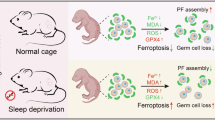
Abstract
Sleep deprivation (SD) has been demonstrated to cause male reproductive dysfunction; however, its underlying mechanisms remain unclear. In this work, we conduct Mendelian randomization analysis, which indicates a significant association between sleep disorders and male infertility in humans. To explore the potential mechanism, ten-week-old male Sprague-Dawley rats are subjected to continuous SD for five days, showing significantly reduced epididymal sperm concentration and motility compared to the control group. SD treatment also decreases serum testosterone levels and epididymal sperm transit time in male rats. Histological analysis reveals reproductive system damage, while bulk and single-cell RNA sequencing highlight that SD significantly alters transcriptomes and induces differentially expressed genes with significant heterogeneity across three segments of the epididymis. Gene ontology analysis indicates that SD upregulates inflammatory response genes, especially in the more inflammation-sensitive cauda epididymis. Moreover, SD activates immune cells and causes cytokines and chemokines to accumulate in the cauda epididymis. However, recovery sleep mitigates this damage. Our findings reveal that continuous SD disrupts the epididymal immunological microenvironment, lowering sperm quality and potentially contributing to male infertility.
Similar content being viewed by others
Introduction
Sleep is a fundamental physiological need that supports both physical and mental health. However, insomnia is prevalent among adults and teenagers worldwide. A study conducted in Germany using the generally accepted cut-off of Pittsburgh Sleep Quality Index (PSQI) > 5 showed that 36% of the general population experienced poor sleep, with females reporting more sleep issues than males1. After evaluating elements of PSQI, a recent study found that 20% of teenagers in the Chinese mainland experienced sleep issues2. Various factors, including anxiety and depression3, SARS-CoV-2 infection4, and substance abuse, contribute to poor sleep quality5.
A short sleep duration, defined as less than six hours of sleep per night6, has been linked to an increased risk of several conditions, including type 2 diabetes7, cardiovascular disease8, obesity9, hypertension10, dementia11, and higher mortality rates12. However, the mechanisms by which insufficient sleep adversely affects health are complex and not fully understood. Sleep deprivation (SD) might trigger a cytokine-storm-like syndrome in mammals, marked by increased levels of cytokines such as interleukin (IL)-6 and IL-17 and proinflammatory factors such as C-X-C motif chemokine ligand 1 (CXCL1) and 2 (CXCL2), leading to neutrophil recruitment from the bone marrow into the bloodstream13. High-quality sleep is essential for preventing oxidative stress. SD can alter the redox status of certain sleep-regulating neurons and increase reactive oxygen species levels in the gut, as demonstrated by studies on Drosophila14,15. Oxidative stress has been implicated in the SD-induced pathogenesis of male reproductive dysfunction16. This is attributable to the fact that spermatozoa membranes are rich in polyunsaturated fatty acids, rendering them susceptible to oxidative damage17,18. The effects of SD on the male reproductive system and the mechanisms by which SD modulates spermatogenesis remain elusive; hence, further studies on its pathogenesis are urgently needed. Continuous SD has been proven to induce testicular damage and result in low sperm quality16. The epididymis, crucial for sperm maturation, is where spermatozoa from the testes acquire motility19. Single-cell RNA sequencing (scRNA-seq) analysis has revealed segment-specific gene expression patterns in epididymal cells, suggesting that different sections of the epididymis have distinct roles in regulating sperm maturation20. Studies examining whether SD induces epididymal damage and its underlying molecular mechanisms at the single-cell level remain scarce.
The present study elucidates the mechanisms underlying the effects of SD on the male reproductive system using a continuous SD-treated rat model. After five days of SD, significant decreases in sperm count and motility were observed in male rats, along with a reduction in serum testosterone levels. Further, scRNA-seq analysis revealed that SD led to a significant upregulation of genes encoding cytokines and chemokines in the cauda epididymis. This upregulation was accompanied by an increase in the number of macrophages in the cauda epididymis with continuous SD, suggesting that SD-induced immune activation contributes to the decline in sperm quality. Additional experiments demonstrate that recovery sleep could improve sperm quality and mitigate SD-induced epididymal damage. These findings complement the Mendelian randomization analysis results, which verify a significant correlation between sleep disorders and male infertility. Together, these findings provide insights into the detrimental effects of SD on the male reproductive system and offer valuable theoretical support for addressing the negative effects of sleep disorders on male reproductive health.
Results
Mendelian randomization
To investigate the causal relationship between sleep disorders and male infertility, Mendelian randomization analyses were conducted using genome-wide association studies (GWAS) data. Five distinct Mendelian randomization approaches confirmed that sleep disorders were a risk factor for male infertility (Fig. 1A). Data from 93 genetic variants associated with sleep disorders were extracted from the male infertility GWAS dataset. In the funnel plot, the causal effects showed close symmetry, indicating significant causality for all analyzed single nucleotide polymorphisms (SNPs) (Fig. 1B). Using the inverse variance weighted method, a significant association was identified between sleep disorders and male infertility risk, with an odds ratio of 1.152 (P = 0.028, Fig. 1C). A sensitivity analysis of the Mendelian randomization findings was performed utilizing the leave-one-out method, where each SNP was systematically excluded, and the meta-effect of the remaining SNPs was recalculated. As illustrated in Fig. 1D, the exclusion of individual SNPs resulted in minimal variations in the overall error bars, consistently positioned to the right of zero. This observation underscores the robustness of the results. Genetic evidence supports the link between frequent sleep disorders and the increased risk of male infertility.
A The scatter plot of the causal relationship between sleep disorders and male infertility uses five different methods for analysis: inverse variance weighted, MR-Egger, weighted median, weighted mode, and simple mode. B The funnel plot displays the overall heterogeneity of the Mendelian randomization estimates for the effect of sleep disorders on male infertility. C The forest plot of the causal relationship between each SNP and the risk of male infertility. D The forest plot of the leave-one-out method for sensitivity analysis.
