Abstract
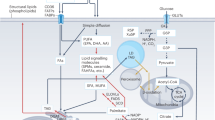
The perception that intracellular lipolysis is a straightforward process that releases fatty acids from fat stores in adipose tissue to generate energy has experienced major revisions over the last two decades. The discovery of new lipolytic enzymes and coregulators, the demonstration that lipophagy and lysosomal lipolysis contribute to the degradation of cellular lipid stores and the characterization of numerous factors and signalling pathways that regulate lipid hydrolysis on transcriptional and post-transcriptional levels have revolutionized our understanding of lipolysis. In this review, we focus on the mechanisms that facilitate intracellular fatty-acid mobilization, drawing on canonical and noncanonical enzymatic pathways. We summarize how intracellular lipolysis affects lipid-mediated signalling, metabolic regulation and energy homeostasis in multiple organs. Finally, we examine how these processes affect pathogenesis and how lipolysis may be targeted to potentially prevent or treat various diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Unger, R. H. & Scherer, P. E. Gluttony, sloth and the metabolic syndrome: a roadmap to lipotoxicity. Trends Endocrinol. Metab. 21, 345–352 (2010).
Walther, T. C., Chung, J. & Farese, R. V. Lipid droplet biogenesis. Annu. Rev. Cell Dev. Biol. 33, 491–510 (2017).
Whitehead, R. H. A note on the absorption of fat. Am. J. Physiol. Content 24, 294–296 (1909).
Vaughan, M., Berger, J. E. & Steinberg, D. Hormone-sensitive lipase and monoglyceride lipase activities in adipose tissue. J. Biol. Chem. 239, 401–409 (1964).
Osuga, J.-I. et al. Targeted disruption of hormone-sensitive lipase results in male sterility and adipocyte hypertrophy, but not in obesity. Proc. Natl Acad. Sci. USA 97, 787–792 (2000).
Haemmerle, G. et al. Hormone-sensitive lipase deficiency in mice causes diglyceride accumulation in adipose tissue, muscle, and testis. J. Biol. Chem. 277, 4806–4815 (2002).
Zimmermann, R. et al. Fat mobilization in adipose tissue is promoted by adipose triglyceride lipase. Science 306, 1383–1386 (2004).
Jenkins, C. M. et al. Identification, cloning, expression, and purification of three novel human calcium- independent phospholipase A2 family members possessing triacylglycerol lipase and acylglycerol transacylase activities. J. Biol. Chem. 279, 48968–48975 (2004).
Lass, A. et al. Adipose triglyceride lipase-mediated lipolysis of cellular fat stores is activated by CGI-58 and defective in Chanarin–Dorfman syndrome. Cell Metab. 3, 309–319 (2006).
Kienesberger, P. C., Oberer, M., Lass, A. & Zechner, R. Mammalian patatin domain containing proteins: a family with diverse lipolytic activities involved in multiple biological functions. J. Lipid Res. 50, 63–68 (2009).
Schweiger, M. et al. The C-terminal region of human adipose triglyceride lipase affects enzyme activity and lipid droplet binding. J. Biol. Chem. 283, 17211–17220 (2008).
Ahmadian, M. et al. Desnutrin/ATGL is regulated by AMPK and is required for a brown adipose phenotype. Cell Metab. 13, 739–748 (2011).
Pagnon, J. et al. Identification and functional characterization of protein kinase A phosphorylation sites in the major lipolytic protein, adipose triglyceride lipase. Endocrinology 153, 4278–4289 (2012).
Zhang, X. et al. An epistatic interaction between Pnpla2 and Lipe reveals new pathways of adipose tissue lipolysis. Cells 8, 395 (2019).
Brejchova, K. et al. Distinct roles of adipose triglyceride lipase and hormone-sensitive lipase in the catabolism of triacylglycerol estolides. Proc. Natl Acad. Sci. USA 118, e2020999118 (2021).
Ohno, Y., Kamiyama, N., Nakamichi, S. & Kihara, A. PNPLA1 is a transacylase essential for the generation of the skin barrier lipid ω-O-acylceramide. Nat. Commun. 8, 14610 (2017).
Soni, K. G. et al. Coatomer-dependent protein delivery to lipid droplets. J. Cell Sci. 122, 1834–1841 (2009).
Ellong, E. N. et al. Interaction between the triglyceride lipase ATGL and the Arf1 activator GBF1. PLoS ONE 6, e21889 (2011).
Wang, T. et al. OSBPL2 is required for the binding of COPB1 to ATGL and the regulation of lipid droplet lipolysis. iScience 23, 101252 (2020).
Cai, M. et al. FAM134B promotes adipogenesis by increasing vesicular activity in porcine and 3T3-L1 adipocytes. Biol. Chem. 400, 523–532 (2019).
Fischer, J. et al. The gene encoding adipose triglyceride lipase (PNPLA2) is mutated in neutral lipid storage disease with myopathy. Nat. Genet. 39, 28–30 (2007).
Missaglia, S., Coleman, R., Mordente, A. & Tavian, D. Neutral lipid storage diseases as cellular model to study lipid droplet function. Cells 8, 187 (2019).
Hirano, K.-I., Ikeda, Y., Zaima, N., Sakata, Y. & Matsumiya, G. Triglyceride deposit cardiomyovasculopathy. N. Engl. J. Med. 359, 2396–2398 (2008).
Haemmerle, G. et al. Defective lipolysis and altered energy metabolism in mice lacking adipose triglyceride lipase. Science 312, 734–737 (2006).
Haemmerle, G. et al. ATGL-mediated fat catabolism regulates cardiac mitochondrial function via PPAR-α and PGC-1. Nat. Med. 17, 1076–1085 (2011).
Attané, C. et al. A beta cell ATGL-lipolysis/adipose tissue axis controls energy homeostasis and body weight via insulin secretion in mice. Diabetologia 59, 2654–2663 (2016).
Heine, M. et al. Lipolysis triggers a systemic insulin response essential for efficient energy replenishment of activated brown adipose tissue in mice. Cell Metab. 28, 644–655.e4 (2018).
Ong, K. T., Mashek, M. T., Bu, S. Y., Greenberg, A. S. & Mashek, D. G. Adipose triglyceride lipase is a major hepatic lipase that regulates triacylglycerol turnover and fatty acid signaling and partitioning. Hepatology 53, 116–126 (2011).
Schreiber, R. et al. Hypophagia and metabolic adaptations in mice with defective ATGL-mediated lipolysis cause resistance to HFD-induced obesity. Proc. Natl Acad. Sci. USA 112, 13850–13855 (2015).
Mottillo, E. P., Bloch, A. E., Leff, T. & Granneman, J. G. Lipolytic products activate peroxisome proliferator-activated receptor (PPAR) α and δ in brown adipocytes to match fatty acid oxidation with supply. J. Biol. Chem. 287, 25038–25048 (2012).
Tang, T. et al. Desnutrin/ATGL activates PPARδ to promote mitochondrial function for insulin secretion in islet β cells. Cell Metab. 18, 883–895 (2013).
Khan, S. A. et al. ATGL-catalyzed lipolysis regulates SIRT1 to control PGC-1α/PPAR-α signaling. Diabetes 64, 418–426 (2015).
Najt, C. P. et al. Lipid droplet-derived monounsaturated fatty acids traffic via PLIN5 to allosterically activate SIRT1. Mol. Cell 77, 810–824.e8 (2020).
Hofer, P. et al. Fatty acid-binding proteins interact with comparative gene identification-58 linking lipolysis with lipid ligand shuttling. J. Biol. Chem. 290, 18438–18453 (2015).
Recazens, E., Mouisel, E. & Langin, D. Hormone-sensitive lipase: sixty years later. Prog. Lipid Res. 82, 101084 (2021).
Albert, J. S. et al. Null mutation in hormone-sensitive lipase gene and risk of type 2 diabetes. N. Engl. J. Med. 370, 2307–2315 (2014).
Farhan, S. M. K. et al. A novel LIPE nonsense mutation found using exome sequencing in siblings with late-onset familial partial lipodystrophy. Can. J. Cardiol. 30, 1649–1654 (2014).
Morigny, P. et al. Interaction between hormone-sensitive lipase and ChREBP in fat cells controls insulin sensitivity. Nat. Metab. 1, 133–146 (2019).
Zimmermann, R. et al. Decreased fatty acid esterification compensates for the reduced lipolytic activity in hormone-sensitive lipase-deficient white adipose tissue. J. Lipid Res. 44, 2089–2099 (2003).
Pajed, L. et al. Advanced lipodystrophy reverses fatty liver in mice lacking adipocyte hormone-sensitive lipase. Commun. Biol. 4, 323 (2021).
