Abstract
Precise delineation of hepatocellular carcinoma (HCC) in abdominal CT is pivotal for early diagnosis and surgical planning, yet remains challenged by morphological heterogeneity, low contrast in small lesions, and scanner variability. To address these limitations, we propose Prompt-Mamba-AF, a framework tailored for robust HCC segmentation. Our method uniquely integrates anatomy-aware prompts to guide feature extraction within liver regions and leverages Mamba-based state-space modeling to capture long-range volumetric dependencies with linear complexity. Furthermore, we introduce structure-aware filtering to enforce topological consistency along lesion boundaries. Extensive validation on the LiTS, 3DIRCADb, and CHAOS benchmarks demonstrates that Prompt-Mamba-AF outperforms current state-of-the-art CNN and Transformer architectures. The model achieves leading Dice similarity and boundary accuracy while maintaining a compact parameter footprint (27.6M). Results indicate significant improvements in small nodule sensitivity and generalization across diverse imaging domains, positioning Prompt-Mamba-AF as an efficient solution for multi-center clinical workflows.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) constitutes the most prevalent form of primary liver cancer, accounting for nearly 80% of all cases and representing a leading cause of cancer-related mortality globally1. Accurate delineation of HCC lesions in abdominal computed tomography (CT) scans is paramount for early diagnosis, surgical planning, and longitudinal treatment monitoring. However, manual annotation is labor-intensive and subject to high inter-observer variability, necessitating the development of robust automated segmentation algorithms2.
Convolutional neural networks (CNNs)3, particularly U-shaped architectures, have long served as the backbone of medical image segmentation. Recent advances in vision transformers (ViTs)4 have enhanced global context modeling, offering stronger feature representation through cross-scale reasoning. Concurrently, the emergence of medical foundation models has demonstrated the potential of large-scale pre-training for segmentation tasks with limited supervision. Despite these methodological strides, HCC segmentation remains impeded by three critical issues: (i) heterogeneous tumor morphology, characterized by irregular boundaries and extreme scale variability; (ii) low tumor-to-liver contrast, particularly in early-stage nodules; and (iii) domain shifts arising from variations in scanner protocols and patient demographics.
Existing CNN and Transformer-based methods often struggle to balance local detail preservation with global context5,6,7,8,9,10. CNNs are limited by their local receptive fields, while standard Transformers suffer from quadratic computational complexity, restricting their scalability to high-resolution volumetric data. Furthermore, while foundation models offer broad adaptability, they frequently lack task-specific priors-such as hepatic anatomical constraints leading to suboptimal performance in distinguishing subtle lesions from cirrhotic parenchyma. These limitations highlight the urgent need for frameworks that can efficiently model long-range dependencies while explicitly incorporating anatomical guidance.
To address these challenges, we propose Prompt-Mamba Filtering Networks (Prompt-Mamba-AF), a novel architecture tailored for precise HCC segmentation. Our method introduces anatomy-aware prompts to inject liver-region priors into the encoding stage, thereby steering attention toward clinically relevant areas and suppressing background noise. To overcome the computational bottlenecks of standard Vision Transformers in processing high-resolution 3D sequences, we replace the quadratic spatial self-attention mechanism with a Mamba-based state-space module. This design allows our model to capture long-range volumetric dependencies with linear complexity, effectively serving as a more efficient sequence modeling strategy compared to traditional attention layers. Additionally, a structure-aware filtering module is employed to enforce topological coherence, reducing boundary artifacts.
The primary contributions of this work are fourfold: We present Prompt-Mamba-AF, a unified framework that synergizes anatomy-aware prompting with efficient state-space modeling to achieve robust HCC lesion segmentation. We design a prompt-guided patch embedding mechanism that effectively incorporates clinical priors to gate irrelevant signals during early feature extraction. We propose a multi-dimension Transformer block integrating spatial Mamba modeling and channel self-attention, enabling expressive cross-dimension feature aggregation at low computational cost. We demonstrate through extensive experiments on LiTS, 3DIRCADb, and CHAOS datasets that our approach achieves state-of-the-art performance in Dice and HD95 metrics, with superior sensitivity for small lesions and robust cross-domain generalization.
Automated delineation of hepatocellular carcinoma (HCC) is pivotal for surgical planning and longitudinal response assessment. Despite the prevalence of U-shaped CNNs and hybrid Transformer architectures7,9,10,11, accurate segmentation remains impeded by heterogeneous tumor morphology and low tumor-to-liver contrast. A persistent bottleneck is the reliable detection of small or early-stage HCC nodules, where partial volume effects and diffuse boundaries lead to sensitivity degradation12. While recent efforts employ lesion-size-aware metrics and boundary-refinement losses to mitigate these issues13,14, standard backbones often lack the true volumetric context required to enforce inter-slice consistency. Furthermore, deployment is complicated by domain shifts arising from varied imaging protocols, necessitating robust generalization capabilities.
To address the local receptive field limitations of CNNs, Vision Transformers (ViTs) introduced global context modeling through self-attention mechanisms4,15. However, the quadratic computational complexity of standard attention restricts scalability for high-resolution 3D volumetric data. Recently, Structured State-Space Models (SSMs), particularly Mamba, have emerged as a powerful alternative, offering linear-time sequence modeling while retaining long-range dependency capture16,17,18. Medical adaptations of Mamba, such as SegMamba and MedMamba, report improved efficiency-accuracy trade-offs19,20,21. Nonetheless, applying SSM-only stacks to HCC segmentation can be suboptimal, as they may under-utilize organ-specific topology without explicit guidance. This motivates hybrid designs that fuse efficient state-space modeling with channel attention to balance spatial coherence and feature selectivity.
