probabilities were relatively dispersed across all classes, with many samples clustered near the decision boundary, indicating uncertainty in classification. In contrast, the HD-CNN model yielded probabilities close to 0 or 1 for most samples, resulting in clearly distinguishable class predictions with higher confidence. The clarity of class separation can be more precisely examined in the graphs shown in Supplementary Fig. 3.
a–c Classification results of the 1D CNN model on the 5-fold cross-validation dataset. a Scatter plots showing predicted class probabilities for HC, LC, and GC groups. b Confusion matrix for 3-class classification. c One-vs-rest ROC curves and AUC values with 95% confidence intervals for each class. d–f Corresponding results of the HD-CNN model on the 5-fold cross-validation dataset. d Scatter plots of predicted probabilities, e confusion matrix, and f one-vs-rest ROC curves and AUC values of the HD-CNN model. Radar plots comparing accuracy, precision, recall, and F1-score between two models for g training dataset, and h validation dataset.
Figure 5b, e presents the confusion matrices for both models. The 1D CNN achieved an accuracy of 71.6% for HC, 79.8% for LC, and 80.3% for GC. However, it misclassified 14.9% of LC samples as HC and 16.4% of GC samples as LC, indicating that distinguishing between abnormal classes remains challenging for a flat classifier. The HD-CNN significantly improved classification performance, with 82.1% accuracy for HC, 84.0% for LC, and 88.1% for GC. Misclassification rates between LC and GC were notably reduced. This result indicates that the hierarchical approach is effective in resolving inter-class ambiguity by simplifying the classification task into sequential decisions and improving class separability.
Receiver operating characteristic (ROC) curve analysis was also conducted to further assess the model’s classification performance. One-vs-rest analysis was performed for each class, and the average performance across the 5-fold validation, along with the corresponding 95% confidence intervals, was shown in Fig. 5c, f. The 1D CNN yielded AUC values of 0.75 for HC, 0.87 for LC, and 0.91 for GC. In contrast, the HD-CNN achieved higher AUCs across all classes: 0.89 for HC (95% CI: 0.84–0.94), 0.92 for LC (95% CI: 0.88–0.97), and 0.89 for GC (95% CI: 0.85–0.92).
Figure 5g, h shows radar plots comparing four key performance metrics, including accuracy, precision, recall, and F1-score. The plots demonstrate that the HD-CNN consistently outperformed the 1D CNN on both training and validation datasets. Especially in the validation dataset, the performance gap was particularly evident in recall and F1-score, reflecting the HD-CNN’s improved capability to correctly identify positive cases and maintain balanced classification across all three classes.
Compared to single-path classification models such as 1D CNNs, the HD-CNN architecture offers a more structured and flexible approach to handling complex multi-class problems. While 1D CNNs extract temporal features through end-to-end learning and stacked convolutional layers, they often face challenges in capturing inter-class differences. In particular, the changes in VOC composition in exhaled breath are often subtle, making the boundaries between classes less distinct and more difficult to separate. In contrast, HD-CNN decomposes the classification task into a two-stage hierarchy, where a coarse classifier first separates HCs from cancer patients, followed by a fine classifier that distinguishes between cancer subtypes. This hierarchical separation allows each classifier to focus on a simpler decision boundary, which improves robustness and interpretability. Furthermore, by isolating cancer subtype classification from healthy and abnormal discrimination, the model reduces confusion between similar classes and enhances sensitivity for minority classes.
For more comparative evaluation, we implemented additional baseline models: (i) a ResNet with residual blocks and global average pooling, and (ii) a Transformer encoder with multi-head self-attention and position encodings. All models used the same input dataset, normalization, class-weighting, optimizer, learning-rate scheduler, and 5-fold validation as the HD-CNN. Hyperparameters were tuned within the same search ranges as the HD-CNN. Supplementary Table 1 summarizes model parameters, classification accuracy, and F1-score for the training and validation datasets. HD-CNN achieved 84.7% accuracy and F1 = 0.85, whereas ResNet achieved 78.4% accuracy and F1 = 0.78, and the Transformer achieved 79.3% accuracy and F1 = 0.79. This result indicates that hierarchical coarse-to-fine inference provides additional discriminative power over non-hierarchical backbones despite comparable model sizes.
