Abstract
Adipose tissue-derived mesenchymal stromal/stem cells (ASCs) possess regenerative potential. Obesity induces a pro-inflammatory environment that compromises their function. Here we investigate how obesity affects ASC biology, focusing on primary cilia. Our data show that obesity alters ASC gene expression, particularly in pathways related to the extracellular matrix, transforming growth factor-β (TGFβ) signaling, cell motility, and differentiation. The gene levels of regulatory factor X2 (RFX2) and adenylate cyclase 3 (ADCY3), important for ciliary biogenesis, are downregulated in obese ASCs. TGFβ treatment significantly decreases the expression of RFX2 and ADCY3 in lean ASCs. Knockdown of ADCY3 reduces primary cilium length, whereas pharmacological activation restores it and improves cell motility. These results suggest that obesity impairs ASC ciliary function, contributing to defective adipogenesis and reduced regenerative capacity. Restoring ADCY3 activity partially rescues ciliary integrity and cellular function, highlighting the role of primary cilia in ASC dysfunction and offering potential therapeutic targets for obesity-related disorders.
Similar content being viewed by others
Introduction
Adipose tissue (AT) has emerged as one of the most promising sources of mesenchymal stem cells (MSCs), owing to their abundance, ease of accessibility, low immunogenicity, regenerative potential, and relatively minimal ethical concerns1,2. ASCs, AT-derived MSCs, offer great promise for regenerative medicine. They can modulate the immune response, stimulate angiogenesis, and differentiate into multiple cell lineages to maintain or repair damaged tissues and organs, with subcutaneous AT being the most practical and clinically relevant source of ASCs2. Accordingly, they have been extensively studied and partially approved for therapeutic applications targeting different diseases3,4. Latest advances have enabled innovative strategies to reprogram human somatic cells into chemically induced pluripotent stem cells (hCiPSCs)5 enabling the first-in-human phase I clinical trial for the treatment of type 1 diabetes with autologous transplantation of hCiPSCs-islets derived from abdominal subcutaneous ASCs6.
ASCs influence the local and systematic environment through immune modulation and tissue homeostasis, mediated by direct cell-cell interactions and the secretion of multiple bioactive factors7,8. They secrete a variety of soluble and vesicular factors that remodel the extracellular matrix (ECM), a process crucial for their own function and regulation9,10,11. The ECM, in turn, interacts with cells through transmembrane receptors that relay external cues and activate intracellular signaling pathways involving the actin cytoskeleton12. ASCs detect environmental stimuli through surface receptors, notably via a specialized, immotile sensory organelle called the primary cilium13,14,15. The primary cilium mediates diverse extracellular signals, acting as a transduction platform for a multitude of pathways, regulating proliferation, migration, and differentiation, and maintaining tissue homeostasis, growth, and development16,17,18. Ciliary signaling relies on various surface receptors within its membrane, including G protein-coupled receptors (GPCRs), along with their downstream effectors and second messengers13,19.
The dramatic rise in obesity prevalence presents a significant global challenge, as it is linked to various diseases, including cardiovascular diseases, type 2 diabetes, multiple types of cancer, and an overall higher mortality rate20,21. It is well established that obesity causes changes in ASC’s multipotency, proliferation, differentiation, and migration capacity, as well as energy metabolism by reducing their functionalities7,22, suggesting that the health status of the donor should be considered to ensure the safe use of ASCs. Epigenetic changes of ASCs from individuals with obesity23 as well as an obesogenic epigenetic memory function of the AT were reported24. Moreover, we have studied ASCs derived from subcutaneous, visceral, and mammary AT from donors with obesity14,25,26,27,28. We have shown that obesity has adverse effects on subcutaneous and visceral ASCs, which are defective in ciliary biogenesis and Hedgehog signaling14,15,26. However, the molecular mechanisms by which ASCs are dysfunctional and their cilia are defective in obesity are not well investigated. Given that reactivation of ciliated cells and ciliary functions may offer a strategy to combat obesity and its related diseases15, this study further characterizes ASCs from subcutaneous AT of donors with obesity. We performed whole-transcriptome analysis to identify deregulated genes and signaling pathways, with a focus on the functional integrity of primary cilia.
Results
Obesity leads to transcriptional reprogramming of ASCs
To investigate the effects of obesity on subcutaneous ASCs in more detail, total RNA was extracted from five individual ASC samples per group: lean (BMI < 25, ASCsub25), obese 35 (BMI ≥ 35, ASCsub35), and obese 40 (BMI ≥ 40, ASCsub40), and subjected to whole-genome RNA sequencing (RNA-seq). The Venn diagram (Fig. 1A) highlights overlapping and unique genes of the three groups. The volcano plots (Fig. 1B) show significantly deregulated genes (p < 0.05). Compared to ASCsub25, ASCsub35 showed 738 significantly up- and 767 downregulated genes (Fig. 1B, left panel), whereas ASCsub40 demonstrated 811 significantly up- and 768 downregulated genes (Fig. 1B, right panel). Gene Ontology (GO) terms are categorized into three main ontological domains: biological process (BP), cellular component (CC), and molecular function (MF). Interestingly, our analysis revealed that obesity significantly altered the “signaling receptor regulator activity” of the MF, “ECM” of the CC, and “ECM structure organization” of the BP in both ASCsub35 and ASCsub40 populations (Fig. 1C, D). In the ASCsub35 group, four GO terms were directly associated with the ECM. To better illustrate the extent of gene overlap among these ECM-related GO terms (Fig. S1A), we used the web-based tool InteractiVenn29. The ASCsub40 group also enriched ECM-related GO terms, along with additional differentially expressed genes related to cell-matrix adhesion and response to transforming growth factor-β (TGFβ) (Fig. 1D and Fig. S1B). Moreover, the “Kyoto Encyclopedia of Genes and Genomes” (KEGG) pathway analysis highlighted the most significantly upregulated genes in both groups of obese ASCs, specifically, in ECM-receptor interaction, cell adhesion molecules (CAMs), regulation of the actin cytoskeleton, and focal adhesion (FA), compared to lean ASCs (Fig. 1E, F).
Total RNAs were extracted from each ASC sample of individual donors with a body mass index (BMI) below 25 (ASCsub25, n = 5), equal or above 35 (ASCsub35, n = 5), and equal or above 40 (ASCsub40, n = 5) for transcriptomic analysis. A Venn diagram of co-expressed genes. B Volcano plots of significantly differentially expressed genes of ASCsub35 vs. ASCsub25 (left plot) and ASCsub40 vs. ASCsub25 (right plot). Upregulated genes are depicted in red, downregulated genes in green, and non‑changed genes in blue. p < 0.05. Significantly enriched “Gene Ontology” (GO) pathways are presented for ASCsub35 compared to ASCsub25 (C) and for ASCsub40 compared to ASCsub25 (D), sub-clustered into three ontological domains: molecular function (MF), cellular component (CC), and biological process (BP). The adjusted p value (padj) is displayed by a color code. Significantly enriched “Kyoto Encyclopedia of Genes and Genomes” (KEGG) pathways (except for PI3K-Akt ASCsub40 vs. ASCsub25) are presented for ASCsub35 compared to ASCsub25 (E) and for ASCsub40 compared to ASCsub25 (F). The p value is indicated by a color code. Abbreviations: neg. negative, PI3K-Akt phosphoinositide 3-kinase - protein kinase B, TGFβ transforming growth factor-β.
Altered ECM genes in obese ASCs
Obesity-induced AT expansion requires ECM remodeling, which is associated with hypoxia and inflammation30. These conditions, in turn, upregulate ECM-related gene expression, particularly collagens, leading to its excessive accumulation and increased rigidity in obese AT30. ECM-related genes that are differentially expressed in both obese groups (ASCsub35 and ASCsub40), compared to ASCsub25, are depicted in Fig. S1C. Notably, they abundantly expressed various collagen genes (COL), including COL11A1, COL12A1, COL4A6, and COL5A2 (Fig. S1D). In addition, the expression of crucial ECM regulator genes was enhanced (Fig. S1E), such as ADAMTS12 (a disintegrin and metallopeptidase with thrombospondin type 1 motif 12), HMCN1 (hemicentin 1), MMP16 (matrix metallopeptidase 16), and HAPLN1 (hyaluronan and proteoglycan link protein 1) (Fig. S1E). Importantly, ANOS1 (anosmin-1), a TGFβ-regulated gene involved in endothelial cell migration and angiogenesis31, GAS6 (growth arrest-specific 6), which is related to cell adhesion, obesity, insulin resistance, and inflammation32, and WISP1 (WNT1-inducible signaling pathway protein 1), which is an adipokine associated with obesity, insulin resistance, and fibrosis, and a downstream target of TGFβ33, were highly expressed in obese ASCs (Fig. S1F). The enhanced expression of COL11A1, COL5A2, and GAS6 was corroborated with RT-PCR (Fig. S1G). These results highlight that ECM-related genes, particularly regulated by TGFβ, are deregulated in obese ASCs, which could lead to alterations of the ECM composition, structure, regulation, and function. This may, in turn, negatively affect the properties and functionalities of ASCs.
