Abstract
Selective fetal growth restriction (sFGR) in monochorionic diamniotic twins (MCDA) reflects placental dysfunction, but trophoblast adaptation mechanisms remain unclear. Using single-cell RNA sequencing on placental tissues from three paired sFGR, we demonstrate that villous cytotrophoblasts (VCT) in growth-restricted placentas undergo a transition from VCT_TP63, which expresses barrier-associated TP63/SOX6 and maintains cytoskeletal integrity, to VCT_LDHA, a metabolically reprogrammed phenotype marked by LDHA/YY1/RELA activation. Trajectory analysis shows diminished syncytial precursors, suggesting impaired fusion capacity. Immune profiling identifies depleted TREM2+ Hofbauer macrophages and expanded interferon-responsive natural killer (NK) cells. Cell-cell interaction mapping demonstrates enhanced Interferon Gamma (IFNG)-Interferon Gamma Receptor 1 (IFNGR1)-Signal Transducer and Activator of Transcription 1 (STAT1) signaling between VCT_LDHA and immune cells, alongside weakened VCT_TP63-stromal crosstalk. This study defines a maladaptive triad of metabolic stress, inflammation, and structural disintegration in sFGR, contributing to sFGR pathogenesis.
Similar content being viewed by others
Introduction
The placenta is a highly specialized organ essential for sustaining pregnancy and coordinating fetal growth. Its dysfunction is a central driver of pregnancy-related complications such as preeclampsia, fetal growth restriction (FGR), and preterm birth1,2. Among these, selective fetal growth restriction (sFGR) affects ~10–15% of monochorionic diamniotic (MCDA) twin pregnancies, primarily due to unequal placental territory allocation and intertwin vascular anastomoses, which predispose to discordant nutrient and oxygen supply between cotwins3,4. sFGR significantly elevates the risks of perinatal morbidity and long-term neurodevelopmental impairment, and its clinical management remains challenging, particularly in cases lacking effective in utero interventions5.
FGR is broadly attributed to three major etiological domains: maternal factors (e.g., hypertensive disorders), fetal factors (e.g., aneuploidy), and placental factors6. However, in singleton pregnancies, it is inherently difficult to disentangle the relative contributions of placental dysfunction from maternal and fetal confounders. Importantly, MCDA twin pregnancies complicated by sFGR provide a unique intra-placental comparison model, where cotwins share an identical maternal environment and nearly identical fetal genetic background. This design enables the isolation of localized placental pathology as the principal determinant of growth discordance, offering a powerful system to specifically interrogate trophoblast maladaptation and microenvironmental remodeling.
Previous investigations into sFGR have largely focused on gross placental morphology, Doppler hemodynamics, and bulk transcriptomic or proteomic profiling, which have identified dysregulation of oxidative stress pathways, angiogenic signaling, and nutrient transport7,8. Nonetheless, the intrinsic cellular heterogeneity of the placenta substantially limits the interpretive resolution of bulk omics approaches, impeding precise identification of pathological driver cell populations and their crosstalk with the microenvironment. Single-cell sequencing technologies have revolutionized our ability to resolve complex cellular ecosystems and have led to important advances in understanding preeclampsia, early-onset FGR, and other placental disorders9,10,11,12. Despite these advances, no studies to date have systematically applied single-cell transcriptomic profiling to MCDA twin pregnancies complicated by sFGR using an intra-placental control design, leaving the cellular and molecular basis of trophoblast maladaptation incompletely defined.
In this context, we conducted a comprehensive single-cell transcriptomic investigation of paired placental samples from MCDA twin pregnancies discordant for fetal growth. Leveraging a paired analysis strategy under a shared maternal and fetal background, we deeply characterized the heterogeneity and remodeling of villous cytotrophoblast (VCT) subpopulations. By integrating transcriptional regulatory network reconstruction, sublineage fate trajectory analysis, and ligand–receptor interaction profiling, this study delineates multilayered regulatory circuits driving VCT fate reprogramming, immune–stromal crosstalk, and maladaptive trophoblast plasticity in sFGR placentas. Our findings may provide molecular insights into trophoblast-centric pathogenesis and identify potential entry points for future therapeutic targeting.
Materials and methods
Study design and sample collection
This study was conducted at the International Peace Maternity and Child Health Hospital, Shanghai Jiao Tong University, with approval from the institutional ethics committee (GKLW 2019-72). Informed consent was obtained from all participants. All ethical regulations relevant to human research participants were followed. A total of three pairs of MCDA twin pregnancies complicated by sFGR were enrolled. The diagnosis of sFGR was based on the Delphi consensus criteria, defined as either an estimated fetal weight (EFW) below the 3rd percentile in one twin or meeting at least two of four criteria: EFW below the 10th percentile, abdominal circumference below the 10th percentile, umbilical artery pulsatility index above the 95th percentile in the smaller twin, or intertwin EFW discordance ≥25%. Immediately after delivery, we collected paired placental samples corresponding to the twin placental territories: sFGR_Smaller (placental territory of the smaller birth-weight twin meeting sFGR criteria) and sFGR_Larger (placental territory of the larger birth-weight twin). All samples were histologically confirmed as monochorionic placentas, and those with major structural abnormalities were excluded. Detailed clinical characteristics of the three twin pairs are provided in Supplementary Table 1.
Placental sampling: to minimize placental tissue heterogeneity, a five-point sampling method was employed. The sampling points included: the umbilical cord root, the central fetal side of the placenta, the marginal fetal side, the central maternal side, and the marginal maternal side. Each sampling point was ~1 cm² in size to ensure uniformity across all locations. All samples were collected by the same researcher at the same time point to ensure consistency and standardization, thereby reducing potential spatial variability. For single-cell RNA sequencing, the five samples were pooled, homogenized, and then subjected to sequencing, allowing for a comprehensive representation of placental tissue at the cellular level. All tissues were processed within 30 min of collection for single-cell dissociation.
Tissue dissociation and single-cell suspension preparation
Placental tissue samples were transported on ice in sterile culture dishes containing 10 mL 1× Dulbecco’s Phosphate-Buffered Saline (DPBS, Thermo Fisher Scientific, Cat. no. 14190144) to remove residual storage solution. Samples were minced on ice and enzymatically dissociated at 37 °C in DPBS containing 0.1% Type I Collagenase (Sigma, Cat. no. C0130), 10 μg/mL DNase I (Sigma, Cat. no. 11284932001), and 5% fetal bovine serum (FBS, Thermo Fisher Scientific, Cat. no. SV30087.02), under gentle agitation (50 rpm) for approximately 40 min. Dissociated cells were collected at 20 min intervals to maximize yield and viability. Single-cell suspensions were sequentially filtered through a 40 μm nylon strainer, treated with 1× Red Blood Cell Lysis Solution (Thermo Fisher Scientific, Cat. no. 00-4333-57), and washed with DPBS containing 2% FBS.
Single-cell RNA sequencing library preparation and sequencing
Single-cell suspensions were processed using the Chromium Single Cell 3′ v3 platform (10× Genomics) following the manufacturer’s protocols (CG000204 Rev B). Briefly, individual cells were partitioned into Gel Bead-In-Emulsions (GEMs) containing barcoded oligonucleotides with unique molecular identifiers (UMIs). After cell lysis and reverse transcription, cDNA molecules were tagged with cell-specific barcodes and UMIs. Libraries were generated through second-strand synthesis, adaptor ligation, and amplification, enriching for 3′ transcript ends. Quality and concentration of libraries were assessed using the High Sensitivity DNA Kit (Agilent Bioanalyzer 2100) and Qubit High Sensitivity DNA Assay Kit (Thermo Fisher Scientific). Libraries were sequenced on an Illumina NovaSeq 6000 platform using paired-end 150 bp reads.