SD impairs the male reproductive system
To explore the effects of SD on the male reproductive system, adult male rats were subjected to SD for five days using a modified multiple-platform method (Fig. 2A). SD resulted in a significant loss of body weight in rats (Fig. 2B, Supplementary Table 1), but no morphological differences were observed in both the testis and epididymis between SD-treated and control rats (Fig. 2C, Supplementary Fig. 1C, D). To further evaluate the effects of SD on the reproductive system of male rats, seminiferous tubules at the same cycle stage of control and SD groups were selected. The 14-stage seminiferous tubules were complete in both rat groups (Supplementary Fig. 1E). The distribution for the stages of the seminiferous epithelium cycle in testicular cross-sections from control and SD-treated rats was also quantitatively compared. The 14 stages of the cycle were categorized as I–VI, VII–VIII, IX–X, XI–XIII, and XIV. The statistical results showed that the frequency of I–VI tubules in the testis of the SD group was increased; however, there was no significant difference overall in the distribution across the stage of the cycle groupings between the control and SD-treated rats (Supplementary Fig. 1F, G). We then evaluated the quality of sperm isolated from the cauda epididymis using the computer-assisted semen assay (CASA) system and found sperm concentration and motility of SD-treated rats were significantly reduced compared to the controls (Fig. 2H, I). It is consistent with statistical analysis that the sperm number in all regions of the epididymis showed a significant decrease (Supplementary Table 2). Histological analyses via hematoxylin and eosin (H&E) staining of testicular sections revealed no differences in seminiferous tubule diameters between each group, but the seminiferous tubular lumen was significantly enlarged following SD treatment, while the epithelial height was significantly reduced, as compared to the control group (Fig. 2D, Supplementary Fig. 1H–J). Furthermore, the SD group presents an increase in testicular lymphatic spaces (Supplementary Fig. 1F). Terminal deoxynucleotidyl transferase-mediated deoxyuridine triphosphate nick end labeling (TUNEL) assays revealed that SD caused significant increases in the proportion of TUNEL-positive cells in the epididymis, and no significant apoptosis was observed in testes of the SD and control groups (Fig. 2E). Interestingly, histological analyses showed that the epididymis of SD-treated adult rats exhibited a severe reduction of spermatozoa (Fig. 2F, G). In addition, the serum testosterone level of the SD group was significantly lower than that of the control group (Fig. 2J). Importantly, sperm transit time in the caput-corpus and cauda epididymis of adult rats after five days of SD showed a significant decrease (Supplementary Fig. 2). Taken together, these results indicate that SD has a potential negative impact on male reproductive health.
A Schematic diagram for the experimental modified multiple-platform SD model. Elements of this image are freely available from Servier Medical Art (https://smart.servier.com/). B Effect of SD on body weight (5 replicates per group). C Morphological images of the testes and epididymides from control and SD group rats (3 replicates per group). Scale bars: 10 mm D Hematoxylin and eosin (H&E) staining of the testicular sections from control and SD rats; black, blue, and green arrows indicate the diameter of the seminiferous tubule, diameter of the seminiferous lumen, and epithelial height of the seminiferous tubule, respectively. Scale bars: 100 µm. E A TUNEL assay was performed with Alexa Fluor 488 (green), and cell nuclei were labeled with DAPI (blue). Scale bars: 100 µm. F H&E staining of the epididymal sections from control and SD rats; the SD group shows significantly reduced sperm density within the lumens (black arrow). Scale bars: 100 µm. G The histogram represents the proportion of lumens with abnormal sperm counts in the epididymis of control and SD rats. More than 290 lumens were counted for each rat (3 replicates per group). H, I SD significantly reduces epididymal sperm concentration and motility (7 replicates per group). J Serum testosterone level in control and SD groups (7 replicates per group). Data are presented as the mean ± standard deviation values. Statistical analysis was performed with an unpaired t-test; ns > 0.05, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. ns, not significant.
SD altered the transcriptome in the epididymis of rats
To further investigate how SD affects sperm quality, bulk RNA sequencing was conducted to identify the transcriptomic profile of testicular and epididymal tissues of SD-treated male rats. The correlation heatmap of testicular tissues showed consistent relationships between the SD and control groups. Differential expression analysis showed that only 5 genes were downregulated or upregulated (Supplementary Fig. 1A, B). Furthermore, we assessed SD-induced transcriptome profile-related changes in the caput, corpus, and cauda epididymis. The correlation heatmap illustrated the relationship between each group and the datasets for different epididymal tissues (Fig. 3B). Differentially expressed genes (DEGs) in different regions of the epididymis were subjected to principal component analysis (PCA), which helped to effectively distinguish SD-treated and control samples along PC1 (Fig. 3A). Volcano plots were used to analyze the expression of DEGs. Notably, 2061 genes were upregulated and 1178 genes were downregulated in the caput epididymis, and 998 genes were upregulated and 525 genes were downregulated in the corpus epididymis, while 155 genes were upregulated and 888 genes were downregulated in the cauda epididymis (Fig. 3C). These results suggest that SD has a significant negative impact on the epididymis.
A, B Principal component analysis (PCA) and Spearman’s rank correlation coefficient of RNA sequencing data from the caput, corpus, and cauda epididymis of control and SD-treated rats, with 3 replicates per group. C Volcano plots showing downregulated and upregulated expressed genes in the caput, corpus, and cauda epididymis of SD-treated rats compared to controls (3 replicates per group).
Effects of SD on the immunological milieu of the cauda epididymis
A gene ontology (GO) analysis of SD-induced expression levels of DEGs based on P-values was performed for cauda epididymal tissues. The results revealed enrichment in biological processes related to inflammatory response regulation (Fig. 4A). Importantly, GO analysis identified upregulated genes that were significantly enriched in signaling pathways related to cytokine response pathways (IL-1 and IL-17) (Fig. 4B). Furthermore, IL-6, a key mediator of cytokine storms, was upregulated in the cauda epididymis after SD treatment (Fig. 4B), indicating an increase in cytokine levels with SD progression. In addition, the DEGs in the caput epididymis and the corpus epididymis assigned to inflammatory responses were not as significant as in the cauda epididymis (Supplementary Fig. 3A–D).