Deregulated genes of the transforming growth factor β pathway in ASCs derived from donors with obesity
It is well established that TGFβ is increased during obesity34,35. This signaling pathway is crucial for regulating cell growth, differentiation, apoptosis, and migration36. To address if the TGFβ pathway is altered in our obese ASC cohorts, we further examined its related genes in RNA-seq data (Fig. 2A, B). Interestingly, three genes were enhanced expressed in both obese groups, CDKN2B (cyclin-dependent kinase inhibitor 2B, p15), ITGB5 (integrin beta 5), and THBS1 (thrombospondin 1) (Fig. 2A–C). Notably, ITGB537 and THBS138 are both able to activate the latent TGFβ1. In comparison with ASCsub25, ASCsub35 cells highly expressed several genes promoting the TGFβ pathway, such as LPXN (leupaxin), LTBP1, encoding the latent TGFβ binding protein 1, participating in TGFβ1 activation via integrins39, and the TGFβ1-activating transcription factor gene SOX9, encoding SRY-box transcription factor 9 (Fig. 2D). In contrast, TGFBR3 (TGFβ receptor 3, betaglycan), the most abundantly expressed TGFβ-receptor lacking kinase activity, which enhances TGFβ signaling and mediates interactions with other proteins40, however, was reduced in ASC35sub cells (Fig. 2E). Similarly, TAB1 (TGFβ activated kinase 1 binding protein 1) and CLEC3B (C-type lectin domain family 3 member B, tetranectin) were also significantly downregulated in ASCsub35 (Fig. 2E). Looking more closely at the ASCsub40 group, we observed further altered genes (Fig. 2B). In particular, ADAM17 (a disintegrin and metallopeptidase 17), which promotes epithelial to mesenchymal transition and fibrosis via TGFβ signaling41, was increased. Additionally, ASPN (asporin), an important protein in the ECM linked to collagen fibrillogenesis42, and LTBP3 (latent TGFβ binding protein 3) that can bind fibrillin and TGFβ39, also showed increased expression. Unexpectedly, SMAD3 (mothers against decapentaplegic homolog 3), a prominent downstream target of TGFβ, was downregulated (Fig. 2F). These results strongly indicate that regulators and downstream effectors of TGFβ signaling are deregulated in obese ASCs. Since KEGG pathway analysis revealed an upregulation of actin cytoskeleton regulation and focal adhesion, and considering that TGFβ can influence cytoskeletal dynamics by regulating actin filament organization43, we next focused on the altered genes within motility and migration pathways.
The heatmaps depict significantly (p < 0.05) differentially expressed transforming growth factor-β (TGFβ) genes with the gene ontology terms GO:0030511, 0030512, and 0071560 of ASCsub35 vs. ASCsub 25 (A) and of ASCsub40 vs. ASCsub 25 (B). Significantly enriched genes have a red color code and reduced genes are encoded in blue. Red and blue colored genes are sorted alphabetically. C–F Truncated violin plots present selected genes that are expressed differentially. Values reflect the fragments per kilobase per million mapped fragments (fpkm) of genes from the RNA‑seq data. Each violin plot displays the median (central dashed line) and quartiles (upper and lower dotted lines).
Motility and migration genes are affected by obesity
ASCs have been reported to be able to migrate into injured areas, differentiate, and repair damaged tissue2. Taking into account that donor obesity may affect their therapeutic potential7, we took a closer look at the differentially expressed genes related to motility and migration, which are significantly altered in both groups (ASCsub35 and ASCsub40) compared to ASCsub25 (Fig. 3A). Intriguingly, the genes CDH1 (cadherin 1 or E-cadherin, an epithelial cell marker) and CDH2 (N-cadherin, a mesenchymal cell marker) were increasingly expressed (Fig. 3B). Cadherins, transmembrane glycoproteins, play a major role in tissue homeostasis, cell-cell adhesion, and epithelial to mesenchymal transition, where E-cadherin expression is commonly lost44. Increased CDH1 could result from hypoxia, which is a hallmark of AT in obesity15, whereas enhanced CDH2, could be induced by various signaling pathways, including TGFβ44. FN1 (fibronectin) and LRP12 (low-density lipoprotein receptor related protein 12) were also significantly increased in ASCs derived from donors with obesity (Fig. 3C). Fibronectin is a major ECM glycoprotein involved in cell migration, tissue repair, and adipogenesis, and it interacts with several cell surface receptors, linked to integrin and focal adhesion dynamics12,45. Enhanced fibronectin may result in excessive accumulation and rigidity of ECM elements in obese AT30. LRP12, an endogenous transmembrane inhibitor for integrin activation, binds to the cytoplasmic tail of integrin α4 (ITG4A), and disturbs the association of talin (integrin-actin linker) with integrin β1 or β7, keeping integrin in its inactive state46. Besides LRP12, ITG4A is also enhanced in the transcriptomic data (Fig. 3A), both could contribute to the reduced migration capacity. The genes that were downregulated included CXCL12 (CXC motif chemokine 12), IFITM1 (interferon induced transmembrane protein 1), and PGF (placental growth factor) (Fig. 3D). Interestingly, the reduced expression of CXCL12 is contributed to impaired migration of ASCs47. Additional RT-PCR analyses were performed and validated an upregulation of CDH1, and downregulation of CXCL12 as well as PGF in ASCs from donors with obesity (Fig. 3E). These data provide an explanation why obese ASCs are hampered in motility and migration capability7,14.
A The heatmap shows significantly (p < 0.05) differentially expressed motility and migration genes with the gene ontology terms GO:0016477, 0048870, and 200145 in both ASC subgroups of donors with obesity. Significantly enriched genes have a red color code and reduced genes are encoded in blue. Red and blue colored genes are sorted alphabetically. B–D Truncated violin plots present selected motility and migration genes that are differentially expressed. Values reflect the fragments per kilobase per million mapped fragments (fpkm) of genes from the RNA‑seq data. Each violin plot displays the median (central dashed line) and quartiles (upper and lower dotted lines). E Relative gene levels of CDH1 (cadherin 1), CXCL12 (C-X-C motif chemokine ligand 12), and PGF (placental growth factor) are shown for lean and obese ASC subgroups (ln25, ob35 and ob40). The results were obtained from five individual samples in each group and presented as relative quantification (RQ) with SEM. GAPDH served as endogenous control. Ordinary one-way ANOVA followed by Dunnett’s multiple comparisons test was used to assess statistical significance relative to ln25 (CDH1). For non-Gaussian data (CXCL12), the Kruskal-Wallis test followed by Dunn’s post hoc test was applied. If variance homogeneity was not met (PGF), Welch’s ANOVA with Dunnett’s T3 multiple comparisons test was used. *p < 0.05.
Deregulated genes of ciliary biogenesis in obese ASCs
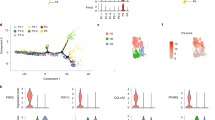
The primary cilium is one of the most important mediators of extracellular signals and the cell has several pathways through which the length of the primary cilium can be modulated48. We have reported that primary cilia are crucial for various functionalities of ASCs, including their motility and migration, and that obesity shortens cilia rendering them dysfunctional14,15. To further underline that observation, we stained ASCs from ASCsub25, ASCsub35, and ASCsub40 for the ciliary markers acetylated tubulin (ace tubulin, green) and ARL13B (ADP-ribosylation factor-like GTPase 13B, red) (Fig. 4A), and measured the cilium length. The analysis showed that primary cilia of ASCsub35 as well as of ASCsub40 were shortened (Fig. 4B). We used CiliaCarta to identify known ciliary genes49 within the transcriptomic data. Cilium-related genes were significantly altered in both groups (Fig. 4C), and even more in ASCsub35 (Fig. 4D). In particular, the gene expression of GLI1 (glioma-associated oncogene homolog 1), which encodes an important regulator of the Hedgehog signaling pathway that is exclusively transduced through the primary cilium50, and MYH10, encoding the myosin heavy chain 10, required for centriole migration during cilium biogenesis51, was enhanced (Fig. 4E), whereas the genes PKIG (protein kinase inhibitor gamma), a protein kinase A inhibitor leading to increased intracellular cAMP levels52, and SNX10, encoding the sorting nexin 10, involved in the regulation of the vesicular trafficking and primary cilia membrane extension53, were downregulated (Fig. 4F). Compared to ASCsub25, ASCsub35 showed significant higher gene levels of MYO7A (myosin 7 A) and HHIP (Hedgehog interacting protein), genes linked to ciliopathies54 and Hedgehog signaling55 (Fig. 4G). Of particular interest, the gene levels of two crucial ciliary regulators, adenylate cyclase 3 (ADCY3), and clusterin associated protein 1 (CLUAP1 or IFT38, intraflagellar transport protein 38) were reduced in obese ASCs (Fig. 4H). Further RT-PCR analysis underlined that GLI1 was enhanced, whereas SNX10, ADCY3, and CLUAP1 were decreased in obese ASCs (Fig. 4I).
A Subcutaneous ASCs (ASCsub) were stained for the ciliary markers acetylated α-tubulin (ace tubulin, green) and ARL13B (ADP-ribosylation factor-like GTPase 13B, red), and DNA (DAPI, blue). Representatives are shown. Scale: 5 μm. B Cilium lengths were measured and are presented as scatter dot plots (mean ± SD). The results are based on at least six individual samples per group (ASCsub25, n = 6 individual individual samples, 180 cilia; ASCsub35, n = 6, 178 cilia; and ASCsub40, n = 7, 208 cilia) and statistically analyzed. C, D The heatmaps depict significantly (p < 0.05) differentially expressed genes from CiliaCarta49 in both ASC subgroups of donors with obesity, or in ASCsub35 vs. ASCsub 25, respectively. Significantly enriched genes have a red color code and reduced genes are encoded in blue. Red and blue colored genes are sorted alphabetically. E–H Truncated violin plots present selected ciliary genes, which are differentially expressed. Values reflect the fragments per kilobase per million mapped fragments (fpkm) of genes from the RNA‑seq data. Each violin plot displays the median (central dashed line) and quartiles (upper and lower dotted lines). I Relative gene levels of GLI1 (glioma-associated oncogene homolog 1), SNX10 (sorting nexin 10), ADCY3 (adenylate cyclase 3), and CLUAP1 (clusterin associated protein 1) were corroborated with RT-PCR. The results are obtained from five individual samples in each group and presented as relative quantification (RQ) with SEM. GAPDH was used as endogenous control. Ordinary one-way ANOVA followed by Dunnett’s multiple comparisons test was used to assess statistical significance relative to ln25 (I). For non-Gaussian data (B, I, GLI1), the Kruskal-Wallis test followed by Dunn’s post hoc test was applied. *p < 0.05, ***p < 0.001.