To investigate the possibility of sensor optimization and array reduction, we analyzed the relative contribution of each sensor using SHapley Additive exPlanations (SHAP) values derived from both CNN and HD-CNN models. As shown in Supplementary Fig. 5, the two models exhibited highly consistent patterns of sensor importance, indicating that both architectures relied on similar feature sources within the multimodal array. Based on this analysis, the top 10 sensors showing the highest mean |SHAP values| were selected for further evaluation. Using only these ten most informative sensors, we retrained the HD-CNN model to assess its classification capability under a reduced-array condition. As presented in Supplementary Fig. 6a, the confusion matrix demonstrates that the optimized model maintained strong discriminative performance across all three classes, achieving accuracies of 74.6% for HC, 81.9% for LC, and 83.6% for GC. The one-vs-rest ROC curves in Supplementary Fig. 6b show corresponding AUCs of 0.82 (95% CI: 0.76–0.88) for HC, 0.9 (95% CI: 0.89–0.94) for LC, and 0.88 (95% CI: 0.81–0.94).
Although the reduced-sensor model maintained recognizable class-specific patterns, its overall validation accuracy and ROC–AUC values were lower than those obtained using the full sensor array. The performance degradation was primarily attributed to the loss of complementary information among heterogeneous sensor modalities. These results suggest that while a smaller subset can extract key diagnostic features to a certain extent, comprehensive pattern representation and generalization capability are better preserved when the full multimodal sensor array is employed.
Performance comparison between coarse and fine classifiers
For further analysis of hierarchical structure on classification performance, we tested three different coarse–fine configurations in the HD-CNN model (Fig. 6). In all configurations, the hierarchical design allowed the model to split the original three-class problem into two sequential binary tasks. However, the order of class separation and the role of each classifier differed, which influenced both overall accuracy and the balance between class-specific performance. In the first structure (Fig. 6a), the coarse classifier was trained to separate HC from cancer patients (CP = LC + GC), followed by a fine classifier to distinguish between LC and GC. This configuration yielded the best fine classification performance, with 95.1% accuracy for LC and 98.1% for GC. The coarse classifier achieved 82.1% accuracy for HC and 85.9% for CP. The clear advantage of this structure lies in the ability of the coarse classifier to isolate healthy individuals. This separation allows the second-stage classifier to focus on the distinction between cancer subtypes, which reduces task complexity and improves classification accuracy. In particular, the fine classifier that distinguishes between LC and GC achieves remarkably high accuracy. This strong performance significantly enhances the stability and reliability of the model.
a Coarse classifier distinguishes HC from CP (LC + GC), and a fine classifier differentiates LC from GC. b Coarse classifier separates GC from non-GC (HC + LC), and fine classifier distinguishes HC from LC. c Coarse classifier separates LC from non-LC (HC + GC), and the fine classifier distinguishes HC from GC.
In the second structure (Fig. 6b), the coarse classifier was designed to distinguish GC from non-GC (HC + LC), and the fine classifier classified HC and LC. While the coarse classifier maintained good performance for GC (88.5%), the fine classifier struggled with distinguishing HC from LC, showing 77.2% and 84.4% accuracy, respectively. Separating GC at the coarse level improves its classification accuracy, but the model shows reduced performance in distinguishing HC from LC, likely due to overlapping breath profiles.
In the third structure (Fig. 6c), the coarse classifier was trained to distinguish LC from non-LC (HC + GC), followed by a fine classifier to differentiate between HC and GC. The coarse classification achieved 78.7% accuracy for LC and 85.9% for non-LC. In the fine stage, the model reached 88.2% accuracy for HC and 85.0% for GC. While this configuration maintained relatively balanced performance across both stages, it showed lower fine-stage accuracy compared to the first structure, particularly in separating HC from GC. This result suggests that the similarity between HC and GC breath profiles may hinder precise discrimination.
Overall, these results confirm that the design of the hierarchical structure has a critical influence on classification outcomes. The configuration where the coarse classifier first isolates HC, followed by fine classification of LC and GC (Fig. 6a), consistently produced the most accurate and interpretable results. Additionally, this approach aligns with the clinical goal of screening by first distinguishing healthy individuals and then identifying the specific type of cancer in those classified as positive. The flexibility of the HD-CNN framework allows for such task-specific adaptation, enhancing its potential utility in real-world multi-cancer screening applications.