Data processing
scRNA-seq data analysis was performed by NovelBio Co., Ltd. with NovelBrain Cloud Analysis Platform. We applied fastp (v0.21.0) with default parameter filtering the adaptor sequence and removed the low-quality reads to achieve the clean data. To quantify the gene expression of the single-cell data, we used CellRanger v7.1.0 along with human genome GRCh38 (Ensembl annotation version 104). Cells contained over 500 expressed genes and mitochondria UMI rate below 30% passed the cell quality filtering and mitochondria genes were removed in the expression table.
Seurat package (version:4.1.0, https://satijalab.org/seurat/) was used for cell normalization and regression based on the expression table according to the UMI counts of each sample and percent of mitochondria rate to obtain the scaled data. For batch correction using Seurat’s canonical correlation analysis (CCA) integration, the following parameters were strictly implemented: Prior to integration, the SelectIntegrationFeatures function was used to retain HVFs conserved across ≥50% of batches, resulting in a shared set of 3000 features for cross-batch alignment. These features were input to FindIntegrationAnchors, which identifies “anchor” cells between batches based on CCA similarity. Critical parameters for anchor detection included k.anchor = 30, dim = 1:30, and k.weight = 100. Following integration, data were scaled with scale data, and dimensionality reduction was performed using PCA (npcs = 50) and UMAP (dims = 1:10, n.neighbors = 30) to validate batch mixing. PCA was constructed based on the scaled data with top 2000 high variable genes and top 10 principals were used for tSNE construction and UMAP construction. Utilizing graph-based cluster method, we acquired the unsupervised cell cluster result based the PCA top 10 principal and we calculated the marker genes by FindAllMarkers function with wilcox rank sum test algorithm under following criteria:1. Log2FC > 0.25; 2. P < 0.05; 3. min.pct > 0.1. In order to identify the cell type detailed, the clusters of same cell type were selected for re-tSNE analysis, graph-based clustering and marker analysis.
Data analysis
Gene Ontology (GO) analysis
To elucidate the biological implications of the DEGs and marker genes, GO analysis was performed. GO annotations were downloaded from NCBI, the Gene Ontology database (http://www.geneontology.org/) and UniProt (http://www.UniProt.org/). Fisher’s exact test was applied to identify the significant GO categories, and an FDR was used to correct the p values.
Pathway analysis
Pathway analysis was used to explore the significant pathways of the DEGs and marker genes based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) database. Fisher’s exact test was used to identify significant pathways, and the threshold of significance was defined by the p value and FDR.
Gene enrichment analysis
For gene enrichment analysis, Fisher’s exact test was applied to calculate the P value for each gene set. Raw P values were adjusted for multiple hypothesis tests using the Benjamini and Hochberg method. Such enrichment analysis was applied to annotations including GO (v2.5.13), the Kyoto Encyclopedia of Genes and Genomes (KEGG, 201900613), the Hallmark (h.all.7.0., https://www.gsea-msigdb.org/gsea/msigdb/index.jsp), Chromosome location gene set (https://ftp.ncbi.nlm.nih.gov/gene/DATA/GENE_INFO/Mammalia/) and customized gene sets. A customized 41-signature gene set including immune-, cytokine- and neurobiology-related terms were collected from the CellphoneDB database. To characterize the relative activation of a given gene set, we performed QuSAGE (2.16.1) analysis (Yaari et al.).
SCENIC analysis
To assess the regulatory strength of TFs, we applied the SCENIC (v0.11.2) (Aibar et al.) workflow, which is a computational method for the construction of regulatory networks and identification of different cell states from scRNA-seq data, using the 20-thousand motif database for RcisTarget and GRNboost. The network of top 20 importance target genes were presented with Cytoscape (v3.8.2).
Pseudotime analysis
Single-cell trajectory analysis was performed with Monocle2 (http://cole-trapnell-lab.github.io/monocle-release) utilizing DDR-Tree and default parameters to determine the dramatic translational relationships among cell types and clusters. Before Monocle analysis, marker genes of the Seurat clustering result and raw expression counts of the cell that passed filtering were selected. Branch expression analysis modeling (BEAM analysis) was utilized for branch fate-determined gene analysis based on pseudotime analysis.
CytoTrace
Cellular Trajectory Reconstruction Analysis using gene Counts and Expression (CytoTRACE, v0.3.3), a computational method, was applied to predict the differentiation state of cells from single-cell RNA-sequencing data through counting the number of expressed genes per cell.
Cell–cell communication analysis
To enable a systematic analysis of cell–cell communication molecules, we applied cell communication analysis based on the CellPhoneDB(v1.1.0), a public repository of ligands, receptors and their interactions. Membrane secreted and peripheral proteins of the cluster was annotated. Significant mean and Cell Communication significance (P < 0.05) was calculated based on the interaction and the normalized cell matrix achieved by Seurat Normalization.
CellChat
Cell communication analysis was performed using the R package CellChat (v1.1.3) with default parameters. The CellChatDB human was used for analysis. All two groups of samples were normalized together, and then each group was extracted and analyzed and compared in parallel. This is with the assumption that they were sharing cell types.
Differential gene expression analysis
To identify differentially expressed genes among samples, the function FindMarkers with wilcox rank sum test algorithm was used under following criteria:1. Log2FC > 0.25; 2. P < 0.05; 3. min.pct > 0.1 which was more conducive to obtain different functional genes among samples.
Statistical analysis
The proportion of cell types among groups were performed using paired two-tailed Student’s t test. All statistical analyses and presentations were performed with R versions 3.6.2. Statistical significance was set at P < 0.05.
Quantitative PCR validation
Key transcription factors were validated by quantitative PCR (qPCR). Total RNA was extracted from the same placental samples (TRIzol, Invitrogen), reverse transcribed into cDNA (PrimeScript RT Kit, Takara), and quantified using SYBR Green-based qPCR (Applied Biosystems). GAPDH/ACTB served as the internal control, and relative expression levels were calculated using the 2−ΔΔCT method. The sequences of all qPCR primers used are provided in Supplementary Table 2.
Multiplex immunofluorescence staining
Formalin-fixed paraffin-embedded placental sections (4 μm) were deparaffinized, rehydrated, and subjected to antigen retrieval in citrate buffer (pH 6.0). After blocking with 5% bovine serum albumin, slides were incubated overnight at 4 °C with primary antibodies against TP63 (CST, #39692, 1:500), SEMA6D (abcam, ab198745, 1:100), LDHA (CST, #3582S, 1:200), S100A9 (CST, #73425, 1:100), and CDH1 (CST, #14472S, 1:200). Sequential detection was performed using HRP-conjugated secondary antibodies and tyramide signal amplification (TSA) fluorophores (Shanghai Huilan Biotechnology Co., Ltd., catalog no. RC0086plus-45RM); image analysis software: CaseViewer (Servicebio).), with microwave stripping between cycles. Nuclei were counterstained with DAPI, and slides were mounted with antifade medium. Multispectral images were acquired using CaseViewer (Servicebio).
Statistics and reproducibility
Statistical analyses were performed using R (version 3.6.2), with statistical significance defined as P < 0.05. For single-cell RNA sequencing, placental tissues from three MCDA twin pregnancies complicated by sFGR were analyzed, with paired sFGR_Smaller and sFGR_Larger placental territories sampled from each pregnancy (n = 3 pregnancies, 6 placental territories). The analysis of differential cell-type proportions between the paired territories was performed using paired two-tailed Student’s t tests, appropriate for the paired design of the study. Normality of the differences in cell proportions was assessed using the Shapiro–Wilk test, which was performed on each cell type across the three pairs of samples. Results indicated that the majority of the P-values were >0.05, suggesting that the differences in cell proportions followed a normal distribution.