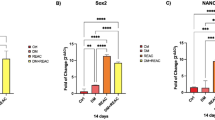
Reduced adipogenic differentiation of obese ASCs
Functional cilia are important for adipogenic differentiation of MSCs and ASCs56,57 and as previously described by our group, shortened cilia of ASCs from obese donors were unable to respond dynamically to osteogenic differentiation stimuli14. To examine the capability of adipogenic differentiation, ASCs derived from lean donors (Fig. S2B, 1st and 2nd panels) or from donors with obesity (3rd and 4th panels) were incubated without or with adipogenic differentiation medium for 14 days for further analyses (Fig. S2A). Treated ASCs were stained with the lipid droplet dye BODIPYTM, the cytoskeleton marker phalloidin, and the DNA dye DAPI for evaluating adipocytes. In comparison with control ASCs (Fig. S2B, 2nd panels), obese ASCs showed much less adipogenic differentiation capacity upon stimulation (Fig. S2B, 4th panels). These results were further underscored by brightfield images and Oil Red O staining (Fig. S2C, D). Evaluation revealed that only 16.99% of ASCs derived from donors with obesity were differentiated, compared to 56.48% of ASCs form lean donors (Fig. S2E). Moreover, total RNAs from these ASCs were extracted for gene analysis. Compared to lean ASCs, the expression of key adipogenic differentiation genes58,59, including the early differentiation gene PPARG (peroxisome proliferator-activated receptor gamma), and its late effectors, ADIPOQ (adiponectin), FABP4 (fatty acid-binding protein 4), and LPL (lipoprotein lipase), was significantly reduced (Fig. S2F). To meet the tri-lineage differentiation criteria defined by the International Society for Cellular Therapy (ISCT)60, we also performed osteogenic and chondrogenic differentiation for 21 days. Both differentiation capacities were reduced in ASCs derived from donors with obesity, as indicated by decreased expression of the master transcription factors RUNX2 (runt-related transcription factor 2) for osteogenesis and SOX9 for chondrogenesis (Fig. S2G, H). These data underscore the observation that obesity impairs the differentiation capability of ASCs, as we have reported14,28.
Decreased ADCY3 in ASCs derived from donors with obesity
After systematically analyzing transcriptomic data, characterizing deregulated genes and signaling pathways, and specifying corresponding defective functionalities, we paid high attention to the two crucial regulators of ciliary biogenesis, CLUAP1/IFT38 and ADCY3. CLUAP1/IFT38, a key component of the IFT-B peripheral subcomplex, is essential for cilium assembly and maintenance61. Cluap1-deficient mice were not viable, and Cluap−/− mouse embryonic fibroblasts failed to form primary cilia and were defective in Hedgehog signaling62,63. To examine this regulator in more detail, we stained ASCs with the CLUAP1 antibody, acetylated α-tubulin, and DAPI (Fig. 5A) for further analysis. Microscopic analysis revealed that CLUAP1 localized to the base and the tip of the cilia, as reported63. Its protein intensity along the primary cilium was further evaluated via a line-scan analysis14,64. This evaluation found that the protein levels of axonemal CLUAP1 were reduced in obese ASCs at the base (Fig. 5A, B). Moreover, ADCY3, a transmembrane enzyme, is regulated by various GPCRs and catalyzes the synthesis of cAMP, a critical intracellular second messenger in signaling pathways that broadly impact cellular and ciliary functions65,66. Intriguingly, ADCY3 was enriched at primary cilia of ASCs, which was more obvious at ASCs derived from lean donors (Fig. 5C). Further line-scan analysis showed that ADCY3 protein levels in the ciliary region were significantly reduced along the entire cilium in obese ASCs (Fig. 5D), suggesting that diminished ADCY3 may contribute to cilium shortening in these cells. To test the functional involvement of ADCY3 in regulating ciliary length, lean ASCs were transfected with either control siRNA (sicon) or siRNA against ADCY3. Knockdown efficiency was confirmed by RT-PCR (Fig. 5G). Further analysis revealed that ADCY3 knockdown significantly reduced ciliary length (Fig. 5E, F), reaching levels comparable to those observed in ASCs from donors with obesity (Fig. 4B). These findings strengthen the involvement of ADCY3 in maintaining the length of primary cilia.
A Cells were stained for the ciliary markers clusterin associated protein 1 (CLUAP1, red) and acetylated α-tubulin (ace tubulin, green), and DNA (DAPI, blue). Representatives are shown for measurements of primary cilium staining of CLUAP1. Scale: 5 µm. B Line-scan analyses of axonemal CLUAP1 are shown for ASCs. Each point of the curve represents the mean fluorescence intensity ±SD and the results are based on three individual samples (n = 3, 73 cilia), normalized to the cilium length in percentage; a.u., arbitrary unit. C Cells were stained for the ciliary markers ADCY3 (adenylate cyclase 3, red) and acetylated α-tubulin (ace tubulin, green), and DNA (DAPI, blue). Representatives are shown for measurements of the primary cilium staining of ADCY3. Scale: 5 µm. D Line-scan analyses of axonemal ADCY3 are shown for ASCs. Each point of the curve represents the mean fluorescence intensity ±SD and the results are based on three individual samples (n = 3, 72 cilia), normalized to the cilium length (in percent); a.u., arbitrary unit. Statistical significance was assessed using the Kruskal-Wallis test followed by Dunn’s post hoc test to compare lean and obese groups at each 10% interval of relative ciliary length (0–100%). *p < 0.05, **p < 0.01, ***p < 0.001; ns, not significant. E–G Lean ASCs were treated with 30 nM of control siRNA (sicon) or with siRNA against ADCY3 for 48 h. E Primary cilia were stained for the ciliary marker acetylated α-tubulin (ace tubulin, green) and ARL13B (ADP-ribosylation factor-like GTPase 13B, red), and DNA (DAPI, blue). Representatives are shown. Scale: 5 µm. F The lengths of individual cilia were measured and are presented as scatter dot plots (mean ± SD). The results are based on three individual samples per group (n = 3, 135 cilia) and statistically analyzed. G Relative gene levels of ADCY3 are shown. The results were obtained from three individual samples in each group and presented as relative quantification (RQ) with SEM. GAPDH served as endogenous control. Unpaired Mann–Whitney U test (F) or Student’s t test was used (G). *p < 0.05, ***p < 0.001.
TGFβ drives ciliary shortening and gene suppression similar to the obese phenotype
We further investigated potential mechanisms underlying impaired ciliation in ASCs from individuals with obesity. Members of the regulatory factor X (RFX) family are among the most well-established transcriptional regulators of primary cilium biogenesis and regulate target genes such as IFT17267,68. Based on the data from transcriptomic profiling, ASCs express the transcription factor genes RFX1, RFX2, and RFX3 (Fig. 6A). Although not significantly altered in the transcriptome data, RFX2 emerged as a promising candidate due to its significantly reduced expression in obese ASCs, as demonstrated by RT-PCR (Fig. 6B).
A Truncated violin plots of the members of the regulatory factor X (RFX) family (RFX1, RFX2, and RFX3). Values reflect the fragments per kilobase per million mapped fragments (fpkm) of genes from the RNA‑seq data. Each violin plot displays the median (central dashed line) and quartiles (upper and lower dotted lines). B Relative gene levels of RFX2 were corroborated with RT-PCR. The results are obtained from five individual samples for the lean and four individual samples of the obesity group, and presented as relative quantification (RQ) with SEM. GAPDH was used as endogenous control. C–G Treatment of lean ASCsub with TGFβ for 48 h. C Primary cilia of lean ASCsub were stained for the ciliary markers acetylated α-tubulin (ace tubulin, green) and ARL13B (ADP-ribosylation factor-like GTPase 13B, red), and DNA (DAPI, blue). Representatives are shown. Scale: 5 μm. D Cilium lengths were measured and are presented as scatter dot plots (mean ± SD). The results are based of three individual samples per group. E–G Relative gene levels of RFX2, ADCY3 (adenylate cyclase 3), IFT88 (intraflagellar transport), IFT172, COL1A1 (collagen 1A1), and SERPINE1 (serpin family E member 1) are shown. The results were obtained from three individual samples in each group and presented as relative quantification (RQ) with SEM. GAPDH served as endogenous control. Statistical significance was analyzed with the Student’s t test. *p < 0.05, ***p < 0.001.
To examine whether the elevated TGFβ signaling observed in obese ASCs contributes to ciliary shortening and reduced expression of ciliary genes, we treated lean ASCsub cells with TGFβ for 48 h. This treatment led to a measurable reduction in ciliary length (Fig. 6C, D). In parallel, RT-PCR analysis showed that TGFβ significantly downregulated the expression of the ciliary transcription factor gene RFX2, the enzyme ADCY3, and two intraflagellar transport genes, IFT88 and IFT172, which are essential for cilium assembly (Fig. 6E, F). To affirm treatment efficiency, we also assessed the expression of established TGFβ target genes, COL1A1 and SERPINE1, which were significantly upregulated, confirming pathway activation (Fig. 6G). These findings indicate that TGFβ may promote ciliary shortening in obesity partly through the suppression of critical ciliary regulator genes, including RFX2, which may in turn lead to reduced expression of ADCY3, though further work is required to decipher how TGFβ reduces the expression of these important ciliary regulators.
