Abstract
Organic biocrystals support essential functions or drive pathologies in numerous living organisms. Here we focus on the nucleation of hematin crystals, which form in malaria parasites as a part of their heme detoxification pathway. Suppression of hematin crystal nucleation has proven the most productive strategy to treat malaria, yet little is known about the relevant molecular mechanisms. We show that crystal nucleation can be suppressed and fine-tuned via the properties of a population of precursors that host nonclassical hematin crystal nucleation. The addition of modifiers selectively invokes one of three outcomes: suppressed nucleation, faster nucleation, or no effect. We demonstrate that β-hematin crystal nuclei form within mesoscopic hematin-rich clusters and that the impacts of the modifiers on crystal nucleation parallel their activity towards the nucleation precursors. Molecular simulations reveal that modifiers’ activities derive from their interactions with the hematin monomers, dimers, and larger agglomerates. Collectively, these observations support a general method to control crystal nucleation that relies on solute-modifier interactions and their consequences for the nucleation precursors. The proposed rationale offers a powerful tool to control nucleation in areas that employ tailored crystalline materials and helps to understand how crystal assemblies with elaborate superstructures appear in nature.
Similar content being viewed by others
Introduction
Hematin (Fig. 1a) is an iron-containing pigment formed after oxidation of heme, the main prosthetic group of hemoglobin1. Hematin accumulates in the digestive vacuole of malaria parasites after catabolism of hemoglobin, where it is sequestered as hemozoin crystals (Fig. 1b) as a part of the parasites’ heme detoxification mechanism2,3. Hematin represents an example of organic biocrystals, which in humans can trigger several pathologies, e.g., cholesterol in atherosclerosis and gallstones, urate in gout, or L-cystine in kidney stones4,5, but in other organisms carry out essential functions6,7,8,9. In contrast to inorganic biominerals, such as calcite or apatite, whose nucleation, growth, and assembly into superstructures have been explored in depth10,11,12, the molecular mechanisms of organic biocrystallization has received less attention. Organic biocrystals often form in organic solvents, both in vivo13,14,15,16,17 and in vitro18, but the modification of the crystallization mechanisms by the dominance of van der Waals forces, whose strength, range, and directionality diverge from those of the hydrogen bonds and polar attraction that regulate water-containing solutions19,20,21, have largely been overlooked.
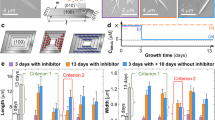
a The structure of hematin: teal, C; red, O; blue, N; and gold, Fe; H atoms are omitted. b An electron micrograph of biogenic hematin crystals, called hemozoin, isolated from Plasmodium falciparum CamWT strain. c A scanning electron micrograph (SEM) image of a laboratory-grown β-hematin crystal, isostructural with hemozoin2. d Hematin molecules in β-hematin link by Fe-O-C(O) bonds and H-bonds between two carboxyl groups, highlighted with solid and dashed lines, respectively; elements are color coded as in panel (a). e The structures of the modifiers tested for their effects on crystal nucleation. Groups that may protonate or deprotonate are colored in blue and red, respectively. f Evolutions of the autocorrelation functions \({g}_{2}\left(\tau \right)\) of light scattered from a 0.50 mM hematin solution in the absence of modifier (left) and in the presence of 10 μM H-ARS (middle) and 4 μM PY (right). See Supplementary Figs. S1, S2 for additional \({g}_{2}\left(\tau \right)\) evolutions. g The effects of H-ARS, CQ, MQ and PY on the amplitudes of the autocorrelation functions at two hematin concentrations: 0.28 mM (left) and 0.50 mM (right).
Here we focus on β-hematin crystals, the synthetic analogue of hemozoin2, and the first stage of β-hematin crystallization, nucleation of new crystals22. Numerous industrial and biomedical tasks require regulation of crystal nucleation23,24,25,26,27, whereas in nature microcomponents are known to mold the elaborate structures of bones, shells, and exoskeletons through precise control of the time and location of crystal formation28,29,30. To date, how modifiers impact crystal nucleation in natural and engineered systems remains elusive30,31,32.
Classical nucleation theory (CNT) provides a commonly applied framework to understand crystal nucleation and how foreign compounds may affect it33,34,35,36,37. The classical approach assumes that solute molecules assemble directly into a crystal nucleus. A crucial limitation of CNT is that at the commonly realized supersaturations, at which the concentrations exceed the solubility by about two-fold, the CNT-predicted nucleation barrier would be high and the resulting nucleation rate may be exceedingly low. Hematin crystallization presents one of a few systems, for which the classical nucleation barrier can be estimated, owing to the known value of surface free energy γ = 25 ± 2 mJ m2 of the interface between a crystallizing solution and a hematin crystal38,39. With this γ, the classical nucleation barrier \(\Delta {G}^{* }=\alpha {\Omega }^{2}{\gamma }^{3}\Delta {\mu }^{-2}\) ≈ 10−17 J (\(\alpha\) ≈ 30, shape factor33,36; \(\Omega\) = 0.708 nm3, molecular volume in the crystal2; \(\Delta \mu\) ≈ \({k}_{B}T\), excess of the solution chemical potential over that of the crystal, approximately equal to the thermal energy at the assumed supersaturation level). This barrier exceeds the thermal energy \({k}_{B}T\) by about 2500-fold. Flat heterogeneous substrates, proposed to accelerate classical crystal nucleation33,36, would lower that barrier by two-fold; crevices in rough substrates, by about 10-fold, keeping the barrier sufficiently high to completely shut off nucleation that would follow a classical pathway.
An additional constraint of CNT is that it provides a limited set of means for crystal nucleation control. CNT highlights three variables that govern the rate of crystal nucleation: the solute concentration, the excess chemical potential \(\Delta \mu\), and the surface free energy γ35,36,40. This \(\Delta \mu\), which relates to the concentrations of solute and third components, such as antisolvents, is a coarse regulator of nucleation. Modest concentration variations, directly or by additives that bind the solute in complexes, often completely suppress nucleation or invoke dramatic shifts to unwanted aggregate structures and morphologies36,41. Efforts to modify the free surface free energy γ by adsorbates are constrained by the Gibbs adsorption rule, a corollary of the second law of thermodynamics, which allows adsorbed modifiers only to lower γ33; accordingly, some surfactants were found to enhance nucleation rates42. An additional means to control crystal nucleation rate relies on foreign surfaces22,28,29,30,36. Classical theory predicts that heterogeneous substrates lower the nuclei surface areas and the nucleation barrier33,34 and drive faster nucleation36. Thus, CNT provides several pathways to enhance crystal nucleation, but only suggests a single coarse means to suppress it, by reducing the solute concentration22,36,43.
Here we show that the nucleation of β-hematin selects a nonclassical pathway, in which crystal nucleation is hosted by mesoscopic solute-rich clusters, and in this way its inhibition circumvents the constraints implied by CNT. The surface free energy \(\gamma\) of crystal nuclei hosted by solute-rich clusters may be about 10-fold lower than that of nuclei suspended in the solution and thus the barriers for nonclassical nucleation are lowered by about 1000-fold, which makes crystal nucleation possible44,45. Metastable mesoscopic clusters—solute-rich, dense-liquid droplets a few tens of nanometers across—have emerged as ubiquitous precursors in nonclassical, two-step crystal nucleation. Such clusters were demonstrated to exist well inside the one-phase region of protein solutions and concentrate the solute to near-crystalline densities, thereby minimizing the free-energy penalty for building a critical nucleus46,47,48,49,50. Direct imaging later revealed that protein crystal nucleation events occur inside these droplets44,45. This concept has since been generalized: mesoscale clusters have been implicated in crystallization of small organic molecules, pharmaceuticals and even porphyrins bonded by van der Waals forces, where tuning solution preparation, pH or additives can modulate cluster populations and thus nucleation kinetics51,52,53,54. We show that the nonclassical pathway of β-hematin crystal nucleation not only allows it to proceed but also provides means to suppress or enhance it by applying compounds that control the volume of the mesoscopic precursors. Importantly, we unravel the molecular interactions between hematin and the modifiers that support or suppress cluster formation.
Results and discussion
Suppression and enhancement of crystal nucleation by antimalarial compounds
We tested the effects on β-hematin (Fig. 1c, d) nucleation of four compounds known to suppress the growth of existing β-hematin: a heme-artesunate adduct (H-ARS), chloroquine (CQ), mefloquine (MQ), and pyronaridine (PY)38,39,55,56,57 (Fig. 1e). As a solvent we choose octanol saturated with citric buffer at pH 4.8 (CBSO), which we, in previous papers15,39, have found to mimic the composition of the lipid nanospheres, suspended in the parasites’ digestive vacuoles, where hemozoin nucleation was found to occur13,14,15. This solvent contains ca. 5 wt. % water58 and dissolves both hematin and the tested modifiers15,59,60. In the presence of water, the carboxyl groups of H-ARS may deprotonate and charge negatively, whereas the nitrogen-containing functional groups of CQ, MQ and PY may protonate and acquire positive charge (Fig. 1e).