Comparison with previous studies and related works
Previous studies have demonstrated the potential of breath analysis for cancer detection using various types of breath sensors and analytical frameworks, as summarized in Table 1. Our previous study demonstrated that a multimodal gas sensor array integrating multimodal sensors could diagnose LC with exceptionally high accuracy (92.3%) through a 1D CNN model22. However, when the same model architecture was applied to datasets containing multiple diseases, the diagnostic accuracy decreased markedly, indicating that the single-stage CNN structure was limited in handling inter-disease heterogeneity and complex class relationships.
In the present study, we developed an HD-CNN designed to overcome these limitations and achieve dual-cancer classification for LC and GC. Although the overall classification accuracy of the HD-CNN (84.7%) was slightly lower than that of the previous binary LC model, the hierarchical framework maintained stable performance across three classes and exhibited greater potential scalability for future multi-cancer screening. These results suggest that the hierarchical coarse-to-fine structure can balance generalization and sensitivity, providing robustness even when applied to more complex diagnostic tasks.
To date, published breath-based multi-disease diagnostic studies have typically relied on single-type sensor arrays37,38,39,40. These sensors must be fabricated and operated under controlled laboratory conditions. In contrast, our system employs a heterogeneous multimodal sensor array capable of capturing a wider range of metabolic information, improving reproducibility and expanding the coverage of the breath patterns. Moreover, the proposed platform features a relatively simple measurement protocol and compact hardware configuration, which facilitate its potential translation to practical clinical environments through further device miniaturization and modular integration.
Unlike previous studies that utilized statistical approaches such as discriminant factor analysis (DFA), the proposed framework adopts a deep-learning-based real-time pattern recognition strategy. The deep learning model autonomously learns temporal and nonlinear dependencies in sensor responses without explicit feature engineering. Furthermore, the HD-CNN structure successfully achieved three-class classification from sensor responses. This model enables adaptive customization of the network according to disease similarity or hierarchical grouping, which is advantageous for future expansion to broader multi-cancer screening. In summary, while our previous model focused on high-accuracy single-cancer detection, the current HD-CNN platform demonstrates an extensible framework capable of stable performance and scalable integration into next-generation breath-based diagnostic systems.
In summary, we developed a hierarchical deep learning model combined with a multimodal gas sensor array for breath-based screening of multiple cancer types. By integrating the SMO, EC, and PID sensors, the distinct breath patterns were captured. We constructed an HD-CNN model composed of identical CNN architectures for both coarse and fine classifiers, and performed dual-cancer classification through a sequential coarse–fine classification process. The model achieved class-wise accuracies of 82.1% for HC, 84.0% for LC, and 88.1% for GC, and AUCs of 0.89, 0.92, and 0.89, respectively. In comparison, the baseline 1D CNN model showed lower accuracies of 71.6% for HC, 79.8% for LC, and 80.3% for GC, with corresponding AUCs of 0.75, 0.87, and 0.91. The HD-CNN demonstrated superior performance in handling inter-class ambiguity, especially in distinguishing between cancer subtypes. We further analyzed the impact of different coarse–fine configurations and found that isolating HC in the coarse stage, followed by LC and GC fine classification, produced the most accurate and interpretable results. These results validated the potential of our platform as a practical and noninvasive tool for early cancer detection. Future work will focus on expanding the sample size, incorporating additional diseases such as chronic obstructive pulmonary disease (COPD), and validating the system in larger clinical settings.
Methods
Design and development of a breath analyzer
The breath analysis device used in this study was based on our previously developed multimodal sensing platform, which integrates SMO, EC, and PID sensors within a flow-optimized chamber. The same chamber design and sensor configuration as in our earlier work were adopted to maintain uniform gas distribution and multimodal chemical selectivity. Specifically, the chamber geometry was retained as a rectangular structure with chamfered corners, previously verified via fluid dynamics simulation to minimize dead volume and ensure homogeneous flow across sensor positions. Sensor selection for breath analysis prioritizes complementary selectivity (a mixture of sensors differentially sensitive to relevant gas groups), wide dynamic range spanning ppb to ppm, and high reproducibility. Based on these requirements, the analyzer incorporates 20 commercially available sensors that, in combination, provide broad coverage of exhaled VOCs through complementary detection modalities.