Modulation of adenylate cyclase activity affects cilium length and ASC motility
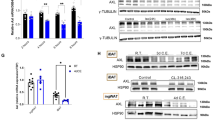
To further investigate the role of ADCY3 and cAMP signaling in ciliary regulation, ASCs from lean donors were treated with the ADCY inhibitor SQ22536, which blocks cAMP signaling69, whereas ASCs from donors with obesity were incubated with an ADCY activator NKH477 (colforsin dapropate hydrochloride), a water-soluble forskolin derivate, leading to an increase in cAMP levels70. The effect of both compounds was proven by intracellular cAMP measurements via ELISA (Fig. 7A, B). To test the hypothesis that ADCY3 activity or cAMP availability affects primary cilium length in ASCs, we performed immunofluorescence staining with the ciliary markers acetylated α-tubulin and ARL13B to assess changes in cilium length (Fig. 7C, F). Primary cilia in ASCs, derived from lean donors and treated with two different concentrations of SQ22536 for 24 and 48 h, became shorter (Fig. 7D, E). Instead, ASCs derived from donors with a BMI above 35 and treated with NKH477 displayed up to 24% increase in cilium length (Fig. 7G, H). As these treatments did not influence cell viability (Fig. S3A), the findings strongly suggest that the deregulated ADCY3/cAMP signaling pathway contributes to shorter cilia in obese ASCs. As controls, lean ASCs were also treated with NKH477, and obese ASCs with SQ22536. Neither treatment affected cell viability (Fig. S3B). While activation of ADCY3 with NKH477 increased slightly the cilium length in lean ASCs (8%, Fig. S3C), inhibition with SQ22536 hardly changed it in obese ASCs (Fig. S3D).
A, B The effect of the ADCY inhibitor SQ22536 (SQ) or its activator NKH477 (NKH) was proven by the measurement of intracellular concentrations of acetylated cAMP via a direct cAMP ELISA. The results are presented as mean ± SD. Student’s t test was used. * p < 0.05. ASCs were treated with the ADCY inhibitor SQ22536 (SQ) (C–E) or its activator NKH477 (NKH) (F–H) for 24 or 48 h with indicated concentrations. Primary cilia of ASCsub were stained for the ciliary markers acetylated α-tubulin (ace tubulin, green) and ARL13B (ADP-ribosylation factor-like GTPase 13B, red), and DNA (DAPI, blue). Representatives are shown. Scale: 5 μm (C, F). Cilium lengths were measured and are presented as scatter dot plots (mean ± SD) (D, E and G, H). The results are based on three individual samples per group (n = 3, 150 cilia) and statistical significance was analyzed with the Kruskal-Wallis test followed by Dunn’s post hoc test. ***p < 0.001. Abbreviations: ADCY3 adenylate cyclase 3, cAMP cyclic adenosine monophosphate, con control, DMSO dimethyl sulfoxide, ELISA enzyme-linked immunosorbent assay, min minutes.
The motility of MSCs/ASCs is vital for the development, maintenance, and repair of body-wide tissues71, and the primary cilium is involved in cell motility and migration72. To further investigate the restoration of the cilium length on the migratory ability of ASCs, time-lapse microscopy with single cell tracking was performed with lean ASCs in absence or presence of the ADCY inhibitor SQ22536, and with obese ASCs without or with its activator NKH477 (Fig. 8A, B). The velocity of untreated obese ASCs (ob con) was significantly decreased compared to lean ASCs (ln DMSO) (Fig. 8C, 1st vs. 3rd row). Relative to lean ASCs, obese ASCs had a reduced velocity of 44% (0.322 ± 0.082 vs. 0.181 ± 0.064 μm/min), which was raised by 54% following the stimulation with NKH477 (0.278 ± 0.080 μm/min) (Fig. 8C, 4th row, ob NKH). As a further control, lean ASCs treated with the ADCY inhibitor SQ22536 (ln SQ) showed a reduction of 34% (0.212 ± 0.066 μm/min) (Fig. 8C, 2nd row), compared to its untreated control (ln DMSO). Similar results were observed for the accumulated distance, with 215 ± 55 for lean con, 121 ± 43 μm for obese con, 185 ± 52 μm for obASCs treated with NKH477, and 142 ± 44 μm for lnASCs with SQ22536 (Fig. 8D). The directionality was significantly increased by 10% in obASCs upon NKH477 treatment (Fig. 8E). As controls, treatment of lean ASCs with NKH477 resulted in only a slight, but non-significant, improvement in velocity and accumulated distance, whereas obese ASCs treated with SQ22536 showed a slight, yet non-significant, reduction in these parameters (Fig. S3E–H). These data highlight that reduced ADCY3 is responsible, at least partially, for shortened cilia and that its activation rescues the cilium length in obese ASCs and, consequently, the motility of obese ASCs.
Analysis of cell motility. A Experimental schedule of treatment with the ADCY inhibitor SQ22536 (SQ) or its activator NKH477 (NKH). B Representative trajectories are depicted for individual cells (n = 1, 50 cells in each group). The velocity (C), the accumulated distance (D), and the directionality (E) of ASC motility are shown as mean ± SD (n = 3, 150 cells pooled from three experiments). Statistical significance was analyzed with the Kruskal-Wallis test followed by Dunn’s multiple comparison test. *p < 0.05; ***p < 0.001; n.s., not significant. F Schematic illustration of the proposed working model. Obesity-associated factors, such as TGFβ, are able to shortened primary cilia in ASCs associated with deregulated pathways and genes/proteins, fueling inflammation and metabolic dysfunction and facilitating the progression of obesity. Treatment with NHK477 reactivates the ADCY3-cAMP signaling axis, prolongs the primary cilium, improves cell motility, and thus possibly prevents the progression of obesity. Abbreviations: ADCY3 adenylate cyclase 3, ADIPOQ adiponectin, cAMP cyclic adenosine monophosphate, CDH1 cadherin-1, COL collagen, CXCL12 CXC motif chemokine 12, DMSO dimethyl sulfoxide, FABP4 fatty acid-binding protein 4, FN1 fibronectin, HH hedgehog, ITGB5 integrin beta 5, ln lean, LPL lipoprotein lipase, ob obese, PGF placental growth factor, PPARG peroxisome proliferator-activated receptor gamma, TGFβ transforming growth factor-β, THBS1 thrombospondin 1.
Discussion
We have previously reported that ASCs are impaired in various functions and properties during obesity14,15. The present study deepens the molecular understanding of the extensive transcriptomic and functional alterations of ASCs from donors with obesity. In particular, we show that ASCs isolated from obese donors exhibit an altered ECM and TGFβ gene profile, along with abnormal ciliary biogenesis and shortened primary cilia, impaired differentiation capacity, and reduced cell motility.
Interestingly, the observed altered pathways and phenotypes are intriguingly linked. Enhanced TGFβ release is well characterized in obesity34,35. In addition, obesity-associated ECM stiffness is thought to promote F-actin assembly, which, in turn, facilitates TGFβ release, activating TGFβ signaling43. On the other hand, actin polymerization and stress fiber formation are known to impair ciliogenesis73. Notably, TGFβ signaling inhibits adipogenesis by suppression of PPARγ74. Moreover, ciliary receptors are also critical for the activation of adipogenesis in preadipocytes75 and functional cilia are necessary for adipogenic and osteogenic differentiation of MSCs and ASCs14,56,57, whereas inhibition of ciliary elongation by IFT88 siRNA reduced the adipogenic differentiation capacity56. Functional ASCs require properly regulated ECM and ciliary biogenesis9,15. Notably, obesity changes the composition, structure, and function of the ECM of AT, which has been suggested to play a major role in defining the severity of obesity76. Therefore, the intertwined relationship between defective ECM, enhanced TGFβ signaling, and altered ciliary biogenesis could lead to impaired differentiation capacity of obese ASCs. In support of these observations, our results show that obesity leads to transcriptional alterations of ASCs, particularly genes involved in various aspects of the ECM and actin cytoskeleton. Both ASCsub35 and ASCsub40 abundantly express various collagen genes (COL). Interestingly, altered collagens contribute to AT dysfunction and/or metabolic diseases and are often highly expressed in obese tissue30. Increased expression of COL4A1, COL5A2, and COL12A1 was also found in the transcriptomic analysis of subcutaneous AT of obese individuals77. The expressed collagens shifted from collagen types I, III, and V to the types IV and VI during the differentiation of 3T3-L1 preadipocytes into adipocytes30,78. The downregulation of FN-integrin α5 interactions has also been reported to be necessary for adipogenesis12. In fact, we show here an increased expression of FN1 in obese ASCs with impaired adipocyte differentiation capability. Furthermore, the primary cilium and ciliary cAMP play a critical role in directing the cell fate of adipocyte progenitor cells in multiple AT types79. The cilium length of human ASCs/MSCs dynamically changed during adipogenic differentiation, likely due to ciliary trafficking of various receptors, while blocking this elongation impaired adipogenic induction56,57. These data underline the importance of the primary cilium in the adipogenic capacity and cell fate of ASCs.