Hermo, L. et al. Alterations in the testis of hormone sensitive lipase-deficient mice is associated with decreased sperm counts, sperm motility, and fertility. Mol. Reprod. Dev. 75, 565–577 (2008).
Tornqvist, H. & Belfrage, P. Purification and some properties of a monoacylglycerol-hydrolyzing enzyme of rat adipose tissue. J. Biol. Chem. 251, 813–819 (1976).
Karlsson, M., Contreras, J. A., Hellman, U., Tornqvist, H. & Holm, C. cDNA cloning, tissue distribution, and identification of the catalytic triad of monoglyceride lipase. Evolutionary relationship to esterases, lysophospholipases, and haloperoxidases. J. Biol. Chem. 272, 27218–27223 (1997).
Savinainen, J. R. et al. Robust hydrolysis of prostaglandin glycerol esters by human monoacylglycerol lipase (MAGL). Mol. Pharmacol. 86, 522–535 (2014).
Heier, C. et al. Monoacylglycerol lipases act as evolutionarily conserved regulators of non-oxidative ethanol metabolism. J. Biol. Chem. 291, 11865–11875 (2016).
Grabner, G. F., Zimmermann, R., Schicho, R. & Taschler, U. Monoglyceride lipase as a drug target: at the crossroads of arachidonic acid metabolism and endocannabinoid signaling. Pharmacol. Ther. 175, 35–46 (2017).
Chon, S.-H., Zhou, Y. X., Dixon, J. L. & Storch, J. Intestinal monoacylglycerol metabolism: developmental and nutritional regulation of monoacylglycerol lipase and monoacylglycerol acyltransferase. J. Biol. Chem. 282, 33346–33357 (2007).
Rakhshandehroo, M. et al. Comprehensive analysis of PPARα-dependent regulation of hepatic lipid metabolism by expression profiling. PPAR Res. 2007, 26839 (2007).
Rajasekaran, D. et al. Staphylococcal nuclease and tudor domain containing 1 (SND1 protein) promotes hepatocarcinogenesis by inhibiting monoglyceride lipase (MGLL). J. Biol. Chem. 291, 10736–10746 (2016).
Thomas, G. et al. The serine hydrolase ABHD6 is a critical regulator of the metabolic syndrome. Cell Rep. 5, 508–520 (2013).
Pribasnig, M. A. et al. α/β hydrolase domain-containing 6 (ABHD6) degrades the late endosomal/lysosomal lipid bis(monoacylglycero)phosphate. J. Biol. Chem. 290, 29869–29881 (2015).
Grabner, G. F. et al. Metabolic disease and ABHD6 alter the circulating bis(monoacylglycerol)phosphate profile in mice and humans. J. Lipid Res. 60, 1020–1031 (2019).
Poursharifi, P., Madiraju, S. R. M. & Prentki, M. Monoacylglycerol signalling and ABHD6 in health and disease. Diabetes, Obes. Metab. 19, 76–89 (2017).
Tang, Z. et al. Enhanced monoacylglycerol lipolysis by ABHD6 promotes NSCLC pathogenesis. EBioMedicine 53, 102696 (2020).
Grüner, B. M. et al. An in vivo multiplexed small-molecule screening platform. Nat. Methods 13, 883–889 (2016).
Tardelli, M. et al. Lack of monoacylglycerol lipase prevents hepatic steatosis by favoring lipid storage in adipose tissue and intestinal malabsorption. J. Lipid Res. 60, 1284–1292 (2019).
Douglass, J. D. et al. Global deletion of monoacylglycerol lipase in mice delays lipid absorption and alters energy homeostasis and diet-induced obesity. J. Lipid Res. 56, 1153–1171 (2015).
Taschler, U. et al. Monoglyceride lipase deficiency in mice impairs lipolysis and attenuates diet-induced insulin resistance. J. Biol. Chem. 286, 17467–17477 (2011).
Chanda, P. K. et al. Monoacylglycerol lipase activity is a critical modulator of the tone and integrity of the endocannabinoid system. Mol. Pharmacol. 78, 996–1003 (2010).
Schlosburg, J. E. et al. Chronic monoacylglycerol lipase blockade causes functional antagonism of the endocannabinoid system. Nat. Neurosci. 13, 1113–1119 (2010).
Nomura, D. K. et al. Endocannabinoid hydrolysis generates brain prostaglandins that promote neuroinflammation. Science 334, 809–813 (2011).
Grabner, G. F. et al. Deletion of monoglyceride lipase in astrocytes attenuates lipopolysaccharide-induced neuroinflammation. J. Biol. Chem. 291, 913–923 (2016).
Nomura, D. K. et al. Monoacylglycerol lipase regulates a fatty acid network that promotes cancer pathogenesis. Cell 140, 49–61 (2010).
Zhang, J. et al. Monoacylglycerol lipase: a novel potential therapeutic target and prognostic indicator for hepatocellular carcinoma. Sci. Rep. 6, 35784 (2016).
Deng, H. & Li, W. Therapeutic potential of targeting α/β-Hydrolase domain-containing 6 (ABHD6). Eur. J. Med. Chem. 198, 112353 (2020).
Frühbeck, G., Méndez-Giménez, L., Fernández-Formoso, J.-A., Fernández, S. & Rodríguez, A. Regulation of adipocyte lipolysis. Nutr. Res. Rev. 27, 63–93 (2014).
Yogosawa, S., Mizutani, S., Ogawa, Y. & Izumi, T. Activin receptor-like kinase 7 suppresses lipolysis to accumulate fat in obesity through downregulation of peroxisome proliferator-activated receptor γ and C/EBPα. Diabetes 62, 115–123 (2013).
Roy, D. et al. Coordinated transcriptional control of adipocyte triglyceride lipase (Atgl) by transcription factors Sp1 and peroxisome proliferator–activated receptor γ (PPARγ) during adipocyte differentiation. J. Biol. Chem. 292, 14827–14835 (2017).
Kim, J. Y., Tillison, K., Lee, J.-H., Rearick, D. A. & Smas, C. M. The adipose tissue triglyceride lipase ATGL/PNPLA2 is downregulated by insulin and TNF-alpha in 3T3-L1 adipocytes and is a target for transactivation by PPARγ. Am. J. Physiol. Endocrinol. Metab. 291, E115–E127 (2006).
Stenson, B. M. et al. Liver X receptor (LXR) regulates human adipocyte lipolysis. J. Biol. Chem. 286, 370–379 (2011).
Kulcenty, K., Holysz, M. & Trzeciak, W. H. SF-1 (NR5A1) expression is stimulated by the PKA pathway and is essential for the PKA-induced activation of LIPE expression in Y-1 cells. Mol. Cell. Biochem. 408, 139–145 (2015).
Gambo, Y., Matsumura, M. & Fujimori, K. Triiodothyronine enhances accumulation of intracellular lipids in adipocytes through thyroid hormone receptor α via direct and indirect mechanisms. Mol. Cell. Endocrinol. 431, 1–11 (2016).
Fujimoto, Y. et al. TFE3 controls lipid metabolism in adipose tissue of male mice by suppressing lipolysis and thermogenesis. Endocrinology 154, 3577–3588 (2013).
Czajkowski, M. T., Holysz, M. & Trzeciak, W. H. Induction of hormone-sensitive lipase/cholesteryl esterase gene expression by C/EBPα independently of the PKA pathway in the adrenocortical Y-1 cells. Steroids 104, 118–121 (2015).
Kaltenecker, D. et al. Adipocyte STAT5 deficiency promotes adiposity and impairs lipid mobilisation in mice. Diabetologia 60, 296–305 (2017).
Kaltenecker, D. et al. STAT5 is required for lipid breakdown and beta-adrenergic responsiveness of brown adipose tissue. Mol. Metab. 40, 101026 (2020).
Shi, S. Y. et al. Adipocyte-specific deficiency of Janus kinase (JAK) 2 in mice impairs lipolysis and increases body weight, and leads to insulin resistance with ageing. Diabetologia 57, 1016–1026 (2014).
Hong, S. et al. Phosphorylation of beta-3 adrenergic receptor at serine 247 by ERK MAP kinase drives lipolysis in obese adipocytes. Mol. Metab. 12, 25–38 (2018).
Greenberg, A. S. et al. Stimulation of lipolysis and hormone-sensitive lipase via the extracellular signal-regulated kinase pathway. J. Biol. Chem. 276, 45456–45461 (2001).
Hjelholt, A. J. et al. Temporal patterns of lipolytic regulators in adipose tissue after acute growth hormone exposure in human subjects: A randomized controlled crossover trial. Mol. Metab. 29, 65–75 (2019).