In this study, several engineering enhancements were introduced to improve device robustness and measurement reliability. First, a heating control module was redesigned to address instability issues observed in earlier setups, where overheating during thermal desorption of Tenax tubes led to material degradation and signal inconsistency. The new module employs closed-loop feedback to precisely regulate the heating profile, thereby protecting sorbent integrity and ensuring consistent VOC release. Second, to mitigate the effects of ambient temperature fluctuations, the gas chamber was enclosed in a thermally insulated housing. This structural modification minimized temperature-induced baseline drift and improved signal stability during extended operation in clinical environments. Third, an FPGA-based ADC board was implemented for high-speed, high-resolution signal acquisition. Compared to previous microcontroller-based systems, the FPGA architecture allowed stable, synchronized sampling across multiple sensor channels and enhanced compatibility with future sensor integration. Each sensor is sampled by the ADC (ADS8556, Texas Instruments) chip at 50 Hz; signals are routed through the FPGA (XC3S200AN-4FTG256C, AMD Xilinx), and one output value is produced once per second as the 1-s average of those samples. The output update rate is set somewhat faster than the intrinsic response time of MOS gas sensors (typically several seconds), because the aim is to characterize trend patterns in a multimodal gas sensor array exposed to mixed gases rather than to report instantaneous concentrations of specific analytes. The FPGA provides 195 user I/O pins, supports operating frequencies up to ~250 MHz, and achieves data transfer rates of >622 Mb/s per pin, enabling future expansion to additional sensors and facilitating the capture of fine transient features that may arise in high-dimensional sensor-array data. The ADC offers six input channels, allowing simultaneous digitization of up to six analog sensor signals. Finally, the power supply system was upgraded to a multi-channel, noise-isolated configuration. This modification enabled independent and efficient power delivery to each sensor type while minimizing electrical interference, particularly important for sensitive EC channels.
These targeted upgrades address the key limitations of the previous system, resulting in an enhanced breath analysis platform capable of reproducible, high-fidelity VOC profiling suitable for deep learning-based cancer screening.
Study participants
This study was conducted in accordance with the Declaration of Helsinki and received approval from the Institutional Review Board of Seoul National University Bundang Hospital (IRB No. E-1208-167-004). All participants were fully informed about the study procedures, risks, and benefits, and informed consent was obtained from all individuals prior to sample collection. A total of 206 participants were prospectively recruited from Seoul National University Bundang Hospital within the period from September 1, 2022, to October 31, 2022. Breath samples from all participant groups (HC, LC, and GC) were collected in a randomized and interleaved schedule throughout the study period. Participants were recruited concurrently, and sampling sessions for all groups were conducted during overlapping time windows under identical laboratory and environmental conditions. The study cohort consisted of 67 HCs, 78 patients with LC, and 61 patients with GC. The HC group consisted of individuals aged over 18 years without any known history of malignancy or current treatment for respiratory or gastrointestinal diseases. Subjects with active infections, metabolic disorders, or systemic conditions that could affect breath composition were excluded.
LC patients were enrolled based on the following inclusion criteria: (i) histological confirmation of LC from either a primary tumor or a metastatic lesion; (ii) presence of intrabronchial lesions visible through bronchoscopy; (iii) chest computed tomography evidence of central lesions located within the inner one-third of the pulmonary hilum; or (iv) peripheral lesions located beyond the outer one-third of the hilum, with a tumor diameter greater than 2 cm. Patients who had coexisting metabolic diseases (e.g., diabetes mellitus), chronic respiratory diseases such as COPD, pneumonia, or other active pulmonary infections were excluded to avoid confounding effects on exhaled breath composition. In addition, patients diagnosed with minimally invasive adenocarcinoma, corresponding to stage T1a(mi)N0M0, were also excluded.
GC patients were selected based on the following criteria: (i) histological confirmation via endoscopic biopsy, and (ii) presence of peritoneal metastasis confirmed by imaging or diagnostic laparoscopy in the case of GC. The exclusion criteria included patients with early-stage tumors confined to the mucosal layer, such as intramucosal carcinoma; those who had undergone curative endoscopic submucosal dissection prior to surgery; and individuals with coexisting metabolic disorders.
