Abstract
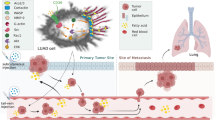
Lymph node metastasis (LNM) is significantly associated with distant metastasis and poor prognosis in patients with breast cancer (BC). However, the mechanism underlying LNM in BC remains unclear, limiting the development of effective anti-metastatic strategies. Given the lipid-enriched lymphatic microenvironment, we identify the fatty acid transporter cluster of differentiation 36 (CD36) via TCGA-based screening. Aberrant expression of CD36 is positively correlated with LNM and poor prognosis in clinical specimens. BC cells with high expression of CD36 confers anoikis-resistance exposed to lymph or palmitic acid treatment. Furthermore, CD36 drives LNM in both popliteal LNM and orthotopic BC mouse models through yes-associated protein (YAP)-TEADs signaling pathway. Elevated CD36 faciliates the uptake of fatty acids and induces the PPARα-CYP7A1-bile acid biosynthesis pathway, leading to the nucleus translocation and activation of YAP by suppressing the YAP phosphorylation at Ser127. Activated YAP-TEADs complexes directly bind to the BCL2 promoter to upregulate Bcl-2 expression. Collectively, our findings identify CD36 as a critical regulator of LNM in BC and reveal a CD36-PPARα-CYP7A1-YAP regulatory axis that may provide an effective therapeutic window for preventing metastasis in BC.

Similar content being viewed by others
Background
Breast cancer (BC) is the most common and harmful cancer in women worldwide, accounting for 11.6% of the incidence rate and 6.9% of the mortality rate1. The death of most BC patients is caused by tumor metastasis, which includes local invasion, vascular metastasis, implantation metastasis, and lymph node metastasis (LNM)2. Acting as a foothold for further dissemination of cancer, LNM presents the most frequent metastatic way in BC3,4, especially in luminal BC rather than in triple-negative BC (TNBC)5,6. Sentinel LNM is also an important risk indicator for further distant metastasis and poor prognosis7,8, and the number of positive lymph nodes is closely related to the risk of distant metastasis9,10. Previous studies have focused on distant metastasis rather than on the mechanism by which cancer cells survive and grow within the lymphatic system11. Hence, there is an urgent need to explore novel biomarkers and elucidate the mechanisms underlying LNM.
Tumor metastasis is a multi-step process that includes local invasion, intravasation to lymphatic or blood vessels, extravasation, and distant colonization12. Within the circulatory system, metastatic tumor cells are eliminated by a special type of programmed cell death termed anoikis due to the detachment from the extracellular matrix (ECM)13. During the process of metastasis, acquisition of anoikis resistance endows tumor cells to survive within circulation in an anchorage-independent manner, and is considered the critical prerequisite for metastatic initiation and dissemination14,15,16. There are multiple strategies by which tumor cells could acquire anoikis resistance, including alterations in fatty acid and glucose metabolism, epithelial-mesenchymal transition, reactive oxygen species signaling, and aberrantly activated anti-apoptotic or pro-survival pathway17,18,19. Nevertheless, the precise mechanisms underlying anoikis resistance in metastatic tumor cells, especially within the lipid-enriched environment20,21, still remain unclear.
According to the lipid-enriched characteristics of the lymphatic system and the crucial role of lipid metabolism on tumorigenesis and LNM22, we performed bioinformatics screening on the TCGA database and found that FAs transport pathways were significantly correlated with LNM in BC. Furthermore, among the multiple fatty acid receptors, CD36 exhibited the strongest correlation with lymph node staging, suggesting that CD36 may be a key functional molecule regulating lymph node metastasis in breast cancer. Cluster of differentiation 36 (CD36) is a multifunctional glycoprotein that is identified as a receptor for various ligands, including fatty acids (FAs), oxidized low-density lipoprotein (oxLDL), cholesterol and thrombospondin-1 (TSP-1). According to different ligands, CD36 plays crucial roles in lipid homeostasis, atherosclerosis, angiogenesis, and tumorigenesis23. In the present study, we aimed to identify the effects of CD36 on promoting LNM in BC and elucidate the underlying mechanism.
Methods
Clinical specimens
This study utilized medical samples and records from two cohorts: 77 human breast cancer (BC) samples with complete medical records were collected from January 1, 2009, to December 31, 2015, at Sun Yat-sen University Cancer Center. Additionally, 80 cases were collected from January 1, 2017, to December 31, 2024, at The Seventh Affiliated Hospital of Sun Yat-sen University. Additionally, the tissue microarrays were purchased from Shanghai Xinchao Biotech Co., Ltd (HBre-Duc060CD-01, 5 cases of adjacent tissue, and 6 cases of normal breast tissue). In total, there were 157 cases of BC, 5 cases of adjacent tissue, and 6 cases of normal breast tissue. All patients’ informed consent has been obtained before surgery, and the use of medical records and histological sections has been approved by the Ethics Committee of Sun Yat-Sen University (B2024-088-01), and Shanghai Xinchao Biotech Co., Ltd (YB M-05-01). All procedures were performed under consensus agreements and in accordance with the Chinese Ethical Review Committee. The study methodologies conformed to the standards set by the Declaration of Helsinki. The clinical and biological characteristics of the patients were described in Table S1.
Cell lines and culture
Human normal mammary epithelial cells MCF-10A was provided by Prof. Erwei Song (Sun Yat-sen Memorial Hospital, China). Human breast cancer cell line T-47D, MDA-MB-231, MCF-7, and HCC1937 were provided by Prof. Haihe Wang (ZhongShan School of Medicine, Sun Yat-sen University, China). 4T1 was purchased from ATCC. All cell lines were maintained in DMEM supplemented with 10% FBS and were incubated at 37 °C under a humidified atmosphere with 5% CO2.
Quantitative Real-Time PCR
Total RNA was isolated from cultured cells using TRIzol reagent, following the manufacturer’s protocol. Reverse transcription was performed with 500 ng of total RNA using the PrimeScript RT Reagent Kit (Takara, Japan). Quantitative real-time PCR (qPCR) was carried out on a Bio-Rad CFX 96 fluorescence quantitative PCR instrument (Bio-Rad, USA) with SYBR® Green I dye (Takara, Japan). Relative mRNA expression levels were calculated using the comparative cycle threshold method. Primer sequences for target genes are listed in Table S4.
Western blot analysis
Total protein from tumor tissues and cultured cells was lysed in SDS lysis buffer (Beyotime, Shanghai, China, #P0013G) supplemented with 1 mM phenylmethanesulfonyl fluoride (Beyotime, #ST506). Protein concentration was quantified using a bicinchoninic acid (BCA) assay kit (KeyGen, Nanjing, China, #KGP902). Nuclear extracts were obtained using the NE-PER Nuclear and cytoplasmic extraction reagents (Thermo Scientific, Waltham, MA, USA, 78833) according to the manufacturer’s instructions. Mitochondrial extracts and cytoplasmic extracts were collected by using Cell Mitochondrial Isolation Kit (C3601, Beyotime). Proteins were separated by SDS-PAGE and transferred to 0.45 μm polyvinyl difluoride (PVDF) membranes (Millipore) according to standard immunoblotting protocols. After blocking with 5% of defatted milk in TBST (20 mM of Tris-HCl pH 7.4, 500 mM of NaCl and 0.1% of Tween-20) for 1 h at room temperature, the membranes were incubated with the following primary antibodies: CD36 (#14347, 1:1000), Caspase 9 (#9502, 1:1000), Cytochrome-C (#4272, 1:1000), YAP (#14074, 1:1000), p-YAP (Ser127) (#4911, 1:1000) from Cell Signaling Technology; Cox-Ⅳ (ab14744, 1:5000), Histone H3 (ab1791, 1:5000), β-tublin (ab179513, 1:5000), CTGF (ab6992, 1:1000) from Abcam; Bcl-2 (sc-7382, 1:1000), Bak (sc-832, 1:1000), Bax (sc-6236, 1:1000) from Santa Cruz; Bim (AF1573, 1:1000) from Beyotime; Bid (MB0118, 1:1000) from bioworld; β-actin (A5441, 1:5000) from Sigma-Aldrich (St. Louis, USA); Ubiquitin Rabbit mAb (A19686, 1:1000) from ABclonal; Lamin B1 Ab (AB0054, 1:5000) from Abways. HRP-conjugated anti-rabbit IgG (Cell Signaling Tech, #7074) and anti-mouse IgG (Sigma-Aldrich, AP308P) were used as secondary antibodies. Proteins were determined using ECL Plus Reagent (Millipore, WBKLS0100) and visualized using the Bio-Rad ChemiDoc Touch Imaging System.
Lentiviral infection and plasmids transfection
The luciferase-overexpressing lentivirus (pLenti-CBh-3FLAG-luc2-tCMV-mNeonGreen-F2A-Blasticidin), CD36-overexpressing lentivirus (pSLenti-EF1-EGFP-F2A-Puro-CMV-CD36-HA-WPRE), CD36-interfering lentivirus (pCLenti-U6-shCD36-CMV-Puro-WPRE), YAP1-overexpressing lentivirus (pCLenti-EF1-Hygro-CMV-YAP1-3xFLAG-WPRE), YAP1-interfering lentivirus (pCLenti-U6-shYAP1-CMV-Hygro-WPRE) and negative control lentivirus were purchased from OBiO Technology (Shanghai, China). CD36-overexpression pcDNA3.1 (+) plasmid was constructed and conserved by our lab. For lentivectors infection experiments, cells were infected with lentivirus followed by drug selection for 1 week (1 μg/mL puromycin, 50 μg/mL hygromycin B, or 2 μg/mL blasticidin). For transfections experiments, plasmids were performed at ~60% confluency using Lipofectamine 3000. After 48 h, confirmation of overexpression was carried out using real-time quantitative PCR (RT-qPCR) and western blotting.
