Abstract
RegIIIα is an antibacterial protein primarily operating in the digestive tract to defend against bacterial infection through direct bactericidal activity. A previous study proposed that RegIIIα forms hexameric pores on the membrane of Gram-positive bacteria, leading to cell lysis. These RegIIIα hexamers can further assemble into filaments, diminishing RegIIIα activity. However, the high-resolution structure of RegIIIα assembly remains elusive, impeding the comprehension of the molecular mechanisms underlying RegIIIα function. In this study, we determined the cryo-electron microscopy (cryo-EM) structure of RegIIIα filaments formed in vitro at a resolution of 2.2 Å. Our structure reveals a similar subunit arrangement but a distinct subunit orientation compared to the previously reported low-resolution model of RegIIIα filaments. Through structural analysis and biochemical assays, we identified two essential interfaces for RegIIIα assembly, offered a potential explanation for the necessity of lipids in RegIIIα assembly, and elucidated the inhibitory mechanism of the pro-segment of RegIIIα. Collectively, our study presents the first near-atomic structure of filaments formed by C-tyle lectin containing proteins, providing structural insights into RegIIIα assembly that are closely related to its physiological functions and regulations.
Similar content being viewed by others
Introduction
RegIIIα, a C-type lectin domain (CTLD) containing protein1,2, belongs to the third subclass of the Reg family3,4,5 and functions as an antibacterial agent against gram-positive bacteria in the intestine6. Previous studies have revealed that RegIIIα recognizes its targets by binding to peptidoglycan and further disrupts bacterial cells6,7. The removal of the pro-segment (residues E27-R37) of RegIIIα through proteolytic cleavage is required to initiate its bactericidal activity8,9. A subsequent study proposed a working model underlying antibacterial activity; RegIIIα forms hexameric pores to permeabilize the bacterial membrane and induce cell death8. After long-term incubation with lipids, these hexamers can further stack into filaments resembling those found in pancreatic secretions10,11, which is considered a self-preservation strategy as filament formation reduces the cytotoxicity of RegIIIα8.
The previous cryo-EM study of the RegIIIα filament resulted in a 9-Å map, and a filament model was proposed by docking the crystal structure of the RegIIIα monomer into this map8. However, the suboptimal resolution of this map has impeded the exploration of detailed subunit-subunit interactions within the filament, thereby leaving the molecular mechanism of RegIIIα hexamer assembly and membrane permeabilization unresolved. In this study, we determined the cryo-EM structure of RegIIIα filaments at a resolution of 2.2 Å. This structure differs from the previous docking model in terms of subunit orientation. Structural analysis revealed the interfaces responsible for RegIIIα filament formation and ligand binding, as well as the inhibitory mechanism of the pro-segment. Our study presents, to our knowledge, the first near-atomic-resolution structure of filaments formed by C-tyle lectin containing proteins, offering crucial insights for exploring the physiological functions of RegIIIα.
Results
Filament preparation and cryo-EM structure determination
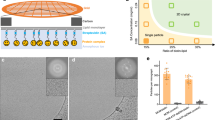
Purified RegIIIα was incubated with or without trypsin at either acidic or neutral pH levels. The SDS-PAGE assays indicated that the molecular weight of RegIIIα incubated with trypsin was smaller than that of RegIIIα without trypsin (Fig. 1a and Supplementary Fig. 1), suggesting the removal of the pro-domain through proteolytic processing6,9. Negative stain electron microscope (EM) analysis of the incubated samples revealed filament formation of RegIIIα after trypsin cleavage at a pH level of 7.4 (Fig. 1b). No filaments were observed in samples without trypsin cleavage, while only a few filaments were observed in samples after trypsin cleavage at pH 5.5 (Fig. 1b). Cryo-EM data was collected for filaments grown in pH 7.4, and two-dimensional (2D) classification of the cryo-EM dataset revealed a single filament species (Supplementary Fig. 2). Cryo-EM structure was determined for this filament species with a resolution of 2.2 Å (Fig. 2a, Supplementary Fig. 3, Supplementary Fig. 4a). An atomic model was built using a previously reported crystal structure of RegIIIα (PDB 4MTH)8 as the initial model. Detailed data collection and processing statistics are summarized in Table 1.
