Abstract
Parkinson’s disease (PD) is the second most common neurodegenerative disease in the world. Although most cases are sporadic and occur later in life, 10–15% of cases are genetic. Loss-of-function mutations in the ring-between-ring E3 ubiquitin ligase parkin, encoded by the PRKN gene, cause autosomal recessive forms of early onset PD. Together with the kinase PINK1, parkin forms a mitochondrial quality control pathway that tags damaged mitochondria for clearance. Under basal conditions, parkin is inhibited and compounds that increase its activity have been proposed as a therapy for PD. Recently, several naturally occurring hyperactive parkin variants were identified, which increased mitophagy in cultured cells. Here, we validate the hyperactivities of these variants in vitro and compare the levels of activity of the variants to those of the wild-type and the well-characterized hyperactive variant, W403A. We also study the effects of mutating the parkin ACT (activating element) on parkin activity in vitro. This work advances our understanding of the pathogenicity of parkin variants and is an important first step in the design of molecules to increase parkin activity.
Similar content being viewed by others
Introduction
Parkinson’s Disease (PD) is globally the second-most common neurodegenerative disorder after Alzheimer’s Disease1. The major pathophysiological features of PD is the loss of dopaminergic neurons in the substantia nigra pars compacta region of the brain, and the presence of Lewy bodies mainly made of unfolded α-synuclein aggregates, which results in both motor and non-motor symptoms2.
While most cases of PD occur spontaneously and later in life, 10–15% of PD cases occur earlier and are called early onset Parkinson’s disease (EOPD). These are generally genetic and frequently caused by autosomal recessive mutations in the genes PRKN (PARK2) or PINK1 (PARK6)3, which encode the proteins parkin, an E3 ubiquitin ligase, and PTEN-induced kinase 1 (PINK1), respectively. Parkin and PINK1 together regulate a mitochondrial quality control pathway, by which proteins on the surface of damaged mitochondria are ubiquitinated to mark the mitochondria for destruction by mitophagy (autophagy of mitochondria)4. Dysregulation of this pathway has been linked to PD pathogenesis4,5,6,7,8.
Parkin belongs to the ring-between-ring (RBR) group of E3 ligases, which function through a RING/HECT hybrid mechanism9,10,11. RBR E3 ligases are characterized by three zinc-coordinating RING domains: RING1, In-Between-RING (IBR), and RING210,12. In parkin, the RBR domain is preceded by another zinc-coordinating RING0 domain and an N-terminal ubiquitin-like domain (Ubl). There is also a short alpha helix between the IBR and RING2 domains, known as the repressor element of parkin (REP)13 (Fig. 1a). Under basal conditions, parkin is autoinhibited. The RING0 domain blocks the catalytic cysteine (Cys431) on RING2 while the Ubl domain and the REP block the E2-binding site on RING113,14,15.
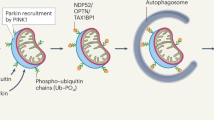
In cells, parkin is activated when PINK1 accumulates on the outer membrane of damaged or depolarized mitochondria5. PINK1 plays a dual role, first phosphorylating ubiquitin molecules to recruit parkin to mitochondria, and secondly phosphorylating parkin to activate its ubiquitin ligase activity5,16,17,18,19. These steps generate Ser65 phosphorylated ubiquitin (pUb) and Ser65 phosphorylated parkin (pParkin). Both are necessary for parkin translocation and activation. pUb recruits parkin to the surface of damaged mitochondria by binding the parkin RING1 domain. This in turn promotes the release of the Ubl domain from RING1 and its phosphorylation by PINK120,21,22,23,24. The phosphorylation of the Ubl also enhances RING1 binding to pUb, preventing parkin dissociation from the mitochondrial surface. Thus, Ubl and pUb compete for binding to the RING1 domain of parkin, operating in a parallel switch mechanism to activate parkin20. Upon phosphorylation, the Ubl domain (now pUbl) binds to the RING0 domain and releases the RING2 domain25,26. Binding of pUbl to RING0 is promoted by a region of parkin between Ubl and RING0 called the activating element of parkin (ACT)26,27. Once parkin is in the active conformation, both the E2-binding site and the catalytic cysteine are accessible and able to catalyze ubiquitination of mitochondrial surface proteins. The added ubiquitins are in turn phosphorylated by PINK1, leading to the recruitment of more parkin molecules as part of a positive feedback loop22,28.
Although parkin mutations have long been associated with PD, the high incidence of sporadic disease has made it difficult to categorize rare sequence variants as pathological or benign. A 2019 study by Yi et al. sought to classify rare parkin missense variants based on their activity in a cellular assay of mitophagy29. Fifty-one missense variants in parkin found in both the general population and PD patients were classified according to their frequency in PD patients and cell-based assays of mitophagy and expression levels. Several non-pathogenic variants were found to exhibit higher levels of mitophagy than wild-type parkin and could rescue mitophagy when combined with common pathogenic parkin variants29. Understanding how these variants act may reveal therapeutic strategies for increasing parkin activity as a treatment for PD.