A–D Gene Ontology (GO) analysis of biological processes for identifying the potential functions of differentially expressed genes in the cauda epididymis.
The other effects of SD on the cauda epididymis were analyzed. The top differentially downregulated genes in the cauda epididymis were associated with biological processes such as cell-cell junction organization, tight junction organization, and tight junction assembly (Fig. 4C). GO analysis showed that genes associated with sperm capacitation and motility were also downregulated (Fig. 4D).
Cell type annotation of scRNA-seq clusters in the cauda epididymis
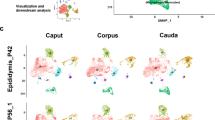
Based on the aforementioned data, scRNA-seq analysis was performed to identify distinct cell populations and elucidate the molecular mechanisms underlying the effects of SD on cauda epididymal cells. After quality control, a total of 8081 and 8071 cauda epididymal cells from the control and SD-treated groups, respectively, were visualized and categorized into ten cell clusters (Fig. 5A). Using established cell markers, these cell clusters were found to be comprised of fibroblasts, principal cells, basal cells, myoid cells, neuronal cells, clear cells, endothelial cells, spermatids, T cells, and monocytes (Fig. 5B, C).
A Combined results of Uniform Manifold Approximation and Projection (UMAP) and clustering analysis on single-cell transcriptomes of the cauda epididymis of the sleep-deprived (SD) (n = 8071) and control (n = 8081) rats. Each dot represents a single cell color-coded based on the cell cluster and experimental group, as indicated in the figure legend. B The heatmap displays the expression of up to 200 representative genes across cell clusters. C Representative gene expression patterns across each cell cluster, with independent expression scales for each gene.
SD affected the T cell and monocyte subgroups in the cauda epididymis
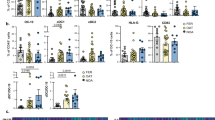
According to bulk RNA sequencing data, genes associated with neutrophils were upregulated after SD, suggesting a potential change in the distribution of leukocyte cell types during SD. To further delineate subpopulations of T cells and monocytes, we used a higher shared nearest neighbor clustering resolution. Four subpopulations of T cells (effector CD8 + T cells, effector memory CD8 + T cells, CD4 + T cells, and naive CD8 + T cells), and two subpopulations of monocytes (dendritic cells and macrophages) were revealed (Fig. 6A, Supplementary Fig. 4A). We analyzed the DEGs of these cell clusters to uncover the mechanism by which important molecules regulate the effects of SD on male fertility. When compared to other cell types, macrophages (29 upregulated and 116 downregulated DEGs) exhibited the most significant changes in gene expression, suggesting that this cell type responded differently to SD (Fig. 6B, Supplementary Fig. 4B). Specifically, Cxcl1 and Cxcl2 were substantially upregulated after SD in macrophages (Fig. 6B, D). Chemokine genes such as C-C motif chemokine ligand 2 (Ccl2), Ccl4, Ccl5, and Ccl7 were also upregulated (Fig. 6B). Furthermore, we found an increase in macrophage percentages (8.9% in the control group vs. 11.5% in the SD group) after SD treatment (Fig. 6C). Notably, Granzyme A (Gzma), Gzmb, and Gzmk expression levels were significantly higher in the SD group, specifically in effector CD8 + T cells (Fig. 6B, Supplementary Fig. 4B). Bulk RNA sequencing revealed the enrichment of upregulated genes in response to the IL-6 and IL-17 signaling pathways. Moreover, we found that Interleukin-6 receptor (Il-6r) and Interleukin-17 receptor A (Il-17ra) were widely expressed in cauda epididymal cells (Supplementary Fig. 5).
A Uniform Manifold Approximation and Projection (UMAP) and clustering analysis of the four subgroups identified in the T cell cluster and two subgroups in the monocyte cluster. Each cell is colored based on its group and the identity of the cell cluster as indicated in the figure legend. B Heatmap showing pseudobulk expression levels for identified cell clusters in the control and SD groups. C The proportion of immune cells identified in the T cell and monocyte clusters in both control and sleep deprivation (SD) groups. D Expression levels of Cxcl1 and Cxcl2 in the T cell and monocyte subgroups from each group.
SD altered cell-cell communication patterns in the cauda epididymis
CellChat was used to analyze the communication of different cell types in the cauda epididymis identified by scRNA-seq. We found that the SD group has both a higher quantity and stronger intensity of cell-cell communications compared to the control group (Fig. 7A). We also observed the fibroblasts in the SD group showed the strongest trend of enhanced intercellular communication, particularly with T cells. These results suggest that fibroblasts in the cauda epididymis play a central role in the increased number and strength of cell interactions after SD (Fig. 7B).
A Comparison of the number of inferred interactions and interaction strength between SD-treated and control cauda epididymal cells. The left bar chart shows the total number of inferred cell-cell interactions, while the right bar chart displays the average interaction strength for the SD and control groups. B Differential cell-cell interactions and interaction strength between cell types. The left panel illustrates the differential number of interactions between cell types, and the right panel depicts the differential interaction strength. The nodes represent different cell types, and the direction of the arrows indicates the direction of cell communication (from the initiating cell to the receiving cell). Red arrows signify an increase in communication quantity or strength in the SD group compared to the control group, while blue arrows indicate the opposite. The thickness of the arrows represents the magnitude of the respective change.
Improving sleep alleviates the SD-induced abnormal epididymal phenotype
A sleep recovery experiment was conducted to further assess whether SD directly induces inflammation and reduces sperm quality in the cauda epididymis. Male rats subjected to SD for five days were transferred to the same culture environment as the control group immediately. A phenotypic analysis was performed after 28 days. Histopathological analysis revealed that the epididymal cytoarchitecture was intact in the recovery group, with the sperm density in the lumen being comparable to that of the control group (Fig. 8A). In addition, sleep recovery partly restored sperm concentration and motility in male rats previously subjected to SD (Fig. 8B, C). Whole transcriptome profiles of the caput, corpus, and cauda epididymis from sleep recovery and control groups were analyzed via bulk RNA sequencing. PCA and correlation heatmaps showed the relationships between each dataset, and the analysis results indicated that sleep recovery mitigated the negative effects of SD on the male reproductive system (Fig. 8D, E).