We evaluated the crystal nucleation rate by dynamic light scattering (DLS), tracking the evolution of the autocorrelation function \({g}_{2}\left(\tau \right)\) of light scattered by the solution (where τ is the lag time). At early crystallization times, the autocorrelation functions were close to negligeable and revealed a homogeneous solution and the lack of any supramolecular assemblies (Fig. 1f, Supplementary Figs. S1, S2). In solutions with concentration C lower than that of crystal solubility Ce (Ce = 0.12 mM at the temperature of the experiments, 22 °C)39,59, \({g}_{2}\left(\tau \right)\) never deviated from the initial near-zero value. When C > Ce, a prominent shoulder in \({g}_{2}\left(\tau \right)\) appeared and grew (Fig. 1f, Supplementary Figs. S1, S2) within several minutes of solution preparation. We concluded that the growing shoulder manifests relatively slow nucleation of β-hematin. These crystals, driven by Brownian collisions with the solvent, create regions of high number density N. The density fluctuations scatter the incident light and when they inevitably dissipate, the scattered intensity decays and the intensities scattered at times separated by longer τ decorrelate49,61,62. Thus, the characteristic time of the \({g}_{2}\left(\tau \right)\) shoulder characterizes the crystallites’ diffusion coefficient63, which, in turn, scales with the crystallite size64. The amplitude A1, roughly the \({g}_{2}\left(\tau \right)\) drop between short and long lag times τ, is proportional to the number and size of the nucleated crystallites and their refractive index difference with the solution. With this, faster A1 gains indicate faster β-hematin nucleation rates. The evolutions of A1 recorded in growth solutions with and without modifiers H-ARS, CQ, MQ, and PY (Fig. 1f, g, Supplementary Figs. S1, S2) reveal that consistently, at two hematin concentrations, H-ARS enhances the nucleation rate (H-ARS-boosted β-hematin nucleation is also seen by atomic force microscopy, AFM, Supplementary Fig. S3); the nucleation rate is indifferent to CQ and MQ; and, intriguingly, crystal nucleation is nearly fully suppressed by PY.
The considerable rate of β-hematin crystal nucleation at concentrations that exceed the solubility by about two-fold and four-fold (Fig. 1f, g, Supplementary Figs. S1, S2) falls outside of the CNT prediction of fully stalled nucleation and suggests that nucleation of β-hematin may follow a nonclassical mechanism. A common feature of nonclassical nucleation is a crucial role for solute assemblies that precede crystals and host and facilitate crystal nucleation11,44,65. The role played by nucleation precursors provides an additional control pathway—by manipulating the properties of the precursor population.
Nucleation hosted by amorphous precursors
To explore the pathway by which β-hematin crystals nucleate, we imaged supersaturated hematin solutions by negative-staining transmission electron microscopy (TEM). In this method, a solution droplet is deposited on an amorphous carbon film and treated with uranyl acetate solution at room temperature66,67. Existing particles in the solution are decorated by the stain, highlighting surface features and patterns68. The stain scatters electrons strongly and creates negative contrast, whereby the particles appear bright against the stain.
Electron micrographs reveal a population of compact-shaped particles with sizes ranging from 70 to 300 nm (Fig. 2a, b, d, e). As hematin is the only component that may aggregate in the examined solutions at room temperature, we identify the particles as hematin aggregates. Electron diffraction patterns indicate that the particle structures are amorphous (Fig. 2c and Supplementary Fig. S4). We observe elongated objects with parallel sharp edges that emerge from a particle (Fig. 2d). The similarity of the sharp-edged objects’ shape to the shapes of β-hematin crystals (Fig. 1b, c) helps us to identify them as β-hematin. The nucleation of these crystals appears to have been assisted by the hosting particle, analogously to crystal nucleation of organic and inorganic compounds within amorphous precursors44,65,69,70,71,72,73.
a, b, d, e Negative staining electron micrographs of amorphous particles and crystals in hematin solutions. c Electron diffraction pattern of the particle in (b) reveals an amorphous signature and lack of order. f Electron diffraction pattern of the particle in (e) exhibit two peaks at 1.24 nm−1 from the image center, highlighted in red circles, which corresponds to crystal lattice spacing of about 8 Å, superimposed on an amorphous pattern. Red arrows in (d and e) indicate crystals, identified from their elongated shapes with sharp parallel edges and, in (e), from its diffraction pattern in (f). In (a, b, and d) yellow arrows point at amorphous particles that do not hold crystals, identified from irregular shapes and diffraction patterns in (c), for particle in (b), and Supplementary Fig. S4f, for particle in (a). In (d and e), orange arrows identify particles that host, in (e), or initiate, in (d), crystals. g Schematic of crystal nucleation hosted by a disordered precursor. h, i Time-resolved AFM monitoring of the evolution of an amorphous particle supported on a large β-hematin crystal. h Evolution of the particle shape at the times indicated on the images. i The height profiles along dashed lines in panel (h); for images at 30, 60, 120, and 360 min see Supplementary Fig. S5.
Select particles exhibit sharp edges that appear to belong to elongated crystals, again likely β-hematin, partially submerged in the particle (Fig. 2e). Supporting this identification of the crystal held within the particle, the diffraction pattern of the particles exhibits peaks at 1.24 nm−1 from the image center (Fig. 2f) superimposed on the amorphous pattern. These peaks corresponds to lattice spacing 8 Å, close to the c lattice parameter of β-hematin crystals, 8.040 Å ref. 2. The (001) reflections are the only ones present in the diffraction pattern; this selection may be due to anisotropic disorder, which weakens the diffraction from the other lattice planes. Some diffraction patterns exhibit hexagonally distributed peaks at 1.98 nm−1 (Fig. 2f and Supplementary Fig. 4f), indicating the presence of crystals with hexagonally symmetric planes spaced 5.05 Å apart. This lattice spacing precludes the possibility that these are the (111) planes of the ubiquitous NaCl crystals, which are spaced at 3.26 Å, and the origin of these peaks remains unclear.
In further support of the role of amorphous particles in crystal nucleation (Fig. 2g), AFM observations reveal rounded particles supported on the surface of a large β-hematin crystal (Fig. 2h, i, Supplementary Fig. S5). The particle shape, compressed normally to the substrate (Fig. 2i), is consistent with a liquid phase that spills along the crystal surface. The particle volume, ca 2 × 105 nm3, is similar to the volume of particles seen with TEM (Fig. 2a–e). Over time, the particle facets and attains sharp edges, indicating a disorder-to-order transition into a crystal (Fig. 2h, i).
Mesoscopic hematin-rich clusters and their response to nucleation inhibitors
For guidance on how the amorphous nucleation precursors may be affected by the four modifiers, we explore their formation mechanism. We monitored hematin solutions with oblique illumination microscopy (OIM) (Supplementary Fig. S6)46,49,50,74,75,76. OIM revealed aggregates (Fig. 3a, Supplementary Fig. S6b, c) that are not present in pure solvent and at low hematin concentration CH and whose number concentration \({N}_{c}\) monotonically increases with CH (Fig. 3e). This steep \({N}_{c}({C}_{H})\) correlation indicates that they do not represent inert heterogeneities and are likely reversible aggregates comprised of hematin: their concentration Nc declines ca. 15-fold in response to a two-fold reduction of hematin concentration (Fig. 3e)50. The aggregate’s size distribution is relatively narrow with an average diameter d of ca. 70 nm (Fig. 3a). These sizes are similar to the sizes of the particles observed by TEM (Fig. 2a, b, d, e); their average volume is of order 105 nm3, similar to the value measured by AFM (Fig. 2h, i). With the molecular volume of hematin ca. 1 nm3, aggregates of such volume may hold ca. 50,000 loosely packed hematin molecules. Both d and Nc are steady for at least one hour (Fig. 3b, c) and the aggregate size is independent of hematin concentration (Fig. 3d). These behaviors stand in contrast to expectations for domains of typical liquid condensates77,78 or crystals, which are generated by a first-order phase transition and whose nucleation and growth persist and d and Nc increase with time79,80.
a Number density Nc distribution of the diameters of clusters determined by oblique illumination microcopy (Supplementary Fig. S6) in a solution with CH = 0.11 mM. Error bars represent the standard deviation from the average of five independent measurements. b, c Evolutions of the average cluster diameter, in (b), and number density, in (c), in a hematin solution with CH = 0.22 mM. d, e Correlations between average cluster diameter, in (d), and cluster number density, in (e), and hematin concentration. Error bars in (b–e) represent the standard deviations from the averages of five independent measurements; horizontal lines in (b–d) represent the average cluster diameters and number densities. The curve in panel (e) is a guide to the eye. f Schematic of the formation mechanism of mesoscopic hematin-rich clusters (blue disk) owing to accumulation of hematin dimers (oval).
On the other hand, the behaviors of the hematin aggregates cohere with previously observed signature behaviors of mesoscopic solute-rich clusters of other organics52,53 and numerous proteins46,47,48,49,50. We conclude that the hematin aggregates are mesoscopic hematin-rich clusters. According to recent models of the mesoscopic clusters, they exist owing to the accumulation of transient solute dimers (Fig. 3f)49,63,81. The cluster size is determined by the balance between the lifetime of the transient dimers and their rate of outward diffusion from the cluster core63,82. The amount of solute captured in the clusters, and the related number of clusters and cluster population volume are governed by thermodynamic equilibrium between the clusters and the bulk solution63,82,83, which enforces a quasi-exponential increase of Nc with the solute concentration, as seen with hematin (Fig. 3e). The mesoscopic clusters of hematin appear to follow the predictions of the dimer-accumulation model. Their sizes, close to those of the β-hematin nucleation precursors (Fig. 2), suggest a nonclassical pathway wherein amorphous precursors originate as mesoscopic hematin-rich clusters. The role of mesoscopic hematin-rich clusters in β-hematin nucleation suggests that the observed enhancement and suppression of crystal nucleation by modifiers (Fig. 1f, g, Supplementary Figs. S1, S2) may be due to modifier-driven variations of cluster properties.