Cannulation of lymph duct in rat
The steps of lymphatic cannulation in SD rats are carried out according to the video of Jove website (https://www.jove.com/v/52389/the-mesenteric-lymph-duct-cannulated-rat-model-application-to), The method of cannulation of mesenteric lymphatic ducts is briefly described as follows: Place the animal with its right side facing toward the operator. Open the abdominal muscle wall from 4–5 mm lateral to the midline to the right flank with a small pair of surgical scissors. Retract the small intestine under the left abdominal muscle wall and keep it in place using 2–3 pieces of sterile gauze saturated with normal saline. Locate the superior mesenteric lymph duct: a vessel ~0.5–1 mm in diameter that is perpendicular to the right kidney and immediately rostral and parallel to the mesenteric artery. After isolating the lymph duct, make a small hole in the lymph duct with either micro-scissors. Ensure that the lymph cannula is completely filled with anti-coagulant solution (using a syringe and 25 G needle) with no air gaps. Insert the lymph cannula ~2–4 mm inside the mesenteric lymph duct via the small hole with the aid of a small pair of forceps. Observe the lymph cannula for a couple of minutes. Observe a gradual flow of intestinal lymph from the free collecting end of the cannula. If the cannulation is successful, secure the cannula by placing a small drop of veterinary adhesive over the entrance hole into the lymph duct. Remove the gauze pieces that were placed in the abdominal cavity carefully and place the small intestine in its original position. Place a lymph collection tube containing anti-coagulant at the end of the cannula to collect the free flowing lymph.
Detection of lipid contents
Triglycerides (TG) and free fatty acids (FFA) of lymph in SD rat were examined by using Triglyceride Assay Kit (ETGA-200) and free Fatty acid detection kit (EFFA-100) from Bioassay Systems specifically according to the manufacturer’s instructions. To examine lipid contents of xenograft tumor tissue, TG, FFA, total cholesterol (TC), and total bile acid (TBA) were examined by Hitachi Biochemical Analyzer. Pre-treatment for lipid composition detection of xenograft tumor is described as follows: 50 mg of fresh tumor tissues was collected. After adding 500 μL of PBS containing 5% Triton X-100, the tissues were homogenized and sonicated on ice, and then centrifuged at 12,000 rpm for 10 min. The supernatants were collected and tested by Hitachi Biochemical Analyzer.
Immunohistochemistry staining
Formalin-fixed, paraffin-embedded tissue sections were deparaffinized, rehydrated, and subjected to antigen retrieval in 0.01 M citrate buffer (pH 6.0) using a high-pressure method. Endogenous peroxidase activity was blocked with 3% H2O2 for 20 min, followed by incubation with 5% goat serum to reduce non-specific binding. Next, the following primary antibodies were used: CD36 (CST, 14347, 1: 100), YAP (CST, 14074, 1: 100). After incubating with primary antibodies at 4°C overnight, On the second day, the slides were treated with HRP-conjugated secondary antibody and the antigen-antibody complex was visualized by incubation with the DAB kit. Sections were photographed through a slide scanner (Axio Scan. Z1, ZEISS). The protein expression level was determined by the staining index (SI). The SI is calculated by stained tumor cell proportions score multiplies staining intensity score. The stained tumor cell proportions were graded as follows: 0, <5% positive tumor cells; 1, 5–25% positive tumor cells; 2, 26–50% positive tumor cells; 3, 51–75% positive tumor cells; 4, >75% positive tumor cells. Staining intensity was scored as follows: 0, negative staining (no staining); 1, weak staining (light yellow); 2, moderate staining intensity (brown); 3, positive staining (yellow). In this study 157 breast cancer cases were divided into 68 cases with high CD36 expression and 89 cases with low expression of CD36 according to the median score.
Popliteal lymph node metastasis model
Four to five weeks old female athymic nude mice (BALB/c, nu/nu, 16−20 g) were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China, license number: SCXK (Beijing) 2016-0006) and raised in North Campus Laboratory Animal Center of Sun Yat-sen University (license number: SYXK (Guangdong) 2019-0209). All of the experiment procedures were reviewed and approved by the Institutional Animal Care and Use Committee of Sun Yat-sen University. Popliteal lymph node metastasis model was conducted according to previous reference24, which is briefly described as follows: The BC cell lines MDA-MB-231 and T-47D were previously infected by luciferase-overexpressing lentivirus (pLenti-CBh-3FLAG-luc2-tCMV-mNeonGreen-F2A-Blasticidin) and indicated lentivirus to generate stable cell lines. 2 × 106 BC cell lines constructed above were subcutaneously inoculated into the foot-pads of mice. Twenty-eight days after the tumors in the foot-pads reached a volume of ~100 mm3, the foot-pads and popliteal lymph nodes of mice were imaged with the IVIS Spectrum Imaging System (Perkin Elmer) after intraperitoneal injection of luciferin (Promega, 150 ug/g mice weight). The mice were then sacrificed, and tumors and popliteal lymph nodes were dissected and weighted. Volume of tumors and lymph nodes was calculated by the following formula: volume (mm3) = (length × width2)/2. Last, tumors and popliteal lymph nodes were enucleated and embedded in parafn. Serial 5 μm sections were analyzed by green fuorescent protein (GFP) imaging and immunohistochemistry (IHC) with anti-CD36 or YAP antibodies. The images were captured using an AxioVision Rel.4.6 system (Carl Zeiss, Jena, Germany). The work has been reported in accordance with the ARRIVE guidelines (Animals in Research: Reporting In Vivo Experiments)25.
Orthotopic breast cancer model establishment
Female BALB/c mice (5–7 weeks old, weighing 18-25 g) were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd. (license number:SCXK (Beijing) 2021-0006). The mice were housed in the Experimental Animal Center of the North Campus of Sun Yat-sen University (license number: SYXK (Guangdong) 2024-0081). All experimental procedures were reviewed and approved by the Institutional Animal Care and Use Committee of Sun Yat-sen University.The breast cancer cell line 4T1 was initially transduced with a lentivirus overexpressing luciferase (pLenti-CBh-3FLAG-luc2-tCMV-mNeonGreen-F2A-Blasticidin) to generate a stable polyclonal population. Stable clones were selected and maintained in medium containing 10 μg/mL Blasticidin S hydrochloride (HY-103401, MCE). Subsequently, this luciferase-expressing stable cell line was further infected with either a Cd36-overexpressing lentivirus or a corresponding negative control lentivirus. A suspension containing 1 × 106 cells from the aforementioned engineered cell lines, mixed with Matrigel, was orthotopically inoculated into the fourth mammary fat pad of BALB/c mice to establish the orthotopic breast cancer model. Three weeks after tumor cell inoculation, D-luciferin (Beyotime, 150 mg/kg mouse body weight) was administered intraperitoneally. Imaging of the mammary region and inguinal lymph nodes was performed using an IVIS Spectrum imaging system (PerkinElmer). Following imaging, the mice were euthanized, and the tumors and inguinal lymph nodes were dissected and weighed. Tumor and lymph node volumes were calculated using the formula: Volume (mm³) = (Length × Width²)/2. Finally, the harvested tumors and lymph nodes were fixed, embedded in paraffin, and sectioned into consecutive 5 μm slices. Green Fluorescent Protein (GFP) signaling was visualized in these sections using fluorescence imaging.
Mitochondrial staining and live cell imaging
MitoLite™ Red FX600 dye (AAT Bioquest, USA, #22677) was used to indicate changes in mitochondrial membrane potential. According to the manufacturer’s instructions, the dye was diluted into 1:500 and added to the cell wells. After incubated for 1 h in a 5% CO2, 37 °C incubator, cell wells were washed three times with PBS and supplemented culture medium and indicated treatments for live cell imaging. 594 nm (Red) and phase contrast microscope field (white light) were taken under the live cell imager (BioTek) for 48 h. The curve of mean fluorescence intensity of MitoLite red fluorescence was statistically analyzed with the instrument software.
Cell viability assay
The viability of BC cells was examined through Cell Counting Kit-8 (CCK-8) assay (Dojindo, CK04), following the manufacturer’s protocol. Briefly, 5000 BC cells were seeded in 100 μL of growth medium per well in ultra-low adhesion 96-well plates (Corning, #3474). Cells were treated differently according to the experimental requirements, and 10 μL of CCK8 was added to each well after 12, 24, 34, 48 h. After 2 h of incubation at 37°C in a humidified incubator containing 5% CO2, the OD value was determined by absorbance at 450 nm using the Sunrise microplate reader (TECAN, Mäannedorf, Switzerland).
Apoptosis Assay
Apoptosis was determined using flow cytometry with a commercial Annexin V-FITC Apoptosis kit (Vazyme Biotech, Nanjing, China) according to the manufacturer’s protocol. 5 × 105 BC cells were seeded in ultra-low adhesion 6-well plates (Corning, #3471) for suspension culture to simulate an anoikis environment, and treated with vehicle or fatty acid for 24 h. Subsequently, cells were collected and washed in 4 °C PBS, and then trypsinized gently using a trypsin solution. After centrifugation to remove the trypsin, the cells were resuspended in binding buffer containing Annexin V-FITC and propidium iodide (PI), then incubated for 15 min at room temperature in the dark and subsequently analyzed on a Beckman CytoFLEX flow cytometer (Beckman Coulter, Inc., Brea, CA, USA).
