Abstract
Serine racemase (SR) dysregulation associates with brain aging and Alzheimer’s disease (AD), as both a deficiency and an excess of D-serine can impact synaptic neurotransmission and the integrity of synapses. Neuronal SR decreases with aging, while glial SR is upregulated in AD. However, the role of SR in microglia involved in AD remains elusive. Here, Srr knockdown/knockout in microglia enhanced whereas overexpression of SR inhibited phagocytosis. Lipopolysaccharide-treated Srr−/− microglia upregulated anti-inflammatory factors—an effect blocked by histone lactylation inhibition. Conditional microglial Srr knockout (5×FAD;Lyz2cre;Srrfl/fl) improved spatial memory and reduced amyloid plaques (male-specific) in 5×FAD mice, with elevated lactylation of histone H3 lysine 18 (H3K18la), pyruvate kinase M2, and arginase1 in plaque-associated microglia. Cerebral D-amino acid oxidase and microglial SR and H3K18la were more prominent in males. Collectively, microglia-specific Srr deletion reprograms microglia toward an anti-inflammatory phenotype and enhanced phagocytic capacity partialy mediated by histone lactylation, thereby mitigating AD neuropathology and improving cognitive function—where sex-specific modulation of D-serine contributes to these beneficial effects. Overall, this study delineates the functional roles of microglial SR in phagocytosis, inflammatory responses, and learning-memory behaviors in AD-related models, thereby implicating microglial SR as a potential therapeutic target for AD.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is an irreversible neurodegenerative disorder characterized by the accumulation of misfolded proteins in both extracellular and intracellular environments within the brain, as well as significant loss of synapses and neurons1. The survival and death of neurons in AD brains, along with the underlying mechanisms, are central questions in the field of AD research. Chronic low-grade neuroinflammation, characterized by the activation of microglia—resident immune cells—plays a crucial role in neurodegeneration and significantly contributes to the progression of AD2,3,4,5.
Most of the microglia present in an unperturbed brain originate from the yolk sac during embryonic development and migrate to various regions both before and after birth, including the CNS and the retina, where they remodel synapses and maintain homeostasis within their resident niche6,7. With highly dynamic processes, microglia scrutinize the brain parenchyma for both endogenous and exogenous insults and cues8. Regarding their roles in AD, microglia participate in the elimination of amyloid β-peptide (Aβ) but also make contributions to senile plaques in both mouse models of AD and human AD brains9,10,11,12. Through the phagocytosis of extracellular Aβ, microglia limit the accumulation of amyloid plaques during the early stages of disease process13,14. However, in the later stages, they appear to shift towards immune tolerance, potentially due to metabolic reprogramming that favors aerobic glycolysis15,16. Besides surveying the CNS, microglia strengthen synapses by releasing neurotrophic factors17. In response to inflammatory stimuli, they also modulate synaptic activity through the release of excitatory neurotransmitters, including glutamate and D-serine, which is a product of serine racemase (SR)4,18,19.
D-serine is an agonist at the binding site on NMDA receptors commonly referred to as the “glycine site.” It is produced through chiral rearrangement of L-serine by serine racemase (SR), utilizing 5’-phosphate-pyridoxal as a cofactor20. The expression of SR is affected inversely during healthy aging versus the development of AD. For instance, SR expression decreases in neurons due to hypermethylation of the Srr promoter or depletion of nuclear factor erythroid 2-related factor 2 (Nrf2), a transcription factor that protects against oxidative stress, over the prolonged course of aging21,22. Conversely, in both animal models of AD and postmortem human AD brains, SR expression increases19,23. The incremental effects observed in AD brain are correlated with the induction of activator protein 1 (AP-1) in response to inflammatory stimuli in microglia24. Similarly, the expression of SR in neurotoxic astrocytes also increases25, potentially involving a comparable mechanism. A decrease in D-serine compromises long-term potentiation in aged rodents26, while an increase in D-serine due to amyloid accumulation impairs synaptic function in an AD-related animal model27. By collaborating with glutamate, SR exacerbates cell death by enhancing excitotoxicity through D-serine production19,28. Systemic deletion of SR mitigates neuronal death in animal models of ischemia29, AD30, and diabetic retinopathy31. However, SR is widely expressed in both neurons and glial cells, and the relevant cell type for the neuroprotection observed after SR deletion remains unclear. In neurons, SR expression is beneficial for neurotransmission and long-term potentiation26,32, whereas selective knockout of SR in astrocytes or microglia protect synapse and improve cognitive function in traumatic injury model33,34. In AD brain, reactive astrocytes with elevated levels of SR are neurotoxic25, while microglia release D-serine in response to Aβ induction, exacerbating neuronal death19,27,35. Here, we sought to further define the role of microglial SR in AD.
Results
Srr knockdown or deletion in microglia enhanced whereas overexpression of SR inhibited phagocytosis
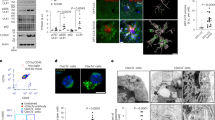
To study whether Srr knockdown alters phagocytosis of microglia, we established BV2 cells with stable knockdown of Srr. The BV2 cells that were infected with mCherry-expressing lentivirus expressing Srr-specific siRNA were screened based on knockdown efficacy. The cell lines exhibiting highest knockdown efficiency were designated as BV2 (336), while BV2 cells stably expressing scrambled siRNA served as control cell lines, referred to as BV2 (NC). The efficacy of transduction was validated with examination under fluorescence microscope, and the knockdown efficacy was validated with western blot against SR (Fig. 1A and Supplementary Fig. 1). By incubating BV2 cells with non-toxic levels of FITC-labeled Aβ oligomer (0.25 μM) for 36 h, we observed a significantly higher percentage of green fluorescence puncta in BV2 (336) cells than in BV2 (NC) cells (Fig. 1B). The percentage of green puncta in BV2 (336) cells peaked at 6 h and gradually decreased; however, the percentage of puncta remained higher than in BV2 (NC) cells, even at the end of observation period, 36 h (Fig. 1B). We further incubated BV2 cells with unlabeled Aβ oligomers (0.25 μM) for 48 h and examined with anti-Aβ immunofluorescence in the cells. Compared to BV2 (NC) cells, the fluorescence intensities of anti-Aβ immunofluorescence in BV2 (336) cells were much higher than those in BV2 (NC) cells at multiple time points, 12–36 h, although the difference was smaller at 48 h (Fig. 1C). We further examined the mechanism of elevated phagocytosis in BV2 (336). Compared to BV2 (NC) cells, BV2 (336) cells expressed higher levels of CD36 mRNA and protein (Supplementary Fig. 2A, B); however, knockdown of CD36 in BV2 (336) cells increased phagocytosis (Supplementary Fig. 2C–E). Furthermore, Beclin1 levels were higher in BV2 (336) cells compared to BV2 (NC), while LC3II and p62 levels were similar in the two lines (Supplementary Fig. 3A). This suggested that BV2 (336) cells had enhanced autophagy initiation without differences in maturation of autophagosomes into degradative autolysosomes. Additionally, Trem2 mRNA levels were not different in BV2 (336) cells compared to BV2 (NC) cells (Supplementary Fig. 3B). To investigate the impact of SR overexpression in microglia on phagocytosis, we utilized an SR-overexpression plasmid harboring an internal ribosomal entry site (IRES), which enabled independent expression of a downstream GFP reporter. Overexpression of SR inhibited phagocytosis monitor with fluorescently labeled microbeads (Fig. 1D, Supplementary Fig. 4, and Fig. 1E).
