Abstract
Pigs are vital to global agriculture, and infectious diseases cause significant economic losses. Leukocytes provide a critical window into the genetic regulation of pig immune traits. However, understanding of these mechanisms within specific immune cell types remains insufficient. Here, we integrate 11 immune traits and systematically map the regulatory landscapes of expression quantitative trait loci (eQTLs), splicing QTLs (sQTLs), and alternative polyadenylation QTLs (apaQTLs) in porcine peripheral blood mononuclear cells (PBMCs) and neutrophils to uncover cell type-specific patterns. These molecular QTLs (molQTLs) exhibit strong cell-type specificity and preferentially regulate genes involved in cross-cell communication that are linked to core immunity, thereby shaping immune phenotypes through intercellular networks. Furthermore, we identify 588 molQTLs that colocalize with genome-wide association study signals for phagocytic capacity. Among these, 60.3% of apaQTLs independently modulate immune traits, including the variant rs330263631. Experiments confirm that rs330263631 modulates mRNA stability and expression levels of the TXNDC15 by dynamically selecting polyadenylation sites and altering the length of the 3′ untranslated region. This work systematically delineates the PBMC- and neutrophil-specific genetic architecture underlying immune regulation in pigs and provides a molecular foundation for deciphering the genetic mechanisms of porcine immune traits.
Similar content being viewed by others
Introduction
As one of the most important agricultural animals globally, pigs account for 30% of global meat consumption and are considered the ideal biomedical model due to their physiological and immunological similarities to humans. However, the swine industry suffers annual economic losses exceeding $17 billion due to bacterial and viral infectious diseases1,2. Amid increasing restrictions on antibiotic use and growing demands for sustainable farming, enhancing the pig immune capacity and disease resistance has become an urgent priority for the industry.
Genome-wide association studies (GWAS) have emerged as a powerful approach to unravel the genetic regulation of immune traits in pigs3,4,5. Notably, most GWAS signals reside in non-coding regions6,7,8,9, exerting phenotypic effects through molecular mechanisms such as gene expression regulation (eQTLs), alternative splicing (sQTLs), and alternative polyadenylation QTLs (apaQTLs)10,11,12. The ongoing Pig Genotype-Tissue Expression (GTEx) project has identified thousands of molecular quantitative trait loci (molQTLs) from multiple porcine immune tissues (e.g., whole blood, spleen, and thymus), significantly advancing our understanding of complex trait regulatory networks13. However, existing studies using bulk tissues (e.g., whole blood) reveal that molQTLs explain less than 30% of the heritability for immune traits, suggesting critical roles for cell type-specific regulatory mechanisms and dynamic microenvironmental interactions in immune genetic regulation14.
Peripheral blood serves as a critical window for monitoring host immune dynamics, and the immunological parameters it contains are important indicators for assessing immune competence and disease resistance15. Among these, hematological parameters such as white blood cell count and the proportions of lymphocytes and neutrophils (Neu) are commonly used as biomarkers of pathological or subpathological states. In addition, circulating immune factors, including interleukins (IL), interferons (IFN), and tumor necrosis factor-alpha (TNFα), play essential roles in antiviral defense, antitumor activity, and immune regulation16. These indicators are directly influenced by the composition and functional state of peripheral blood immune cells. Based on morphological and density characteristics, immune cells in peripheral blood can be classified into mononuclear cells and polymorphonuclear leukocytes. Peripheral blood mononuclear cells (PBMCs), encompassing various mononuclear cell populations, are easily accessible and widely used in swine immunology research because they systematically reflect immune regulation, inflammatory responses, and tissue repair processes17,18. In contrast, neutrophils (Neu), the most abundant polymorphonuclear leukocytes in circulation (approximately 70%), execute rapid defense mechanisms such as phagocytosis and NETosis19,20. However, their short half-life (6–8 h), high RNase activity, and technical challenges in isolation have hindered functional studies, as traditional single-cell sequencing methods often fail to adequately capture their biological characteristics21,22. Therefore, this study aims to systematically identify molQTLs in PBMCs and Neu to decipher the cell type-specific genetic regulatory mechanisms underlying key swine immune traits, such as blood parameters, cytokine levels, and phagocytic capacity.
Here, we performed multi-dimensional genetic profiling of PBMCs and neutrophils from 134 healthy Yorkshire pigs, systematically mapping eQTLs, sQTLs, and apaQTLs. Integration of co-expression networks with immune traits revealed that cell-specific molQTLs target genes associated with lymphocyte/Neu proportions and intercellular communication. Enrichment analysis showed that PBMCs and Neu molQTLs are significantly enriched in genomic regions associated with porcine immune traits (P = 1.2 × 10⁻⁵). Colocalization analysis further identified 588 molQTLs showing significant colocalization with phagocytosis capacity GWAS signals (PPH4 > 0.7). Notably, over 60% of apaQTLs, including rs330263631 targeting the TXNDC15, demonstrated independent regulatory functions. 3′ rapid amplification of cDNA ends (3′RACE) experiments confirmed that rs330263631 regulates TXNDC15 expression levels by dynamically selecting its polyadenylation sites. Importantly, TXNDC15 has been established to participate in redox homeostasis and anti-apoptotic processes during immune regulation. In summary, our study provides crucial resources for deciphering the genetic basis of porcine immune traits.
Result
Data analysis summary
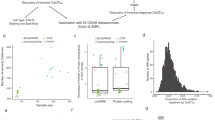
This study aimed to systematically dissect the cell type-specific genetic regulation of porcine immune traits using a multi-omics framework, as outlined in Fig. 1A. Whole genome sequencing data with an average coverage depth of 11.81 ± 1.5× from all whole-blood samples were analyzed. A total of 14,757,757 high-quality polymorphic SNPs were retained for downstream analyses, with SNP distribution across chromosomes illustrated in Fig. 1B. For transcriptomic profiling, RNA-seq data from peripheral blood mononuclear cells (PBMCs) and neutrophils (Neu) yielded an average of 20,974,970 and 20,510,516 mapped reads per sample, respectively, with mapping rates of 86.0% and 77.0% (Supplementary Fig. 1A). Following rigorous filtering (removal of samples from males, samples with low-mapping-rates, and samples with genotype concordance below 90%) (Supplementary Fig. 1B), from paired samples collected from 158 Yorkshire pigs, 134 PBMC samples and 125 Neu samples were retained after quality control for subsequent analysis (Supplementary Data 1). Molecular phenotypes were defined using strict criteria, including 10,833 genes, 10,396 alternative splicing clusters, and 10,621 polyadenylation (polyA) sites, which were subjected to molecular quantitative trait loci (molQTLs) mapping. Principal component analysis (PCA) revealed pronounced divergence between PBMCs and Neu across three molecular layers: gene expression profiles, alternative splicing patterns, and polyadenylation signatures. This separation underscores the distinct regulatory landscapes characterizing two immune cell types (Fig. 1C).
A The workflow of this study includes the detection of complete blood counts (n = 11) and immune markers (TNFα, IFNα, and IFNγ) for all samples, as well as the identification strategy for eQTL, sQTL, and apaQTL from 134 Peripheral blood mononuclear cells (PBMCs) and 125 Neutrophils (Neu) samples. We used a multi-omics association strategy to identify promising candidate genes and pathogenic variants. B The distribution of SNPs, gene expression levels, alternative splicing event (AS event), and alternative polyadenylation (APA) sites across the 18 autosomes of pigs. C Principal component analysis (PCA) based on gene expression levels, alternative splicing ratios, and APA site usage. Sample are colored by immune cell type (Neu, n =125; PBMC, n = 134).
Transcriptomic profiling of PBMCs and neutrophils unveils regulatory modules associated with pig immunological traits
To elucidate the transcriptional regulatory mechanisms underlying immune traits in pigs, we conducted transcriptome association analyses of 23,012 genes across 259 samples (134 PBMCs and 125 Neu) (Fig. 2A, B and Supplementary Fig. 2A). Our results revealed extensive genetic associations in PBMCs, with lymphocyte percentage (LYMPH%, 1851 genes, 8.04%) and white blood cell count (1213 genes, 5.27%) showing significant correlations (False Discovery Rate, FDR < 0.05; Fig. 2B and Supplementary Data 2). Notably, 27.3% of genes (6293 genes) in Neu exhibited specific associations with Neu proportion (Neu%, Fig. 2B, Supplementary Data 3), including the myeloid differentiation master regulator GFI1 (FDR = 1.44 × 10−8) and the effector molecule S100A12 (FDR = 1.73 × 10−5)23,24.