Detection of mitochondrial membrane potential
Mitochondrial membrane potential (MMP) was determined using JC-10 (Solarbio, Beijing, China, CA1310), a cationic and lipophilic dye that selectively enters mitochondria. In general, JC-10 accumulates in the mitochondrial matrix where it forms red fluorescent aggregates. In apoptotic and necrotic cells, JC-10 diffuses out of the mitochondria and changes to monomeric forms and stains cells with green fluorescence, indicating depolarization of MMP and mitochondrial apoptosis. According to the manufacturer’s protocol, 5 × 105 BC cells treated with suspended palmitic acid (PA, 800 μm) conditions for 24 h were collected and strained with JC-10 for 20 min at 37 °C incubator. After washed with washing buffer, the JC-10 fluorescents were imaged using Leica microscope (DMI8). The aggregate/monomer ratio was quantitated and was proportional to the MMP.
Immunofluorescence staining
After fixed in 4% paraformaldehyde, cells were blocked with goat serum at 37 °C for 1 h. They were incubated with mouse CD36 antibody (Santa Cruz, sc-7309, 1: 100) and rabbit YAP antibody (CST, 14074, 1: 100) at 4 °C overnight and then were incubated with anti-mouse 594 (Thermo Fisher, A21203) and anti-rabbit 488 (Thermo Fisher, A21206) at 37 °C for 1 h after three times washing. Finally, DAPI (Sigma-Aldrich, Tokyo, Japan, D9542) was used to stain the cell nucleus. Cells were visualized under Olympus BX51.
Dual-luciferase reporter assay
Luciferase assay was performed using the Dual Luciferase Reporter Assay Kit (Promega, Madison, WI, USA) according to manufacturer’s protocol. The human BCL2 promoter region spanning nucleotides −2100 to 0 (relative to the TSS) were cloned into pGL3-basic luciferase reporter plasmid to generate full-length BCL2 luciferase reporters. The truncated fragments of BCL2 promoter with predicted binding site (BCL2-PS) spans nucleotides −695 to −296, and the mutant site spans nucleotides −433 to −421 within BCL2-PS region (BCL2-PS-Mut). The full-length BCL2, BCL2-PS, and BCL2-PS-Mut pGL3 plasmids were all constructed by Guangzhou Hanyi Biotech Co., Ltd.
Chromatin immunoprecipitation (ChIP) assay
2 × 106 T-47D cells in a 100 mm dish were treated with 1% formaldehyde to cross-link proteins to DNA. The cell lysates were sonicated to shear the DNA into 100-1000 bp fragments. YAP antibody (CST, 14074, 1:200) or IgG antibody was mixed with the sonicated product and incubated overnight on a shaker at 4 °C. Following reverse cross-linking of protein–DNA complexes to free the DNA, PCR was carried out. The primers used in this study are listed in the Table S4.
Cycloheximide chase assay
T-47D or MDA-MB-231 cells were plated on six-well culture plates and transient transfection procedure was performed as described above. After transfection for 24 h, CHX (Genview, Beijing, China, AC466) were added to cells at a final concentration of 10 μg/ml to block new protein synthesis. MG132 is a specific proteasome inhibitor (Selleck Chemicals, Houston, USA, S2619). Cells were then collected at the indicated time points for western blotting analysis.
Immunoprecipitation (IP)
MDA-MB-231 cells were transfected with a CD36-expressing plasmid (CD36 plasmid) or a control plasmid (CON plasmid). CD36-overexpressing cells were subsequently treated with the YAP inhibitor verteporfin (HY-B0146, MCE) or vehicle control (DMSO). To stabilize ubiquitinated proteins, three groups were treated with the proteasome inhibitor MG132 (10 µM) for 12 h prior to cell lysis. Seed cells in 10 cm culture dishes. After treatment to a specific time point, discard the supernatant, wash with PBS for 3 times, aspirate the residual PBS, and place the culture dishes on ice to maintain low temperature.Add 500 μL of weak RIPA lysis buffer (BL652A, Biosharp), scrape the cells with a cell scraper, and place them on a 4 °C shaker for slow shaking to lyse overnight.Centrifuge at 12,000 rpm at 4 °C for 20 min, then take the supernatant for quantification.Take 1000 μg of protein, add the corresponding IP-competent antibody at a ratio of 1:100, and incubate overnight on a 4 °C shaker with slow shaking. For each sample, prepare 20 μL of Protein A/G PLUS Agarose(sc-2003,Santa Cruz Biotechnology), wash it with 300 μL of weak RIPA lysis buffer, centrifuge at 2000 rpm at 4 °C for 2 min, remove the supernatant, and repeat this washing step 3 times. Add the washed Protein A/G PLUS Agarose to the sample that has been incubated with the YAP antibody overnight, and incubate again overnight.Centrifuge at 2000 rpm at 4 °C for 5 min, remove the supernatant, add 300 μL of weak RIPA lysis buffer for washing (repeat 3 times), and then lyse the precipitate with 1 × SDS loading buffer for WB detection.
Bioinformatics Analysis
199,764 BC cases of Surveillance, Epidemiology, and End Results (SEER) data was downloaded from SEER database and extracted by SEER*Stat software. TCGA BC data was downloaded from LinkOmics database for analysis. Cancer Cell Line Encyclopedia (CCLE) database was downloaded from the CCLE official website (https://portals.broadinstitute.org/ccle/data). GEO dataset (GSE59232) was obtained by R software using GEOquery package. R (R version 4.0.0) packages used in this study were listed as follow: limma was used for differential expression analyses; Gene Set Variation Analysis (GSVA) was used to estimate variation in pathway and biological process enrichment through the samples of a expression data set. EnhancedVolcano and pheatmap were performed for volcano map and heat map plotting. JASPAR database (http://jaspar.genereg.net/) was used to predict the possibility of the BCL2 promoter region (2100 bp upstream of the gene) binding to TEADs family transcription factors, as well as its specific binding sites.
scRNA-seq data analysis: The raw single-cell RNA sequencing data were downloaded in GEO database (GSE176078) and processed using the R package Seurat (version 3.1.1) on R platform (version 4.2.1). After passing quality control, cells were merged into one count matrix and normalized using the NormalizeData and ScaleData functions of Seurat. Variable genes were identified using the “FindVariableGenes” function. RunPCA function was performed on the variable genes, and the top principal components were selected for downstream analysis. FindClusters function was used for cell clustering and FindAllMarkers function was applied for identifying the markers of clusters.
Statistical analysis
The variability of the data was expressed as mean ± standard deviation (mean ± SD) and was assessed with Student’s t-test between two groups. One-way analysis of variance (One-way ANOVA) was used for comparison of multiple groups, and least significant difference t-test (LSD-t) was used for pairwise comparison between groups. Categorical variables were compared using the Chi-squared test. The correlation of two continuous variables was analyzed by Pearson correlation, and the correlation analysis of variables involving rank data was analyzed by Spearman correlation analysis. The above analysis was completed using SPSS 23.0 software. Kaplan-Meier survival curves and their Log-rank statistical analysis were performed using GraphPad Prism 8. Statistical significance was defined at p < 0.05.
Results
CD36 is highly expressed in BC and positively correlated with N stage and poor prognosis
To clarify the relationship between lymph node metastasis and distant metastasis and prognosis in BC, we analyzed the correlation of lymph node stage with distant metastasis and prognosis of patients from the TCGA and SEER databases. The results showed that lymph node stage was positively correlated with the incidence of distant metastasis and poor prognosis, corroborating the importance of LNM (Fig. S1A–D). Based on the lipid-enriched characteristics of the lymphatic system, which were also validated in SD rats (Fig. S1E–G), we conducted a correlation analysis between N stage and lipid/glucose metabolism level assessed by Gene Set Variation Analysis (GSVA). The volcano plot exhibited that several fatty acids transport pathways presented strong correlation with N stage, whereas some glucose metabolism pathways were negatively correlated with lymph node staging (Fig. S1H). Further, among several fatty acid receptors, CD36 exhibited the strongest correlation with N stage (Fig. S1I), suggesting that CD36 may be a key molecule mediating LNM in BC. Next, immunohistochemistry (IHC) was performed to detect CD36 expression in BC clinical specimens. As shown in Fig. 1A, B, the expression of CD36 in BC tumor tissue was significantly higher than that in normal and adjacent tissues. Consistent with TCGA database analysis, the expression of CD36 was positively correlated with lymph node staging and the number of positive lymph nodes (Fig. 1C–E). In addition, CD36 expression was also associated with the occurrence of distant metastasis and tumor stage (Fig. 1F, G), and showed no significantly correlation with T stage and patient age (Fig. S1J, K). Importantly, the Kaplan-Meier survival curve showed that the progression-free survival time and overall survival time of breast cancer patients with high CD36 expression were significantly shorter than those of patients with low CD36 expression (Fig. 1H, I). These results demonstrated that the aberrant elevation of CD36 was positively correlated with LNM and poor prognosis of BC.
A Representative images of CD36 IHC staining of normal, adjacent specimens and BC specimens in a BC Tissue Microarray (top, 100× magnification; bottom, 200× magnification). Scale bar, 100 μm and 50 μm. B Statistical analysis of CD36 staining in normal (n = 6), adjacent (n = 5), and BC (n = 157) specimens. C Representative images of CD36 IHC staining at different N stages (50x magnification). Scale bar, 100 μm. D Statistical analysis of CD36 staining at different N stages (N0: n = 34, N1: n = 59, N2: n = 40, N3: n = 24). E Correlation analysis of CD36 expression with the number of positive lymph node (LN) in 157 BC patient specimens (Spearman). F Statistical analysis of CD36 staining in non-metastatic specimens (n = 123) and metastatic BC specimens (n = 32). G Statistical analysis of CD36 staining at different clinical stages (Stage Ⅰ-Ⅱ: n = 73, Stage Ⅲ-Ⅳ: n = 67). H, I Progression-free survival analysis and overall survival analysis of 157 BC patients with low versus high CD36 expression. Bars correspond to mean ± standard deviation (SD). *p < 0.05; ***p < 0.001.