Here, we compared the activities and phosphorylation rates of five hyperactive variants: P37L, V224A, R234Q, R256C, and M458L with wild-type parkin and a well-characterized hyperactive variant W403A13 (Fig. 1a, b). We conducted autoubiquitination, ubiquitin vinyl-sulfone (UbVS), kinase assays, isothermal calorimetry (ITC), and differential scanning fluorimetry experiments. Three of the variants showed increased autoubiquitination activity. We also assessed the importance of the ACT region and observed that deletion of the entire ACT region decreased ubiquitination activity in vitro but the effect was small compared to the PD mutation K161N. Notably, loss of Arg104 which had been proposed to cause disease26 had no effect on parkin autoubiquitination.
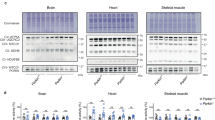
a Domain organization of parkin and mutations studied. b Positions of the mutations in the autoinhibited parkin structure (PDB 5C1Z). c Time-course of the autoubiquitination activity of unphosphorylated GST-parkin. The fraction of mono-ubiquitinated parkin was quantified and shown under the gel. d Quantification of percentage increase in monoubiquitinated GST-parkin over time and S.E.M. from three replicate experiments (n = 3). The activity levels of the controls (WT and W403A) are shown only on the first graph for clarity. e Statistical comparison of p-values for the increase in monoubiquitinated GST-parkin. *p < 0.05; NS, not significant.
Methods
Mutagenesis of parkin constructs
Single-point mutations were performed on full-length wild-type human parkin28 cloned into BamHI-XhoI sites of the pGEX-6P-1 vector (Cytiva) using the QuickChange Lightning Site-Directed Mutagenesis Kit (Agilent). The primers used are listed in Supplementary Table S2.
Expression and purification
Protein expression of GST-tagged human parkin and Ubl domains, GST-tagged Tribolium castaneum (Tc) PINK1, human His6-E1, human His6-E2 (UbcH7), and GST-tagged human ubiquitin (Ub) S65A was performed in BL21 (DE3) E. coli cells as described previously28.
In brief, GST-tagged proteins were purified using glutathione-Sepharose resin (Cytiva) and eluted with 20 mM reduced glutathione. For all experiments except the autoubiquitination assay, the GST tags of GST-parkin and GST-Ub S65A were cleaved with 3C protease overnight at 4 °C. His6-tagged proteins were purified using TALON Co2+ affinity resin (Takara Bio) and eluted with 500 mM imidazole. All proteins were further purified by size exclusion chromatography (SEC) (Cytiva) in SEC buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM TCEP). Purified proteins were verified by SDS-PAGE analysis (Supplementary Fig. S1). Protein concentrations were determined using UV absorbance.
Preparation of phosphorylated proteins
Ubiquitin was phosphorylated by incubation of 350 μM of commercial bovine ubiquitin (Sigma-Aldrich) with 11.6 μM GST-tagged PINK1 in a reaction buffer containing 5 mM ATP, 10 mM MgCl2, 50 mM Tris-HCl pH 8.0, 100 mM NaCl, 0.1% CHAPS and 1 mM DTT, at 30 °C for 3 h. The reaction was verified on a Phos-tag (ApexBio CA) Tris/tricine SDS-PAGE gel and dialyzed against 20 mM Tris-HCl pH 8.7 overnight at 4 °C (Supplementary Fig. S2). The phosphorylated ubiquitin (pUb) was purified as described previously by Wauer et al.24 and buffer-exchanged into SEC buffer. To generate phosphorylated parkin (pParkin) for activity assays, 5 μM parkin in complex with pUb (pParkin:pUb) was incubated with 0.2 μM PINK1 in SEC buffer supplemented with 5 mM ATP, 10 mM MgCl2, and 10 μM pUb at 30 °C for 3 h28.
Autoubiquitination assay of hyperactive parkin variants
Autoubiquitination assays were performed at room temperature for 1 h by incubating 3 μM GST-parkin or GST-pParkin:pUb with 50 nM His6-E1, 2 μM UbcH7, and 100 μM ubiquitin S65A in 15 mM Tris-HCl pH 8.0, 85 mM NaCl, 5 mM ATP, and 10 mM MgCl2. Reactions were stopped by the addition of 5× SDS-PAGE loading buffer containing 100 mM DTT and visualized on Tris/glycine SDS-PAGE gels. Reaction mixtures lacking E1 were used as negative controls. Experiments for each sample were performed in triplicate.