Ciliopathies are often accompanied by obesity, and associated with fibrosis, ECM accumulation, and enhanced expression of TGFβ, suggesting that the impairment of primary cilia may be involved in the pathogenesis of obesity80,81. Obesity-associated factors shortens its length and impairs its functionalities in ASCs14. However, the molecular mechanisms are not entirely defined. It has been described that RFXs are well-established transcription factors of ciliary biogenesis67,68. We report here that RFX2 is significantly downregulated in ASCs from donors with obesity, which may lead to reduced expression of ADCY3. Importantly, the treatment of lean ASCs with TGFβ reduces ciliary length as well as the gene expression of the transcription factor RFX2, its downstream genes ADCY3, and IFT88 and IFT172, which are essential for cilium assembly. In line with our data, various components of the TGFβ signaling machinery have been reported to be present at the primary cilium82. In further support, TGFβ treatment resulted in reduced ciliary length by upregulating histone deacetylase 6 activity in human osteoblasts83, or suppressed the levels of IFT88 mRNA and protein of murine chondrocytes84. These data clearly demonstrate that TGFβ is vital for ciliary biogenesis. Therefore, obesity-associated increase in TGFβ contribute to impaired primary cilia formation in obese ASCs, thereby compromising their functional properties.
ADCY3 is downregulated in obese ASCs. It has been revealed that three types of adenylate cyclase localize to the primary cilium (ADCY3, ADCY5, and ADCY6)85. These enzymes catalyze the synthesis of the second messenger cAMP, which is a central player in ciliary signaling86. ADCY3 is localized at primary cilia of neurons87, primary epithelial kidney cells88, human fibroblasts or fibroblast-like synoviocytes89,90, and MLO-Y4 osteocyte-like cells91. Distinct human ADCY3 genetic variants have been linked to the pathogenesis of monogenic and polygenic obesity as well as type 2 diabetes92. Moreover, mice heterozygous and homozygous null for Adcy3 exhibited increased susceptibility to obesity associated with insulin or leptin resistance93,94. Interestingly, the inhibition of Adcy3 in neurons, and consequently ciliary signaling, increased the body weight of mice95. In line with these observations, we report here that ADCY3 is localized at primary cilia of ASCs, its abundance is significantly reduced throughout the primary cilium in obese ASCs, and siRNA against ADCY3 reduced the cilium length of lean ASCs to a level comparable to that in obese ASCs. Altogether, our data clearly suggest that reduced RFX2 and ADCY3 could contribute to defective cilia of obese ASCs. These findings also highlight that the TGFβ-RFX-ADCY network is of importance in ciliary biogenesis.
Interestingly, it has been supposed that the stimulation of the ADCY3 activity may be a useful treatment approach for obesity65. We demonstrate here that treatment with the ADCY3 activator NKH477 restores the cilium length of ASCs derived from donors with obesity, whereas reducing cAMP levels by inhibiting ADCY3 results in shorter cilia in lean ASCs. In support of our data, using optogenetic tools, Hansen et al. showed that an increase in ciliary cAMP lengthened the primary cilium of the mouse kidney cell line IMCD396. In addition, Ansari et al. demonstrated that reduced ciliary cAMP production or Adcy3 knockout led to primary cilium shortening in IMCD3 cells, which was restored after treatment with the cAMP analog dibutyryl-cAMP97. Our data, in combination with reported results, highlight that the ADCY3-cAMP signaling axis is a central component of ciliary signaling, which is disturbed in ASCs derived from donors with obesity.
The primary cilium is involved in the control of cell motility80,98, in neuronal cells as well as non-neuronal, including endothelial cells, fibroblasts, and MSCs66,99, especially ASCs14,26. Primary cilium signaling can influence the migratory response by affecting cytoskeletal reorganization, cell polarity, adhesion dynamics, directionality, and speed98. Studies show that the primary cilium senses and responds to ECM changes and that its formation, orientation, and length dynamics may act as a switch for regulating cellular responsiveness during migration98. Interestingly, genetic ablation of the primary cilium, or knockdown of Adcy3 caused migratory defects of neurons with reduced migratory speed99. We show here that ASCs derived from donors with obesity exhibit shorter cilia, reduced levels of RFX2 and ADCY3, and impaired cell migration. These defects can be partially remedied by targeting ADCY3 and enhancing intracellular cAMP levels. Treatment with an ADCY activator improves the velocity, accumulated distance, and directionality of impaired ASCs from donors with obesity.
Conclusions
This study highlights that obesity induces transcriptional reprogramming of ASCs, leading to alterations in ECM composition and ciliary biogenesis. The impairment of primary cilia in obese ASCs, particularly through downregulation of key regulators like ADCY3, probably through the transcription factor RFX2 that is regulated by TGFβ, may contribute to defective differentiation capacity and cell motility, potentially fueling the progression of obesity (Fig. 8F). Moreover, the reduced cAMP signaling in obese ASCs, which can be partially restored through the activation of the ADCY3-cAMP signaling axis, further underscores the critical role of primary cilia in regulating ASC function. Although further investigation is needed, these findings suggest that obesity-induced changes in ECM, TGFβ signaling, ciliary biogenesis, and cAMP signaling may impair ASC differentiation and migration, highlighting potential therapeutic targets for obesity-related disorders.
Materials and methods
Ethics approval and consent to participate
This work was approved by the Ethics Committee of the Johann Wolfgang-Goethe University Hospital Frankfurt (reference number: 375/11, title: Comparison of metabolic pathways in normal and preeclampsia pregnancies, approved newly on 23th November 2021) and informed written consent was obtained from all donors. The Ethics Committee acts in accordance with the Helsinki Declaration. All ethical regulations relevant to human research participants were followed.
ASC isolation, treatment, transfection, and differentiation
Subcutaneous AT samples were taken from women undergoing cesarean section. Previous studies have shown that the reproductive state has minimal impact on the differentiation capacity of ASCs100. All participants showed no evidence of impaired glucose tolerance, as confirmed by the 50 g oral glucose tolerance test (oGTT50) and, if indicated, an oGTT75 or hemoglobin A1c. Participant information is listed in Table 1.
ASCs were isolated as reported25,101. The isolated cells were resuspended in DMEM with pyruvate, 4.5 g/L D-glucose and L-glutamine (Gibco, Carlsbad) containing 20% fetal bovine serum, 1% penicillin/streptomycin (Sigma-Aldrich), and 1 µg/ml amphotericin-B (Sigma-Aldrich), and cultured under standard cell culture conditions. All ASCs were routinely tested for mycoplasma (Minerva Biolabs GmbH).
ASCs were treated with the ADCY inhibitor SQ22536 (sc-201572, Santa Cruz Biotechnology, Inc., Heidelberg), dissolved in dimethyl sulfoxide (DMSO, Thermo Fisher Scientific), the ADCY activator NKH477 (sc-204130, Santa Cruz), or TGFβ (#100-21-2UG, PeproTech). Concentrations and time points are indicated in each experiment. To match the maximal concentration of SQ22536, DMSO was used at a final concentration of 0.05% (v/v).
ASCs were transfected with 30 nM of control small interfering RNA (AllStars Negative Control siRNA, Qiagen, Hilden, #1027281) or siRNA targeting ADCY3 (Santa Cruz Biotechnology, Inc., sc-29600) using the 4D-NucleofectorTM Core Unit with X Unit (AmaxaTM P1 Primary Cell 4d- NucleofectorTM Kit, #V4XP-102) with the program DO-101 (Lonza, Maastricht).
Adipogenic differentiation of ASCs was performed as reported25. Briefly, ASCs were cultured with optimized differentiation medium for the generation of adipocytes (StemMACSTM AdipoDiff Media, #130-091-677, Miltenyi Biotec, Gladbach) up to 14 days. Cells were fixed and stained with BODIPYTM 493/503 (D3922, Invitrogen) and phalloidin (Phalloidin-Atto 550, Sigma-Aldrich), or with Oil Red O (Sigma-Aldrich) and hematoxylin (Sigma-Aldrich) to analyze adipogenic differentiation. For osteogenic and chondrogenic differentiation, ASCs were incubated with StemMACSTM OsteoDiff Media (#130-091-678) or ChondroDiff Media (#130-091-679, Miltenyi Biotec) for 21 days28.
Surface marker measurement
The cell surface markers of ASCs were measured via FACS with FACSCaliburTM (BD Biosciences, Heidelberg). Cells were trypsinized with 0.25% trypsin (Sigma-Aldrich), harvested, and fixed for 15 min with ice-cold 2% paraformaldehyde. After washing twice with PBS, the cells were stained with the following antibodies: PE-conjugated anti-human cluster of differentiation 44 (CD44) (#130-113-342, clone REA690, lot 5240507442, MACS Miltenyi Biotec, Bergisch Gladbach), PE-conjugated anti-human CD73 (#550257, lot 4149735, BD-Pharmingen, Heidelberg), FITC-CD90 (#11-0909-42, clone eBio5E10, lot 2648893, eBioscience, Frankfurt), PE-conjugated anti-human CD105 (#323206, clone 43A3, lot B418585, BioLegend, San Diego), PerCP-Cy5.5-conjugated anti-human CD14 (#45-0149-42, clone 61D3, lot 2785933, eBioscience), APC-conjugated anti-human CD31 (#17-0319-42, clone WM-59, lot 2702043, eBioscience), and FITC-conjugated anti-human CD34 (#343504, clone 581, lot B407007, eBioscience). Non-stained ASCs were used as negative controls. All ASCs were characterized for the positive (CD44, CD73, CD90, CD105) and the negative markers (CD14, CD31, CD34), as listed in Table 2. An example for the gating strategy is provided in Supplementary Information (Fig. S4). The passages P3 to P8 of isolated ASCs were used for experiments.
