Abstract
XRN1 (5’-3’ exoribonuclease 1) degrades RNA from the 5′ → 3′ direction and utilizes both single- and double-stranded RNA as substrates. XRN1 plays a critical role in mRNA turnover as well as regulating the cellular response to viral infection. XRN1 also protects the cell by preventing endogenous double-stranded RNA accumulation. XRN1 was identified as a putative vulnerability in a subset of cancer cell lines through analysis of publicly available CRISPR data. The role of XRN1 was explored using a set of non-small cell lung cancer cell lines with differential predicted XRN1 dependency to validate XRN1 as an oncology target. In predicted sensitive cell lines, XRN1 knockout reduced proliferation, increased apoptosis and activated the pPKR and MDA5 dsRNA sensing pathways. To facilitate drug discovery targeting XRN1, a suite of biochemical and biophysical assays was developed. These assays were used to characterize adenosine-3’,5’-bisphosphate (pAp), a non-selective nuclease inhibitor, as a nanomolar inhibitor of XRN1. Additionally, the crystal structure of human XRN1 was solved with pAp bound, demonstrating distinct interactions for the compound in the XRN1 active site. These studies provide a strong foundation for the discovery of potent, selective inhibitors of XRN1 as a novel approach to cancer therapeutics.
Similar content being viewed by others
Introduction
High levels of cytosolic double-stranded RNAs (dsRNAs) are known to be immunogenic and thus activate innate dsRNA sensing pathways. Canonically, these pathways are triggered upon viral infection of a human cell, resulting in an antiviral response that includes increases in interferon (IFN), stimulation of downstream interferon signaling pathways and, in some cases, apoptosis and cell death1,2. Endogenous dsRNA can also activate these pathways, a process called “viral mimicry”3,4. Engagement of these pathways to increase dsRNA burden and induce apoptosis as a potential treatment for cancer is an area of active research; efforts to date have focused on epigenetic regulation (e.g. DNA4,5 and protein6,7 (including histone) methylation), RNA splicing8 and RNA editing9. In addition, decreasing degradation of immunogenic dsRNA, thus increasing dsRNA burden within the cell, may also be a viable path to engage the desired signaling pathways.
XRN1 is a 5’ to 3’ processive exoribonuclease10,11 and can degrade both single- and double-stranded RNA substrates, making XRN1 a critical component of multiple essential cellular functions. For example, XRN1 is directly involved in mRNA turnover; mRNA substrates of XRN1 are deadenylated and decapped to expose a 5’ monophosphorylated overhang conducive to XRN1-mediated degradation12,13. XRN1 can also degrade viral RNA, and multiple viruses have developed novel, XRN1-resistant RNA structures and motifs in order to avoid degradation by the exonuclease14,15. Therefore, XRN1 is a key component of cellular viral defense pathways.
Recently, exploitation of viral mimicry pathways for oncology has been an area of active research, and XRN1 in particular has been implicated as a potential target in this approach16,17. Specifically, for cancer cell lines with high levels of interferon signaling, loss of XRN1 results in an increase of endogenous dsRNA within the cell which activates the dsRNA sensor protein kinase R (PKR) through dimerization and autophosphorylation of the enzyme (pPKR). pPKR, in turn, initiates multiple signaling pathways, including apoptotic pathways that result in cell death18. The MDA5 dsRNA sensor is also activated by increased endogenous dsRNA; a conformational change in MDA5 upon dsRNA binding leads to activation of mitochondrial antiviral signal protein (MAVS), an adapter protein, and expression of Type I interferon genes19. Engagement of the pPKR and MDA5 pathways thus leads to cell death and proinflammatory signaling, both of which are critical in the setting of viral infection. In cancer cells, the removal of XRN1-mediated maintenance of dsRNA levels can be co-opted to increase dsRNA above the threshold that activates the dsRNA sensors pPKR and MDA5, triggering downstream signaling pathways and inducing cell death.
Demonstration of the dependence of XRN1 sensitivity on interferon signaling has been previously documented. Cells lacking elevated Type I interferon signaling have been shown to be indifferent to XRN1 knockout but are rendered XRN1-dependent with treatment of agents known to increase levels of dsRNA16 or by treating cells with directly with IFNβ17. Conversely, cells dependent on XRN1 activity were made XRN1-independent by treatment with a JAK1/2 inhibitor known to decrease cellular PKR levels17. Thus, XRN1 is a viable oncology target for cancers with high levels of interferon signaling. Intriguingly, XRN1 was also identified as a potential cancer immunotherapy target through an in vivo CRISPR screen20 as well as in directed studies with the target21 with silencing of the gene resulting in tumor growth suppression in immunocompetent but not immunodeficient mice.
In this report, data further supporting the role of XRN1 as a therapeutic target in oncology are presented. Non-small cell lung cancer (NSCLC) cell lines predicted to be either dependent on or insensitive to XRN1 based on publicly available screening data were utilized to explore effects of XRN1 knockout on proliferation and known cell signaling pathways; these results confirmed previous reports validating XRN1 as a viable target for further exploration. Hence, biochemical and biophysical assays were developed to enable the identification and characterization of small molecule inhibitors of XRN1. These assays were then utilized to characterize a known nucleoside-based inhibitor of XRN1, adenosine 3’,5’-bisphosphate (pAp). Surprisingly, pAp was found to be a nanomolar inhibitor of XRN1 and a nanomolar binder to both XRN1 and XRN2. Finally, the first published crystal structure of human XRN1 was solved in complex with pAp, showing important differences in pAp binding interactions to XRN1 when compared to another bimetal nuclease enzyme. The suite of assays described here and the structural insights gleaned from the co-crystal structure of the XRN1:pAp complex provide a solid foundation for further chemical biology and drug discovery efforts aimed at the identification and development of small molecule inhibitors of XRN1 for the treatment of cancer.
Results
Validation of XRN1 as a cancer target in non-small cell lung cancer
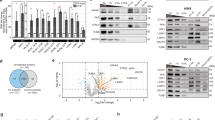
A subset of cancer cell lines is predicted to be dependent on XRN1 for proliferation, as assessed by publicly available pooled CRISPR screen data22 (https://depmap.org/portal; Fig. 1a); however, until recently16,17,21, XRN1 had not been explored as a prospective cancer target. To more fully assess the potential for XRN1 as a target of interest in oncology, XRN1 CRISPR knockout was performed in non-small cell lung cancer (NSCLC) cell lines. NCI-H1650 and NCI-H1703 cells, shown to be sensitive to XRN1 loss in pooled screens (Supplementary Table 1), exhibited decreased colony formation upon XRN1 CRISPR knockout (KO) when compared to cells transduced with a non-targeting control sgRNA (Fig. 1b). In contrast, cells predicted to be insensitive to XRN1 KO, NCI-H838 and NCI-H1944, proliferated normally upon XRN1 KO. A positive control sgRNA which induced KO of POL2RL was effective in reducing proliferation across all four NSCLC cell lines. Proliferation was also assessed by Cell TiterGlo at 12 days post sgRNA transduction (Fig. 1c). Both NCI-H1650 and NCI-H1703 cells were dependent on XRN1, consistent with the colony formation results, while NCI-H1944 and NCI-H838 cell proliferation was equivalent to a non-targeting control. Apoptosis was assessed in a pair of NSCLC lines using a Caspase 3/7 luminescent assay (Fig. 1d); XRN1-dependent NCI-H1703 but not NCI-H838 cells exhibited robust apoptosis upon XRN1 KO. Based on these results, XRN1 is essential for proliferation in a subset of NSCLC cell lines which validates XRN1 as a potential oncology target.
a Pooled screen (Depmap 23Q4) results for XRN1 across 1100 cancer cell lines. b Colony formation in sensitive (red) and insensitive (blue) NSCLC cell lines after CRISPR knockout (KO) of XRN1 or a positive control. c Normalized proliferation of NSCLC cell lines after 12 days of CRISPR KO of XRN1 (gray bars) or positive control POL2RL (white bars). Error bars represent standard deviation of 3 replicates. d Apoptosis (normalized to uninfected control cells) as assessed by a caspase 3/7 luminescent assay in NCI-H1703 or NCI-H838 cell lines after 7 days of XRN1 CRISPR knockout (black bars: Non-targeting control; gray bars: XRN1 knockout). Error bars represent standard deviation of 3 replicates.