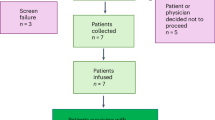
For quantitative PCR (qPCR) validation, RNA was extracted from paired placental samples (n = 7 MCDA twin pregnancies), with each sample representing both the sFGR_Smaller and sFGR_Larger territories from the same pregnancy. Data were analyzed by the 2−ΔΔCt method, using ACTIN as the internal control, and differences between paired samples were assessed using paired two-tailed Student’s t tests. All experiments were conducted under standardized conditions to ensure reproducibility and reduce variability.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Cellular composition and proportional changes in placentae from MCDA twin pregnancies complicated by sFGR
To characterize the cellular composition and alterations in placentae from MCDA twin pregnancies complicated by sFGR, we performed scRNA-seq on placental tissues from three twin pairs (Table 1 & Fig. 1A). We recovered a total of 175,160 cells from the six placental samples, with an average of ~29,193 cells per sample. The overall mean sequencing depth was 14,935 reads per cell. The dataset exhibited a broad spectrum of transcriptional abundance and complexity, with the median UMI counts per cell averaging 3920 and the median genes per cell averaging 1538 across all samples.
A Schematic overview of the experimental workflow. Placental tissues were collected from monochorionic diamniotic (MCDA) twin pregnancies complicated by selective fetal growth restriction (sFGR). Single-cell RNA sequencing was performed using 10× Genomics platform, followed by paired analysis to compare growth-restricted (sFGR_Smaller) and appropriately grown (sFGR_Larger) cotwin placental samples under a shared maternal and fetal background. Downstream analyses included unsupervised clustering, transcriptional regulatory network inference, pseudotime trajectory reconstruction, and ligand–receptor interaction profiling. B Uniform manifold approximation and projection (UMAP) visualization of all cells (n = 175,160) from the integrated dataset. Cells are colored by annotated clusters, including villous cytotrophoblast (VCT), syncytiotrophoblast (SCT), extravillous trophoblast (EVT), cycling VCT (VCT_Cycling), fusing VCT (VCT_Fusing), Hofbauer cells, monocytes, neutrophils, stromal cells, B cells, T cells, and NK cells. C UMAP embeddings for each individual sample (3 pairs of sFGR_Larger and sFGR_Smaller placentas), demonstrating robust reproducibility of the clustering across biological replicates. D Dot plot of canonical marker gene expression used for cluster annotation. The size of each dot represents the proportion of cells expressing each marker gene in a given cluster, and the color intensity indicates average scaled expression. E Bar plot showing the composition of major cell clusters in sFGR_Larger and sFGR_Smaller samples. F Heatmap summarizing relative abundance of each cell cluster in sFGR_Larger versus sFGR_Smaller placentas. Color scale represents the fold change in proportion (log2 scale) between the two groups. Paired comparative analysis of cell proportions for G VCT, H VCT_Cycling, and I VCT_Fusing subpopulations between sFGR_Larger and sFGR_Smaller placentas in each twin pair (n = 3 biologically independent twin pairs). Lines connect matched samples; statistical significance was assessed using paired t-tests. *Error bars indicate mean ± s.e.m. *P < 0.05, *P < 0.01, ns not significant.
A total of 12 distinct cell clusters were identified, encompassing trophoblast cells, immune cells, and stromal cells. The trophoblast clusters included villous cytotrophoblasts (VCT), cycling VCT (VCT_Cycling), fusing VCT (VCT_Fusing), extravillous trophoblasts (EVT), and syncytiotrophoblasts (SCT). The immune clusters comprised T cells, B cells, monocytes, neutrophils, natural killer (NK) cells, and Hofbauer macrophages (Fig. 1B–E).
General intergroup comparisons revealed significant changes across all major cell populations (Fig. 1F). Notably, we focused on VCT subpopulations but not SCT or EVT for two primary reasons: (1) SCT, being multinucleated giant cells, are largely underrepresented in 10x single-cell sequencing platforms due to technical limitations; and (2) the collected specimens were restricted to villous tissue, excluding decidua, thereby accounting for the limited proportion of EVT. The VCT_Fusing cluster can to some extent reflect SCT precursor activity.
Analysis of immune cell composition revealed elevated proportions of pro-inflammatory immune subtypes (including B cells, T cells, monocytes, NK cells, and neutrophils) in sFGR_Smaller placentae, indicating an activated inflammatory microenvironment. In contrast, stromal cells decreased in abundance, suggesting compromised structural support within the placenta.
Leveraging the intrinsic paired nature of MCDA twin samples, we further conducted pairwise analyses of cell-type proportions. Only VCT, VCT_Cycling, and VCT_Fusing displayed consistent directional changes across the paired samples, with VCT and VCT_Fusing consistently reduced (Fig. 1G, I), and VCT_Cycling consistently increased in the sFGR_Smaller placentas (Fig. 1H), reaching statistical significance for VCT (P = 0.025, log2FC = −0.42) and VCT_Cycling (P = 0.006, log2FC = 1.36). These findings highlight these VCT subpopulations as potential common cell groups central to sFGR pathophysiology.
Global shifts in VCT subpopulation proportions and functional remodeling
Comprehensive subclustering of trophoblast lineage cells identified five major VCT subpopulations: VCT_TP63, VCT_LDHA, VCT_TP63_Cycling, VCT_LDHA_Cycling, and VCT_Fusing, in addition to EVT and SCT clusters (Fig. 2A). Among these, VCT_TP63 and VCT_LDHA represented distinct progenitor subtypes characterized by high expression of TP63 and LDHA, respectively (Fig. 2C). Distribution analysis revealed that sFGR_Smaller placentas exhibited a significant reduction in the proportion of VCT_TP63 cells (P = 0.006, log2FC = −2.5) and a corresponding expansion of VCT_LDHA (P = 0.0005, log2FC = 2.5) and VCT_LDHA_Cycling (P = 0.03, log2FC = 0.93) subpopulations, indicating a marked shift toward metabolically reprogrammed trophoblast compartments under growth-restricted conditions (Fig. 2B, C).
A UMAP visualization of all trophoblast lineage cells, showing further subclustering into VCT subsets (VCT_TP63, VCT_LDHA), VCT_Cycling, VCT_Fusing, and EVT and syncytiotrophoblast (SCT) clusters. B UMAP colored by sFGR_Larger and sFGR_Smaller samples, illustrating the distribution of trophoblast subsets across the two groups. C Feature plots depicting the expression of TP63 and LDHA, demonstrating distinct spatial segregation of structural (TP63-high) and metabolic (LDHA-high) VCT subpopulations. D Gene Ontology (GO) enrichment analysis of marker genes defining VCT_TP63 (blue) and VCT_LDHA (orange) subclusters, highlighting differences in cytoskeletal organization, signal transduction, and mitochondrial metabolic processes. E GO enrichment analysis of marker genes in the VCT_Cycling subpopulation. F GO enrichment analysis of marker genes in the VCT_Fusing subpopulation. G Venn diagrams showing the overlap of differentially expressed genes (DEGs) identified in the three paired samples for VCT_TP63 upregulated DEGs (left), VCT_TP63 downregulated DEGs (second), VCT_LDHA upregulated DEGs (third), and VCT_LDHA downregulated DEGs (right). H KEGG pathway enrichment analysis of DEGs in VCT_TP63 between sFGR_Smaller and sFGR_Larger samples, with top panel representing upregulated DEGs and bottom panel representing downregulated DEGs. I KEGG pathway enrichment analysis of DEGs in VCT_LDHA between sFGR_Smaller and sFGR_Larger samples, with top panel representing upregulated DEGs and bottom panel representing downregulated DEGs.
Functional annotation of marker genes highlighted clear biological specialization within each subpopulation. Specifically, VCT_TP63 marker genes were enriched in pathways related to cytoskeletal organization, GTPase activity, chromatin remodeling, and transcriptional activation, underscoring its structural and progenitor maintenance functions (Fig. 2D). By contrast, VCT_LDHA marker genes showed significant enrichment in mitochondrial respiratory chain assembly, electron transport, oxidative phosphorylation, and metabolic stress adaptation processes, reflecting a metabolically active phenotype (Fig. 2D).
The VCT_Cycling subpopulation exhibited marker gene enrichment in pathways including mitotic cell cycle regulation, RNA splicing, mitochondrial translation, and chromosome segregation (Fig. 2E), consistent with active proliferation and biosynthetic capacity. In parallel, VCT_Fusing marker genes were enriched in autophagy, apoptotic regulation, endoplasmic reticulum stress response, and protein transport processes (Fig. 2F), suggesting readiness for differentiation and syncytial fusion under stress conditions.