To test whether the responses of crystal nucleation to modifiers correlate with those of the mesoscopic hematin-rich clusters, we monitored how the cluster population reacts to increasing concentrations of H-ARS, CQ, MQ, and PY. In compliance with the transient dimer model of cluster formation, the addition of the modifiers does not affect the clusters size (Fig. 4a). These additives, however, invoke disparate responses of the cluster number density Nc: H-ARS boosts Nc, CQ and MQ do not affect it, whereas PY suppresses the clusters—nearly completely at 4 μM (Fig. 4b). Remarkably, the responses of the cluster population volumes to the four modifiers run parallel to the responses of crystal nucleation, up to the complete suppression of cluster formation and β-hematin nucleation by 4 μM of PY (Figs. 1c and 4b), coherently with the role of clusters as crystal nucleation sites (Fig. 2). Collectively, the TEM observation of cluster-assisted nucleation and the parallel trends of additive activity on the cluster population and crystal nucleation support a mechanism of nucleation control employing additives that manipulate the amorphous nucleation precursors.
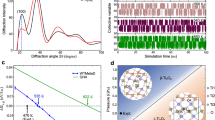
a, b OIM results for solutions with CH = 0.18 mM. The responses of cluster diameters, in (a), and cluster number densities, in (b), to the modifiers. Error bars represent the standard deviation from the average of five independent measurements; horizontal dashed lines represent values in the absence of modifiers. c, e Molecular dynamics (MD) simulation snapshots. c Inhomogeneous distribution of water and octanol in the solvent. Water and octanol O atoms are represented with blue and red spheres, respectively, water H atoms, with silver spheres. The aliphatic chains of octanol are omitted for clarity. d Correlation function g2(τ) and intensity distribution function G(τ) obtained by dynamic light scattering (DLS) from CBSO. e Water accumulates around the carboxyl groups of hematin and H-ARS and the nitrogens of CQ, MQ and PY. Water oxygens represented with blue spheres; water-rich regions are highlighted with a blue tint. Atom color code: C, teal; O, red; N, blue; F, pink; and H, silver. Octanol molecules are omitted for clarity.
Notably, the diverse impacts of the modifiers on the nucleation precursors and the numbers of nucleated crystals do not correlate with the potent inhibition of the growth of hematin crystals by these same modifiers38,39,55,56,57. We posit that the discrepancy of the activity of the modifiers is rooted in the distinct interactions of the modifier molecules with the sites on the surface of a growing crystal and with the mesoscopic hematin-rich clusters. Thus, H-ARS and MQ associate to the kinks along the β-hematin growth steps and activate an inhibition mechanism called kink blocking, whereas CQ and PY adsorb on the terraces between steps and act as step pinners39,55,56. Clearly, the prenucleation clusters, which are amorphous (Fig. 2), do not exhibit kinks, steps, and terraces; the interactions of these modifiers with the hematin-rich clusters are discussed below. Interestingly, the nucleation enhancement, enforced by H-ARS, was found to be a part of a powerful mode of irreversible inhibition of β-hematin crystallization, whereby newly nucleated hematin nanocrystals incorporate into larger crystals, which strains the crystal lattice and suppresses growth57.
How do the modifiers control mesoscopic hematin clusters?
We used a combination of molecular dynamics (MD) simulations and experiments to assign the modifiers’ control of the cluster population to their activity towards hematin monomers, dimers, and large agglomerates. As a first step in elucidating this activity, we explore the state of the solution components: water, octanol, hematin, and modifiers. Molecular simulations84,85 reveal that the solvent is not homogeneous. Instead, water droplets of size several nanometers form, stabilized by H-bonds and enriched with octanol hydroxyl groups (Fig. 4c)86. In support of these computational findings, DLS characterization of CBSO solvent reveals the presence of a low concentration of inhomogeneities of two sizes: ca. 3 nm and ca. 110 nm (Fig. 4d). No inhomogeneities are observed in separate neat octanol and citric buffer solutions (Supplementary Fig. S7), indicating that the inhomogeneities detected in CBSO are droplets of the minority component, water. The smaller of their two sizes is similar to that predicted in simulations, the larger size is likely the result of coalescence and maturation of smaller droplets over laboratory timescales. When hematin or modifiers are present in the solution, water-rich regions envelop the polar groups of hematin and modifiers that have the capacity to form H-bonds with water (Fig. 4e): carboxyl in hematin and H-ARS and nitrogen in the secondary and ternary amine groups of CQ, MQ and PY, in the quinoline groups of CQ and MQ, and the benzo-naphthyridine group of PY (Fig. 1e). It has been suggested that the water-enriched regions wrapped around the hematin carboxyl groups may crucially enable their coordination to the iron and hydrogen bonding to other carboxyl groups that support the β-hematin structure (Fig. 1d)15,87.
The accumulation of water facilitates the dissociation of the carboxyl groups of hematin and H-ARS (Fig. 1a, e) by stabilizing the resulting COO− and H+ ions. The pKa (Ka, acidity constant) values of these groups in water are ca. 5, close to the pH of the citric buffer in the solvent, so that approximately half of the carboxyl groups in both hematin and H-ARS are ionized. The resulting electrostatic repulsion would extend to long distances owing to the low dielectric constant of the solvent outside of the water nanodroplets. Consistently with the repulsion between H-ARS and hematin, UV–vis spectroscopy tests revealed that H-ARS and hematin do not bind (Supplementary Fig. S8). We propose that this repulsion excludes the hematin molecules from a relatively large volume around each H-ARS molecule. The reduction of the volume accessible to the hematin molecules, in turn, lowers the hematin entropy and boosts its chemical potential and the driving force for its cluster formation, an effect similar to crowding and depletion in biological, colloid, and polymer solutions88,89. The resulting increase in cluster population (Fig. 4b) empowers substantially faster β-hematin nucleation (Fig. 1f, g).
To understand the cluster indifference to CQ and MQ and the suppression of cluster formation by PY we perform MD simulations of the collective behaviors of hematin molecules and their response to the presence of CQ, MQ, or PY. The results reveal three crucial elements of the mechanism of mesoscopic aggregation of solute molecules: hematin forms weakly-bound dimers, whose characteristics fit the requirements of the cluster formation mechanism (Fig. 3f); CQ, MQ and PY increase the stability of these dimers; and PY uniquely induces the assembly of disordered hematin agglomerates, which lowers the driving force for cluster formation (Fig. 5a–j). Independent experimental tests support the latter two computational results (Fig. 5k).
a, b, e, h Two-dimensional potential of mean force \(F(\theta ,r)\) between two hematin molecules in CBSO in the absence of modifier, a, and in the presence of CQ, MQ, and PY in (b, e, and h), respectively. Parameters θ and r are the angle between the vectors normal to the porphyrin planes of the hematin molecules and the distance between their hydroxyl O atoms, respectively. Values of \(F\) were computed from well-tempered metadynamics simulations. The local minima in \(F(\theta ,r)\) correspond to dimers i, ii, iii, and iv, whose structures are shown in the outlays of panel (a), with atom color codes: C, in cyan; O, in red, N, in blue; Fe, in gold; and H, in silver. The equilibrium constants KD for hematin dimerization are shown in each plot. The F scale bar is shown in panel (a). c, f, i Representative snapshots at 3 μs of simulation time of hematin aggregates in the vicinity of molecules of CQ (blue sticks in (c)), MQ (red sticks in (f)), and PY (green sticks in (i)). The evolution of the numbers of hematin molecules captured in aggregates per molecule of d CQ, g MQ, and j PY. k Intensity distribution functions G(τ) obtained by DLS from a 0.28 mM hematin solution in CBSO in the presence of 4 μM PY at 14, 135, 237, 389, and 501 min after solution preparation. The peaks at ca. 5 ms are due to the population of crystals in Fig. 1b; and peaks at 300 and 1000 ms reveal the formation of 1–3 μm loose agglomerates holding thousands of hematin molecules. The corresponding autocorrelation functions are shown in Supplementary Fig. S13a.
MD simulations model hematin dissolved in CBSO at pH 4.8, in which water clusters (Fig. 4e) stabilize dissociation of one of its carboxyl groups. The potential of mean force F acting between two hematin molecules in modifier-free solution reveals that hematin may form four weekly-bonded dimers (Fig. 5a). The deepest minimum in \(F(\theta ,r)\) (θ, angle between the porphyrin planes of the hematin molecules; r, distance between the hydroxyl oxygens capping the porphyrin irons) is occupied by a roughly centrosymmetric dimer i (Fig. 5a), in which the hydroxylated iron atoms and the carboxyl groups are oriented in opposite directions. In the second most stable structure, ii (Fig. 5a), the two hematin molecules face each other with their respective irons. The hydroxylated irons protrude from the porphyrin planes and enforce two orientations with distinct tilt angles θ. In dimer iii (Fig. 5a), the porphyrin planes of the constituent hematin molecules adopt parallel orientations with the iron of one of them facing the back of the other. Dimers i, ii, and iii (Fig. 5a) appear to be supported by π−π stacking interactions between the porphyrin cores of the two hematin molecules. Dimer iv is similar to i, but the gap between the two hematin molecules is populated by several octanols resulting in greater r and weaker bonding. Notably, dimers i – iv are distinct from two strongly bound dimers found in aqueous solutions90: the μ-oxo dimer, supported by a covalent Fe–O–Fe bond, and the head-to-tail dimer, similar to the crystal structural element (Fig. 1d). The overall dimerization equilibrium constant, computed by integrating the dimer and monomer areas of \(F(\theta ,r)\)85,91, \({K}_{D}\) = 54 ± 1 pM−1 (Supplementary Table S1, Supplementary Figs. S9, S10). Importantly, the potential minima corresponding to the dimers i – iii fall within the range of several kBT units required for the formation of mesoscopic solute-rich clusters81. The presence of four distinct hematin dimers, each with its own interaction potential, complies with the premise of a modified dimer model of cluster formation92.