Aberrant CD36 promotes lymph node metastasis of BC cells in vivo
To further verify the abnormal expression of CD36 in clinical subjects, we assessed the expression of CD36 in breast cancer cell lines. Compared with normal breast epithelial cell line MCF-10A, CD36 expression in breast cancer cells was significantly elevated (Fig. S2A). Previous study have reported that TNBC has the lowest lymph node metastasis rate compared with other molecular subtypes of breast cancer5,6, which is consistent with our TCGA database analysis (Fig. S2B, C). Notably, CD36 expression in luminal breast cancer cell lines was also higher than that in TNBC via WB and qPCR experiments (Fig. S2A, D). Consistent with these findings, analysis of the TCGA and CCLE databases revealed similar trends (Fig. S2E, F). Furthermore, single-cell RNA sequencing (scRNA-seq) analysis indicated that CD36 expression in epithelial cell subpopulations was markedly lower in TNBC samples compared to luminal samples (Fig. S2G, H). Intriguingly, CD36 expression was significantly higher in luminal epithelial and precursor cells from LNM-positive cases than in those from LNM-negative cases (Fig. S2I). We next conducted a popliteal LNM model to fully characterize the role of CD36 in LNM of BC, We utilized in vivo imaging system IVIS to track the labeled metastatic BC cells. Compared with the control group, the number of metastasized cancer cells in the popliteal lymph nodes (LNs) was significantly increased in the CD36-overexpressing MDA-MB-231 (231-CD36) group (Fig. 2A). The popliteal LNs in the 231-CD36 group exhibited dramatic increases in volume and weight upon dissection compared with those in the 231-EGFP group (Fig. 2B, C; Fig. S2J). Furthermore, immunofluorescence (IF) staining confirmed a significantly higher incidence of LN metastasis in the 231-CD36 group than in the 231-EGFP group (Fig. 2D, E). Likewise, in the CD36-knockdown T-47D (T-47D-shCD36) group, the luciferase signal intensity in the popliteal LNs was markedly reduced relative to the T-47D-shNC group (Fig. 2F). Consistently, upon dissection, the volume and weight of the LNs, as well as the metastasis incidence determined by IF, were all significantly decreased (Fig. 2G–J; Fig. S2K). In addition, an orthotopic mammary fat pad model using 4T1 cells was established to better recapitulate clinical metastasis patterns. The results showed that in the Cd36-overexpressing 4T1 (4T1-Cd36) group, the luciferase signal in the inguinal LNs was significantly enhanced (Fig. 2K), along with increased LN volume, weight, and inguinal LN metastasis rate (Fig. 2M–O; Fig. S2L). Taken together, these in vivo findings demonstrate that CD36 promotes lymph node metastasis in breast cancer. Taken together, these results confirmed that CD36 induced LNM of BC in vivo.
A, F, K Indicated BC cells which had metastasized to the homolateral popliteal (A, F) or inguinal (K) lymph nodes were imaged by live imaging with IVIS system after luciferin injection, red arrows showing the metastasized BC cells which were labeled with luciferase beforehand. B, G, L Representative photographs of the resected xenograft tumors and popliteal lymph nodes from nude mice (B, G), and the resected orthotopic tumors and inguinal lymph nodes from BALB/c mice (L), respectively. C, H, M Violin plot representing the volume of lymph nodes from the indicated group. D, I, N BC cells which had metastasized to the homolateral popliteal (D, I) or inguinal (N) lymph nodes were imaged by fluorescence microscope. The metastasized BC cells were labeled with GFP beforehand. 40× and 200× magnification. Scale bar: 1 mm and 200 μm. (E, J, O) The contingency plot showed the frequency of lymph node metastasis of each group (Chi-squared test). *p < 0.05, **p < 0.01.
Elevated CD36 induces anoikis-resistance of BC cells within lymph or palmitic acid condition
To investigate how CD36 mediates LNM in BC, we tested the adaptability of BC cells with different CD36 expression in lymph according to its lipid-enriched condition. The results showed that within the lymph environment, the MMP and cellular status of T-47D cell with high CD36 expression maintained stably, while MDA-MB-231 cell, whose CD36 expression was absent, exhibited striking reductions of MMP and apoptosis (Fig. 3A; Fig. S3A). To further explore the key lipid components in lymph that induce apoptosis of BC cells, we simulated lymph with different ratios of saturated fatty acid palmitic acid (PA) and monounsaturated fatty acid oleic acid (OA) and repeated the experiment. The live cell imaging results showed that under PA-treated conditions, T-47D cells had longer maintenance of MMP and better cellular status than MDA-MB-231 cells, in line with lymph treatment (Fig. 3B; Fig. S3B). Consistent results were obtained by using the CCK8 method to tested cells viability after 48 h of treatment with lymph and PA (Fig. S3C). However, there was no significant difference between the two cells lines under the others fatty acids treatments (Fig. S3D). Previous studies have reported that tumor cells suspended in the circulatory system are prone to anoikis26. Based on the above results, we speculated that CD36 may induce anoikis-resistance in BC cells in a high-fat lymphatic environment. We used ultra low-adherent plates for BC cells detached culture and evaluated the viability and apoptosis of BC cell lines by CCK8 assay and Annexin Ⅴ/PI staining. The results exhibited that T-47D and HCC1937 cells, which have sufficient CD36 expression, stably maintained cell viability under PA conditions, while CD36 absent MDA-MB-231 and MCF-7 cells presented notable decreases in cell viability and apoptosis (Fig. S3E–G). Furthermore, CD36 overexpression significantly enhanced the cellular viability and anoikis-resistance of BC cells (Fig. 3C, D; Fig. S3H), and the CD36-knockdowned cells exhibited the opposite effect (Fig. 3E, F; Fig. S3I). In addition, soft agar assays showed that CD36 promotes anchorage-independent growth of BC cells (Fig. 3G, H). Collectively, we concluded that CD36 confers resistance to anoikis in BC cells.
A, B Representative images of MitoLite staining of MDA-MB-231 and T-47D cells treated with SD rat lymph (A) or 800 μM PA (B) (100× magnification). Scale bar: 200 μm. Curves of MitoLite mean intensity of of 231 and T-47D treated with SD rat lymph (A) or 800 μM PA (B). C, E The viability of indicated detached EGFP/CD36-infected or shNC/shCD36-infected BC cells treated with palmitic acid (PA, 800 μm) by using CCK8 (n = 5). D, F Flow cytometric analyses of indicated NC/CD36-transfected (D) or shNC/shCD36-1, 2, 3-transfected (F) BC cells apoptosis under suspended palmitic acid (PA, 800 μm) conditions for 24 h. Apoptosis percentages were plotted as mean ± S.D of three independent experiments. G, H Representative images of soft agar assays of indicated BC cells. Colony numbers were plotted as mean ± S.D of three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, n.s. not significant.
CD36 inhibits mitochondrial apoptosis of BC cells treated with palmitic acid
Previous studies have reported that anoikis of tumor cells is mainly dependent on the mitochondrial apoptosis or the death receptor apoptosis pathway18. To explore the pathway by which CD36 is involved, we performed GSVA analysis of apoptotic pathways in the TCGA database to screen for CD36-related apoptotic pathways. The results of the volcano plot and correlation matrix analysis showed that the expression level of CD36 was significantly negatively correlated with multiple mitochondrial apoptosis pathways (Fig. S4A). The results of JC-10 assay showed that the MMP was stably maintained in CD36-transfected 231 cells and was depolarizated in CD36 knockdown T-47D cells (Fig. 4A, B). Consistent results were also observed in MCF-7 and HCC1937 cells (Fig. S4B). Furthermore, the results of WB showed that overexpression of CD36 resulted in the dramatic upregulation of cleaved caspase 9 (Fig. 4C; Fig. S4C) and higher levels of cytochrome c (Cyto-C) released from the mitochondria to the cytosol as compared with control within PA condition. Conversely, CD36-silenced T-47D cells exhibited reduced cleavage of caspase 9 and release of Cyto-C (Fig. 4C, D).
A, B Mitochondrial membrane potential was detected by JC-10 assay. In normal cells, JC-10 concentrates in the mitochondrial matrix where it forms red fluorescent aggregates. While in apoptotic and necrotic cells, JC-10 changes to the monomeric form and stains cells in green fluorescence. Indicated NC/CD36-transfected 231(A) and shNC/shCD36-infected T-47D cells (B) were treated with 800 μM PA. 200× magnification. Scale bar: 75 μm. Ratios of JC-10 fluorescence were plotted as mean ± S.D of three independent experiments. C Indicated detached BC cells were treated with vehicle (Veh) or 800 μm palmitic acid for 24 h. Caspase 9 was examined by WB to analyze mitochondrial caspase activation. D Cells were fractionated to mitochondrial and cytosol subfraction after undergoing the same treatments above. Cytochrome C (Cyto-C) protein in mitochondrial and cytosol subtraction were detected by WB assay. β-actin was loaded as a control to cytosol subfraction, Cox-IV was loaded as a control to mitochondrial subfraction. E−H Relative mRNA and protein level of several Bcl-2 family proteins in indicated detached BC cells treated with vehicle (Veh) or 800 μm palmitic acid by RT-qPCR (E, F) and WB analysis (G). H WB analysis of the expression of Bcl-2 from xenograft models tumors. Tumors formed by EGFP-Luc-/CD36-Luc-infected MDA-MB-231 cells. Bars represent the mean ± SD of three independent experiments, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, n.s. not significant.
The Bcl-2 family controls Cyto-C release and the balance of mitochondrial membrane permeability27, thus we further tested whether the Bcl-2 family was involved in the effect of CD36. Impressively, we found that BCL2 mRNA levels were both significantly up-regulated by CD36 in both 231 and T-47D cells, while other molecules remained unchanged (Fig. 4E, F; Fig. S4D). Moreover, CD36 increased the protein expression of Bcl-2 in BC cells (Fig. 4G) and animal models (Fig. 4H), whereas silencing CD36 had the opposite effects (Fig. S4E). Collectively, CD36 induced the mRNA and protein levels of Bcl-2 in BC cells.