To define robust differential expression signatures, we applied a stringent paired-sample strategy, wherein differentially expressed genes (DEGs) were required to be consistently identified in all three matched sFGR_Larger/sFGR_Smaller pairs to be retained for downstream analysis (Fig. 2G). This approach aimed to minimize inter-sample variability and isolate reproducible transcriptional alterations associated with growth restriction.
KEGG enrichment analysis of DEGs in VCT_TP63 revealed that genes upregulated in sFGR_Smaller placentas were predominantly involved in lipid metabolic processes, ferroptosis, and regulation of lipolysis and steroidogenesis, suggesting an adaptive metabolic reprogramming response (Fig. 2H, top panel). In contrast, downregulated genes were enriched in pathways related to extracellular matrix organization, cell adhesion, tight junction assembly, Rap1 and Apelin signaling, and protein processing in the endoplasmic reticulum (Fig. 2H, bottom panel), indicating compromised structural stability and barrier function.
Within the VCT_LDHA compartment, upregulated DEGs in sFGR_Smaller placentas were associated with mitochondrial oxidative phosphorylation, reactive oxygen species metabolism, proteostasis networks, and neurodegenerative disease-related pathways (Fig. 2I, top panel), reflecting sustained metabolic stress and impaired protein homeostasis. Conversely, downregulated genes were enriched in focal adhesion, actin cytoskeleton regulation, ECM-receptor interactions, and PI3K-Akt and Hippo signaling pathways (Fig. 2I, bottom panel), suggesting attenuated migratory and integrative capacities.
Together, these findings delineate a dual pattern of quantitative and functional remodeling in the VCT compartment of sFGR placentas: selective depletion and destabilization of the structural VCT_TP63 subset, coupled with compensatory expansion and metabolic reprogramming of VCT_LDHA and VCT_LDHA_Cycling subsets. This reorganization reflects an adaptive yet maladaptive trajectory, whereby trophoblast progenitors activate metabolic stress pathways at the expense of cytoskeletal integrity and adhesion competence, potentially impairing placental development and fetal support under growth-restrictive conditions.
Intra-subpopulation heterogeneity and differentiation dynamics of VCT_TP63 and VCT_LDHA subclusters
Fine-grained subclustering of VCT_TP63 and VCT_LDHA cells revealed substantial intra-subpopulation heterogeneity, highlighting distinct adaptive programs engaged by different subsets under sFGR pathological stress (Fig. 3A, B).
A UMAP visualization of VCT_TP63 and VCT_LDHA clusters further subdivided into ten transcriptionally distinct subclusters (five for each main cluster). Each subcluster is labeled (VCT_TP63_1–5 and VCT_LDHA_1–5) and colored separately. B Dot plot displaying selected marker gene expression for each VCT subcluster. Dot size represents the percentage of cells expressing the gene; color intensity reflects average scaled expression. C GO enrichment analysis of marker genes for each VCT_TP63 subcluster, highlighting subcluster-specific functional features such as chromatin organization, RNA splicing, and epithelial development. D GO enrichment analysis of marker genes for each VCT_LDHA subcluster, demonstrating enrichment of mitochondrial translation, oxidative phosphorylation, and stress response pathways in distinct subsets. E Diffusion pseudotime trajectory reconstruction for all VCT subclusters (Monocle 2). Cells are colored by pseudotime progression, suggesting a continuous differentiation continuum from VCT_TP63 subclusters toward VCT_LDHA and VCT_fusing states. F State assignment of individual cells along the inferred trajectory. Five discrete states are inferred, with VCT_TP63 subclusters predominantly occupying early states and VCT_fusing occupying the terminal state. G Trajectory overlaid with subcluster labels to illustrate branch allocation. Notably, VCT_TP63 subclusters distribute along early pseudotime, while VCT_LDHA subclusters align with later pseudotime and bifurcation toward fusing trophoblast. H Trajectory visualization for each subcluster individually, showing the spatial distribution of each subset along the differentiation manifold. I Trajectory visualization of individual samples (SC01, SC03, SC06) and their respective sFGR_Larger/sFGR_Smaller components to confirm consistency across biological replicates and paired twin samples.
VCT_TP63 cells were subdivided into five transcriptionally distinct subclusters, each characterized by specific functional signatures (Fig. 3C). VCT_TP63_1 was enriched for embryonic placenta development, response to vascular endothelial growth factor stimulus, actin cytoskeleton remodeling, and cell differentiation, suggesting involvement in early trophoblast growth and angiogenic integration. VCT_TP63_2 displayed enrichment in GTPase regulation, cell migration, focal adhesion assembly, Wnt signaling, and fatty acid metabolism, reflecting structural integration and metabolic adaptability. VCT_TP63_3 showed enrichment in actin cytoskeleton organization, TGF-β signaling, wound healing, viral defense, and hypoxia response, indicating a role in stress adaptation. VCT_TP63_4 was associated with cAMP-mediated signaling, multicellular development, hormonal responses (notably progesterone), angiogenesis, and TGF-β pathways, suggesting endocrine responsiveness and vascular development regulation. VCT_TP63_5 exhibited enrichment in mitochondrial translational elongation, ATP synthesis, electron transport chain activity, mRNA splicing, and oxidative stress response, reflecting metabolic homeostasis and energy balance maintenance. Transcription factor network analysis further identified SOX6 and RUNX1 as prominent regulators in VCT_TP63_1 and VCT_TP63_2 subsets, respectively.
Similarly, VCT_LDHA subclusters demonstrated diverse biological features (Fig. 3D). VCT_LDHA_1 was enriched in GTPase activity regulation, angiogenesis, cell adhesion, and EGFR signaling, suggesting roles in trophoblast migration and microenvironmental integration. VCT_LDHA_2 displayed enrichment for mRNA splicing, spliceosomal snRNP assembly, calcium signaling regulation, and mitochondrial ATP synthesis. VCT_LDHA_3 was characterized by Type I interferon signaling, antiviral defense, and ISG15 conjugation, with IRF9 and STAT1 identified as key transcriptional regulators, indicating an activated innate immune profile. VCT_LDHA_4 showed strong enrichment in mitochondrial electron transport, oxidative phosphorylation, ATP synthesis, and mRNA processing, reflecting pronounced metabolic activation. VCT_LDHA_5 was enriched for estrogen biosynthesis, positive regulation of inflammatory response, Toll-like receptor signaling, and NF-κB activity modulation, indicating combined endocrine and inflammatory regulatory functions.
Diffusion pseudotime trajectory analysis (Fig. 3E–G) reconstructed a continuous differentiation continuum extending from VCT_TP63 subclusters through VCT_LDHA subsets toward the VCT_Fusing population. Along this trajectory, VCT_TP63 subclusters predominantly occupied early pseudotime states (States 1–3), consistent with progenitor-like transcriptional programs, whereas VCT_LDHA subclusters and VCT_Fusing cells were mainly distributed across later pseudotime states (States 4–5), reflecting progressive differentiation and metabolic activation.
At the early stages of differentiation, VCT_TP63_1 and VCT_TP63_2 subclusters localized at the initial end of the trajectory, demonstrating the highest differentiation potential and primitive features, with enrichment of cell migration, adhesion regulation, and tissue morphogenesis pathways. Notably, VCT_TP63_1 exhibited characteristics consistent with early progenitor reserve states, while VCT_TP63_2 represented an intermediate transition between progenitors and more functionally mature VCT.
During mid-stage differentiation, VCT_TP63_5, VCT_TP63_3, and VCT_TP63_4 progressively occupied the developmental trunk, characterized by activation of steroidogenesis, oxidative stress buffering, and apoptotic response pathways, reflecting metabolic adaptations and stress balancing mechanisms accompanying lineage progression. In the late-mid stages, subclusters such as VCT_LDHA_1, VCT_LDHA_5, and VCT_LDHA_4 became prominent, exhibiting enriched mitochondrial energy metabolism, oxidative phosphorylation, mRNA splicing, and protein homeostasis regulation, indicating continued adaptation to developmental energy demands and oxidative challenges.