CQ is a weak diprotic base, which at pH 4.8 carries two positive charges located on the quinoline and diethyl-amine nitrogens93. By analogy to CQ, we assume that MQ acquires a proton on the piperidine nitrogen and PY acquires three, one on a naphthyridine nitrogen and two on the pyrrolidine nitrogens. The presence of CQ, MQ, and PY substantially strengthens overall dimerization: KD increases four to seven-fold (Fig. 5b, e, h). CQ and PY enforce a transformation of the octanol-mediated dimer iv (Fig. 5a) into a T-shaped dimer, in which the angle θ between the two porphyrin planes is ca. π/2 rad (Fig. 5b, c, h), whereas MQ mostly stabilizes the centrosymmetric dimer i (Fig. 5e, f). The high stability of hematin dimers in the vicinity of CQ and MQ molecules would trap a modifier molecule with two hematin molecules, which is consistent with the formation of H2CQ and H2MQ complexes (H, hematin) observed by UV–vis spectroscopy55. Since the concentrations of CQ and MQ in the solutions, where clusters form (Fig. 3), are ca. 100× lower than that of hematin, the complexation with hematin would completely deactivate these two modifiers, but only lower the hematin concentration by ca. 2%. Such a small decrease in free solute concentration is insufficient to detectably affect the cluster population. Thus, CQ and MQ remain passive bystanders to mesoscopic hematin-rich cluster formation and β-hematin nucleation.
The average number of hematin molecules in complexes stabilized by CQ and MQ remain approximately two or three (Fig. 5c, d, f, g, Supplementary Figs. S11, S12) over simulation times as long as 3 μs. By contrast, PY induces larger agglomerates that may contain more than one PY molecule and bind considerably greater amounts of hematin (Fig. 5i, j). This is consistent with the distribution functions (and the respective correlation functions in Supplementary Fig. S13) of light scattered from hematin solution in the presence of PY reveal the presence of large agglomerates of sizes several microns (Fig. 5k) in addition to the population of ca. 40 nm crystals (Fig. 1f). Despite their large size, the signal from these large scatterers is weak (Supplementary Fig. S13), indicating a low refractive index that may be due to loose packing, as suggested by MD simulations (Fig. 5i). These agglomerates are only seen at longer times, when they have grown sufficiently large to scatter light detectably. In the agglomerates, the binding of hematin to PY and to other hematin molecules appears random, consistent with the unchanged hematin UV–vis spectrum in the presence of PY55. Agglomerates are not seen in pure hematin solutions or in the presence of CQ or MQ (Supplementary Fig. S14). Even if loosely packed, PY-induced agglomerates of such size would capture thousands of hematin molecules per one PY molecule. The resulting lower concentration of free hematin would substantially lower the driving force for cluster formation, the cluster number density Nc, and the rate of crystal nucleation. Notably, the sequestration of hematin into PY-stabilized agglomerates may suppress β-hematin crystal nucleation that follows a classical mechanism. This pathway, however, is proscribed for β-hematin by its very high nucleation barriers.
Conclusions
In summary, we demonstrate that crystal nucleation can be suppressed and fine-tuned through solute-modifier interactions that control the properties of the nucleation precursors. The molecular forces that support the formation of the nucleation precursors and direct their suppression by modifiers are typical for most organic systems. Thus, it is likely that the nucleation control mechanisms that we find will apply to other biological and synthetic organic crystals. The generality of the mesoscopic solute-rich clusters as sites for nucleation of crystals of all classes of materials suggest that similar mechanisms may universally operate to control nucleation in all areas that rely on crystalline materials, from catalyst synthesis to pharmaceuticals production, where nonclassical pathways are pervasive. In a broader context, our results shed light on how elaborate crystal assemblies appear in tissues and rocks. The molecular interactions that we highlight may serve as the basis for a broadly applicable strategy to predict how a modifier may impact crystal nucleation and design nucleation modifiers de novo, including by in silico modifier screening, and in this way shortcut the search for compounds that suppress or enhance nucleation.
Methods
Materials
The following compounds were purchased from Sigma Aldrich: hematin porcine (≥98%), citric acid (anhydrous, ≥99.5%), sodium hydroxide (anhydrous, ≥98%), n-octanol (anhydrous, ≥99%), artesunate (anhydrous, ≥98.0%), chloroquine diphosphate (≥98%), mefloquine hydrochloride (anhydrous, ≥98.0%), pyronaridine tetraphosphate (\(\ge \,\)95%), hypoxanthine, saponin, sodium dodecylsulfate (SDS), sorbitol, sodium bicarbonate (NaHCO3), hypoxanthine, and HEPES buffer (N-2-hydroxyethylpiperazine-N-2-ethane sulfonic acid). The following chemical were purchased from Thermo Fisher Scientific: RPMI 1640 media supplemented with L-glutamine and gentamycin. All reagents were used as received without further purification unless otherwise noted. Deionized (DI) water was produced by a Millipore reverse osmosis ion exchange system (Rios-8 Proguard 2–MilliQ Q-guard).
Solution preparation
Citric buffer at pH 4.8 was prepared by dissolving 50 mM of citric acid in DI water and titrating the solution, under continuous stirring, with the addition of 0.10 M NaOH to reach the desired pH as verified before each experiment using an Accumet Basic pH meter (Thermo Fisher Scientific). Fresh buffers were prepared every month and stored at ambient conditions. We placed 5 mL of citric buffer (pH 4.8) in direct contact with n-octanol at room temperature and allowed 30 min for equilibration. The upper portion of the two-phase system was decanted and denoted as citric buffer-saturated octanol (CBSO). Hematin solutions were prepared by dissolving hematin powder in 8 mL of freshly made CBSO and heating to 70 °C for 7–9 h. The solution was filtered through a 0.2 μm nylon membrane filter (Thermo Fisher Scientific) and the concentration was determined using a previously reported39 extinction coefficient of 3.1 ± 0.1 cm−1 mM−1 at wavelength λ = 594 nm.
For this study, we used four modifiers: heme-artesunate adduct (H-ARS), chloroquine (CQ), mefloquine (MQ), and pyronaridine (PY). The synthesis of H-ARS and preparation of its solutions is discussed below. Solid MQ were added to CBSO and the solutions reached the desired concentration after 2–4 days. CQ and PY were added in excess to CBSO and stored in the dark for 30–45 days, allowing the concentrations to approach the respective solubilities60. All modifier solutions were filtered through 0.2 μm nylon membrane filters and the concentrations were determined by UV–vis spectrometry using a Beckman DU 800 spectrophotometer and extinction coefficients and wavelengths listed in Ketchum et al.60.
Synthesis of hematin adduct of artesunate
Hematin solutions were prepared by a modified method a procedure similar to the one above, in which n-octanol was substituted with n-butanol. Sodium dithionite at ca. 1 mM and artesunate (ARS) at ca. 1 mM were dissolved in DI water and n-butanol, respectively. The heme solution was filtered with a 0.2 μm nylon membrane filter and placed in contact with the dithionite solution in a glass vial to yield a net molar ratio of 1:2:5 hematin: ARS: sodium dithionite. The vial was sealed under flow of nitrogen gas to create an inert atmosphere. The reaction involved the reduction of iron(III) to iron(II) in hematin by dithionite. The system was maintained at 50 °C using a hotplate (Thermo Scientific Super-Nuova Multi-Position Digital Stirring Hotplates). The aqueous and organic phases were rigorously mixed by shaking for ca. 30 s until the color of the solution changed from dark green to pink, indicating the reduction of hematin (containing Fe(III)) to heme (containing Fe(II)). The mixture was kept under static conditions for at least 30 min to allow for the separation of the organic and aqueous phases, after which the artemisinin solution was injected into the organic (top) fraction. The reaction between heme and artemisinin happened immediately after the injection, as judged by the instantaneous change in color from pink to orange. After allowing ca. 30 min for the reaction to reach completion, the organic layer was collected for later purification of the product, heme – artesunate adduct (or H-ARS).
Further details about the procedures to purify and identify H-ART and H-ARS by mass spectroscopy are provided in Ma et al.56.
Light scattering characterization of β-hematin nucleation in hematin solutions
Dynamic light scattering data was collected by an ALV instrument (ALV-GmbH, Germany), which includes ALV goniometer, a He-Ne laser with wavelength 632.8 nm, and an ALV-5000/EPP Multiple tau Digital Correlator. A focused laser beam shines on a cylindrical cuvette with diameter 10 mm so that the illuminated volume is ca. 300 nL. Normalized intensity correlation functions \({g}_{2}(q,\tau )\) were collected at a fixed scattering angle of 90° for 60 s. The characteristic diffusion time \({\tau }_{1}\) for hematin crystallites and the respective amplitude \({A}_{1}\) (which is proportional to the intensity scattered by the crystallites) were calculated by fitting \({g}_{2}(q,\tau )\) with an exponential relation47,62 \({g}_{2}\left(q,\tau \right)-1={\left[{A}_{1}\exp \left(-^{\tau} /{\tau }_{1}\right)+\varepsilon (\tau )\right]}^{2}\), where \(\varepsilon (\tau )\) is background noise in the correlation function. The intensity distribution function \(G(\tau )\), corresponding to each correlation function \({g}_{2}(\tau )\), was computed employing the CONTIN inverse Laplace transform algorithm94.