As previously described, experiments in colorectal, breast and lung cell lines have demonstrated that XRN1 is important for controlling endogenous cytosolic dsRNA levels and that loss of XRN1 leads to an elevation of dsRNA burden that triggers the PKR and MDA5 innate immune pathways16,17. PKR activation leads to autonomous tumor cell death via downstream phosphorylation of eIF2α, while MDA5-mediated induction of Type I interferons can induce an anti-tumor immune response. In these studies, XRN1 KO induced a robust upregulation of pPKR, consistent with activation of dsRNA sensing in both XRN1-dependent NSCLC cell lines tested (Fig. 2a)18. Among the tested cell lines, pPKR induction is restricted to NCI-H1650 and NCI-H1703 XRN1-dependent cell lines and is not observed in NCI-H1838 and NCI-H1944 cell lines, consistent with the lack of anti-proliferative activity upon XRN1 KO in these cell lines, providing additional validation of the hypothesis for XRN1 importance in select NSCLC cells. Interestingly, total PKR levels are variable across the panel of lung cell lines tested and total PKR levels do not appear to correlate to XRN1 dependence. In addition to pPKR induction, Type I interferon gene expression was robustly upregulated in NCI-H1650 and NCI-H1703 cell lines, consistent with activation of the MDA5 pathway (Fig. 2b). In contrast, changes in IFNα1 and IFNβ1 gene expression were not detected in the insensitive NCI-H838 and NCI-H1944 cell lines. Together these data validate the hypothesis that targeting XRN1 has the potential for monotherapy activity in a subset of solid tumor cell lines via activation of dsRNA sensing leading to cell death.
a XRN1, pPKR, total PKR and loading control western blot for NCI-H1650, NCI-H1703, NCI-H838, and NCI-H1944 cell lines transduced with non-targeting (NT), XRN1, or POL2RL sgRNA for 7 days. b RT-qPCR for Type I interferon genes, IFNA1 and IFNB1, 7 days post CRISPR transduction (black bars: Non-targeting control; gray bars: XRN1 knockout). Error bars represent standard deviation of three replicates. c Proliferation of NCI-H838 cells (black bars: Non-targeting control; gray bars: XRN1 knockout) following 96 hours of stimulation with exogenous IFNβ, as assessed by a luminescent assay. Error bars represent standard deviation of three replicates. d Model for XRN1 in cancers with elevated dsRNA burden and rationale for monotherapy and combination with checkpoint inhibitors.
Because XRN1 dependent cell lines were shown to have intrinsically elevated Type I interferon signaling16,17, it was hypothesized that addition of exogenous IFNβ to the NCI-H838 cell line would render this previously insensitive cell line dependent on XRN1. NCI-H838 cells transduced with XRN1 sgRNA or non-targeting (NT) controls were stimulated with IFNβ for 96 hours and assessed for proliferation; the cells transduced with XRN1 sgRNA exhibited decreased proliferation compared to IFNβ-stimulated NT controls (Fig. 2c). Phosphorylated eIF2α (peIF2α), which is downstream of PKR activation, was elevated in XRN1 KO, IFNβ stimulated NCI-H838 cells, consistent with activation of the dsRNA response (Supplementary Fig. 1). These results suggest that elevated exogenous interferon can sensitize otherwise insensitive cells to activation of the dsRNA response in upon XRN1 loss. Because immunotherapeutic agents such as checkpoint inhibitors lead to elevation of cytokine signaling, including Type I interferons, in the tumor microenvironment, these results provide a rationale for the combination of XRN1 inhibition and checkpoint inhibitors in the treatment of cancer23. Thus, these data demonstrate that XRN1 is a validated target for oncology drug discovery, and that XRN1 inhibition has the potential to be effective as both a monotherapy in tumors with elevated dsRNA burden and in combination with immunotherapeutic agents (Fig. 2d).
XRN1 protein construct design
Target validation efforts have indicated that XRN1 catalytic activity is important for antiproliferative activity17; thus, an inhibitor of XRN1 enzymatic activity could be utilized as a therapeutic agent to target XRN1 sensitive tumors. Therefore, efforts to enable high throughput hit-finding and characterization of small molecules modulators of XRN1 enzymatic activity were undertaken. Multiple constructs of full-length human XRN1 were expressed and purified using both mammalian and insect cell expression systems, but the final proteins from these purifications suffered from degradation or cleavage, resulting in protein of insufficient quality for assay development. Utilization of N-and C-terminal affinity tags and western blotting techniques indicated the degradation occurred at the C-terminus, a region of XRN1 shown to interact with proteins directly involved in mRNA decapping activities24 that is predicted to be unstructured25,26. Therefore, a C-terminal truncation construct was designed (human XRN1 1-1191-FLAG, hereafter referred to as hXRN1) that includes the catalytic domain, PAZ (Piwi-Argonaute-Zwille), KOW (Kyrpides-Ouzounis-Woese), winged helix and SH3 (Src homology 3) domains as well as a C-terminal FLAG tag (Supplementary Fig. 2a). Purification of hXRN1 from mammalian cells resulted in protein of high purity and sufficient yield to enable potential drug discovery efforts (Supplementary Fig. 2b). To ensure nuclease activity seen in biochemical assays was due to enzymatic activity from XRN1, a variant protein mutating a key active site metal-chelating residue, Asp 208, to alanine (hereafter referred to as hXRN1-D208A) was also produced (Supplementary Fig. 2c); an analogous variant was previously shown to be an inactivating mutation in yeast XRN127.
Development of XRN1 nuclease activity assay
With high-quality protein in hand, development, optimization and kinetic characterization of an enzymatic assay for XRN1 was initiated. A fluorescence-based assay utilizing a 3’ FAM-linked RNA substrate hybridized to a DNA sequence containing a 5’ TAMRA fluorescent quencher (FQ) had been previously identified as suitable for measuring XRN1 nuclease activity28 and was the basis for assay optimization efforts (Fig. 3a). Monophosphate was selected as the RNA 5’ end group to mimic the cellular substrate. RNA and DNA reagents were annealed to produce a duplex RNA/DNA hybrid with a 13 nucleotide 5’ RNA overhang. An Iowa Black FQ, a “dark quencher” that absorbs light broadly and emits the absorbed energy as heat and not light, was used instead of TAMRA to increase signal to background in the assay. Initial testing of hXRN1 protein with the RNA/DNA hybrid substrate indicated the protein was enzymatically active, and optimization of buffer conditions was performed with a focus on conditions appropriate for high-throughput screening efforts. Given that XRN1 is a metal-dependent nuclease, it was unsurprising that magnesium chloride was necessary for activity. Detergent and protein carrier were desired in the final buffer to minimize potential non-specific binding. An RNase inhibitor was added at concentrations previously shown to be sufficient for RNA stability against non-specific RNA hydrolases29. ETDA was found to fully inhibit the hydrolysis reaction, and 80 mM was utilized as the stop condition for the assay. 20 mM HEPES pH 7.5, 50 mM NaCl, 5 mM MgCl2, 1 mM DTT, 0.01% BSA, 0.004 U/μL RNaseOUT and 0.01% Tween-20 was found to be optimal for hXRN1 enzyme activity.
a Schematic of the XRN1 biochemical assay: enzyme (yellow), RNA (red), DNA (blue), fluorescent label (FAM, orange), Iowa Black fluorescent quencher (FQ, black). Partially created in BioRender (2025) https://BioRender.com/e62n951. b hXRN1, but not hXRN1-D208A, has catalytic activity. Error bars represent standard deviation of 6 replicates. c SPR sensorgrams of ssDNA (5’-AC*TCACTCACTCACCAAAAAAAAAAAAACC-3’; * = phosphorothioate) binding to hXRN1-Avi indicate dose dependent binding and slow return to baseline. d Data plotted from SPR sensorgrams of ssDNA-hXRN1-Avi binding is not sigmoidal and does not fit a 1:1 binding model. e SPR sensorgrams of ssDNA binding to hXRN1-D208A-Avi is dose dependent binding with rapid dissociation. f ssDNA binding to hXRN1-D208A-Avi fits well to 1:1 binding model (KD = 412 ± 96 nM).