A BV2 cells with stable knockdown of Srr were established. Images were captured using LEICA DMi8 microscope system (LEICA) with a 10× objective. Bar: 50 μm for all panels. Lysates containing equal amounts of protein (20–40 μg) were subjected to western blot analysis to examine SR levels, with α-tubulin used as a loading control. Student’s t-test was employed for statistical comparison. SR protein levels in BV2 (336) cells transduced with lentivirus expressing Srr shRNA (n = 6) were significantly lower than those in cells transduced with lentivirus expressing scrambled shRNA (n = 6) (t[6.164] = 8.432, p = 0.00013). Representative blots for SR andα-tubulin were shown. The sample sizes (n) refers to the number of dishes used. B Non-toxic levels of FITC-Aβ (0.25 μM) were incubated with BV2 (NC) and BV2(336) for 6, 12, 24, and 36 h to examine Aβ phagocytosis. Images were captured. Bar: 50 μm for all panels. Student’s t-test was used to compare the differences in green puncta in BV2 cells at the indicated time points. Mann–Whitney U test was applied to compare the differences at the 24-h time point. The green puncta in BV2(336) (n = 6 at 6 h, n = 8 at 12 h, n = 6 at 24 h, n = 6 at 36 h) were higher than those in BV2(NC) (n = 8 at 6 h, n = 7 at 12 h, n = 5 at 24 h, n = 6 at 36 h) (t[12] = 7.826 at 6 h, p = 0.0000047; t[8.188] = 7.776 at 12 h, p = 0.0000047; p = 0.004 at 24 h; t[6.021] = 5.079 at 36 h, p = 0.004). C Aβo (0.25 μM) was incubated with BV2(NC) and BV2(336) cells for 6, 12, 24, 36, and 48 h. Aβ (green) and nuclei (blue) was co-stained to examine Aβ phagocytosis. Images were captured with a scale bar at 30 μm shown for all panels. Student’s t-test was used to compare differences in fluorescence intensities in BV2 cells at the indicated time points. Mann–Whitney U test was applied to compare the differences at the 6-h time point, where no significant differences were observed. The fluorescence intensities in BV2(336) cells (n = 3 at 12, 24, 36, and 48 h) were higher than those in BV2(NC) cells (n = 3 at 12, 24, 36, and 48 h) (t[4] =16.194 at 12 h, p = 0.000085; t[4] =11.964 at 24 h, p = 0.00028; t[4] =13.894 at 36 h, p = 0.00015; 48 h, t[4] = 4.558 at 48 h, p = 0.010). D Lysates of BV2 cells transfected with empty vector (IRES-eGFP) or an equal mass of Srr-expressing plasmid (Srr-IRES-eGFP) were subjected to western blot analysis to examine SR levels, with α-tubulin used as a loading control. Student’s t-test was employed for statistical comparison. SR protein levels in BV2 (Srr-IRES-eGFP) cells (n = 3) were significantly higher than those in BV2 (IRES-eGFP) cells (n = 3) (t[4] = 3.168, p = 0.034). Representative blots for SR and α-tubulin were shown. The sample sizes (n) refers to the number of dishes used. E Red fluorescent latex beads were incubated with BV2 cells transfected either with IRES-eGFP or Srr-IRES-eGFP plasmid for 2 h. The images of SR-expressing BV2 cells (green fluorescence), of beads (red fluorescence) and bright field were acquired. Beads⁺ and eGFP cells/eGFP cells (%) was defined as the percentage of eGFP-expressing cells that had phagocytosed the beads in the total eGFP-expressing cells, and the percentage was used to represent the phagocytic capacity of the cells. Images in columns 1–4 are presented with a scale bar of 100 μm, while column 5 displays magnified views of the squared areas in column 4 (scale bar = 50 μm). Student’s t-test revealed a significant difference between groups (t(24) = 6.889, p = 0.0000045). Error bars denote the variability of the data.
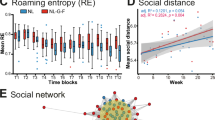
To confirm the findings in BV2 cell line, we employed microglia cultured from WT and Srr−/− mice. The microglia cultured from Srr−/− mice showed greater phagocytosis of Aβ oligomers compared to those cultured from WT mice (Fig. 2A). We further utilized fluorescently labeled microbeads to evaluate the phagocytic capacity of primary cultured microglia. Notably, Srr−/− microglia displayed significantly enhanced phagocytosis of microbeads compared with WT microglia (Fig. 2B).
A Primary microglia was cultured from the mice with systemic knockout of Srr (Srr−/−) or wild-type (WT). FITC-Aβ (0.25 μM) was incubated with microglia for 6, 12, 24, 36, and 48 h. Images were captured with a scale bar of 100 μm shown for all panels. Student’s t-test was used to compare the numbers of microglia phagocytosing FITC-Aβ at all time points, 6–48 h. At 6 h time point, the percentages of microglia phagocytosing FITC-Aβ in Srr−/− microglia (n = 14) were higher than those in WT microglia (n = 11), t[23] = 4.776, p = 0.0000081. Meanwhile, the FITC-Aβ intensity in Srr−/− microglia (n = 14) increased compared to WT microglia (n = 11), t[23] = 2.179, p = 0.04. At 36-h time point, there were more Srr−/− microglia (n = 15) which phagocytosed FITC-Aβ than WT microglia (n = 12), t[25] = 2.192, p = 0.038. No significant differences in the percentage of positive cells ingesting FITC-Aβ were observed between Srr−/− and WT microglia for 12, 24, and 48 h (Srr−/− microglia (n = 14), WT (n = 11), t[23] = 0.359, p = 0.723 at 12 h; Srr−/− (n = 13), WT (n = 13), t[24] = 0.714, p = 0.482 at 24 h; Srr−/− (n = 12), WT (n = 12), t[22] = 0.358, p = 0.723 at 48 h). FITC-Aβ intensities in Srr−/− microglia were not different from those in WT microglia for 12, 24, 36, and 48 h (t[23] = 0.890 at 12 h, p = 0.382; t[24] = 0.652, p = 0.521 at 24 h; t[25] = 0.898, p = 0.378 at 36 h; t[17.827] = 1.360, p = 0.191 at 48 h). BF, bright field. B Primary microglial cultures derived from Srr⁻/⁻ and wild-type (WT) mice were co-incubated with latex beads for 2 h. To assess microglial phagocytic capacity, images of red-fluorescent beads and corresponding bright-field views were acquired. Phagocytic activity was quantified as the percentage of microglia that had internalized the beads. All panels in column 1 were presented with a scale bar of 100 μm, whereas column 2 displayed magnified views of the squared areas in column 1 (scale bar = 50 μm). Student’s t-test was performed to compare phagocytic capacity between the two groups, and the results demonstrated that Srr⁻/⁻ microglia exhibited significantly higher phagocytic capacity than WT microglia (t(19.525) = 6.269, p = 0.0000045). Error bars denote the variability of the data.
Srr −/− microglia exhibited higher Arg1 and Il10 mRNA levels after LPS treatment
To further investigate the pathophysiology of microglia following the deletion of Srr, we tested inflammation-related responses in primary microglia from WT and Srr−/− mice. A Cre-loxP system generated to knockout Srr conditionally in microglia by crossing Lyz2cre mice with floxed Srr to delete the gene in lymphoid cells, including microglia. Neonatal offspring were used to produce mixed-glia cultures from which microglia were purified. LPS (25 ng/mL) was applied for 24 h, and RNA was harvested and subjected to RT-qPCR. Both WT and Srr−/− microglia were responsive to LPS treatment. The mRNA levels of Arg1 and Il10 were significantly higher in Srr−/− microglia compared to WT microglia after LPS treatment. In contrast, Tnf mRNA levels remained comparable between WT and Srr−/− microglia after LPS treatment (Fig. 3A–D). Studies indicate that aerobic glycolysis and histone lactylation, particularly at lysine 12 of histone H4 (H4K12), exacerbate microglia-mediated neuroinflammation in AD brain15,16. A study reported that lactylation of lysine 18 of H3 promotes Arg1 transcription36. We investigated whether Srr deletion modified the transcription of Arg1 and Il10 in microglia through histone lactylation. As histone lactylation can be mediated by p30036, we treated with LPS in the presence of the p300 inhibitor C646. To confirm that Arg1 induction in response to LPS is specific to histone lactylation and not acetylation, we tested the effects of oxamate, an inhibitor of lactate dehydrogenase A (LDHA). Both C646 and oxamate prevented LPS from elevating Arg1 mRNA levels in Srr−/− microglia higher than those reached in WT cells (Fig. 3E). Addition of C646 or oxamate before LPS treatment also blunted the induction of Il10 mRNA in Srr−/− microglia (Fig. 3F). Interleulin-1β (IL-1β) has been reported to regulate histone lactylation37. We next explored how Srr−/− microglia respond to IL-1β treatment. IL-1β treatment resulted in an increased trend of Il10 mRNA, albeit without significant difference (Supplementary Fig. 5).
The primary microglia were cultured from Srr−/− and WT mice, and treated with LPS (25 ng/mL) for 24 h at DIV 3–4 w. A After treatment, RNA was extracted from microglia and reverse transcription quantitative PCR (RT-qPCR) was performed. One-way ANOVA followed by Bonferroni test was used to compare Arg1 mRNA levels in WT or Srr−/− microglia; F(3, 8) = 40.362, p = 0.0000035. Arg1 mRNA increased under the treatment by LPS (n = 3) compared with sham treatment (n = 3) in Srr−/− microglia, p = 0.00011. Under LPS treatment, Arg1 mRNA levels in Srr−/− cells (n = 3) were more than those in WT cultures (n = 3), p = 0.000037. B Il10 mRNA levels were elevated under LPS treatment either in WT (n = 3 for sham or LPS treatment, p = 0.005) or in Srr−/− (n = 3 for sham or LPS treatment, p = 0.000028). Il10 mRNA levels were significantly higher in Srr−/− microglia compared to those in WT cells under LPS treatment, p = 0.005. One-way ANOVA followed by Bonferroni test was used for comparisons; F (3, 8) = 51.887, p = 0.000014. C Tnf mRNA levels under LPS treatment (n = 3) were higher than those with sham treatment in WT microglia (n = 3), p = 0.010. No differences observed in Srr−/− microglia between LPS and sham treatment (n = 3 for each), p = 0.671. One-way ANOVA followed by Bonferroni test was used for comparisons; F (3, 8) = 10.262, p = 0.004. D Nos2 mRNA levels were not different either in Srr−/− cells (n = 3) or WT cells under LPS treatment (n = 3 for each), p = 1.000. One-way ANOVA followed by Tamhane’s test was used for comparison; F (3, 8) = 5.401, p = 0.025. E Srr−/− and WT microglia were pre-treated with an LDHA inhibitor oxamate (Oxa., 10 mM) or an p300 inhibitor C646 (20 μM) for 4 h, then treated with LPS (50 ng/mL) for 24 h. RNA was extracted and RT-qPCR was performed to examine Arg1 and Il10 mRNA levels. One-way ANOVA followed by Bonferroni test was used to compare Arg1 mRNA levels in WT cells; F (3, 10) = 2.432, p = 0.125. One-way ANOVA followed by Bonferroni test was used to compare Arg1 mRNA levels in Srr−/− cells, F (3, 14) = 11.626, p = 0.000043. Arg1 mRNA levels in Srr−/− microglia increased under LPS treatment (n = 5 for sham or LPS treatment, p = 0.00003). Arg1 mRNA levels were significantly reduced with Oxa. (n = 4) or C646 (n = 4) pre-treatment compared to LPS treatment alone in Srr−/− microglial cultures; LPS+Oxa. vs. LPS, p = 0.029, LPS + C646 vs. LPS, p = 0.010. F Il10 mRNA levels were elevated in Srr−/− cells under LPS treatment (n = 5 for sham or LPS treatment, p = 0.032). One-way ANOVA followed by Bonferroni test was used to compare Il10 mRNA levels in WT cells; F (3, 12) = 1.632, p = 0.234. One-way ANOVA followed by Tamhane’s test was used to compare Il10 mRNA levels in Srr−/− cells, F (3, 15) = 6.935, p = 0.004. Error bars denote the variability of the data.