A Proportion of five immune cell types across 134 samples. B Genes significantly associated (FDR < 0.05) with partial immune traits in pigs (white blood cell count (WBC), neutrophil proportion (Neu%), lymphocyte proportion (LYMPH%), monocyte proportion (Mono%), IFNγ, and IFNα) as calculated by the linear regression model. Results for other immune phenotypes are provided in Supplementary Fig. S2 A. C Application of Weighted Gene Co-expression Network Analysis (WGCNA) to identify modules associated with immune traits in PBMCs and Neu gene expression profiles, with color indicating the strength of correlation between module genes and phenotypes (Neutrophil number: Neu_N, LYMPH number: LYMPH_N). D Immune trait-associated module genes and the results of their enrichment analysis (FDR < 0.05). Overlap genes indicates genes identified by both WGCNA and the linear regression model as associated with immune traits. E Hub genes associated with LYMPH% and Neu% are shown. F IL18 expression level is significantly correlated with Neu% (n = 124) (left panel); expression pattern of IL18 across all cell types, with TAU values indicating cell-type specificity (right panel).
Weighted gene co-expression network analysis (WGCNA) identified 17 distinct modules associated with immune parameters (ranging from 63 to 6432 genes) (Fig. 2C and Supplementary Fig. 2B–D). Specifically, modules M8, M12, M13, and M16 (PBMC-associated) showed correlation with LYMPH% and LYMPH_Number (LYMPH_N), whereas modules M1, M2, and M11 (Neu-associated) were correlated with Neu percentage (Neu%) and Neu absolute count (Neu_N) (Fig. 2C and Supplementary Fig. 2E). These modules demonstrated > 50% overlap with the linear model results (Fig. 2D). Functional analysis demonstrated that genes linked to LYMPH% and LYMPH_N were enriched in adaptive immune processes (e.g., Th17 cell differentiation and IL-17 signaling)25,26, whereas those associated with Neu% and Neu_N were predominantly engaged in innate immunity and TNF signaling (Fig. 2D), including key players such as the pro-inflammatory cytokine IL-18, chemokine CXCR2, and transcription factor RELA (Fig. 2E). Notably, IL-18 demonstrated a strong positive correlation with Neu% (P = 1.63 × 10−9) and has been established as a marker gene for Neu27,28 (Fig. 2F). Furthermore, we found that several conserved modules (M4, M5, M6) were enriched in fundamental metabolic processes and contained classical housekeeping genes, such as RPL13A and RPL4. These findings suggest that core metabolic functions are evolutionarily conserved across immune cell types, while functional specialization likely arises from a limited set of cell type-specific regulatory genes.
Genetic regulatory features of molecular quantitative trait loci (molQTLs) in pig PBMCs and neutrophils
To investigate the genetic determinants of multilayer transcriptional features in PBMCs and Neu, we systematically identified molecular quantitative trait loci (molQTLs), including expression QTLs (eQTLs), splicing QTLs (sQTLs), and alternative polyadenylation QTLs (apaQTLs). Specifically, we identified 3183 lead eQTLs, 2217 lead sQTLs, and 774 lead apaQTLs in PBMCs, and 2676, 1398, and 381 in neutrophils (Fig. 3A). Conditional analysis indicated that more than 10% of genes are associated with multiple independent molQTLs (Fig. 3B). Median cis-heritability estimates for gene expression, alternative splicing, and alternative polyadenylation (APA) were 0.18, 0.26, and 0.17, respectively (Fig. 3C), with sQTLs demonstrating the strongest regulatory effects, and Neu displayed a higher proportion of large-effect sQTLs (|effect size| ≥1; Supplementary Data 4), suggesting potent splicing regulation as a potential mechanism underlying immune plasticity.
A Number and overlap of eQTLs, sQTLs, and apaQTLs identified in PBMCs (n = 134) and Neu (n = 125). B Proportion of independent eQTLs, sQTLs, and apaQTLs in PBMCs and Neu as determined by conditional analysis. C cis-heritability of eQTLs, sQTLs, and apaQTLs in PBMCs and Neu. D Genomic distribution of eQTLs, sQTLs, and apaQTLs around transcription start sites (TSS), splice sites, and selective polyadenylation sites. E Genomic annotation categories and effect sizes of eQTLs, sQTLs, and apaQTLs identified in PBMCs and Neu. The horizontal axis represents the effect size of molQTLs. F eQTLs and apaQTLs showing opposite regulatory effects at the same genetic loci (n = 108) in PBMCs and Neu. G Impact of molQTLs (rs326112779 and rs322830411) on transcription factor (TF) motif sequences and their target gene expression.
Genomic enrichment analysis revealed that all identified molQTLs were significantly clustered near transcriptional start/termination sites and functional regions (including splice donor/acceptor sites and 3′UTRs) (Fig. 3D, E). Although 60.5% of eQTLs were located more than 100 kb from the transcription start site (TSS) of their target genes, only 5.8% of these distal eQTLs exhibited large effect sizes. This indicates that distal regulatory variants collectively influence gene expression through weak but coordinated mechanisms, whereas strong-effect eQTLs are preferentially enriched in core functional elements such as promoters and splice sites. Furthermore, we found that only 0.67%–1.14% of molQTLs were shared between PBMCs and Neu (Supplementary Fig. 3A), with the vast majority (> 80%) exhibited cell type-specific patterns. These cell type-specific QTLs were predominantly located in distal regions relative to the TSS and demonstrated stronger effect sizes than shared QTLs (Supplementary Fig. 3B). Meanwhile, we observed that 108 apaQTLs showed inverse regulatory directions compared to eQTLs at the same loci (Fig. 3F), suggesting that SNPs may regulate gene expression levels by altering the selection of polyadenylation sites and generating unstable mRNA subtypes (such as 3′UTRs shortening).
Enrichment analysis demonstrated that cell-specific molQTLs preferentially target binding sites of immune-related transcription factors (TFs), including IRF4, TCF4, and ZNF692, and modulate molecular phenotypes by disrupting or remodeling TF binding motifs (Fig. 3G and Supplementary Fig. 4A). Furthermore, molQTL regions were significantly enriched for RNA-binding protein (RBP) binding sites (e.g., PUM2, SRSF1; P < 0.01), with 3′UTRs length showing a positive correlation with RBP binding site density (Supplementary Fig. 4B, C).
Cell type specific molQTLs reveal genetic regulation of immune function and cross-cell communication
In this study, we defined the target genes regulated by significant eQTLs, sQTLs, and apaQTLs as eGenes (expression genes), sGenes (splicing genes), and apaGenes (alternative polyadenylation genes), respectively (FDR < 0.05). Specifically, 3189 eGenes, 2222 sGenes, and 959 apaGenes were found in PBMCs, compared with 2685, 1400, and 472 in Neu (Fig. 4A). Over 50% of these genes exhibited significant cell-type specificity. Functional enrichment analysis revealed that PBMC-specific eGenes were primarily involved in adaptive immune processes (e.g., T and B lymphocyte activation and antigen processing/presentation), while Neu-specific eGenes were significantly enriched in innate immune pathways (e.g., FcγR-mediated phagocytosis and endocytosis, FDR < 0.05; Fig. 4B).
A Numbers of overlapping and cell type-specific molGenes (eGenes, sGenes, and apaGenes) identified in PBMCs and Neu. B Functional enrichment analysis results of target genes regulated by PBMCs- and Neu-specific molQTLs. C Enrichment analysis for immune trait-associated eGenes in PBMCs and Neu. Significant eGenes were identified by overlapping results from the linear regression model and WGCNA (odds ratios calculated using Fisher’s exact test; * indicates significant enrichment: P < 0.05). D Cell type-specific regulatory networks linking phenotype-associated eQTLs, eGenes, and pathways. Representative SNPs (rsIDs) are shown, prioritized for their regulation of genes significantly associated with porcine neutrophil percentage (Neu%). Pathways are derived from enrichment analysis of WGCNA module genes. Arrow direction indicates the regulatory relationship; SNP colors denote the direction of effect (red: positive, green: negative). Genes are grouped by pathway (black circles: innate immune response; light brown circles: IFN signaling pathway). E Identification and characterization of cyto-cis and cyto-trans genes (defined in main text) in PBMCs and Neu, along with their regulating eQTLs. The upper panel shows the proportion of cyto-cis eGenes associated with immune phenotypes in each cell type, and the lower panel shows the corresponding proportion for cyto-trans eGenes. Color intensity represents the proportion of phenotype-associated eGenes.
Further analysis revealed significant overlap (P < 0.05) between these cell-specific molGenes and co-expression module genes corresponding to immune cell proportions, including multiple Neu%-related genes involved in innate immune response, such as OAS2 (rs337766841), VNN1 (rs322713915), and TRIM21 (rs346057197) (Figs. 4C, D). For instance, PBMC-specific eGenes overlapped with modules (M8, M12, M13, and M16) related to LYMPH% and LYMPH_N, whereas Neu-specific eGenes overlapped with modules (M1, M2, and M11) associated with Neu% and Neu_N (Upper-left panel). Notably, most overlapping genes were highly expressed in their respective immune cells (defined as cyto-cis genes)29, suggesting that genetic variants differentially regulate immune functions through cell type-specific regulatory networks (Fig. 4E, Upper-right panel).