Importantly, VCT_LDHA_3 was strongly enriched in the later stages of the trajectory, marked by robust type I interferon-mediated immune responses and antiviral defense programs, suggesting progressive involvement of immune activation and chronic stress signaling in regulating terminal trophoblast differentiation. Ultimately, all VCT subpopulations converged toward the VCT_Fusing terminal cluster, forming the syncytiotrophoblast precursor pool that marks the completion of syncytial differentiation stages. Individual subclusters demonstrated distinct spatial distributions along the differentiation manifold (Fig. 3H), further supporting their functional divergence. Notably, assessments across biological replicates and paired sFGR_Larger/sFGR_Smaller samples confirmed high reproducibility of the trajectory structure and subcluster allocation (Fig. 3I).
Collectively, these findings delineate a complex landscape of trophoblast adaptation in sFGR placentas, wherein subsets within VCT_TP63 and VCT_LDHA manifest specialized programs spanning metabolic activation, inflammatory regulation, stress response, structural integration, and endocrine signaling. This heterogeneity underscores a multifaceted adaptive strategy whereby trophoblast progenitors balance metabolic demands, differentiation progression, and microenvironmental interactions under pathological constraint, offering mechanistic insights and potential therapeutic entry points for placental dysfunction.
Transcriptional regulatory features underlying VCT subpopulation remodeling
To further elucidate the upstream regulatory landscape driving VCT subcluster remodeling in sFGR placentas, we performed SCENIC analysis to infer transcription factor (TF) regulon activity and reconstruct regulatory networks (Fig. 4A–C). The heatmap of regulon activity revealed that key TFs such as SOX6, TEAD1, TCF7L2, YY1, RELA, and HCFC1 displayed distinct activation patterns across refined VCT_TP63 and VCT_LDHA subclusters (Fig. 4A). Specifically, SOX6, TEAD1, and TCF7L2 were predominantly active within VCT_TP63 subsets, while YY1, RELA, and HCFC1 showed strong enrichment in VCT_LDHA subclusters, indicating transcriptional programs aligned with progenitor maintenance or metabolic stress adaptation.
A Heatmap showing the relative activity of transcription factor (TF) regulons across the refined VCT_TP63 and VCT_LDHA subclusters. Regulon activity was inferred by SCENIC analysis, with key TFs including SOX6, TEAD1, TCF7L2, YY1, RELA, and HCFC1 demonstrating distinct subcluster-specific activation patterns. B Regulatory network reconstruction of representative TFs enriched in VCT_TP63 subclusters. Nodes denote TFs and predicted target genes, with edge color intensity reflecting regulatory confidence scores. UMAP feature plots depict the spatial expression distribution of representative TFs including SOX6, TEAD1, and TCF7L2 across the VCT subsets. C Regulatory network reconstruction of representative TFs enriched in VCT_LDHA subclusters. Notably, YY1, RELA, and HCFC1 regulons were prominent in VCT_LDHA clusters. UMAP feature plots demonstrate their specific enrichment patterns within the metabolic-stress-oriented VCT_LDHA subpopulations.
Regulatory network reconstruction highlighted that in the VCT_TP63 system, an integrated module comprising TFAP2C, FOS, RUNX1, TEAD1, CUX1, GRHL2, NR3C1, and BACH1 governed trophoblast differentiation, epithelial barrier stability, polarity-adhesion maintenance, and oxidative stress response (Fig. 4B). For example, RUNX1 and GRHL2 were central to polarity regulation and integration capacity, whereas TEAD1 and BACH1 coordinated Wnt signaling and stress adaptation. In contrast, the VCT_LDHA regulatory landscape was dominated by STAT1–IRF9 (pro-inflammatory amplification), E2F1 (cell cycle and protein homeostasis), YY1, SMARCB1, HCFC1, and RELA (metabolic stress and chromatin remodeling) (Fig. 4C). Notably, YY1 and RELA formed a hub coordinating persistent activation of inflammatory and metabolic pathways, reflecting the metabolic-stress-oriented fate of VCT_LDHA cells.
Diffusion mapping of TF expression across UMAP embeddings further demonstrated that SOX6 expression localized primarily within early-stage VCT_TP63 subclusters, whereas YY1, RELA, and HCFC1 were enriched in metabolically active VCT_LDHA subpopulations (Fig. 4B, C, right panels).
In concert, these analyses define a multilayered transcriptional framework underlying VCT subcluster remodeling in sFGR placentas, integrating epigenetic regulation, metabolic adaptation, stress response, and structural maintenance. This regulatory architecture provides mechanistic insights into trophoblast dysfunction and suggests potential targets for therapeutic intervention in placental insufficiency.
Pro-inflammatory remodeling of the immune microenvironment and pathological alterations in specific immune subpopulations
Single-cell transcriptomic analysis revealed that, in addition to trophoblast functional reprogramming, placental tissues from sFGR pregnancies exhibited a pronounced remodeling of the immune microenvironment characterized by a globally activated pro-inflammatory state (Fig. 5A). Multiple innate and adaptive immune subpopulations showed increased abundance in sFGR_Smaller (growth-restricted) placentas, suggesting sustained immune activation locally. To identify the most consistently altered subsets, we conducted paired comparative analyses across all immune clusters (Fig. 5B).
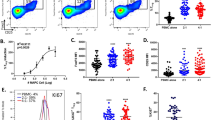
A UMAP plots illustrating refined subclusters of placental immune cells, including T cell subsets (CD4Tcm1, CD4Tcm2, CD4Tem, CD8Tem, Treg, MAIT, and gdT), Hofbauer macrophages (Hofbauer1–4), monocytes (Mono1–4), neutrophils (Neu1–5), and NK cell subsets (NK1–4 and NKT). Each dot represents an individual cell colored by cluster identity. B Paired proportion analysis of immune subclusters comparing sFGR_Larger and sFGR_Smaller placental samples across the three twin pairs. The barplots display mean proportions ± SEM, and connecting lines trace individual sample pairs. Notably, consistent trends in abundance were observed in Hofbauer4, NK3, Neu3, and Neu5 across all pairs, with Hofbauer4 and NK3 reaching statistical significance (*P < 0.05). C Gene ontology enrichment analysis of marker genes defining the significantly altered immune subclusters (Hofbauer4 and NK3). Hofbauer4 cells exhibited enrichment in mitochondrial translation, fatty acid metabolism, and apoptotic pathways, while NK3 cells were characterized by antiviral responses, type I interferon signaling, and apoptotic regulation.
Within the macrophage compartment, the Hofbauer4 subcluster demonstrated a stable and statistically significant reduction in the sFGR_Smaller placentas (P = 0.048, log2FC = −1.22). Functional enrichment of its marker genes highlighted central roles in mitochondrial function and metabolic regulation, including neutrophil degranulation, amino acid metabolism, mitochondrial translational elongation and termination, oxidative phosphorylation, and ATP synthesis (Fig. 5C, left panel). These features suggest that Hofbauer4 macrophages are key regulators of mitochondrial energy homeostasis and oxidative stress buffering in the placental environment. Their depletion under growth-restricted conditions likely reflects compromised metabolic adaptation capacity, which may further impair tissue resilience to adverse intrauterine stress.
Conversely, within the NK cell compartment, the NK3 subcluster exhibited significant expansion in the sFGR_Smaller placentas (P = 0.044, log2FC = 1.09). This subpopulation was transcriptionally characterized by robust enrichment of antiviral defense and type I interferon response programs, including viral genome suppression, interferon-alpha/beta signaling, apoptosis modulation, and glycolytic regulation (Fig. 5C, right panel). KEGG pathway analysis further associated NK3 cells with multiple viral infection pathways and metabolic stress processes. This expansion likely represents a compensatory activation of innate immune surveillance and antiviral defense mechanisms but may also exacerbate local inflammation and trophoblast dysfunction. Notably, other innate immune populations including Neu3 and Neu5 neutrophil subclusters displayed consistent but non-significant trends toward increased abundance in sFGR_Smaller placentas across twin pairs, suggesting broader engagement of pro-inflammatory granulocyte responses under pathological conditions.