a SDS-PAGE of recombinant RegIIIα with or without the treatment of trypsin. The uncropped gel was displayed in Fig. S1. Representative negative stain EM images of (b) wild type RegIIIα with or without the treatment of trypsin and incubated in indicated pH levels, (c) RegIIIα mutants with trypsin treatment and incubated in pH 7.4, or (d) RegIIIα H50A with trypsin treatment and incubated in indicated pH levels.
a cryo-EM map of RegIIIα filaments, with six subunits being colored. IF, interface. b The helical net of RegIIIα filaments. c Structural alignment of one subunit in the RegIIIα filament (blue) with the pre-reported crystal structure (white, PDB 4MTH). d Sequence of RegIIIα with secondary structures annotated upper the corresponding region. e Comparison of the RegIIIα filament structure reported here with the low-resolution map previously reported (EMD-5795). Black arrows indicate the handedness of the 6-start helix.
Cryo-EM structure of RegIIIα filament
Within the filament, all RegIIIα subunits are symmetrically related to each other with a 1-start helical symmetry and a C3 symmetry (Fig. 2a, b). Specifically, three identical RegIIIα subunits form a homotrimer with a C3 symmetry, and multiple homotrimers stack on top of each other with a 1-start helical symmetry to construct the filament.
All RegIIIα subunits within the filament exhibit an identical structure to each other and to the previously reported crystal structure of the RegIIIα monomer (PDB ID 4MTH8, Fig. 2c). Each RegIIIα subunit contains the entire C-type lectin domain (CTLD, residues 38-175, Fig. 2d). The N- and C-termini of each RegIIIα subunit are positioned at the center of the filament, leaving no room for flexible residues beyond the visible core region (Supplementary Fig. 4b). This observation indicates that the pro-segment must be cleaved before filament formation, suggesting a molecular mechanism involving the proteolytic activation of RegIIIα (see Discussion).
Comparison with the previously reported low-resolution model
Comparing our map with the previously reported low-resolution map of RegIIIα filaments8, we observed that these two maps closely resemble each other in subunit arrangements, albeit with reversed handedness (Fig. 2e). Specifically, the 6-start helix in the cryo-EM map determined here is left-handed (Fig. 2a), whereas the corresponding helix in the low-resolution map is right-handed (Fig. 2e). By employing rigid-body docking of the filament structure into the handedness-reversed low-resolution map, we ascertained that the model aligned well with the map, positioning all subunits of the model within in the blobs of the map (Fig. 2e). As information regarding handedness is absent in transmission electron microscopy, determining the handedness of the maps at low resolutions poses a challenge, unless the resolution allows for the observation of the handedness of an α-helix. Based on these observations, it is reasonable to infer that the RegIIIα filaments adopt a similar structure in this study and the previous study8.
Further comparison between the previous RegIIIα filament model, proposed by docking the crystal structure of the RegIIIα monomer into the low-resolution map of the RegIIIα filament8, and the filament structure determined here revealed that they differ from each other in subunit orientations. Specifically, in the previous model, the N- and C- termini of RegIIIα are facing outward from the filament center, and the loop 2 is facing inward, contrasting with the orientation of our structure (Supplementary Fig. 4b). These observations indicate the need to re-evaluate the proposed antibacterial mechanism of RegIIIα in light of the revised hexamer model (see discussion).
Subunit-subunit interfaces within the filament
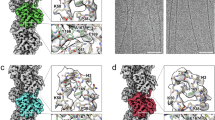
Based on the 1-start helical symmetry and C3 symmetry described above, all inter-subunit interfaces within the RegIIIα filament can be categorized into two types, designated as IF1 (interface 1) and IF2 (Fig. 2a). Three sets of IF1 and IF2 are responsible for stacking one homo-trimer of RegIIIα to another homo-trimer (Fig. 2a). The interactions in IF1 are established by a long-distance salt-bridge between Asp63 and Lys93, hydrogen bonds between the side chain of Gln69 and the main chain of Ala46, and the insertion of Leu66 into the hydrophobic pocket formed by Phe87, Leu91, and the Cβ-Cε of Lys45 (Fig. 3a, left panel). On the other hand, the interactions in IF2 entail a salt-bridge between Glu114 and Lsy70, a salt-bridge between Glu118 and Lsy59, hydrogen bonds between the side chain of Asn129 and the main chains of Leu54 and Leu56, hydrogen bonds between the side chain of Arg71 and the main chains of Ser125 and Asp126, and the insertion of Val127 into hydrophobic pocket formed by Gly43, Ala53, and Phe55 (Fig. 3a, right panel). In addition to these direct subunit-subunit interactions, additional densities are observed in both IF1 and IF2, potentially indicating unidentified ligand binding could further facilitate subunit-subunit interactions (Fig. 3b, described in another subsection below).