Moreover, we identified several atypical immune cell percentage associated transcriptional modules within the co-expression network. Specifically, the PBMC-specific module M3 demonstrated a significant positive correlation with Neu% (Pearson’s r = 0.45, P = 1.73 × 10−5), while the Neu-specific module M10 showed a notable association with LYMPH% (Pearson’s r = 0.40, P = 5.66 × 10−5) (Fig. 2C). Notably, over 40% of genes in these modules were regulated by eQTLs (Fig. 4E, lower left panel). Further analysis revealed that although 27.2% of module M3 genes (PBMCs group) and 30.3% of module M10 genes (Neu group) were associated with Neu% or LYMPH%, respectively, their expression levels were relatively low in the corresponding target cells (classified as cyto-trans genes) (Fig. 4E, lower right panel). These indicates that potential trans-cellular regulatory interactions among immune cells are primarily embedded within modules displaying atypical cross-cell associations.
Functional enrichment analysis further supported our speculation. We found that PBMC-derived cyto-trans genes were significantly enriched in the Toll-like receptor signaling pathway and phagocytosis-related pathways, suggesting their potential role in Neu regulation. Conversely, Neu-derived cyto-trans genes showed enrichment in apoptosis and the complement system (Supplementary Fig. 5), indicating that Neu may regulate lymphocyte function through mechanisms such as the secretion of regulatory mediators. These results demonstrate that molQTLs may mediate their regulatory effects on distal cell types through modulating genes involved in specific intercellular communication pathways, despite all cells being derived from the circulating blood.
Cross-immune tissue comparison highlights the strong association between cell specific molQTLs and immune traits
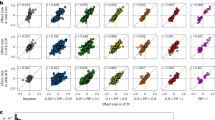
To systematically elucidate the shared patterns of genetic regulatory effects in porcine immune tissues, we integrated eQTL and sQTL data from 11 immune tissues in the pig GTEX project. This analysis identified 21,903 significant eQTL-gene pairs and 30,926 sQTL-intron cluster pairs (LFSR < 0.05). Our results revealed a high degree of concordance in regulatory direction across tissues: 51.5% of eQTLs (11,282/21,903) and 70.4% of sQTLs (21,771/30,926) showed consistent effect directions between any two immune tissues (Supplementary Data 5). The majority of molQTLs exhibited significant tissue specificity, with 66.5% of eQTL-eGene pairs and 67.2% of sQTL-sGene pairs detected in 1-2 tissues (Fig. 5A and Supplementary Fig. 6A, B). In contrast, a small fraction of eQTLs (4.8%) and sQTLs (3.3%) were shared across multiple tissues, with sQTLs demonstrating significantly stronger tissue specificity than eQTLs (Fig. 5A). Notably, functionally or anatomically related tissues preferentially shared molQTLs. Approximately 52.3% and 35% of eQTLs and sQTLs in PBMCs and Neu, respectively, were shared with functionally or anatomically related tissues, while functionally related tissues (e.g., whole blood) exhibited even higher sharing rates (up to 70%; Supplementary Fig. 6C). These findings suggest that the regulatory sharing patterns of molQTLs are closely linked to tissue physiological functions. Given the relatively modest overall contribution of sQTLs to gene expression and their pronounced tissue specificity (only 3.1% shared across tissues), we subsequently focused on elucidating the dynamic regulatory mechanisms of eQTLs to uncover the core genetic regulatory networks of the immune system.
A Proportion of eQTLs and sQTLs shared across 1-2, 3-8, and 9-13 tissues at LFSR < 0.05, based on data from 11 immune tissues in pig GTEx and PBMCs and Neu from this study. B Shared and tissue-specific eQTL–eGene pairs among PBMCs, Neu, and 11 other immune tissues. C Enrichment ratio of PBMC- and Neu specific eQTL target genes in porcine immune trait-associated genes (identified through overlapping results from linear regression model and WGCNA) compared with whole blood (error bars indicate 95% confidence intervals). D Enrichment ratio of PBMC- and Neu-specific eQTL target genes in porcine immune trait-associated genes (identified through overlapping results from linear regression model and WGCNA) compared with all other tissues (error bars indicate 95% confidence intervals). E Functional comparison of specific and shared genes in PBMCs and Neu compared with whole blood (FDR < 0.05). F Illustration of Neu-specific eQTLs (brown dots) and eQTLs shared with whole blood but exhibiting opposite regulatory effect directions (pink dots), with rs324196757 highlighted in red as an example. Inset shapes indicate the tissue/cell type of QTL discovery: circles for whole blood and triangles for Neu.
Although a substantial number of eQTLs were detected in whole blood and other immune tissues (10,227 and 12,557 pairs, respectively), numerous masked associations (8467 and 8768, respectively) were still identified through cell type-specific analyses, such as in PBMCs compared with Neu. Notably, approximately 52.5% of PBMC-specific and 55.3% of Neu-specific eQTLs were absent in bulk whole blood data (Supplementary Data 6) (Fig. 5B), highlighting the critical need for cell type-resolved QTL mapping to capture tissue-specific regulatory signals.
By categorizing eQTLs into cell type-specific (restricted to PBMCs/Neu) and cross-tissue shared groups, we uncovered distinct functional profiles. Compared to cross-tissue shared eGenes, cell type-specific molQTLs demonstrated significantly higher enrichment for immune hub genes, indicating cell type-specific molQTLs preferentially targeted immune hub genes associated with Neu% (e.g., NCF2, TLR8, CSF3R) or LYMPH% (e.g., CD6, PIK3R1, CD5) (Fig. 5C, D). Genes corresponding to these cell type-specific eQTLs exhibited stronger evolutionary constraints and larger effect sizes at the sequence level, and were enriched in core immune regulatory pathways (Supplementary Fig. 6D, E), including FcγR-mediated phagocytosis (involving MYO10, ARPC3) and apoptosis signaling (involving TUBA1B, MAP2K2) (Fig. 5E and Supplementary Fig. 6F). In contrast, cross-tissue shared eQTLs predominantly regulated fundamental metabolic processes such as arginine and proline metabolism (Fig. 5E and Supplementary Fig. 6F). We further identified tissue-specific inverse regulatory effects at key loci—for instance, the PRKCG variant (rs324196757) exhibited opposing eQTL directions in Neu compared with whole blood, modulating FcγR signaling to influence Neu phagocytic capacity (Fig. 5F). Collectively, these findings underscore the unique value of cell type-specific molQTL analyses in elucidating immune regulatory networks.
Integration of immune cell molQTLs with GWAS identifies variants and genes associated with phagocytosis
Our investigation revealed that cell type-specific molQTLs were significantly enriched in genomic regions associated with immune functions and hematological parameters (Fisher’s exact test, P = 1.20 × 10−5, Odds Ratio = 1.70), with 29 molQTLs overlapping these regions (Fig. 6A and Supplementary Data 7). These molQTLs likely influence host immune competence by modulating the abundance or functional states of key immune cells. For example, the PBMC-specific eQTL rs705486648 (3_46423678_T_C) that was significantly enriched in LYMPH% associated QTL regions and the Neu-specific eQTL rs81476484 (6_61200357_G_T) enriched in Neu% related QTL intervals. Notably, eight of these immune-trait-associated molQTLs exhibited significant overlap with GWAS signals for Porcine Reproductive and Respiratory Syndrome (PRRS) and African Swine Fever (ASF) (Supplementary Fig. 7A), including the PBMC-specific eQTL rs346036024 (12_59163357_T_C) that was enriched in Mono% related QTL regions, which is consistent with previous findings that monocytes/macrophages serve as key target cells for PRRSV replication and immune evasion, collectively suggesting that PBMCs- and Neu-specific molQTLs may constitute the potential genetic basis for disease susceptibility in swine populations.
A The PBMCs- and Neu- eQTLs, sQTLs, and apaQTLs show significant enrichment in QTL regions associated with porcine health and immune traits. QTL names are provided in Supplementary Data 7. B Examples of apaQTLs (rs330263631) colocalized with GWAS signals related to pig phagocytosis (PPH4 = 0.92). C Agarose gel electrophoresis of 3′RACE PCR products under different rs330263631 alleles (n = 4 independent biological samples). D The rs330263631 regulates the selection of long and short transcripts of TXNDC15 (analyzed by transcript-specific qPCR; left panel: T allele promotes long transcript expression; right panel: C allele promotes short transcript expression). Bar height represents the mean expression level of transcripts across four independent biological samples (n = 4) within each genotype group, with error bars indicating standard deviation. E The short transcript of TXNDC15 is expressed at a higher level than the long transcript in both CC and TT genotypes of rs330263631 (analyzed by transcript-specific qPCR; n = 4 independent biological samples). F The C allele at the rs330263631 locus may promote the generation of short 3′UTRs transcripts to increase mRNA levels, thereby ultimately regulating cellular phagocytic capacity, whereas the T genotype exhibits the opposite effect. The right panel in F shows representative images of phagocytosis assays for different genotypes: pink arrows indicate phagocytic cells, and blue arrows indicate apoptotic cells. The CC genotype demonstrates stronger phagocytic capacity, characterized by a greater number of phagocytic cells (pink) and fewer unengulfed apoptotic cells (blue).