A Hematoxylin and eosin (H&E) staining of cauda epididymis sections from the control and recovery sleep group rats, showing no significant change in sperm density within the lumen. Scale bars: 200 µm. B, C Histograms of sperm concentration and motility in control and recovery sleep group rats. Data are presented as the mean ± standard deviation values (3 replicates per group). Statistical analysis was performed with an unpaired t-test. ns > 0.05, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. ns, not significant. D, E Principal component analysis (PCA) and Spearman’s rank correlation coefficient analysis of RNA sequencing data from the caput, corpus, and cauda epididymis of control and recovery sleep group rats (3 replicates per group).
Discussion
A growing number of individuals are experiencing sleep disorders, largely due to the rapid development of the economy and accelerated pace of life. These disorders have garnered attention because of their impact on semen quality, making it essential to comprehensively assess how SD affects male reproduction and identify factors contributing to male fertility. Understanding these factors might guide the identification of therapeutic targets for this clinical condition.
In this study, we first explored the genetic link between sleep disorders and male infertility using Mendelian randomization and found that frequent sleep disorders increased the risk of male infertility. A related study conducted by Liu et al. involving 981 male participants found that individuals who slept for less than 6 h per night exhibited significantly reduced sperm count, motility, and survival rates compared to those with average (7–8 h per night) or extended (>9 h per night) sleep durations21. Similarly, Demirkol et al. also identified a negative correlation between sleep disorders and sperm concentration in 104 shift workers22. Collectively, these findings indicated that sleep disorders markedly impair sperm quality, which is consistent with our findings. Our animal model-based study indicated that five consecutive days of SD significantly reduced the sperm concentration and motility in rats, and resulted in a breakdown of testicular and epididymal tissue structures, compared to controls. Moreover, the accelerated sperm transit time in all regions of the epididymis may be associated with the decrease in sperm reserves in the epididymis of rats subjected to five days of SD. Rapid sperm transit through the epididymis often affects the maturation process of epididymal sperm23. These results support previous findings demonstrating that SD can lower semen quality and impair male reproductive function16. To delve deeper into the mechanisms by which SD affects sperm quality, we generated a comprehensive transcriptome profile of testicular and region-specific epididymal genes of rats subjected to SD. It provides valuable insight into the effect of SD on the male reproductive system. Interestingly, the expression levels of DEGs in the epididymis after SD were more significant than those in the testis. The present findings regarding SD-induced alterations in the expression profiles of testicular genes are not in agreement with those of a previous study, which showed that SD led to a significant change in the expression of genes in the testis24. The discrepancy in the results regarding the impact of SD on DEGs in the testis could probably be attributed to differences in the selection of experimental animal strains and selected SD models. In this study, ten-week-old male Sprague-Dawley rats were subjected to continuous SD, while in the study by Chen et al., eight-week-old Wistar rats were used, and a chronic sleep restriction model was employed, allowing rats to sleep for 6 h every day24. Chronic sleep restriction has been shown to trigger hyperphagia, leading to weight gain25. Our findings were consistent with those of Hamed et al., who showed that continuous SD results in weight loss16. These findings reflect the complexity of the effects of SD on biological function. Research has shown that sleep loss can affect the metabolic function and the regulatory function of hormone secretion in adults26. In this study, serum testosterone levels were significantly lower in the SD group, which was consistent with previous research16. Furthermore, we found significant weight loss in the rats after five days of continuous SD. Underweight is one of the important factors affecting fertility status and is associated with reduced levels of fertility-related hormones27.
In addition, our findings indicate that the cellular immune response in the cauda epididymis was significantly altered by SD. GO analysis suggests that upregulated genes were highly enriched in pathways associated with responses to IL-1 and IL-17, with Il-6 being strongly expressed after SD treatment. IL-1, the first cytokine reported to be associated with sleep, can mediate innate immunity and induce IL-6 generation. The most prevalent SD-induced cytokines, IL-6 and IL-17, also act as key mediators of human cytokine storms13,28. Upregulated genes in the cauda epididymis were the second highest enriched in response to lipopolysaccharides (LPS), which was consistent with previous findings showing that acute sleep loss induced LPS ligation of toll-like receptor 4 or CD14, leading to significantly increased IL-6 and TNF-α production in peripheral blood monocyte populations in humans29. The increase of IL-6 may account for the weight loss observed in this study. AMP-activated protein kinase can be activated by IL-6, which also promotes lipolysis and glucose disposal30. Notably, SD-induced DEGs showed significant heterogeneity across epididymis regions. The epididymis is more susceptible than the testis to inflammation and autoimmunity as previously reported31,32. Multiple cell types of immunological importance are present throughout the epididymal epithelium; by contrast, the seminiferous epithelium lacks immune cells33. Macrophages and dendritic cells are found throughout the epididymis34. In the absence of inflammation, dendritic cells and macrophages are numerous, mainly in the caput epididymis35. The possible reason is that these cells have the potential to inhibit sperm autoimmunity31. Studies of bacterial epididymo-orchitis showed that the cauda epididymis exhibits significantly more severe inflammatory responses and leukocytic infiltrates than the caput and corpus epididymis36,37, which indicates the cauda epididymis shows stronger immune responses and susceptibility to inflammatory damage than other regions of the epididymis38, possibly because of its greater risk of infection while ascending along the urethra39. Importantly, the blood-testis and blood-epididymis barriers share common characteristics, and impaired blood-tissue barriers in the reproductive tract can lead to spermatogenesis disorders. Previous studies have proposed that sleep loss can increase the permeability of these barriers and decrease tight junction protein expression40. Although we demonstrated a reduction in the expression of genes related to proteins of the blood-epididymis barriers, our study focuses on immunoregulation and not on the integrity of the barriers as the cause of sperm alterations.