Convergently, these findings define a dual-axis immune remodeling in sFGR placentas characterized by the loss of metabolically supportive Hofbauer4 macrophages and the compensatory expansion of interferon-activated NK3 cells. This reorganization of the immune niche likely interacts synergistically with trophoblast stress adaptation and functional impairment, contributing to disrupted placental perfusion and nutrient exchange as an integrated component of sFGR pathogenesis.
Dysregulated crosstalk between villous cytotrophoblast subpopulations and the placental microenvironment in sFGR
Comprehensive cell–cell communication analysis revealed marked remodeling of trophoblast interactions with stromal and immune compartments in sFGR placentas (Fig. 6A). Notably, interactions between VCT_TP63 and stromal cells—including smooth muscle cells (SMC), fibroblasts, and endothelial cells—were significantly attenuated in growth-restricted (sFGR_Smaller) samples. In contrast, VCT_LDHA demonstrated broadly enhanced interactions with innate and adaptive immune cells, particularly Hofbauer macrophages, monocytes, and NK cells, indicating a reorganization of the trophoblast microenvironmental niche under pathological conditions.
A Network visualization of differential interaction strength between villous cytotrophoblast (VCT) subtypes (VCT_TP63, VCT_LDHA) and stromal (smooth muscle cells (SMC), fibroblasts, endothelial), myeloid (Hofbauer cells, monocytes, neutrophils), and lymphoid (T, NK) cell populations in sFGR_Smaller compared with sFGR_Larger placental samples. Edges represent ligand–receptor interactions; red indicates increased interaction strength and blue indicates decreased interaction strength in sFGR_Smaller. B–D Bubble plots of representative ligand–receptor pairs mediating the interactions between VCT_TP63 and stromal cell subtypes. B SMC-derived ligands interacting with VCT_TP63 receptors. C Fibroblast-derived ligands interacting with VCT_TP63 receptors. D Endothelial-derived ligands interacting with VCT_TP63 receptors. Dot size represents the statistical significance of each interaction (–log10 P value), and color scale indicates the mean expression level. Notable reductions in adhesion-related and Wnt/Notch signaling interactions were observed in sFGR_Smaller. E Bubble plot of representative ligand–receptor pairs mediating the interactions between immune cells (Hofbauer macrophages, monocytes, NK cells) and VCT_LDHA. sFGR_Smaller placentas demonstrated enhanced pro-inflammatory, including increased HLA-E–NKG2A/NKG2C and VEGFA–FLT1 signaling. SMC smooth muscle cell, NK natural killer cell, NRP1/2 neuropilin 1/2, FLT1 VEGF receptor 1.
In the stromal compartment, the reduction in SMC–VCT_TP63 crosstalk was driven by decreased signaling via TGFβ1–TGFβ receptor complexes, ADGRE5–CD55, and VEGFA–FLT1 ligand–receptor pairs (Fig. 6B). These interactions are implicated in maintaining trophoblast differentiation, modulating local immune tolerance, and promoting angiogenic support. Fibroblast–VCT_TP63 interactions showed diminished engagement of WNT5A–FZD5/LRP6, VEGFA–FLT1/NRP1, and DHEAsulfate–SULT2B, indicating compromised regulation of Wnt-dependent migratory cues, metabolic adaptation, and extracellular matrix dynamics (Fig. 6C). Endothelial–VCT_TP63 signaling was weakened through the loss of adhesion- and growth-regulatory axes including FN1–integrin complexes (αVβ1/β5/β6/β4), IGF1–IGF1R, and NECTIN2–JAM3, reflecting impaired trophoblast–vascular interface integrity and structural maintenance (Fig. 6D).
Conversely, the immune compartment demonstrated significant activation of trophoblast–immune communication in sFGR_Smaller placentas. Specifically, VCT_LDHA displayed enhanced engagement with Hofbauer macrophages, monocytes, and NK cells via upregulated HLA-E–NKG2A/NKG2C, VEGFA–FLT1, CXCL10–CXCR3, and TGFβ1–TGFβR ligand–receptor interactions (Fig. 6E). These signaling axes collectively orchestrate pro-inflammatory and metabolic stress responses, including immune recognition modulation, antiviral defense, and reactive oxygen species adaptation. In particular, the increased HLA-E–NKG2 interactions indicate heightened trophoblast immunogenicity or stress-induced ligand expression, potentially fueling chronic immune activation.
Validation of trophoblast subpopulation signatures and regulatory hubs
To verify the single-cell findings, we performed multiplex immunofluorescence (mIF) and qPCR validation in an expanded cohort. mIF staining confirmed a significant reduction of VCT_TP63 markers (TP63, SEMA6D) (Fig. 7A) and a concomitant expansion of VCT_LDHA markers (LDHA, S100A9) (Fig. 7B) in sFGR_Smaller placental villi. Furthermore, qPCR analysis of key transcription factors identified by SCENIC demonstrated that progenitor-associated regulators SOX6 and NR3C1 were significantly downregulated, while metabolic stress and inflammatory hubs including YY1, RELA, and HCFC1 were markedly upregulated in growth-restricted placentas (Fig. 7C). These experimental results corroborate the predicted trophoblast fate reprogramming and provide molecular evidence for the pathogenesis of sFGR.
Representative multiplex immunofluorescence staining of placental villi from sFGR_Larger (A) and sFGR_Smaller (B) cotwins. TP63 and SEMA6D were used as principal markers of the VCT_TP63 subpopulation; LDHA and S100A9 as key markers of the VCT_LDHA subpopulation; CDH1 as a general VCT marker. Nuclei were counterstained with DAPI (blue). C Quantitative RT-PCR validation of representative transcription factors and subpopulation markers in paired placental samples (n = 7 twin pairs; each dot represents one pregnancy). Data are presented as mean ± s.d. and were analyzed using two-tailed paired Student’s t tests. The results demonstrate downregulation of SOX6 and NR3C1 and upregulation of YY1, RELA and HCFC1 in sFGR_Smaller placentas. *P < 0.05.
Overall, these findings delineate a dual-axis remodeling of trophoblast crosstalk in sFGR: a substantial loss of stromal support and adhesion-regulatory signaling from SMC, fibroblasts, and endothelial cells to VCT_TP63, juxtaposed with a compensatory or maladaptive increase in pro-inflammatory immune interactions targeting VCT_LDHA. This imbalance may exacerbate trophoblast dysfunction and microenvironmental disorganization, ultimately impairing placental structural integrity and nutrient exchange capacity (Fig. 8).
Schematic summary of three interconnected axes of placental dysfunction in sFGR: structural dysfunction (depletion of structurally supportive VCT_TP63 cells and loss of barrier integrity), metabolic disorder (expansion of VCT_LDHA metabolic-stress cells with upregulated oxidative phosphorylation, ROS metabolism and glycolysis), and immune imbalance (immune microenvironment remodeling and formation of a pro-inflammatory milieu). These processes reinforce each other, resulting in placental insufficiency and ultimately sFGR.
Discussion
This study represents the first single-cell transcriptomic analysis of MCDA twin placentas with sFGR. By leveraging paired placental samples, we systematically delineated trophoblast subpopulation dynamics, functional remodeling, transcriptional regulation, and microenvironmental interactions. Compared to previous studies based on singleton samples9,10,11,12, the MCDA twin design minimizes confounding from fetal and maternal genetic background, thereby enhancing the robustness and interpretability of the findings. Prior investigations have established that trophoblast invasion defects13, mitochondrial dysfunction14,15, chronic local inflammation16, oxidative stress17, and oxygen-regulated villous development 18 are central to FGR pathophysiology, but detailed insights into VCT subpopulation heterogeneity and differentiation trajectories have remained limited.