Colocalization analysis revealed that 588 molQTLs (6.93%) in PBMCs and Neu showed significant colocalization (posterior probability for colocalization, PPH4 > 0.7) with GWAS signals associated with phagocytic capacity (Supplementary Data 8), a proportion that was significantly higher than the genomic background rate (binomial test, P < 1 × 10−5). This proportion was significantly higher than that observed in whole blood molQTLs (3.50%, 350 loci), demonstrating that cell type-specific molQTL data are more effective for identifying potential causal variants related to porcine immune traits. Among colocalized regions, 30.7% (181/588) showed significant associations with key immune phenotypes, including LYMPH% (15 loci), Neu% (10 loci), and IFNα levels (17 loci) (Supplementary Data 9), underscoring the utility of cell type-specific molQTLs in pinpointing functional SNPs. Notably, 60.3% of apaQTLs (38/63) independently influenced phagocytic traits without overlapping with eQTLs/sQTLs (Supplementary Fig. 7B), revealing the unique role of APA in immune regulation. For instance, the apaQTL rs330263631 (2_137069919_C_T) modulated APA events in TXNDC15, a critical regulator of antiviral defense, and showed strong correlations with diverse immune phenotypes (Fig. 6B and Supplementary Data 9)30,31.
To validate the APA-mediated regulatory mechanism, we performed 3′RACE experiments on TXNDC15. The results revealed that the apaQTL rs330263631 specifically influenced the polyadenylation site selection of TXNDC15 (Fig. 6C). Individuals carrying the C allele exhibited decreased PDUI values, indicating a preference for shorter 3′UTRs transcripts, whereas those with the T allele predominantly retained longer 3′UTRs isoforms (Fig. 6D and Supplementary Data 10). This genetic variation led to a 3.38-fold increase in the short/long 3′UTRs ratio in C allele carriers (Fig. 6E), consistent with enhanced usage of the proximal PAS. Across both genotypes, the expression levels of short 3′UTRs transcripts were higher than those of their long counterparts for the two genes examined (Fig. 6E and Supplementary Data 10), aligning with the expected mechanism whereby shorter 3′UTRs confer greater mRNA stability. Therefore, we hypothesize that the C allele at locus rs330263631 may promote preferential selection of the proximal polyadenylation signal (PAS), as indicated by reduced PDUI values, thereby increasing the production of short 3′UTR transcripts to elevate mRNA levels and consequently regulate cellular phagocytic capacity (Fig. 6F and Supplementary Fig. 7C).
Discussion
This study has constructed a genetic regulatory map for PBMCs and neutrophils, systematically revealing the cell type-specific architecture of genetic regulation and its associations with immune traits. Our results demonstrate that the cell-type specificity of genetic regulation is closely linked to cellular functional specialization and cross-cell communication mechanisms, providing novel insights into the genetic basis of porcine immune traits.
While interpreting our findings, it is also necessary to carefully define the boundaries and applicability of this study. First, the low proportion of male individuals in the analysis cohort may have limited our ability to fully capture sex-dimorphic molecular regulatory features related to porcine immune and disease traits32,33. Second, although the current sample size is sufficient to detect most moderate- to strong-effect genetic loci, the power to detect rare variants and weak-effect signals remains limited; and expanding the cohort size would enhance the resolution and statistical power of molQTLs detection. Third, this study primarily focused on two major immune populations, PBMCs and neutrophils. Consequently, the cell type-specific regulatory patterns identified here remain limited compared to the single-cell-resolution molQTLs analyses conducted in other species34,35. Finally, due to the lack of gene expression data before and after pathogen stimulation, we were unable to validate the role of the identified genetic loci in dynamic immune responses. Therefore, the targets discovered in this study are more applicable to genetic breeding for enhancing herd immunity. It is worth noting, however, that disease prevention targets may also hold therapeutic potential, as exemplified by genes such as IL18, GCLC, and CD200 in the context of Porcine Reproductive and Respiratory Syndrome36,37,38. Consequently, the causal genes and loci identified in this study require further experimental validation to clarify their specific regulatory roles in porcine disease traits.
Although several molQTLs in pigs have been reported recently13,39,40,41, systematic dissection of cell type-specific genetic architectures underlying immune traits, particularly in key innate immune cells like neutrophils, has remained scarce. To address this gap, we performed an integrated analysis of gene expression, alternative splicing, and polyadenylation in neutrophils and PBMCs. Our results revealed that immune cell-specific genetic variants are significantly enriched in distinct biological pathways: for instance, T and B cell activation and other adaptive immune pathways in PBMCs42, and FcγR-mediated phagocytosis in neutrophils43,44. Compared to the pig GTEx data, our study identified over one-third of novel eGenes, with the most notable findings in neutrophils. This discrepancy primarily stems from three aspects: first, tissue differences—pig GTEx primarily used whole blood, whose mixed cellular composition inevitably masks cell type-specific regulatory signals45,46; second, SNP coverage differences, although pig GTEx effectively detected eQTLs via RNA-seq, potential functional SNPs in non-transcribed regions may have been missed, thereby affecting the comprehensiveness of eQTL detection; and third, developmental stage differences—the pig GTEx cohort encompassed all growth stages, with adult individuals accounting for over 60%, whereas piglets constituted only 4%. Thus, future eQTL mapping in more refined tissue types and critical developmental stages will be an important direction.
Through genetic architecture analysis of multi-layered molecular phenotypes, we uncovered both similarities and differences across regulatory layers. Our study found that sQTLs exhibited the highest heritability and effect sizes among all molecular phenotypes, a phenomenon also reported in humans and other species47. This may be because pre-mRNA splicing is a process precisely catalyzed by the spliceosome and is more directly under strong cis-regulatory control48,49. In contrast, gene expression levels integrate multi-layered regulation from transcription initiation to mRNA degradation, making them more susceptible to interference from “environmental noise,” such as transient cellular states, resulting in relatively lower average heritability50. Notably, neutrophils demonstrated higher heritability across all molecular phenotypes compared to PBMCs. This can be explained by the intrinsic biological properties of these two cell types: Neutrophils are a terminally differentiated population whose core transcriptional programs are highly consolidated under homeostatic conditions and strictly governed by genetic instructions31. In contrast, PBMCs are a heterogeneous population comprising lymphocytes and monocytes, whose inherent cellular diversity introduces significant “environmental noise” into the analysis. This finding corroborates the diluting effect of cellular heterogeneity on heritability estimates observed in human studies51,52, demonstrating that in tissues with mixed cell types (such as whole blood), eQTL effects are masked by cellular heterogeneity, whereas cell type-specific analyses enable the detection of more numerous and stronger genetic effects. Therefore, our results suggest that studies based on mixed cell populations may systematically underestimate the true strength of genetic regulation within specific cell types.
Integrated analysis of GWAS and molQTLs colocalization helps elucidate potential causal genes for immune traits. In this study, we identified 694 cis-molQTL colocalized with GWAS signals related to phagocytic traits in both PBMCs and neutrophils, including LY9, CXCL10, FCMR, and PDCD1. Among these, LY9 functions as a costimulatory receptor that promotes intercellular adhesion and transduces activation signals, thereby positively modulating the immune responses and viability of immune cells53; CXCL10 recruits lymphocytes to synergize immune responses54,55; FCMR acts as a negative regulator in host immune modulation by suppressing the activation and migratory capacity of myeloid cells, including dendritic cells, monocytes/macrophages, and others56; and PDCD1 maintains self-tolerance and prevents excessive inflammation by eliciting inhibitory signals that terminate immune responses57. Additionally, we identified a novel candidate gene, TXNDC15, in PBMCs. This gene encodes a thioredoxin-like protein that may play a role in immune homeostasis by inhibiting RIPK1-dependent apoptosis and inflammatory responses58.
In summary, the genetic regulatory map we constructed provides a crucial resource for deciphering the molecular mechanisms of porcine immune cells and serves as a key resource for elucidating the genetic basis of complex immune traits. Our findings underscore the importance of dissecting genetic regulation within relevant cell types and offer new perspectives for deepening the understanding of the evolution and regulatory mechanisms of the porcine immune system.
Materials and methods
Sample information
The samples were randomly collected from 158 healthy Yorkshire pigs between 68 and 75 days old (average 70 days old, Supplementary Data 1). Health status was determined by experienced veterinarians based on the absence of clinical signs of disease (e.g., lethargy, anorexia, cough, or diarrhea) for a minimum of 2 weeks prior to blood collection, normal rectal temperature (<39.5 °C). All pigs (134 females and 24 males) were housed under similar environmental conditions, with identical feeding regimens and vaccination programs. Blood samples were collected via the anterior vena cava and were divided into EDTA anticoagulant tubes and serum separator tubes for subsequent analysis. The samples were transported in insulated boxes with cold packs and were delivered to the laboratory within 3 h of collection. Cell viability was determined by Trypan Blue exclusion using an automated cell counter and was consistently > 95% prior to downstream processing. We have complied with all relevant ethical regulations for animal use. All animal experiments were conducted in strict accordance with the regulations and guidelines established by the Animal Welfare Committee of China Agricultural University (permit number: DK996).