Since SR shows nuclear localization in neurons, we speculate that SR presence in the nucleus may interfere with p300. Indeed, SR immunofluorescence exhibited a strong overlap with p300; this overlap was significantly diminished by LPS treatment (Supplementary Fig. 6).
Conditional knockout of Srr in the microglia of 5×FAD mice increased spatial learning memory and improved brain pathology
To investigate the effects of specifically knocking out Srr in the microglia of AD-related mice, we crossed the Cre-loxP system described above into 5×FAD mice. Since most microglia originate from yolk sac hematopoiesis and are maintained through self-renewal without input from circulating monocytes, Lyz2-directed expression of Cre recombinase would be expected to permanently delete Srr in microglia. In contrast, bone marrow-derived myeloid cells are replaced within 4 weeks due to continuous turnover38. The 5×FAD;Lyz2cre;Srrfl/fl mice were subjected to the Morris water maze at 9–11 months of age, comparing them to controls that lacked the Cre transgene (5×FAD;Srrfl/fl). Lyz2cre significantly knocks down Srr in 5×FAD;Lyz2cre;Srrfl/fl mice (Fig. 4A). By days 5 and 6 of training, 5×FAD;Lyz2cre;Srrfl/fl females exhibited significantly less latency in finding the platform (Fig. 4B, C). In the probe trial on day 7, female 5×FAD;Lyz2cre;Srrfl/fl mice showed greater time in the target quadrant compared to female 5×FAD;Srrfl/fl mice counterparts (Fig. 4D). Deletion of Srr did not affect the frequency to cross platform (Fig. 4E), but increased locomotor activity in female mice (Fig. 4F, G). With regard to Morris water maze tests in male mice, 5×FAD;Lyz2cre;Srrfl/fl mice demonstrated a higher frequency of platform crossings compared to male 5×FAD;Srrfl/fl mice, although the residency time and escape latency were similar between the two genotypes (Supplementary Fig. 7).
Female 5×FAD;Lyz2cre;Srrfl/fl mice, in which Srr was conditionally knocked out in microglia, and the control mice, 5×FAD;Srrfl/fl (aged 9–11 months) were subjected to Morris water maze test to assess spatial learning ability. A Paraffin-embedded brain sections from 5×FAD;Lyz2cre;Srrfl/fl mice (male, n = 2) and 5×FAD;Srrfl/fl mice (male, n = 1) were immunostained for SR (green) and Iba1 (red); DAPI (blue) was used to visualize nuclei. Images were captured with a LEICA Stellaris5 microscope (10× objective); scale bar = 1000 μm for column 1, and columns 2–5 are magnified views. B Representative swimming trajectories for the mice are depicted. (C) During the training session, 5×FAD;Lyz2cre;Srrfl/fl mice (n = 9) took less time to reach the submerged platform (escape latency) than female 5×FAD;Srrfl/fl mice (n = 12) on the 5th and 6th days (5th, p = 0.041, 6th, p = 0.012). Mann–Whitney U test was used to compare escape latency at each time point. D In the probe test, 5×FAD;Lyz2cre;Srrfl/fl mice spent more time in the target quadrant than 5×FAD;Srrfl/fl mice on the 7th day (p = 0.028). Mann–Whitney U test was used to compare the percentage of time spending in quadrants. E In the probe test, the number of platform crossings did not differ between the two strains of mice (p = 0.082). Mann–Whitney U test was used to compare the frequencies of platform crossing. F In the probe test, 5×FAD;Lyz2cre;Srrfl/fl mice exhibited a longer swim distance than 5×FAD;Srrfl/fl mice. Student’s t test was used to compare the swim distance, yielding t[19] = 2.992, p = 0.007. G In the probe test, 5×FAD;Lyz2cre;Srrfl/fl mice exhibited significantly faster swimming speed than 5×FAD;Srrfl/fl mice. Student’s t test was used to compare the swim speed, yielding t[19] = 2.992, p = 0.007. Error bars denote the variability of the data.
We further examined amyloid plaques in various brain regions. In the dentate gyrus, the number, total fluorescence intensity, and the area of Aβ plaques were lower in male 5×FAD;Lyz2cre;Srrfl/fl mice compared to those in male 5×FAD;Srrfl/fl mice (Fig. 5A, B). In contrast, no significant difference were observed in the mean fluorescence intensity or mean area of individual Aβ plaques between two groups (Fig. 5B). Similar differences were also observed in the cerebral cortex, thalamus, and hypothalamus (Supplementary Fig. 8). However, the characteristics of Aβ plaques were comparable between female 5×FAD;Lyz2cre;Srrfl/fl mice and female 5×FAD;Srrfl/fl mice (Supplementary Fig. 9).
The mouse brains were processed for paraffin-embedded sections, which were subjected to staining with Aβ antibody 6E10. A Representative immunofluorescence images of the hippocampi are depicted. The 6E10 staining (red) was used to visualize Aβ plaques, Iba1 (green) to label activated microglia, and DAPI (blue) to mark cell nuclei. Images were captured with ZeissLSM880 confocal laser microscope system (Zeiss) under a 40× objective. Bar: 500 μm for the first column. The images were magnified in the second column with a scale bar of 100 μm. B In dentate gyri, 5×FAD;Lyz2cre;Srrfl/fl mice (n = 3) exhibited fewer Aβ plaques compared to 5×FAD;Srrfl/fl mice (n = 3). Student’s t test was used to compare the numbers of Aβ plaques in dentate gyri; t[4] = 9.214, p = 0.001. The 5×FAD;Lyz2cre;Srrfl/fl mice showed a significant reduction in the total fluorescence intensity (int.) of Aβ plaques relative to 5×FAD;Srrfl/fl control. Student’s t test was used to compare the total fluorescence intensity (int.) of Aβ plaques; t[4] = 2.789, p = 0.049. The total area of Aβ plaques was less in 5×FAD;Lyz2cre;Srrfl/fl mice than that in 5×FAD;Srrfl/fl control mice. Student’s t test was used for comparison; t[4] = 5.208, p = 0.006. There were no significant differences in the mean fluorescence intensity (int.) or mean area of Aβ plaques between 5×FAD;Lyz2cre;Srrfl/fl mice and 5×FAD;Srrfl/fl control (n = 3). Student’s t test was used to compare the mean fluorescence intensity (int.) with t[4] = 0.769, p = 0.485, and the mean area of Aβ plaques with t[4] = 0.082, p = 0.939. A significantly higher number of Iba1-positive microglia was present in the dentate gyri of 5×FAD;Lyz2cre;Srrfl/fl mice relative to 5×FAD;Srrfl/fl control mice. Student’s t test was used for comparison; t[4] = 6.379, p = 0.003. Error bars denote the variability of the data.
Conditional knockout of Srr in the microglia of 5×FAD mice elevated lactylation of H3K18, arginase 1, and pyruvate kinase M2
In primary microglia cultures, addition of C646 and oxamate both suppressed the LPS-induced elevation of Arg1 mRNA and the reduction in amplitude was similar. Further, because Arg1 is specifically impacted by H3K18 lactylation, we speculated that Srr deletion increased arg1 through enhancing H3K18 lactylation, contributing to improved AD-like phenotype and brain pathology at some extent. To test this possibility, we examined the levels of histone lactylation in AD mice. Compared with those in 5×FAD;Srrfl/fl mice, the microglia in the hippocampi of 5×FAD;Lyz2cre;Srrfl/fl mice contained significantly higher levels of H3K18 lactylation and arginase 1 surrounding amyloid plaques (Fig. 6A, B). In contrast, the H4K12 lactylation levels surrounding amyloid plaques was at a similar level in both strains of mice (Fig. 6C). We also examined pyruvate kinase M2 (PKM2), an enzyme synthesizing pyruvate from phosphoenolpyruvate. Similar to H3K18 lactylation, PKM2 levels in peri-plaque hippocampal microglia of 5×FAD;Lyz2cre;Srrfl/fl mice were significantly higher than those in 5×FAD;Srrfl/fl mice (Fig. 6D).