Our results demonstrated that the VCT lineage is not a homogeneous population but rather comprises distinct subclusters with specialized functional signatures. The structural-supportive VCT_TP63 subpopulation showed a significant reduction in sFGR smaller twin placentas, characterized by enrichment of pathways regulating cell adhesion, extracellular matrix remodeling, epithelial barrier maintenance, and stemness-related transcription factors, including GTPase activity regulation, TGF-β signaling, and pluripotency maintenance programs. In contrast, the metabolic-stress VCT_LDHA subpopulation was markedly expanded in growth-restricted placentas and exhibited pronounced functional reprogramming, reflected by enrichment in mitochondrial oxidative phosphorylation, reactive oxygen species metabolism, mRNA splicing, and protein homeostasis regulation pathways, suggesting a compensatory metabolic adaptation to hypoxic and nutrient-restricted conditions.
Pseudotime trajectory analysis revealed a continuous dynamic process in which VCT differentiation progresses from early stem-like subclusters (VCT_TP63_1 and VCT_TP63_2) through intermediate transitional states (VCT_TP63_3, VCT_TP63_4, VCT_TP63_5) toward late-stage metabolically active subclusters (VCT_LDHA_1, VCT_LDHA_2) and finally converges on the VCT_Fusing terminal state. Along this trajectory, early VCT_TP63 subpopulations exhibited epithelial maintenance and proliferative signatures, whereas later VCT_LDHA subpopulations showed progressive upregulation of metabolic adaptation and inflammatory response markers, implying that trophoblasts under pathological stress may accelerate fate drift to compensate for functional deficits. This model aligns with recent systematic reviews and mechanistic studies highlighting mitochondrial dysfunction and metabolic imbalance in FGR placentas19,20,21.
Furthermore, our findings may also intersect with the concept of placental ageing, which has been implicated in FGR and other placental insufficiencies. Accelerated ageing, characterized by telomere shortening, mitochondrial dysfunction, and senescence-associated secretory phenotypes (SASPs), could contribute to the metabolic stress and inflammatory milieu we observed in sFGR placentas22,23. The depletion of VCT_TP63 progenitors and expansion of metabolically stressed VCT_LDHA subsets may reflect an accelerated trophoblast turnover and exhaustion, akin to premature ageing. Previous studies have shown that placental ageing is marked by increased oxidative stress and mitochondrial dysfunction24, which aligns with our findings of metabolic reprogramming and impaired trophoblast function in sFGR. Additionally, the loss of structural VCT_TP63 cells could reflect a premature decline in trophoblast regenerative capacity, exacerbating placental dysfunction in the context of FGR. Future studies integrating markers of cellular senescence and oxidative damage could further elucidate the role of placental ageing in sFGR pathogenesis.
Transcription factor network analysis revealed a coordinated regulatory reprogramming in sFGR, where stemness- and barrier-related regulators (e.g., SOX6, NR3C1) were suppressed, while oxidative stress and metabolic adaptation regulators (e.g., YY1, RELA, HCFC1) were activated, underscoring the importance of transcriptional plasticity in disease progression. These factors have been previously implicated in trophoblast invasion, inflammatory signaling, and oxidative stress responses. For example, YY1 knock-down impaired extravillous trophoblast outgrowth by downregulating MMP2 in first-trimester villous explants25, and NF-κB/RELA activation has been linked to altered trophoblast invasion and placental spiral-artery remodeling26,27. These findings suggest that YY1 and RELA may serve as potential biomarkers for placental dysfunction in sFGR. In our study, TP63 emerged as a well-established transcriptional regulator of trophoblast stem/progenitor maintenance and differentiation. Previous research has highlighted TP63’s critical role in the regulation of trophoblast progenitors, with studies showing that TP63 expression is essential for maintaining trophoblast stemness and promoting trophoblast differentiation during placental development28,29. Given its role in trophoblast biology, TP63 may serve as a valuable biomarker of trophoblast progenitor health in the placenta, and its expression could potentially be used to monitor placental dysfunction, particularly in the context of sFGR. On the other hand, SOX6 is another transcriptional regulator with a potential role in trophoblast differentiation, although its function in human placental tissue remains less well-characterized and requires further validation.
In the immune microenvironment, we observed significant immune remodeling in sFGR placentas, characterized by a dual remodeling pattern in the immune microenvironment. On the one hand, Hofbauer macrophage subpopulations associated with mitochondrial protein homeostasis were decreased, which may impair the trophoblast environment and alter trophoblast differentiation and invasion30. On the other hand, NK cell subpopulations enriched in type I interferon signaling and antiviral programs were expanded, promoting an inflammatory environment in the placenta31,32. These immune alterations were mirrored in the trophoblast-stromal cell interactions, as cell–cell communication analysis revealed attenuated adhesive and morphogenetic ligand–receptor interactions, such as FN1-integrin and Wnt-related ligands, suggesting impaired trophoblast invasion and remodeling. Concurrently, we observed enhanced immune-trophoblast crosstalk involving chemokine and interferon pathways, including HLA-E–NKG2 and CXCL10–CXCR3, which are known to promote inflammation and immune activation. These findings suggest that local inflammation and metabolic stress jointly shape trophoblast dysfunction in sFGR. Such immune dysregulation at the maternal-fetal interface is consistent with previous reports of immune alterations in FGR and preeclampsia26,27,28,29,30.
Moreover, these changes in immune landscape, particularly the expansion of NK cells and the loss of Hofbauer cells, could contribute to altered trophoblast function through both secreted factors and direct cell-to-cell interactions. Elevated pro-inflammatory cytokines, such as TNF-α and IFN-γ, secreted by NK cells, have been shown to inhibit trophoblast invasion and disrupt placental vascular remodeling33. Additionally, direct interactions between NK cells and trophoblasts, mediated by receptors such as NKG2D and KIRs, could further compromise trophoblast function by inducing apoptosis or limiting trophoblast differentiation34,35. These combined immune alterations, along with changes in trophoblast-stromal cell signaling, likely contribute to the pathophysiology of sFGR, impairing placental development and fetal growth. Further studies are needed to explore how these immune-trophoblast interactions influence placental function and contribute to adverse pregnancy outcomes.
This study has several limitations: While our sample size (n = 3 pairs) is constrained by the rarity of well-phenotyped MCDA twin placentas with sFGR, the paired design provides intrinsic control for genetic and environmental confounders, enhancing the reliability of our findings. Nevertheless, the modest sample size may limit the generalizability of some immune subpopulation trends (e.g., Neu3, Neu5) that showed consistent but non-significant changes. Validation in larger cohorts is warranted. As noted, multinucleated STBs are challenging to capture in single-cell RNA sequencing, which typically isolates individual cells rather than multinucleated ones. In our study, we utilized single-cell RNA sequencing to profile the transcriptomes of placental cells. However, because STBs are large and multinucleated, they are difficult to isolate as single units using standard single-cell RNA-seq methods. As a result, these cells are underrepresented in our dataset. The VCT_Fusing cluster, which we identified, likely represents an earlier stage of trophoblast fusion, but does not fully capture the mature, multinucleated STBs. Future research using alternative approaches, such as targeted enrichment or single-cell approaches designed to better capture multinucleated cells, could provide a more comprehensive understanding of STB biology. Moreover, it is important to note that our study design is cross-sectional, and thus temporal or causal relationships between VCT state shifts and the onset of sFGR cannot be established. Future longitudinal studies incorporating mid-gestation chorionic villus sampling, serial Doppler ultrasounds, and cell-free RNA/mtDNA profiling from maternal plasma or amniotic fluid could help determine whether VCT transcriptional drift precedes overt growth discordance and hemodynamic changes.
In summary, our study using MCDA twin paired single-cell RNA sequencing provides the first systematic characterization of trophoblast subpopulation dynamics, transcriptional regulation, and disrupted interactions with stromal and immune cells in sFGR placentas, culminating in a tripartite pathogenesis model that integrates structural, metabolic, and immune axes of placental dysfunction (Fig. 8). These findings highlight the combined roles of early progenitor depletion and metabolic-stress-driven trophoblast expansion, laying an important foundation for future mechanistic research and informing potential strategies for early detection and targeted intervention.