In recent years, scRNA-seq has been used to characterize cellular heterogeneity and gene expression patterns in epididymal tissues20. Building on the above experimental data, scRNA-seq was performed to evaluate cell-specific transcriptional changes and biological processes in the cauda epididymis of rats subjected to SD for five days. T cells and monocytes were subdivided into six clusters via uniform manifold approximation and projection (UMAP) analysis using well-defined markers. It is well-known that intraepithelial macrophages and cytotoxic lymphocytes, also known as CD8 + T cells, play a crucial role in preventing infections in the epididymis41. Monocytes can differentiate into macrophages upon recruitment into tissues. Our study identified significantly upregulated chemokine genes (Ccl2, Ccl4, Ccl5, Ccl7, Cxcl1, and Cxcl2) in macrophages after rats were subjected to SD, aligning with previous findings showing that continuous SD induced severe systemic inflammation in mammals13. As reported previously, certain chemokines exhibit chemotactic activity on monocytes42,43,44, while tissue macrophages synthesize CXCL1 and CXCL2 and then recruit neutrophils during tissue inflammation45. Macrophages, abundant in the interstitial area of the epididymis, are implicated in the phagocytosis of senescent and excess sperm41,46. Our findings demonstrate that macrophages within the cauda epididymis play a key role in mediating SD-induced inflammation and sperm phagocytosis. Meanwhile, we found that the SD treatment had significantly increased levels of granzymes expression in effector CD8 + T cells. Granzymes play an important role in promoting apoptosis in infected cells47,48. Furthermore, we found that Il-6r and Il-17ra were widely expressed in epididymal tissues. The IL-6/IL-6R signaling pathway activates the expression of different genes critical for inflammation, while the IL-17/IL-17R axis is essential for tissue homeostasis49,50. We also observed that SD significantly affected the communication of fibroblasts with other types of cells in the cauda epididymis. Tissue-resident fibroblasts are a crucial cell type in controlling the activation of an immune response51. Our analysis revealed that fibroblasts play an important role in the SD-induced epididymal immune response.
To further explore the impact of continuous SD on the male reproductive system, we analyzed region-specific epididymal RNA-seq data for rats in the recovery sleep group. PCA and Spearman’s rank correlation coefficient analysis showed no significant differences between the control and recovery sleep groups. Some reversible damage to the male reproductive system is hypothesized to have been induced by SD, according to our study. These findings are supported by a recent report showing that recovery sleep progressively restores the number of lymphocytes and monocytes to normal levels52. These findings could have therapeutic implications for shift workers and other populations frequently experiencing sleep deprivation. Evidence indicates that shift workers have higher white blood cell counts53, and chronic sleep deficiency is associated with an increased risk of reproductive health issues54.
In summary, our study demonstrated that SD triggers immune responses in the cauda epididymis, potentially contributing to low sperm quality. These findings highlight the importance of future studies on the spatial immune microenvironment of the epididymis and its role in sleep insufficiency-related male reproductive dysfunction.
Materials and methods
Animals
Ten-week-old specific pathogen-free male Sprague-Dawley rats were purchased from the animal research center at Zhejiang University. All animals were housed in the same room under specific pathogen-free conditions. Rats were randomly categorized into the control, SD-treated, and recovery groups. The body weights of the rats showed no significant differences between groups. The number of animals used was consciously minimized in this study (control group (n = 14), SD group (n = 12), recovery group (n = 5)). All rats were euthanized by CO2 asphyxiation using a slow displacement of chamber air (20–30%/min) from a CO2 gas tank with a gas regulator.
SD treatment
The SD treatment of rats lasted for five days and was performed using a modified multiple-platform method55. Briefly, rats were placed in a cage that contained platforms (6 cm in diameter, 1 cm above the water surface). The rats could move freely between the platforms with food and water available. The control rats without SD were placed into the same sleep restriction cage, which was not filled with water. Rats from the recovery group were subjected to five days of SD and then kept in a cage for twenty-eight days without performing interventions. Blinding is used during the conduct of the experiment and data analysis. There were no exclusions in this study.
H&E staining
Testicular and cauda epididymis tissues of control, SD-treated, and recovery rats were collected, fixed in 4% paraformaldehyde (P0099, Beyotime) for 24 h, embedded in paraffin wax, and sectioned at 3 µm. Sections were mounted onto glass microscope slides and stained with H&E (60524ES60, Yeasen) using standard procedures.
Histopathological study
Light microscopy was used to analyze the sections stained with H&E and capture images. Over two hundred distinct seminiferous tubules per testis were chosen for histological analysis to assess the seminiferous tubule epithelial height as well as the diameter of the seminiferous tubules and lumen.
TUNEL staining
Testicular and epididymal tissues were fixed in 4% paraformaldehyde, were then gradient dehydrated, and were embedded in optimal cutting temperature compound. The samples were sectioned and stored at −80 °C. The TUNEL Apoptosis Assay Kit (C1086, Beyotime) was used to perform TUNEL staining according to the manufacturer’s instructions. Images were obtained with an immunofluorescence microscope (Olympus, Tokyo, Japan). ImageJ software (US National Institutes of Health, Bethesda, MD, USA) was used to analyze and count the number of TUNEL-positive cells56.
Assessment of sperm parameter
The caudal epididymis was removed from the rats in each group, and epididymal tissues were incubated in Tyrode’s salt solution (T2397, Sigma-Aldrich) for 15 min at 37 °C. The incubation liquid was added to Tyrode’s solution and mixed gently for semen analysis. The sperm concentration and motility were measured using a CASA system (Hamilton Throne IVOS II, Lancaster, United States).
Daily sperm production and sperm transit time in the epididymis analysis
As previously reported, homogenization-resistant testicular spermatids and sperm in the caput-corpus and cauda epididymis were counted57. In order to calculate daily sperm production (DSP), the total number of homogenization-resistant spermatids per testis was divided by 6.1 days, which is the number of days that these spermatids are present in the seminiferous epithelium. The transit time via the caput-corpus epididymis or cauda epididymis was calculated by dividing the quantity of sperm in the caput-corpus epididymis or cauda epididymis by the DSP23.