RNA extraction, transcriptomic analysis (RNA-seq), and real-time PCR (RT-PCR)
For transcriptomic analysis, total RNAs were extracted from each ASC sample (cells at P4) of five different donors per condition with EXTRACTME TOTAL RNA Kit (#EM09.2.-250, 7Bioscience, Neuenburg). A total amount of 1.5 μg RNA per sample was forwarded to Novogene Co., Ltd. (London) and used as starting material for RNA sample preparation28. The Venn diagram, volcano plots, violin plots, and pathway analyses were either generated by Novogene Co. Ltd. or by re-analyzing the data using Microsoft®Excel® (Redmond, Washington) and GraphPad Prism 8 or 10 (GraphPad software Inc., San Diego). The Revigo tool102 was used to reduce redundancy among GO term analyses, where a selection of significantly enriched pathways are shown. Additional Venn diagrams (Fig. S1A, B) were performed with the web-based tool InteractiVenn29. CiliaCarta was used to identify known ciliary genes49.
For RT-PCR, reverse transcription was done using GoScriptTM Reverse Transcription Mix, Random Primers (#A2801, Promega, Madison). The probes for ADCY3 (Hs01086502_m1), ADIPOQ (Hs00605917_m1), CDH1 (Hs01023895_m1), CLUAP1 (Hs01032009_m1), COL1A1 (Hs00164004_m1), COL11A1 (Hs01097664_m1), COL5A2 (Hs00893878_m1), CXCL12 (Hs00171022_m1), FABP4 (Hs01086177_m1), GAPDH (Hs02758991_g1), GAS6 (Hs01090305_m1), GLI1 (Hs00171790_m1), IFT88 (Hs00901755_m1), IFT172 (Hs00404485_m1), LPL (Hs00173425_m1), PGF (Hs00182176_m1), PPARG (Hs01115513_m1), RFX2 (Hs01100925_m1), RUNX2 (Hs01047973_m1), SERPINE1 (Hs00167155_m1), SNX10 (Hs00203362_m1), and SOX9 (Hs00165814_m1) were from Thermo Fisher Scientific (Dreieich). RT-PCR was performed with a QuantStudio 3 (Applied Biosystems by Thermo Fisher Scientific). All results were shown as relative quantification (RQ)103,104. RQ values were calculated using the ΔΔCt method. Mean and standard error of the mean (SEM) were calculated from biological replicates based on the individual RQ values (n = 3 to 5).
Indirect immunofluorescence staining, microcopy, and signal intensity measurement
Indirect immunofluorescence staining and microscopic evaluation were performed105,106. Cells were fixed with 4% paraformaldehyde containing 0.2% Triton X-100 for 15 min at room temperature. To evaluate the cilium length, and the relative intensity of ciliary proteins, ASCs were stained with mouse monoclonal antibody targeting acetylated α-tubulin (#T6793, clone 6-11B-1, lot 0000298764, Sigma-Aldrich), rabbit polyclonal antibody against ARL13B (#17711-1-1AP, lot 00142161, Proteintech, Herford), rabbit polyclonal ADCY3 (#19492-1-AP, lot 00046273, Proteintech), and rabbit polyclonal CLUAP1 (#17470-1-AP, lot 00025316, Proteintech). FITC- and Cy3-conjugated secondary antibodies (Jackson ImmunoResearch, Pennsylvania) and DAPI (4’, 6-diamidino-2- phenylindole-dihydrochloride, Roche, Mannheim) were used. Confocal laser scanning microscopy (CLSM, Leica, Wetzlar) was conducted using a Leica CTR 6500 system, employing Z-stack imaging with a HCXPI APO CS 63.0×/1.4 oil-immersion objective. Z-stacks were acquired at 0.5 µm intervals. Representative images were generated by overlaying individual confocal Z-sections utilizing maximum intensity projection. All experiments, unless otherwise indicated, were independently performed with ASCs isolated from at least three different donors. Relative fluorescence intensities of ADCY3 and CLUAP1 were measured using line-scan-based analysis with ImageJ and the ImageJ plugin Plot Roi Profile (National Institutes of Health, Bethesda)14,64. The mean values of 72 or 73 cilia from three individual ASC samples in each group were obtained within the intervals and were plotted to GraphPad Prism 8 (GraphPad software Inc.).
cAMP ELISA
Cells were treated with 100 µM of SQ22536 or 50 nM of NKH477 for 30 min, respectively. cAMP (3’,5’-cyclic adenosine monophosphate) was extracted from ASCs using 0.1 M HCl for 20 min. The intracellular concentrations of acetylated cAMP in the lysates were then determined using a direct cAMP ELISA kit (#ADI-901-066A, Enzo Life Sciences GmbH, Lörrach).
Cell viability assay
Cell viability was assessed using the CellTiter-Blue® assay (#G808B, Promega, Walldorf) following the manufacturer’s protocol.
Cell motility evaluation via time-lapse microscopy
Cells were seeded into 24-well plates with a low confluency (30-40%) and were imaged at 5-min time intervals for 13 h. Time-lapse imaging was performed with an AxioObserver.Z1 microscope (Zeiss, Göttingen, Germany), imaged with an AxioCam MRc camera (Zeiss, Göttingen, Germany) provided with an environmental chamber to maintain proper conditions (37 °C, 5% CO2)101. Time-lapse movies were analyzed using ImageJ 1.49i software (National Institutes of Health) with the manual tracking plugin. Tracks were derived from raw data points, and the velocity, accumulated distance, and directionality were calculated using the Chemotaxis and Migration Tool (Ibidi GmbH, Gräfelfing), and depicted by GraphPad Prism 8 (GraphPad software Inc.). Fifty random cells per experiment were analyzed and independently performed with ASCs isolated from three individual samples.
Statistics and reproducibility
Outliers were identified using Grubbs’ test (GraphPad QuickCalcs, San Diego). Statistical analyses were performed using GraphPad Prism 8 (GraphPad software Inc.). The normality of data distribution was assessed using the Shapiro-Wilk test. For comparisons between two groups, an unpaired Student’s t test was applied for normally distributed data, and the Mann–Whitney U test for non-normally distributed data. To compare three or more groups, one-way ANOVA was performed, followed by multiple comparisons tests: Tukey’s (to compare all groups), Dunnett’s (to compare all groups versus a control), or Sidak’s (to compare selected groups). If the assumption of homogeneity of variances was violated (Brown-Forsythe test), Welch’s ANOVA followed by Dunnett’s T3 multiple comparisons test was used. For non-normally distributed data, the Kruskal-Wallis test followed by Dunn’s multiple comparisons test was applied. A p < 0.05 was considered statistically significant. All experiments were performed using ASCs from independent donors, with n representing the number of individual patients from whom cells were isolated. Unless otherwise specified, experiments were conducted with n = 3. For transcriptomic analysis, ASCs from n = 5 individual samples were used. Exact n numbers are indicated in the corresponding figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA sequencing data have been deposited in the NCBI Gene Expression Omnibus (GEO) under accession number GSE308101. The source data behind the graphs in the paper are included in the Supplementary Data 1 file. All other data supporting the findings of this study are available from the corresponding author upon request.
References
Mazini, L., Ezzoubi, M. & Malka, G. Overview of current adipose-derived stem cell (ADSCs) processing involved in therapeutic advancements: flow chart and regulation updates before and after COVID-19. Stem Cell Res. Ther. 12, 1 (2021).
Si, Z. et al. Adipose-derived stem cells: sources, potency, and implications for regenerative therapies. Biomed. Pharmacother. 114, 108765 (2019).
Jovic, D. et al. A brief overview of global trends in MSC-based cell therapy. Stem Cell Rev. Rep. 18, 1525–1545 (2022).
Fernandez-Garza, L. E., Barrera-Barrera, S. A. & Barrera-Saldana, H. A. Mesenchymal stem cell therapies approved by regulatory agencies around the World. Pharmaceuticals 16, https://doi.org/10.3390/ph16091334 (2023).
Liuyang, S. et al. Highly efficient and rapid generation of human pluripotent stem cells by chemical reprogramming. Cell Stem Cell 30, 450–459.e459 (2023).
Wang, S. et al. Transplantation of chemically induced pluripotent stem-cell-derived islets under abdominal anterior rectus sheath in a type 1 diabetes patient. Cell 187, 6152–6164.e6118 (2024).
Louwen, F., Ritter, A., Kreis, N. N. & Yuan, J. Insight into the development of obesity: functional alterations of adipose-derived mesenchymal stem cells. Obes. Rev. https://doi.org/10.1111/obr.12679 (2018).
Al-Ghadban, S. & Bunnell, B. A. Adipose tissue-derived stem cells: immunomodulatory effects and therapeutic potential. Physiology 35, 125–133 (2020).
Yen, B. L. et al. Three-dimensional spheroid culture of human mesenchymal stem cells: offering therapeutic advantages and in vitro glimpses of the in vivo state. Stem Cells Transl. Med. 12, 235–244 (2023).
Ritter, A. et al. Subcutaneous and visceral adipose-derived mesenchymal stem cells: commonality and diversity. Cells 8, https://doi.org/10.3390/cells8101288 (2019).
Trigo, C. M., Rodrigues, J. S., Camoes, S. P., Sola, S. & Miranda, J. P. Mesenchymal stem cell secretome for regenerative medicine: where do we stand? J. Adv. Res. https://doi.org/10.1016/j.jare.2024.05.004 (2024).
Longstreth, J. H. & Wang, K. The role of fibronectin in mediating cell migration. Am. J. Physiol. Cell Physiol. 326, C1212–C1225 (2024).
Hilgendorf, K. I., Myers, B. R. & Reiter, J. F. Emerging mechanistic understanding of cilia function in cellular signalling. Nat. Rev. Mol. Cell Biol. 25, 555–573 (2024).
Ritter, A. et al. Primary cilia are dysfunctional in obese adipose-derived mesenchymal stem cells. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2017.12.022 (2018).
Ritter, A., Louwen, F. & Yuan, J. Deficient primary cilia in obese adipose-derived mesenchymal stem cells: obesity, a secondary ciliopathy? Obes. Rev. 19, 1317–1328 (2018).