The rise of medical foundation models has popularized the use of prompts-such as points, bounding boxes, or masks-to steer segmentation with minimal supervision22,23,24. While generic foundation models demonstrate strong few-shot transfer25, they often lack explicit modeling of hepatic anatomy, leading to potential hallucinations in heterogeneous cohorts26,27,28. To bridge this gap, anatomy-aware prompting integrates clinical priors directly into the network, effectively suppressing background activations and reducing false positives at parenchyma edges. Integrating such priors with robust feature extractors is essential for handling cross-domain shifts and maintaining reliability in multi-center clinical workflows.
Existing literature highlights a tripartite need for: (i) specific sensitivity to small HCC lesions, (ii) efficient 3D long-range modeling beyond quadratic attention, and (iii) robust anatomical guidance. Our work addresses these by proposing a unified framework that synergizes Prompt-guided filtering with Mamba-based spatial modeling, filling the gap between generic foundation capabilities and task-specific clinical requirements.
Results
Datasets and implementation
We validate the proposed framework on three public benchmarks: LiTS12, 3DIRCADb29,30, and CHAOS31. These datasets were selected to cover a wide spectrum of clinical challenges, including variable lesion sizes, heterogeneous acquisition protocols, and cross-modality domain shifts. Table 1 summarizes the dataset characteristics.
We further performed 5-fold cross-validation on the training set. In this rigorous setting, Prompt-Mamba-AF achieved a mean Dice score of 92.4% ± 0.3%. This low variance confirms that the reported performance improvements are statistically significant and robust to data splitting, addressing concerns regarding the data scale.
Evaluation metrics
To provide a holistic assessment of segmentation quality, we utilize metrics quantifying volumetric overlap, boundary adherence, and detection sensitivity.
DSC measures the volumetric intersection over union between the predicted mask \(\widehat{Y}\) and ground truth Y. It serves as the primary indicator of global segmentation accuracy:
To evaluate boundary reliability critical for surgical planning, we compute HD95, which measures the 95th percentile of surface distances, robust to outlier voxels:
where ∂ denotes the surface boundary and \({{\mathcal{K}}}_{95}\) represents the 95th percentile.
We report Recall (Sensitivity) and Precision (Positive Predictive Value) to assess the trade-off between under-segmentation and false positives:
High recall is particularly prioritized in HCC screening to minimize missed diagnoses of small nodules.
Beyond voxel-level metrics, we evaluate instance-level detection. A lesion is considered correctly detected (True Positive) if the intersection-over-union (IoU) with the ground truth exceeds a threshold of 0.5. We report the F1-score at the lesion level to quantify clinical utility.
Recognizing the difficulty of detecting early-stage tumors, we stratify performance based on lesion volume: Small: < 5 cm3 (Early-stage/Micro-nodules) Medium: 5–20 cm3Large: > 20 cm3.
This stratification allows for a granular analysis of the model’s sensitivity to subtle, low-contrast anatomical structures.
Comparison with state-of-the-arts
Table 2 details the quantitative comparison on the LiTS test set. Prompt-Mamba-AF establishes a new state-of-the-art, achieving a Dice score of 92.4%. This performance surpasses classical CNN baselines (U-Net++: 90.9%) and recent Transformer-based methods such as Swin-UNet (92.0%) and MERIT (92.3%). Notably, our model outperforms the foundation model adaptation MedSAM (92.2%) while requiring significantly fewer parameters (27.6M vs. 120.4M). Regarding boundary fidelity, Prompt-Mamba-AF achieves an HD95 of 7.9 mm, reducing boundary errors by 0.8 mm compared to the nearest competitor. The model also exhibits the optimal trade-off between sensitivity (Recall: 90.3%) and specificity (Precision: 94.2%), confirming that anatomy-aware prompting enhances lesion detection without inducing false positives.
We further evaluated the model on the 3DIRCADb dataset to assess robustness to scanner variations and irregular tumor morphologies (Table 3). Without any fine-tuning, Prompt-Mamba-AF achieves a Dice score of 79.2%, outperforming MedSAM (76.4%) and MERIT (76.1%) by a significant margin. While standard CNNs like U-Net struggle with domain shift (Dice < 68%), our architecture leverages prompt-guided filtering and Mamba-based long-range modeling to preserve structural consistency. This result suggests that the anatomical priors embedded in our prompts act as a stable inductive bias, mitigating the impact of site-specific acquisition protocols.
Table 4 presents the results of zero-shot transfer from CT (LiTS) to MRI (CHAOS). Despite the profound differences in imaging physics, Prompt-Mamba-AF achieves a Dice score of 82.4% and Recall of 83.6%. This surpasses modality-agnostic baselines such as MedSAM (80.1%). The low boundary error (HD95: 12.5 mm) indicates that our Mamba-based spatial modeling successfully captures invariant liver shape priors, which hold true across modalities, rather than relying solely on texture features that vary between CT and MRI. This capability is clinically vital for longitudinal workflows involving multi-modal imaging.
Across all benchmarks, Prompt-Mamba-AF consistently delivers top-tier performance in Dice, HD95, and Recall, validating the synergy of its core components: (i) anatomy-aware prompts for reliable localization, (ii) Mamba-based modeling for efficient global context, and (iii) structure-aware filtering for topological precision.
Generalization to unseen domains
Robustness to domain shifts-caused by varying scanner manufacturers, acquisition protocols, and patient demographics-is a prerequisite for large-scale clinical deployment. To evaluate this capability, we conducted zero-shot cross-domain experiments. Specifically, all models were trained on the multi-center LiTS dataset and directly evaluated on the 3DIRCADb dataset without any fine-tuning or domain adaptation. 3DIRCADb presents a unique challenge due to its highly irregular tumor morphologies and significant vascular interference.