Kopchick, J. J., Berryman, D. E., Puri, V., Lee, K. Y. & Jorgensen, J. O. L. The effects of growth hormone on adipose tissue: old observations, new mechanisms. Nat. Rev. Endocrinol. 16, 135–146 (2020).
El-Merahbi, R. et al. The adrenergic-induced ERK3 pathway drives lipolysis and suppresses energy dissipation. Genes Dev. 34, 495–510 (2020).
Magnusson, B., Svensson, P. A., Carlsson, L. M. S. & Sjöholm, K. Activin B inhibits lipolysis in 3T3-L1 adipocytes. Biochem. Biophys. Res. Commun. 395, 373–376 (2010).
Bu, Y. et al. Insulin regulates lipolysis and fat mass by upregulating growth/differentiation factor 3 in adipose tissue macrophages. Diabetes 67, 1761–1772 (2018).
Zhu, H. J. et al. The effect of myostatin on proliferation and lipid accumulation in 3T3-L1 preadipocytes. J. Mol. Endocrinol. 54, 217–226 (2015).
Modica, S. et al. Bmp4 promotes a brown to white-like adipocyte shift. Cell Rep. 16, 2243–2258 (2016).
Boon, M. R. et al. BMP7 activates brown adipose tissue and reduces diet-induced obesity only at subthermoneutrality. PLoS ONE 8, e74083 (2013).
Guo, T. et al. Adipocyte ALK7 links nutrient overload to catecholamine resistance in obesity. Elife 3, e03245 (2014).
Li, F., Yang, H., Duan, Y. & Yin, Y. Myostatin regulates preadipocyte differentiation and lipid metabolism of adipocyte via ERK1/2. Cell Biol. Int. 35, 1141–1146 (2011).
Lee, S.-J. Targeting the myostatin signaling pathway to treat muscle loss and metabolic dysfunction. J. Clin. Invest. 131, e148372 (2021).
Langin, D. & Arner, P. Importance of TNFα and neutral lipases in human adipose tissue lipolysis. Trends Endocrinol. Metab. 17, 314–320 (2006).
Li, Y. et al. A global perspective on FOXO1 in lipid metabolism and lipid-related diseases. Prog. Lipid Res. 66, 42–49 (2017).
Chakrabarti, P. & Kandror, K. V. FoxO1 controls insulin-dependent adipose triglyceride lipase (ATGL) expression and lipolysis in adipocytes. J. Biol. Chem. 284, 13296–13300 (2009).
Barthel, A., Schmoll, D. & Unterman, T. G. FoxO proteins in insulin action and metabolism. Trends Endocrinol. Metab. 16, 183–189 (2005).
Ow, J. R. et al. Remodeling of whole-body lipid metabolism and a diabetic-like phenotype caused by loss of CDK1 and hepatocyte division. eLife 9, e63835 (2020).
Saline, M. et al. AMPK and AKT protein kinases hierarchically phosphorylate the N-terminus of the FOXO1 transcription factor, modulating interactions with 14-3-3 proteins. J. Biol. Chem. 294, 13106–13116 (2019).
van der Heide, L. P. & Smidt, M. P. Regulation of FoxO activity by CBP/p300-mediated acetylation. Trends Biochem. Sci. 30, 81–86 (2005).
Chakrabarti, P. et al. SIRT1 controls lipolysis in adipocytes via FOXO1-mediated expression of ATGL. J. Lipid Res. 52, 1693–1701 (2011).
Jung, S. M. et al. Non-canonical mTORC2 signaling regulates brown adipocyte lipid catabolism through SIRT6–FoxO1. Mol. Cell 75, 807–822.e8 (2019).
Sun, C. et al. Adipose SNAIL1 regulates lipolysis and lipid partitioning by suppressing adipose triacylglycerol lipase expression. Cell Rep. 17, 2015–2027 (2016).
Chakrabarti, P., English, T., Shi, J., Smas, C. M. & Kandror, K. V. Mammalian target of rapamycin complex 1 suppresses lipolysis, stimulates lipogenesis, and promotes fat storage. Diabetes 59, 775–781 (2010).
Paolella, L. M. et al. mTORC1 restrains adipocyte lipolysis to prevent systemic hyperlipidemia. Mol. Metab. 32, 136–147 (2020).
Zhang, X. et al. Sustained activation of autophagy suppresses adipocyte maturation via a lipolysis-dependent mechanism. Autophagy 16, 1668–1682 (2020).
Chakrabarti, P. et al. Insulin inhibits lipolysis in adipocytes via the evolutionarily conserved mTORC1–Egr1–ATGL-mediated pathway. Mol. Cell. Biol. 33, 3659–3666 (2013).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Kumar, A. et al. Fat cell-specific ablation of rictor in mice impairs insulin-regulated fat cell and whole-body glucose and lipid metabolism. Diabetes 59, 1397–1406 (2010).
Caron, A., Richard, D. & Laplante, M. The roles of mTOR complexes in lipid metabolism. Annu. Rev. Nutr. 35, 321–348 (2015).
Steinberg, D. & Huttunen, J. K. The role of cyclic AMP in activation of hormone-sensitive lipase of adipose tissue. Adv. Cycl. Nucleotide Res. 1, 47–62 (1972).
Sveidahl Johansen, O. et al. Lipolysis drives expression of the constitutively active receptor GPR3 to induce adipose thermogenesis. Cell 184, 3502–3518.e33 (2021).
Kimmel, A. R. & Sztalryd, C. The perilipins: major cytosolic lipid droplet-associated proteins and their roles in cellular lipid storage, mobilization, and systemic homeostasis. Annu. Rev. Nutr. 36, 471–509 (2016).
Carper, D. et al. Atrial natriuretic peptide orchestrates a coordinated physiological response to fuel non-shivering thermogenesis. Cell Rep. 32, 108075 (2020).
Yu, L., Li, Y., Grisé, A. & Wang, H. CGI-58: versatile regulator of intracellular lipid droplet homeostasis. Adv. Exp. Med. Biol. 1276, 197–222 (2020).
Gandotra, S. et al. Perilipin deficiency and autosomal dominant partial lipodystrophy. N. Engl. J. Med. 364, 740–748 (2011).
Gandotra, S. et al. Human frame shift mutations affecting the carboxyl terminus of perilipin increase lipolysis by failing to sequester the adipose triglyceride lipase (ATGL) coactivator AB-hydrolase-containing 5 (ABHD5). J. Biol. Chem. 286, 34998–35006 (2011).
Wang, H. et al. Unique regulation of adipose triglyceride lipase (ATGL) by perilipin 5, a lipid droplet-associated protein. J. Biol. Chem. 286, 15707–15715 (2011).
Granneman, J. G., Moore, H. P., Mottillo, E. P., Zhu, Z. & Zhou, L. Interactions of perilipin-5 (Plin5) with adipose triglyceride lipase. J. Biol. Chem. 286, 5126–5135 (2011).
Pollak, N. M. et al. The interplay of protein kinase A and perilipin 5 regulates cardiac lipolysis. J. Biol. Chem. 290, 1295–1306 (2015).
Keenan, S. N. et al. Perilipin 5 S155 phosphorylation by PKA is required for the control of hepatic lipid metabolism and glycemic control. J. Lipid Res. 62, 100016 (2021).
Kuramoto, K. et al. Deficiency of a lipid droplet protein, perilipin 5, suppresses myocardial lipid accumulation, thereby preventing type 1 diabetes-induced heart malfunction. Mol. Cell. Biol. 34, 2721–2731 (2014).
Kolleritsch, S. et al. Low cardiac lipolysis reduces mitochondrial fission and prevents lipotoxic heart dysfunction in perilipin 5 mutant mice. Cardiovasc. Res. 116, 339–352 (2020).
Patel, S., Yang, W., Kozusko, K., Saudek, V. & Savage, D. B. Perilipins 2 and 3 lack a carboxy-terminal domain present in perilipin 1 involved in sequestering ABHD5 and suppressing basal lipolysis. Proc. Natl Acad. Sci. USA 111, 9163–9168 (2014).
Petersen, M. C. & Shulman, G. I. Mechanisms of insulin action and insulin resistance. Physiol. Rev. 98, 2133–2223 (2018).
DiPilato, L. M. et al. The role of PDE3B phosphorylation in the inhibition of lipolysis by insulin. Mol. Cell. Biol. 35, 2752–2760 (2015).
Xia, W. et al. Loss of ABHD15 impairs the anti-lipolytic action of insulin by altering PDE3B stability and contributes to insulin resistance. Cell Rep. 23, 1948–1961 (2018).