a Subunit-subunit interactions in IF1 (left) and IF2 (right). Mainchains and sidechains involved in interactions are displayed in sticks, labeled, and colored in blue (subunit n), cyan (subunit n + 1), and magenta (subunit n”+1). Residues involved in interactions with only mainchains are labeled with asterisks. b Atomic model of 6 neighboring RegIIIα subunits, with extra densities at subunit-subunit interfaces displayed in orange envelops. Detailed illustrations of the extra densities at IF1 (c) and IF2 (d), with nearby residues displayed. e Locations of Lsy42 and His50 at the subunit-subunit interface. The side chains of Lys42 and His50 are displayed in spheres.
Mutagenesis study confirmed key residues for RegIIIα filament assembly
In order to explore the contribution of IF1 and IF2 in RegIIIα filament assembly, two mutations, L66W and Q69R, were designed to disrupt IF1, respectively, and one mutation, V127Y, was designed to disrupt IF2. Specifically, both L66W and V127Y disrupt the interfaces through steric hinderance with the hydrophobic pockets on the opposing subunit, whereas Q69R generates static repulsion with Lys45 from the other subunit (Fig. 3a). When RegIIIα mutants were incubated with trypsin at pH 7.4, no filaments were observed under EM (Fig. 1c), suggesting that disrupting either interface of IF1 and IF2 is sufficient to abolish the filament formation of RegIIIα.
Moreover, we found His50 in each RegIIIα subunit is adjacent to Lsy42 from another subunit, situated at the interface of three subunits, in a crowded environment where there are no negatively charged amino acids to neutralize the positive charges of these two amino acids (Fig. 3e). These observations may explain the pH preference in RegIIIα filament formation. Specifically, it is plausible that the pKa of His50 is between 5.5 and 7.4. Therefore, at pH 5.5, His50 is protonated and positive charged, causing static repulsion with Lys42 and inhibiting filament assembly. In contrast, at pH 7.4, His50 is deprotonated and without a positive charge, which should accommodate the positive charge of Lys42 and allow filament formation. To validate this hypothesis, we incubated RegIIIα H50A with trypsin at pH 5.5 and 7.4. Filaments with similar amount were observed at both pH levels, suggesting that His50 is responsible for the pH preference of RegIIIα filament formation.
Additional densities in RegIIIα filaments suggest ligand binding
Additional densities observed in both IF1 and IF2 are likely a result of ligand binding (Fig. 3b–d). These additional densities are encompassed by multiple residues with diverse properties, including negatively charged residues (i.e., Glu118 and Glu121), positively charged residues (i.e., Lys70 and Arg71), hydrogen bond donors and acceptors (i.e., Asn136 and Tyr169), and hydrophobic residues (i.e., Ala67 and Phe173) (Fig. 3d). Collectively, these residues create suitable binding sites for ligands, enabling high selectivity and stability in ligand binding. The potential interactions between the ligand and subunits participating the interfaces could enhance the subunit-subunit interface, thereby stabilizing the assembly of the RegIIIα filament. Identifying these ligands should aid future studies of the molecular mechanism of RegIIIα assembly. The shape of these additional densities suggests that these ligands are likely to possess an elongated molecular conformation (Fig. 3d). A plausible assumption is that these ligands could be lipids, as a previous study suggested that the addition of phospholipids can promote RegIIIα filament formation in vitro8. However, the suboptimal quality of these additional densities hinders the definitive identification of the ligands in this study.