CD36 induced the activation of Hippo-YAP signaling
To uncover the underlying mechanism of CD36 leading to LNM and anti-anoikis, we first analyzed the correlation between the expression of CD36 and multiple reported key molecules of lymph node metastasis28. Correlation analysis showed that CD36 was significantly positively correlated with various molecules of the Hippo-YAP pathway (Fig. 5A), which plays a critical role in lymph node metastasis and anoikis-resistance20,29,30. Moreover, IHC staining of clinical specimens exhibited that CD36 expression was significantly correlated with YAP (Fig. 5B), and 231-CD36 xenografts presented higher YAP expression compared with 231-EGFP group (Fig. 5C, D). In BC cell lines, the expression of YAP was higher with highly expressed CD36 than in cell lines with low CD36 expression (Fig. S5A). What’s more, the expression of YAP was significantly elevated in CD36-transfected 231 and MCF-7 cells, but down-regulated in CD36-silenced T-47D cells (Fig. 5E; Fig. S5B, C). However, the mRNA levels remained unchanged after CD36 transfection or knockdown (Fig. 5F). Therefore, we explored the regulation of CD36 on the post-translational modification of YAP. Studies have reported that YAP phosphorylation at Ser127 (p-YAP S127) blocks YAP from entering the nucleus, leading to the cytoplasmic retention, ubiquitination, and ultimately proteasomal degradation31,32. Based on these studies, we performed WB analysis of phosphorylated and total YAP in BC cells at 1 and 24 h after treatment with PA. The results showed that CD36 substantially inhibited p-YAP S127 level both at the cells levels at 1 h (Fig. 5G) and at the animal xenografts level (Fig. S5D). To further assess the effect of CD36 on YAP protein stability, we used cycloheximide (CHX) to inhibit new protein synthesis. We observed dramatic degradation of YAP in CD36-knockdowned groups compared to control groups. When applying a specific inhibitor of the 26S proteasome, MG132, the suppressive effects of silencing CD36 on YAP were mostly eliminated (Fig. S5E), indicating that YAP level was indeed maintained by CD36 through the proteasome-mediated protein degradation pathway. Further, CD36 remarkably increased the nuclear translocation of YAP, as shown by immunofluorescence (Fig. 5H; Fig. S5F) and nuclear extraction assays (Fig. S5H, I) whereas knockdown of CD36 reduced YAP nuclear signals (Fig. 5I; Fig. S5J). As expected, in vitro and in vivo assays showed that CD36 markedly promoted YAP signaling downstream genes (Fig. S5D, J, K), validating that CD36 induced the activation of the YAP pathway.
A Volcano plot exhibits different lymph node metastasis-associated molecules in rank correlation with CD36 expression in TCGA database. (|Rank correlation (Pearson)| > 0.1; p < 10E-5). B Immunohistochemistry staining of YAP in 77 BC specimens collected from Sun Yat-sen University Cancer Center. Three representative cases are shown. 50× magnification and 200×magnification; Scale bar: 100 μm and 50 μm. Correlation analysis of YAP expression with CD36 expression in BC specimens is shown in the upper left corner (Spearman). n = 77, R = 0.331, p = 0.003. C, D IHC staining (A) and quantification (B) of the expression of YAP from Xenograft Models tumors. Tumors formed by CD36-infected 231, vector-transfected 231 cells. Bars correspond to mean ± standard deviation (SD). E WB analysis of the expression of YAP in indicated CD36-infected, CD36-knockdown, and vector control cell lines treated with vehicle or palmitic acid (800 μm). F Relative mRNA level of YAP1 in indicated BC cell lines treated with vehicle or palmitic acid (800 μm) for 24 h. Bars correspond to mean ± standard deviation (SD), n = 3, n.s.: not significant. G WB analysis of total YAP and phosphorylated YAP S127 expression in the indicated CD36-infected, CD36-knockdown, and vector control cells treated with vehicle (Veh) or palmitic acid (PA) for 1 h. H, I Immunofluorescence staining of nuclear YAP expression in the indicated cells treated with palmitic acid (PA, 800 μm) for 24 h. BCs cells were stained with CD36 (red), YAP (green), and DAPI (blue). Scale bar: 50 μm. Bars represent the mean ± SD of three independent experiments; *p < 0.05, **p < 0.01, ***p < 0.001, n.s. not significant.
CD36 inhibits mitochondrial apoptosis via the binding of YAP-TEADs
To verify the vital role of YAP in CD36-induced LNM in BC, we conducted a popliteal LNM model for the rescue experiments. As shown in Fig. 6A, B, silencing YAP1 significantly decreased the volume and weight of PLN in 231-CD36 group. Moreover, the IVIS system and IF assays exhibited that the increased incidence of LNM induced by CD36 was substantially attenuated by knockdown of YAP1 (Fig. 6C, D; Fig. S6A). To Investigate whether the regulatory effect of CD36 on YAP protein stability is YAP-TEAD interaction-dependent, CHX chase and ubiquitination assay were performed. The results showed that CD36 overexpression attenuates ubiquitin-mediated degradation of YAP, which is rescued by verteporfin treatment, a pharmacological inhibitor that disrupts YAP-TEAD binding (Fig. S6B–D). Further, cellular rescue experiments to validate whether CD36 induced anoikis-resistance by activating the YAP pathway were applied. It showed that with knockdown of YAP1 or treatment with verteporfin, the inhibitor targeting the binding of YAP-TEAD, the ability of anti-anoikis and MMP maintenance was reduced significantly (Fig. 6E, F), but markedly recovered in CD36-silenced T-47D cells after YAP1 overexpression (Fig. 6G, H). Moreover, CD36-transfected 231 cells treated with YAP1 knockdown or verteporfin showed markedly elevated the release of Cyto-C from mitochondria to cytoplasm (Fig. 6I). Taken together, these data demonstrated that CD36-induced LNM and anoikis-resistance through the binding of YAP-TEAD.
A BC xenografts and popliteal lymph nodes from nude mice implanted with indicated 231 cells respectively. B Violin plot representing the volume and weight of each group lymph nodes. The results are mean ± SD. The number of each group was 4. C The BC cells that had metastasized to the homolateral popliteal lymph nodes were imaged by live imaging with IVIS system after luciferin injection, red arrows showing the metastasized BC cells which were labeled with luciferase beforehand. The number of each group was 4. D The contingency plot showed the frequency of lymph node metastasis in each group. n = 4 for each group (Chi-squared test). *p < 0.05, p = 0.1573. E−H Flow cytometric analyses (E, G) and JC-10 MMP detection (F, H) of MDA-MB-231 cells (E, F) and T-47D (G, H) apoptosis processed with indicated treatments under suspended palmitic acid (PA, 800 μm) conditions for 24 h. I MDA-MB-231 cells were fractionated to mitochondrial and cytosol subfractions after undergoing the indicated treatments. Cytochrome C (Cyto-C) protein in mitochondrial and cytosol subtractions were detected by WB assay. β-actin was loaded as a control to cytosol subfraction, Cox-IV was loaded as a control to mitochondrial subfraction. Bars =represent the mean ± SD of three independent experiments. *p < 0.05, ** p < 0.01, ***p < 0.001.
CD36 upregulates BCL2 expression via the direct transcriptional regulation of YAP-TEADs
Studies have shown that YAP promotes cell survival by regulating the expression of the Bcl-2 family members, including Bim and BMF33,34. However, the direct transcriptional regulation of Bcl-2 by YAP has not been reported. Using the GEO database, we found that Bcl-2 mRNA levels was remarkably higher in shNC group than that in shYAP1 group (Fig. S7A, B). To investigate whether YAP-TEADs acts as transcriptional cofactors - transcription factors (TFs) and directly mediates the up-regulation of Bcl-2 induced by CD36, WB and qPCR assays were performed. The results showed that the elevated Bcl-2 levels induced by CD36 were substantially inhibited after YAP knockdown or verteporfin treatment at both the protein and mRNA level (Fig. 7A, C). In contrast, Bcl-2 expression in T-47D-shCD36 cells with YAP overexpression was significantly restored compared with T-47D-shCD36 cells (Fig. 7B, D). Moreover, we predicted that BCL2 promoter region has multiple binding sites for TEADs in the JASPAR database (Table S2). Overexpression of YAP1 significantly increased BCL2 promoter-driven reporter activity, which was attenuated by verteporfin treatment (Fig. 7E). To identify the specific binding site, we mutated the BCL2-predicted site (PS) fragments according to the predicted site with the highest confidence (Fig. 7F; Fig. S7C). This mutation eliminated the YAP1-mediated increase in BCL2-PS promoter reporter activity (Fig. 7G). ChIP DNA enrichment was evaluated by PCR, which indicated that YAP could directly bind to the BCL2-PS fragments (Fig. 7H, I). Collectively, our results indicated that CD36 induces anoikis-resistance and LNM of BC through the activation of YAP-TEADs, which upregulates Bcl-2 by directly binding to the promoter of BCL2 gene.