The mouse brains were processed for paraffin-embedded sections and subjected to examination of lactylation of H4K12 and H3K18 using immunofluorescence, as well as PKM2. A Representative images are depicted for the microglia expressing lactylated histones in microglia surrounding Aβ plaques. H3K18 lactylation (green) and Iba1 (red) were co-stained to indicate the approximate levels of H3K18 lactylation in activated microglia, while thioflavin S (TS, blue) was used to visualize Aβ plaques. Images were captured with ZeissLSM880 confocal laser microscope system (Zeiss) under 40× objective, with a scale bar of 100 μm for the 1st column. The images were magnified in the 2nd–5th columns with a scale bar of 10 μm. The fluorescence intensity of H3K18 lactylation in activated microglia surrounding Aβ plaques was significantly higher in male 5×FAD;Lyz2cre;Srrfl/fl mice (n = 2, cell = 133) compared to male 5×FAD;Srrfl/fl (n = 2, cell = 124). Mann–Whitney U test was used to compare the fluorescence intensity of H3K18 lactylation; p-value was approximately 0. B Co-immunostaining of Arg1 (green) and Iba1 (red) for evaluating Arg1 levels in activated microglia. Scale bar: 50 μm (column 1), 10 μm (columns 2–5). Arg1 fluorescence intensity in Aβ plaque-associated activated microglia was significantly elevated in male 5×FAD;Lyz2cre;Srrfl/fl mice (n = 1, cell = 86) compared to male 5×FAD;Srrfl/fl (n = 1, cell = 102); Mann–Whitney U test, p-value was approximately 0. C H4K12 lactylation (green) and Iba1 (red) were co-stained to indicate the approximate levels of H4K12 lactylation in activated microglia. Bar: 100 μm for the 1st column, and 10 μm for the 2nd–5th columns. No differences in the fluorescence intensity of H4K12 lactylation were observed between male 5×FAD;Lyz2cre;Srrfl/fl mice (n = 1, cell = 39) and male 5×FAD; Srrfl/fl (n = 2, cell = 102). Mann–Whitney U test was used for comparison; p = 0.197. D PKM2 (green) immunostaining revealed the in situ expression of PKM2 in activated microglia surrounding Aβ plaques. Bar: 100 μm for the 1st column and 10 μm for the 2nd–5th columns. The fluorescence intensity of PKM2 in activated microglia surrounding Aβ plaques was significantly higher in male 5×FAD;Lyz2cre;Srrfl/fl mice (n = 1, cell = 90) compared to male 5×FAD;Srrfl/fl (n = 1, cell = 53). Mann–Whitney U test was used for comparison; p-value was approximately 0. Error bars denote the variability of the data.
Sex-specific expression of SR, DAAO, and H3K18la in 5×FAD;Srr fl/fl mice
To determine the sex-specific expression of SR and DAAO, which influence D-serine availability and consequently affect cognitive function, brain sections were analyzed. SR antibody staining revealed robust labeling in the CA1 region, consistent with SR expression predominantly in post-synaptic neurons in the hippocampus39. Co-localization of SR and Iba1 staining showed significantly stronger co-staining in male 5×FAD;Srrfl/fl mice compared to female 5×FAD;Srrfl/fl mice (Fig. 7A). Staining of DAAO in the cerebral cortex also revealed significantly stronger staining in male 5×FAD;Srrfl/fl mice (Fig. 7B), with a similar effect observed in the hippocampus. H3K18la may indicate an upregulated response by promoting an anti-inflammatory response. Similarly, co-localization of H3K18la and Iba1 staining showed significantly stronger co-staining in male 5×FAD;Srrfl/fl mice compared to female 5×FAD; Srrfl/fl mice (Fig. 7C).
A Representative images of SR (green) and Iba1 (red) immunostaining in the hippocampus of 5×FAD: Srr fl/fl mice; DAPI (blue) was used to visualize nuclei. Images captured with LEICA Stellaris5 microscope (20× objective); scale bar = 100 μm (rows 1–2); row 3 = magnified views. SR fluorescence intensity in activated microglia was significantly higher in males (n = 1, 102 cells) than females (n = 1, 86 cells); Mann–Whitney U test, p-value was approximately 0. B Representative images of DAAO (green) and DAPI (blue) to evaluate DAAO levels in the cerebral cortex of 5×FAD;Srrfl/fl mice. Scale bar = 50 μm. DAAO fluorescence intensity was significantly higher in males (n = 1, 33 cells) than females (n = 1, 48 cells); Mann–Whitney U test, p = 0.002. C Co-immunostaining of H3K18 lactylation (green) and Iba1 (red) in the hippocampus to evaluate H3K18la levels in activated microglia of 5×FAD;Srrfl/fl mice. Scale bar = 50 μm. H3K18la fluorescence intensity in Aβ plaque-associated microglia was significantly higher in males (n = 1, 47 cells) than females (n = 1, 20 cells); Mann–Whitney U test, p-value was approximately 0. Error bars denote the variability of the data.
Discussion
In this manuscript, we demonstrated that deletion of Srr increased phagocytosis of Aβ oligomer and enhanced transcription of anti-inflammatory factors, Arg1 or IL-10, whereas blockade of histone lactylation abrogate upregulation of transcription of Arg1. We further demonstrated that selective deletion of Srr in the microglia of 5×FAD mice significantly mitigated AD-like neuropathology and behavior deficit. Further, we found that SR deficiency in microglia reduced lactate production in response to LPS stimulation (Supplementary Fig. 10). Combined with the in vivo data from Alzheimer’s disease (AD)-like mice—where microglial PKM2 expression was elevated following SR deletion—these findings suggest that a greater proportion of pyruvate is directed toward oxidative phosphorylation rather than lactate production. Our studies have shown that deletion or inhibition of SR in diverse cell types and tissues—including the retinal pigment epithelium40 and the retina41—leads to the accumulation of L-serine, which is an agonist of PKM242. Given that lactate generation is a hallmark of anaerobic glycolysis, we consider that SR deletion shifts microglial metabolic phenotype from anaerobic glycolysis toward oxidative phosphorylation in response to LPS treatment or in AD mice. Taken together with the upregulated expression of anti-inflammatory mediators and enhanced phagocytic capacity observed in LPS-challenged SR-deficient microglia, these findings support our hypothesis that microglial SR deletion mitigates AD-like neuropathology and associated behavioral deficits, which is mediated at least in part by histone lactylation-dependent suppression of neuroinflammation and metabolic reprogramming, although direct experimental evidence for the latter mechanism remains limited owing to technical constraints. A previous study indicates that selective deletion of microglial SR reduces extrasynaptic NMDAR-mediated excitotoxicity and enhances phagocytic capacity in TBI model33. Analogously, this intervention improves behavioral outcomes in our AD-like mouse model, at least partially via the same molecular mechanism.
The precise mechanism by which Srr deletion enhanced H3K18 lactylation is unclear. Nuclear SR immunofluorescence exhibited a strong overlap with p300, which was mitigated by LPS treatment. Consistent with this, absence of SR significantly increased transcription of Arg1 or Il10 in Srr−/− microglia compared to those in WT microglia under LPS treatment. However, we did not find a direct interaction between nuclear SR and p300, nor did we find evidence of interaction between SR and CREB-binding protein (CBP) using protein-protein interaction prediction tools, such as the publicly available bioinformatics resource (https://zhanggroup.org/PEPPI/) and BioGrid. Thus, interference of p300 lactylation by nuclear SR occurs through an indirect mechanism, which presents a significant challenge for investigation. Targeted deletion of SR in microglia elevates H3K18 lactylation, likely through the relief of steric hindrance imposed on p300 by endogenous SR. Notably, this observation stands in contrast to the diminished lactate synthesis associated with microglial SR ablation. We assume that the stimulatory effect of SR deletion on p300-dependent lactylation activity overrides the impact of reduced lactate availability.
Selective deletion of Srr in the microglia of female 5×FAD mice resulted in significant improvements in escape latency, time spent in the targeted quadrant, and frequency of platform crossings. In contrast, deletion of Srr in the microglia of male 5×FAD only increased the frequency of platform crossings but not the other indices used to measure spatial learning. The less obvious effect observed in male mice may be attributed to well-documented effects of testosterone on the activity of D-amino acid oxidase (DAAO); elevated DAAO activity has been consistently found in male mice across six different strains when compared to their female counterparts43,44,45. We also documented elevated DAAO levels in cerebral cortex of male mice compared to female mice. Regarding the sex-specific effects on cognitive function, we found that deletion of microglial SR exerted a more pronounced improvement in female mice than in male mice within an AD-related mouse model. In contrast, in the traumatic brain injury (TBI) model, the beneficial effect of microglial SR deletion was more prominent in male mice33. A plausible explanation for this discrepancy lies in the distinct characteristics of the inflammatory responses in the two models: TBI involves an acute inflammatory reaction, in which estrogen exerts a prominent neuroprotective effect. By comparison, the AD model is associated with a slow-progressing, chronic inflammatory process, whereas the neuroprotective role of estrogen is less pronounced. Thus, the basal neuroprotective capacity of estrogen may underlie the sex-specific differences observed between these two models. Consistent with its more pronounced beneficial effects specifically in female mice within chronic inflammatory disease models, SR deletion in a schizophrenia mouse model—given that chronic inflammation is widely recognized to be a risk factor for schizophrenia—results in attenuated photoreceptor responses in male mice, whereas female mice exhibit no such alterations46. This cross-model consistency further validates the notion that estrogen exerts robust neuroprotective effects in female mice, a mechanism that underpins the sex-specific phenotypic outcomes induced by SR ablation.