Atomic Force Microscopy (AFM)
Experiments were performed with a Multimode Nanoscope VIII atomic force microscopes from Digital Instruments. AFM images were collected in tapping mode (i.e. light engage) using Olympus TR800PSA probes (silicon nitride, Cr/Au coated 5/30, 0.15 N m−1 spring constant) with a frequency of 32 kHz. Images were obtained using scan sizes of 0.3–20 μm, scan rates of 0.5–2.5 s−1, 256 scan lines, and various scan angles depending on the orientation of the crystal substrate. The temperature in the liquid cell reached a steady value of 27.8 ± 0.1 °C within 15 min of imaging39. This value was higher than room temperature owing to heating by operation of the AFM scanner. The density of hematin crystal substrates grown on glass disks was monitored with an optical microscope to ensure an equivalent number of crystals for all samples (i.e. minimize potential depletion of free hematin and growth inhibitor due to high total surface area of crystals). The glass slides were mounted on AFM sample disks (Ted Pella) and the samples were placed on the AFM scanner. Supersaturated hematin solutions in CBSO were prepared less than 2 h in advance. The growth solution was loaded into the AFM liquid cell using a 1 mL disposable polypropylene syringe (Henck Sass Wolf), which is tolerant of organic solvents. After loading, the system was left standing for 10–20 min to thermally equilibrate. The crystal edges in optical micrographs were identified to determine the orientation and the crystallographic directions on the upward-facing (100) crystal surface. The crystals were kept in contact with the solution for 0.5–1.5 h to allow their surface features to adapt to the growth conditions.
The scan direction was set parallel to the [001] crystallographic direction and AFM images were collected for 3–5 h. The solution in the AFM liquid cell was exchanged every 30 min to maintain an approximately constant hematin (and inhibitor) concentration. For studies of antimalarials, growth solutions were replaced with ones containing a selected inhibitor concentration. For each assay, the crystal substrates were first allowed to equilibrate (ca. 30 min) in growth solution without added inhibitor prior to addition of solutions containing the inhibitor. For studies assessing irreversible inhibition, a series of solutions with varying hematin and/or inhibitor concentrations were supplied to the AFM liquid cell at periodic imaging times.
AFM observations reveal that in the presence of H-ARS, numerous nanocrystals of size 100–300 nm, indicated with a red arrow, appear in the view field (Fig. 1d). The nanocrystals orient randomly, indicating that they did not form on the surface, where the periodic force field of the underlying lattice may have directed them into alignment, but rather in the solution bulk. They reach the surface likely by Brownian diffusion. The slow migration through the solution allows them to grow to the observed sizes.
Imaging of β-hematin crystals by scanning electron microscopy (SEM)
The crystallization container was capped and placed in an incubator at 23 °C with minimal exposure to light. Crystallization proceeded for 16 days. The microscope slides were rinsed with DI water and dried with pure ethanol. For SEM imaging, the slides and the crystal attached to them were coated with 15 nm gold.
Transmission electron microscopy (TEM) imaging with negative staining
To prepare hematin solution samples we used 2% aqueous uranyl acetate (UA) solution. Briefly, 3 µl of the hematin solution were applied to the grids pre-cleaned in a Solarus 950 plasma cleaner (Gatan), incubated for 20 s, and blotted with filter paper. The grids were immediately stained with UA, the excess of the liquid was blotted with filter paper and the process was repeated. The air-dried grids were transferred into a JEM 2100 electron microscope (JEOL Ltd.) operating at 200 keV equipped with a LaB6 electron gun and US4000 Ultrascan digital camera (Gatan). Total electron dose/image was ~80 electrons/Å2. Image pixel size was 2.87 Å on the specimen scale.
TEM imaging of intracellular hemozoin
Purified digestive vacuoles were isolated. Samples were fixed in the presence of 0.1% malachite green and 0.1% malachite green was again added during osmium post-fixation95. Field emission in lens EM (LEO 1550) was performed on sucrose-purified hemozoin, washed in 50 mM Tris/HCl (pH 8.0), then fixed in the presence of Malachite Green before electron micrograph processing96. The selectively fluorescent, hydrophobic phenoxazone dye, Nile Red (N 3013; Sigma), was used as a neutral lipid probe on SPHz and vacuole preparations before fixation on glass slides95. Images were obtained using a Nikon Eclipse E800 at ×1000 magnification13.
Oblique illumination microscopy
Hematin solutions were monitored with oblique illumination microscopy (OIM), also known as Brownian microscopy or particle tracking46,74,75,76. In this method, a green laser illuminates a thin solution layer at an oblique angle such that the incident beam avoids the lens of a microscope positioned above the sample (Supplementary Fig. S6a)75, This method enables the detection of nano- and microscale objects through light scattered at wavevectors of order µm−1. The scattered intensity is proportional to the sixth power of the scatterers’ sizes; thus, in a solution containing objects of varying size the scattering signal is dominated by larger particles. This feature makes this technique particularly well suited to characterize the size and number distribution of the aggregates that appear as bright spots in OIM micrographs (Supplementary Fig. S6b). The spots are counted by a custom-made image package from Nanosight. The concentration of the observed aggregates is determined from the number of spots in a frame and the observed volume \(V=120 \times 80 \times 5\) μm3 refs. 74,97. OIM records the Brownian trajectory of each particles in the image plane (Supplementary Fig. S6c) and calculates diffusion coefficient from correlation between the mean squared displacement \({d}_{{n}}^{{2}}\) and the lag time ∆t97.
where D is the diffusion coefficient of the observed aggregate.
For all OIM measurements we used the Nanosight LM10-HS microscope (Malvern Panalytical Inc). Hematin solution was loaded into a cuvette with volume about 0.3 mL and depth about 0.5 mm. The wavelength of illumination was 532 nm. Temperature was set at 22 °C. Movies were acquired over 30 s. We found that objects recorded for times shorter than 1 s were interference spots from two or more clusters tracked for significantly longer times. This observation was supported by the estimate that a cluster with diffusivity D2 ≈ 10−12 m2s−1 would be detectable in a focal plane with depth 5 μm for about 25 s. Therefore, we excluded these short-duration objects from the determination of the cluster parameters. Five movies from distinct solution volumes were collected in each tested sample. The numbers of clusters in each range of sizes were averaged.
Tests of complexation of hematin with H-ARS
Hematin solutions with initial volume V0 = 8 mL and CH,0 = 0.3 mM were titrated with H-ARS by adding 0.100 mL aliquots of a solution of the H-ARS in CBSO. This addition was repeated up to 7 times. At each titration step, a 0.3 mL sample of the H-ARS – hematin mixture was decanted and a UV–vis spectrum was collected.
The spectra of hematin in the presence of H-ARS were similar to those in pure hematin solutions (Supplementary Fig. S7). They exhibit two peaks at 400 and 594 nm. The height of both peaks decreased upon addition of all H-ARS–containing solutions. Observed changes in the absorbance upon titration may be associated with three phenomena: dilution of the hematin concentration by the newly added solution volume Vx, decrease of the concentration of free hematin due to complexation with the added H-ARS, and the formation of H-ARS – hematin complexes that absorb in the same wavelength range. To account for the solution dilution, we correct the recorded absorbance Ax to account for the increased solution volume from V0 to V0 + Vx
where CH,0 is the hematin concentration prior to H-ARS addition and CH,T is the analytical concentration of hematin, including the amounts present as H-ARS – hematin complexes, after H-ARS addition, \({C}_{H,T}={C}_{H,0}{V}_{0}{\left({V}_{0}+{V}_{x}\right)}^{-1}\).
After accounting for dilution, the height of the peaks at 400 and 594 nm were independent of the concentrations of added H-ARS, which exhibits weak absorbance at 400 nm. Hence, we chose to monitor changes in the absorbance at 594 nm as a function of the total H-ARS concentration to characterize the H-ARS – hematin complexation equilibrium. The choice of this wavelength offers an additional advantage. The weaker absorbance at 594 nm allowed us to use titration data with CH,0 = 0.3 mM, close to the hematin concentrations tested in all experiments reported in the manuscript.
As an indicator of H-ARS-hematin binding, we use the dependence on the total H-ARS concentration cD of the ratio of Acorr at 594 nm to the absorbance of the pure hematin solution A0 at the same wavelength
where l = 1 cm is the optical path length. Combining Eqs. (1) and (2) yields
Equation (4) announces that if \(\frac{{A}_{{corr}}}{{A}_{0}}\) is independent of the added H-ARS (as seen in Fig. S8b), then the absorbance decrease to Ax is fully attributable to dilution from CH,0 to CH,T and no hematin has been sequestered in a complex.