Malicki, J. J. & Johnson, C. A. The cilium: cellular antenna and central processing unit. Trends Cell Biol. 27, 126–140 (2017).
Anvarian, Z., Mykytyn, K., Mukhopadhyay, S., Pedersen, L. B. & Christensen, S. T. Cellular signalling by primary cilia in development, organ function and disease. Nat. Rev. Nephrol. 15, 199–219 (2019).
Gopalakrishnan, J. et al. Emerging principles of primary cilia dynamics in controlling tissue organization and function. EMBO J. 42, e113891 (2023).
Derderian, C., Canales, G. I. & Reiter, J. F. Seriously cilia: a tiny organelle illuminates evolution, disease, and intercellular communication. Dev. Cell 58, 1333–1349 (2023).
Aune, D. et al. Body mass index, abdominal fatness, fat mass and the risk of atrial fibrillation: a systematic review and dose-response meta-analysis of prospective studies. Eur. J. Epidemiol. 32, 181–192 (2017).
Bhaskaran, K., Dos-Santos-Silva, I., Leon, D. A., Douglas, I. J. & Smeeth, L. Association of BMI with overall and cause-specific mortality: a population-based cohort study of 3.6 million adults in the UK. Lancet Diabetes Endocrinol. 6, 944–953 (2018).
Yang, H., Li, C., Li, Y., Tai, R. & Sun, C. Adipose-derived stem cells and obesity: the spear and shield relationship. Genes Dis. 10, 175–186 (2023).
Eirin, A. et al. Obesity-driven mitochondrial dysfunction in human adipose tissue-derived mesenchymal stem/stromal cells involves epigenetic changes. Cell Death Dis. 15, 387 (2024).
Hinte, L. C. et al. Adipose tissue retains an epigenetic memory of obesity after weight loss. Nature https://doi.org/10.1038/s41586-024-08165-7 (2024).
Ritter, A. et al. Characterization of adipose-derived stem cells from subcutaneous and visceral adipose tissues and their function in breast cancer cells. Oncotarget 6, 34475–34493 (2015).
Ritter, A. et al. Restoration of primary cilia in obese adipose-derived mesenchymal stem cells by inhibiting Aurora A or extracellular signal-regulated kinase. Stem Cell Res. Ther. 10, 255 (2019).
Ritter, A. et al. Adipose tissue-derived mesenchymal stromal/stem cells, obesity and the tumor microenvironment of breast cancer. Cancers 14, https://doi.org/10.3390/cancers14163908 (2022).
Ritter, A. et al. Cancer-educated mammary adipose tissue-derived stromal/stem cells in obesity and breast cancer: spatial regulation and function. J. Exp. Clin. Cancer Res. 42, 35 (2023).
Heberle, H., Meirelles, G. V., da Silva, F. R., Telles, G. P. & Minghim, R. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 16, 169 (2015).
Jaaskelainen, I. et al. Collagens regulating adipose tissue formation and functions. Biomedicines 11, :biomedicines11051412 [pii];biomedicines-11-01412 [pii]; https://doi.org/10.3390/biomedicines11051412 (2023).
Matsushima, S. et al. Anosmin-1 activates vascular endothelial growth factor receptor and its related signaling pathway for olfactory bulb angiogenesis. Sci. Rep. 10, 188 (2020).
Su, S. C. et al. Growth arrest-specific 6 modulates adiponectin expression and insulin resistance in adipose tissue. J. Diabetes Investig. 12, 485–492 (2021).
Singh, K. & Oladipupo, S. S. An overview of CCN4 (WISP1) role in human diseases. J. Transl. Med. 22, 601 (2024).
Ghaben, A. L. & Scherer, P. E. Adipogenesis and metabolic health. Nat. Rev. Mol. Cell Biol. 20, 242–258 (2019).
Fain, J. N., Tichansky, D. S. & Madan, A. K. Transforming growth factor beta1 release by human adipose tissue is enhanced in obesity. Metabolism 54, 1546–1551 (2005).
Deng, Z. et al. TGF-beta signaling in health, disease, and therapeutics. Signal. Transduct. Target Ther. 9, 61 (2024).
Munger, J. S. & Sheppard, D. Cross talk among TGF-beta signaling pathways, integrins, and the extracellular matrix. Cold Spring Harb. Perspect. Biol. 3, a005017 (2011).
Gutierrez, L. S. & Gutierrez, J. Thrombospondin 1 in metabolic diseases. Front. Endocrinol. 12, 638536 (2021).
Robertson, I. B. et al. Latent TGF-beta-binding proteins. Matrix Biol. 47, 44–53 (2015).
Vander Ark, A., Cao, J. & Li, X. TGF-beta receptors: In and beyond TGF-beta signaling. Cell Signal 52, 112–120 (2018).
Sisto, M., Ribatti, D. & Lisi, S. ADAM 17 and epithelial-to-mesenchymal transition: the evolving story and its link to fibrosis and cancer. J. Clin. Med. 10, https://doi.org/10.3390/jcm10153373 (2021).
Fan, R., Yan, X. & Zhang, W. Relationship between asporin and extracellular matrix behavior: a literature review. Medicine 101, e32490 (2022).
Melchionna, R., Trono, P., Tocci, A. & Nistico, P. Actin cytoskeleton and regulation of TGFbeta signaling: exploring their links. Biomolecules 11, https://doi.org/10.3390/biom11020336 (2021).
Kaszak, I. et al. Role of cadherins in cancer-a review. Int. J. Mol. Sci. 21, https://doi.org/10.3390/ijms21207624 (2020).
Naba, A. Mechanisms of assembly and remodelling of the extracellular matrix. Nat. Rev. Mol. Cell Biol. 25, 865–885 (2024).
Huang, M. et al. LRP12 is an endogenous transmembrane inactivator of alpha4 integrins. Cell Rep. 42, 112667 (2023).
Kato, T. et al. SDF-1 improves wound healing ability of glucocorticoid-treated adipose tissue-derived mesenchymal stem cells. Biochem. Biophys. Res. Commun. 493, 1010–1017 (2017).
Macarelli, V., Leventea, E. & Merkle, F. T. Regulation of the length of neuronal primary cilia and its potential effects on signalling. Trends Cell Biol. 33, 979–990 (2023).
van Dam, T. J. P. et al. CiliaCarta: an integrated and validated compendium of ciliary genes. PLoS ONE 14, e0216705 (2019).
Jing, J. et al. Hedgehog signaling in tissue homeostasis, cancers, and targeted therapies. Signal. Transduct. Target Ther. 8, 315 (2023).
Hong, H., Kim, J. & Kim, J. Myosin heavy chain 10 (MYH10) is required for centriole migration during the biogenesis of primary cilia. Biochem. Biophys. Res. Commun. 461, 180–185 (2015).
Hoy, J. J., Salinas Parra, N., Park, J., Kuhn, S. & Iglesias-Bartolome, R. Protein kinase A inhibitor proteins (PKIs) divert GPCR-Galphas-cAMP signaling toward EPAC and ERK activation and are involved in tumor growth. FASEB J. 34, 13900–13917 (2020).
Chen, Y. et al. A SNX10/V-ATPase pathway regulates ciliogenesis in vitro and in vivo. Cell Res. 22, 333–345 (2012).
Wolfrum, U., Liu, X., Schmitt, A., Udovichenko, I. P. & Williams, D. S. Myosin VIIa as a common component of cilia and microvilli. Cell Motil. Cytoskeleton 40, 261–271 (1998).
Bishop, B. et al. Structural insights into hedgehog ligand sequestration by the human hedgehog-interacting protein HHIP. Nat. Struct. Mol. Biol. 16, 698–703 (2009).
Dalbay, M. T., Thorpe, S. D., Connelly, J. T., Chapple, J. P. & Knight, M. M. Adipogenic differentiation of hMSCs is mediated by recruitment of IGF-1r onto the primary cilium associated with cilia elongation. Stem Cells 33, 1952–1961 (2015).
Forcioli-Conti, N., Lacas-Gervais, S., Dani, C. & Peraldi, P. The primary cilium undergoes dynamic size modifications during adipocyte differentiation of human adipose stem cells. Biochem. Biophys. Res. Commun. 458, 117–122 (2015).
Khine, H. E. E., Sungthong, R., Sritularak, B., Prompetchara, E. & Chaotham, C. Untapped pharmaceutical potential of 4,5,4’-trihydroxy-3,3’-dimethoxybibenzyl for regulating obesity: a cell-based study with a focus on terminal differentiation in adipogenesis. J. Nat. Prod. 85, 1591–1602 (2022).
Kamble, P. G. et al. Proof-of-concept for CRISPR/Cas9 gene editing in human preadipocytes: deletion of FKBP5 and PPARG and effects on adipocyte differentiation and metabolism. Sci. Rep. 10, 10565 (2020).
Dominici, M. et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 8, 315–317 (2006).
Katoh, Y. et al. Overall architecture of the intraflagellar transport (IFT)-B complex containing Cluap1/IFT38 as an essential component of the IFT-B peripheral subcomplex. J. Biol. Chem. 291, 10962–10975 (2016).
Lamri, L. et al. Ciliogenesis-coupled accumulation of IFT-B proteins in a novel cytoplasmic compartment. Genes Cells 24, 731–745 (2019).
Botilde, Y. et al. Cluap1 localizes preferentially to the base and tip of cilia and is required for ciliogenesis in the mouse embryo. Dev. Biol. 381, 203–212 (2013).
He, M. et al. The kinesin-4 protein Kif7 regulates mammalian Hedgehog signalling by organizing the cilium tip compartment. Nat. Cell Biol. 16, 663–672 (2014).