It is notable that conditional knockout of Srr reduced Aβ plaque load in male 5×FAD mice in the hippocampus, cerebral cortex, thalamus, and hypothalamus. Moreover, knockdown or deletion of Srr enhanced the phagocytosis of Aβ oligomers in the BV2 cell line and primary microglia. The clearance of Aβ plaques in female 5×FAD mice was less evident. Sex-dependent differences in several aspects of microglial activity have been widely documented, particularly in the context of AD47,48. Specifically, male microglia in vitro exhibit greater migratory capacity than female microglia under both basal and stimulated conditions. Conversely, female microglia demonstrate higher levels of basal and stimulated phagocytic activity compared to male microglia in response to interferon-γ49. Thus, general sex-dependent differences in microglial activity may have contributed to the variations in amyloid load we observed between male and female Srr−/− mice. Notably, amyloid plaque burden increases during the menopausal transition, attributed to the stimulatory effect of follicle-stimulating hormone on the C/EBPβ-δ-secretase pathway50,51. Thus, the true extent of amyloid plaque burden may also be masked by these hormonal regulatory effects, as the mice utilized for spatial memory assessments were in the premenopausal stage.
Notably, the improvement in spatial learning was not correlated with the densities of amyloid plaques in the hippocampi of female mice. The incorrelation is consistent with numerous previous reports indicating that tau pathology is more tightly associated with cognitive decline, while Aβ deposition is also detectable in the brains of healthy individuals52,53,54. Alternatively, the disjunction between Aβ deposition and behavior may simply indicate that SR activation and the consequent D-serine production act downstream of Aβ accumulation to interfere with synaptic activity and behavior. In this sense, a requirement for perturbation of SR and D-serine could potentially explain, at least in part, the relatively poor correlation between plaque load and cognition52. Preservation of normal SR/D-serine levels in some individuals, despite amyloid accumulation, may represent one aspect of resilience against dementia.
Microglia play neuroprotective roles at the early stages of disease; however, they adopt destructive functions at the later stages when aerobic glycolysis reprograms them toward immune tolerance15. This reprogramming can exacerbate neuroinflammation by activating glycolytic enzymes16. In the brain regions associated with neuritic amyloid plaques, microglia that switch to TREM2-APOE signaling exhibit a transcriptional phenotype indicative of neurodegeneration, along with a decrease in Aβ clearance and altered cell-cell communication following aducanumab treatment55,56,57. Notably, we observed elevated levels of H3K18 lactylation and PKM2 in microglia surrounding the plaques, supporting the notion that some of these microglia undergo compensatory responses. Consequently, the selective deletion of Srr in microglia resulted in increased levels of H3K18 lactylation and PKM2 levels in terminally inflammatory microglia surrounding amyloid plaques, suggesting potential therapeutic avenues for AD. TREM2 signaling is elevated in disease-associated microglia55. However, we did not detect alteration of TREM2 and CD36 levels by SR depletion in vitro, indicating that a different, yet uncharacterized, pathway was responsible for any adaptive effects of limiting D-serine production. Overall, we present a therapeutic strategy aiming at reprogramming terminally inflammatory microglia associated with pathological states into anti-inflammatory microglia, thereby highlighting its therapeutic potential in AD.
Methods
Materials
TRIzol reagent (cat #15596-018), donkey antirabbit IgG (H + L) cross-adsorbed secondary antibody, Alexa Fluor® 488 (cat#A21206, RRID# AB_2535792) or anti-mouse IgG (H + L) cross-adsorbed secondary antibody, and Alexa Fluor® 594 (cat#A11012, RRID#AB_14135) were purchased from Invitrogen. B27 (cat# 17504-044) was from Gibco. HiScript II first-strand cDNA synthesis kit (cat# R212-02) was from Vazyme. The Power SYBR Green PCR master mix (cat# 4367659) was from ABI. Penicillin–streptomycin (cat#C0222), amyloid β-peptide (1–42) (cat#P9001), lipopolysaccharide (cat#ST1470), DL-dithiothreitol (cat#ST041), the enhanced BCA protein assay kit (cat#P0010), enhanced BCA protein assay kit (cat #P0010), normal goat serum (cat #C0265), and antifade mounting medium with DAPI (cat #P0131) were purchased from Beyotime Biotechnology, Inc (Shanghai, China). Mouse anti-Aβ antibody (6E10; RRID: AB_2564652) was purchased from BioLegend (San Diego, CA, USA). SR antibody (RRID: AB_399440) was bought from BD Biosciences (San Jose, CA, USA). NaF (cat#100196 08) was from Sinopharm. Na3VO4 (cat#S6508-10G), protease inhibitor cocktails (cat#p8340), Phenylmethanesulfonyl fluoride (PMSF; cat #ST506), DL-dithiothreitol (cat #ST041), protease inhibitor cocktail (cat #P1005), fluorescence-labeled microbeads (1 μm in diameter, cat#L2778) were purchased from Sigma. Neg-50 frozen-section medium was purchased from Thermo Scientific (Rockford, IL, USA). Nitrocellulose (NC) membranes (0.22 µm, cat #66485) was purchased from PALL Corporation (New York, NY, USA). Clarity Western ECL Substrate was bought from Bio-Rad (Hercules, CA, USA). Dulbecco’s modified Eagle’s medium (DMEM) and fetal bovine serum (FBS) were purchased from Thermo Fisher Scientific (Shanghai, China). Mouse IL-1β (cat #50101-MNAE) was purchased from Sino Biological (Beijing, China). Srr−/− mice were created from Biocytogen through the CRISPR/Cas9 technique, and this strain of mouse was used in our research group58.
Data reporting
No statistical methods were used to predetermine sample size. The sample size was determined based on our previous work59,60. There were no sample size differences between the beginning and the end of experiments. The investigators were blinded as to the genotype of the mice. The experiments were simply randomized.
Experimental design
Primary microglial cultures were prepared from postnatal day 1 mice, while behavioral assays were performed on 9–11-month-old adult mice. All the data acquired during experiments were included for analysis. To minimize the number of animals used, the examination of Aβ plaques in the brain was performed after behavior test. To minimize potential confounding factors affecting behavioral assays, mice were habituated to the test environment via repeated exposure to the swimming pool prior to testing. Primary outcome, including phagocytosis, inflammatory response, and behavior tests, were assessed. The numbers of animal used, animal allocation to different experiments were indicated in Table 1.
Microglia culture
Neonatal mice at postnatal day 1 were used. The mice were euthanized on ice, sterilized with 75% ethanol, and then decapitated. The skull was opened using a surgical scissor, and the brain was removed and transferred to dissection media. Under a stereo microscope, the meninges were carefully removed and the cerebral cortex was isolated and digested in 0.05% trypsin at 37 °C for 10 min. The tissues were dissociated using a fire-polished glass pipette, and trypsinization was terminated with the addition of minimum essential medium and fetal bovine serum. The dissociated tissues were subjected to centrifugation at 200 × g for 10 min. After centrifugation, the tissues were resuspended, and the supernatant was filtered through a membrane with a pore size of 40 μm. The filtrate was transferred to a 75-cm2 cell culture flask and cultured in DMEM supplemented with 10% FBS and a penicillin-streptomycin (PS) mixture at 37 °C in a cell incubator with 5% CO2. After three days, the medium was replaced with fresh DMEM supplemented with 10% FBS and the PS mixture, and this process was repeated every seven days until the entire culture period lasted three to four weeks. During the culture period, the cultures were examined for health under a phase-contrast microscope on a weekly basis. Astrocytes, characterized with a polygonal shape, cover the bottom of the culture flask, while microglia, which are spherical and exhibit light refraction, form clusters and rest on the layer of astrocytes. For harvesting microglia, the culture flasks were gently tapped and shaken for 30 min on a rotary shaker until most microglia detached from astrocytes. After collecting microglia, they were plated on 6-well plates and cultured in DMEM plus 10% FBS. The cell purity was confirmed with CD11b staining, reaching ~99% purity.
Construction of a BV2 cell line with stable knockdown of Srr
BV2 cells were purchased from GeFan Biotech Company (Shanghai, cat# GFC063), which was validated with CD11b staining in our research group. Prior to experimentation, the cell line was verified to be free of mycoplasma contamination. To establish an Srr-knockdown BV2 cell line, a culture was transduced with a lentiviral vector carrying sequences for Srr shRNA (GGCGCAATCTCTTCTTCAAAT) under the control of a U6 snRNA promoter in the backbone of pGPU6/mCherry/Puro was used for lentivirus package (OBIO Tech, Inc, Shanghai). In these plasmids, the encoding genes for Srr shRNAs and mCherry fluorescence protein were transcribed as independent transcripts. Before transduction, BV2 cells were plated onto poly-D-lysine-coated 96-well plates at a density of 10,000 cells per well and cultured in DMEM supplemented with 10% FBS and PS mixture. The cells were infected with a specific amount of virus basing on the following formula: virus per well = (cell number × multiplicity of infection (MOI)/virus titer) × 103. Before transduction, the culture media were switched to pre-warmed Opti-MEM and the virus was added to various culture wells according to MOI values of 10, 15, 25, and 40. After 2 h, fresh culture media were added to culture well, and within 24 h, the media were replaced with new culture media. Following an infection of 72 to 96 h, appropriate MOI was selected based on the intensity of mCherry. Puromycin (2 μg/mL) was added to the culture wells until the BV2 cells reached confluence. After detaching BV2 cells, they were subjected to selection using fluorescence-activated cell sorting (FACS), utilizing mCherry fluorescence. The sorted BV2 cells were further selected with puromycin. After five cycles of FACS and puromycin selection, the percentage of mCherry-positive cells exceeded 90%. The knockdown efficacy was validated with western blot. The mCherry fluorescence intensity was comparable between BV2 cells transfected with scrambled siRNA and those transfected with SR-specific siRNA.