Blood parameter and cytokine measurement
Complete blood counts (CBC) and cytokine concentrations were measured for all samples. Specifically, a total of two mL of anticoagulated blood was collected for complete blood count (CBC). The CBC analysis provided a total of 11 immune cell parameters: the total white blood cell count (WBC), the percentages of the five leukocyte types (neutrophils, NEUT%; lymphocytes, LYMPH%; monocytes, MONO%; eosinophils, EO%; basophils, BASO%), and their respective absolute counts (NEUT_N, LYMPH_N, MONO_N, EO_N, BASO_N). Additionally, blood collected in plain tubes was left at room temperature for four hours, then centrifuged at 2000–3000 rpm for approximately 20 min to collect the supernatant. Using competitive double-antibody sandwich enzyme-linked immunosorbent assay (ELISA), three immune indicators (IFNα, IFNγ, and TNFα) were measured. Samples were diluted 1:5 and processed according to the manufacturer’s instructions, including standard curve construction, incubation, washing, color development, termination, and measurement steps. The absorbance of the test samples was compared to that of the standard, and the concentrations of the target analytes were calculated by fitting a four-parameter curve. Each sample was measured in triplicate.
Isolation of Peripheral blood mononuclear cells (PBMCs) and neutrophils (Neu)
Combining the physicochemical differences between PBMCs and Neu, PBMCs and Neu were simultaneously isolated from paired whole-blood samples collected from all samples. Specifically, four mL of EDTA-treated blood was diluted with PBS buffer and carefully layered with a cell separation solution to ensure a clear interface between the two liquids. The centrifuge was set to 15,000 rpm and spun for 10 min, causing the blood to separate into four distinct layers due to differences in cell density. The middle layer (the white cell layer) was collected, and the cells were resuspended and washed using phosphate-buffered saline (PBS) containing 10% fetal bovine serum (PBS with 10% FBS). The cells were resuspended, centrifuged at 450 × g at room temperature for 10 min, and the supernatant was discarded. This washing step was repeated two times to obtain purified PBMCs. Meanwhile, the lower cell layer was collected, and red blood cells were lysed using a red blood cell lysis buffer. The remaining cells were washed twice with 10% FACS buffer to obtain purified neutrophils.
RNA/DNA extraction and sequencing
Genomic DNA was extracted from blood samples using the TIANamp Genomic DNA Kit (TIANGEN, China) according to the manufacturer’s instructions. DNA quality and concentration were assessed using a Qubit 2.0 Fluorometer (ThermoFisher Scientific). A total of 158 samples were sent to Novogene Bioinformatics Technology Co., Ltd. (Beijing, China) for library preparation and shotgun genome sequencing. Briefly, sequencing libraries were constructed from 0.2 μg of genomic DNA per sample. The DNA was fragmented by sonication to an average size of 350 bp, followed by end-repair, A-tailing, adapter ligation, and PCR amplification. The qualified libraries were sequenced on the DNBSEQ-T7 platform to generate 150 bp paired-end reads. For data quality control, the raw sequencing reads in FASTQ format were processed using Fastp (v0.23.1) with the following criteria: (1) removal of reads containing adapter sequences; (2) removal of reads with more than 10% uncertain bases (N); and (3) removal of reads where over 50% of the bases had a Phred quality score below 5. A total of 158 samples that passed both the initial sample QC and this sequencing data QC were included in the subsequent analysis.
Total RNA was directly extracted from PBMCs and Neu samples using TRIzol reagent (Invitrogen Life Technologies). Before sequencing, RNA concentration was measured using a Qubit 2.0 Fluorometer (ThermoFisher Scientific). RNA integrity and concentration were assessed using the RNA Nano 6000 Assay Kit on the Bioanalyzer 2100 system (Agilent Technologies, CA, USA). Total RNA samples (307 samples, PBMCs = 157, Neu = 150) (meeting the quality control thresholds (total RNA > 200 ng and RNA Integrity Number > 7.0) were used for subsequent library construction. RNA sequencing libraries were prepared using the NEBNext® Ultra™ II RNA Library Prep Kit for Illumina® (NEB #E7775L) following the manufacturer’s instructions. The resulting libraries were sequenced on an Illumina NovaSeq X Plus platform with a 150 bp paired-end read configuration (PE150).
To mitigate potential confounding effects of sex, subsequent weighted gene co-expression network analysis (WGCNA) and molecular quantitative trait loci (molQTL) mapping were restricted to female individuals. The molQTLs analysis included expression QTLs (eQTLs), splicing QTLs (sQTLs), and alternative polyadenylation QTLs (apaQTLs), as detailed in the dedicated section below. After RNA quality control and preprocessing, the final analytical cohort comprised transcriptomes from 134 PBMCs and 125 neutrophil samples, all from females. All statistical analyses in this study were based on measurements from these independent biological replicates (n = 134 pigs).
DNA alignment and variant calling
First, the raw DNA sequencing data were processed using fastp (v0.19.4) with default parameters to remove adapter sequences, low-quality bases, and trim poly-G tails59. Next, sequencing reads were aligned to the pig reference genome (Sscrofa11.1) using the mem algorithm in BWA (v0.7.17). The alignment results were then sorted by coordinate using samtools (v1.18) to generate sample-specific sorted BAM files and their corresponding indices60. Duplicate reads were marked and removed from the BAM files using the MarkDuplicates command in Picard (v2.25.7)61, producing deduplicated BAM files.
Subsequently, base quality scores were recalibrated using the BaseRecalibrator command in GATK to improve the accuracy of downstream variant calling. Coverage analysis was performed on each sample’s BAM file using GATK’s DepthOfCoverage tool to calculate genome coverage at various depth thresholds. Variant calling was conducted using HaplotypeCaller, generating gVCF files for each sample. The resulting gVCF files were combined using the CombineGVCFs tool and genotyped with GenotypeGVCFs. Finally, SNPs with a missing rate below 10% and a minor allele frequency (MAF) greater than or equal to 5% were retained for further analysis.
Expression, splicing, and 3′UTRs quantification
We performed quality control on RNA sequencing data using fastp (v0.19.4) to remove adapter sequences, low-quality bases, poly-A tails, and poly-G tails59, and Ensemble gene annotation (Sscrofa11.1.v105) was conducted using STAR (v2.5.3a)62. Gene expression quantification was performed using Subread (v2.0.6) and StringTie (v2.2.1) to obtain raw read counts and TPM values18,63. Only genes with expression levels ≥ 0.1 TPM and at least 6 reads present in ≥20% of the samples were retained. The filtered expression data were inverse normal transformed and quantile-normalized for subsequent eQTL mapping.
To infer alternative splicing events, LeafCutter (v0.2.9) was used for the identification and quantification of splice variants64. The ‘prepare_genotype_table.py’ script was then used to calculate intron excision ratios, excluding introns utilized in fewer than 40% of individuals or showing no variation. The standardized and quantile-normalized intron excision ratios were used as percent spliced-in (PSI) values for each sample.
For inferring distal polyadenylation site usage ratios, the DaPars algorithm was employed to calculate polyadenylation site usage index (PDUI) values from RNA-seq data65. The PDUI value is defined as the expression level of the distal poly(A) site isoform divided by the total expression levels of both distal and proximal poly(A) site isoforms. In this study, we focused exclusively on APA events within the 3′UTRs regions and excluded events with fewer than 30 reads.
Identity by state (IBS) analysis
To ensure accurate sample-source matching, we initially performed SNP calling from transcriptomic data followed by IBS analysis to assess genotype concordance between RNA sequencing and whole-genome sequencing datasets. Only samples demonstrating high genotype consistency (IBS > 90%) were included in subsequent analyses (Supplementary Fig. 1A).
Gaussian mixture model determines the gene expression states of PBMCs and neutrophils
In this study, we used a Gaussian Mixture Model (GMM) to fit the log-transformed expression data of the PBMCs and Neu groups, assuming that gene expression states can be classified into two categories: high expression and low expression66. By fitting the model, we extracted the mean and standard deviation of the low-expression component for each cell type and calculated a conservative low-expression threshold based on the 95th percentile of the normal distribution. Finally, we determined the expression state (expressed or not expressed) of each gene in PBMCs and Neu based on this threshold.