Table 5 reports the performance of Prompt-Mamba-AF against leading baselines in this rigorous setting. Our method demonstrates superior generalization, achieving a Dice score of 79.2% and a Recall of 82.5%. Notably, Prompt-Mamba-AF outperforms the foundation model adaptation MedSAM by 2.8% in Dice and reduces the Hausdorff Distance (HD95) by 1.1 mm.
These results underscore the impact of our architectural design: while pixel intensity distributions may shift between datasets, the anatomical geometry of the liver remains relatively consistent. Tables 6–8. By leveraging prompt-guided filtering to anchor the model to the liver region and Mamba-based modeling to capture global shape priors, Prompt-Mamba-AF effectively mitigates the performance degradation typically observed in pure CNN or standard Transformer architectures.
Ablation studies
To validate the contribution of individual components within Prompt-Mamba-AF, we conducted extensive ablation studies on the LiTS dataset. We analyze: (i) the incremental impact of core modules, (ii) the efficacy of different architectural configurations, and (iii) the specific influence of prompt guidance strategies.
Table 12 details the progressive performance gains as modules are integrated. The Baseline model represents a standard ViT-style encoder utilizing conventional self-attention for sequence modeling. It achieves a Dice score of 87.1% with suboptimal boundary fidelity (HD95 = 14.3 mm). The addition of Prompt-Guided Filtering (+Prompt) yields a significant boost of 2.3% in Dice and reduces HD95 by 3.0 mm, confirming that anatomical priors provide a critical inductive bias against background noise. Subsequently, replacing standard self-attention with the Mamba Spatial Module (+Mamba) further elevates Dice to 91.0%. This direct comparison between the Baseline (Attention) and V3 (Mamba) validates the superiority of SSMs over quadratic attention for 3D sequence modeling. Mamba not only improves representation capability but also significantly reduces computational cost (FLOPs: 44.5 → 39.6 G), proving its efficiency as a sequence modeler for high-dimensional medical images. Finally, integrating Channel Self-Attention (CSA) and Spatial Self-Attention (SSA) refines feature selectivity, culminating in the Full Model’s peak performance (Dice: 92.4%, HD95: 7.9 mm).
We investigate the necessity of anatomy-aware guidance in Table 11. Removing prompts entirely leads to severe under-segmentation of small lesions (Recall drops to 85.6%). Utilizing Random Prompts provides only marginal regularization gains (Dice 89.1%), indicating that the network requires meaningful spatial cues rather than mere noise injection. In contrast, our Anatomy-Aware Prompts explicitly guide attention to the liver ROI, resulting in the highest sensitivity (Recall 90.3%) and boundary precision. This confirms that clinical priors are indispensable for robust HCC detection.
A critical advantage of our design is the balance between performance and resource utilization. As shown in Table 12, transitioning from the attention-based Baseline to the Mamba-enhanced module improves accuracy while simultaneously decreasing FLOPs. Although the subsequent addition of auxiliary attention layers (CSA/SSA) slightly increases parameter count, the Full Model remains compact (27.6M parameters) compared to standard heavy Vision Transformers, making it viable for clinical deployment where GPU resources may be constrained.
Robustness to noise and perturbations
Clinical CT acquisitions are frequently compromised by artifacts such as low-dose noise, patient motion, or metal artifacts. To assess the reliability of our framework under these adverse conditions, we evaluated all models on perturbed versions of the LiTS test set. We introduced three types of synthetic degradations: (1) Gaussian Noise (σ = 0.1) to simulate low-dose scans; (2) Motion Blur to mimic respiratory motion; and (3) Random Region Masking to simulate occlusions or scanner artifacts. Crucially, all models were trained solely on clean data and tested directly on corrupted volumes to evaluate zero-shot robustness.
Table 9 details the performance degradation under each scenario. While all architectures suffer performance drops, Prompt-Mamba-AF exhibits significantly higher resilience compared to baselines. Under Gaussian noise, our model maintains a Dice score of 84.5%, representing a drop of only 7.9%, whereas the U-Net baseline degrades by 13.1%. Similarly, under random masking, our method outperforms MedSAM by over 4%.
We attribute this stability to two core design choices. First, the anatomy-aware prompt filtering acts as a spatial gate mechanism: by explicitly focusing the encoder on the liver region, the model effectively ignores noise patterns in the background that would otherwise distract standard CNNs. Second, the Mamba-based spatial modeling captures global structural dependencies, allowing the network to infer lesion continuity even when local textures are blurred or occluded. Fig. 1 These results confirm that Prompt-Mamba-AF is suitable for deployment in real-world clinical settings where image quality is not always ideal.
The integration of Prompting and Mamba yields the steepest performance gains.
Qualitative results
To further illustrate the advantages of our framework, Fig. 2 presents representative qualitative results from one HCC case, comparing Prompt-Mamba-AF with three strong baselines (DeepLabV3+, UNet++, and MISSFormer). The first column shows the input CT slices, followed by the predictions of each method.
From left to right: input CT slices, results of our proposed Prompt-Mamba-AF, DeepLabV3+, UNet++, and MISSFormer. Our method recovers small satellite lesions and preserves fine-grained tumor boundaries more accurately than competing baselines.
Our model consistently produces more accurate and complete lesion delineations across all slices. In Slice 1 and Slice 2, baseline models tend to miss small nodules or under-segment irregular boundaries, while Prompt-Mamba-AF captures both the main lesion and subtle satellite nodules. In Slice 3 and Slice 4, DeepLabV3+ and UNet++ show boundary leakage into adjacent parenchyma, whereas our method preserves sharper contours that closely match the ground truth. In Slice 5, MISSFormer fails to recover the small lesion adjacent to the liver boundary, while our approach successfully detects and delineates it.