Phagocytosis analysis
Phagocytosis analysis of Aβ was conducted in the constructed BV2 cell line or primary microglia. FITC-Aβ ammonium salt was purchased from Bachem, Inc (cat# 4033502). Before use, FITC-Aβ was dissolved in 0.1% ammonia water (Chendu Kelong Chemical Company, China) to 1 mg/mL, diluted to 100 μM, and separately packaged. All the procedures were conducted in a clean bench. Primary microglia isolated from WT or Srr−/− mice were cultured in poly-D-lysine (0.01 mg/mL)-coated culture plates at a density of 100,000 cells per well. After four days, new culture media were replaced and 0.25 μM FITC-Aβ was used to treat the cultures for 6–48 h. At 6, 12, 24, 48 h, the fluorescence images were acquired under immunofluorescence microscope. Following a protocol61, microglial phagocytosis was also examined with a fluorescent latex beads of 1 μm diameter. Briefly, the beads were pre-opsonized in FBS for 1 h at 37 °C. Subsequently, microglial conditioned culture media were replaced with beads-containing DMEM and then incubated at 37 °C for 1 h. The cultures were thoroughly washed five times with ice-cold PBS and then the cells were fixed using 4% paraformaldehyde for 15 min. The images were acquired under fluorescent microscope.
Cloning of SR-expressing plasmid and transfection
The Srr-expressing plasmid was constructed with reference to our previous work20. Briefly, the Srr-encoding cDNA fragment, derived from the RNA template of N2A cells, was amplified using the following primers: forward primer, 5′-ATGTGTGCTCAGTACTGCATCT-3′; reverse primer, 5′-CTAAACAGAAACCGTCTGGTAAGGAG-3′. Using the purified PCR product as the template, a second round of PCR was performed to synthesize the target fragment with the desired restriction sites introduced. Additionally, a Kozak sequence was incorporated to enhance translation efficiency. The primers for this amplification were designed as follows: the forward primer was 5′-AAGATCTCCGAGGCCACCATGTGTGCTCAGTACTGCATCT-3′, with the BglII restriction site and the Kozak consensus sequence underlined, respectively. The reverse primer was 5′-CAGAAACCGTCTGGTAAGGAGAGTACCC AATACGACGTCCCAGACTCGCTTAACTGCAGTCTAGA-3′, carrying the PstI and XbaI restriction sites (both underlined), respectively. After PCR amplification, the resulting product was ligated into the pMD18-T vector. Positive clones were screened by DNA sequencing, followed by bacterial clone propagation for plasmid extraction. The recombinant pMD18-T vector was subjected to sequential restriction digestion with BglII and PstI. The released Srr fragment was then subcloned into the IRES-eGFP vector that had been digested with the same pair of restriction enzymes. The ligation product was transformed into DH5α competent cells. The plasmid extracted from the propagated positive bacterial colonies was designated as the srr-expressing plasmid.
BV2 cells were seeded in 24-well plates at a density of 50,000 cells/well and cultured in DMEM medium supplemented with 10% fetal bovine serum (FBS). After 24 h incubation, the cells were transiently transfected with 500 ng/well of empty vector (pIRES-EGFP, Clontech) or an equal mass of Srr-expressing plasmid (Srr-IRES-eGFP) using lipofectamineTM 3000 reagent. The transfected cells were further cultured for 48 h prior to subsequent experiments.
Establishment of 5×FAD;Lyz2cre;Srr fl/fl mice
The 5×FAD mice expressing human APP and PSEN1 transgenes with a total of five AD-linked mutations were kindly provided by Professor Guiquan Chen in Model Animal Research Center at Nanjing University. Heterozygous Srrfl/+ mice were purchased from Biocytogen (Beijing, China) and crossed with C57BL/6 J mice to generate Srrfl/+ offspring, which were then backcrossed to produce Srrfl/fl mice. Lyz2cre mice were purchased from Cyagen.cn (Suzhou, China). To create 5×FAD;Lyz2cre;Srrfl/fl mice, 5×FAD mice were crossed with Srrfl/fl mice and backcrossed to generate 5×FAD;Srrfl/fl mice, which were crossed with Lyz2cre mice and backcrossed to produce 5×FAD;Lyz2cre;Srrfl/fl mice. 5×FAD; Srrfl/fl mice were used as control mice.
Genotyping
Genotyping was conducted through polymerase chain reaction (PCR) analyses of genomic DNA extracted from tail biopsies. APP Swedish mutations were identified with one primer pair that included a forward primer, 5’-AGGACTGACCACTCGACCAG-3’, and a reverse primer, 5’-CGGGGGTCTAGTTCTGCAT-3′, yielding a product length of 377 bp. The Florida and London mutations were identified with a forward primer, 5’-CTAGGCCACAGAATTGAAAGATCT-3’, and a reverse primer, 5’-GTAGGTGGAAATTCTAGCATCATCC-3’, yielding a product length of 324 bp. PSEN1 mutations were identified with a forward primer, 5’-AATAGAGAACGGCAGGAGCA-3’, and a reverse primer, 5’-GCCATGAGGGCACTAATCAT-3’, producing a product length of 608 bp. The 5’ loxP site were identified with a forward primer, 5’-ATCTGAAGGACGTCTTTAGTCCCTT-3’, and a reverse primer, 5’-GCTGATCAGGTAAGCGCCATGTTAG-3’. This results in a product of 519 bp, indicating the presence of 5’ loxP, or alternatively 365 bp, which signifies the absence of 5’ loxP. The 3’ loxP site were identified with a forward primer, 5’-GTGGCCACTCACCCCTGTAATCTAA-3’, and a reverse primer, 5’-CTGCTGGATTCTCTGGAGTGAGCAT-3’. This reaction produces a product of 474 bp, indicating the presence of 3’ loxP, or alternatively 384 bp, which signifies the absence of 3’ loxP. Lyz2cre were identified with a PCR reaction that included two forward primers: 5’-CCCAGAAATGCCAGATTACG-3’ for detecting cre mutation and 5’-TTACAGTCGGCCAGGCTGAC-3’ for the non-mutated form, along with a common reverse primer, 5’-CTTGGGCTGCCAGAATTTCTC-3’. This reaction produces a product of 700 bp, indicating the presence of cre, or alternatively 350 bp, signifies the absence of cre.
Morris water maze
The spatial learning was assessed using SLY-WMS Morris Water Maze System (Beijing Sunny Instruments Co. Ltd) consisting of a frame grabber, a video camera, a water pool, and an analysis software. The differences between 5×FAD;Srrfl/fl mice and 5×FAD;Lyz2cre;Srrfl/fl were compared in each gender, respectively. Mice of approximately same age and weight were used and the operator was blinded to the group assignment; only the corresponding author, who was not engaged in experimental procedures, was aware of group allocation throughout all experimental stages. The water pool, with a diameter of 1.2 m, was filled with water maintained at 24 °C by a thermostat. A non-toxic white paint was added to render the water opaque. A black curtain placed 80 cm away that held three salient visual cues surrounded the water maze. Mice were randomly trained in four quadrants of the water maze and were allowed a maximum of 60 s to find a hidden platform (10-cm diameter, 1 cm under water). Mice were trained for 6 days (4 trials per day, 10-min interval between trial). On day 7, mice were given a 60-s probe test to test their spatial memory.
Immunofluorescence and thioflavin S staining
The experimental mice were deeply anesthetized via intraperitoneal injection of pentobarbital sodium (50 mg/kg body weight). After deep anaesthetization, the mice were transcardially perfused with ice-cold PBS, followed by perfusion with 4% paraformaldehyde (PFA) dissolved in PBS. The isolated brains were post-fixed in 4% PFA overnight. The brain tissues were dehydrated using an automatic dehydrator (Leica HistoCore PEARL) and then embedded in paraffin. The tissue block was sectioned into coronal slices (5 µm) using a microtome (Leica HistoCore BIOCUT) and stored until use. Antigen retrieval was conducted in 0.01 M sodium citrate solution (pH 6.0) at 100 °C for 40 min. After they had been washed in PBS three times (5 min each) at room temperature (RT), the sections were blocked using 5% normal donkey serum in PBS containing 0.2% Triton X-100 for 1 h at room temperature. The sections were incubated with Aβ antibody (6E10, 1:200), Iba1 (1:300), H3K18la (1:500) overnight at 4 °C. Slides were stained with 2 × 10−11 g/mL thioflavin S in the dark at room temperature for 8 min, followed by three washes with PBS in the dark. After thioflavin S staining, the sections were incubated with secondary antibodies (Alexa Fluor 594 goat anti-rabbit or Alexa Fluor 488 goat anti-mouse IgG, 1:500, Life Technologies Corporation, Carlsbad, CA, USA) for 1 h at RT, then washed three times with PBS. The sections were subsequently mounted on coverslips (Citotest Labware Manufacturing Co., Ltd., Jiangsu, China), and the images were acquired with a laser confocal microscope (Zeiss 880).