Vannucci, S. J., Klim, C. M., Martin, L. F. & LaNoue, K. F. A1-adenosine receptor-mediated inhibition of adipocyte adenylate cyclase and lipolysis in Zucker rats. Am. J. Physiol. 257, E871–E878 (1989).
Ceddia, R. B. The role of AMP-activated protein kinase in regulating white adipose tissue metabolism. Mol. Cell. Endocrinol. 366, 194–203 (2013).
Mottillo, E. P. et al. Lack of adipocyte AMPK exacerbates insulin resistance and hepatic steatosis through brown and beige adipose tissue function. Cell Metab. 24, 118–129 (2016).
Chen, G., Zhou, G., Lotvola, A., Granneman, J. G. & Wang, J. ABHD5 suppresses cancer cell anabolism through lipolysis-dependent activation of the AMPK/mTORC1 pathway. J. Biol. Chem. 296, jbc.RA120.014682 (2021).
Xie, H. et al. Adipose triglyceride lipase activity regulates cancer cell proliferation via AMP-kinase and mTOR signaling. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1865, 158737 (2020).
Rohm, M. et al. An AMP-activated protein kinase-stabilizing peptide ameliorates adipose tissue wasting in cancer cachexia in mice. Nat. Med. 22, 1120–1130 (2016).
Lord, C. C., Thomas, G. & Brown, J. M. Mammalian alpha beta hydrolase domain (ABHD) proteins: lipid metabolizing enzymes at the interface of cell signaling and energy metabolism. Biochim. Biophys. Acta 1831, 792–802 (2013).
Boeszoermenyi, A. et al. Structure of a CGI-58 motif provides the molecular basis of lipid droplet anchoring. J. Biol. Chem. 290, 26361–26372 (2015).
Sanders, M. A., Zhang, H., Mladenovic, L., Tseng, Y. Y. & Granneman, J. G. Molecular basis of ABHD5 lipolysis activation. Sci. Rep. 7, 42589 (2017).
Sanders, M. A. et al. Endogenous and synthetic ABHD5 ligands regulate ABHD5–perilipin interactions and lipolysis in fat and muscle. Cell Metab. 22, 851–860 (2015).
Radner, F. P. W. et al. Growth retardation, impaired triacylglycerol catabolism, hepatic steatosis, and lethal skin barrier defect in mice lacking comparative gene identification-58 (CGI-58). J. Biol. Chem. 285, 7300–7311 (2010).
Lefèvre, C. et al. Mutations in CGI-58, the gene encoding a new protein of the esterase/lipase/thioesterase subfamily, in Chanarin-Dorfman syndrome. Am. J. Hum. Genet. 69, 1002–1012 (2001).
Schweiger, M., Lass, A., Zimmermann, R., Eichmann, T. O. & Zechner, R. Neutral lipid storage disease: genetic disorders caused by mutations in adipose triglyceride lipase/PNPLA2 or CGI-58/ABHD5. Am. J. Physiol. Endocrinol. Metab. 297, E289–E296 (2009).
Kien, B. et al. ABHD5 stimulates PNPLA1-mediated O-acylceramide biosynthesis essential for a functional skin permeability barrier. J. Lipid Res. 59, 2360–2367 (2018).
Ohno, Y., Nara, A., Nakamichi, S. & Kihara, A. Molecular mechanism of the ichthyosis pathology of Chanarin–Dorfman syndrome: stimulation of PNPLA1-catalyzed ω-O-acylceramide production by ABHD5. J. Dermatol. Sci. 92, 245–253 (2018).
Yang, A., Mottillo, E. P., Mladenovic-Lucas, L., Zhou, L. & Granneman, J. G. Dynamic interactions of ABHD5 with PNPLA3 regulate triacylglycerol metabolism in brown adipocytes. Nat. Metab. 1, 560–569 (2019).
Montero-Moran, G. et al. CGI-58/ABHD5 is a coenzyme A-dependent lysophosphatidic acid acyltransferase. J. Lipid Res. 51, 709–719 (2010).
Jebessa, Z. H. et al. The lipid droplet-associated protein ABHD5 protects the heart through proteolysis of HDAC4. Nat. Metab. 1, 1157–1167 (2019).
Russell, L. & Forsdyke, D. R. A human putative lymphocyte G0/G1 switch gene containing a CpG-Rich island encodes a small basic protein with the potential to be phosphorylated. DNA Cell Biol. 10, 581–591 (1991).
Heckmann, B. L., Zhang, X., Xie, X. & Liu, J. The G0/G1 switch gene 2 (G0S2): regulating metabolism and beyond. Biochim. Biophys. Acta 1831, 276–281 (2013).
Yang, X. et al. The G0/G1 switch gene 2 regulates adipose lipolysis through association with adipose triglyceride lipase. Cell Metab. 11, 194–205 (2010).
Zhang, X., Heckmann, B. L., Campbell, L. E. & Liu, J. G0S2: A small giant controller of lipolysis and adipose-liver fatty acid flux. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 1862, 1146–1154 (2017).
Lu, X., Yang, X. & Liu, J. Differential control of ATGL-mediated lipid droplet degradation by CGI-58 and G0S2. Cell Cycle 9, 2719–2725 (2010).
Cerk, I. K. et al. A peptide derived from G0/G1 switch gene 2 acts as noncompetitive inhibitor of adipose triglyceride lipase. J. Biol. Chem. 289, 32559–32570 (2014).
Zhang, X. et al. Identification of an intrinsic lysophosphatidic acid acyltransferase activity in the lipolytic inhibitor G0/G1 switch gene 2 (G0S2). FASEB J. 33, 6655–6666 (2019).
Zandbergen, F. et al. The G0/G1 switch gene 2 is a novel PPAR target gene. Biochem. J. 392, 313–324 (2005).
Heckmann, B. L. et al. Liver X receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 switch gene 2 expression. JCI Insight 2, e88735 (2017).
Heier, C. et al. G0/G1 switch gene 2 regulates cardiac lipolysis. J. Biol. Chem. 290, 26141–26150 (2015).
Heckmann, B. L. et al. Defective adipose lipolysis and altered global energy metabolism in mice with adipose overexpression of the lipolytic Inhibitor G0/G1 switch gene 2 (G0S2). J. Biol. Chem. 289, 1905–1916 (2014).
Ma, T. et al. Mice lacking G0S2 are lean and cold-tolerant. Cancer Biol. Ther. 15, 643–650 (2014).
El-Assaad, W. et al. Deletion of the gene encoding G0/G 1 switch protein 2 (G0s2) alleviates high-fat-diet-induced weight gain and insulin resistance, and promotes browning of white adipose tissue in mice. Diabetologia 58, 149–157 (2015).
Zhang, X. et al. Targeted disruption of G0/G1 switch gene 2 enhances adipose lipolysis, alters hepatic energy balance, and alleviates high-fat diet-induced liver steatosis. Diabetes 63, 934–946 (2014).
Denko, N. et al. Epigenetic regulation of gene expression in cervical cancer cells by the tumor microenvironment. Clin. Cancer Res. 6, 480–487 (2000).
Gimm, T. et al. Hypoxia-inducible protein 2 is a novel lipid droplet protein and a specific target gene of hypoxia-inducible factor-1. FASEB J. 24, 4443–4458 (2010).
Mattijssen, F. et al. Hypoxia-inducible lipid droplet-associated (HILPDA) is a novel peroxisome proliferator-activated receptor (PPAR) target involved in hepatic triglyceride secretion. J. Biol. Chem. 289, 19279–19293 (2014).
de la Rosa Rodriguez, M. A. & Kersten, S. Regulation of lipid droplet homeostasis by hypoxia inducible lipid droplet associated HILPDA. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 1865, 158738 (2020).
DiStefano, M. T. et al. The lipid droplet protein hypoxia-inducible gene 2 promotes hepatic triglyceride deposition by inhibiting lipolysis. J. Biol. Chem. 290, 15175–15184 (2015).
Zhang, X. et al. Inhibition of intracellular lipolysis promotes human cancer cell adaptation to hypoxia. eLife 6, e31132 (2017).
Padmanabha Das, K. M. et al. Hypoxia-inducible lipid droplet-associated protein inhibits adipose triglyceride lipase. J. Lipid Res. 59, 531–541 (2018).
de la Rosa Rodriguez, M. A. et al. Hypoxia-inducible lipid droplet-associated induces DGAT1 and promotes lipid storage in hepatocytes. Mol. Metab. 47, 101168 (2021).
Dijk, W. et al. Hypoxia-inducible lipid droplet–associated Is not a direct physiological regulator of lipolysis in adipose tissue. Endocrinology 158, 1231–1251 (2017).