Exhaled breath sample collection
We collected exhaled breath samples from a total of 222 cases, comprising 67 HCs, 94 samples from 78 patients with LC, and 61 samples from patients with GC. In the LC group, multiple breath samples were acquired from some individuals to capture intra-individual variability. The breath collection protocol was uniformly applied across all participants to minimize pre-analytical variation and ensure comparability of sensor responses.
The day before breath collection, participants were advised to consume a light dinner with minimal seasoning to reduce dietary interference. On the day of sampling, a fasting period of at least 4 h was strictly maintained. To prevent oral contamination, participants refrained from brushing their teeth within 2 h prior to breath collection. Instead, they rinsed their mouths thoroughly with 200 mL of sterilized distilled water immediately before sampling. After performing at least five deep inhalation and exhalation cycles, 3 L of exhaled breath was collected into a Tedlar bag. The collected breath sample was transferred to a desorption tube in the breath analyzer within 2 h of collection.
The workflow of the breath sampling procedure is illustrated in Supplementary Fig. 4. After breath collection into a Tedlar bag, the sample was adsorbed onto a desorption tube (Tenax tube) using a controlled flow system (Supplementary Fig. 4a). The measurement chamber was subsequently purged with N2 gas to eliminate any residual VOCs from prior measurements (Supplementary Fig. 4b). To prevent hygiene issues during sampling, we did not reuse the Tedlar bag after transferring the sample to the desorption tube. High-purity N2 was used as the purge and baseline gas to establish an inert, oxygen-free reference environment within the sensor chamber. This minimized undesired oxidation or moisture-induced surface reactions on the sensor materials and provided a stable, reproducible baseline resistance prior to each measurement. Next, the adsorbed compounds were then thermally desorbed by a heating lamp and introduced into the gas chamber, where VOCs were detected using the multimodal sensor array (Supplementary Fig. 4c). As the heated exhaled breath sample is transported into the chamber, it is cooled by the room-temperature carrier gas. Inside the chamber, the sample environment is stabilized at approximately 40 °C with a relative humidity close to zero. Finally, the used Tenax tubes were cleaned and recycled for subsequent use (Supplementary Fig. 4d).
Measurement and analysis of gas sensor response
The voltage signals of the multimodal sensor array were automatically acquired and recorded using a custom-designed measurement software. The sensor response was calculated by normalizing the sensor signal as follows:
where Rbreath refers to the sensor values measured during the entire sampling process, and Rair denotes the baseline signal recorded immediately before the desorption of exhaled breath samples from the Tenax tube.
Deep learning-based analysis and evaluation
Deep learning models of this study were constructed on the open-source library PyTorch version 2.7 (Meta, USA). The deep learning models were trained and validated on a high-performance computing platform equipped with an RTX Titan GPU (NVIDIA, USA). The models evaluated in this study included a previously established 1D CNN model and three hierarchical deep neural network (HD-CNN) models, each incorporating a different coarse classifier structure. All models were trained using randomly assigned 5-fold cross-validation.
The 1D CNN model contains seven layers, including two convolutional layers, two fully connected layers, and one dropout layer. The input to the model is a 2D response map reshaped to (19, 1800), corresponding to 19 sensor channels and 1800 s. The first convolutional layer applies 4 filters with a kernel size of (19, 300) and a stride of 100, effectively capturing local features. The second convolutional layer for nonlinear transformation applies 8 filters with a kernel size of (1, 1). The output from the convolutional layers is flattened and passed to a fully connected layer with 32 neurons. A hidden layer with a 50% drop rate follows. The final output layer consists of three neurons for multi-class classification, using a softmax activation function. All convolutional and fully connected layers are followed by batch normalization and a LeakyReLU activation.
The HD-CNN model architecture introduces a structurally hierarchical design consisting of two independent classifiers: a coarse classifier and a fine classifier. Each classifier adopts the same base architecture as the 1D CNN model, comprising two convolutional layers, two fully connected layers, and a dropout layer. While structurally identical to the 1D CNN, the two classifiers are trained separately. The coarse classifier first performs binary classification, then the fine classifier is selectively activated to differentiate between the remaining two classes. A dedicated probability integration layer combines the outputs of both classifiers to produce final predictions across three classes.
Model weights were initialized using He initialization for convolutional layers and zero initialization for linear layer biases. Training was conducted using the Adam optimizer (learning rate = 0.0002, weight decay = 0.01) and the cross-entropy loss function. All models were trained for up to 500 epochs with a batch size of 64, and a StepLR scheduler was applied to reduce the learning rate by a factor of 0.1 every 100 epochs.