Determination of intracellular lactate
Intracellular lactate levels were quantified using a L-lactate assay kit (Abcam, UK, cat#AB65330). Prior to measurement, the culture medium was discarded, and the cultured microglia were gently rinsed twice with PBS. Subsequently, 200 μL of Extraction Solution 1 (provided in the kit) was added. Cells were lysed by ultrasonication on ice (power: 300 W; working time: 3 s; interval: 7 s; total duration: 3 min). The lysates were centrifuged at 12,000 × g for 10 min at 4 °C. Then, 0.8 mL of the resulting supernatant was transferred to a new tube, and 0.15 mL of Extraction Solution 2 was added. The mixture was centrifuged again under the same conditions (12,000 × g, 10 min, 4 °C), and the final supernatant was collected for measurement. A standard curve was prepared by serially diluting a 100 μmol/mL standard solution with distilled water to create a series of working standards with concentrations of 1, 0.625, 0.3125, 0.15625, 0.078, and 0.039 μmol/mL. The reaction system was divided into four groups: Sample Tube, Control Tube, Standard Tube, and Blank Tube. The components were added as specified below: Sample Tube: 50 μL sample + 200 Μl Reagent 1 + 50 μL Reagent 2 + 100 μL Reagent 4; Control Tube: 50 μL sample + 50 μL distilled water + 200 μL Reagent 1 + 100 μL Reagent 4; Standard Tube: 50 μL standard + 200 μL Reagent 1 + 50 μL Reagent 2 + 100 μL Reagent 4; Blank Tube: 50 μL distilled water + 200 μL Reagent 1 + 50 μL Reagent 2 + 100 μL Reagent 4. All mixtures were thoroughly vortexed and incubated in a 37 °C water bath for exactly 20 min. After the initial incubation, 30 μL of Reagent 5 and 300 μL of Reagent 3 were added to each tube. The tubes were then incubated at 37 °C for an additional 20 min in the dark. Following this, the mixtures were centrifuged at 10,000 rpm for 10 min at 25 °C. The supernatant was carefully discarded, and the resulting precipitate was dissolved in 1 mL of absolute ethanol. The absorbance of each solution was measured at 570 nm. The absorbance values were denoted as A Sample, A Control, A Standard, and A Blank, respectively. The net absorbance was calculated using the following formulas: ΔA Sample = A Sample − A Control, ΔA Standard = A Standard − A Blank, A standard curve was plotted with the standard concentrations on the x-axis and the corresponding ΔA Standard values on the y-axis, yielding the regression equation y = kx + b. The lactate concentration (x, in μmol/mL) of the samples was determined by substituting ΔA Sample into this equation. Finally, the intracellular lactate content was normalized to the total protein concentration.
Real-time quantitative PCR
Primary WT or Srr−/− microglia isolated from the aforementioned glial cultures were cultured in DMEM supplemented with 10% FBS and a PS mixture in poly-D-lysine-coated 24-well plates at a density of 100,000 cells per well. After four days, the culture medium was changed to DMEM, followed by sham treatment or lipopolysaccharide (LPS; 25 ng/mL) treatment for 24 h. If the cells remain normal, TRIzol reagent was used to extract total RNA, which was quantified using a NanoDrop 2000 spectrophotometer (ThermoFisher). Reverse transcription of the total RNA was conducted with the HiScript III RT SuperMix for qPCR (+gDNA wiper). The RT-qPCR was conducted using the Taq Pro Universal SYBR qPCR Master Mix in the QuantStudio 5 Real-Time PCR System, with Gapdh serving as an internal control. The following primers were used in the PCR reaction: Arg1, sense, 5′-GTGAAGAACCCACGGTCTGT-3′, anti-sense, 5′-GCCAGAGATGCTTCCAACT G-3′; Il10, sense, 5’-TGCTATGCTGCCTGCTCTTA-3’, anti-sense, 5’-ATGTTGTCC AGCTGGTCCTT-3’; Tnf, sense, 5’-TATGTCTCAGCCTCTTCTCATTCC-3’, anti-sense, 5’-TGAGTGTGAGGGTCTGGGCCC-3’; Nos2, sense, 5’-CAAGCACC TTGGAAGAGGAG-3’, anti-sense, 5’-AACGCCAAACACAGCATACC-3’; Gapdh, sense, 5′-ATGGGGAAGGTGAAGGTCG-3′, and anti-sense, 5′-GGGGTCATTGAT GGCAACAATA-3′. The relative levels of target genes were analyzed using the comparative 2− ΔΔCT method.
Statistics and reproducibility
All the data are all presented as mean ± SEM. Cell culture data were averaged from at least three independent biological replicates, with each independent culture represents designated as a single replicate. For in vivo immunofluorescence analyses, data were averaged from more than nine non-overlapping, randomly selected fields of view per individual specimen. Data were examined with Shapiro–Wilk test for normality distribution. If data display normal distribution, the comparisons between two groups were performed by Student’s t test, and the differences among multiple groups were analyzed with one-way ANOVA followed by Bonferonni when homogeneity of variance was present versus Tamhane’s post hoc test when heterogeneity of variance; otherwise, Mann–Whitney U-test was used to compare the differences between two groups and Kruskal–Wallis for differences among multiple groups (SPSS15.0.1; SPSS, Inc., Chicago, IL, USA). Differences were considered statistically significant if p < 0.05.
Ethics declaration
All experimental procedures following Animal Research Reporting of the In Vivo Experiments (ARRIVE) Guidelines and the protocols used were also approved by the Wenzhou Medical University Committee on the Use and Care of Animals (wydw2020-0056). The mice were provided with water and food ad libitum in a standard pathogen-free facility with automatic illumination on a 12-h on/off cycle at Wenzhou Medical University. C57BL/6 J mice of both sexes were employed in the present study.
Data availability
All the figures in the main manuscript and supplemental information were also deposited in the figshare repository at https://doi.org/10.6084/m9.figshare.30252505. The numerical source data for the graphs were provided in Supplemental Data 1. The uncropped gels were provided as Supplementary Figs. in Supplementary information pdf. The newly generated plasmid was deposited in a community repository, WESTLAKE LABORATORIES, with the link, https://wekwikgene.wllsb.edu.cn and the barcode# 0002358, the name, Srr-IRES-GFP.
References
Mattson, M. P. Pathways towards and away from Alzheimer’s disease. Nature 430, 631–639 (2004).
Cherry, J. D., Olschowka, J. A. & O’Banion, M. K. Neuroinflammation and M2 microglia: the good, the bad, and the inflamed. J. Neuroinflammation 11, 98 (2014).
Heneka, M. T. et al. NLRP3 is activated in Alzheimer’s disease and contributes to pathology in APP/PS1 mice. Nature 493, 674–678 (2013).
Barger, S. W. & Basile, A. S. Activation of microglia by secreted amyloid precursor protein evokes release of glutamate by cystine exchange and attenuates synaptic function. J. Neurochem. 76, 846–854 (2001).
Streit, W. J., Mrak, R. E. & Griffin, W. S. Microglia and neuroinflammation: a pathological perspective. J. Neuroinflammation 1, 14 (2004).
Murenu, E., Gerhardt, M. J., Biel, M. & Michalakis, S. More than meets the eye: the role of microglia in healthy and diseased retina. Front. Immunol. 13, 1006897 (2022).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Prinz, M., Jung, S. & Priller, J. Microglia biology: one century of evolving concepts. Cell 179, 292–311 (2019).
Sosna, J. et al. Early long-term administration of the CSF1R inhibitor PLX3397 ablates microglia and reduces accumulation of intraneuronal amyloid, neuritic plaque deposition and pre-fibrillar oligomers in 5XFAD mouse model of Alzheimer’s disease. Mol. Neurodegener. 13, 11 (2018).
Spangenberg, E. et al. Sustained microglial depletion with CSF1R inhibitor impairs parenchymal plaque development in an Alzheimer’s disease model. Nat. Commun. 10, 3758 (2019).
Sheng, J. G., Griffin, W. S., Royston, M. C. & Mrak, R. E. Distribution of interleukin-1-immunoreactive microglia in cerebral cortical layers: implications for neuritic plaque formation in Alzheimer’s disease. Neuropathol. Appl. Neurobiol. 24, 278–283 (1998).
Mrak, R. E. & Griffin, W. S. Interleukin-1 and the immunogenetics of Alzheimer disease. J. Neuropathol. Exp. Neurol. 59, 471–476 (2000).
Grubman, A. et al. Transcriptional signature in microglia associated with Abeta plaque phagocytosis. Nat. Commun. 12, 3015 (2021).
Michaud, J. P., Bellavance, M. A., Prefontaine, P. & Rivest, S. Real-time in vivo imaging reveals the ability of monocytes to clear vascular amyloid beta. Cell Rep. 5, 646–653 (2013).