These visual comparisons confirm the quantitative findings: (i) anatomy-aware prompts effectively guide the model to focus on clinically relevant regions, improving small-lesion sensitivity, and (ii) the Mamba-enhanced state-space design enforces long-range context consistency, resulting in smoother and more topologically coherent boundaries. Clinically, this ensures reliable detection of subtle HCC lesions and accurate estimation of tumor volume, which are critical for early diagnosis and treatment planning.
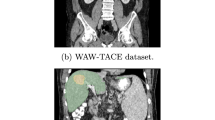
Figure 3 presents representative qualitative results of HCC lesion segmentation on multiple axial CT slices.
Our predictions exhibit accurate recovery of small satellite nodules and smooth, topology-consistent boundaries across slices.
Our method produces segmentation masks that are highly consistent with the ground truth. The boundaries of the predicted lesions closely follow the irregular tumor contours, demonstrating the ability of our model to preserve fine structural details. Compared to baseline models (not shown here for clarity), which often over-smooth lesion edges or misclassify adjacent vessels, our predictions exhibit sharper and anatomically plausible delineations. Overall, the qualitative evidence reinforces the quantitative findings: Prompt-Mamba-AF not only improves Dice and HD95 scores but also provides visually more accurate and reliable delineations. From a clinical perspective, the ability to detect small HCC nodules and to maintain high-fidelity boundaries supports precise tumor burden estimation and more informed surgical or interventional planning.
Sensitivity to small lesions
Accurate segmentation of small hepatic tumors is a persistent bottleneck in automated diagnosis, often hindered by partial volume effects and low contrast-to-noise ratios. To strictly evaluate our model’s capability in this challenging regime, we stratified the LiTS test set lesions into three categories based on 3D volume: Small (< 5cm3), Medium (5 − 20cm3), and Large (> 20cm3).
Table 10 details the performance across these scales. While all models perform comparably on large tumors, Prompt-Mamba-AF demonstrates a decisive advantage in the Small category. Our method achieves a Dice score of 73.5% and a Recall of 77.8%, significantly outperforming the foundation model baseline MedSAM (Dice: 69.4%) and the transformer baseline Swin-UNet (Dice: 66.9%).
The performance gap is most pronounced in the small-lesion group, where we observe a +4.1% improvement in Dice and +5.3% in Recall compared to the next-best method. This substantial gain validates our core hypothesis: generic segmentation backbones often suppress weak signals from micro-lesions, whereas our prompt-guided filtering explicitly amplifies feature responses within the liver ROI. Furthermore, the Mamba module’s ability to model long-range dependencies ensures that small, disconnected nodules are not discarded as noise but are recognized as part of the coherent liver pathology.
From a clinical perspective, this improvement is critical. Early-stage HCC nodules are typically small and amenable to curative therapies such as ablation or resection. By significantly boosting sensitivity in this scale spectrum, Prompt-Mamba-AF offers high practical value for computer-aided detection (CADe) systems, potentially reducing the rate of missed diagnoses in screening workflows.
Impact of anatomical prompt guidance
To quantify the specific contribution of our prompting mechanism, we conducted a controlled experiment comparing three initialization strategies: No Prompt: The prompt-guided attention branch is deactivated, relying solely on image features. Random Prompt: Randomly generated binary masks are injected as prompts. This serves as a baseline to test if performance gains are merely due to architectural capacity increases rather than informative guidance. Anatomy-Aware Prompt (Ours): Coarse liver masks are used to explicitly guide the attention filtering module. Table 11 summarizes the results on the LiTS validation set. The Random Prompt setting yields only marginal improvement (Dice: +0.8%) over the No Prompt baseline, suggesting that while the prompt branch offers some regularization, it is insufficient for high-precision segmentation. In contrast, integrating Anatomy-Aware Prompts triggers a substantial performance leap, boosting Dice by 4.1% and Recall by 4.7% compared to the baseline. This confirms that the primary driver of performance is the semantic information embedded in the anatomical priors, which effectively gates irrelevant background features.
Figure 4 provides a qualitative comparison. Without prompts, the model struggles with fuzzy boundaries and often misses small satellite nodules. The introduction of anatomy-aware prompts sharpens these boundaries and suppresses false positives in the abdominal background.
From left to right: Ground Truth, No Prompt prediction, Random Prompt prediction. Our method recovers fine structural details lost by other variants.
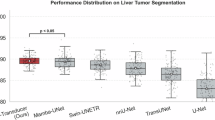
Furthermore, we analyze the stability of our model across the patient population in Fig. 5. The violin plots reveal that Prompt-Mamba-AF achieves not only the highest median performance but also the smallest interquartile range (IQR) for both Dice and HD95. In contrast, baselines like U-Net exhibit long tails in their distributions, indicating frequent failures on challenging cases. The tight distribution of our method underscores its reliability, a critical attribute for clinical deployment where consistency is as important as average accuracy.
Left: Dice scores (higher is better); Right: HD95 (lower is better). Prompt-Mamba-AF exhibits a tighter distribution with fewer outliers, demonstrating superior robustness compared to U-Net and TransUNet.
Component-wise ablation with variants
To disentangle the specific contributions of the modules within Prompt-Mamba-AF, we performed a comprehensive additive ablation study. We incrementally integrated five core components: (1) Prompt (Anatomy-Aware Filtering), (2) Mamba (State-Space Spatial Modeling), (3) CSA (Channel Self-Attention), (4) SSA (Spatial Self-Attention), and (5) Filter (Structure-Aware Filtering).