Using the optimized conditions for hXRN1 and the RNA/DNA hybrid substrate, the relative enzymatic activities of the hXRN1 and hXRN1-D208A proteins were tested (Fig. 3b). As expected, the active site mutant, hXRN1-D208A, was inactive at all concentrations tested. Assay linearity for hXRN1 was tested at multiple assay concentrations and timepoints; reactions proceeded linearly at concentrations between 156 and 1250 pM within the timecourse of the experiment. A final protein concentration of 250 pM was selected for a 60-minute assay reaction time. Steady-state enzyme kinetic analysis was then performed by titrating the RNA/DNA hybrid substrate at multiple concentrations and timepoints; initial velocities were calculated from data collected during the linear region of the progress curves. Fitting of the data to the Michalis-Menten equation yielded a substrate KM value of 0.4 ± 0.03 nM for hXRN1 (Supplementary Fig. 3), while kcat was found to be 0.21 ± 0.02 min−1. A final RNA/DNA substrate concentration of 1.5 nM was selected as the value is within 4-fold of the KM values determined for each enzyme, allowing any hit finding activities to be performed under balanced assay conditions.
Development of XRN1 and XRN2 SPR assays
Orthogonal assays are a critical component of a successful hit validation funnel30, and SPR can play a key role in hit validation activities by confirming compound binding potency and reversibility as well as provide information on mechanism of inhibition. Therefore, development of an XRN1 SPR assay was desired. C-terminal Avi-tagged constructs of hXRN1 and hXRN1-D208A (hXRN1-Avi and hXRN1-D208A-Avi, respectively) were produced. hXRN1-Avi and hXRN1-D208A-Avi were captured on a streptavidin-coated chip. To enable robust data generation, assay buffer conditions were modified to minimize non-specific binding and included increasing the salt concentration as well as removal of protein components (BSA and RNaseOUT); Tween-20 and 1% DMSO were well tolerated but not required in the SPR assay buffer.
To validate the assay for future hit validation activities, a single-stranded 30-mer DNA containing a non-hydrolysable phosphorothioate bond between the second and third nucleotides at the 5’ end (phosphorothioate designated with *; 5’-AC* TCA CTC ACT CAC CAA AAA AAA AAA AAC C-3’, hereafter referred to as ssDNA) was synthesized as a positive control reagent. Single-stranded DNA was selected over RNA for these experiments due to potential non-specific degradation of RNA during the experiment by RNase enzymes31. hXRN1 has exonuclease activity for DNA10, therefore, a non-hydrolysable bond was included in the construct to create a stable protein-substrate complex in case of enzymatic activity in situ. Indeed, ssDNA showed concentration-dependent binding to hXRN1-Avi (Fig. 3c), but the subsequent binding curve did not fit well with a 1:1 binding model (Fig. 3d). As XRN1 is a processive enzyme10, it was hypothesized that the ssDNA substrate could bind to hXRN1-Avi and undergo a single catalytic reaction cycle. This would result in a reaction product inhibitor, a 29-mer single stranded DNA with a non-hydrolysable bond, bound to hXRN1-Avi; this process would be expected to result in an atypical SPR binding curve. In contrast, the catalytically dead hXRN1-D208A-Avi binding curves showed typical behavior expected from binding and release of the ssDNA ligand (Fig. 3e), with a KD of 0.412 ± 0.096 µM (Fig. 3f), indicating that enzyme activity during the SPR experiment likely contributes to the shape of the binding curves for the active XRN1 enzyme.
An assay for XRN2, a member of the 5’-3’ exoribonuclease family that is highly similar to XRN1 in overall structure and active site residues32, was required to understand selectivity between these related enzymes; thus, an SPR binding assay for XRN2 was developed. Using the knowledge gained from XRN1 protein engineering and guided by the C. elegans XRN2 crystal structure33, a construct removing predicted unstructured C-terminal residues was designed for XRN2 containing the catalytic domain (human XRN2 1-790-FLAG, hereafter referred to hXRN2; Supplementary Fig. 2a). Purification of hXRN2 from mammalian cells similarly produced high quality protein (Supplementary Fig. 2d) that was catalytically active (Supplementary Fig. 4), ensuring the construct was suitable for selectivity assessments. A C-terminal Avi-tagged construct of hXRN2 (hereafter referred to as hXRN2-Avi) was then produced. To enable efficient selectivity screening, identical buffer conditions to hXRN1-Avi were utilized for the XRN2 SPR assay.
In vitro characterization of pAp
With in vitro biochemical and biophysical assays established, identification of a small molecule tool compound to validate the assay suite in advance of any hit finding activities was desired. Adenosine 3,5 bisphosphate (pAp; Fig. 4a) is a nucleotide analog and was previously identified as an inhibitor of XRN1 enzymatic activity28,34; additionally, it has shown to inhibit multiple nuclease enzymes including XRN2 and DXO34,35. Based on these publications, pAp was expected to be a high μM to mM inhibitor of XRN1 enzymatic activity. Therefore, it was a surprise to find that pAp was a very potent inhibitor of and binder to XRN1 using both enzymatic and SPR assays, with an IC50 value of 36 ± 10 nM for hXRN1 (Fig. 4b) and a KD value of 12 ± 1 nM for hXRN1-Avi (Fig. 4c). Additionally, pAp was equally potent on hXRN2-Avi with a KD value of 14 ± 4 nM (Fig. 4d).
a Chemical structure of pAp. b pAp is an inhibitor of hXRN1 (IC50 = 36 ± 10 nM). Error bars represent standard deviation of 6 replicates. pAp binds to c hXRN1-Avi (KD = 12 ± 1 nM) and d hXRN2-Avi (KD = 14 ± 4 nM). e pAp does not bind to the metal-binding site mutant hXRN1-D208A-Avi. f pAp IC50 curves shift right with increasing amounts of RNA substrate. Error bars represent standard deviation of 2 replicates. g The near-linear increases in pAp IC50 values as RNA concentration is increased (α = 23) is indicative of a mixed-competitive mechanism with the oligonucleotide substrate.
The crystal structure of pAp bound to the nuclease DXO (PDB code 6AIX) previously revealed the nucleotide analog bound to the bimetal active site of the enzyme35. If pAp bound to XRN1 in the same manner, the active site mutant D208A would be expected to ablate binding affinity. Indeed, no binding was observed for pAp to hXRN1-D208A-Avi by SPR up to 0.2 μM (Fig. 4e), supporting the hypothesis of pAp binding to the bimetal active site of XRN1. Additional experiments were performed to assess the inhibition modality of pAp for XRN1 by measuring the IC50 of pAp at different substrate concentrations (Fig. 4f ) as a function of the concentration of substrate over KM ([S]/KM)36,37. The resulting curvilinear ascending plot (α = 23; Fig. 4g) is indicative of mixed-competitive inhibition. This result is consistent with a compound that competes with oligonucleotide substrate for binding to the bimetal binding site of XRN1 but does not rule out other substrate-competitive mechanisms.