Correlations between PBMCs/neutrophil gene expression and immune traits
In accordance with Johnson’s method67, we performed association analysis between gene expression and all immune traits using edgeR, with all validated technical covariates fully incorporated into the model (Supplementary Data 2 and 3). The analysis proceeded as follows: First, the raw count matrix was loaded into the edgeR object, and low-expression genes were filtered using the filterByExpr function. Next, the relationship between gene expression and all immune traits was modeled using the estimateDisp function, incorporating batch effects, RNA integrity number (RIN), and other technical covariates in the design matrix to control for potential confounding effects. The glmQLFit function was then used to fit a quasi-likelihood negative binomial generalized log-linear model, followed by a quasi-likelihood F-test (glmQLFTest) to assess the correlation between each gene and the immune traits. Finally, the Benjamini-Hochberg (BH) method was applied to correct the P values of all genes for multiple testing. For these significant associations, the model coefficient (β) was extracted to quantify the effect size and direction of the correlation.
WGCNA analysis
Weighted Gene Co-expression Network Analysis (WGCNA) was used to construct gene co-expression networks for PBMCs and Neu. Genes were assigned to specific modules based on their expression relationships. To ensure the networks adhered to a scale-free topology (R² > 0.8), a soft threshold power of β = 7 was selected. The topological overlap matrix (TOM) was calculated from the weighted adjacency matrix, and modules were defined using the hierarchical clustering dendrogram of the topological overlap, with a minimum module size of 35 and a merging threshold of 0.25. The module-trait association was analyzed using a comprehensive matrix of all measured traits. For clarity, this study primarily presents results for major immune cell percentages and key cytokines, as percentage data more accurately reflect immune cell composition. Complete analytical results for all traits are provided in Supplementary Fig. 2. The proportions of four immune cell types and the concentrations of three cytokines were used as trait matrices to assess the correlation between different modules. Hub genes for each module were then identified based on module membership (MM) > 0.8.
For each immune trait (e.g., lymphocyte proportion, cytokine levels), we independently selected genes using two approaches: (1) a linear model implemented in edgeR (False Discovery Rate, FDR < 0.05), and (2) WGCNA, retaining genes from significant trait-correlated modules with a module membership (MM) > 0.8. The final, high-confidence gene set for each trait was defined as the intersection of the genes identified by both methods. Correlations between module eigengenes and traits are reported as Pearson’s correlation coefficients (r) with associated exact p values.
Functional analysis of genes
We performed Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway and Gene Ontology (GO) enrichment analyses using the online tool KOBAS 3.0 (http://bioinfo.org/kobas/). The enrichment results were considered statistically significant at FDR of < 0.0568,69,70.
Mapping of molQTLs
We performed mapping for three types of molQTLs: eQTL, sQTL, and apaQTL. These analyses were performed using all chromosomes in pigs. To control for population effects, the first five principal components (PCs) of the genotypes were included in the molQTLs identification. To identify hidden factors and other biological influences, we performed principal component analysis (PCA) to uncover potential covariates affecting gene expression, splicing levels, and polyadenylation site usage71. In the analysis, different numbers of principal PCs were used as confounders for the gene expression, splicing levels, and polyadenylation site usage in the PBMCs and Neu groups. As PCs might not completely capture all technical variations, we included RIN as a covariate to control for biases introduced by differences in RNA quality72,73 (Supplementary Fig. 1C).
Subsequently, cis-molQTLs analysis was performed using the linear model in TensorQTL (v1.0.3), assessing the association between gene expression levels, intron excision rates, and distal74 polyadenylation site usage index within a ± 1 Mb window and SNPs39. The analysis incorporated the identified potential variables and known covariates, following the same pipeline as pig GTEX75. The linear model used in this study is as follows:
Y: Molecular phenotype (e.g., gene expression level, intron excision rate, polyadenylation site usage index), SNP: Genotype locus being tested (encoded as 0/1/2, representing allele dosage). \({{PC}}_{1}^{({geno})}\)-\({{PC}}_{5}^{({geno})}\): To control for confounding effects due to population structure and cryptic relatedness in the molecular phenotype association analysis, we included the top five principal components derived from genotype data in the linear model, which effectively capture allele frequency correlations resulting from shared ancestry. RIN: RNA integrity number. \({{PC}}_{1}^{({pheno})}\)-\({{PC}}_{m}^{({pheno})}\): m PCs extracted from phenotype data (expression/splicing/apa matrices) via PCA, included to account for hidden confounding factors (Supplementary Fig. 1D).
Additionally, the Ensemble Variant Effect Predictor (VEP) was used to compare the standardized molQTLs effect sizes with functional annotation categories, including non-coding regions, introns, alternative splicing, and 3’UTR regions76.
Significant molecular quantitative trait genes (eGenes, sGenes, apaGenes) were identified through a multi-step thresholding procedure. An initial FDR correction (BH method, FDR < 0.05) was applied to beta-approximated p-values. The significance boundary identified from this step was used to determine an empirical p-value threshold, which was subsequently converted to a gene-specific nominal p-value threshold based on the fitted Beta distribution parameters. Final significance required associations to pass both the nominal and empirical p-value thresholds. The statistical test was a two-sided linear regression.
Heritability estimation
The cis-heritability (h²-cis) for gene expression, splicing, and APA was estimated using the GREML (Genome-based Restricted Maximum Likelihood) method as implemented in the GCTA software77. Briefly, a genetic relationship matrix (GRM) was constructed from all common SNPs within the cis-window (±1MB). Variance components were then estimated using REML, with h²-cis calculated as the proportion of phenotypic variance explained by the cis-SNPs. The model included covariates to account for population stratification and technical factors.
Enrichment of molQTL target genes among immune trait-associated genes
To examine the enrichment of shared molGenes between immune trait-related genes and PBMCs or Neu, we tested the high-confidence immune trait-related genes identified by the intersection of significant associations (linear model, FDR < 0.05) and high network connectivity (WGCNA, MM > 0.8). These traits included lymphocyte proportion, neutrophil proportion, monocyte proportion, white blood cell count, and concentrations of TNFα, IFN-γ, and IFNα. We then performed a two-sided Fisher’s exact test (function in R) to analyze the enrichment of these immune trait-related genes among the PBMCs and Neu molGenes.
Comparison of PBMCs/neutrophil and Pig GTEx molQTLs with mash
We used the multivariate adaptive shrinkage (mash) method from the MashR package (v0.2.69) to evaluate the tissue-sharing patterns of PBMCs and Neu molQTLs with pig GTEx eQTLs and sQTLs75. To broadly assess the sharing of genetic regulatory effects, we compared our molQTLs to eQTLs and sQTLs from all 11 tissues available in the pig GTEx dataset. These included canonical immune tissues (e.g., spleen, lymph node, blood, macrophage) and gastrointestinal tissues (e.g., colon, duodenum, ileum, jejunum, large and small intestine), the latter being of interest due to the abundance of gut-associated lymphoid tissue and the importance of the mucosal immune system.
To further explore the regulatory effects of PBMC- and Neu-specific molQTLs, we classified the molQTLs into different categories: PBMCs- and Neu-specific molQTLs and shared molQTLs vs. blood, and PBMCs- and Neu-specific molQTLs and shared molQTLs vs. other immune tissues (excluding blood). We then analyzed various regulatory features of the specific molQTLs. We calculated the overlap between specific molQTLs target genes and cross-tissue shared molQTLs target genes with immune trait-associated genes and computed the 95% confidence intervals using a binomial distribution. To investigate the influence of molQTLs on gene regulation, we compared the genetic effect sizes between cell type-specific molQTLs and cross-tissue shared molQTLs. Furthermore, to evaluate the biological constraints on their target genes, we compared the probability of being loss-of-function intolerant (pLI) scores between genes targeted by cell type-specific molQTLs and those targeted by shared molQTLs.
Enrichment of molQTLs in immune trait-associated genomic regions
To investigate the enrichment of PBMC and neutrophil molQTLs within genomic regions associated with pig immune traits, we compiled two independent sets of immune-related loci: (1) 5009 health-related QTLs from AnimalQTLdb (categorized as blood parameters, disease susceptibility, immune capacity, and pathogens/parasites), and (2) 240 lead GWAS signals for 26 immune traits from the pig GTEx database (Supplementary Data 7). Enrichment of our molQTLs within these immune trait-associated regions was assessed using a two-sided Fisher’s exact test (using the fisher.test function in R), with the background set defined as all SNPs tested in our molQTL analysis (n = 18,217,567). All significantly enriched regions were subsequently visualized.
In addition, to statistically test whether cell type-specific molQTLs preferentially target immune hub genes compared to non-cell type-specific molQTLs, we performed an enrichment analysis. Specifically, we compared the fraction of immune-trait-associated genes within PBMC- or neutrophil-specific eGenes against the fraction within cross-tissue shared eGenes (which serve as a background set of non-cell type-specific regulatory events). Significance was assessed using Fisher’s exact test.