Serum testosterone analysis
After the rats were euthanized by CO2 asphyxiation, the blood sample was collected and centrifuged for 15 min at 3000 rpm. Testosterone levels were measured by an enzyme-linked immunosorbent assay kit (MM-0577R1, FEIYUBIO) according to the manufacturer’s instructions.
RNA sequencing
Total RNA was isolated from the testis, caput epididymis, corpus epididymis, and cauda epididymis of rats using Trizol reagent (15596018, Invitrogen). RNA integrity was evaluated using 1% agarose gel electrophoresis, and RNA quality was assessed using the Agilent 2100 Bioanalyzer (Agilent Technologies, United States). The mRNA was enriched using Oligo (dT) beads, broken into short fragments, and reverse transcribed into cDNA using the NEBNext Ultra RNA Library Prep Kit for Illumin, according to the manufacturer’s instructions. Double-stranded cDNA fragments were ligated to Illumina sequencing adapters, amplified by PCR, and purified using AMPure XP beads. The cDNA library was then sequenced using an Illumina Novaseq 6000 by Gene Denovo Biotechnology Co. (Guangzhou, China). Clean reads were obtained using fastp (v0.23) with default parameters58 and mapped to the mRatBN7.2 (rn7) genome using hisat2 (v2.2.1). Samtools (v1.19.2)59 was used to calculate read coverage and depth. Stringtie (v2.2.1)60 was used to count the reads mapped to each gene, and transcripts per million mapped reads were calculated for each gene. DESeq2 (V1.42.1)61 was used to identify DEGs with a threshold of |log2FoldChange | >2 and padj < 0.01. DEGs were visualized using EnhancedVolcano (v1.20.0). Raw counts were variance-stabilized transformed using DESeq2, and variance-stabilized data were used to calculate Spearman’s correlation coefficients for each sample, with correlations visualized using pheatmap (v1.0.12).
Cell suspension preparation and scRNA-seq library construction
The cell suspension was prepared according to a standard procedure20. Briefly, tissues were washed three times using Hank’s balanced salt solution, minced, and digested. A 40 µm sterile strainer (352340, Corning Incorporated) was used to remove cell debris and impurities. Cells were collected by centrifugation at 500 g at 4 °C for 5 min, resuspended in PBS, and counted with a TC20 automated cell counter (Bio-Rad, United States). Cell suspensions were loaded onto a 10 × Genomics GemCode single-cell instrument to generate single-cell gel beads-In-EMlusion (GEMs). The gel beads in GEMs were dissolved to release barcode sequences, and barcoded full-length cDNAs were reverse transcribed and amplified via PCR for library construction. The purified scRNA-seq libraries were sequenced on an Illumina HiSeq X Ten system with 150 bp paired-end reads.
scRNA-seq data analysis
Raw sequencing data were converted to the fastq format using cellranger mkfastq (10x Genomics, v7.2.0). scRNA-seq reads were aligned to the mRatBN7.2 (rn7) reference genome and quantified using cellranger count. After collecting the digital gene expression count matrix, we performed quality control to the cells based on the distribution of genes detected and unique molecule identifiers, mitochondrial transcripts percent of each cell for all experiments. Further data processing was performed using the Seurat (v5.0.2) package62. Cells with detected gene counts between 200 and 6000 and mitochondrial gene transcripts ≤5% were retained, yielding 8,081 control and 8,071 cells of SD group. Raw counts were normalized using “LogNormalize” and the top 2000 highly variable genes were selected for dimensionality reduction and clustering. The harmony algorithm was applied to minimize batch effects and behavioral conditions during clustering. UMAP dimensionality reduction and clustering were performed using the built-in functions in Seurat, and cell clusters were manually annotated based on the characteristic genes in each cluster. Following the identification of major cell types in the epididymis, immune cell clusters were further isolated for additional UMAP and clustering analyses. DEGs between single-cell clusters were identified using the Wilcoxon test, with the threshold set at |log2(FoldChange)| > 1 and the P value from the Wilcoxon test <0.01. To compare overall gene expression levels between control and sleep-deprived groups across cell types, the PseudobulkExpression() method was used to compute pseudobulk expression levels for each cell cluster, followed by visualization using heatmaps.
Analysis of cell–cell communications
Differential cell-cell communication analysis was performed using the CellChat (v1.6.0) R package to investigate the differential communication patterns between different cell types in the cauda epididymis of SD and control groups.
Mendelian randomization analysis
Mendelian randomization was performed to infer causality using genetic variations. Robust genetic instruments were identified through extensive GWAS. Exposure data (GWAS-ID: GCST90042815) from 43,474 individuals with sleep disorders and 13,741 controls were sourced from the GWAS Catalog (https://gwas.mrcieu.ac.uk/). Outcome data (GWAS-ID: finngen_R10_N14_MALEINFERT) for 1429 individuals with male infertility and 130,139 controls were obtained from the FinnGen database (https://www.finngen.fi/en). The TwoSampleMR package of R software (version 4.3.1) was used to extract SNP data associated with sleep disorders as exposure factors, and the risk of male infertility as the outcome was analyzed. The link between sleep disorders and male infertility risk was evaluated using inverse variance weighted analysis. Additionally, sensitivity analysis was performed via a leave-one-out analysis.
Statistics and reproducibility
Statistical analyses were performed using GraphPad Prism 8 (GraphPad Software, California, USA). Statistical analysis was performed with an unpaired t-test. Data were presented as mean ± standard deviation values from at least three replicates. Significance was indicated as follows: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data were submitted to the National Center for Biotechnology Information and are accessible with BioProject ID: PRJNA1107966. The source data underlying the graphs in the manuscript can be found in Supplementary Data 1. All data of this study are available from the corresponding author upon reasonable request.
References
Hinz, A. et al. Sleep quality in the general population: psychometric properties of the Pittsburgh Sleep Quality Index, derived from a German community sample of 9284 people. Sleep Med. 30, 57–63 (2017).
Xu, Z. et al. Sleep quality of Chinese adolescents: distribution and its associated factors. J. Paediatr. Child Health 48, 138–145 (2012).