Ubiquitin vinyl sulfone assay
2 μM parkin or pParkin:pUb was incubated with 10 μM of ubiquitin-vinyl sulfone (UbVS) (R&D Systems) in SEC buffer at room temperature for 30 min. Reactions were stopped by the addition of 5× SDS-PAGE loading buffer and visualized on SDS-PAGE gels. Negative controls were made without the addition of UbVS. Experiments for each sample were performed in triplicate.
Parkin phosphorylation
Kinase assays of full-length parkin were performed at 30 °C by incubating 3 μM parkin and 0.1 μM PINK1 in SEC buffer supplemented with 5 mM ATP and 10 mM MgCl2, either in the presence or absence of 5 μM pUb and visualized on Phos-tag Tris/glycine SDS-PAGE gels stained by Coomassie blue. For kinase assays of the isolated Ubl domain, 50 μM Ubl (residues 1–76) and 1 μM PINK1 were incubated in SEC buffer supplemented with 5 mM ATP and 10 mM MgCl2 at 30 °C, and visualized on Phos-tag Tris/tricine SDS-PAGE gels. The time-zero (T0) reaction controls lacked ATP.
Isothermal titration calorimetry (ITC)
ITC measurements of pUb binding to wild-type and V224A parkin were carried out on a VP-ITC titration calorimeter (Microcal) at 20 °C in SEC buffer. Results were analyzed using ORIGIN v7 software (MicroCal) and fit to a single binding site model.
Differential scanning fluorimetry
20 μl samples containing 5 μM wild-type or P37L Ubl domain (residues 1–76) and Protein Thermal Shift™ dye (Applied Biosystems, Thermo Fisher) were prepared in a 96-well plate, in SEC buffer or the provided commercial buffer. Samples were heated from 25 to 99 °C at a rate of 1 °C/min, and fluorescence signals were monitored by the StepOne Plus quantitative real-time PCR system (Life Technologies). Data were analyzed using Thermal Shift software (Life Technologies). The maximum change of fluorescence with respect to temperature was used to determine the melting temperature (Tm) of the proteins. Experiments for each sample were performed in triplicate.
Autoubiquitination assay of parkin ACT variants
Autoubiquitination assays were performed at room temperature for 5 min by incubating 2.7 μM pParkin:pUb with 50 nM His6-E1, 60 μM UbcH7, and 50 μM ubiquitin and 50 mM MgCl2. Reactions were stopped by the addition of 5× SDS-PAGE loading buffer containing 100 mM DTT and visualized on Tris/glycine SDS-PAGE gels. Reaction mixtures lacking ATP were used as negative controls. Experiments were performed in duplicate.
Analysis of SDS-PAGE gels
SDS-PAGE gels were stained with Coomassie Blue, imaged using ChemiDoc imaging system (Bio-Rad) and quantified using ImageJ30.
Statistical analysis
Two-tailed, unpaired t-tests were performed on the data from the autoubiquitination assays, UbVS assays and kinase assays of the full-length proteins. P-values less than 0.05 were considered to indicate statistically significant differences, while p-values above 0.05 were considered to indicate statistically insignificant differences. This was followed with a Bonferroni correction with an adjusted significance level of 0.05/6 = 0.0083 (and 0.05/9 = 0.0056 for the ACT experiment) to reduce the incidence of false positives.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Activity of hyperactive variants
We carried out autoubiquitination assays of hyperactive GST-parkin variants to study their activity. In the absence of canonical substrates in vitro, E3 ligases can autoubiquitinate themselves, which can be monitored by a loss of the unmodified ligase and the formation of higher molecular weight polyubiquitinated conjugates. In the absence of phosphorylation, little ligase activity was observed with wild-type parkin: only a faint band of mono-ubiquitinated parkin was visible (Fig. 1c). Parkin W403A was used as a positive control. The variant, which lies in the REP region of parkin, interferes with REP binding to RING1, increasing the basal level of ligase activity13. Three of the variants, R234Q, R256C, and M458L, showed increased autoubiquitination activity with 5 to 7% mono-ubiquitination after 1 h. While much less than the level of W403A (24.5% mono-ubiquitination), the activity was significantly higher than the 1 to 2% autoubiquitination observed with wild-type parkin (Fig. 1d).
When phosphorylated and pUb bound, parkin becomes highly active and heavily ubiquitinated (Fig. 2). We quantified the amount of activity based on the decrease in unmodified parkin since the amount of mono-ubiquitinated parkin decreases at later time points due to formation of polyubiquitinated species. All the variants, including W403A, showed activity similar to wild-type parkin. Unmodified protein was decreased by roughly 50% after 5 min followed, at later time points, by the formation of a broad smear of high molecular weight, polyubiquitinated protein.
a Time-course of the autoubiquitination activity of phosphorylated GST-parkin with pUb bound. The decrease of unmodified parkin band present at different times of reaction is shown below the gel. b Quantification of the decrease in unmodified phosphorylated GST-parkin:pUb over time and S.E.M. from three replicate experiments (n = 3). The activity levels of the controls (pWT and pW403A) are shown only on the last graph for clarity.