Molecular dynamics simulations
We performed all-atom molecular dynamics (MD) simulations using GROMACS 2021.498 to investigate the clustering of water around hematin and the antimalarial drugs. To mimic the experimental conditions, simulations of hematin and the drug compounds were performed in a 5% wt. water in n-octanol solution using protonation states for each species consistent with a pH of 4.8. Under these conditions, hematin and its derivative drug H-ARS are partially deprotonated, whereas the other three drug compounds are partially protonated, resulting in the following net charges for each of the species: hematin(−1), H-ARS (−2), CQ(+2), MQ(+1), and PY(+3). Water was simulated using the TIP3P99 model, and n-octanol, hematin, and the drug compounds were modeled via the Generalized AMBER force field (GAFF)100,101 with restrained electrostatic potential (RESP) charges derived at B3LYP/6-31G(d) theory level using the PyRED program102. GROMACS compatible topology files for the hematin monomer and H-ARS were prepared by modifying the hematin dimer model that we used in our previous work86 and substituting missing atom types with parameters from the GAFF, whereas the topology files for CQ, MQ, and PY were prepared using the ACPYPE package103 with the calculated RESP charges. The MD trajectories were propagated using the leap-frog algorithm with a 2 fs integration time step. All bonds were constrained using the LINCS104 algorithm to remove the degrees of freedom associated with fast bond vibrations and facilitate the use of this large time step, and water was modeled as rigid body using the SETTLE105 algorithm. Van der Waals interactions were truncated using a 1 nm cutoff, and periodic boundary conditions were applied in all three dimensions. Coulomb interactions were handled with the particle mesh Ewald method106, using a cutoff of 1 nm for the real-space contributions and an error tolerance of 10−5.
To investigate water cluster formation, MD simulations were performed of systems containing one hematin monomer or a single molecule of a selected drug compound. The hematin or drug molecules were placed in a 6.5 × 6.5 × 6.5 nm3 cubic simulation box and solvated with 380 water and 1000 n-octanol to produce a 5% wt. water in n-octanol solution. An appropriate number of sodium (Na) or chloride (Cl) ions were added to neutralize the total charge of the system. The energy of each system was minimized using the steepest decent algorithm with a stopping criterion of 1000 kJ/mol/nm for the maximum force on an atom. The systems were then equilibrated in the canonical (NVT) ensemble for 10 ns at 300 K using a Bussi-Donadio-Parrinello thermostat107 with a time constant of 0.1 ps. Following equilibration, 120 ns production simulations in the isothermal-isobaric (NPT) ensemble were performed at 300 K and 1 bar by applying a Parrinello-Rahman barostat108 with a time constant of 2 ps. Trajectory data was saved every 10 ps over the last 100 ns of each NPT simulation for subsequent analysis.
To study hematin dimerization, well-tempered metadynamics (WTmetaD)109 simulations were performed of two hematin monomers and a single molecule of a selected drug compound in a 5% wt. water in n-octanol solution using the PLUMED 2. 8.014,110 free energy plug-in for GROMACS 2021.498. The initial configurations were prepared and energy minimized, as described above, and then equilibrated by performing sequential 10 and 20 ns simulations in the NVT (300 K) and NPT (300 K and 1 bar) ensembles, respectively. Equilibrated configurations were used to initialize a set of three independent WTmetaD13 simulations at 300 k and 1 bar for each system. Hematin dimerization in the presence of the drug compounds was characterized using several collective variables (CVs): \(r\) measures the separation between the two hematin monomers and is defined as the distance between the oxygens (O) in the hydroxyl groups that are connected to the iron (Fe) cores on different monomers (Supplementary Fig. S15); \(\theta\) characterizes the relative orientation of the two hematin monomers and is defined as the angle formed by the vectors passing through Fe-O bonds on each monomer (Supplementary Fig. S15); S (=d1 sin ϕ) is the radial distance of a selected central atom on the drug compound from the vector \(\vec{{{\boldsymbol{r}}}}\) joining the two hematin monomers (Supplementary Fig. S15); and N (=d1 cos ϕ) is the axial distance of the selected central atom on the drug compound along the vector \(\vec{{{\boldsymbol{r}}}}\) from one of the hematin monomers. Sampling was performed using CVs {\(r\), \(\theta\)} and {\(r\), \(S,{N}\)} to study hematin dimerization in the absence and presence of drug compounds, respectively. To limit the exploration of CV regions, upper quadratic walls were placed at \(r=3.1\) nm, \({d}_{1}=3.1\) nm, and \({d}_{2}=3.1\,{{\rm{nm}}}\) using a spring constant of \(k=5000\) kJ mol−1 nm−2 for each wall. A bias factor of 10 was used in the WTmetaD simulations and Gaussian potentials of height 1 kJ/mol with widths of 0.1 nm and 0.1 rad were deposited every 1 ps.
Data from the WTmetaD simulations was analyzed using the reweighting method proposed by Tiwary et al.111 to obtain estimates of the potential of mean force. The equilibrium dimerization constant for hematin, \({K}_{D}\), was calculated from two-dimensional potential of mean force \(F\left(r,\theta \right)\) via85,91
where \({P}_{b}\) and \({P}_{u}\) are the probabilities of the bound (dimerized) and unbound (disassociated) states, respectively, \({v}^{\varnothing }=({{c}^{\varnothing }{N}_{{Av}}})^{-1}\)=1.66 nm3 is the standard volume of a single molecule, \({c}^{\varnothing }=1\,{{\rm{mol}}}/{{\rm{L}}}\) is the standard concentration, \({N}_{{Av}}\) is the Avogadro’s number, \({k}_{B}\) is the Boltzmann constant, T is the temperature, \(\beta ={\left({k}_{B}T\right)}^{-1}\), R=3.0\(\,{{\rm{nm}}}\) is the maximum value of \(r\) considered in the integration, and rb is the position of the dividing surface along the order parameter \(r\) that delineates the bound and unbound states. A value of \({r}_{b}\)=1.5 nm was chosen in the calculations to consider contributions from all the distinct types of hematin dimers observed in the simulations (Supplementary Table S2). In each case, the equilibrium dimerization constants were computed by averaging values from three independent WTmetaD simulations (Supplementary Table S3), using the standard deviation to estimate the associated statistical uncertainty.
Lastly, we also performed MD simulations to study the effect of CQ, MQ, and PR on hematin clustering. These simulations were performed using larger systems containing 50 hematin, 4 drug, 1520 water, and 4000 n-octanol molecules, resulting in ca 5% wt. water in n-octanol solution with ~66.83 and ~5.35 mM concentrations of hematin and drug compounds, respectively. Following addition of neutralizing salt ions and energy minimization, the systems were equilibrated by performing sequential 20 ns simulations in the NVT (300 K) and NPT (300 K and 1 bar) ensembles. A 3 μs NPT simulation was then performed for each system, saving trajectory data every 1 ps for subsequent analysis.
Statistics and reproducibility
Statistics and reproducibility of experimental measurements
DLS correlation functions from modifier-free and modifier-enriched hematin solutions, in which β-hematin nucleates, were collected over 60 s. As crystal nucleation occurs over time scales of ca. 60 min (Fig. 1) we used single g2s as a measure of nucleation progress. Averaging over 10 sequential g2s, as done to characterize steady aggregates, would obscure nucleation evolution. To judge the reliability and reproducibly of the data, we carried out data collection several times. As nucleation is intrinsically stochastic, and data are collected from a small volume, each correlation function is unique and slightly deviates from those collected under identical conditions. The overall trends are fully reproducible as seen by comparing the signal from nucleating crystals (Fig. 1, Supplementary Figs. S1, S2, S13, S14), or from water nanodroplets (Fig. 4d in this study and Fig. 1A in Olafson et al.112.) We present individual unaveraged correlation and distribution functions.
Atomic force microcopy presents unique images of structures and their evolutions, which cannot be averaged. All crystal nucleation behaviors observed by AFM and discussed in the paper, however, were reproduced multiple times. We collected three image sequences that reveal the transformation of amorphous particles into β-hematin. We collected more than 10 image sequences that show enhance nucleation of β-hematin in the presence of H-ARS.
In OIM measurements, the scatterers’ trajectories were measured from 30 s movies recorded at 25 frames per second. Each movie contained ca. 100 diffusing scatterers. We collected five movies at each set of conditions and averaged the five cluster size distributions \({N}_{c}(d)\), the average cluster diameter \(d\), and the average cluster number density \({N}_{c}\), evaluated from the individual movies. The error bars in Figs. 3 and 4 of the main manuscript represent the standard deviation from these averages.
Statistics and reproducibility of the MD simulations
Quantities from the simulations were computed by averaging values from saved MD trajectory data. The analysis of water clustering was performed by averaging over frames saved every 10 ps from 100 ns trajectories, whereas the analysis of hematin clustering was performed by averaging over frames saved every 1 ps from 3 \(\mu s\) trajectories. Statistical uncertainties were estimated by computing the standard error using the effective number of independent samples determined from the autocorrelation time obtained from time-series analysis113. The potentials of mean force and dimerization constants were computed by averaging over three independent replicas and using the standard deviations to estimate the associated statistical uncertainties.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets generated during the current study are available as Supporting Information.
References
Poulos, T. L. Heme enzyme structure and function. Chem. Rev. 114, 3919–3962 (2014).
Pagola, S., Stephens, P. W., Bohle, D. S., Kosar, A. D. & Madsen, S. K. The structure of malaria pigment β-haematin. Nature 404, 307–310 (2000).
Weissbuch, I. & Leiserowitz, L. Interplay between malaria, crystalline hemozoin formation, and antimalarial drug action and design. Chem. Rev. 108, 4899–4914 (2008).
Poloni, L. N. & Ward, M. D. The materials science of pathological crystals. Chem. Mater. 26, 477–495 (2014).
Shtukenberg, A. G., Ward, M. D. & Kahr, B. Crystal growth with macromolecular additives. Chem. Rev. 117, 14042–14090 (2017).
Minsky, A., Shimoni, E. & Frenkiel-Krispin, D. Stress, order and survival. Nat. Rev. Mol. Cell Biol. 3, 50–60 (2002).
Addadi, L., Kronik, L., Leiserowitz, L., Oron, D. & Weiner, S. Organic crystals and optical functions in biology: knowns and unknowns. Adv. Mater. 36, e2408060 (2024).