Colocalization of PBMCs and neutrophils molQTLs with GWAS summary statistics
We performed colocalization analysis between PBMCs and Neu molQTLs with published GWAS signals for porcine phagocytosis capacity78. The analysis was performed using the coloc R package (v.4.0.4) to determine whether the GWAS SNPs mediate their phenotypic associations by regulating molQTLs79. All analyses were conducted under the default prior settings. We interpreted the colocalization results based on the posterior probabilities for the five canonical hypotheses (H0-H4) tested by coloc. Specifically, the posterior probability for hypothesis H4 (PPH4) represents the probability that both traits share a single causal variant80. In this study, events with PPH4 > 0.7 were defined as successfully colocalized. Regional visualization was carried out using ggplot2, and the linkage disequilibrium (LD) between the identified causal SNPs and other SNPs was calculated using PLINK (v.1.90). To evaluate the statistical significance of molQTL-GWAS signal colocalization, we performed an enrichment analysis. We first calculated the baseline proportion of genome-wide background loci showing significant colocalization (PPH4 > 0.7) with the target GWAS trait. Using this baseline proportion as the random expectation, we conducted a binomial test under the null hypothesis that the observed colocalization proportion does not exceed the random expectation. The results of the binomial test are reported with the exact P value. For individual colocalization events, the posterior probability for hypothesis 4 (PPH4) is reported.
Experimental validation of APA events by 3′-RACE and qRT-PCR
To experimentally validate the polyadenylation mechanism regulated by apaQTL rs330263631, as suggested by colocalization analysis, we focused on TXNDC15, a gene involved in redox homeostasis and anti-apoptotic processes. 3′ rapid amplification of cDNA ends (3′RACE) was performed using a 3′RACE kit (Sangon Biotech, Shanghai, China) to identify different poly(A) sites of TXNDC15. Sanger sequencing was used to validate the qualified 3′RACE products. To quantify the functional impact of this APA event on transcript abundance, specific primers were designed for each transcript variant. Quantitative reverse transcription PCR (qRT-PCR) was performed, and relative expression levels were calculated using the 2− ΔΔCT method, with GAPDH as an internal reference gene. The sequences of the long and short 3′UTR isoforms of TXNDC15, along with GAPDH, are provided in Tables 1 and 2. Statistical comparison between groups was performed using a two-sided Student’s t-test.
Statistics and reproducibility
All statistical analyses were performed using GraphPad Prism 7.0 software. Error bars in the figures represent the standard error of the mean (SEM). The number of biological replicates is indicated by n in the figure legends. Statistical significance was defined as FDR < 0.05 (BH correction). Comparisons between two groups were performed using two-tailed t-tests. Correlation analysis results are reported as Pearson’s correlation coefficients (r). Enrichment analyses (e.g., gene set overlap) were assessed using two-sided Fisher’s exact tests. In omics analyses (e.g., eQTL mapping), FDR control was applied using the Benjamini-Hochberg method, with FDR < 0.05 considered significant. All analyses in this study were based on measurements from independent biological replicates (the final analytical cohort comprised n = 134 pigs).
Reagents
Oligonucleotide sequences used for 3′RACE and qRT-PCR validation are listed in Tables 1 and 2. Antibodies and kits used for cytokine measurement were as follows: Porcine IFNα ELISA Kit (Enzyme-linked Biotechnology Co., Ltd, #ml0023760, used at 1:5 dilution); Porcine IFNγ ELISA Kit (Enzyme-linked Biotechnology Co., Ltd, #ml002333, used at 1:5 dilution); Porcine TNFα ELISA Kit (Enzyme-linked Biotechnology Co., Ltd, #ml002360, used at 1:5 dilution). All cells analyzed were primary PBMCs or neutrophils isolated directly from study subjects as described in the PBMCs and neutrophil isolation section.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (RNA-seq: CRA032459, CRA032460, https://ngdc.cncb.ac.cn/gsa) and Genome Variation Map (WGS: GVM001201, GVM001202, https://ngdc.cncb.ac.cn/gvm/) at the National Genomics Data Center (NGDC), part of the China National Center for Bioinformation / Beijing Institute of Genomics, Chinese Academy of Sciences. Source data for Fig. 1B, C are available in the publicly accessible raw data. Data for Fig. 2 can be found in the publicly accessible raw data, Supplementary Data 2 and 3. Figure 3A–E source data are provided in Supplementary Data 4 and the raw data, while data for Fig. 3F–G are in Supplementary Data 5. For Fig. 4, panels A–C utilize data from Supplementary Data 5 and; panels D and E are derived from Supplementary Data 4. Figure 5 source data are available in Supplementary Data 6. Figure 6A data are provided in Supplementary Data 7; Fig. 6B data in Supplementary Data 8 and 9; and Fig. 6C–E data in Supplementary Data 10. All data are available from the corresponding author upon reasonable request.
Code availability
This study did not generate any new code. All the code used relied on publicly available software packages. Detailed analytical workflows are available from the corresponding author upon reasonable request.
References
Lunney, J. K. et al. Importance of the pig as a human biomedical model. Sci. Transl. Med. 13, eabd5758 (2021).
Uruen, C., Garcia, C., Fraile, L., Tommassen, J. & Arenas, J. How Streptococcus suis escapes antibiotic treatments. Vet. Res. 53, 91 (2022).
Ballester, M. et al. Genetic architecture of innate and adaptive immune cells in pigs. Front. Immunol. 14, 1058346 (2023).
Dauben, C. M. et al. Genome-wide associations for immune traits in two maternal pig lines. BMC Genom. 22, 717 (2021).
Roth, K. et al. Multivariate genome-wide associations for immune traits in two maternal pig lines. BMC Genomics 24, 492 (2023).
Liu, C. et al. Unveiling the genetic mechanism of meat color in pigs through GWAS, multi-tissue, and single-cell transcriptome signatures exploration. Int. J. Mol. Sci. 25, 3682 (2024).
Wang, X. et al. Gwas of reproductive traits in large white pigs on chip and imputed whole-genome sequencing data. Int. J. Mol. Sci. 23, 13338 (2022).
Wu, P. et al. A combined GWAS approach reveals key loci for socially-affected traits in Yorkshire pigs. Commun. Biol. 4, 891 (2021).
Zhang, Y. et al. Genetic correlation of fatty acid composition with growth, carcass, fat deposition and meat quality traits based on gwas data in six pig populations. Meat Sci. 150, 47–55 (2019).
Albert, F. W. & Kruglyak, L. The role of regulatory variation in complex traits and disease. Nat. Rev. Genet. 16, 197–212 (2015).
Olayinka, O. A., O’Neill, N. K., Farrer, L. A., Wang, G. & Zhang, X. Molecular quantitative trait locus mapping in human complex diseases. Curr. Protoc. 2, e426 (2022).
Umans, B. D., Battle, A. & Gilad, Y. Where are the disease-associated eqtls?. Trends Genet. 37, 109–124 (2021).
Teng, J. et al. A compendium of genetic regulatory effects across pig tissues. Nat. Genet. 56, 112–123 (2024).
Schmiedel, B. J. et al. Impact of genetic polymorphisms on human immune cell gene expression. Cell 175, 1701–1715 (2018).
Vosa, U. et al. Large-scale cis- and trans-eqtl analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
Heyer, C. M. et al. The impact of phosphorus on the immune system and the intestinal microbiota with special focus on the pig. Nutr. Res. Rev. 28, 67–82 (2015).
Ozanska, A., Szymczak, D. & Rybka, J. Pattern of human monocyte subpopulations in health and disease. Scand. J. Immunol. 92, e12883 (2020).
Sun, L., Su, Y., Jiao, A., Wang, X. & Zhang, B. T cells in health and disease. Signal Transduct. Target. Ther. 8, 235 (2023).
Herre, M., Cedervall, J., Mackman, N. & Olsson, A. K. Neutrophil extracellular traps in the pathology of cancer and other inflammatory diseases. Physiol. Rev. 103, 277–312 (2023).
Kolaczkowska, E. & Kubes, P. Neutrophil recruitment and function in health and inflammation. Nat. Rev. Immunol. 13, 159–175 (2013).
Scheel-Toellner, D. et al. Reactive oxygen species limit neutrophil life span by activating death receptor signaling. Blood 104, 2557–2564 (2004).
Summers, C. et al. Neutrophil kinetics in health and disease. Trends Immunol. 31, 318–324 (2010).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one fastq preprocessor. Bioinformatics 34, i884-i890 (2018).
Li, H. et al. The sequence alignment/map format and samtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The genome analysis toolkit: a mapreduce framework for analyzing next-generation dna sequencing data. Genome Res. 20, 1297–1303 (2010).
Dobin, A. et al. Star: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. Featurecounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Pertea, M. et al. Stringtie enables improved reconstruction of a transcriptome from rna-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Li, Y. I. et al. Annotation-free quantification of RNA splicing using leafcutter. Nat. Genet. 50, 151–158 (2018).
Xia, Z. et al. Dynamic analyses of alternative polyadenylation from rna-seq reveal a 3’-utr landscape across seven tumour types. Nat. Commun. 5, 5274 (2014).
Dubovik, T. et al. Interactions between immune cell types facilitate the evolution of immune traits. Nature 632, 350–356 (2024).