Discussion
In this study, we prepared RegIIIα filaments with recombinant proteins and determined their cryo-EM structure at a resolution of 2.2 Å. Our structure revises a previously proposed filament model in term of subunit orientation and provides essential molecular details in understanding the mechanism of RegIIIα assembly and activity.
Our structural analysis revealed two subunit-subunit interfaces, IF1 and IF2, that are essential for the assembly of RegIIIα. Disrupting either interface through site mutation abolished RegIIIα filament formation (Fig. 1c). However, structural analysis suggested that the primary components of these interfaces are not direct subunit-subunit interactions but rather interactions potentially mediated by ligand binding, as indicated by additional densities in the cry-EM map (Fig. 3a–d). Therefore, these ligands, exhibiting shapes reminiscent of lipids, might play a crucial role in the assembly of RegIIIα. This observation is consistent with previous findings indicating that RegIIIα filament formation is lipid-dependent8. We hypothesize that lipids could initiate RegIIIα assembly by directly mediating subunit-subunit interactions. Notably, no lipids were intentionally introduced during the preparation of RegIIIα filaments used for cryo-EM structure determination; the presence of potential lipids may have originated from the membranes of E. coli cells utilized for expressing RegIIIα proteins. Further studies are required to remove these lipids from RegIIIα and incubate lipid-free RegIIIα with various types of lipids to investigate the lipid specificity in triggering RegIIIα filament formation.
In the filament structure presented here, six adjacent RegIIIα subunits form a hexamer reminiscent of that observed in the low-resolution map8 (Fig. 2e). This hexamer has been postulated as the functional state of RegIIIα, given its shape and size, which are suitable for creating pores in the bacterial membrane to induce cell leakage. The docking model suggested that the outer surface of the hexamer contains positively charged residues capable of interacting with the negatively charged phospholipids in the lipid bilayer8. Although our high-resolution structure indicates a reversed subunit orientation in the hexamer compared to the previous model, the updated model maintains consistent shape and size, with positively charged residues such as Arg135 and Lys159 present on the outer surface of the hexamer. These findings suggest that the hypothesis of the hexamer serving as a pore-forming structure remains relevant. However, given that the interfaces necessary for hexamer assembly are the same as those required for filament assembly, it remains puzzling how RegIIIα determines its fate between these two assembly types. This puzzle is further complicated by the fact that the hexamer is proposed to represent a functional antibacterial state, while the filament is suggested to be a protective inhibitory state, as filament formation was observed to partially inhibit RegIIIα activity8.
The pro-segment is known to inhibit RegIIIα activity12 and previous hypothesis suggested that this inhibition occurs by the pro-segment blocking the interaction of the N-terminus of RegIIIα with lipids8. However, our high-resolution structure has revealed a different orientation of the N-terminus of RegIIIα towards the center rather than the outer surface of the hexamer. This finding indicates that the pro-segment does not inhibit RegIIIα activity by blocking interactions between RegIIIα and lipids. Instead, it is likely that the pro-segment hinders RegIIIα activity by impeding hexamer formation, as the space in the center of the hexamer is inadequate to accommodate the uncleaved pro-segment of six subunits (Supplementary Fig. 4b). It is worth noting that we cannot fully exclude the possibility that the previously reported hexameric model of RegIIIα indeed exhibits the opposite subunit orientation relative to the structure presented here, since the filaments were generated under different conditions in the previous study8.
In this study, we observed a pH preference in RegIIIα filament formation, with enhanced filament formation occurring at a pH level of 7.4 but not at 5.5 (Fig. 1b). Through our structural analysis and mutagenesis study, we have identified the residue that responsible for this pH preference as His50 of RegIIIα. Since this residue is also involved in hexamer assembly, it is reasonable to infer that the activity of RegIIIα is also influenced by pH through the same mechanism. Further studies can be conducted to investigate the physiological significance of this pH preference. The pH-independent mutant, H50A, identified in this study, can serve as a valuable tool for these studies.