Wu, L., Shen, C., Seed, A. M., Ostenson, C. G. & Gu, H. F. Adenylate cyclase 3: a new target for anti-obesity drug development. Obes. Rev. 17, 907–914 (2016).
Stoufflet, J. & Caille, I. The primary cilium and neuronal migration. Cells 11, https://doi.org/10.3390/cells11213384 (2022).
Choksi, S. P., Lauter, G., Swoboda, P. & Roy, S. Switching on cilia: transcriptional networks regulating ciliogenesis. Development 141, 1427–1441 (2014).
Chung, M. I. et al. RFX2 is broadly required for ciliogenesis during vertebrate development. Dev. Biol. 363, 155–165 (2012).
Emery, A. C., Eiden, M. V. & Eiden, L. E. A new site and mechanism of action for the widely used adenylate cyclase inhibitor SQ22,536. Mol. Pharm. 83, 95–105 (2013).
Pavan, B., Biondi, C. & Dalpiaz, A. Adenylyl cyclases as innovative therapeutic goals. Drug Discov. Today 14, 982–991 (2009).
Labour, M. N., Riffault, M., Christensen, S. T. & Hoey, D. A. TGFbeta1 - induced recruitment of human bone mesenchymal stem cells is mediated by the primary cilium in a SMAD3-dependent manner. Sci. Rep. 6, 35542 (2016).
Collinson, R. & Tanos, B. Primary cilia and cancer: a tale of many faces. Oncogene 44, 1551–1566 (2025).
Smith, C. E. L., Lake, A. V. R. & Johnson, C. A. Primary cilia, ciliogenesis and the actin cytoskeleton: a little less resorption, a little more actin please. Front. Cell Dev. Biol. 8, 622822 (2020).
Lee, M. J. Transforming growth factor beta superfamily regulation of adipose tissue biology in obesity. Biochim. Biophys. Acta Mol. Basis Dis. 1864, 1160–1171 (2018).
Hilgendorf, K. I. et al. Omega-3 fatty acids activate ciliary FFAR4 to control adipogenesis. Cell 179, 1289–1305 e1221 (2019).
Lawler, H. M. et al. Adipose tissue hypoxia, inflammation, and fibrosis in obese insulin-sensitive and obese insulin-resistant subjects. J. Clin. Endocrinol. Metab. 101, 1422–1428 (2016).
Henegar, C. et al. Adipose tissue transcriptomic signature highlights the pathological relevance of extracellular matrix in human obesity. Genome Biol. 9, R14 (2008).
Mor-Yossef Moldovan, L. et al. Cell shape alteration during adipogenesis is associated with coordinated matrix cues. J. Cell Physiol. 234, 3850–3863 (2019).
Scamfer, S. R., Lee, M. D. & Hilgendorf, K. I. Ciliary control of adipocyte progenitor cell fate regulates energy storage. Front. Cell Dev. Biol. 10, 1083372 (2022).
Nishimura, Y., Kasahara, K., Shiromizu, T., Watanabe, M. & Inagaki, M. Primary cilia as signaling hubs in health and disease. Adv. Sci. 6, 1801138 (2019).
Seeger-Nukpezah, T. & Golemis, E. A. The extracellular matrix and ciliary signaling. Curr. Opin. Cell Biol. 24, 652–661 (2012).
Christensen, S. T., Morthorst, S. K., Mogensen, J. B. & Pedersen, L. B. Primary cilia and coordination of receptor tyrosine kinase (RTK) and transforming growth factor beta (TGF-beta) Signaling. Cold Spring Harb. Perspect. Biol. 9, https://doi.org/10.1101/cshperspect.a028167 (2017).
Ehnert, S. et al. TGF-beta(1) impairs mechanosensation of human osteoblasts via HDAC6-mediated shortening and distortion of primary cilia. J. Mol. Med. 95, 653–663 (2017).
Kawasaki, M. et al. TGF-beta suppresses Ift88 expression in chondrocytic ATDC5 Cells. J. Cell Physiol. 230, 2788–2795 (2015).
Somatilaka, B. N. et al. Ankmy2 Prevents Smoothened-independent Hyperactivation Of The Hedgehog Pathway Via Cilia-regulated Adenylyl Cyclase Signaling. Dev. Cell 54, 710–726 e718 (2020).
Hilgendorf, K. I., Johnson, C. T. & Jackson, P. K. The primary cilium as a cellular receiver: organizing ciliary GPCR signaling. Curr. Opin. Cell Biol. 39, 84–92 (2016).
Bishop, G. A., Berbari, N. F., Lewis, J. & Mykytyn, K. Type III adenylyl cyclase localizes to primary cilia throughout the adult mouse brain. J. Comp. Neurol. 505, 562–571 (2007).
Nikonova, A. S. et al. Nedd9 restrains renal cystogenesis in Pkd1-/- mice. Proc. Natl. Acad. Sci. USA 111, 12859–12864 (2014).
Halbritter, J. et al. Defects in the IFT-B component IFT172 cause Jeune and Mainzer-Saldino syndromes in humans. Am. J. Hum. Genet 93, 915–925 (2013).
Ou, Y. et al. Adenylate cyclase regulates elongation of mammalian primary cilia. Exp. Cell Res. 315, 2802–2817 (2009).
Duffy, M. P., Sup, M. E. & Guo, X. E. Adenylyl cyclase 3 regulates osteocyte mechanotransduction and primary cilium. Biochem. Biophys. Res. Commun. 573, 145–150 (2021).
Fitzpatrick, M. & Solberg Woods, L. C. Adenylate cyclase 3: a potential genetic link between obesity and major depressive disorder. Physiol. Genom. 56, 1–8 (2024).
Tong, T., Shen, Y., Lee, H. W., Yu, R. & Park, T. Adenylyl cyclase 3 haploinsufficiency confers susceptibility to diet-induced obesity and insulin resistance in mice. Sci. Rep. 6, 34179 (2016).
Wang, Z. et al. Adult type 3 adenylyl cyclase-deficient mice are obese. PLoS ONE 4, e6979 (2009).
Siljee, J. E. et al. Subcellular localization of MC4R with ADCY3 at neuronal primary cilia underlies a common pathway for genetic predisposition to obesity. Nat. Genet. 50, 180–185 (2018).
Hansen, J. N. et al. Nanobody-directed targeting of optogenetic tools to study signaling in the primary cilium. Elife 9, https://doi.org/10.7554/eLife.57907 (2020).
Ansari, S. S. et al. Sonic Hedgehog activates prostaglandin signaling to stabilize primary cilium length. J. Cell Biol. 223, https://doi.org/10.1083/jcb.202306002 (2024).
Veland, I. R., Lindbaek, L. & Christensen, S. T. Linking the primary cilium to cell migration in tissue repair and brain development. Bioscience 64, 1115–1125 (2014).
Stoufflet, J. et al. Primary cilium-dependent cAMP/PKA signaling at the centrosome regulates neuronal migration. Sci. Adv. 6, https://doi.org/10.1126/sciadv.aba3992 (2020).
Ng, L. W. et al. Adipose-derived stem cells from pregnant women show higher proliferation rate unrelated to estrogen. Hum. Reprod. 24, 1164–1170 (2009).
Ritter, A., Friemel, A., Kreis, N. N., Louwen, F. & Yuan, J. Impact of Polo-like kinase 1 inhibitors on human adipose tissue-derived mesenchymal stem cells. Oncotarget 7, 84271–84285 (2016).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Muschol-Steinmetz, C. et al. B-cell lymphoma 6 promotes proliferation and survival of trophoblastic cells. Cell Cycle 15, 827–839 (2016).
Kreis, N. N. et al. Function of p21 (Cip1/Waf1/CDKN1A) in Migration and Invasion of Cancer and Trophoblastic Cells. Cancers 11, https://doi.org/10.3390/cancers11070989 (2019).
Steinhauser, K. et al. Deficiency of RITA results in multiple mitotic defects by affecting microtubule dynamics. Oncogene 36, 2146–2159 (2017).
Kreis, N. N. et al. Potential involvement of RITA in the activation of Aurora A at spindle poles during mitosis. Oncogene https://doi.org/10.1038/s41388-019-0716-7 (2019).
Acknowledgements
This work was funded by the German Research Foundation (Deutsche Forschungsgemeinschaft (DFG), project number 438690235 to J.Y.). We are grateful to our donors and our clinical team for making this study possible. We thank Drs. He and Anderson, Sloan-Kettering Institute, for kindly providing us the ImageJ plug-in Plot Roi Profile for analyzing the fluorescence intensity in cilia. We sincerely thank Mr. Dominic Menger (Georg-Speyer-Haus, Institute for Tumor Biology and Experimental Therapy, Frankfurt) for providing access to the confocal laser scanning microscope (CLSM).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
N.N.K. designed and performed experiments, analyzed data, and draw, modified and improved the manuscript. A.F. performed experiments, analyzed the data and modified the manuscript. A.R. analyzed data and did critical reading. A.E.H. has acquired and informed donors. E.S. collected the samples. F.L. conceived the project. J.Y. conceived the project and draw the first manuscript draft and did critical reading. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
AI disclouser
The authors declare that they have not use AI-generated work in this manuscript. During the revision process of this work, the authors used OpenAI’s language model (ChatGPT) to improve the readability and clarity of the manuscript. The authors subsequently reviewed and edited the content as necessary and take full responsibility for the final version of the manuscript.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Tiago Dantas and Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kreis, NN., Friemel, A., Ritter, A. et al. In-depth analysis of obesity-associated changes in adipose tissue-derived mesenchymal stromal/stem cells and primary cilia function. Commun Biol 8, 1462 (2025). https://doi.org/10.1038/s42003-025-08986-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08986-w