Johnson, K. E. et al. Human milk variation is shaped by maternal genetics and impacts the infant gut microbiome. Cell Genom 4, 100638 (2024).
Gene ontology consortium: going forward. Nucleic Acids Res. 43, D1049-D1056 (2015).
Bu, D. et al. Kobas-i: intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res. 49, W317–W325 (2021).
Kanehisa, M. & Goto, S. Kegg: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Zhou, H. J., Li, L., Li, Y., Li, W. & Li, J. J. Pca outperforms popular hidden variable inference methods for molecular qtl mapping. Genome Biol. 23, 210 (2022).
Gallego Romero, I. et al. RNA-seq: impact of RNA degradation on transcript quantification. BMC Biol. 12, 42 (2014).
Garrido-Martin, D., Borsari, B., Calvo, M., Reverter, F. & Guigo, R. Identification and analysis of splicing quantitative trait loci across multiple tissues in the human genome. Nat. Commun. 12, 727 (2021).
Taylor-Weiner, A. et al. Scaling computational genomics to millions of individuals with GPUs. Genome Biol. 20, 228 (2019).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Urbut, S. M., Wang, G., Carbonetto, P. & Stephens, M. Flexible statistical methods for estimating and testing effects in genomic studies with multiple conditions. Nat. Genet. 51, 187–195 (2019).
Hang, L. et al. The determination of porcine mononuclear/macrophage phagocytic capacity and estimation of genetic parameters. Chinese. J. of Anim. Sci. 58, 68–72 (2022).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Fraszczak, J. & Moroy, T. The transcription factors gfi1 and gfi1b as modulators of the innate and acquired immune response. Adv. Immunol. 149, 35–94 (2021).
Yao, Z. et al. Neutrophil infiltration characterized by upregulation of s100a8, s100a9, s100a12 and cxcr2 is associated with the co-occurrence of Crohn’s disease and peripheral artery disease. Front. Immunol. 13, 896645 (2022).
Futosi, K., Fodor, S. & Mocsai, A. Neutrophil cell surface receptors and their intracellular signal transduction pathways. Int. Immunopharmacol. 17, 638–650 (2013).
Hua, Z. & Hou, B. Tlr signaling in B-cell development and activation. Cell. Mol. Immunol. 10, 103–106 (2013).
Yang, T. et al. Transcriptome of porcine pbmcs over two generations reveals key genes and pathways associated with variable antibody responses post prrsv vaccination. Sci. Rep. 8, 2460 (2018).
Zheng, L. et al. The CD8α-πilralpha interaction maintains cd8(+) t cell quiescence. Science 376, 996–1001 (2022).
Itell, H. L., Humes, D. & Overbaugh, J. Several cell-intrinsic effectors drive type i interferon-mediated restriction of hiv-1 in primary cd4(+) t cells. Cell Rep. 42, 112556 (2023).
Shaheen, R. et al. Characterizing the morbid genome of ciliopathies. Genome Biol. 17, 242 (2016).
Klein, S. L. & Flanagan, K. L. Sex differences in immune responses. Nat. Rev. Immunol. 16, 626–638 (2016).
Traglia, M., Bout, M. & Weiss, L. A. Sex-heterogeneous snps disproportionately influence gene expression and health. PLoS Genet. 18, e1010147 (2022).
Yazar, S. et al. Single-cell eqtl mapping identifies cell type-specific genetic control of autoimmune disease. Science 376, eabf3041 (2022).
Wang, F. et al. Endothelial cell heterogeneity and microglia regulons revealed by pig cell landscape at single-cell level. Nat. Commun. 13, 3620 (2022).
Zhao, H. et al. Acquisition of different transcriptional shear mrna and biological function of porcine interleukin 18 binding protein in prrsv infection. mBio 15, e0064024 (2024).
Liu, X. et al. Xanthohumol inhibits prrsv proliferation and alleviates oxidative stress induced by prrsv via the Nrf2-Hmox1 axis. Vet. Res. 50, 61 (2019).
Elmore, M. R. et al. Respiratory viral infection in neonatal piglets causes marked microglia activation in the hippocampus and deficits in spatial learning. J. Neurosci. 34, 2120–2129 (2014).
Crespo-Piazuelo D. et al. Identification of transcriptional regulatory variants in pig duodenum, liver, and muscle tissues. GigaScience 12, giad042 (2022).
Xu, Z. et al. Integrating eqtl and genome-wide association studies to uncover additive and dominant regulatory circuits in pig uterine capacity. Animal 19, 101599 (2025).
Liu, Y. et al. Trait correlated expression combined with eqtl and ase analyses identified novel candidate genes affecting intramuscular fat. BMC Genomics 22, 805 (2021).
Zhang C. et al. Comprehensive genome and transcriptome analysis identifies slco3a1 associated with aggressive behavior in pigs. Biomolecules. 13, 1381 (2023).
Aleman, O. R. & Rosales, C. Human neutrophil fc gamma receptors: different buttons for different responses. J. Leukoc. Biol. 114, 571–584 (2023).
Aleman, O. R., Mora, N., Cortes-Vieyra, R., Uribe-Querol, E. & Rosales, C. Differential use of human neutrophil fcgamma receptors for inducing neutrophil extracellular trap formation. J. Immunol. Res. 2016, 2908034 (2016).
Yao, D. W., O’Connor, L. J., Price, A. L. & Gusev, A. Quantifying genetic effects on disease mediated by assayed gene expression levels. Nat. Genet 52, 626–633 (2020).
Zhang, J. & Zhao, H. Eqtl studies: from bulk tissues to single cells. J. Genet. Genomics. 50, 925–933 (2023).
Liu, S. et al. A multi-tissue atlas of regulatory variants in cattle. Nat. Genet. 54, 1438–1447 (2022).
Li, Y. I. et al. RNA splicing is a primary link between genetic variation and disease. Science 352, 600–604 (2016).
Khokhar, W. et al. Genome-wide identification of splicing quantitative trait loci (sqtls) in diverse ecotypes of Arabidopsis thaliana. Front. Plant Sci. 10, 1160 (2019).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Niu, F., Wang, D. C., Lu, J., Wu, W. & Wang, X. Potentials of single-cell biology in identification and validation of disease biomarkers. J. Cell. Mol. Med. 20, 1789–1795 (2016).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Kim-Hellmuth S. et al. Cell type-specific genetic regulation of gene expression across human tissues. Science. 369, eaaz8528 (2020).
Ogishi, M. et al. Human ly9 governs cd4(+) t cell ifn-gamma immunity to Mycobacterium tuberculosis. Sci. Immunol. 10, eads7377 (2025).
Meyer, L. et al. Identification of interferon-stimulated genes with modulated expression during hepatitis e virus infection in pig liver tissues and human heparg cells. Front. Immunol. 14, 1291186 (2023).
Hwang, S. et al. Interleukin-22 ameliorates neutrophil-driven nonalcoholic steatohepatitis through multiple targets. Hepatology 72, 412–429 (2020).
Kubli, S. P. et al. Fcmr regulates mononuclear phagocyte control of anti-tumor immunity. Nat. Commun. 10, 2678 (2019).
James, E. S. et al. Pdcd1: a tissue-specific susceptibility locus for inherited inflammatory disorders. Genes Immun. 6, 430–437 (2005).
Li, P. et al. A prospective study on the regulation of osteoarthritis risk through inflammatory pathways in clonal hematopoiesis. GeroScience https://doi.org/10.1007/s11357-025-01843-y (2025). Online ahead of print.
Acknowledgements
The authors would like to thank Ning Ding, Shifeng Tong, Hang Li and all the members of the Laboratory of Animal Molecular and Quantitative Genetics, China Agricultural University, for their support and assistance. Simultaneously, special thanks are extended to Senior Student Wenhui Guo from the team of Teacher Shuyang Yu at the College of Biological Sciences for her guidance in the operation and data analysis of flow cytometry experiments. In addition, Supported by High-performance Computing Platform of China Agricultural University. In addition, this study was funded by the National Key R&D Program of China (2023YFD1300400), Science and Technology Innovation 2030-Major Project (2023ZD04046, 2023ZD0407106).
Author information
Authors and Affiliations
Contributions
J.Y.Y. collected and analyzed all the data. S.Q.C. and Y.J.T. participated in results discussion and proved several important suggestions. L.L.W. and X.N.W. assisted in performing some of the experiments and gathered literature. H.T.L. contributed to the biological interpretation of results. F.P.M., Q.Y.Z., and K.X. provided important scientific input. Y.Y. and C.D.W. conceived and designed the study and revised the manuscript. Other authors participated in the sample collection. All the authors contributed to manuscript revisions, read and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Qiao Fan and Mengtan Xing.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, J., Chen, S., Tang, Y. et al. Integrated analysis of GWAS and molQTLs reveals cell-specific genetic variants in the porcine immune system. Commun Biol 9, 408 (2026). https://doi.org/10.1038/s42003-026-09605-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09605-y