As an alternative to autoubiquitination, we turned to a ubiquitin vinyl sulfone (UbVS) assay to characterize the hyperactive parkin variants. UbVS is a C-terminally modified ubiquitin derivative that chemically cross-links with the parkin active site cysteine (Cys431) and can be used as a measure of parkin activation31. In its inactive state, parkin is in a closed conformation and the cysteine is unreactive. Upon activation, the catalytic domain detaches from the rest of the protein, exposing the cysteine. Appearance of the cross-linked parkin-Ub adduct can be used to measure this aspect of parkin activation independently of other aspects such as E2 enzyme binding or aminolysis14,22.
SDS-PAGE gels were used to monitor appearance of the parkin-Ub adduct in the hyperactive variants compared to wild-type and the W403A variant (Fig. 3). As before, unphosphorylated W403A variant showed the most activity with 6% of the total protein forming cross-links. Wild-type parkin and the other variants showed very little cross-linked protein indicating that they were mostly in a closed conformation in the absence of activation. UbVS assays with phosphorylated parkin in complex with pUb showed a large increase in cross-linking for both wild-type and variants. The W403A variant protein showed the most activity followed by four of the five hyperactive variants. Replicate experiments showed the same pattern of activity but with large variations in the extent of cross-linking.
a UbVS assay of unphosphorylated parkin. b UbVS assay of phosphorylated parkin with pUb bound. The percentage of cross-linked parkin-Ub over the total parkin is shown below the gels. c Quantification of the percentage of parkin-Ub adduct observed after 30 min of reaction with UbVS for both unphosphorylated parkin and phosphorylated parkin:pUb and S.E.M. from three replicate experiments (n = 3).
Phosphorylation of hyperactive parkin variants by PINK1
As enhanced phosphorylation could account for parkin hyperactivity in cells, we examined the ability of the hyperactive variants to act as substrates in a PINK1 kinase assay (Fig. 4). Assays were carried out in the absence and presence of pUb and a 30-fold excess of parkin over PINK1. In the absence of pUb, roughly 50% of parkin was phosphorylated after 60 min (Fig. 4a, b).
a, b Time course of parkin phosphorylation by PINK1 in the absence of pUb. c, d Time course of parkin phosphorylation by PINK1 in the presence of pUb. Representative phos-tag gels are shown in panels (a) and (c) along with the percentage of phosphorylated parkin measured by densitometry. Quantification of percentage increase in phosphorylated parkin over time and S.E.M. from three replicate experiments are shown in panels (b) and (d) (n = 3). The phosphorylation levels of the controls (WT and W403A) are shown only on the last graphs for clarity.
The binding of pUb to parkin enhances its phosphorylation by PINK1. pUb decreases the affinity of the RING1 domain for the Ubl domain and leads to its partial release21,32. Our assays with the hyperactive variants reproduced this observation. All the proteins reached close to 100% phosphorylation after 45 min (Fig. 4c, d). Again, no significant differences were observed between wild-type parkin and the parkin variants.
Isothermal titration calorimetry of hyperactive V224A variant
Among the hyperactive variants reported in Yi et al., V224A showed the highest mitophagic activity in cells and was the only hyperactive variant not found in PD patients29. Notably, it partially rescued the mitophagy defects of some pathogenic mutations when combined in the same protein29. Since Val224 is located in the pUb high-affinity binding pocket, the mutation could directly affect the binding of pUb to parkin and contribute to parkin activation. Accordingly, ITC experiments were performed to compare the binding affinities of pUb to wild-type and V224A parkin (Fig. 5). A KD of 200 nM was determined for the binding of pUb to wild-type parkin, while a KD of 270 nM was determined for the binding of pUb to V224A parkin. These values are not significantly different, indicating that the two constructs bind pUb with similar affinities.
The data shown here was obtained from a single experiment (n = 1) with a statistical analysis of the fitting. Thermodynamic parameters from the fittings are shown in the lower panel.
Thermal stability of P37L Ubl domain
The P37L variant in the parkin Ubl domain has been reported to be possibly pathogenic and associated with PD29,33,34. Multiple missense mutations in the parkin Ubl domain have been found to unfold the Ubl domain or decrease its stability33. To test if the stability of the Ubl domain is altered in the P37L variant, we carried out a differential scanning fluorimetry (thermal shift) assay on the isolated wild-type and P37L Ubl domains. The average Tm was found to be 58.1 °C for the wild-type Ubl and 58.4 °C for P37L Ubl (Fig. 6a). The absence of a significant difference suggests that the P37L mutation does not disrupt the Ubl structure.
a Differential scanning fluorimetry traces show unfolding at 58.1 ± 0.2 °C for the WT domain and 58.4 ± 0.2 °C for the P37L variant. Three replicate experiments were performed (n = 3). b Time course of PINK1 phosphorylation of the wild-type and mutated Ubl domains analyzed on a Coomassie-stained phos-tag gel. A single experiment was performed for this test (n = 1).