DiStefano, M. T. et al. Adipocyte-specific hypoxia-inducible gene 2 promotes fat deposition and diet-induced insulin resistance. Mol. Metab. 5, 1149–1161 (2016).
Hofer, P., Taschler, U., Schreiber, R., Kotzbeck, P. & Schoiswohl, G. The lipolysome—a highly complex and dynamic protein network orchestrating cytoplasmic triacylglycerol degradation. Metabolites 10, 147 (2020).
Smith, A. J. et al. Physical association between the adipocyte fatty acid-binding protein and hormone-sensitive lipase: a fluorescence resonance energy transfer analysis. J. Biol. Chem. 279, 52399–52405 (2004).
Gillilan, R. E., Ayers, S. D. & Noy, N. Structural basis for activation of fatty acid-binding protein 4. J. Mol. Biol. 372, 1246–1260 (2007).
Mahadevan, S. & Tappel, A. L. Lysosomal lipases of rat liver and kidney. J. Biol. Chem. 243, 2849–2854 (1968).
Grumet, L. et al. Lysosomal acid lipase hydrolyzes retinyl ester and affects retinoid turnover. J. Biol. Chem. 291, 17977–17987 (2016).
Sheriff, S., Du, H. & Grabowski, G. A. Characterization of lysosomal acid lipase by site-directed mutagenesis and heterologous expression. J. Biol. Chem. 270, 27766–27772 (1995).
Warner, T. G., Dambach, L. M., Shin, J. H. & O’Brien, J. S. Purification of the lysosomal acid lipase from human liver and its role in lysosomal lipid hydrolysis. J. Biol. Chem. 256, 2952–2957 (1981).
Ameis, D., Merkel, M., Eckerskorn, C. & Greten, H. Purification, characterization and molecular cloning of human hepatic lysosomal acid lipase. Eur. J. Biochem. 219, 905–914 (1994).
Rajamohan, F. et al. Crystal structure of human lysosomal acid lipase and its implications in cholesteryl ester storage disease. J. Lipid Res. 61, 1192–1202 (2020).
Zechner, R., Madeo, F. & Kratky, D. Cytosolic lipolysis and lipophagy: two sides of the same coin. Nat. Rev. Mol. Cell Biol. 18, 671–684 (2017).
Sando, G. N. & Henke, V. L. Recognition and receptor-mediated endocytosis of the lysosomal acid lipase secreted by cultured human fibroblasts. J. Lipid Res. 23, 114–123 (1982).
Haka, A. S. et al. Macrophages create an acidic extracellular hydrolytic compartment to digest aggregated lipoproteins. Mol. Biol. Cell 20, 4932–4940 (2009).
Singh, R. K. et al. Degradation of aggregated LDL occurs in complex extracellular sub-compartments of the lysosomal synapse. J. Cell Sci. 129, 1072–1082 (2016).
Leake, D. S. Does an acidic pH explain why low density lipoprotein is oxidised in atherosclerotic lesions? Atherosclerosis 129, 149–157 (1997).
Pajed, L. et al. Hepatocyte-specific deletion of lysosomal acid lipase leads to cholesteryl ester but not triglyceride or retinyl ester accumulation. J. Biol. Chem. 294, 9118–9133 (2019).
Burton, B. K. et al. A phase 3 trial of sebelipase alfa in lysosomal acid lipase deficiency. N. Engl. J. Med. 373, 1010–1020 (2015).
Ikonen, E. Cellular cholesterol trafficking and compartmentalization. Nat. Rev. Mol. Cell Biol. 9, 125–138 (2008).
Goldstein, J. L. & Brown, M. S. A century of cholesterol and coronaries: from plaques to genes to statins. Cell 161, 161–172 (2015).
Radović, B. et al. Lysosomal acid lipase regulates VLDL synthesis and insulin sensitivity in mice. Diabetologia 59, 1743–1752 (2016).
Huang, S. C. C. et al. Cell-intrinsic lysosomal lipolysis is essential for alternative activation of macrophages. Nat. Immunol. 15, 846–855 (2014).
Schlager, S. et al. Lysosomal lipid hydrolysis provides substrates for lipid mediator synthesis in murine macrophages. Oncotarget 8, 40037–40051 (2017).
Duta-Mare, M. et al. Lysosomal acid lipase regulates fatty acid channeling in brown adipose tissue to maintain thermogenesis. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 1863, 467–478 (2018).
Singh, R. et al. Autophagy regulates lipid metabolism. Nature 458, 1131–1135 (2009).
Ouimet, M. et al. Autophagy regulates cholesterol efflux from macrophage foam cells via lysosomal acid lipase. Cell Metab. 13, 655–667 (2011).
Cui, W. et al. Lipophagy-derived fatty acids undergo extracellular efflux via lysosomal exocytosis. Autophagy 17, 690–705 (2021).
Schulze, R. J., Sathyanarayan, A. & Mashek, D. G. Breaking fat: the regulation and mechanisms of lipophagy. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 1862, 1178–1187 (2017).
Mijaljica, D., Prescott, M. & Devenish, R. J. Microautophagy in mammalian cells: revisiting a 40-year-old conundrum. Autophagy 7, 673–682 (2011).
Schulze, R. J. et al. Direct lysosome-based autophagy of lipid droplets in hepatocytes. Proc. Natl Acad. Sci. USA 117, 32443–32452 (2020).
Schott, M. B. et al. Lipid droplet size directs lipolysis and lipophagy catabolism in hepatocytes. J. Cell Biol. 218, 3320–3335 (2019).
Kaushik, S. & Cuervo, A. M. Degradation of lipid droplet-associated proteins by chaperone-mediated autophagy facilitates lipolysis. Nat. Cell Biol. 17, 759–770 (2015).
Fred Dice, J. Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem. Sci. 15, 305–309 (1990).
Chiang, H. L., Terlecky, S. R., Plant, C. P. & Dice, J. F. A role for a 70-kilodaton heat shock protein in lysosomal degradation of intracellular proteins. Science 246, 382–385 (1989).
Du, H., Duanmu, M., Witte, D. & Grabowski, G. A. Targeted disruption of the mouse lysosomal acid lipase gene: long-term survival with massive cholesteryl ester and triglyceride storage. Hum. Mol. Genet. 7, 1347–1354 (1998).
Du, H. et al. Lysosomal acid lipase-deficient mice: depletion of white and brown fat, severe hepatosplenomegaly, and shortened life span. J. Lipid Res. 42, 489–500 (2001).
Patrick, A. D. & Lake, B. D. Deficiency of an acid lipase in wolman’s disease. Nature 222, 1067–1068 (1969).
Burke, J. A. & Schubert, W. K. Deficient activity of hepatic acid lipase in cholesterol ester storage disease. Science 176, 309–310 (1972).
Bernstein, D. L., Hülkova, H., Bialer, M. G. & Desnick, R. J. Cholesteryl ester storage disease: review of the findings in 135 reported patients with an underdiagnosed disease. J. Hepatol. 58, 1230–1243 (2013).
Wild, P. S. et al. A genome-wide association study identifies LIPA as a susceptibility gene for coronary artery disease. Circ. Cardiovasc. Genet. 4, 403–412 (2011).
Schweiger, M. et al. Adipose triglyceride lipase and hormone-sensitive lipase are the major enzymes in adipose tissue triacylglycerol catabolism. J. Biol. Chem. 281, 40236–40241 (2006).
Pichery, M. et al. PNPLA1 defects in patients with autosomal recessive congenital ichthyosis and KO mice sustain PNPLA1 irreplaceable function in epidermal omega-O-acylceramide synthesis and skin permeability barrier. Hum. Mol. Genet. 26, 1787–1800 (2017).
Hirabayashi, T. et al. PNPLA1 has a crucial role in skin barrier function by directing acylceramide biosynthesis. Nat. Commun. 8, 14609 (2017).
Grall, A. et al. PNPLA1 mutations cause autosomal recessive congenital ichthyosis in golden retriever dogs and humans. Nat. Genet. 44, 140–147 (2012).
Onal, G. et al. Impairment of lipophagy by PNPLA1 mutations causes lipid droplet accumulation in primary fibroblasts of Autosomal Recessive Congenital Ichthyosis patients. J. Dermatol. Sci. 93, 50–57 (2019).
Wilson, P. A., Gardner, S. D., Lambie, N. M., Commans, S. A. & Crowther, D. J. Characterization of the human patatin-like phospholipase family. J. Lipid Res. 47, 1940–1949 (2006).
Baulande, S., Lasnier, F., Lucas, M. & Pairault, J. Adiponutrin, a transmembrane protein corresponding to a novel dietary- and obesity-linked mRNA specifically expressed in the adipose Lineage. J. Biol. Chem. 276, 33336–33344 (2001).