Lemcoff, T. et al. Brilliant whiteness in shrimp from ultra-thin layers of birefringent nanospheres. Nat. Photonics 17, 485–493 (2023).
Deis, R. et al. Genetic control over biogenic crystal morphogenesis in zebrafish. Nat. Chem. Biol. 21, 383–392 (2024).
Smeets, P. J. M., Cho, K. R., Kempen, R. G. E., Sommerdijk, N. A. J. M. & De Yoreo, J. J. Calcium carbonate nucleation driven by ion binding in a biomimetic matrix revealed by in situ electron microscopy. Nat. Mater. 14, 394–399 (2015).
Du, J. S., Bae, Y. & De Yoreo, J. J. Non-classical crystallization in soft and organic materials. Nat. Rev. Mater. 9, 229–248 (2024).
De Yoreo, J. J. et al. Crystallization by particle attachment in synthetic, biogenic, and geologic environments. Science 349, aaa6760 (2015).
Pisciotta, J. M. et al. The role of neutral lipid nanospheres in Plasmodium falciparum haem crystallization. Biochem. J. 402, 197–204 (2007).
Egan, T. J. Haemozoin formation. Mol. Biochem. Parasitol. 157, 127–136 (2008).
Vekilov, P. G., Rimer, J. D., Olafson, K. & Ketchum, M. Lipid or aqueous medium for hematin crystallization? CrystEngComm 17, 7790–7800 (2015).
Lambou-Gianoukos, S. & Heller, S. J. Lithogenesis and bile metabolism. Surg. Clin. North Am. 88, 1175–1194 (2008).
Mughal, J., Katkoori, V. R., Nidorf, S. M., Manu, M. & Abela, G. S. The formation of cholesterol crystals and embolization during myocardial infarction. Am. Heart J. 51, 100509 (2025).
Myerson, A. S. Handbook of industrial crystallization, 3rd ed. (Cambrifge University Press, 2019).
Hu, Y., Liang, J. K., Myerson, A. S. & Taylor, L. S. Crystallization monitoring by Raman spectroscopy: simultaneous measurement of desupersaturation profile and polymorphic form in flufenamic acid systems. Ind. Eng. Chem. Res. 44, 1233–1240 (2005).
Bonnett, P. E., Carpenter, K. J., Dawson, S. & Davey, R. J. Solution crystallization via a submerged liquid-liquid phase boundary: oiling out. Chem. Commun. 2003, 698–699, (2003).
Chakrabarti, R. G. & Vekilov, P. G. Attraction between permanent dipoles and London dispersion forces dominate the thermodynamics of organic crystallization. Cryst. Growth Des. 20, 7429–7438 (2020).
De Yoreo, J. J. & Vekilov, P. G. Principles of crystal nucleation and growth. Rev. Mineral. Geochem. 54, 57–93 (2003).
Samuel, R. E., Salmon, E. D. & Briehl, R. W. Nucleation and growth of fibres and gel formation in sickle cell haemoglobin. Nature 345, 833–835 (1990).
Zhou, J. et al. Observing crystal nucleation in four dimensions using atomic electron tomography. Nature 570, 500–503 (2019).
Tang, W. et al. Tautomerism unveils a self-inhibition mechanism of crystallization. Nat. Commun. 14, 561 (2023).
Van Driessche, A. E. S. et al. Molecular nucleation mechanisms and control strategies for crystal polymorph selection. Nature 556, 89–94 (2018).
Zhu, G. et al. Self-similar mesocrystals form via interface-driven nucleation and assembly. Nature 590, 416–422 (2021).
Aizenberg, J., Black, A. J. & Whitesides, G. M. Control of crystal nucleation by patterned self-assembled monolayers. Nature, 398, 495–498, (1999).
Weiner, S. & Dove, P. M. An overview of biomineralization processes and the problem of the vital effect. Rev. Mineral. Geochem. 54, 1–29 (2003).
Aizenberg, J., Tkachenko, A., Weiner, S., Addadi, L. & Hendler, G. Calcitic microlenses as part of the photoreceptor system in brittlestars. Nature 412, 819–822 (2001).
Mosquera-Giraldo, L. I. et al. Crystallization inhibition properties of cellulose esters and ethers for a group of chemically diverse drugs: experimental and computational insight. Biomacromolecules 19, 4593–4606 (2018).
Mallette, A. J., Seo, S. & Rimer, J. D. Synthesis strategies and design principles for nanosized and hierarchical zeolites. Nat. Synth. 1, 521–534 (2022).
Gibbs, J. W. On the equilibrium of heterogeneous substances, First Part. Trans. Connect. Acad. Sci. 3, 108–248 (1876).
Gibbs, J. W. On the equilibrium of heterogeneous substances (concluded). Trans. Connect. Acad. Sci. 3, 343–524 (1878).
Volmer, M. Kinetik der Phasenbildung (Steinkopff, 1939).
Kashchiev, D. Nucleation. Basic theory with applications. (Butterworth, 2000).
Kashchiev, D. Analysis of experimental data for the nucleation rate of water droplets. J. Chem. Phys. 125, 044505 (2006).
Ma, W., Lutsko, J. F., Rimer, J. D. & Vekilov, P. G. Antagonistic cooperativity between crystal growth modifiers. Nature 577, 497–501 (2020).
Olafson, K. N., Ketchum, M. A., Rimer, J. D. & Vekilov, P. G. Mechanisms of hematin crystallization and inhibition by the antimalarial drug chloroquine. Proc. Natl Acad. Sci. 112, 4946–4951 (2015).
Zel’dovich, Y. B. Theory of new phase formation: Cavitation. Acta Physicochimica URSS 18, 1–22 (1943).
Hoffmann, J. et al. The unexpected dominance of secondary over primary nucleation. Faraday Discuss. 235, 109–131 (2022).
Rosenfeld, G., Servaty, R., Teichert, C., Poelsema, B. & Comsa, G. Layer-by-layer growth of Ag on Ag(111) induced by enhanced nucleation: a model study for surfactant-mediated growth. Phys. Rev. Lett. 71, 895–898 (1993).
De Yoreo, J. J. & Legg, B. A. What atoms do when they get together. Nat. Chem. 12, 883–885 (2020).
Vekilov, P. G. Nonclassical Nucleation. in Crystallization via Nonclassical Pathways Volume 1: Nucleation, Assembly, Observation & Application Vol. 1358 ACS Symposium Series Ch. 2, 19–46 (American Chemical Society, 2020).
Kaissaratos, M., Filobelo, L. & Vekilov, P. G. Two-step crystal nucleation is selected because of a lower surface free energy barrier. Cryst. Growth Des. 21, 5394–5402 (2021).
Vorontsova, M. A., Chan, H. Y., Lubchenko, V. & Vekilov, P. G. Lack of dependence of the sizes of the mesoscopic protein clusters on electrostatics. Biophys. J. 109, 1959–1968 (2015).
Safari, M. S., Vorontsova, M. A., Poling-Skutvik, R., Vekilov, P. G. & Conrad, J. C. Differential dynamic microscopy of weakly scattering and polydisperse protein-rich clusters. Phys. Rev. E 92, 042712 (2015).
Chan, H. Y. & Lubchenko, V. A mechanism for reversible mesoscopic aggregation in liquid solutions. Nat. Commun. 10, 2381 (2019).
Safari, M. S. et al. Anomalous dense liquid condensates host the nucleation of tumor suppressor p53 Fibrils. iScience 12, 342–355 (2019).
Yang, D. S. et al. Mesoscopic protein-rich clusters host the nucleation of mutant p53 amyloid fibrils. Proc. Natl Acad. Sci. 118, e2015618118 (2021).
Yerragunta, M. et al. A dual growth mode unique for organic crystals relies on mesoscopic liquid precursors. Commun. Chem. 7, 190 (2024).
Warzecha, M., Safari, M. S., Florence, A. J. & Vekilov, P. G. Mesoscopic solute-rich clusters in olanzapine solutions. Cryst. Growth Des. 17, 6668–6676 (2017).
Warzecha, M., Florence, A. J. & Vekilov, P. G. The ambiguous functions of the precursors that enable nonclassical modes of olanzapine nucleation and growth. Crystals 11, 738 (2021).
Barua, H., Svärd, M., Rasmuson, Å. C., Hudson, S. P. & Cookman, J. Mesoscale clusters in the crystallisation of organic molecules. Angew. Chem. Int. Ed. 63, e202312100 (2024).
Olafson, K. N., Nguyen, T. Q., Rimer, J. D. & Vekilov, P. G. Antimalarials inhibit hematin crystallization by unique drug–surface site interactions. Proc. Natl Acad. Sci. 114, 7531–7536 (2017).
Ma, W. et al. A second mechanism employed by artemisinins to suppress Plasmodium falciparum hinges on inhibition of hematin crystallization. J. Biol. Chem. 296, 100123 (2021).
Ma, W. et al. Nonclassical mechanisms to irreversibly suppress β-hematin crystal growth. Commun. Biol. 6, 783 (2023).
Segatin, N. & Klofutar, C. Thermodynamics of the Solubility of Water in 1-Hexanol, 1-Octanol, 1-Decanol, and Cyclohexanol. Monatshefte fur Chem. 135, 241–248 (2004).
Ketchum, M. A., Olafson, K. N., Petrova, E. V., Rimer, J. D. & Vekilov, P. G. Hematin crystallization from aqueous and organic solvents. J. Chem. Phys. 139, 121911 (2013).