Phosphorylation of P37L Ubl domain by PINK1
We next asked if the P37L mutation would affect the rate of phosphorylation of the isolated Ubl domain. A kinase assay performed on the Ubl domain alone revealed no difference in the phosphorylation rates of wild-type and P37L Ubl domain, mirroring results with the full-length proteins. Both wild-type and mutated Ubl reached 100% phosphorylation within 5 min (Fig. 6b). The much faster phosphorylation of the isolated Ubl domain compared to full-length parkin reflects the inaccessibility of the phosphorylation site in the compact, autoinhibited conformation of parkin.
Autoubiquitination assay of ACT variants
A mutation in a region following the Ubl domain, termed the activating element (ACT), has been proposed to cause early onset PD. To test how the ACT region affects parkin activity, six different point mutants, R104A, L102K, T103A, V105K, D106A, and L107K, and a truncation lacking the entire ACT region (Δ101-109) were generated (Fig. 7). The variants R104A and Δ101-109 were generated to replicate the experiments performed by Gladkova et al.26 with additional variants designed to test the importance of specific ACT residues. The proteins where phosphorylated (Supplementary Fig. S3) and the autoubiquitination activity of the variants was compared to that of wild-type, and K161N, a PD mutation that disrupts the RING0 phosphate binding site and diminishes parkin activity25. The K161N variant is classified as definitely pathogenic by MDSgene but is only likely pathogenic in clinical settings according to the ACMG guidelines. Parkin activity was quantified by the decrease in unmodified parkin. While none of the variants showed activity levels as low as unphosphorylated wild-type parkin (4% at 5 min) or the K161N mutant (32%), several showed markedly less activity than phosphorylated wild-type parkin. When phosphorylated, the variants L107K andΔ101-109 showed roughly 50% of wild-type activity. Variants L102K, V105K showed slight impairments, while R104A, T103A, and D106A showed activity indistinguishable from phosphorylated wild-type parkin. An independent experiment with the R104A and truncation ∆101-109 mutants showed identical results (Supplementary Fig. S4).
a Time-course of the autoubiquitination activity of phosphorylated parkin with pUb bound. b Sequence alignment of parkin ACT elements from different species. c Quantification of the decrease of unmodified phosphorylated parkin over time from two replicate experiments (n = 2). The activity levels of the controls (WT, pWT, and pK161N) are shown only on the first graph for clarity.
Discussion
Most cases of genetic autosomal recessive PD involve loss-of-function mutations in parkin and PINK1. These cases cause mostly early onset forms of PD13,35. Many parkin mutations found in PD patients decrease parkin activity29,36, so increasing or derepressing parkin activity could delay the age of onset or rescue the effects of the loss-of-function mutations. Understanding the effects of mutations that enhance parkin activity offers insight into structural mechanisms of parkin activation and could be used to design small molecules to mimic the effects of hyperactive variants. Current efforts in drug discovery have shown that small molecules can indeed increase parkin activity in vitro37.
Parkin is basally autoinhibited and activated in a multistep process that involves domain and structural rearrangements. Disruption of inhibitory contacts by synthetic mutations, F146A, W403A, C457S, and F463A, greatly increases the autoubiquitination activity of unphosphorylated parkin13,38. Of the naturally occurring variants studied here, none activated parkin to a similar degree, but several showed enhanced basal activity of unphosphorylated parkin.
The P37L variant, which is located in parkin Ubl domain, was previously reported by Aguirre et al. to slightly increase parkin activity when phosphorylated34 but decrease activity in cells. Interestingly, other studies have associated the variant with PD and classified it as a pathogenic mutant with impaired Ub-charging activity29,33,39. In our in vitro autoubiquitination and UbVS assays, P37L parkin displayed levels of activity similar to wild-type, regardless of whether or not it was phosphorylated (Figs. 1–3). There is also a lack of consensus about the structural consequences of the variant. Using circular dichroism, Safadi et al. reported that the P37L variant increases the thermal stability of the Ubl domain by 6 °C33 while computational modeling predicted a decrease in stability39. Using differential scanning fluorimetry, we found no significant difference in the stability of the domain in the P37L variant, compared to the wild-type (Fig. 6). Studies with the kinase PINK1 detected no effect of the mutation on phosphorylation of the Ubl domain either alone or in full-length parkin (Figs. 4 and 6). The consequences of the P37L mutation thus remain an enigma. In organello approaches, such as those pioneered by Vranas40, are needed to bridge the complexity of cellular assays and the simplicity and defined composition of in vitro assays.