Data availability
All main training results are presented in the main manuscript and Supplementary Information. Due to the inclusion of personally identifiable information, the raw and preprocessed input data are subject to access restrictions and are securely managed by Seoul National University Bundang Hospital. Researchers wishing to access the data must obtain approval from the hospital’s Institutional Review Board (IRB) and receive authorization from the corresponding author.
Code availability
The code used for training the CNN and HD-CNN models is available at https://github.com/danharu-cpu/multi-cancer-screening.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
World Health Organization. Cancer Fact Sheet. https://www.who.int/news-room/fact-sheets/detail/cancer (World Health Organization, 2025).
International Agency for Research on Cancer. Global Cancer Observatory: Cancer Today. https://gco.iarc.fr/today (IARC, 2022).
Tan, N. et al. Global, regional, and national burden of early-onset gastric cancer. Cancer Biol. Med. 21, 667–678 (2024).
Tang, F. H. et al. Recent advancements in lung cancer research: a narrative review. Transl. Lung Cancer Res. 14, 975–990 (2025).
Behera, B. et al. Electronic nose: a non-invasive technology for breath analysis of diabetes and lung cancer patients. J. Breath Res. 13, 024001 (2019).
Güntner, A. T. et al. Breath sensors for health monitoring. ACS Sens. 4, 268–280 (2019).
Vasilescu, A. et al. Exhaled breath biomarker sensing. Biosens. Bioelectron. 182, 113193 (2021).
Hanna, G. B. et al. Accuracy and methodologic challenges of volatile organic compound-based exhaled breath tests for cancer diagnosis JAMA Oncol. 5, e182815 (2019).
Gao, Y. et al. Volatile organic compounds in exhaled breath: applications in cancer diagnosis and predicting treatment efficacy. Cancer Pathog. Ther. 3, 411–419 (2025).
Amor, R. E., Morad, N. K. & Barash, O. Breath analysis of cancer in the present and the future. Eur. Respir. Rev. 28, 190002 (2019).
Heng, W. et al. A smart mask for exhaled breath condensate harvesting and analysis. Science 385, 954–961 (2024).
Chang, J. E. et al. Analysis of volatile organic compounds in exhaled breath for lung cancer diagnosis using a sensor system. Sens. Actuators B Chem. 255, 800–807 (2018).
Lee, B. et al. Breath analyzer for real-time exercise fat burning prediction: oral and alveolar breath insights with CNN. ACS Sens. 10, 2510–2519 (2024).
Del Orbe, D. V. et al. Breath analyzer for personalized monitoring of exercise-induced metabolic fat burning. Sens. Actuators B Chem. 369, 132192 (2022).
Saalberg, Y. & Wolff, M. VOC breath biomarkers in lung cancer. Clin. Chim. Acta 459, 5–9 (2016).
Nooreldeen, R. & Bach, H. Current and future development in lung cancer diagnosis. Int. J. Mol. Sci. 22, 8661 (2021).
Gasparri, R. et al. Volatile signature for the early diagnosis of lung cancer. J. Breath Res. 10, 016007 (2016).
Kim, D., Lee, J., Park, M. K. & Ko, S. H. Recent developments in wearable breath sensors for healthcare monitoring. Commun. Mater. 5, 41 (2024).
Moura, P. C., Raposo, M. & Vassilenko, V. Breath volatile organic compounds (VOCs) as biomarkers for the diagnosis of pathological conditions: a review. Biomed. J. 46, 100623 (2023).
Ates, H. C. & Dincer, C. Wearable breath analysis. Nat. Rev. Bioeng. 1, 80–82 (2023).
Lee, B. et al. Breath analysis system with convolutional neural network (CNN) for early detection of lung cancer. Sens. Actuators B Chem. 409, 135578 (2024).
Gharra, A. et al. Exhaled breath diagnostics of lung and gastric cancers in China using nanosensors. Cancer Commun. 40, 273–278 (2020).
Liu, B. et al. Lung cancer detection via breath by electronic nose enhanced with a sparse group feature selection approach. Sens. Actuators B Chem. 339, 129896 (2021).
Schuermans, V. N. E. et al. Pilot study: detection of gastric cancer from exhaled air analyzed with an electronic nose in chinese patients. Surg. Innov. 25, 429–434 (2018).