Ketchum, M. A., Lee, A. M., Vekilov, P. G. & Rimer, J. D. Biomimetic assay for hematin crystallization inhibitors: a new platform to screen antimalarial drugs. Cryst. Growth Des. 17, 197–206 (2017).
Petsev, D. N., Thomas, B. R., Yau, S.-T. & Vekilov, P. G. Interactions and aggregation of apoferritin molecules in solution: effects of added electrolytes. Biophys. J. 78, 2060–2069 (2000).
Safari, M. S., Poling-Skutvik, R., Vekilov, P. G. & Conrad, J. C. Differential dynamic microscopy of bidisperse colloidal suspensions. npj Microgravity 3, 21 (2017).
Pan, W., Vekilov, P. G. & Lubchenko, V. The origin of anomalous mesoscopic phases in protein solutions. J. Phys. Chem. B 114, 7620–7630 (2010).
Einstein, A. Zur Theorie der Brownschen Bewegung. Ann. der Phys. 19, 371–381 (1906).
De Yoreo, J. A perspective on multistep pathways of nucleation. In Crystallization via Nonclassical Pathways Volume 1: Nucleation, Assembly, Observation & Application Vol. 1358 ACS Symposium Series Ch. 1, 1-17 (American Chemical Society, 2020).
Hayat, M. A. Principles and techniques of electron microscopy: biological applications (Cambridge University Press, 2000).
Scarff, C. A., Fuller, M. J. G., Thompson, R. F. & Iadanza, M. G. Variations on negative stain electron microscopy methods: tools for tackling challenging systems. J. Vis. Exp. 6, 57199 (2018).
Bacher, A. et al. Electron microscopy of decorated crystals for the determination of the crystallographic rotation and translation parameters in large protein complexes. J. Mol. Biol. 225, 1065–1073 (1992).
Houben, L., Weissman, H., Wolf, S. G. & Rybtchinski, B. A mechanism of ferritin crystallization revealed by cryo-STEM tomography. Nature 579, 540–543 (2020).
Yamazaki, T. et al. Two types of amorphous protein particles facilitate crystal nucleation. Proc. Natl Acad. Sci. 114, 2154–2159 (2017).
Tsarfati, Y. et al. Crystallization of organic molecules: nonclassical mechanism revealed by direct imaging. ACS Cent. Sci. 4, 1031–1036 (2018).
Harano, K. et al. Heterogeneous nucleation of organic crystals mediated by single-molecule templates. Nat. Mater. 11, 877–881 (2012).
Yuwono, V. M., Burrows, N. D., Soltis, J. A. & Penn, R. L. Oriented aggregation: formation and transformation of mesocrystal intermediates revealed. J. Am. Chem. Soc. 132, 2163–2165 (2010).
Maes, D. et al. Do protein crystals nucleate within dense liquid clusters? Acta Crystallogr. Sect. F. 71, 815–822 (2015).
Li, Y., Lubchenko, V. & Vekilov, P. G. The use of dynamic light scattering and brownian microscopy to characterize protein aggregation. Rev. Sci. Instrum. 82, 053106 (2011).
Filipe, V., Hawe, A. & Jiskoot, W. Critical evaluation of nanoparticle tracking analysis (NTA) by NanoSight for the measurement of nanoparticles and protein aggregates. Pharm. Res. 27, 796–810 (2010).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Feric, M. et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell 165, 1686–1697 (2016).
Chaikin, P. M. & Lubensky, T. C. Principles of condensed matter physics (Cambridge Unversity Press, 1995).
Debenedetti, P. G. Metastable liquids (Princeton Univ Pr, 1996).
Chan, Ho. Y., Lankevich, V., Vekilov, PeterG. & Lubchenko, V. Anisotropy of the coulomb interaction between folded proteins: consequences for mesoscopic aggregation of lysozyme. Biophys. J. 102, 1934–1943 (2012).
Safari, M. S., Byington, M. C., Conrad, J. C. & Vekilov, P. G. Polymorphism of lysozyme condensates. J. Phys. Chem. B 121, 9091–9101 (2017).
Li, Y., Lubchenko, V., Vorontsova, M. A., Filobelo, L. & Vekilov, P. G. Ostwald-like ripening of the anomalous mesoscopic clusters in protein solutions. J. Phys. Chem. B 116, 10657–10664 (2012).
Verma, L. et al. How to identify the crystal growth unit. Isr. J. Chem. 61, 1–11 (2021).
Warzecha, M. et al. Olanzapine crystal symmetry originates in preformed centrosymmetric solute dimers. Nat. Chem. 12, 914–920 (2020).
Verma, L., Vekilov, P. G. & Palmer, J. C. Solvent structure and dynamics near the surfaces of β-hematin crystals. J. Phys. Chem. B 125, 11264–11274 (2021).
Kapishnikov, S., Hempelmann, E., Elbaum, M., Als-Nielsen, J. & Leiserowitz, L. Malaria pigment crystals: the achilles’ heel of the malaria parasite. ChemMedChem 16, 1515–1532 (2021).
Zhou, H.-X., Rivas, G. & Minton, A. P. Macromolecular crowding and confinement: biochemical, biophysical, and potential physiological consequences*. Annu. Rev. Biophys. 37, 375–397 (2008).
Binder, K., Virnau, P. & Statt, A. Perspective: the Asakura Oosawa model: a colloid prototype for bulk and interfacial phase behavior. J. Chem. Phys. 141, 140901 (2014).
Gorka, A. P., de Dios, A. & Roepe, P. D. Quinoline drug–heme interactions and implications for antimalarial cytostatic versus cytocidal activities. J. Med. Chem. 56, 5231–5246 (2013).
De Jong, D. H. et al. Determining equilibrium constants for dimerization reactions from molecular dynamics simulations. J. Computational Chem. 32, 1919–1928 (2011).
Lutsko, J. F. Mechanism for the stabilization of protein clusters above the solubility curve: the role of non-ideal chemical reactions. J. Phys. Condens. Matter 28, 244020 (2016).
Martin, R. E. et al. Chloroquine transport via the malaria parasite’s chloroquine resistance transporter. Science 325, 1680–1682 (2009).
Provencher, S. W. CONTIN: a general purpose constrained regularization program for inverting noisy linear algebraic and integral equations. Comp. Phys. Commun. 27, 229–242 (1982).
Vielemeyer, O., McIntosh, M. T., Joiner, K. A. & Coppens, I. Neutral lipid synthesis and storage in the intraerythrocytic stages of Plasmodium falciparum. Mol. Biochem. Parasitol. 135, 197–209 (2004).
Noland, G. S., Briones, N. & Sullivan, D. J. Jr The shape and size of hemozoin crystals distinguishes diverse Plasmodium species. Mol. Biochem. Parasitol. 130, 91–99 (2003).
Vorontsova, M. A., Vekilov, P. G. & Maes, D. Characterization of the diffusive dynamics of particles with time-dependent asymmetric microscopy intensity profiles. Soft. Matter. 12, 6926–6936 (2016).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 25, 247–260 (2006).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Computational Chem. 25, 1157–1174 (2004).
Vanquelef, E. et al. R.E.D. Server: a web service for deriving RESP and ESP charges and building force field libraries for new molecules and molecular fragments. Nucleic Acids Res. 39, W511–517 (2011).
Sousa da Silva, A. W. & Vranken, W. F. ACPYPE - AnteChamber PYthon Parser interfacE. BMC Res Notes 5, 367 (2012).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. J. Computational Chem. 18, 1463–1472 (1997).
Miyamoto, S. & Kollman, P. A. Settle: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Computational Chem. 13, 952–962 (1992).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Barducci, A., Bussi, G. & Parrinello, M. Well-tempered metadynamics: a smoothly converging and tunable free-energy method. Phys. Rev. Lett. 100, 020603 (2008).
Bonomi, M. et al. PLUMED: a portable plugin for free-energy calculations with molecular dynamics. Comput. Phys. Commun. 180, 1961–1972 (2009).
Tiwary, P. & Parrinello, M. A time-independent free energy estimator for metadynamics. J. Phys. Chem. B 119, 736–742 (2015).
Olafson, K. N., Rimer, J. D. & Vekilov, P. G. Growth of large hematin crystals in biomimetic solutions. Cryst. Growth Des. 14, 2123–2127 (2014).
Grossfield, A. et al. Best practices for quantification of uncertainty and sampling quality in molecular simulations [Article v1.0]. Living J. Comput. Mol. Sci. 1, 5067 (2018).
Acknowledgements
We thank Lynn Taylor and Anatoly Kolomeisky for thought-provoking discussions and Monika Warzecha for valuable suggestions on the manuscript. This work was supported by the National Institutes of Health (Award No. R01 AI150763), the National Science Foundation (Award No. DMR-2128121), and the Welch Foundation (Awards E-2170, E-1882, and E-1794, and the Welch Center for Advanced Bioactive Materials Crystallization, Award V-E-0001).
Author information
Authors and Affiliations
Contributions
P.G.V. conceived this work and designed experiments. W.M., W.P., and H.J.L. carried out light scattering measurements. W.M. performed OIM characterization. W.M. and M.B.S. carried out EM imaging. L.V. and J.C.P. carried out MD simulations. D.J.S. observed hemozoin crystals. P.G.V., J.C.P., J.D.R., and D.J.S. wrote the manuscript. All authors discussed the results and edited the paper. W.M. and L.V. contributed equally.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, W., Verma, L., Lee, HJ. et al. A nonclassical pathway of β-hematin crystal nucleation enables its suppression by antimalarials. Commun Chem 8, 246 (2025). https://doi.org/10.1038/s42004-025-01612-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-025-01612-0