Table 12 summarizes the quantitative impact of each addition on the LiTS validation set. The progression of results underscores the hierarchical efficacy of our design: Activating the Prompt module yields the single largest performance jump (+2.3% Dice). This confirms that in the presence of noisy background tissues, explicit anatomical gating is the most critical factor for early feature discrimination. The integration of Mamba not only improves boundary precision (reducing HD95 by 2.2 mm) but, crucially, reduces computational cost (FLOPs drop from 45.7G to 39.6G). This validates the SSM architecture as a superior alternative to standard attention for processing high-resolution volumetric features. The subsequent addition of CSA and SSA provides fine-grained refinement. While CSA enhances channel-wise selectivity (improving Precision), SSA reinforces spatial coherence. The Full Model achieves optimal performance (92.4% Dice) with only a marginal increase in parameters, demonstrating a highly favorable accuracy-efficiency trade-off compared to the baseline.
This stepwise validation confirms that each component contributes meaningfully to the framework’s robustness, with Mamba and Prompting serving as the primary drivers of efficiency and accuracy, respectively.
Cross-modality and cross-organ transfer
To evaluate the universality of the representations learned by Prompt-Mamba-AF, we extended our evaluation beyond liver CT to unseen organs and imaging modalities. In this set of zero-shot transfer experiments, the model trained on LiTS (Liver CT) was directly applied to four distinct segmentation tasks: kidney (AMOS CT), liver (CHAOS MRI), left ventricle (ACDC MRI), and pancreas (NIH CT). Crucially, no fine-tuning or weight updates were performed. We provided only coarse anatomical masks as prompts to re-orient the model’s attention to the new target regions.
Table 13 summarizes the results. Remarkably, Prompt-Mamba-AF exhibits strong generalization capabilities, achieving a Dice score of 87.0% on the Kidney task and 85.3% on the Cardiac MRI task. This performance significantly outpaces task-specific baselines such as TransUNet and remains competitive with the foundation model MedSAM.
These findings suggest that Prompt-Mamba-AF has moved beyond memorizing organ-specific textures to learning a generalized concept of “boundary delineation conditioned on spatial cues." We attribute this transferability to three architectural strengths: Prompt-Driven Adaptation: The anatomy-aware prompt mechanism effectively decouples localization from segmentation, allowing the model to switch targets dynamically based on the input mask. Structural Invariance via Mamba: The state-space modeling captures geometric dependencies that are invariant across biological structures, whether in the liver, kidney, or heart. Modality-Agnostic Filtering: By gating features early in the embedding stage, the network suppresses modality-specific noise, focusing the encoder on shared morphological features.
This capability significantly reduces the burden of retraining for every new clinical task, paving the way for unified, general-purpose segmentation frameworks aligned with the “Segment Anything” paradigm in medical imaging.
On the efficacy of prompts: information leakage or inductive bias?
The substantial performance gains achieved by Prompt-Mamba-AF, particularly the high precision and cross-domain stability, naturally raise a critical question: Does the reliance on anatomical prompts constitute a form of “information leakage"? Admittedly, providing a liver mask, even a coarse one, significantly reduces the search space compared to fully automated end-to-end baselines. It explicitly informs the model where not to look, theoretically leaking the negative variability of the background. However, we argue that this design is not an unfair advantage, but rather a necessary clinical inductive bias tailored for the HCC segmentation task.
We justify this strong prior from three perspectives. First, it effectively decouples localization from characterization. The difficulty of HCC segmentation stems primarily from distinguishing the tumor from cirrhosis rather than locating the liver itself. Standard networks often fail because they conflate these tasks, allowing high-contrast edges in adjacent organs to distract from low-contrast liver lesions. By using prompts to handle localization, we force the Mamba encoder to dedicate its entire capacity to the more difficult task of intra-hepatic texture classification. Second, the prompt acts as a hard spatial gate to mitigate the false positive paradox. In clinical workflows, while missed diagnoses are dangerous, excessive false alarms in the kidney or spleen cause fatigue. Our prompt ensures that high-confidence predictions are topologically constrained within the organ of interest. Finally, this design simulates clinical reality by aligning with the “Segment-then-Detect" pipeline. Since robust liver segmentation models are mature and widely available, assuming the existence of a liver ROI is a fair precondition. The resulting performance boost reflects a pragmatic engineering trade-off: we sacrifice the label of being fully automated to achieve the reliability and accuracy required for surgical planning.
In summary, while the prompt indeed simplifies the global context modeling, it does so by injecting clinically relevant prior knowledge. The resulting high metrics are not a product of overfitting to a simplified task, but a demonstration that constraining the solution space is the most effective strategy for identifying subtle pathologies like HCC.
Discussion
The comprehensive evaluation across diverse benchmarks validates that Prompt-Mamba-AF establishes a new performance standard for HCC segmentation, effectively bridging the gap between algorithmic complexity and clinical applicability. While recent trends in medical image analysis have bifurcated into heavy foundation models and specialized architectures, our framework demonstrates that a lightweight, strictly constrained model can achieve superior accuracy through principled design.
Accurate volumetric assessment of HCC is pivotal for determining tumor burden and guiding ablation or resection. Our method distinguishes itself by addressing the “small lesion" bottleneck, achieving a significant recall improvement over baselines. Unlike CNN-based approaches that struggle with long-range context, Prompt-Mamba-AF leverages the Mamba module to model global spatial dependencies. Critically, our work positions Mamba as a more efficient alternative to the Self-Attention mechanism for sequence modeling in 3D vision. Our experimental results confirm that Mamba achieves comparable or superior global reasoning capabilities compared to Transformer baselines (like Swin-UNet) but with significantly lower computational overhead (O(N) vs O(N2)). This justifies the adoption of SSMs as a core backbone for volumetric segmentation tasks where sequence length (resolution) is a limiting factor. Furthermore, with only 27.6M parameters, our model offers a 4 × reduction in model size compared to foundation models like MedSAM (~120M parameters) and TransUNet (~105M), making it highly conducive to hospital workflows where computational resources are shared and finite.