Huang, J. et al. Noninvasive diagnosis of gastric cancer based on breath analysis with a tubular surface-enhanced Raman scattering sensor. ACS Sens. 7, 1439–1450 (2022).
Yang, H. Y. et al. Breath biopsy of breast cancer using sensor array signals and machine learning analysis. Sci. Rep. 11, 103 (2021).
Liu, J. et al. A novel non-invasive exhaled breath biopsy for the diagnosis and screening of breast cancer. J. Hematol. Oncol. 16, 63 (2023).
Khan, M. S. et al. Improving multi-organ cancer diagnosis through a machine learning ensemble approach. In Proc. 7th International Conference on Electronics, Communication and Aerospace Technology (ICECA) 1075–1082 (IEEE, 2023).
Zhou, M. et al. Comparison of perioperative outcomes of selective arterial clipping guided by near-infrared fluorescence imaging using indocyanine green versus undergoing standard robotic-assisted partial nephrectomy: a systematic review and meta-analysis. Int. J. Surg. 110, 1234–1244 (2024).
Furuhashi, T. et al. Review of cancer cell volatile organic compounds: their metabolism and evolution. Front. Mol. Biosci. 11, 1499104 (2025).
Danzi, F. et al. To metabolomics and beyond: a technological portfolio to investigate cancer metabolism. Signal Transduct. Target. Ther. 8, 137 (2023).
Ali, S. et al. State-of-the-art challenges and perspectives in multi-organ cancer diagnosis via deep learning-based methods. Cancers 13, 5546 (2021).
Kumar, Y. et al. Automating cancer diagnosis using advanced deep learning techniques for multi-cancer image classification. Sci. Rep. 14, 25006 (2024).
Xiao, Y. et al. A semi-supervised deep learning method based on stacked sparse auto-encoder for cancer prediction using RNA-seq data. Comput. Methods Programs Biomed. 153, 99–105 (2018).
Yan, Z. et al. HD-CNN: Hierarchical deep convolutional neural networks for large scale visual recognition. In Proc. IEEE International Conference on Computer Vision (ICCV) 2740–2748 (IEEE, 2015).
Peng, G. et al. Diagnosing lung, breast, colorectal, and prostate cancers from exhaled breath using gold nanoparticle sensors. Br. J. Cancer 103, 542–551 (2010).
Nakhleh, M. K. et al. Diagnosis and classification of 17 diseases from 1404 subjects via pattern analysis of exhaled molecules. ACS Nano 11, 112–125 (2017).
Gharra, N. et al. Exhaled breath analysis of lung and gastric cancers in a large clinical trial using a nanosensor array. Cancer Commun. 40, 273–278 (2020).
Xie, J. et al. Surface-enhanced Raman spectroscopy coupled with deep learning for non-invasive detection of lung and gastric cancers from exhaled breath. Spectrochim. Acta A Mol. Biomol. Spectrosc. 314, 124181 (2024).
Acknowledgements
B.L. and J.L. contributed equally to this work. This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (No. 2021M3H4A4079271). It was also supported by the K-Sensor Development Programs (RS-2022-00154853 and RS-2022-00154855) funded by the Ministry of Trade, Industry and Energy (MOTIE, Korea). In addition, this research was supported by the National Research Council of Science & Technology (NST) grant by the Korean government (MSIT) (No. GTL25061-000). This work was also supported by the Commercialization Promotion Agency for R&D Outcomes (COMPA) grant funded by the Korean government (Ministry of Science and ICT) (RS-2023-00304776).
Author information
Authors and Affiliations
Contributions
B.L. and J.L. contributed equally to this work and are co-first authors. B.L. and J.L. designed and performed the experiments, conducted data analysis and preprocessing, developed the deep learning framework, and analyzed the model training results. B.L. and J.L. drafted the manuscript. H.N. performed experimental validation and manuscript review. H.B. and J.J. conducted clinical interpretation of results and manuscript revision. I.P., S.J., and D.L. conceived and supervised the project. All authors discussed the results and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, B., Lee, J., Noh, H. et al. Advanced breath analysis through hierarchical deep convolutional neural network for multi-cancer screening. npj Digit. Med. 9, 138 (2026). https://doi.org/10.1038/s41746-025-02319-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02319-1