A, B Effects of CD36 on increasing Bcl-2 were blocked after knockdown of YAP and treatment of Verteporfin or recovered after overexpression of YAP1. C, D Effects of CD36 on increasing BCL2 transcription were blocked after knockdown of YAP and treatment of verteporfin or recovered after overexpression of YAP1. E The sequence logo shows the predicted matrix profiles of the highest score. F Luciferase reporter assays of BCL2 promoter transcriptional activity. MCF-7 cells were transfected with YAP1-overexpressing plasmid or empty vector plasmid, BCL2 promoter-luciferase reporter plasmid, and Renilla luciferase plasmid for 48 h, followed by fluorescence detection. Renilla luciferase served as the transfection control. Verteporfin (10 μM) was designed to confirm the effect of YAP-TEADs-binding. G Luciferase reporter assays of mutant YAP-P3 promoter transcriptional activity. MCF-7 cells were infected with YAP1-overexpressing plasmid or empty vector plasmid, wild-type BCL2-PS promoter-reporter, and mutant reporter for 48 h, followed by fluorescence detection. (H) Chip assays were performed to verify BCL2 binding to the BCL2-PS promoter. Lane 1: BCL2 ChIP primer generated PCR product derived from 2% input DNA; Lane 2: BCL2 ChIP primer generated PCR product derived from immunoprecipitated by normal IgG; Lane 3: BCL2 ChIP primer generated PCR product derived from immunoprecipitation by an anti-YAP antibody; Lane 4: CTGF ChIP primer generated PCR product derived from immunoprecipitation by an anti-YAP antibody (I) Statistical analysis of Chip assays. Bars represent the mean ± SD of three independent experiments; *p < 0.05, **p < 0.01, ***p < 0.001.***p < 0.001.
CD36 activates YAP pathway by inducing bile acid synthesis
Previous studies have reported that lipid metabolites play an important role in regulating YAP activation, including fatty acids and bile acids20,30,35,36. Thus, we speculated that CD36 induces the activation of YAP through lipid metabolism. Taken the fatty acids receptor role of CD36 identified by studies and our fatty acids uptake assay (Fig. S8A), we performed transcriptomic and metabolomic analyses on the CCLE database to further explore the role of CD36 on lipid metabolism. The results showed that the expression of CD36 was significantly positively correlated with the cholesterol and bile acid pathways and their metabolites (Fig. 8A; Fig. S8B, C). Intriguingly, overexpression of CD36 in foot-pad xenografts significantly elevated the levels of cholesterol, bile acid, and free fatty acid in tumor tissue (Fig. 8B; Fig. S8D), but had no significant effect on triglyceride content (Fig. S8E). The key enzymes in cholesterol and bile acid synthesis are HMGCR and CYP7A1, respectively, and they are transcriptionally regulated by PPARα37, which is activated by fatty acids38,39 and abnormally activated in lymph node metastases20. Furthermore, WB assays revealed that the expression of PPARα, CYP7A1, and HMGCR were upregulated in CD36-transfected 231 cells, and were conversely attenuated in CD36-silenced T-47D cells (Fig. 8C, D). As expected, 231-CD36 xenografts also exhibited higher expression of PPARα, CYP7A1, and HMGCR than 231-EGFP xenografts (Fig. 8E). Recent studies have reported that bile acids inhibit the phosphorylation and ubiquitination degradation of YAP, inducing its nuclear translocation and activation of YAP20,35,36. To further verify whether YAP is regulated by the PPARα-HMGCR/CYP7A1 pathway induced by CD36, we first performed the correlation analysis in the TCGA database. GSEA results showed that the expression of YAP1 was positively correlated with the PPARα pathway (Fig. S8F), and the expression of CYP7A1, PPARA, and HMGCR (Table S3). In addition, by applying the PPARα inhibitor GW6471, the upregulation of YAP, HMGCR, and CYP7A1 induced by CD36 were substantially reduced (Fig. 8F). IF assays also exhibited that CD36-overexpressed 231 cells treated with GW6471 remarkably inhibited the nucleus translocation of YAP remarkably (Fig. 8G). Similarly, administration of the bile acid inhibitor obeticholic acid (an FXR agonist) also potently blocked this YAP nuclear activation induced by CD36 overexpression (Fig. S8G). Taken together, CD36 induced the activation of the Hippo YAP pathway by upregulating the PPARα-CYP7A1 pathway, and eventually bile acid synthesis.
A GSEA of the genes related to cholesterol storage and bile acids metabolism according to dichotomized CD36 expression (Median) in the TCGA database. NES: 1.8623657 & 1.7081771; Nominal p-value < 0.0001 & 0.006 respectively. B The concentration of total bile acid (TBA) and cholesterol of indicated BC xenografts lysis. C, D WB analysis of the expression of PPARα, HMGCR, and CYP7A1 from EGFP/CD36-infected MDA-MB-231 (C) and shNC/shCD36-infected T-47D (D) cells treated with vehicle (Veh) or palmitic acid (PA, 800 μm). (E) WB analysis of the expression of PPARα, HMGCR, and CYP7A1 from Xenograft Models tumors. F WB analysis of the expression of PPARα, HMGCR, and CYP7A1 in indicated MDA-MB-231 cells treated or no treated with PPARα inhibitor (GW6471, 25 µM). G Immunofluorescence staining of nuclear YAP expression in the indicated cells treated with palmitic acid (PA, 800 μm) for 24 h. BC cells were stained with CD36 (red), YAP (green), and DAPI (blue). Scale bar: 50 μm. *p < 0.05, **p < 0.01, n.s. not significant.
Discussion
CD36 is a membrane glycoprotein with multi-functional properties. By binding to different ligands, CD36 exerts diverse functions on tumorigenesis and development23,40. CD36 activation has been linked to tumor progression by transporting lipid ligands and utilizing lipid metabolism in several types of cancers. Although CD36 has also been reported for the pro-tumoral activity by promoting tamoxifen resistance41, immune suppression42, and motility43, some studies found that the loss of CD36 may contribute to BC indicating the contradictory anti-tumoral effect of CD3644,45. The present study systematically unraveled the function of CD36 in inducing anoikis-resistance and further promoting LNM in BC. Moreover, our results demonstrated that CD36 exerts its effect on anoikis-resistance through the activation of the YAP-TEADs pathway mediated by the up-regulation of bile acid synthesis. These findings identified the critical role of CD36 in LNM of BC, and provided mechanistic insights into exploring therapeutic strategies for metastatic stages in BC based on the lipid metabolic tendency of LNM.
Unlike hematogenous metastasis, LNM appears to rely on lipid metabolic adaptation according to the lipid-enriched lymphatic system28 (Fig. S1E–G). Study reported that a metabolic shift toward fatty acids oxidation occurrs in tumor cells during LNM, which is driven by the activation of the Hippo YAP pathway20. The lncRNA LNMICC promotes lymph node metastasis through fatty acid-binding protein 5-mediated fatty acid uptake in cervical cancer mouse models46. Likewise, our bioinformatic results showed that lipid metabolism pathways were more correlated with lymph nodes staging than glucose metabolism. Furthermore, several fatty acid transmembrane transport pathways exhibited significant correlations with LNM, and CD36 was scanned out according to correlation ranking among multiple FAs receptors (Fig. S1E-G). Further analysis of clinical specimens and the popliteal LNM model demonstrated that aberrantly elevated CD36 was correlated with LNM and poor prognosis (Fig. 1), and promoted LNM in BC (Fig. 2).
Studies have reported that the incidence of LNM in aggressive TNBC is lower than that in luminal BC5,6, which was also validated in the in vitro experiments, TCGA database and scRNA-seq dataset (GSE176078) (Fig. S2A-I). In our study, luminal T-47D cells exhibited higher CD36 expression and incidence of popliteal LNM than TNBC 231 cells (Fig. 2). In addition, research showed that PA significantly induces apoptosis in MDA-MB-231 cells47, which is consistent with our results, indicating that TNBC 231 cells fail to survive within the lipid-enriched lymph compared with luminial T-47D cells, and PA may be the vital apoptosis-inducing FAs in lymph (Fig. 3). Additionally, FAs metabolism was enriched in luminal BC and glycolysis metabolism was enriched in TNBC. Nevertheless, intravenous injections of TNBC cells present high incidence of metastasis in mouse model. Accordingly, we considered that some TNBC rely on aerobic glycolysis, namely the “Warburg effect”, and are inclined to hematogenous metastasis, rather than relying on lipid metabolism to drive LNM. Nonetheless, axillary lymph nodes involvement is still an independent risk factor for poor prognosis in TNBC48. Taken our in vivo experiments (Fig. 2; Fig. S2) together, these findings collectively suggest that while TNBC generally exhibits a preference for glycolytic metabolism and hematogenous metastasis, CD36 overexpression can confer a subset of TNBC cells with enhanced lipid metabolic adaptability, thereby promoting lymph node metastasis through utilization of lipid-rich lymphatic environment.
Emerging evidence highlights CD36’s multifaceted roles in shaping the tumor immune microenvironment. Recent studies demonstrate that CD36 orchestrates immunosuppression polarizing tumor-associated macrophages (TAMs) toward an M2 phenotype and facilitating their recruitment into tumor tissues49. Additionally, CD36 induces ferroptosis and exhaustion in CD8+ T cells, supports regulatory T cells (Tregs) survival and function, and attenuates dendritic cell-mediated anti-tumor immune responses of CD4+ T cells, collectively fostering an immune-evasive niche42,50,51. While previous studies emphasize CD36’s pro-tumoral effects via fatty acid (FA) metabolism-driven proliferation, invasion, and therapy resistance23, our findings unveiled a distinct FA metabolism-independent mechanism in BC (Fig. S8H): CD36 promotes lymphatic metastasis by inducing anoikis resistance via mitochondrial apoptosis regulation. Anoikis resistance is a prerequisite for metastasis and depends on the mitochondrial and death receptor apoptotic pathways18,19. Taken the MMP depolarization we observed in 231 cells treated with lymph (Fig. 3A) and the significant correlation between CD36 expression and mitochondrial-related pathways (Fig. S4A), our cellular assays further demonstrated that CD36 up-regulated Bcl-2 expression and attenuated mitochondrial apoptosis, eventually inducing anoikis resistance (Figs. 3, 4, Figs. S3, S4).