Crystal structure of human XRN1 catalytic domain bound to pAp
A crystal structure of pAp bound to XRN1 would fully validate the compound as a specific binder to XRN1 and unambiguously define the nucleotide inhibitor binding site. Extensive crystallization screening was performed with the hXRN1 construct in the presence and absence of pAp, but no diffraction quality crystals were obtained. Crystal structures of XRN2 from C. elegans33 and C. thermophilum38 have been solved that contain only the catalytic domain of XRN2; therefore an XRN1 crystallography construct was generated containing only the catalytic domain of XRN1 (human XRN1 aa 1-653-FLAG, hereafter referred to as hXRN1x, Supplementary Fig. 2a, e). As the construct was enzymatically active and pAp was a potent binder and inhibitor of the construct (Supplementary Fig. 5), crystallization trials were initiated. Although extensive screening with both apo and pAp-bound hXRN1x was performed, hXRN1x crystals only formed in the presence of pAp.
A 2.1 Å crystal structure of hXRN1x bound to pAp was solved revealing the nucleotide analog bound to the bimetal binding site (Fig. 5a), consistent with the biochemical and biophysical data generated for this tool compound. The asymmetric unit of the crystal contained two hXRN1x molecules in the same overall conformational state with an RMSD value of 0.545 Å2. However, unambiguous electron density corresponding to two magnesium ions and pAp was only observed in the active site of molecule A (Figs. 5b, c) and not in molecule B even though the protein buffer contained 5 mM magnesium ion and saturating concentrations (1 mM) of pAp. Therefore, structural analysis focused on molecule A.
a Ribbon representation of hXRN1x (cyan) – pAp (blue stick) structure. Magnesium ions represented by gray spheres. b 2Fo-Fc (blue mesh, 1.0 σ) and c omit Fo-Fc (green/red mesh, +/− 3.0 σ) electron density maps of pAp unambiguously confirm the presence of the ligand in the XRN1 active site. d A close view of the XRN1-pAp binding site including XRN1 residues that interact directly with phosphate moieties of the ligand. Magenta dashes = 5’ phosphate interactions; yellow dashes = 3’ phosphate “up” orientation interactions; orange dashes = 3’ phosphate “down” orientation interactions; protein view is similar to (b, c). e Superposition of hXRN1x-pAp and Drosophila XRN1-DNA (PDB code 2Y35; gray) structures. Base rings are aligned while shifts are seen for ribose and phosphate groups. f Superposition of hXRN1x-pAp and C. elegans XRN2 (PDB code 7OPK; green) shows good overall structural similarity. g Residue differences (stick representation) in the hXRN1x and XRN2 active sites do not directly interact with pAP. h Comparison of hXRN1x-pAp (3’ “up” confirmation) and mDXO-pAp (PDB code 6AIX; pink) shows conservation of pAp orientation despite lack of structural similarity between proteins.
In molecule A, the magnesium ions are bridged by Glu 176 and a water molecule, with the first magnesium further coordinated by the 3’ phosphate of pAp, Asp 86 and Glu 178, and the second magnesium also coordinated by Asp 292 and an additional water molecule. Unexpectedly, this structure revealed two conformations of the pAp 3’ phosphate at roughly 50% occupancy, each of which makes favorable interactions with the protein and either one or both magnesium ions (Fig. 5d). In the “up” position, the 3’-phosphate interacts with the protein residues Asp 292, Lys 93, Arg 100 and both magnesium ions while in the “down” position, the 3’ phosphate engages with Glu 178 and Asp 35 and with both magnesium ions either through direct or water-mediated interactions. The 5’-phosphate forms extensive interactions with polar residues Lys 93, Gln 97, Arg 100 and Arg 101. The adenine ring forms a π-stacking interaction with His 41; the adenine and ribose also formed water-mediated interactions with the protein.
Superposition of the hXRN1x-pAp structure with the D. melanogaster XRN1-DNA bound structure11 (dXRN1-DNA; PDB code 2Y35) revealed a relatively low RMSD of 2.72 Å2 with 51.3% identity between the XRN1 orthologs, indicating that the exonuclease structural components have been conserved. The “up” conformation of the 3’ pAp phosphate most resembles the position of the equivalent phosphate for the dXRN1-DNA substrate, although the smaller single-nucleotide inhibitor is shifted in position when compared to the larger oligonucleotide (Fig. 5e). This shift results in small changes in the side chain orientation of the π-stacking His 41 residue to optimize the interaction. Additionally, the hXRN1x loop containing Trp 560 undergoes significant conformational change when compared to dXRN1-DNA (Supplementary Fig. 6). Trp 560 interacts with the third nucleotide base in the dXRN1-DNA structure; presumably, the structure of this loop in hXRN1 rearranges in the presence of longer oligonucleotide substrates.
Although no structure of human XRN2 has been solved, the structures of C. elegans XRN233 (PDB code 7OPK) and hXRN1x-pAp superimpose well with an RMSD of 3.89 Å2, with high degree of structural similarity seen in the bimetal active site (Fig. 5f). Comparison human XRN1 and XRN2 sequences reveal an extremely high degree of sequence identity for residues within 6 Å of pAp (24 of 26 residues, 92%). While the two variable residues (XRN1 Gln 42 to XRN2 Pro 61, XRN1 Ser 104 to XRN2 Ala 121) may result in different side chain interactions and/or backbone conformations, both residues are located more than 5.5 Å from the pAp binding site (Fig. 5g) and are not expected to substantially influence pAp binding. Given this high degree of structural and sequence similarities between XRN1 and XRN2, it is unsurprising that pAp is also a very potent inhibitor of hXRN2. In contrast, mouse DXO (hereafter referred to as mDXO) has a completely dissimilar structural fold and pAp binding site when compared to hXRN1 (Supplementary Fig. 7). However, comparison of the pAp binding mode in hXRN1x and mDXO reveals the three-dimensional orientation of the inhibitor is conserved despite distinct interactions between the compound, the catalytic magnesium ions and the protein which arise from differences in XRN1 and DXO tertiary structures (Fig. 5h). The ability of pAp to form interactions with diverse tertiary protein structures is likely critical for its inhibition of multiple bimetal nuclease enzymes.
Discussion
The role of dsRNA sensing and viral mimicry in cancer, previously underappreciated, has been an increasingly active area of research in recent years. It is becoming clear that RNA modifying proteins such as XRN1 play critical roles in suppressing the innate immune response to endogenous dsRNA induction in a subset of cancer cells, and that targeting such proteins may lead to effective targeted oncology therapeutics. This work validates XRN1 as a dependency in a subset of lung cancer cell lines and demonstrates that upon XRN1 loss the PKR and MDA5 dsRNA sensing pathways are activated, leading to cell death. These results are consistent with other recent publications demonstrating that tumor cells with elevated Type I interferon signaling are particularly dependent on XRN116,17. The role of XRN1 in suppressing dsRNA signaling is also consistent with the overlap in Depmap profile of XRN1 and ADAR1, an RNA editor with a clear role in promoting the survival of endogenously high Type I IFN cell lines. These results suggest that an XRN1 inhibitor will have monotherapy potential in tumors with endogenously high Type I IFN signaling and provide a strong rationale for developing inhibitors to XRN1 for potential therapeutic use.
In addition to the cell autonomous effects of XRN1 loss in dependent cell lines, these data also support the hypothesis that XRN1 loss can potentiate the effects of checkpoint inhibitors. Checkpoint inhibitors such as anti-PD1 are known to induce Type I interferon signaling in the tumor microenvironment; this Type I IFN secretion has been linked to resistance to checkpoint inhibition over time23. This work demonstrates that exogenous Type I IFNs render previously insensitive cancer cell lines dependent on XRN1 for proliferation. This result suggests that combining XRN1 inhibition and checkpoint inhibition may be an effective therapeutic strategy to prevent or circumvent resistance to immunotherapy. In addition to conferring autonomous tumor cell sensitivity in the presence of Type I Interferons, XRN1 loss has also been shown to increase antigen presentation and tumor cell infiltration in a syngeneic mouse model21 and to be a CRISPR screen hit sensitizing B16 tumors to anti-PD-120. Additional combination opportunities outside of immunotherapy have also been demonstrated16; both palbociclib and 5-azacytidine have been shown to elevate dsRNA burden and intrinsic XRN1 dependence.