Given that RegIIIα can easily form filament under physiological conditions in vitro, it is plausible that RegIIIα filaments may also be present in vivo. However, it is challenging to validate this hypothesis through extracting filament from tissues like other studies13,14,15,16 because RegIIIα monomer can undergo filament formation within 2 h of incubation after the removal of the pro-segment in vitro (Fig. 1b). Therefore, if pro-segment-free RegIIIα is detected in tissues and RegIIIα filaments are extracted from these tissues, it is difficult to differentiate whether these filaments were formed within the tissues or during the extraction processes. Specifically, the mostly used filament extraction protocols usually include long-time incubation before the sedimentation of filament, such as a 2-h sarkosyl treatment13 or an overnight collagenase digestion14. These processes will give adequate time for RegIIIα to undergo filamentation. In situ methods, such as immunostaining of the tissues with filament-specific antibodies, or cryo-electron tomography (cryo-ET) with a high enough resolution to distinguish RegIIIα filament from other types of filaments, are required to confirm the presence of RegIIIα filament in vivo.
To further investigate the physiological roles of the RegIIIα assembly, we mapped all pathogenic mutations recorded in the UniPort database17 onto our filament structure (Supplementary Fig. 5). We found that several mutations are localized to subunit-subunit interfaces (i.e., K45N, A46D/T, G48V, S90F, A83P, Q69R on IF1, and V127L on IF2) and could therefore impair filament formation. We experimentally confirmed the disruptive effects of Val127 and Gln69 substitutions (Fig. 1c). Moreover, all His50-associated mutations are predicted to be pathogenic or likely pathogenic, which is likely related to His50-mediated filament formation in responding to pH change. Collectively, our analysis suggested that these mutations may compromise RegIIIα function by blocking filament assembly, further supporting the biological significance of RegIIIα polymerization.
In summary, in this study, we prepared RegIIIα filaments with recombinant RegIIIα proteins and determined the first high-resolution cryo-EM structure of these filaments. Our findings provide structural insights into RegIIIα assembly and bring new perspectives to the mechanistic understanding of RegIIIα.
Methods
Expression, purification of RegIIIα and filament preparation
The gene encoding RegIIIα (E27-D175) was cloned from cDNA library and ligated into pET28a expression vector. All RegIIIα mutants were generated by overlapping PCR using wild type as a template. The resulting plasmids were transformed into E. coli BL-21 (DE3) and cultured at 37 °C overnight. The culture was inoculated into 1 L fresh LB medium and cultured to optical density of 0.8 (OD600). The expression of RegIIIα was induced by IPTG at the final concentration of 1 mM and expressed as inclusion bodies for 4 h at 37 °C. RegIIIα and mutants were purified based on the denaturing protocol as previously reported with minor modifications18,19. Briefly, bacterial cells were harvested by centrifugation, resuspended in 50 ml lysis buffer (20 mM Tris-HCl pH 7.5, 10 μM EDTA, 1% Triton X-100) and lysed by sonication. The inclusion-body-containing debris was obtained by centrifugation at 12,000 g for 30 min, after which the pellet was washed 2 times with washing buffer (20 mM Tris-HCl pH 7.5, 10 μM EDTA, 1% Triton X-100, 500 mM NaCl). The resulting inclusion body was resuspended in 20 ml resuspension buffer (100 mM Tris-HCl pH 7.5, 7 M Guanidine-HCl, 150 mM Reduced Glutathione, 2 mM EDTA) and stirred overnight to thoroughly disperse it. Then, the solution was clarified by centrifugation, 5-ml supernatant was drop-wise titrated into 200-ml refolding buffer (50 mM Tris–HCl, pH 8.0, l0 mM KCl, 240 mM NaCl, 2 mM MgCl2, 2 mM CaCl2, 0.5 M guanidine–HCl, 0.4 M sucrose, 0.5 M arginine–HCl, 1 mM reduced glutathione, and 0.1 mM oxidized glutathione) and stand at room temperature 24 h. After that, the refolding buffer was dialyzed against 10-folds of dialysis buffer 1 (25 mM Tris-HCl pH 7.5, 2 mM CaCl2, 25 mM NaCl) twice and dialysis buffer 2 (25 mM MES pH 6.0, 2 mM CaCl2, 25 mM NaCl), sequentially. RegIIIα proteins from dialyzed supernatant was purified using SPFF column and size exclusion chromatography. The elute buffer for SPFF purification is 25 mM MES pH 6.0, 2 mM CaCl2, 300 mM NaCl, and the running buffer for size-exclusion chromatography is 25 mM Tris-HCl pH 7.5, 25 mM NaCl. The purified RegIIIα was concentrated and stored at −80 °C for further use.