The Hippo pathway is an evolutionarily conserved signaling pathway participating in organ development and regeneration, wound healing, immune modulation, and tumor supression52. The Hippo pathway consists of a regulatory network to phosphorylated and inactive YAP, leading to its cytoplasm retention and degradation53. Dysregulation of the Hippo signaling was found in cancer, leading to the nucleus activation of YAP and triggering the expression of pro-tumorigenesis gene programs by binding to TFs, mainly the TEADs family54. YAP-TEADs pathway palys a vital role in inducing anoikis resistance55, reprogramming cancer metabolism30, and promoting LNM20,56 in various cancer including BC. Our study scanned out YAP as the strongest CD36-correlated oncogene involved in LNM (Fig. 5A). Further experiments demonstrated that CD36 inhibited the S127 phosphorylation and degradation of YAP and promoted the nucleus activation of YAP under PA condition. In addition, CD36 overexpression attenuates YAP ubiquitin-proteasome degradation, which is rescued by Verteporfin treatment (Fig. S6B–D). The above results were also validated in clinical samples and animal tumors (Fig. 5 and Fig. S5). Our research reveal that CD36 induces the activation of YAP signaling in BC cells within the FAs-rich lymphatic environment.
Recent studies have reported the regulation of YAP signaling via lipid metabolism. Fatty acids induce the nucleus translocation of YAP by releasing YAP from the β-catenin destruction complex57 and inhibiting YAP phosphorylation on S12758. PA also promotes the transcriptional activity of YAP through the F-actin pathway59 and the palmitoylation of TEADs60. In addition to fatty acids, bile acid is also critical for YAP activation35,36,61,62,63. Bile acid levels is aberrantly elevated in the metastatic LNs, inducing the nucleus activation of YAP in tumor cells20. However, the mechanism underlying the increased levels of bile acid in metastatic LNs remains elusive. Our study showed that CD36 promoted CYP7A1—the rate-limiting enzyme of bile acid synthesis—expression and up-regulated bile acid level (Fig. 8B–E). It has been reported that PPARα serves as an upstream TFs to regulate CYP7A1 expression37,64, and it can be activated by lipid ligands such as long-chain FAs38,39. Furthermore, the PPARα pathway is highly activated in LN metastases20, and is closely associated with metastasis in hepatocellular carcinoma65. Our study showed that CD36 facilitates the uptake of FAs (Fig. S8A) and activates the PPARα-CYP7A1 signaling (Fig. 8B–E). Subsequently, the significant positive correlation between YAP and the PPARα-CYP7A1 pathway was shown by analyzing the TCGA database (Fig. S8F, G). Moreover, our cellular assays demonstrated that by using PPARα inhibitor GW6471, the up-regulation of CYP7A1 and nucleus activation of YAP was substantially inhibited in BC cells treated with PA (Fig. 8F, G). Similarly, as bile acid synthesis inhibitors, FXR agonist obeticholic acid also attenuates CD36-induced YAP nuclear translocation in BC cells (Fig. S8G). Taken together, these findings provide insights into the increased level of bile acid and the activation of YAP induced by CD36 in BC cells within the lymphatic system. Moreover, since systemic inhibition of CD36 or YAP warrants cautions due to the potentially severe side effects and toxicity23,52, PPARα and CYP7A1 might be appealing targets for the treatment of BC.
The YAP pathway inhibits tumor cells apoptosis including anoikis and mitochondrial apoptosis by regulating the Bcl-2 family. In non-small cell lung cancer, YAP reduces the apoptotic response of cancer cells to chemotherapeutic drugs by inhibiting the expression of the pro-apoptotic factor BMF34. In addition, the expression of Bcl-2 and Mcl-1 are promoted by the activation of YAP when melanoma cells detached from ECM and acquire anoikis resistance66. However, the directly transcriptional regulation of Bcl-2 by YAP-TEADs has not been reported. Our results show that CD36 promotes the transcriptional expression of BCL2 in BC cells, depending on the binding of YAP-TEADs within FAs-rich conditioins (Fig. 7A, B). Furthermore, luciferase reporter gene and ChIP assays indicated that YAP-TEADs directly bind to the BCL2 promoter region and identify their specific binding site (Fig. 7F-I). Our research supplements the regulatory network of YAP-TEADs signaling on the apoptotic pathway.
In conclusion, our study demonstrated that aberrantly elevated CD36 in BC facilitated the uptake of FAs and induced bile acid synthesis by up-regulating PPARα-CYP7A1 signaling, leading to the activation of the YAP-TEADs pathway, and eventually promoting anoikis resistance and LNM (Fig. 8H). In addition to identifying the vital role of CD36 on LNM, our research further strengthens the key role of lipid metabolism in LNM. These lipid metabolic characteristics of the metastatic process-dependent and CD36-PPARα-CYP7A1-YAP regulatory axis may provide an effective therapeutic window for preventing or reducing BC metastasis.
Conclusions
The present study demonstrates that abnormally elevated CD36 in breast cancer (BC) promotes lymph node metastasis (LNM) via the activation of Hippo-YAP signaling mediated anoikis-resistance. Mechanistically, CD36 facilitates the uptake of fatty acids and induces bile acid synthesis by up-regulating PPARα-CYP7A1 signaling, thereby triggering YAP-TEADs pathway. Collectively, the metabolic traits of LNM and identified CD36-PPARα-CYP7A1-YAP regulatory axis may provide therapeutic windows for preventing or reducing metastasis in BC.
Data availability
SEER data was downloaded from SEER database and extracted by SEER*Stat software. The TCGA BC data was downloaded from the LinkOmics database for analysis. The CCLE database was retrieved from BROAD Institute (https://portals.broadinstitute.org/ccle/data). The source data behind the graphs in the main manuscript can be found in “Supplementary Data file”. All data generated or analyzed during this study are included in this article and its supplementary information files.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J. Clin. 74, 229–263 (2024).
Lorusso, G. & Ruegg, C. New insights into the mechanisms of organ-specific breast cancer metastasis[J]. Semin. Cancer Biol. 22, 226–233 (2012).
Stacker, S. A. et al. Lymphangiogenesis and lymphatic vessel remodelling in cancer[J]. Nat. Rev. Cancer 14, 159–172 (2014).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation[J]. Cell 144, 646–674 (2011).
Lu, X. et al. Predicting features of breast cancer with gene expression patterns[J]. Breast Cancer Res. Treat 108, 191–201 (2008).
Harbeck, N. et al. Breast cancer[J]. Nat. Rev. Dis. Primers 5, 66 (2019).
Ferris, R. L. & Kraus, D. H. Sentinel lymph node biopsy versus selective neck dissection for detection of metastatic oral squamous cell carcinoma[J]. Clin. Exp. Metastasis 29, 693–698 (2012).
Morton, D. L. et al. Sentinel-node biopsy or nodal observation in melanoma[J]. N. Engl. J. Med. 355, 1307–1317 (2006).
Tseng, H. S. et al. Tumor characteristics of breast cancer in predicting axillary lymph node metastasis[J]. Med Sci Monit 20, 1155–1161 (2014).
Gnant, M. et al. Identifying clinically relevant prognostic subgroups of postmenopausal women with node-positive hormone receptor-positive early-stage breast cancer treated with endocrine therapy: a combined analysis of ABCSG-8 and ATAC using the PAM50 risk of recurrence score and intrinsic subtype[J]. Ann. Oncol. 26, 1685–1691 (2015).
Obenauf, A. C. & Massague, J. Surviving at a distance: organ-specific metastasis[J]. Trends Cancer 1, 76–91 (2015).
Nathanson, S. D. et al. Breast cancer metastasis through the lympho-vascular system[J]. Clin. Exp. Metastasis 35, 443–454 (2018).
Li, A. E. et al. A role for reactive oxygen species in endothelial cell anoikis[J]. Circ. Res. 85, 304–310 (1999).
Cao, L. et al. Mitogen-activated protein kinase pathway is pivotal for anoikis resistance in metastatic hepatoma cells[J]. Mol. Med. Rep. 9, 1121–1127 (2014).
Simpson, C. D., Anyiwe, K. & Schimmer, A. D. Anoikis resistance and tumor metastasis[J]. Cancer Lett. 272, 177–185 (2008).
Taddei, M. L. et al. Anoikis: an emerging hallmark in health and diseases[J]. J. Pathol. 226, 380–393 (2012).
Lu, J., Tan, M. & Cai, Q. The Warburg effect in tumor progression: mitochondrial oxidative metabolism as an anti-metastasis mechanism[J]. Cancer Lett. 356, 156–164 (2015).
Paoli, P., Giannoni, E. & Chiarugi, P. Anoikis molecular pathways and its role in cancer progression[J]. Biochim. Biophys. Acta. 1833, 3481–3498 (2013).
Kim, Y. N. et al. Anoikis resistance: an essential prerequisite for tumor metastasis[J]. Int. J. Cell Biol. 2012, 306879 (2012).
Lee, C. K. et al. Tumor metastasis to lymph nodes requires YAP-dependent metabolic adaptation[J]. Science 363, 644–649 (2019).
Laerke, H. N. et al. Effect of beta-glucan supplementation on acute postprandial changes in fatty acid profile of lymph and serum in pigs[J]. Int. J. Mol. Sci. 15, 13881–13891 (2014).
Rohrig, F. & Schulze, A. The multifaceted roles of fatty acid synthesis in cancer[J]. Nat. Rev. Cancer 16, 732–749 (2016).
Wang, J. & Li, Y. CD36 tango in cancer: signaling pathways and functions[J]. Theranostics 9, 4893–4908 (2019).
Liu, L. et al. TBL1XR1 promotes lymphangiogenesis and lymphatic metastasis in esophageal squamous cell carcinoma[J]. Gut 64, 26–36 (2015).