Dubuquoy, C. et al. Distinct regulation of adiponutrin/PNPLA3 gene expression by the transcription factors ChREBP and SREBP1c in mouse and human hepatocytes. J. Hepatol. 55, 145–153 (2011).
He, S. et al. A sequence variation (I148M) in PNPLA3 associated with nonalcoholic fatty liver disease disrupts triglyceride hydrolysis. J. Biol. Chem. 285, 6706–6715 (2010).
Pirazzi, C. et al. PNPLA3 has retinyl-palmitate lipase activity in human hepatic stellate cells. Hum. Mol. Genet. 23, 4077–4085 (2014).
Huang, Y., Cohen, J. C. & Hobbs, H. H. Expression and characterization of a PNPLA3 protein isoform (I148M) associated with nonalcoholic fatty liver disease. J. Biol. Chem. 286, 37085–37093 (2011).
Kumari, M. et al. Adiponutrin functions as a nutritionally regulated lysophosphatidic acid acyltransferase. Cell Metab. 15, 691–702 (2012).
Dong, X. C. PNPLA3—a potential therapeutic target for personalized treatment of chronic liver disease. Front. Med. 6, 304 (2019).
Basantani, M. K. et al. Pnpla3/adiponutrin deficiency in mice does not contribute to fatty liver disease or metabolic syndrome. J. Lipid Res. 52, 318–329 (2011).
Chen, W., Chang, B., Li, L. & Chan, L. Patatin-like phospholipase domain-containing 3/adiponutrin deficiency in mice is not associated with fatty liver disease. Hepatology 52, 1134–1142 (2010).
Romeo, S. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40, 1461–1465 (2008).
Wang, Y., Kory, N., BasuRay, S., Cohen, J. C. & Hobbs, H. H. PNPLA3, CGI‐58, and inhibition of hepatic triglyceride hydrolysis in Mice. Hepatology 69, hep.30583 (2019).
BasuRay, S., Smagris, E., Cohen, J. C. & Hobbs, H. H. The PNPLA3 variant associated with fatty liver disease (I148M) accumulates on lipid droplets by evading ubiquitylation. Hepatology 66, 1111–1124 (2017).
Negoita, F. et al. PNPLA3 variant M148 causes resistance to starvation-mediated lipid droplet autophagy in human hepatocytes. J. Cell. Biochem. 120, 343–356 (2019).
Lee, W. C., Salido, E. & Yen, P. H. Isolation of a new gene GS2 (DXS1283E) from a CpG island between STS and KAL1 on xp22.3. Genomics 22, 372–376 (1994).
Lake, A. C. et al. Expression, regulation, and triglyceride hydrolase activity of adiponutrin family members. J. Lipid Res. 46, 2477–2487 (2005).
Gao, J. & Simon, M. Identification of a novel keratinocyte retinyl ester hydrolase as a transacylase and lipase. J. Invest. Dermatol. 124, 1259–1266 (2005).
Kohda, M. et al. A comprehensive genomic analysis reveals the genetic landscape of mitochondrial respiratory chain complex deficiencies. PLoS Genet. 12, e1005679 (2016).
Labonne, J. D. J. et al. Comparative genomic mapping implicates LRRK2 for intellectual disability and autism at 12q12, and HDHD1, as well as PNPLA4, for X-linked intellectual disability at Xp22.31. J. Clin. Med. 9, 274 (2020).
Kornahrens, A. F. et al. Design of benzoxathiazin-3-one 1,1-dioxides as a new class of irreversible serine hydrolase inhibitors: discovery of a uniquely selective PNPLA4 inhibitor. J. Am. Chem. Soc. 139, 7052–7061 (2017).
Gao, J. G. & Simon, M. A comparative study of human GS2, its paralogues, and its rat orthologue. Biochem. Biophys. Res. Commun. 360, 501–506 (2007).
Dupont, N. et al. Neutral lipid stores and lipase PNPLA5 contribute to autophagosome biogenesis. Curr. Biol. 24, 609–620 (2014).
Lange, L. A. et al. Whole-exome sequencing identifies rare and low-frequency coding variants associated with LDL cholesterol. Am. J. Hum. Genet. 94, 233–245 (2014).
Lian, J., Nelson, R. & Lehner, R. Carboxylesterases in lipid metabolism: from mouse to human. Protein Cell 9, 178–195 (2018).
Holmes, R. S. et al. Recommended nomenclature for five mammalian carboxylesterase gene families: human, mouse, and rat genes and proteins. Mamm. Genome 21, 427–441 (2010).
Wang, D. et al. Human carboxylesterases: a comprehensive review. Acta Pharm. Sin. B 8, 699–712 (2018).
Zhao, B., Fisher, B. J., St Clair, R. W., Rudel, L. L. & Ghosh, S. Redistribution of macrophage cholesteryl ester hydrolase from cytoplasm to lipid droplets upon lipid loading. J. Lipid Res. 46, 2114–2121 (2005).
Ghosh, S., St Clair, R. W. & Rudel, L. L. Mobilization of cytoplasmic CE droplets by overexpression of human macrophage cholesteryl ester hydrolase. J. Lipid Res. 44, 1833–1840 (2003).
Xu, Y. et al. Hepatocyte‐specific expression of human carboxylesterase 1 attenuates diet‐induced steatohepatitis and hyperlipidemia in mice. Hepatol. Commun. 4, 527–539 (2020).
Bie, J. et al. Liver-specific transgenic expression of cholesteryl ester hydrolase reduces atherosclerosis in LDLR−/− mice. J. Lipid Res. 55, 729–738 (2014).
Zhao, B. et al. Macrophage-specific transgenic expression of cholesteryl ester hydrolase significantly reduces atherosclerosis and lesion necrosis in Ldlr−/− mice. J. Clin. Invest. 117, 2983–2992 (2007).
Crow, J. A. et al. Inhibition of carboxylesterase 1 is associated with cholesteryl ester retention in human THP-1 monocyte/macrophages. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1781, 643–654 (2008).
Igarashi, M. et al. The critical role of neutral cholesterol ester hydrolase 1 in cholesterol removal from human macrophages. Circ. Res. 107, 1387–1395 (2010).
Dolinsky, V. W., Sipione, S., Lehner, R. & Vance, D. E. The cloning and expression of a murine triacylglycerol hydrolase cDNA and the structure of its corresponding gene. Biochim. Biophys. Acta 1532, 162–172 (2001).
Gilham, D., Alam, M., Gao, W., Vance, D. E. & Lehner, R. Triacylglycerol hydrolase is localized to the endoplasmic reticulum by an unusual retrieval sequence where it participates in VLDL assembly without utilizing VLDL lipids as substrates. Mol. Biol. Cell 16, 984–996 (2005).
Soni, K. G. et al. Carboxylesterase 3 (EC 3.1.1.1) is a major adipocyte lipase. J. Biol. Chem. 279, 40683–40689 (2004).
Wei, E., Gao, W. & Lehner, R. Attenuation of adipocyte triacylglycerol hydrolase activity decreases basal fatty acid efflux. J. Biol. Chem. 282, 8027–8035 (2007).
Gilham, D. et al. Inhibitors of hepatic microsomal triacylglycerol hydrolase decrease very low density lipoprotein secretion. FASEB J. 17, 1685–1687 (2003).
Lehner, R., Lian, J. & Quiroga, A. D. Lumenal lipid metabolism: implications for lipoprotein assembly. Arterioscler. Thromb. Vasc. Biol. 32, 1087–1093 (2012).
Wei, E. et al. Apolipoprotein B and triacylglycerol secretion in human triacylglycerol hydrolase transgenic mice. J. Lipid Res. 48, 2597–2606 (2007).
Lian, J. et al. Liver specific inactivation of carboxylesterase 3/triacylglycerol hydrolase decreases blood lipids without causing severe steatosis in mice. Hepatology 56, 2154–2162 (2012).
Wei, E. et al. Loss of TGH/Ces3 in mice decreases blood lipids, improves glucose tolerance, and increases energy expenditure. Cell Metab. 11, 183–193 (2010).
Lian, J. et al. Ces3/TGH deficiency attenuates steatohepatitis. Sci. Rep. 6, 25747 (2016).
Lian, J., van der Veen, J. N., Watts, R., Jacobs, R. L. & Lehner, R. Carboxylesterase 1d (Ces1d) does not contribute to cholesteryl ester hydrolysis in the liver. J. Lipid Res. 62, 100093 (2021).
Ko, K. W. S., Erickson, B. & Lehner, R. Es-x/Ces1 prevents triacylglycerol accumulation in McArdle-RH7777 hepatocytes. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1791, 1133–1143 (2009).