For filament preparation, RegIIIα protein was diluted with stock buffer (20 mM MES pH 5.5/20 mM Tris-HCl pH 5.5 or 7.4, 25 mM NaCl) to the final concentration of 10 μM, followed by the addition of trypsin at the molar ratio of 20:1 (RegIIIα : trypsin) and incubation at 37 °C for 2 h.
SDS-PAGE assays
Samples were mixed with 25% (v/v) SDS loading buffer, boiled at 100 °C for 10 min, and separated with YoungPAGE 4–20% Bis-Tris precast gels (GenScript, catalog number M00928). The protein was visualized by Coomassie blue staining.
Transmission electron microscopy (TEM)
2.6 μl sample was applied onto 200 mesh glow-discharged carbon-coated Cu grids and sedimented for 2 min. The excess solution was blotted off by filter papers, grids were stained using 3.3 μl 2% uranyl acetate for 1 min and further washed with 3.3 μl 2% uranyl acetate, followed by air-dried for 2 min. The grids were imaged using Talos L120C G2 (Thermofisher, USA).
Cryo-EM data collection and processing
For cryo-EM sample preparation, 2.6 μl of the sample was applied onto a freshly glow discharged Quantifoil 1.2/1.3 200 mesh grid (catalog number N1-C14nCu20-01). The grid was plunge-frozen in liquid ethane using a Vitrobot Mark IV instrument (FEI). Cryo-EM data were collected on a 300-kV Titan Krios transmission election microscope (Thermo Fisher Scientific) equipped with a Falcon 4i detector with a total accumulated dose of 40 e-/Å2 per image, a nominal physical pixel size of 0.932 Å/pixel, and an energy filter of 20 eV. A total of 2044 micrographs were collected for data processing.
Cryo-EM data processing was performed using cryoSPARC v4.0.120. Micrographs were processed with motion correction and CTF estimation in cryoSPARC, and particles were automatically picked with filament tracer. Particle extraction was conducted with a box size of 360 pixels, and 2D classification was performed with the extracted particles. The resulting 2D classes all belonged to a single filament species, except for those represent junk particles. Particles from the optimal 2D classes were selected for helical refinement without helical parameter and initial reference provided. The initial helical parameters were determined as a helical rise of 40.9 Å and a helical twist of 11.52° through automatic helical parameter search using symmetry search utility in cryoSPARC. This set of helical parameters corresponds to that between subunit n and n”+2 along the 6-start helices in the final reconstruction. The second round of helical refinement was performed with this set of helical parameters and the map of the previous helical refinement as a reference, resulting in a map with a resolution of 3.0 Å. Manual inspection revealed a C3 symmetry of this map, and the handedness of this map needed to be flipped according to the handedness of the α-helix in this map. Automatic symmetry search with the handedness-flipped map revealed the updated helical parameters to be a helical rise of 20.5 Å and a helical twist of 54.24°. The final helical refinement was conducted with local CTF-refined particles, the updated helical parameters, and the handedness-flipped map as a reference, resulting in the final map with a resolution of 2.2 Å. The resolutions were all estimated using the 0.143 Fourier shell correlation (FSC) resolution cut-off. Data collection and processing statistics are summarized in Table 1.
Atomic model building
The crystal structure of RegIIIα (PDB 4MTH)8 was manually docked into the final map as a rigid body and adjusted to fit the map using COOT21. The initial model was expanded to contain 12 identical subunits and refined with phenix.real_space_refine22. The final model was validated with MolProbity 23.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM map and atomic model of RegIIIα fibrils have been deposited into the Worldwide Protein Data Bank (wwPDB) and the Electron Microscopy Data Band (EMDB) with accession codes PDB 9WTD and EMD-66219. Any other relevant data is available from the corresponding author upon reasonable request.