Data availability
The single-cell RNA sequencing data generated in this study have been deposited in the Gene Expression Omnibus (GEO) database under accession code GSE314816. The numerical source data for qPCR validation (Fig. 7C) are provided as Supplementary Data 1. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The custom analysis code is not publicly available due to ongoing development and integration with proprietary laboratory information systems. However, the code is available from the corresponding author upon reasonable request for academic, non-commercial purposes.
References
Burton, G. J. & Fowden, A. L. The placenta: a multifaceted, transient organ. Philos. Trans. R. Soc. Lond. B Biol. Sci. 370, 20140066 (2015).
Guttmacher, A. E., Maddox, Y. T. & Spong, C. Y. The Human Placenta Project: placental structure, development, and function in real time. Placenta 35, 303–304 (2014).
Lewi, L. et al. Monochorionic diamniotic twins: complications and management options. Curr. Opin. Obstet. Gynecol. 15, 177–194 (2003).
Khalil, A. et al. ISUOG Practice Guidelines: role of ultrasound in twin pregnancy. Ultrasound Obstet. Gynecol. 47, 247–263 (2016).
Valsky, D. V., Eixarch, E., Martinez, J. M., Crispi, F. & Gratacós, E. Selective intrauterine growth restriction in monochorionic twins: pathophysiology, diagnostic approach and management dilemmas. Semin Fetal Neonatal Med. 15, 342–348 (2010).
Gordijn, S. J. et al. Consensus definition of fetal growth restriction: a Delphi procedure. Ultrasound Obstet. Gynecol. 48, 333–339 (2016).
Delorme-Axford, E., Sadovsky, Y. & Coyne, C. B. The Placenta as a Barrier to Viral Infections. Annu. Rev. Virol. 1, 133–146 (2014).
Tsang, J. C. H. et al. Integrative single-cell and cell-free plasma RNA transcriptomics elucidates placental cellular dynamics. Proc. Natl. Acad. Sci. USA 114, E7786–e7795 (2017).
Liu, Y. et al. Single-cell RNA-seq reveals the diversity of trophoblast subtypes and patterns of differentiation in the human placenta. Cell Res. 28, 819–832 (2018).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353 (2018).
Suryawanshi, H. et al. A single-cell survey of the human first-trimester placenta and decidua. Sci. Adv. 4, eaau4788 (2018).
Pique-Regi, R. et al. Single cell transcriptional signatures of the human placenta in term and preterm parturition. Elife 8, e52004 (2019).
Meinhardt, G. et al. Pivotal role of the transcriptional co-activator YAP in trophoblast stemness of the developing human placenta. Proc. Natl. Acad. Sci. USA 117, 13562–13570 (2020).
Holland, O. J. et al. Placental mitochondrial adaptations in preeclampsia associated with progression to term delivery. Cell Death Dis. 9, 1150 (2018).
Singh, A., Jaiswar, S. P., Priyadarshini, A. & Deo, S. Linking of oxidative stress and mitochondrial DNA damage to the pathophysiology of idiopathic intrauterine growth restriction. Int. J. Health Sci. 17, 15–22 (2023).
Firatligil, F. B. et al. Evaluation of systemic immune-inflammation index for predicting late-onset fetal growth restriction. Arch. Gynecol. Obstet. 310, 433–439 (2024).
Burton, G. J. & Jauniaux, E. Oxidative stress. Best. Pr. Res Clin. Obstet. Gynaecol. 25, 287–299 (2011).
Maltepe, E. & Fisher, S. J. Placenta: the forgotten organ. Annu. Rev. Cell Dev. Biol. 31, 523–552 (2015).
Hu, Y. et al. Mitochondrial dysfunction and oxidative stress in selective fetal growth restriction. Placenta 156, 46–54 (2024).
Jansson, T. & Powell, T. L. Role of placental nutrient sensing in developmental programming. Clin. Obstet. Gynecol. 56, 591–601 (2013).
Kiyokoba, R. et al. Mitochondrial dysfunction-induced high hCG associated with development of fetal growth restriction and pre-eclampsia with fetal growth restriction. Sci. Rep. 12, 4056 (2022).
Kajdy, A. et al. Evidence of placental aging in late SGA, fetal growth restriction and stillbirth-a systematic review. Biomedicines 11, 1785 (2023).
Paules, C. et al. Premature placental aging in term small-for-gestational-age and growth-restricted fetuses. Ultrasound Obstet. Gynecol. 53, 615–622 (2019).
Sultana, Z. et al. Oxidative stress, placental ageing-related pathologies and adverse pregnancy outcomes. Am. J. Reprod. Immunol. 77, https://doi.org/10.1111/aji.12653 (2017).
Tian, F. J. et al. The YY1/MMP2 axis promotes trophoblast invasion at the maternal-fetal interface. J. Pathol. 239, 36–47 (2016).
Socha, M. W. et al. The role of NF-κB in uterine spiral arteries remodeling, insight into the cornerstone of preeclampsia. Int. J. Mol. Sci. 22, 704 (2021).
Armistead, B., Kadam, L., Drewlo, S. & Kohan-Ghadr, H. R. The role of NFκB in healthy and preeclamptic placenta: trophoblasts in the spotlight. Int. J. Mol. Sci. 21, 1775 (2020).
Li, Y., Moretto-Zita, M., Leon-Garcia, S. & Parast, M. M. p63 inhibits extravillous trophoblast migration and maintains cells in a cytotrophoblast stem cell-like state. Am. J. Pathol. 184, 3332–3343 (2014).
Wang, L. J. et al. Functional antagonism between ΔNp63α and GCM1 regulates human trophoblast stemness and differentiation. Nat. Commun. 13, 1626 (2022).
Reyes, L. & Golos, T. G. Hofbauer cells: their role in healthy and complicated pregnancy. Front. Immunol. 9, 2628 (2018).
Wei, X. & Yang, X. The central role of natural killer cells in preeclampsia. Front. Immunol. 14, 1009867 (2023).
Zhou, J., Yan, P., Ma, W. & Li, J. Cytokine modulation and immunoregulation of uterine NK cells in pregnancy disorders. Cytokine Growth Factor Rev. 81, 40–53 (2025).
Gaynor, L. M. & Colucci, F. Uterine natural killer cells: functional distinctions and influence on pregnancy in humans and mice. Front. Immunol. 8, 467 (2017).
Yang, X., Yang, Y., Yuan, Y., Liu, L. & Meng, T. The roles of uterine natural killer (NK) cells and KIR/HLA-C combination in the development of preeclampsia: a systematic review. Biomed. Res. Int. 2020, 4808072 (2020).
Shmeleva, E. V. & Colucci, F. Maternal natural killer cells at the intersection between reproduction and mucosal immunity. Mucosal. Immunol. 14, 991–1005 (2021).
Acknowledgements
This project is supported by the Key Research and Development Program of the Ministry of Science and Technology (2023YFC2705901), Shanghai Municipal Science and Technology Commission (22Y11902300, 23DZ2300300), Shanghai Municipal Health Commission (202440131), Shanghai Jiao Tong University STAR Grant (YG2023ZD26, YG2024QNA59).
Author information
Authors and Affiliations
Contributions
Yan Bi: Conceptualization, Formal analysis, Investigation, Visualization, Writing – original draft, Writing – review & editing. Jiawen Yang: Methodology, Visualization. Xiaoyu Li: Validation, Data curation. Yuhong Lin: Resources, Investigation. Yucheng Hu: Investigation, Methodology. Xiang Ying: Methodology, Software. Li Gao: Resources. Yanlin Wang: Funding acquisition, Project administration, Supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Kaliya Georgieva. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bi, Y., Yang, J., Li, X. et al. Single-cell insights into trophoblast heterogeneity and adaptive dysfunction in selective fetal growth restriction. Commun Biol 9, 387 (2026). https://doi.org/10.1038/s42003-026-09798-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09798-2