V224A parkin is the most active of the variants reported by Yi et al. and, strikingly, possesses the ability to suppress some inactivating mutations when co-expressed in the same polypeptide29. Surprisingly, we did not observe higher autoubiquitination or UbVS activity with either the unphosphorylated or phosphorylated variant. The mutation is located adjacent to the pUb-binding pocket in RING1, which led us to hypothesize that it enhances mitochondrial recruitment through increased affinity for pUb. ITC experiments did not support this hypothesis although it remains possible that an effect would be seen with phosphorylated parkin. Phosphorylation of the Ubl domain leads to conformational changes that increase the affinity of pUb binding and it is possible that this allosteric coupling is enhanced in the V224A variant. As V224A rescues inactivating missense mutations, understanding the mechanism should be a focus of future studies.
The variants R234Q, R256C, and M458L, which displayed increased activity in vitro, are located on autoinhibitory interfaces so explaining the increase in basal autoubiquitination activity is relatively straightforward (Fig. 1). The first reported carrier of the R234Q variant was a woman with PD who was compound heterozygous with another PD mutation (R33Q)41. R234Q and R256C are both in the RING1 domain and part of the RING1:REP interface. Like the W403A variant, they could weaken REP binding to RING1 and increase the fraction of parkin that escapes autoinhibition. Similarly, M458L is in the RING2 domain and part of the RING0:RING2 interface. A decrease in the affinity of RING2 binding would naturally lead to increased basal ubiquitin ligase activity. It is notable that in the activated (phosphorylated) state, none of the variants including W403A showed significantly higher activity than the wild-type. Similar results were recently reported for synthetic, rationally designed parkin variants42. The eleven activating mutations identified all occurred at the RING0:RING2 and REP:RING1 interfaces and destabilized parkin folding. While we have not examined the structural consequences of the variants studied here, it is noteworthy that the artificial intelligence analysis by AlphaMissense found that the R256C variant is benign while the M458L variant is ambiguous43.
Conclusively classifying rare parkin variants as pathogenic, benign, or protective is challenging due to the small number of occurrences, conflicting reports from different studies, variable ages of onset, and the possibility of other factors affecting PD susceptibility. The five variants characterized here appear in a very small percentage of the population. Details about specific information relating to the variants from across various databases, and a summary of our findings from the autoubiquitination assay are provided in Supplementary Table S1. Animal studies of parkin variants are an additional source of variability due to the differences in the expression of PINK1 protein in different species44,45,46. A recent study by Chen et al.45 reported that while PINK1 plays an important neuroprotective role in primates, it does not appear to do so in pigs. Unlike cultured monkey cells, cultured pig cells lacking PINK1 were still found to maintain normal levels of phosphorylated parkin suggesting the existence of alternative phosphorylation and activation pathways. In flies, parkin overexpression has long been known to genetically rescue PINK1 mutants, pointing to PINK1-independent functions of parkin47. In patient-derived dopaminergic neurons, parkin was shown to be phosphorylated by calmodulin-dependent protein kinase 2 (CaMK2) to trigger its recruitment to synaptic vesicles and facilitate endocytosis48. Mutations in parkin led to defective recycling of vesicles and accumulation of toxic oxidized dopamine. Other studies have also implicated parkin in synaptic vesicle recycling and ubiquitination of synaptic membrane proteins49,50.
The ACT element plays a role in the conformational changes between inactive and active parkin by enhancing the binding of the phosphorylated Ubl domain to RING026,27. To explore this, we tested six point mutants and a deletion spanning the ACT (Fig. 7). Of particular interest is Arg104, which was suggested to cause PD when mutated to tryptophan51. Surprisingly, we found no loss of activity in the R104A variant, in disagreement with Gladkova et al. (2018) who observed a near complete loss of activity26. We did find impaired activity with the L107K and Δ101-109 deletion mutants, confirming the importance of the interactions between the ACT and the hydrophobic binding pocket on RING026. The loss of activity was less than the K161N mutant but potentially large enough to be clinically relevant. As the ACT is unstructured in the inactive conformation of parkin, the mutations in this region are unlikely to affect the level of basal activity. Deep mutational scanning of parkin variant expressed in cultured cells found decreased stability of mutations in the ACT consistent with a prediction of a quality control degron in that region52. Taken together, our results indicates that the ACT element enhances but is not required for parkin activation.
In summary, we observed that mutations that destabilize the autoinhibitory interfaces in parkin increase the parkin basal activity but not activity in its activated, phosphorylated state. Targeting these interfaces could be a possible route to increasing parkin activity in cells. With the demonstration that small molecules can activate parkin in vitro37, the identification of compounds that shift parkin toward its active conformation remains a promising direction for the development of drugs in the fight against PD.