References
Wang, L. et al. The regenerating protein 3A: a crucial molecular with dual roles in cancer. Mol. Biol. Rep. 49, 1491–1500 (2022).
Lasserre, C. et al. Structural organization and chromosomal localization of a human gene (HIP/PAP) encoding a C-type lectin overexpressed in primary liver cancer. Eur. J. Biochem 224, 29–38 (1994).
Terazono, K. et al. A novel gene activated in regenerating islets. J. Biol. Chem. 263, 2111–2114 (1988).
Laurine, E. et al. PAP IB, a new member of the Reg gene family: cloning, expression, structural properties, and evolution by gene duplication. Biochim. Biophys. Acta 1727, 177–187 (2005).
Narushima, Y. et al. Structure, chromosomal localization and expression of mouse genes encoding type III Reg, RegIII alpha, RegIII beta, RegIII gamma. Gene 185, 159–168 (1997).
Cash, H. L., Whitham, C. V., Behrendt, C. L. & Hooper, L. V. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 313, 1126–1130 (2006).
Lehotzky, R. E. et al. Molecular basis for peptidoglycan recognition by a bactericidal lectin. Proc. Natl. Acad. Sci. USA 107, 7722–7727 (2010).
Mukherjee, S. et al. Antibacterial membrane attack by a pore-forming intestinal C-type lectin. Nature 505, 103–107 (2014).
Mukherjee, S. et al. Regulation of C-type lectin antimicrobial activity by a flexible N-terminal prosegment. J. Biol. Chem. 284, 4881–4888 (2009).
Ho, M. R. et al. Human pancreatitis-associated protein forms fibrillar aggregates with a native-like conformation. J. Biol. Chem. 281, 33566–33576 (2006).
Gross, J. et al. Isolation, characterization, and distribution of an unusual pancreatic human secretory protein. J. Clin. Invest. 76, 2115–2126 (1985).
Medveczky, P., Szmola, R. & Sahin-Tóth, M. Proteolytic activation of human pancreatitis-associated protein is required for peptidoglycan binding and bacterial aggregation. Biochem. J 420, 335–343 (2009).
Han, J. et al. Structure of human glycoprotein 2 reveals mechanisms underlying filament formation and adaption to proteolytic environment in the digestive tract. PLOS Biol. 23, e3003238 (2025).
Jiang, Y. X. et al. Amyloid fibrils in FTLD-TDP are composed of TMEM106B and not TDP-43. Nature 605, 304–309 (2022).
Schweighauser, M. et al. Structures of α-synuclein filaments from multiple system atrophy. Nature 585, 464–469 (2020).
Schweighauser, M. et al. Age-dependent formation of TMEM106B amyloid filaments in human brains. Nature 605, 310–314 (2022).
The UniProt, C UniProt: the Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 53, D609–D617 (2025).
Hill, J. H. et al. BefA, a microbiota-secreted membrane disrupter, disseminates to the pancreas and increases β cell mass. Cell Metab. 34, 1779–1791.e1779 (2022).
Cash, H. L., Whitham, C. V. & Hooper, L. V. Refolding, purification, and characterization of human and murine RegIII proteins expressed in Escherichia coli. Protein Expr. Purif. 48, 151–159 (2006).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Acknowledgements
This work was supported by the National Natural Science Foundation (NSF) of China (grant nos. 32271276 and 32571420) to Q.C., STI2030-Major Projects 2022ZD0212500 to Q.C, and the Yongxin Young Scholar Program to Q.C. The authors thank for cryo-EM data collection at the Instrument Analysis Center (IAC), Shanghai Jiao Tong University. The authors acknowledge the National Facility for Translational Medicine (Shanghai) for support.
Author information
Authors and Affiliations
Contributions
J.H. performed protein purification and filament preparation. J.H. prepared cryo-EM grids and collected cryo-EM data. J.H. and Q.C. processed the data and determined the structure of RegIIIα fibrils. All authors analyzed the results and wrote the manuscript. Q.C. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Wenjuan Zhang and Lora V. Hooper for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, J., Cao, Q. Structural basis for human RegⅢα filament formation. Commun Chem 9, 117 (2026). https://doi.org/10.1038/s42004-026-01921-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-026-01921-y