Xu, J. et al. Hepatic carboxylesterase 1 is essential for both normal and farnesoid X receptor-controlled lipid homeostasis. Hepatology 59, 1761–1771 (2014).
Quiroga, A. D. et al. Deficiency of carboxylesterase 1/esterase-x results in obesity, hepatic steatosis, and hyperlipidemia. Hepatology 56, 2188–2198 (2012).
Bahitham, W., Watts, R., Nelson, R., Lian, J. & Lehner, R. Liver-specific expression of carboxylesterase 1g/esterase-x reduces hepatic steatosis, counteracts dyslipidemia and improves insulin signaling. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1861, 482–490 (2016).
Li, Y. et al. Carboxylesterase 2 prevents liver steatosis by modulating lipolysis, endoplasmic reticulum stress, and lipogenesis and is regulated by hepatocyte nuclear factor 4 alpha in mice. Hepatology 63, 1860–1874 (2016).
Ruby, M. A. et al. Human carboxylesterase 2 reverses obesity-induced diacylglycerol accumulation and glucose intolerance. Cell Rep. 18, 636–646 (2017).
Xu, Y. et al. Hepatocyte-specific expression of human carboxylesterase 2 attenuates nonalcoholic steatohepatitis in mice. Am. J. Physiol. Liver Physiol. 320, G166–G174 (2021).
Chalhoub, G. et al. Carboxylesterase 2 proteins are efficient diglyceride and monoglyceride lipases possibly implicated in metabolic disease. J. Lipid Res. 62, 100075 (2021).
Maresch, L. K. et al. Intestine-specific overexpression of carboxylesterase 2c protects mice from diet-induced liver steatosis and obesity. Hepatol. Commun. 3, 227–245 (2019).
Goo, Y.-H., Son, S.-H., Kreienberg, P. B. & Paul, A. Novel lipid droplet-associated serine hydrolase regulates macrophage cholesterol mobilization. Arterioscler. Thromb. Vasc. Biol. 34, 386–396 (2014).
Thiel, K. et al. The evolutionarily conserved protein CG9186 is associated with lipid droplets, required for their positioning and for fat storage. J. Cell Sci. 126, 2198–2212 (2013).
Kory, N. et al. Mice lacking lipid droplet-associated hydrolase, a gene linked to human prostate cancer, have normal cholesterol ester metabolism. J. Lipid Res. 58, 226–235 (2017).
Kolkhof, P. et al. A luciferase-fragment complementation assay to detect lipid droplet-associated protein–protein interactions. Mol. Cell. Proteom. 16, 329–345 (2017).
Goo, Y. H., Son, S. H. & Paul, A. Lipid droplet-associated hydrolase promotes lipid droplet fusion and enhances ATGL degradation and triglyceride accumulation. Sci. Rep. 7, 1–13 (2017).
Currall, B. B. et al. Loss of LDAH associated with prostate cancer and hearing loss. Hum. Mol. Genet. 27, 4194–4203 (2018).
Nakajima, K. I. et al. A novel phospholipase A1 with sequence homology to a mammalian Sec23p-interacting protein, p125. J. Biol. Chem. 277, 11329–11335 (2002).
Tani, K., Mizoguchi, T., Iwamatsu, A., Hatsuzawa, K. & Tagaya, M. p125 Is a novel mammalian Sec23p-interacting protein with structural similarity to phospholipid-modifying proteins. J. Biol. Chem. 274, 20505–20512 (1999).
Araki, M. et al. Enzymatic characterization of recombinant rat DDHD2: a soluble diacylglycerol lipase. J. Biochem. 160, 269–279 (2016).
Aso, C. et al. Protein purification and cloning of diacylglycerol lipase from rat brain. J. Biochem. 159, 585–597 (2016).
Inloes, J. M. et al. The hereditary spastic paraplegia-related enzyme DDHD2 is a principal brain triglyceride lipase. Proc. Natl Acad. Sci. U. S. A. 111, 14924–14929 (2014).
Inloes, J. M. et al. Functional contribution of the spastic paraplegia-related triglyceride hydrolase DDHD2 to the formation and content of lipid droplets. Biochemistry 57, 827–838 (2018).
Morikawa, R. K. et al. Intracellular phospholipase A1γ(iPLA1γ) is a novel factor involved in coat protein complex I-and Rab6-independent retrograde transport between the endoplasmic reticulum and the golgi complex. J. Biol. Chem. 284, 26620–26630 (2009).
Schuurs-Hoeijmakers, J. H. M. et al. Mutations in DDHD2, encoding an intracellular phospholipase A1, cause a recessive form of complex hereditary spastic paraplegia. Am. J. Hum. Genet. 91, 1073–1081 (2012).
Gonzalez, M. et al. Mutations in phospholipase DDHD2 cause autosomal recessive hereditary spastic paraplegia (SPG54). Eur. J. Hum. Genet. 21, 1214–1218 (2013).
Citterio, A. et al. Mutations in CYP2U1, DDHD2 and GBA2 genes are rare causes of complicated forms of hereditary spastic paraparesis. J. Neurol. 261, 373–381 (2014).
Maruyama, T. et al. Loss of DDHD2, whose mutation causes spastic paraplegia, promotes reactive oxygen species generation and apoptosis. Cell Death Dis. 9, 1–15 (2018).
Yang, L. et al. Neuronal lipolysis participates in PUFA-mediated neural function and neurodegeneration. EMBO Rep. 21, e50214 (2020).
Yang, C. et al. Rewiring neuronal glycerolipid metabolism determines the extent of axon regeneration. Neuron 105, 276–292.e5 (2020).
Park, C. Y. et al. Genome-wide landscape of RNA-binding protein target site dysregulation reveals a major impact on psychiatric disorder risk. Nat. Genet. 53, 166–173 (2021).
Matoba, N. et al. Common genetic risk variants identified in the SPARK cohort support DDHD2 as a candidate risk gene for autism. Transl. Psychiatry 10, 265 (2020).
Probst, M. R. et al. Human liver arylacetamide deacetylase. Molecular cloning of a novel esterase involved in the metabolic activation of arylamine carcinogens with high sequence similarity to hormone-sensitive lipase. J. Biol. Chem. 269, 21650–21656 (1994).
Trickett, J. I. et al. Characterization of the rodent genes for arylacetamide deacetylase, a putative microsomal lipase, and evidence for transcriptional regulation. J. Biol. Chem. 276, 39522–39532 (2001).
Lo, V. et al. Arylacetamide deacetylase attenuates fatty-acid-induced triacylglycerol accumulation in rat hepatoma cells. J. Lipid Res. 51, 368–377 (2010).
Tiwari, R., Köffel, R. & Schneiter, R. An acetylation/deacetylation cycle controls the export of sterols and steroids from S. cerevisiae. EMBO J. 26, 5109–5119 (2007).
Gibbons, G. F., Islam, K. & Pease, R. J. Mobilisation of triacylglycerol stores. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1483, 37–57 (2000).
Lee, H. et al. Compositional dynamics of the milk fat globule and its role in infant development. Front. Pediatr. 6, 313 (2018).
Van Niel, G., D’Angelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Flaherty, S. E. et al. A lipase-independent pathway of lipid release and immune modulation by adipocytes. Science. 363, 989–993 (2019).
Schneider, M. R. Lipid droplets and associated proteins in sebocytes. Exp. Cell. Res. 340, 205–208 (2016).
Choa, R. et al. Thymic stromal lymphopoietin induces adipose loss through sebum hypersecretion. Science 373, eabd2893 (2021).
Zimmet, P., Alberti, K. G., Magliano, D. J. & Bennett, P. H. Diabetes mellitus statistics on prevalence and mortality: facts and fallacies. Nat. Rev. Endocrinol. 12, 616–622 (2016).
Morigny, P., Houssier, M., Mouisel, E. & Langin, D. Adipocyte lipolysis and insulin resistance. Biochimie 125, 259–266 (2016).
Hoy, A. J. et al. Adipose triglyceride lipase-null mice are resistant to high-fat diet–induced insulin resistance despite reduced energy expenditure and ectopic lipid accumulation. Endocrinology 152, 48–58 (2011).
Schoiswohl, G. et al. Impact of reduced ATGL-mediated adipocyte lipolysis on obesity-associated insulin resistance and inflammation in male mice. Endocrinology 156, 3610–3624 (2015).
Xia, B. et al. Adipose tissue deficiency of hormone-sensitive lipase causes fatty liver in mice. PLoS Genet. 13, 1–17 (2017).
Schweiger, M. et al. Pharmacological inhibition of adipose triglyceride lipase corrects high-fat diet-induced insulin resistance and hepatosteatosis in mice. Nat. Commun. 8, 14859 (2017).