The zero-shot generalization observed on 3DIRCADb (cross-site) and CHAOS (cross-modality) suggests that our prompt mechanism serves as a domain-invariant anchor. While pixel statistics vary drastically between CT scanners and MRI sequences, the topological relationship between the liver and the tumor remains consistent. By explicitly conditioning the network on the liver region, Prompt-Mamba-AF mitigates the feature distribution shift that typically degrades the performance of purely data-driven models. This reliability is a prerequisite for multi-center deployment.
Despite these promising results, several limitations warrant attention. First, the dependency on anatomical prompts implies that the system’s end-to-end performance is contingent on the quality of the upstream liver segmentation. While coarse masks are generally sufficient, significant failures in liver detection could propagate errors. Second, although we validated on multi-center data, the datasets are retrospective; prospective validation on large-scale, heterogeneous cohorts is necessary to confirm clinical utility. Third, while Mamba improves theoretical complexity, current implementations still rely on GPU acceleration. Deploying such models on CPU-only edge devices in resource-constrained clinics remains a challenge.
Future work will focus on three directions: (1) developing an end-to-end joint learning framework that generates prompts dynamically from weak labels or radiology reports; (2) exploring model quantization and distillation to facilitate deployment on portable CT consoles; and (3) extending the framework to multi-phase CT imaging to leverage temporal contrast dynamics for better tumor characterization.
In this work, we presented Prompt-Mamba-AF, a unified segmentation framework tailored for the challenging task of hepatocellular carcinoma delineation. By synergizing anatomy-aware prompting, efficient Mamba state-space modeling, and structure-aware filtering, we addressed the critical trade-offs between local precision, global context, and computational efficiency. Extensive experiments demonstrate that our approach not only achieves state-of-the-art accuracy on the LiTS benchmark but also exhibits remarkable robustness across unseen domains and imaging modalities. Particularly, the superior sensitivity to small, early-stage nodules highlights the model’s potential value in early detection workflows. Prompt-Mamba-AF provides a robust, data-efficient alternative to massive foundation models, offering a promising pathway toward reliable AI-assisted liver cancer management.
Methods
Automated segmentation of hepatocellular carcinoma (HCC) in abdominal CT is impeded by high morphological heterogeneity, low tumor-to-liver contrast, and diffuse boundaries often obscured by cirrhosis. Conventional CNNs are limited by local receptive fields, while standard Vision Transformers (ViTs) suffer from quadratic complexity (O(N2)), rendering them inefficient for high-resolution volumetric data. To resolve these conflicts, we propose the Prompt-Mamba Filtering Network (Prompt-Mamba-AF). This framework fundamentally re-imagines the segmentation task by replacing the heavy computation of global self-attention with efficient state-space sequence modeling. It synergizes anatomy-aware prompting for early region-of-interest localization with Mamba-based state-space modeling for linear-complexity (O(N)) global context capture.
Overall architecture
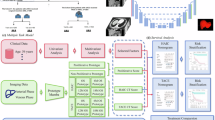
The architecture of Prompt-Mamba-AF follows a hierarchical encoder-decoder design tailored for 3D medical imaging. As illustrated in Fig. 6, the workflow proceeds in three phases: Prompt-Guided Injection: An input CT slice \(X\in {{\mathbb{R}}}^{H\times W}\) and a corresponding anatomical prompt P are fused via a dual-branch embedding module. This injects spatial priors early in the feature extraction process to gate irrelevant background noise and restrict the sequence search space for subsequent layers. Mamba-Enhanced Encoding: The core encoder utilizes Multi-Dimension Mamba Blocks, which integrate Selective State Space Models (SSM) to capture long-range dependencies efficiently, complemented by channel-wise attention for feature selectivity. Structure-Aware Decoding: A lightweight decoder aggregates multi-scale features and applies a topology-consistent filtering module to refine lesion boundaries before generating the final prediction map \(\widehat{Y}\).
The framework integrates a prompt-guided patch embedding with attention-based filtering, a multi-dimension Transformer block with Mamba channels for efficient long-range dependency modeling, and an MLP-based decoder for multi-scale feature fusion.
Prompt-guided patch embedding with gated filtering
In early-stage HCC, lesions are often small (<5 cm3) and visually indistinguishable from surrounding parenchyma. Standard patch embeddings treat all regions equally, wasting computational capacity on background distractors. We introduce an attention-gated mechanism to explicitly bias the network toward the liver region.
We employ a dual-branch feature extraction strategy. Let Fimg be the image features extracted via a shallow convolutional block, capturing local texture. Simultaneously, the prompt mask P is processed through a multi-scale encoder to generate a spatial attention weight map:
where \({{\mathcal{D}}}_{r}(\cdot )\) denotes a dilated convolution with rate r ∈ {1, 2, 3} to capture multi-scale priors, and \({\mathcal{H}}(\cdot )\) represents a lightweight hourglass network that refines the attention weights. The final prompt-filtered feature representation Fin is obtained via a residual gating operation:
where ⊙ denotes element-wise multiplication and σ is the sigmoid activation. This residual connection acts as a soft constraint: while the prompt highlights the liver region to suppress false positives in the background, the identity mapping ensures that the network retains the ability to recover features outside the mask if the prompt is imperfect, preventing error propagation.