Thus, the results presented in this report add to the strong rationale for developing small molecule inhibitors of XRN1 as well as supporting both the monotherapy and combination potential for XRN1 inhibitors. Although complete XRN1 loss is embryonic lethal in mice, heterozygous mice develop normally and are fertile, suggesting that a therapeutic window is possible for XRN1 inhibition39. In addition, there are many marketed cancer therapeutics targeting proteins that are embryonic or early post-natal lethal in mice, such as VEGF, EGFR, and CDK4/640,41,42, which is often attributed to differential dependence between development and during adulthood. Consistent with this finding, these results indicate that multiple cell lines proliferate normally upon XRN1 loss. This is in contrast to XRN2 which appears to be a common essential gene as both CRISPR KO and siRNA demonstrate XRN2 dependency across cancer cell lines22,43. A XRN1 therapeutic, therefore, would need to be highly selective over XRN2 to minimize potential safety issues.
With biological validation of XRN1 as a target for tumors with high interferon signaling levels and/or sensitization to immunotherapeutic agents described here and by others16,17,21, development of an assay suite to enable drug discovery activities, including high throughput screening and compound validation, is required. The enzymatic assay for XRN1 described in this report, optimized under balanced conditions, enables large-scale hit finding activities, compound inhibitory activity assessment through concentration response and determination of the enzymatic mechanism of inhibition (MOI) for a compound of interest. The SPR binding assay provides an orthogonal method to quantify compound potency, selectivity and specificity, as well as to determine reversibility of binding and drug-target residence time information; these data are critical in the early phases of compound validation to filter compounds that are nonspecific inhibitors and prioritize well-behaved compounds for additional experiments. The XRN2 SPR assay provides a rapid assessment of compound selectivity between these closely related nucleases, an important consideration during series optimization. The human XRN1 crystal structure presented provides a platform for initiating crystallization experiments with any prioritized chemical matter resulting from hit finding activities. Importantly, the XRN1x construct utilized for crystallization is enzymatically active and can adopt protein confirmations necessary for catalysis. In addition, the structure includes two magnesium ions bound in the active site. Other ortholog structures of XRN1 have been solved using catalytically inactive variant proteins in which amino acid mutations alter magnesium coordination of the enzyme; structures of these XRN1 variants contain a single magnesium bound in the XRN1 active site11,44. As the proposed mechanism of substrate hydrolysis for XRN1 requires two magnesium ions, it is satisfying to confirm two metal ions bound in the predicted coordination sites within this wild-type protein structure11,44. Additionally, the interactions seen between pAp and the magnesium ions observed in the crystal structure are consistent the with loss of binding seen by SPR of pAp with the D208A mutant, suggesting the bimetal plays a significant role in driving pAp binding.
Utilization of a tool compound to test the robustness of any assay suite prior to initiation of hit finding activities is desirable. pAp was selected for characterization due to previous reports of weak to moderate inhibitory activity against XRN1 and other nuclease enzymes34,35, but the nanomolar potency of pAp for XRN1 in both biochemical and biophysical assays was unexpected. This result not only provided evidence of the ability of a small molecule to inhibit XRN1 enzymatic activity, but also indicated pAp could be a viable starting point for further drug discovery activities. At physiological pH, pAp contains two negative charges due to the phosphate moieties which would limit cell permeability; optimization of a pAp-based series to improve chemical and physical properties while maintaining or increasing XRN1 potency would be required. Nucleotide-based inhibitors such as sofosbuvir and remdesivir are FDA approved treatments for hepatitis C and COVID-19, respectively; these marketed compounds are prodrugs, requiring compound cleavage to the pharmacologically active chemical component after absorption, to improve properties necessary for in vivo activity45. In addition, achieving selectivity for XRN1 over XRN2 is likely required for a successful nucleotide-based molecule to enter the clinic; the structure of human XRN1 with pAp presented here as well as available XRN2 structures provide an excellent start to rational design efforts to improve the selectivity profile of nucleotide-based compounds.
Non-nucleotide compounds could also provide starting points for an XRN1-selective inhibitor. Compounds targeting bimetal active sites have been approved by the FDA, including the polymerase acidic (PA) protein inhibitor baloxavir marboxil for influenza A and B infections, as well as the integrase inhibitor raltegravir for HIV infections. Crystal structures reveal both compounds utilize alcohol and carboxyl groups to interact with the metal ions without utilization of negative charge46,47, therefore, screening of targeted libraries focused on moieties known to interact with metal ions would be a viable approach to identifying chemical starting points. As XRN1 is a processive enzyme48, conformational changes to the enzyme structure are likely necessary to process the RNA substrate during consecutive cycles of nucleotide hydrolysis. As all current structures of XRN1 and XRN2 have a high degree of similarity in their overall conformations, the extent of conformational change during enzyme catalysis is unknown. However, any conformational state may provide additional pockets for inhibitory compounds to bind. The XRN1 enzymatic assay described in this report, due to optimization under near-balanced conditions, provides opportunities to identify compounds that bind to all conformations of the enzyme and enzyme-substrate complex36. As balanced assays are preferred for screening large diversity libraries49, high throughput screening using the XRN1 biochemical assay presented here is a viable approach to identify chemical matter for XRN1 appropriate for further drug discovery efforts. Additionally, if wild-type XRN1 is enzymatically active in the SPR experiment and undergoing the conformational changes necessary for base cleavage to occur, as hypothesized by the SPR results with ssDNA, the enzyme may also be able to adopt conformations on the SPR chip that are stabilized or induced by small molecule inhibitors identified through biochemical screening efforts. Thus, SPR may be able support compound validation efforts through corroborating evidence of compound binding regardless of the enzyme conformational state.
XRN1 is a compelling target for oncology drug discovery efforts. The enzyme has been validated as a target capable of inducing apoptosis for cancer cells lines from various lineages in this and previous reports. Additional literature implicating XRN1 as important in cancer immunotherapy further highlights the potential of an XRN1 inhibitor to impact diverse patient populations. The assay suite developed for XRN1 provides the necessary tools to identify and validate inhibitors of the nuclease, and the discovery of pAp as a potent small molecule inhibitor in vitro indicates that XRN1 is a druggable enzyme. These tools may enable the discovery and optimization of potent, selective and cell active inhibitors of XRN1 that will facilitate further research into the biology of the enzyme and potentially translate into therapeutic options for patients.
Materials and methods
Cell culture and viral transduction
NCI-H1650, NCI-H1703, NCI-H1944, and NCI-H838 non-small cell lung cancer cell lines (ATCC) were cultured in 10% fetal bovine serum (FBS) and RMPI-1640 (Gibco). Cells were transduced with a Cas9-expressing lentivirus (Cellecta) and selected with 100 µg/mL hygromycin. Cells were then infected with sgRNA-containing lentivirus (Supplementary Table 2) and selected with 8 µg/ml puromycin for 3 days.
Proliferation and apoptosis assays
Following viral transduction and selection, cells were replated for proliferation, apoptosis, and colony formation assays. After 12 days of culture, proliferation was assessed by incubation with Cell Titer Glo® (Promega) for 10 min at room temperature. Luminescence was then measured on an Envision (Perkin Elmer). Apoptosis was measured with Caspase3/7 Glo® (Promega); the apoptosis assay was conducted 7 days post-plating and apoptosis induction is reported as fold change compared to uninfected cells. For colony formation, cells were incubated for 14 days, after which cells fixed with 70% ethanol for fifteen minutes, followed by incubation with crystal violet for 15 minutes. Cells were rinsed with PBS and imaged using an Odyssey CLx. Three replicates were assessed for each proliferation and apoptosis endpoint.