Alvaro, P. K., Roberts, R. M. & Harris, J. K. A systematic review assessing bidirectionality between sleep disturbances, anxiety, and depression. Sleep 36, 1059–1068 (2013).
Partinen, M. Sleep research in 2020: COVID-19-related sleep disorders. Lancet Neurol. 20, 15–17 (2021).
Roth, T. et al. Sleep problems, comorbid mental disorders, and role functioning in the National Comorbidity Survey Replication. Biol. Psychiatry 60, 1364–1371 (2006).
He, D., Guo, Z., Mcclure, M. A., Mu, Q. & Jiang, B. Cognitive-behavioral therapy for insomnia with objective short sleep duration phenotype: a systematic review with meta-analysis. Sleep. Med Rev. 67, 101736 (2023).
Cappuccio, F. P., D’Elia, L., Strazzullo, P. & Miller, M. A. Quantity and quality of sleep and incidence of type 2 diabetes: a systematic review and meta-analysis. Diab. Care 33, 414–420 (2010).
Malhotra, A. & Loscalzo, J. Sleep and cardiovascular disease: an overview. Prog. Cardiovasc Dis. 51, 279 (2009).
Ogilvie, R. P. & Patel, S. R. The epidemiology of sleep and obesity. Sleep health 3, 383–388 (2017).
Wang, Y. et al. Relationship between duration of sleep and hypertension in adults: a meta-analysis. J. Clin. Sleep. Med. 11, 1047–1056 (2015).
Vaou, O. E., Lin, S. H., Branson, C. & Auerbach, S. Sleep and dementia. Curr. Sleep. Med Rep. 4, 134–142 (2018).
Grandner, M. A., Hale, L., Moore, M. & Patel, N. P. Mortality associated with short sleep duration: the evidence, the possible mechanisms, and the future. Sleep Med. Rev. 14, 191–203 (2010).
Sang, D. et al. Prolonged sleep deprivation induces a cytokine-storm-like syndrome in mammals. Cell 186, 5500–5516.e5521 (2023).
Vaccaro, A. et al. Sleep loss can cause death through accumulation of reactive oxygen species in the gut. Cell 181, 1307–1328.e1315 (2020).
Kempf, A., Song, S. M., Talbot, C. B. & Miesenböck, G. A potassium channel β-subunit couples mitochondrial electron transport to sleep. Nature 568, 230–234 (2019).
Hamed, M. et al. Glutamine restores testicular glutathione-dependent antioxidant defense and upregulates NO/cGMP signaling in sleep deprivation-induced reproductive dysfunction in rats. Biomed. Pharmacother. 148, 112765 (2022).
Guo, H. et al. Cu-induced spermatogenesis disease is related to oxidative stress-mediated germ cell apoptosis and DNA damage. J. Hazard Mater. 416, 125903 (2021).
Wang, Y. et al. Cadmium exposure during puberty damages testicular development and spermatogenesis via ferroptosis caused by intracellular iron overload and oxidative stress in mice. Environ. Pollut. 325, 121434 (2023).
Cornwall, G. A. New insights into epididymal biology and function. Hum. Reprod. Update 15, 213–227 (2009).
Shi, J. et al. Spatio-temporal landscape of mouse epididymal cells and specific mitochondria-rich segments defined by large-scale single-cell RNA-seq. Cell Discov. 7, 34 (2021).
Liu, M.-M. et al. Sleep deprivation and late bedtime impair sperm health through increasing antisperm antibody production: a prospective study of 981 healthy men. Med. Sci. Monit. Int. Med. J. Exp. Clin. Res. 23, 1842 (2017).
Demirkol, M. K., Yıldırım, A., Gıca, Ş, Doğan, N. T. & Resim, S. Evaluation of the effect of shift working and sleep quality on semen parameters in men attending infertility clinic. Andrologia 53, e14116 (2021).
Fernandez, C. D. B., Porto, E. M., Arena, A. C. & Kempinas, W. D. G. Effects of altered epididymal sperm transit time on sperm quality. Int. J. Androl. 31, 427–437 (2008).
Chen, W. et al. Transcriptional alterations of genes related to fertility decline in male rats induced by chronic sleep restriction. Syst. Biol. Reprod. Med. 66, 99–111 (2020).
Mavanji, V., Teske, J. A., Billington, C. J. & Kotz, C. M. Partial sleep deprivation by environmental noise increases food intake and body weight in obesity-resistant rats. Obesity 21, 1396–1405 (2013).
Leproult, R. & Van Cauter, E. Role of sleep and sleep loss in hormonal release and metabolism. Endocr. Dev. 17, 11 (2009).
FO U., Alumana E. & Joshua P. Correlation of weight loss with infertility following sleep deprivation in albino rats. J. Exp. Res. 4 (2016).
Fajgenbaum, D. C. & June, C. H. Cytokine storm. N. Engl. J. Med. 383, 2255–2273 (2020).
Irwin, M. R., Carrillo, C. & Olmstead, R. Sleep loss activates cellular markers of inflammation: sex differences. Brain Behav. Immun. 24, 54–57 (2010).
Kelly, M., Gauthier, M.-S., Saha, A. K. & Ruderman, N. B. Activation of AMP-activated protein kinase by interleukin-6 in rat skeletal muscle: association with changes in cAMP, energy state, and endogenous fuel mobilization. Diabetes 58, 1953–1960 (2009).
Wijayarathna, R. & Hedger, M. Activins, follistatin and immunoregulation in the epididymis. Andrology 7, 703–711 (2019).
Kohno, S. et al. Immunopathology of murine experimental allergic orchitis. J. Immunol.130, 2675–2682 (1983).
Da Silva, N. et al. A dense network of dendritic cells populates the murine epididymis. Reprod.141, 653 (2011).
Voisin, A. et al. Comprehensive overview of murine epididymal mononuclear phagocytes and lymphocytes: unexpected populations arise. J. Reprod. Immunol. 126, 11–17 (2018).
Duan, Y. G. et al. Characterisation of dendritic cell subsets in chronically inflamed human epididymis. Andrologia 48, 431–440 (2016).