Data availability
All data supporting the findings of this study are available within the article and its Supplementary Information. Unprocessed gel images are presented in Supplementary Figs. S5–S9. Source data for the graphs and charts presented in the main figures are available in the Supplementary Data file. All other data are available from the corresponding author upon reasonable request.
References
Ascherio, A. & Schwarzschild, M. A. The epidemiology of Parkinson’s disease: risk factors and prevention. Lancet Neurol. 15, 1257–1272 (2016).
Balestrino, R. & Schapira, A. H. V. Parkinson disease. Eur. J. Neurol. 27, 27–42 (2020).
Trempe, J. F. & Gehring, K. Mechanisms of mitochondrial quality control mediated by PINK1 and Parkin. J. Mol. Biol. 435, 168090 (2023).
Pickrell, A. M. & Youle, R. J. The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 85, 257–273 (2015).
Matsuda, N. et al. PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J. Cell Biol. 189, 211–221 (2010).
Narendra, D., Tanaka, A., Suen, D.-F. & Youle, R. J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 (2008).
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nat. Rev. Mol. Cell Biol. 12, 9–14 (2011).
Sugiura, A., McLelland, G. L., Fon, E. A. & McBride, H. M. A new pathway for mitochondrial quality control: mitochondrial-derived vesicles. EMBO J. 33, 2142–2156 (2014).
Dove, K. K., Stieglitz, B., Duncan, E. D., Rittinger, K. & Klevit, R. E. Molecular insights into RBR E3 ligase ubiquitin transfer mechanisms. EMBO Rep. 17, 1221–1235 (2016).
Smit, J. J. & Sixma, T. K. RBR E3-ligases at work. EMBO Rep. 15, 142–154 (2014).
Wenzel, D. M., Lissounov, A., Brzovic, P. S. & Klevit, R. E. UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids. Nature 474, 105–108 (2011).
Walden, H. & Rittinger, K. RBR ligase–mediated ubiquitin transfer: a tale with many twists and turns. Nat. Struct. Mol. Biol. 25, 440–445 (2018).
Trempe, J. F. et al. Structure of parkin reveals mechanisms for ubiquitin ligase activation. Science 340, 1451–1455 (2013).
Riley, B. E. et al. Structure and function of Parkin E3 ubiquitin ligase reveals aspects of RING and HECT ligases. Nat. Commun. 4, 1982 (2013).
Wauer, T. & Komander, D. Structure of the human Parkin ligase domain in an autoinhibited state. EMBO J. 32, 2099–2112 (2013).
Kondapalli, C. et al. PINK1 is activated by mitochondrial membrane potential depolarization and stimulates Parkin E3 ligase activity by phosphorylating Serine 65. Open Biol. 2, 120080 (2012).
Shiba-Fukushima, K. et al. PINK1-mediated phosphorylation of the Parkin ubiquitin-like domain primes mitochondrial translocation of Parkin and regulates mitophagy. Sci. Rep. 2, 1002 (2012).
Kane, L. A. et al. PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J. Cell Biol. 205, 143–153 (2014).
Koyano, F. et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 510, 162–166 (2014).
Sauve, V. et al. A Ubl/ubiquitin switch in the activation of Parkin. EMBO J. 34, 2492–2505 (2015).
Kazlauskaite, A. et al. Binding to serine 65-phosphorylated ubiquitin primes Parkin for optimal PINK1-dependent phosphorylation and activation. EMBO Rep. 16, 939–954 (2015).
Ordureau, A. et al. Quantitative proteomics reveal a feedforward mechanism for mitochondrial PARKIN translocation and ubiquitin chain synthesis. Mol. Cell 56, 360–375 (2014).
Kumar, A. et al. Disruption of the autoinhibited state primes the E3 ligase parkin for activation and catalysis. EMBO J. 34, 2506–2521 (2015).
Wauer, T., Simicek, M., Schubert, A. & Komander, D. Mechanism of phospho-ubiquitin-induced PARKIN activation. Nature 524, 370–374 (2015).
Sauvé, V. et al. Mechanism of parkin activation by phosphorylation. Nat. Struct. Mol. Biol. 25, 623–630 (2018).
Gladkova, C., Maslen, S. L., Skehel, J. M. & Komander, D. Mechanism of parkin activation by PINK1. Nature 559, 410–414 (2018).
Fakih, R., Sauvé, V. & Gehring, K. Structure of the second phosphoubiquitin-binding site in parkin. J. Biol. Chem. 298, 102114 (2022).
Sauvé, V. et al. Structural basis for feedforward control in the PINK1/Parkin pathway. EMBO J. 41, e109460 (2022).