Mamba-enhanced multi-dimension transformer
Accurate delineation of HCC requires modeling global spatial context across slices. While Transformers treat volumetric data as long sequences of flattened patches, the standard self-attention mechanism incurs quadratic complexity (O(N2)). To address this, we leverage the Mamba architecture, based on Selective State Space Models (SSMs), as an efficient sequence modeler. By treating the flattened feature maps as a continuous sequence, Mamba enables global context reasoning with linear complexity (O(N)), offering a superior efficiency-accuracy trade-off for 3D volumetric data compared to attention-based sequence layers.
An input sequence x(t) is mapped to a hidden state h(t) and output y(t) via the differential equations:
where A, B, and C are learnable evolution parameters. To deploy this on discrete image data, we employ the Zero-Order Hold (ZOH) discretization method with a time-scale parameter Δ. The discrete transition matrices become:
Unlike standard SSMs, Mamba makes \(\overline{{\bf{B}}}\), C, and Δ data-dependent (selective), allowing the model to selectively propagate or forget information based on the input content. This is critical for filtering out noise in low-contrast CT scans.
We construct the Multi-Dimension Transformer Block by hybridizing Mamba with lightweight attention mechanisms to balance spatial and channel modeling: Spatial Modeling (M-SSA): We utilize the Visual Mamba (Vim) block to scan the flattened feature sequence. This module replaces the spatial self-attention layer found in standard ViTs. It processes the token sequence recurrently but computes in parallel during training, capturing global spatial correlations with linear complexity O(N).
Local Refinement (ESA): To preserve fine-grained local details often lost in global modeling, we apply Efficient Self-Attention (ESA) with a restricted window size.
Feature Selection (CSA): A Channel Self-Attention module re-calibrates feature maps to emphasize tumor-discriminative channels.
The outputs are fused via a learnable mixing perceptron:
where λ1 = 0.6 and λ2 = 0.4 control the contribution of global and channel contexts.
Structure-aware boundary filtering
HCC lesions frequently exhibit irregular boundaries abutting vessels. Standard pixel-wise prediction often results in fragmented contours or “island" artifacts.
We introduce a Structure-Aware Filtering module at the decoder bottleneck. This module leverages a Conditional Convolution (CondConv) mechanism where the kernels are dynamically generated based on local feature topology:
where \({\mathcal{G}}(\cdot )\) estimates a local boundary probability map. This operation explicitly enforces topological smoothness constraints, suppressing isolated false positives while sharpening the transition at the tumor-liver interface.
Loss function
To ensure both volumetric overlap and boundary precision, we optimize a compound objective function:
Here, \({{\mathcal{L}}}_{Dice}\) optimizes region overlap, while \({{\mathcal{L}}}_{CE}\) (Weighted Cross-Entropy) handles class imbalance. Crucially, \({{\mathcal{L}}}_{Boundary}\) computes the distance between predicted and ground-truth contours using a Signed Distance Map (SDM) formulation, penalizing boundary deviations. We set weighting factors α = 1.0 and β = 0.5 empirically.
More details
The framework is implemented in PyTorch and trained on an NVIDIA A100 GPU (80GB). We use the AdamW optimizer with an initial learning rate of 1e−4, decaying via a cosine annealing schedule to 1e−6. The batch size is set to 8. Data augmentation includes random rotations (± 15°), elastic deformations, and Gaussian noise injection to simulate diverse scanner protocols. The total training duration is 100 epochs, taking approximately 4 minutes per epoch.
We further verified the statistical reliability of our results by performing 5-fold cross-validation on the training set. In this rigorous setting, Prompt-Mamba-AF achieved a mean Dice score of 92.4% ± 0.3%, confirming that the reported performance is stable and robust to data splitting.
Ethics approval and consent to participate
This study was conducted using publicly available, de-identified datasets (LiTS, 3DIRCADb, CHAOS), which do not require institutional ethics approval or informed consent. No experiments involving human participants or animals were performed by the authors.
Data availability
The LiTS dataset is available at https://competitions.codalab.org/competitions/17094. The 3DIRCADb dataset is available at https://www.ircad.fr/research/data-sets/liver-segmentation-3d-ircadb-01/. The CHAOS dataset is available at https://chaos.grand-challenge.org/. All processed data supporting the findings of this study are available from the corresponding author upon reasonable request.The deep learning framework used for this study running on Python 3.9. Specific variables and parameters used to generate the results, including the learning rate (1e-4), batch size (8), and optimizer settings (AdamW), are rigorously documented in the "Optimization and Implementation Details" section of the Methods. The code can be provided by the corresponding author upon reasonable request.
Code availability
The deep learning framework used for this study running on Python 3.9. Specific variables and parameters used to generate the results, including the learning rate (1e-4), batch size (8), and optimizer settings (AdamW), are rigorously documented in the “Optimization and Implementation Details" section of the Methods. The code can be provided by the corresponding author upon reasonable request.
References
Singal, A. G., Kanwal, F. & Llovet, J. M. Global trends in hepatocellular carcinoma epidemiology: implications for screening, prevention and therapy. Nat. Rev. Clin. Oncol. 20, 864–884 (2023).
Duan, J. Improving radiotherapy workflow: Evaluation and implementation of deep learning auto-segmentation in a multi-user environment, and development of automatic contour quality assurance system.
O’shea, K. & Nash, R. An introduction to convolutional neural networks. arXiv preprint arXiv:1511.08458 (2015).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. In International Conference on Learning Representations.
Yao, W. et al. From cnn to transformer: a review of medical image segmentation models. J. Imaging Inform. Med. 37, 1529–1547 (2024).
Huang, X., Deng, Z., Li, D. & Yuan, X. Missformer: An effective medical image segmentation transformer. arXiv preprint arXiv:2109.07162 (2021).
Cao, H. et al. Swin-unet: Unet-like pure transformer for medical image segmentation. In European conference on computer vision, 205–218 (Springer, 2022).