Baik, S. H. et al. A breakdown in metabolic reprogramming causes microglia dysfunction in Alzheimer’s disease. Cell Metab. 30, 493–507.e496 (2019).
Pan, R. Y. et al. Positive feedback regulation of microglial glucose metabolism by histone H4 lysine 12 lactylation in Alzheimer’s disease. Cell Metab. 34, 634–648.e636 (2022).
Parkhurst, C. N. et al. Microglia promote learning-dependent synapse formation through brain-derived neurotrophic factor. Cell 155, 1596–1609 (2013).
Barger, S. W., Goodwin, M. E., Porter, M. M. & Beggs, M. L. Glutamate release from activated microglia requires the oxidative burst and lipid peroxidation. J. Neurochem. 101, 1205–1213 (2007).
Wu, S. Z. et al. Induction of serine racemase expression and D-serine release from microglia by amyloid beta-peptide. J. Neuroinflammation 1, 2 (2004).
Zhang, H. et al. Alterations of serine racemase expression determine proliferation and differentiation of neuroblastoma cells. FASEB J. 36, e22473 (2022).
Zhang, H. et al. Reduced serine racemase expression in aging rat cerebellum is associated with oxidative DNA stress and hypermethylation in the promoter. Brain Res. 1629, 221–230 (2015).
Zhang, H., Lu, J. & Wu, S. Sp4 controls constitutive expression of neuronal serine racemase and NF-E2-related factor-2 mediates its induction by valproic acid. Biochim. Biophys. Acta Gene Regul. Mech. 1863, 194597 (2020).
Madeira, C. et al. d-serine levels in Alzheimer’s disease: implications for novel biomarker development. Transl. Psychiatry 5, e561 (2015).
Wu, S. & Barger, S. W. Induction of serine racemase by inflammatory stimuli is dependent on AP-1. Ann. N. Y Acad. Sci. 1035, 133–146 (2004).
Balu, D. T. et al. Neurotoxic astrocytes express the D-serine synthesizing enzyme, serine racemase, in Alzheimer’s disease. Neurobiol. Dis. 130, 104511 (2019).
Turpin, F. R. et al. Reduced serine racemase expression contributes to age-related deficits in hippocampal cognitive function. Neurobiol. Aging 32, 1495–1504 (2011).
Billard, J. M. et al. Early involvement of D-serine in β-amyloid-dependent pathophysiology. Cell. Mol. Life Sci. 82, 179 (2025).
Wu, S. Z., Jiang, S., Sims, T. J. & Barger, S. W. Schwann cells exhibit excitotoxicity consistent with release of NMDA receptor agonists. J. Neurosci. Res. 79, 638–643 (2005).
Mustafa, A. K. et al. Serine racemase deletion protects against cerebral ischemia and excitotoxicity. J. Neurosci. 30, 1413–1416 (2010).
Inoue, R., Hashimoto, K., Harai, T. & Mori, H. NMDA- and beta-amyloid1-42-induced neurotoxicity is attenuated in serine racemase knock-out mice. J. Neurosci. 28, 14486–14491 (2008).
Jiang, H. et al. Loss-of-function mutation of serine racemase attenuates retinal ganglion cell loss in diabetic mice. Exp. Eye Res. 175, 90–97 (2018).
Mothet, J. P. et al. D-serine is an endogenous ligand for the glycine site of the N-methyl-D-aspartate receptor. Proc. Natl. Acad. Sci. USA 97, 4926–4931 (2000).
Arizanovska, D. et al. Cognitive loss after brain trauma results from sex-specific activation of synaptic pruning processes. Brain https://doi.org/10.1093/brain/awaf293 (2025).
Tapanes, S. A. et al. Inhibition of glial D-serine release rescues synaptic damage after brain injury. Glia 70, 1133–1152 (2022).
Ni, X. et al. Regional contributions of D-serine to Alzheimer’s disease pathology in male App(NL-G-F/NL-G-F) mice. Front Aging Neurosci. 15, 1211067 (2023).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575 (2019).
Wang, N. et al. Histone lactylation boosts reparative gene activation post-myocardial infarction. Circ. Res. 131, 893–908 (2022).
Wang, S. et al. TREM2 drives microglia response to amyloid-beta via SYK-dependent and -independent pathways. Cell 185, 4153–4169.e4119 (2022).
Wong, J. M. et al. Postsynaptic serine racemase regulates NMDA receptor function. J. Neurosci. 40, 9564–9575 (2020).
Wu, M. et al. Intravenous injection of l-aspartic acid beta-hydroxamate attenuates choroidal neovascularization via anti-VEGF and anti-inflammation. Exp. Eye Res. 182, 93–100 (2019).
Wang, S. et al. Effects of topic delivery of an inhibitor of serine racemase on laser-induced choroidal vasculopathy. Transl. Vis. Sci. Technol. 13, 24 (2024).
Chaneton, B. et al. Serine is a natural ligand and allosteric activator of pyruvate kinase M2. Nature 491, 458–462 (2012).
Clark, L. C. Jr., Kochakian, C. D. & Fox, R. P. The effect of castration and testosterone propionate on D-amino acid oxidase activity in the mouse. Science 98, 89 (1943).
Konno, R. & Yasumura, Y. Mouse mutant deficient in D-amino acid oxidase activity. Genetics 103, 277–285 (1983).
Endahl, B. R. & Kochakian, C. D. Role of castration and androgens in D-amino acid oxidase activity of tissues. Am. J. Physiol. 185, 250–256 (1956).
Torres Jimenez, N. et al. Electroretinographic abnormalities and sex differences detected with mesopic adaptation in a mouse model of schizophrenia: A and B wave analysis. Invest Ophthalmol. Vis. Sci. 61, 16 (2020).
Villa, A. et al. Sex-specific features of microglia from adult mice. Cell Rep. 23, 3501–3511 (2018).
Ferretti, M. T. et al. Sex differences in Alzheimer disease—the gateway to precision medicine. Nat. Rev. Neurol. 14, 457–469 (2018).
Yanguas-Casas, N. et al. Sex differences in the phagocytic and migratory activity of microglia and their impairment by palmitic acid. Glia 66, 522–537 (2018).
Xiong, J. et al. FSH blockade improves cognition in mice with Alzheimer’s disease. Nature 603, 470–476 (2022).
Xiong, J. et al. FSH and ApoE4 contribute to Alzheimer’s disease-like pathogenesis via C/EBPbeta/delta-secretase in female mice. Nat. Commun. 14, 6577 (2023).
Nelson, P. T. et al. Correlation of Alzheimer disease neuropathologic changes with cognitive status: a review of the literature. J. Neuropathol. Exp. Neurol. 71, 362–381 (2012).
Moscoso, A. et al. Frequency and clinical outcomes associated with tau positron emission tomography positivity. JAMA. 334, 229–242 (2025).
Malpas, C. B. et al. Tau and amyloid-beta cerebrospinal fluid biomarkers have differential relationships with cognition in mild cognitive impairment. J. Alzheimers Dis. 47, 965–975 (2015).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e1217 (2017).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581.e569 (2017).
Millet, A., Ledo, J. H. & Tavazoie, S. F. An exhausted-like microglial population accumulates in aged and APOE4 genotype Alzheimer’s brains. Immunity 57, 153–170.e156 (2024).
Zhou, J. et al. Deletion of serine racemase reverses neuronal insulin signaling inhibition by amyloid-beta oligomers. J. Neurochem 163, 8–25 (2022).
Wang, Y., Lian, M., Zhou, J. & Wu, S. Brain Dicer1 is down-regulated in a mouse model of Alzheimer’s disease via Abeta42-induced repression of nuclear factor erythroid 2-related factor 2. Mol. Neurobiol. 57, 4417–4437 (2020).
Wang, Y. et al. Dicer1 promotes Abeta clearance via blocking B2 RNA-mediated repression of apolipoprotein E. Biochim. Biophys. Acta Mol. Basis Dis. 1867, 166038 (2021).
Lian, H., Roy, E. & Zheng, H. Microglial phagocytosis assay. Bio Protoc. 6, e1988 (2016).
Acknowledgements
This study was supported by National Natural Science Foundation of China for Young Scholars (82301614), and an intramural grant, Integrated Project of State Key Laboratory of School of Optometry and Ophthalmology, Wenzhou Medical University (J02-20190204).
Author information
Authors and Affiliations
Contributions
J. Zhou: Investigation, visualization, and funding acquisition; Y.H. Yang: Investigation and visualization; S.Y. Liu: Investigation and visualization; J. Chen: Investigation; H.J. Liao: Investigation; W.J. Liang: Investigation; Z.W. Zhang: Investigation; Y. Wang: Investigation; Y.M. Liu: Investigation; H. Zhang: Investigation; H.Y. Jiang: Investigation; W.C. Lin: Methodology; J. Qu: Supervision and methodology; S.W. Barger: Revision and discussion; S.Z. Wu: Conceptualization, supervision, and writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Ibrahim Javed and Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, J., Yang, Y., Liu, S. et al. Microglial serine racemase knockout alleviates Alzheimer-like neuropathology and behavioral deficit via lactylation-mediated anti-inflammation. Commun Biol 9, 493 (2026). https://doi.org/10.1038/s42003-026-09772-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09772-y