Kilkenny, C., Browne, W. J., Cuthill, I. C., Emerson, M. & Altman, D. G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research[J]. PLoS Biol. 8, e1000412 (2010).
Frisch, S. M. & Francis, H. Disruption of epithelial cell-matrix interactions induces apoptosis[J]. J. Cell Biol. 124, 619–626 (1994).
Harris, M. H. & Thompson, C. B. The role of the Bcl-2 family in the regulation of outer mitochondrial membrane permeability[J]. Cell Death Differ 7, 1182–1191 (2000).
Bergers, G. & Fendt, S. M. The metabolism of cancer cells during metastasis[J]. Nat. Rev. Cancer 21, 162–180 (2021).
Zanconato, F., Cordenonsi, M. & Piccolo, S. YAP/TAZ at the roots of cancer[J]. Cancer Cell 29, 783–803 (2016).
Zhang, X. et al. The role of YAP/TAZ activity in cancer metabolic reprogramming[J]. Mol. Cancer 17, 134 (2018).
Zhao, B. et al. A coordinated phosphorylation by Lats and CK1 regulates YAP stability through SCF(beta-TRCP)[J]. Genes Dev. 24, 72–85 (2010).
Yu, F. X. et al. Regulation of the Hippo-YAP pathway by G-protein-coupled receptor signaling[J]. Cell 150, 780–791 (2012).
Wang, L. et al. YAP and TAZ protect against white adipocyte cell death during obesity[J]. Nat Commun. 11, 5455 (2020).
Kurppa, K. J. et al. Treatment-induced tumor dormancy through YAP-mediated transcriptional reprogramming of the apoptotic pathway[J]. Cancer Cell 37, 104–122.e12 (2020).
Anakk, S. et al. Bile acids activate YAP to promote liver carcinogenesis[J]. Cell Rep. 5, 1060–1069 (2013).
Liu, R. et al. Conjugated bile acids promote invasive growth of esophageal adenocarcinoma cells and cancer stem cell expansion via sphingosine 1-phosphate receptor 2-mediated yes-associated protein activation[J]. Am. J. Pathol. 188, 2042–2058 (2018).
Marrapodi, M. & Chiang, J. Y. Peroxisome proliferator-activated receptor alpha (PPARalpha) and agonist inhibit cholesterol 7alpha-hydroxylase gene (CYP7A1) transcription[J]. J. Lipid Res. 41, 514–520 (2000).
Mottillo, E. P. et al. Genetically-encoded sensors to detect fatty acid production and trafficking[J]. Mol. Metab. 29, 55–64 (2019).
Nakamura, M. T., Yudell, B. E. & Loor, J. J. Regulation of energy metabolism by long-chain fatty acids[J]. Prog. Lipid Res. 53, 124–144 (2014).
Chen, Y. et al. CD36, a signaling receptor and fatty acid transporter that regulates immune cell metabolism and fate[J]. J. Exp. Med. 219, e20211314 (2022).
Liang, Y. et al. CD36 plays a critical role in proliferation, migration and tamoxifen-inhibited growth of ER-positive breast cancer cells[J]. Oncogenesis 7, 98 (2018).
Wang, H. et al. CD36-mediated metabolic adaptation supports regulatory T cell survival and function in tumors[J]. Nat Immunol 21, 298–308 (2020).
Zaoui, M. et al. Breast-associated adipocytes secretome induce fatty acid uptake and invasiveness in breast cancer cells via CD36 independently of body mass index, menopausal status and mammary density[J]. Cancers 11, 2012 (2019).
Declerck, Y. A. Desmoplasia: a response or a niche?[J]. Cancer Discov. 2, 772–774 (2012).
Seewaldt, V. L. Cancer: destiny from density[J]. Nature 490, 490–491 (2012).
Shang, C. et al. LNMICC promotes nodal metastasis of cervical cancer by reprogramming fatty acid metabolism[J]. Cancer Res 78, 877–890 (2018).
Hardy, S., Langelier, Y. & Prentki, M. Oleate activates phosphatidylinositol 3-kinase and promotes proliferation and reduces apoptosis of MDA-MB-231 breast cancer cells, whereas palmitate has opposite effects[J]. Cancer Res. 60, 6353–6358 (2000).
Houvenaeghel, G. et al. Axillary lymph node micrometastases decrease triple-negative early breast cancer survival[J]. Br. J. Cancer 115, 1024–1031 (2016).
Liu, S. et al. S100A4 enhances protumor macrophage polarization by control of PPAR-gamma-dependent induction of fatty acid oxidation[J]. J. Immunother. Cancer 9, e002548 (2021)..
Xu, S. et al. Uptake of oxidized lipids by the scavenger receptor CD36 promotes lipid peroxidation and dysfunction in CD8(+) T cells in tumors[J]. Immunity 54, 1561–1577.e7 (2021).
Oh, D. S. & Lee, H. K. Autophagy protein ATG5 regulates CD36 expression and anti-tumor MHC class II antigen presentation in dendritic cells[J]. Autophagy 15, 2091–2106 (2019).
Dey, A., Varelas, X. & Guan, K. L. Targeting the Hippo pathway in cancer, fibrosis, wound healing and regenerative medicine[J]. Nat. Rev. Drug Discov. 19, 480–494 (2020).
Yan, F. et al. The posttranslational modifications of Hippo-YAP pathway in cancer[J]. Biochim Biophys Acta Gen Subj 1864, 129397 (2020).
Cunningham, R. & Hansen, C. G. The Hippo pathway in cancer: YAP/TAZ and TEAD as therapeutic targets in cancer[J]. Clin. Sci. 136, 197–222 (2022).
Zhao, B. et al. Cell detachment activates the Hippo pathway via cytoskeleton reorganization to induce anoikis[J]. Genes Dev. 26, 54–68 (2012).
Shen, J. et al. Hippo component YAP promotes focal adhesion and tumour aggressiveness via transcriptionally activating THBS1/FAK signalling in breast cancer[J]. J. Exp. Clin. Cancer Res. 37, 175 (2018).
Azzolin, L. et al. YAP/TAZ incorporation in the beta-catenin destruction complex orchestrates the Wnt response[J]. Cell 158, 157–170 (2014).
Ye, J. et al. JCAD promotes progression of nonalcoholic steatohepatitis to liver cancer by inhibiting LATS2 kinase activity[J]. Cancer Res. 77, 5287–5300 (2017).
Deng, Y. et al. Yap1 plays a protective role in suppressing free fatty acid-induced apoptosis and promoting beta-cell survival[J]. Protein Cell 7, 362–372 (2016).
Chan, P. et al. Autopalmitoylation of TEAD proteins regulates transcriptional output of the Hippo pathway[J]. Nat. Chem. Biol. 12, 282–289 (2016).
Chiang, J. Y. Bile acid metabolism and signaling[J]. Compr. Physiol. 3, 1191–1212 (2013).
De Boer, J. F. et al. New insights in the multiple roles of bile acids and their signaling pathways in metabolic control[J]. Curr. Opin. Lipidol. 29, 194–202 (2018).
Hegyi, P. et al. Guts and gall: bile acids in regulation of intestinal epithelial function in health and disease[J]. Physiol Rev. 98, 1983–2023 (2018).
Kanehisa M. The KEGG Database[J]. https://www.genome.jp/kegg/ (2022).
Li, J. et al. CD147 reprograms fatty acid metabolism in hepatocellular carcinoma cells through Akt/mTOR/SREBP1c and P38/PPARalpha pathways[J]. J Hepatol. 63, 1378–1389 (2015).
Zhao, B. et al. YAP activation in melanoma contributes to anoikis resistance and metastasis[J]. Exp. Biol. Med. 246, 888–896 (2021).
Acknowledgements
This study was supported by The National Natural Science Foundation of China (Grants 82273116, 82203661, 82070888, 82070882, 82100917, 81872165); Key Project of Nature Science Foundation of Guangdong Province, China (Grant 2019B1515120077); National Key R&D Program of China (Grant 2018YFA0800403); Guangdong Special Support Program for Young Top Scientist (Grant 201629046); Guangdong Natural Science Fund (Grant 2019A1515011810, 2021A1515010434, 2022A1515012423, 2023A1515010214 and 2024A1515010149); Fundamental Research Funds for the Central Universities (Grant 50000-31620106); China Postdoctoral Science Foundation (Grant 2021M703679); China Postdoctoral Science Foundation (2022M713594; BX20220360); Guangdong Provincial Health and Health Commission Medical Research Foundation (A2022161); Guangdong Basic and Applied Basic Research Foundation (2022A1515111062; 2022A1515111199); General project of Shenzhen Science and Technology Innovation Commission (JCYJ20220530145003008); Sun Yat-sen University College Student Innovation Training Program (202210661). We also thank Prof. Erwei Song and Haihe Wang who generously shared the BC cell lines for experimentation.
Author information
Authors and Affiliations
Contributions
Xia Yang and Guoquan Gao were responsible for designing and supervising the entire study and revising the manuscript. Zhijian Huang, Xiaoyan Zhang, and Jiajia Huang, and Xiaojun Liu performed research and analyzed the data. Tianxiao Gao, Mingting Liang, Fengyu Yang, Jieye Huang, Gan Xia, and Jiayin Lu contributed with in vivo studies, data analysis. Jizhao Cao, Ti Zhou, Weiwei Qi, and Anton Gao collected the medical records and human tissues and contributed to the discussion.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Marina Holz and Mengtan Xing.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, Z., Zhang, X., Huang, J. et al. Elevated CD36 drives lymph node metastasis of breast cancer by activating Hippo-YAP signaling-mediated anoikis resistance. Commun Biol 9, 80 (2026). https://doi.org/10.1038/s42003-025-09346-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09346-4