Western blot
Cells were rinsed with PBS, then lysed in RIPA buffer in the presence of protease and phosphatase inhibitors (Thermo Fisher) and kept on ice. Following addition of SDS-page loading buffer (Invitrogen), lysates were incubated at 95 °C for 20 min. 20–40 μg of protein was run on a precast tris-acetate gel (Invitrogen) and transferred to PVDF via iBlot (Invitrogen). For XRN1 western blots, gels were incubated in 5% ethanol/water for 5 min before transfer. Membranes were blocked with Odyssey blocking buffer before incubation with primary and secondary antibodies and imaging on an Odyssey CLx. Antibodies used were as follows: XRN1 (Cell Signaling Technology (CST) 70205; 1:1000), β-Actin (CST 3700; 1:2000), total PKR (R&D systems MAB1980; 1:500), phospho-Thr451 PKR (Abcam ab81303; 1:500), IRDye 800CW Goat anti-Rabbit (Li-Cor 926-32211; 1:20,000), and IRDye 680RD Goat anti-Mouse (Licor 926-68070; 1:20,000).
qPCR
RNA was isolated using the RNeasy Plus Mini kit (Qiagen); cDNA synthesis utilized the Superscript III kit (Thermo Fisher). The qPCR reaction was carried out on a Quantstudio (Thermo Fisher) using TaqmanTM master mix. Taqman probes utilized are as follows: IFNB1 FAM (ThermoFisher 4331182, Assay ID:Hs01077958_s1), IFNA1 FAM (ThermoFisher 4331182, Assay ID Hs03044218_g1), 18s VIC (ThermoFisher 4331182, Assay ID: Hs99999901_s1). Three replicates were assessed for each qPCR analysis.
Protein production
All constructs for human (H. sapiens) XRN1 (UniProt entry: Q8IZH2) and XRN2 (UniProt entry: NP_036387) proteins were generated by subcloning from synthesized, full-length XRN1 and XRN2 DNA sequences codon optimized for protein overexpression. Each construct (hXRN1, hXRN1-D208A, hXRN2, hXRN1-Avi, hXRN1-D208A-Avi, hXRN2-Avi, hXRN1x; construct boundaries are described in Results) was cloned into the pTT5 vector (NRC Biotechnology Research Institute) and included either a C-terminal FLAG or FLAG-Avi tag. All XRN1 and XRN2 constructs were expressed in Expi293F cells (Thermo Fisher) after transient transfection using standard methodologies. Cells were harvested at 72 h post-transfection by centrifugation, resuspended in PBS and centrifuged again. Cells were resuspended in lysis buffer and incubated with protease inhibitors (cOmplete EDTA-free Protease Inhibitor Cocktail Tablet, Roche) prior to lysis by homogenization (AH1500; ATS Engineering Limited). After centrifugation to remove cell debris, the lysate was purified by FLAG affinity chromatography. Anti-FLAG affinity gel (Sigma) was pre-equilibrated with Buffer A (25 mM HEPES, 300 mM NaCl, pH 7.5, protease inhibitor (1 tablet/200 ml)) then incubated with the supernatant and loaded in a column. Resin was washed with Buffer A followed by Buffer A with 5 mM ATP and 10 mM MgCl2. Protein was eluted with Buffer A with 250 μg/ml FLAG peptide until no signal was observed in eluent by Coomassie G-250 (Thermo Fisher). Proteins without an Avi tag were then loaded onto a size exclusion column (Superdex 200, GE) pre-equilibrated with Buffer B (25 mM HEPES, 300 mM NaCl, 1 mM DTT, pH 7.5) and run with the same buffer. The target proteins were concentrated by ultra-filtration (Amicon Ultra 30kD MWCO, Millipore) to 18 mg/mL, aliquoted, frozen using liquid nitrogen and stored at −80 °C.
Avi-tagged XRN1 and XRN2 proteins required additional steps in the purification protocol. After elution from the FLAG resin, the proteins were biotinylated using a His-tagged BirA enzyme via previously established methods50. The protein solution was added to a nickel affinity column preincubated in 25 mM HEPES, 300 mM NaCl, pH 7.5 and proteins were eluted with 25 mM HEPES, 300 mM NaCl, 20 mM imidazole, pH 7.5. The proteins were then purified by size exclusion chromatography in the same manner as described above.
XRN1 biochemical assay
Reagents
Substrate Annealing: RNA (5’-pACUCACUCACUCACCAAAAAAAAAAAAACC-FAM-3’) and DNA (5’-IOWABLACKFQ-GGTTTTTTTTTTTTTGG-OH−3’) oligomers were synthesized by Integrated DNA Technologies (Coralville, IA) and purified by high-performance liquid chromatography (HPLC) to greater than 95% purity. The DNA and RNA oligos were resuspended with RNAase/DNAase free or nuclease-free water. RNA and DNA were mixed at a 1:2 ratio with a final RNA concentration of 50 μM. To anneal DNA and RNA, sample was heated to 70 °C in a heat block for 5 min and allowed to cool slowly back to room temperature; samples were then aliquoted and stored at −20 °C.
Assay
The hXRN1 assay was performed in small volume, non-binding, 384-well black colored plates (catalog #781900, Greiner) at a final volume of 50 μL per well. 20 mM HEPES pH 7.5, 50 mM NaCl, 5 mM MgCl2, 1 mM DTT, 0.01% BSA, 0.004 U/μL RNaseOUT (catalog #10777019, Thermo Fischer Scientific) and 0.01% Tween-20 was used as optimized 1X assay buffer. For compound testing, hXRN1 (final concentration (f.c.) = 250 pM) was added using a Multidrop Combi (Thermo Fisher Scientific) and pre-incubated for 15 min at room temperature. Reactions were initiated by adding double-stranded RNA/DNA substrate (f.c. 1.5 nM). Plates were centrifuged and sealed; the reaction proceeded for 60 min. Reactions were stopped with EDTA (f.c. = 80 mM) using an Apricot (SPT Labtech). Plates were centrifuged and then read on an Envision (PerkinElmer) with excitation and emission filters at 485 nM and 535 nm, respectively.
Data analysis
Enzyme kinetics and parameters such as KM and kcat were calculated using Michaelis–Menten fits of steady-state enzyme velocities in GraphPad (San Diego, CA). Scigilian Analyze software (Montreal, QC, Canada) was used to calculate IC50 values and Hill slopes using four-parameter fits to the standard Hill-Langmuir equation. The quality and robustness of the assay were determined by analysis of the Z’ factor. IC50 values for inhibitors were calculated from the midpoint values of concentration–response plots.
SPR assay
Instrument preparation
The Biacore S200 instrument (Cytiva) was primed in 25 mM HEPES pH 7.5, 150 mM NaCl, 1.5 mM MgCl2, the SA series S sensor chip (Cytiva) was docked, and the analysis temperature was set to 20 °C for all subsequent steps.