Yi, W. et al. The landscape of Parkin variants reveals pathogenic mechanisms and therapeutic targets in Parkinson’s disease. Hum. Mol. Genet 28, 2811–2825 (2019).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Borodovsky, A. et al. A novel active site-directed probe specific for deubiquitylating enzymes reveals proteasome association of USP14. EMBO J. 20, 5187–5196 (2001).
Spratt, D. E., Walden, H. & Shaw, G. S. RBR E3 ubiquitin ligases: new structures, new insights, new questions. Biochem. J. 458, 421–437 (2014).
Safadi, S. S., Barber, K. R. & Shaw, G. S. Impact of autosomal recessive juvenile Parkinson’s disease mutations on the structure and interactions of the parkin ubiquitin-like domain. Biochemistry 50, 2603–2610 (2011).
Aguirre, J. D., Dunkerley, K. M., Lam, R., Rusal, M. & Shaw, G. S. Impact of altered phosphorylation on loss of function of juvenile Parkinsonism-associated genetic variants of the E3 ligase parkin. J. Biol. Chem. 293, 6337–6348 (2018).
Oliveras-Salvá, M., Van Rompuy, A. S., Heeman, B., Van den Haute, C. & Baekelandt, V. Loss-of-function rodent models for parkin and PINK1. J. Parkinsons Dis. 1, 229–251 (2011).
Broadway, B. J. et al. Systematic functional analysis of PINK1 and PRKN coding variants. Cells 11, 2426 (2022).
Shlevkov, E. et al. Discovery of small-molecule positive allosteric modulators of Parkin E3 ligase. iScience 25, 103650 (2022).
Tang, M. Y. et al. Structure-guided mutagenesis reveals a hierarchical mechanism of Parkin activation. Nat. Commun. 8, 14697 (2017).
Fiesel, F. C. et al. Structural and functional impact of Parkinson disease-associated mutations in the E3 ubiquitin ligase parkin. Hum. Mutat. 36, 774–786 (2015).
Vranas, M. et al. Selective localization of Mfn2 near PINK1 enables its preferential ubiquitination by Parkin on mitochondria. Open Biol. 12, 210255 (2022).
Hertz, J. M. et al. Low frequency of Parkin, Tyrosine Hydroxylase, and GTP Cyclohydrolase I gene mutations in a Danish population of early-onset Parkinson’s Disease. Eur. J. Neurol. 13, 385–390 (2006).
Stevens, M. U. et al. Structure-based design and characterization of Parkin-activating mutations. Life Sci. Alliance 6, e202201419 (2023).
Cheng, J. et al. Accurate proteome-wide missense variant effect prediction with AlphaMissense. Science 381, eadg7492 (2023).
Yang, W. et al. CRISPR/Cas9-mediated PINK1 deletion leads to neurodegeneration in rhesus monkeys. Cell Res. 29, 334–336 (2019).
Chen, X. S. et al. Comparative analysis of primate and pig cells reveals primate-specific PINK1 expression and phosphorylation. Zool. Res. 45, 242–252 (2024).
Yang, W. et al. PINK1 kinase dysfunction triggers neurodegeneration in the primate brain without impacting mitochondrial homeostasis. Protein Cell 13, 26–46 (2022).
Whitworth, A. J. & Pallanck, L. J. PINK1/Parkin mitophagy and neurodegeneration-what do we really know in vivo? Curr. Opin. Genet Dev. 44, 47–53 (2017).
Song, P. et al. Parkinson’s disease-linked parkin mutation disrupts recycling of synaptic vesicles in human dopaminergic neurons. Neuron 111, 3775–3788 e3777 (2023).
Trempe, J. F. et al. SH3 domains from a subset of BAR proteins define a Ubl-binding domain and implicate parkin in synaptic ubiquitination. Mol. Cell 36, 1034–1047 (2009).
Cao, M., Milosevic, I., Giovedi, S. & De Camilli, P. Upregulation of Parkin in endophilin mutant mice. J. Neurosci. 34, 16544–16549 (2014).
Chaudhary, S. et al. Parkin mutations in familial and sporadic Parkinson’s disease among Indians. Parkinsonism Relat. Disord. 12, 239–245 (2006).
Clausen, L. et al. A mutational atlas for Parkin proteostasis. Nat. Commun. 15, 1541 (2024).
Acknowledgements
This work was supported by Canadian Institutes of Health Research grant FDN-159903 and Michael J. Fox Foundation grant MJFF-019029.
Author information
Authors and Affiliations
Contributions
V.S. and K.G. designed the study. T.S.H., J.L., and R.F. constructed the models, performed assays, and analyzed the results. V.S. and K.G. supervised the study. T.S.H. and K.G. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Weili Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Ibrahim Javed and Benjamin Bessieres. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huq, T.S., Luo, J., Fakih, R. et al. Naturally occurring hyperactive variants of human parkin. Commun Biol 7, 961 (2024). https://doi.org/10.1038/s42003-024-06656-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-024-06656-x