Klein, B. et al. Differential tissue-specific damage caused by bacterial epididymo-orchitis in the mouse. Mol. Hum. Reprod. 26, 215–227 (2020).
Michel, V. et al. Uropathogenic Escherichia coli causes fibrotic remodelling of the epididymis. J. Pathol. 240, 15–24 (2016).
Pleuger C., Silva E. J. R., Pilatz A., Bhushan S. & Meinhardt A. Differential immune response to infection and acute inflammation along the epididymis. Front. Immunol. 11, 599594 (2020).
Stammler, A. et al. Epididymitis: ascending infection restricted by segmental boundaries. Hum. Reprod. 30, 1557–1565 (2015).
Domínguez-Salazar, E. et al. Chronic sleep loss disrupts blood–testis and blood–epididymis barriers, and reduces male fertility. J. Sleep. Res 29, e12907 (2020).
Hedger, M. P. Immunophysiology and pathology of inflammation in the testis and epididymis. J. Androl. 32, 625–640 (2011).
Qian, B.-Z. et al. CCL2 recruits inflammatory monocytes to facilitate breast-tumour metastasis. Nature 475, 222–225 (2011).
Sindhu, S. et al. The cooperative induction of CCL4 in human monocytic cells by TNF-α and palmitate requires MyD88 and involves MAPK/NF-κB signaling pathways. Int. J. Mol. Sci. 20, 4658 (2019).
Shi, C. & Pamer, E. G. Monocyte recruitment during infection and inflammation. Nat. Rev. Immunol. 11, 762–774 (2011).
De Filippo, K. et al. Mast cell and macrophage chemokines CXCL1/CXCL2 control the early stage of neutrophil recruitment during tissue inflammation. Blood, J. Am. Soc. Hematol. 121, 4930–4937 (2013).
Gregory, M. & Cyr, D. G. The blood-epididymis barrier and inflammation. Spermatogenesis 4, e979619 (2014).
Fan, Z., Beresford, P. J., Oh, D. Y., Zhang, D. & Lieberman, J. Tumor suppressor NM23-H1 is a granzyme A-activated DNase during CTL-mediated apoptosis, and the nucleosome assembly protein SET is its inhibitor. Cell 112, 659–672 (2003).
Flemming, A. GZMK+ T cells a hallmark of immune ageing. Nat. Rev. Immunol. 21, 1–1 (2021).
Moseley, T., Haudenschild, D., Rose, L. & Reddi, A. Interleukin-17 family and IL-17 receptors. Cytokine Growth Factor Rev. 14, 155–174 (2003).
Azevedo, A., Cunha, V., Teixeira, A. L. & Medeiros, R. IL-6/IL-6R as a potential key signaling pathway in prostate cancer development. World J. Clin. Oncol. 2, 384 (2011).
Davidson, S. et al. Fibroblasts as immune regulators in infection, inflammation and cancer. Nat. Rev. Immunol. 21, 704–717 (2021).
Lasselin, J., Rehman, J. -u, Åkerstedt, T., Lekander, M. & Axelsson, J. Effect of long-term sleep restriction and subsequent recovery sleep on the diurnal rhythms of white blood cell subpopulations. Brain Behav. Immun. 47, 93–99 (2015).
Nishitani, N. & Sakakibara, H. Subjective poor sleep and white blood cell count in male Japanese workers. Ind. Health 45, 296–300 (2007).
Gibson, M. J., Lawlor, D. A. & Millard, L. A. Identifying the potential causal role of insomnia symptoms on 11,409 health-related outcomes: a phenome-wide Mendelian randomisation analysis in UK Biobank. BMC Med. 21, 128 (2023).
Machado, R. B., Hipólide, D. C., Benedito-Silva, A. A. & Tufik, S. Sleep deprivation induced by the modified multiple platform technique: quantification of sleep loss and recovery. Brain Res. 1004, 45–51 (2004).
Zhou, S. et al. UHRF1 interacts with snRNAs and regulates alternative splicing in mouse spermatogonial stem cells. Stem Cell Rep. 17, 1859–1873 (2022).
Robb, G., Amann, R. & Killian, G. Daily sperm production and epididymal sperm reserves of pubertal and adult rats. Reproduction 54, 103–107 (1978).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. Imeta 2, e107 (2023).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Shumate, A., Wong, B., Pertea, G. & Pertea, M. Improved transcriptome assembly using a hybrid of long and short reads with StringTie. PLoS Comp. Biol. 18, e1009730 (2022).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 1–21 (2014).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Acknowledgements
This study was supported by the Key Research and Development Program of Zhejiang Province (2023C03035), the Zhejiang Provincial Natural Science Foundation of China (Grant No. LQ24H050003), the National Key Research and Development Program of China (2024YFC2706800), the National Natural Science Foundation of China (82371613), the key Research and Development Program of Ningxia Hui Autonomous Region (2021BEG02029), the scientific research Foundation of Nantong Municipal Health Commission (QNZ2024060), the Nantong City Municipal Science and Technology Plan Guiding Projects (JCZ2024013), and the Nantong University Special Research Fund for Clinical Medicine (2024JQ042). We thank Chao Bi from the Core Facilities, Zhejiang University School of Medicine, for her technical support.
Author information
Authors and Affiliations
Contributions
F.S. and Z.L. conceived and supervised the project. Z.L. and S.G. designed and performed the experiments and analyzed experimental data. Z.Y. and Y.L. performed computational analysis. Y.Z., E.X., H.L., and D.X. assisted with the experiments, data analysis, and discussions. Z.L. wrote the manuscript. All authors contributed to and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All animal procedures were approved by the animal research ethics committee of Zhejiang University (No. ZJU20240220). We have complied with all relevant ethical regulations for animal use.
Peer review
Peer review information
Communications Biology thanks Wei Li and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Frank Avila and Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Z., Gong, S., Yu, Z. et al. Sleep deprivation impacts the immunological milieu of epididymis leading to low sperm quality in rats. Commun Biol 8, 644 (2025). https://doi.org/10.1038/s42003-025-08091-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08091-y