Chip preparation
Prior to protein immobilization on the SA series S sensor chip, all flow cells were preconditioned by 3 consecutive injections of 50 mM NaOH in 1 M NaCl per manufacturer instructions. Human XRN1 or XRN2 proteins containing a biotinylated C-terminal Avi tag were used in this study. To immobilize several thousand response units of protein on an active flow cell of the streptavidin-coated (SA) chip via its biotinylated Avi tag, the protein was diluted in running buffer and injected at 5 μL/min. The SA-only reference flow cell was left untreated.
ssDNA binding
Running buffer comprised of sterile filtered 25 mM HEPES pH 7.5, 150 mM NaCl, 1.5 mM MgCl2 +/− 0.05% v/v Tween 20 was prepared; inclusion of detergent had no effect on data quality or resultant KD. A 3-fold dilution series of ssDNA in running buffer with a top concentration of 10 µM was prepared. Several cycles of buffer only association and dissociation were performed to sufficiently equilibrate the sensor chip. ssDNA binding was performed in multi-cycle kinetic mode at 30 μL/min flow rate. Association injection time was set to 60 s and dissociation time was set to 250 s. Flanking buffer blank injections were performed.
pAp binding
Running buffer comprised of 25 mM HEPES pH 7.5, 150 mM NaCl, 1.5 mM MgCl2, 0.05% Tween 20, and 1% DMSO was prepared. Matching buffer without DMSO was prepared for solvent correction solution generation. Solutions for a solvent correction standard curve were generated by spiking DMSO into the running buffer without DMSO, to achieve a range of DMSO concentrations that span 1% DMSO as the midpoint. Running buffer was sterile filtered prior to use. A 2-fold dilution series of pAp in running buffer from a top concentration of 200 nM was prepared. Several cycles of buffer-only association and dissociation were performed to sufficiently equilibrate the sensor chip. Serial injection of solvent correction solutions was performed to allow DMSO signal correction in data analysis. pAp binding was performed in multi-cycle kinetic mode at 30 μL/min flow rate. Association injection time was set to 90 s and dissociation time was set to 180 s. Flanking buffer blank injections were performed.
Data processing
Raw SPR sensorgram data were processed and analyzed in Biacore Insight Evaluation software. The concentration-dependent signal of DMSO on the reference flow cell compared to the active protein flow cell was corrected for in the data through use of a DMSO standard curve (solvent correction). To remove any non-specific binding to the chip from the sensorgram data, signal for pAp binding to the SA-only reference flow cell was subtracted from the pAp binding signal on the human XRN active flow cell. To remove buffer noise from the data, signal from a buffer only sensorgram on the active flow cell was subtracted from the pAp binding sensorgram. The fully processed (solvent corrected, double reference subtracted) pAp sensorgrams were mathematically fit using a standard 1:1 binding kinetic model (A + B ↔ AB). Output parameters from data fitting were the kinetic rate constants ka (on-rate) and kd (off-rate). Binding affinity (KD) was subsequently calculated in the analysis software using the equation KD = kd/ka. Residence time (τ) was manually calculated from the equation τ = 1/kd.
Crystallization and structure determination
hXRN1x was cocrystallized with pAp using the sitting drop vapor diffusion method at 18 °C. pAp (100 mM solution in water) and MgCl2 (100 mM solution in water) were added to hXRN1x (30 mg/mL in 25 mM HEPES, 300 mM NaCl, 1 mM DTT, pH 7.5) to generate a final crystallization sample containing 8 mg/mL hXRN1x, 5 mM MgCl2and 1 mM pAp. Crystallization drops were generated by mixing 2 μL crystallization sample with 1 μL precipitant solution containing 200 mM KF, 20% w/v PEG 3350, pH 7.3 and streak seeding methods were utilized to grow diffraction quality crystals. Crystals were passed through a solution containing 20% ethylene glycol, 80% precipitant solution prior to flash-freezing by immersion in liquid nitrogen. Diffraction data were collected at beamline BL45XU at Spring8 at 1 Å wavelength and 100 °K. Integration, scaling, and merging were performed with XDS51 and Aimless52; resolution limits were defined by I/σI ≥ 2. The structure was solved by molecular replacement in Phaser53 using the crystal structure of D. melanogaster XRN1 (PDB code 2Y35)11. Structure refinement was completed using iterative cycles of refinement and model building using REFMAC554 and COOT55, respectively. Validation of the refined structure was performed at the Protein Data Bank validation server (validate.wwpdb.org)56. Ramachandran statistics for the structure were 97.24% favored, 2.76% allowed, and 0.00% outliers. Data collection and refinement statistics are shown in Table 1. The structure has been deposited in the Protein Data Bank (PDB entry 9E70).
Statistics and reproducibility
Data is presented as mean +/− standard deviation when error bars are present. If not otherwise mentioned in the methods section, analysis was performed in Graphpad or Microsoft Excel. All experiments utilized 2–6 replicates and are representative of multiple independent experiments (typically 3).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data for all graphs can be found in Supplementary Data 1. Unedited uncropped agarose gel images are contained in Supplementary Fig. 2 within the Supporting Information. Unedited and uncropped western blot images are also found in Supporting Information (Supplementary Figs. 8 and 9). The XRN1 crystal structure has been deposited in the Protein Data Bank (PDB entry 9E70). Other requests for source data can be accommodated by the corresponding author upon reasonable request.
References
Katze, M. G., He, Y. & Gale, M. Viruses and interferon: a fight for supremacy. Nat. Rev. Immunol. 2, 675–687 (2002).
Samuel, C. E. Antiviral Actions of Interferons. Clin. Microbiol. Rev. 14, 778–809 (2001).
Chen, R., Ishak, C. A. & De Carvalho, D. D. Endogenous Retroelements and the Viral Mimicry Response in Cancer Therapy and Cellular Homeostasis. Cancer Discov. 11, 2707–2725 (2021).
Roulois, D. et al. DNA-Demethylating Agents Target Colorectal Cancer Cells by Inducing Viral Mimicry by Endogenous Transcripts. Cell 162, 961–973 (2015).
Chiappinelli, K. B. et al. Inhibiting DNA Methylation Causes an Interferon Response in Cancer via dsRNA Including Endogenous Retroviruses. Cell 162, 974–986 (2015).
Griffin, G. K. et al. Epigenetic silencing by SETDB1 suppresses tumour intrinsic immunogenicity. Nature 595, 309–314 (2021).
Sheng, W. et al. LSD1 Ablation Stimulates Anti-tumor Immunity and Enables Checkpoint Blockade. Cell 174, 549–563.e19 (2018).
Bowling, E. A. et al. Spliceosome-targeted therapies trigger an antiviral immune response in triple-negative breast cancer. Cell 184, 384–403.e21 (2021).
Liu, H. et al. Tumor-derived IFN triggers chronic pathway agonism and sensitivity to ADAR loss. Nat. Med. 25, 95–102 (2019).
Stevens, A. 5′-Exoribonuclease 1: Xrn1. In: Methods in Enzymology vol. 342 251–259 (Elsevier, 2001).
Jinek, M., Coyle, S. M. & Doudna, J. A. Coupled 5′ Nucleotide Recognition and Processivity in Xrn1-Mediated mRNA Decay. Mol. Cell 41, 600–608 (2011).
Houseley, J. & Tollervey, D. The Many Pathways of RNA Degradation. Cell 136, 763–776 (2009).
Parker, R. & Song, H. The enzymes and control of eukaryotic mRNA turnover. Nat. Struct. Mol. Biol. 11, 121–127 (2004).
Akiyama, B. M., Eiler, D. & Kieft, J. S. Structured RNAs that evade or confound exonucleases: function follows form. Curr. Opin. Struct. Biol. 36, 40–47 (2016).
Dilweg, I. W., Peer, J. & Olsthoorn, R. C. L. Xrn1-resistant RNA motifs are disseminated throughout the RNA virome and are able to block scanning ribosomes. Sci. Rep. 13, 15987 (2023).
Hosseini, A. et al. Retroelement decay by the exonuclease XRN1 is a viral mimicry dependency in cancer. Cell Rep. 43, 113684 (2024).
Zou, T. et al. XRN1 deletion induces PKR-dependent cell lethality in interferon-activated cancer cells. Cell Rep. 43, 113783 (2024).
Gal-Ben-Ari, S., Barrera, I., Ehrlich, M. & Rosenblum, K. PKR: A Kinase to Remember. Front. Mol. Neurosci. 11, 480 (2019).
Pichlmair, A. et al. Activation of MDA5 Requires Higher-Order RNA Structures Generated during Virus Infection. J. Virol. 83, 10761–10769 (2009).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Ran, X. -B. et al. Targeting RNA Exonuclease XRN1 Potentiates Efficacy of Cancer Immunotherapy. Cancer Res. 83, 922–938 (2023).