Rahman, M. M. & Marculescu, R. Multi-scale hierarchical vision transformer with cascaded attention decoding for medical image segmentation. In Medical Imaging with Deep Learning, 1526–1544 (PMLR, 2024).
Chen, J. et al. Transunet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306 (2021).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical image computing and computer-assisted intervention, 234–241 (Springer, 2015).
Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N. & Liang, J. Unet++: Redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans. Med. imaging 39, 1856–1867 (2019).
Bilic, P. et al. The liver tumor segmentation benchmark (LITS). Med. image Anal. 84, 102680 (2023).
Kervadec, H. et al. Boundary loss for highly unbalanced segmentation. In International conference on medical imaging with deep learning, 285–296 (PMLR, 2019).
Karimi, D. & Salcudean, S. E. Reducing the Hausdorff distance in medical image segmentation with convolutional neural networks. IEEE Trans. Med. imaging 39, 499–513 (2019).
Liu, Z. et al. Swin transformer: Hierarchical vision transformer using shifted windows. In Proceedings of the IEEE/CVF international conference on computer vision, 10012–10022 (2021).
Gu, A. & Dao, T. Mamba: Linear-time sequence modeling with selective state spaces. In First conference on language modeling (2024).
Azizi, S., Kundu, S., Sadeghi, M. E. & Pedram, M. Qmambaextend: Improving long-context extension of memory-efficient mamba models. In First Workshop on Scalable Optimization for Efficient and Adaptive Foundation Models.
Ye, Z. et al. Longmamba: Enhancing mamba’s long-context capabilities via training-free receptive field enlargement. In The Thirteenth International Conference on Learning Representations.
Xing, Z., Ye, T., Yang, Y., Liu, G. & Zhu, L. Segmamba: Long-range sequential modeling mamba for 3d medical image segmentation. In International conference on medical image computing and computer-assisted intervention, 578–588 (Springer, 2024).
Yue, Y. & Li, Z. Medmamba: Vision mamba for medical image classification. arXiv preprint arXiv:2403.03849 (2024).
Ma, J., Li, F. & Wang, B. U-mamba: Enhancing long-range dependency for biomedical image segmentation. arXiv preprint arXiv:2401.04722 (2024).
Ma, J. et al. Segment anything in medical images. Nat. Commun. 15, 654 (2024).
Wu, J. et al. Medsegdiff: Medical image segmentation with diffusion probabilistic model. In Medical Imaging with Deep Learning, 1623–1639 (PMLR, 2024).
Kirillov, A. et al. Segment anything. In Proceedings of the IEEE/CVF international conference on computer vision, 4015–4026 (2023).
Sun, J. et al. Medical image analysis using improved sam-med2d: segmentation and classification perspectives. BMC Med. Imaging 24, 241 (2024).
Xiao, X. et al. Describe anything in medical images. arXiv preprint arXiv:2505.05804 (2025).
Xiao, X. et al. Visual instance-aware prompt tuning. In Proceedings of the 33rd ACM International Conference on Multimedia, 2880–2889 (2025).
Xiao, X. et al. Prompt-based adaptation in large-scale vision models: A survey. arXiv preprint arXiv:2510.13219 (2025).
Soler, L. et al. 3d image reconstruction for comparison of algorithm database. https://www.ircad.fr/research/data-sets/liver-segmentation-3d-ircadb-0113 (2010).
Du, Y., Bai, F., Huang, T. & Zhao, B. Segvol: Universal and interactive volumetric medical image segmentation. Adv. Neural Inf. Process. Syst. 37, 110746–110783 (2024).
Kavur, A. E. et al. Chaos challenge-combined (ct-mr) healthy abdominal organ segmentation. Med. Image Anal. 69, 101950 (2021).
Acknowledgements
This work was supported by grants from the Inner Mongolia Autonomous Region Natural Science Foundation Project Youth Fund (2024QN08001); 2021 Inner Mongolia Autonomous Region’s own institutions’ introduction of high-level talent research support projects; 2022 Inner Mongolia Autonomous Region Talent Development Fund (22nd Batch) High-level Talent Individual Project Funding Project; Hohhot Science and Technology Bureau Applied Research and Development Funds (2023–SHE-14); Science and Technology Project of High-level Clinical Specialty Construction in Hohhot Public Hospital of Inner Mongolia Autonomous Region Health Commission (2023SGGZ029); Guangzhou Basic and Applied Basic Research Funds (grant no. 202201011123); and the Plan on enhancing scientific research in Guangzhou Medical University.
Author information
Authors and Affiliations
Contributions
L.X., H.-Y.C., and Y.-W.C. contributed equally to this work, having full access to all study data and assuming responsibility for the integrity and accuracy of the analyses (Validation, Formal analysis). L.X., C.-Q.G. and J.-Z.Z. conceptualized the study, designed the methodology, and participated in securing research funding (Conceptualization, Methodology, Funding acquisition).H.-Y.C., W.-H.Z., and H.J. carried out data acquisition, curation, and investigation (Investigation, Data curation) and provided key resources, instruments, and technical support (Resources, Software). Y.-W.C., S.J., and X.L. drafted the initial manuscript and generated visualizations (Writing - Original Draft, Visualization). H.L., Y.-N.T., and J.-J.Z. supervised the project, coordinated collaborations, and ensured administrative support (Supervision, Project administration). All authors contributed to reviewing and revising the manuscript critically for important intellectual content (Writing - Review & Editing) and approved the final version for submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xia, L., Chen, HY., Cao, YW. et al. Prompt-mamba filtering networks for accurate hepatocellular carcinoma lesion segmentation in abdominal CT. npj Digit. Med. 9, 193 (2026). https://doi.org/10.1038/s41746-026-02371-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-026-02371-5