DepMap, B. DepMap 23Q4 Public. 21888030738 Bytes Figshare+ https://doi.org/10.25452/FIGSHARE.PLUS.24667905.V2 (2023).
Jacquelot, N. et al. Sustained Type I interferon signaling as a mechanism of resistance to PD-1 blockade. Cell Res. 29, 846–861 (2019).
Chang, C. -T., Bercovich, N., Loh, B., Jonas, S. & Izaurralde, E. The activation of the decapping enzyme DCP2 by DCP1 occurs on the EDC4 scaffold and involves a conserved loop in DCP1. Nucleic Acids Res. 42, 5217–5233 (2014).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Solinger, J. A., Pascolini, D. & Heyer, W. -D. Active-Site Mutations in the Xrn1p Exoribonuclease of Saccharomyces cerevisiae Reveal a Specific Role in Meiosis. Mol. Cell. Biol. 19, 5930–5942 (1999).
Sinturel, F. et al. Real-time fluorescence detection of exoribonucleases. RNA 15, 2057–2062 (2009).
Buker, S. M. et al. A Mass Spectrometric Assay of METTL3/METTL14 Methyltransferase Activity. SLAS Discov. 25, 361–371 (2020).
Renaud, J. -P. et al. Biophysics in drug discovery: impact, challenges and opportunities. Nat. Rev. Drug Discov. 15, 679–698 (2016).
Edelmann, F. T., Niedner, A. & Niessing, D. Production of pure and functional RNA for in vitro reconstitution experiments. Methods 65, 333–341 (2014).
Nagarajan, V. K., Jones, C. I., Newbury, S. F. & Green, P. J. XRN 5′→3′ exoribonucleases: Structure, mechanisms and functions. Biochim. Biophys. Acta BBA Gene Regul. Mech. 1829, 590–603 (2013).
Richter, H., Katic, I., Gut, H. & Großhans, H. Structural basis and function of XRN2 binding by XTB domains. Nat. Struct. Mol. Biol. 23, 164–171 (2016).
Dichtl, B. Lithium toxicity in yeast is due to the inhibition of RNA processing enzymes. EMBO J. 16, 7184–7195 (1997).
Yun, J. -S. et al. Molecular mechanism for the inhibition of DXO by adenosine 3′,5′-bisphosphate. Biochem. Biophys. Res. Commun. 504, 89–95 (2018).
Copeland, R. A. Evaluation of Enzyme Inhibitors in Drug Discovery: A Guide for Medicinal Chemists and Pharmacologists https://doi.org/10.1002/9781118540398. (Wiley, 2013).
Buker, S. M., Boriack-Sjodin, P. A. & Copeland, R. A. Enzyme–Inhibitor Interactions and a Simple, Rapid Method for Determining Inhibition Modality. SLAS Discov. 24, 515–522 (2019).
Overbeck, J. H., Stelzig, D., Fuchs, A. -L., Wurm, J. P. & Sprangers, R. Observation of conformational changes that underlie the catalytic cycle of Xrn2. Nat. Chem. Biol. 18, 1152–1160 (2022).
Takaoka, S. et al. Neuronal XRN1 is required for maintenance of whole-body metabolic homeostasis. iScience 24, 103151 (2021).
Ferrara, N. et al. Heterozygous embryonic lethality induced by targeted inactivation of the VEGF gene. Nature 380, 439–442 (1996).
Linder, M. et al. EGFR controls bone development by negatively regulating mTOR-signaling during osteoblast differentiation. Cell Death Differ. 25, 1094–1106 (2018).
Maurer, B. a. r. b. a. r. a., Brandstoetter, T. a. n. i. a., Kollmann, S. e. b. a. s. t. i. a. n. & Sexl, V. e. r. o. n. i. k. a. & Prchal-Murphy, Michaela. Inducible deletion of CDK4 and CDK6 – deciphering CDK4/6 inhibitor effects in the hematopoietic system. Haematologica 106, 2624–2632 (2020).
Tsherniak, A. et al. Defining a Cancer Dependency Map. Cell 170, 564–576.e16 (2017).
Chang, J. H., Xiang, S., Xiang, K., Manley, J. L. & Tong, L. Structural and biochemical studies of the 5′→3′ exoribonuclease Xrn1. Nat. Struct. Mol. Biol. 18, 270–276 (2011).
Rautio, J., Meanwell, N. A., Di, L. & Hageman, M. J. The expanding role of prodrugs in contemporary drug design and development. Nat. Rev. Drug Discov. 17, 559–587 (2018).
Hare, S. et al. Molecular mechanisms of retroviral integrase inhibition and the evolution of viral resistance. Proc. Natl Acad. Sci. 107, 20057–20062 (2010).
Omoto, S. et al. Characterization of influenza virus variants induced by treatment with the endonuclease inhibitor baloxavir marboxil. Sci. Rep. 8, 9633 (2018).
Stevens, A. Purification and characterization of a Saccharomyces cerevisiae exoribonuclease which yields 5’-mononucleotides by a 5’ leads to 3’ mode of hydrolysis. J. Biol. Chem. 255, 3080–3085 (1980).
Copeland, R. A. Mechanistic considerations in high-throughput screening. Anal. Biochem. 320, 1–12 (2003).
Chan-Penebre, E. et al. A selective inhibitor of PRMT5 with in vivo and in vitro potency in MCL models. Nat. Chem. Biol. 11, 432–437 (2015).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution?. Acta Crystallogr. D. Biol. Crystallogr. 69, 1204–1214 (2013).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of Macromolecular Structures by the Maximum-Likelihood Method. Acta Crystallogr. D. Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Berman, H., Henrick, K. & Nakamura, H. Announcing the worldwide Protein Data Bank. Nat. Struct. Mol. Biol. 10, 980–980 (2003).
Acknowledgements
The authors thank Jim Valentine, Cindy Collins, Alexandra Gardino, Gina Prophete, Zachary Burke and our CRO partners (particularly Deborah Bruce, Clemence Mondielli, Ghislaine Marchand, Bastien Cautain, Corrine Lafon, Yu Xia, Xiangduo Mu, and Shenhua Zhang) for project support, Matthew Lynes for manuscript formatting assistance, and current and former Accent employees for helpful discussions.
Author information
Authors and Affiliations
Contributions
These authors contributed equally: Gordon J. Lockbaum and Maureen M. Lynes. G.L., M.L., P.A.B., K.K., S.G., B.S., L.S., S.S., S.N., S.M.B., S.J.S., K.D., S.J.B., and R.C. contributed to manuscript writing. G.L., M.L., S.S., J.L., N.H., S.G., S.N., K.K., S.Y., E.A.S., S.M.B., B.S., A.C. designed and conducted experiments as well as analyzed data from our CRO partners. S.R. conceptualized the original XRN1 Depmap analysis which was iterated upon by J.W. and A.R.; J.W. and A.R. helped with additional data analysis and visualization for this work. L.S. performed data analysis and manuscript revisions. G.L., M.L., B.S., E.A.S., K.K., S.N., S.M.B., S.R., S.G., K.D., J.L., S.J.B., R.C., S.J.S., and P.A.B. provided scientific oversight as well as contributed to manuscript revision and advice.
Corresponding author
Ethics declarations
Competing interests
The authors are current or former employees of Accent Therapeutics, Inc, where the work was performed.
Peer review
Peer review information
Communications Biology thanks John Gross and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Ingrid Span and Kaliya Georgieva. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lockbaum, G.J., Lynes, M.M., Shen, S.A. et al. Characterization of exoribonuclease XRN1 as a cancer target and identification of adenosine-3’,5’-bisphosphate as a potent enzyme inhibitor. Commun Biol 8, 589 (2025). https://doi.org/10.1038/s42003-025-08005-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08005-y