Girousse, A. et al. Partial inhibition of adipose tissue lipolysis improves glucose metabolism and insulin sensitivity without alteration of fat mass. PLoS Biol. 11, e1001485 (2013).
Perry, R. J. et al. Hepatic acetyl CoA links adipose tissue inflammation to hepatic insulin resistance and type 2 diabetes. Cell 160, 1–14 (2015).
Prentki, M., Matschinsky, F. M. & Madiraju, S. R. M. Metabolic signaling in fuel-induced insulin secretion. Cell Metab. 18, 162–185 (2013).
Peyot, M.-L. et al. Adipose triglyceride lipase is implicated in fuel- and non-fuel-stimulated insulin secretion. J. Biol. Chem. 284, 16848–16859 (2009).
Liu, S. et al. Adipose triglyceride lipase is a key lipase for the mobilization of lipid droplets in human beta cells and critical for the maintenance of Syntaxin1a level in beta cells. Diabetes 69, 1178–1192 (2020).
Fex, M. et al. A beta cell-specific knockout of hormone-sensitive lipase in mice results in hyperglycaemia and disruption of exocytosis. Diabetologia 52, 271–280 (2009).
Roduit, R. et al. A role for hormone-sensitive lipase in glucose-stimulated insulin secretion: a study in hormone-sensitive lipase-deficient mice. Diabetes 50, 1970–1975 (2001).
Pearson, G. L. et al. Lysosomal acid lipase and lipophagy are constitutive negative regulators of glucose-stimulated insulin secretion from pancreatic beta cells. Diabetologia 57, 129–139 (2014).
Zhao, S. et al. α/β-hydrolase domain-6-accessible monoacylglycerol controls glucose-stimulated insulin secretion. Cell Metab. 6, 1–15 (2014).
Scheja, L. et al. Altered insulin secretion associated with reduced lipolytic efficiency in aP2–/– mice. Diabetes 48, 1987–1994 (1999).
Wu, J. W. et al. Fasting energy homeostasis in mice with adipose deficiency of desnutrin/adipose triglyceride lipase. Endocrinology 153, 2198–2207 (2012).
Mitra, S., De, A. & Chowdhury, A. Epidemiology of non-alcoholic and alcoholic fatty liver diseases. Transl. Gastroenterol. Hepatol. 5, 16 (2020).
Kim, D., Touros, A. & Kim, W. R. Nonalcoholic fatty liver disease and metabolic syndrome. Clin. Liver Dis. 22, 133–140 (2018).
Donnelly, K. L. et al. Sources of fatty acids stored in liver and secreted via lipoproteins in patients with nonalcoholic fatty liver disease. J. Clin. Invest. 115, 1343–1351 (2005).
Jaeger, D. et al. Fasting-induced G0/G1 switch gene 2 and FGF21 expression in the liver are under regulation of adipose tissue derived fatty acids. J. Hepatol. 63, 437–445 (2015).
Ahmadian, M. et al. Adipose overexpression of desnutrin promotes fatty acid use and attenuates diet-induced obesity. Diabetes 58, 855–866 (2009).
Lindén, D. et al. Pnpla3 silencing with antisense oligonucleotides ameliorates nonalcoholic steatohepatitis and fibrosis in Pnpla3 I148M knock-in mice. Mol. Metab. 22, 49–61 (2019).
BasuRay, S. PNPLA3-I148M: a problem of plenty in non-alcoholic fatty liver disease. Adipocyte 8, 201–208 (2019).
Reid, B. N. et al. Hepatic overexpression of hormone-sensitive lipase and adipose triglyceride lipase promotes fatty acid oxidation, stimulates direct release of free fatty acids, and ameliorates steatosis. J. Biol. Chem. 283, 13087–13099 (2008).
Ponikowski, P. et al. 2016 ESC Guidelines for the diagnosis and treatment of acute and chronic heart failure: The Task Force for the diagnosis and treatment of acute and chronic heart failure of the European Society of Cardiology (ESC)Developed with the special contribution of the Heart Failure Association (HFA) of the ESC. Eur. Heart J. 37, 2129–2200 (2016).
Groenewegen, A., Rutten, F. H., Mosterd, A. & Hoes, A. W. Epidemiology of heart failure. Eur. J. Heart Fail. 22, 1342–1356 (2020).
Kolwicz, S. C., Purohit, S. & Tian, R. Cardiac metabolism and its interactions with contraction, growth, and survival of cardiomyocytes. Circ. Res. 113, 603–616 (2013).
Szabó, T. et al. Increased catabolic activity in adipose tissue of patients with chronic heart failure. Eur. J. Heart Fail. 15, 1131–1137 (2013).
Cohn, J. N. et al. Plasma norepinephrine as a guide to prognosis in patients with chronic congestive heart failure. N. Engl. J. Med. 311, 819–823 (1984).
Scherbakov, N. et al. Insulin resistance in heart failure: differences between patients with reduced and preserved left ventricular ejection fraction. Eur. J. Heart Fail. 17, 1015–1021 (2015).
Parajuli, N. et al. Atglistatin ameliorates functional decline in heart failure via adipocyte-specific inhibition of adipose triglyceride lipase. Am. J. Physiol. Circ. Physiol. 315, H879–H884 (2018).
Salatzki, J. et al. Adipose tissue ATGL modifies the cardiac lipidome in pressure-overload-induced left ventricular failure. PLoS Genet. 14, e1007171 (2018).
Thiele, A. et al. Pharmacological inhibition of adipose tissue adipose triglyceride lipase by atglistatin prevents catecholamine-induced myocardial damage. Cardiovasc. Res. cvab182 (2021).
Kintscher, U., Foryst-Ludwig, A., Haemmerle, G. & Zechner, R. The role of adipose triglyceride lipase and cytosolic lipolysis in cardiac function and heart Failure. Cell Reports Med. 1, 100001 (2020).
Schiattarella, G. G. et al. Nitrosative stress drives heart failure with preserved ejection fraction. Nature 568, 351–356 (2019).
Baracos, V. E., Martin, L., Korc, M., Guttridge, D. C. & Fearon, K. C. H. Cancer-associated cachexia. Nat. Rev. Dis. Prim. 4, 1–18 (2018).
Petruzzelli, M. & Wagner, E. F. Mechanisms of metabolic dysfunction in cancer-associated cachexia. Genes Dev. 30, 489–501 (2016).
Das, S. K. et al. Adipose triglyceride lipase contributes to cancer-associated cachexia. Science 333, 233–238 (2011).
Fouladiun, M. et al. Body composition and time course changes in regional distribution of fat and lean tissue in unselected cancer patients on palliative care–correlations with food intake, metabolism, exercise capacity, and hormones. Cancer 103, 2189–2198 (2005).
Sun, X. et al. Fat wasting is damaging: role of adipose tissue in cancer-associated cachexia. Front. Cell Dev. Biol. 8, 1–9 (2020).
Petruzzelli, M. et al. A switch from white to brown fat increases energy expenditure in cancer-associated cachexia. Cell Metab. 20, 433–447 (2014).
Tsoli, M. et al. Depletion of white adipose tissue in cancer cachexia syndrome is associated with inflammatory signaling and disrupted circadian regulation. PLoS ONE 9, e92966 (2014).
Kaur, S. et al. Adipose‐specific ATGL ablation reduces burn injury‐induced metabolic derangements in mice. Clin. Transl. Med. 11, e417 (2021).
Samsa, M. M. et al. Dengue virus capsid protein usurps lipid droplets for viral particle formation. PLoS Pathog. 5, e1000632 (2009).
Richards, O. C. & Ehrenfeld, E. in Poliovirus RNA Replication. 89–119 (Springer, 1990).
Laufman, O., Perrino, J. & Andino, R. Viral generated inter-organelle contacts redirect lipid flux for genome replication. Cell 178, 275–289 (2019).
Sturley, S. L. et al. Potential COVID-19 therapeutics from a rare disease: weaponizing lipid dysregulation to combat viral infectivity. J. Lipid Res. 61, 972–982 (2020).
Vance, D. E., Trip, E. M. & Paddon, H. B. Poliovirus increases phosphatidylcholine biosynthesis in HeLa cells by stimulation of the rate-limiting reaction catalyzed by CTP:phosphocholine cytidylyltransferase. J. Biol. Chem. 255, 1064–1069 (1980).
Nchoutmboube, J. A. et al. Increased long chain acyl-Coa synthetase activity and fatty acid import is linked to membrane synthesis for development of picornavirus replication organelles. PLoS Pathog. 9, e1003401 (2013).
Yen, C. L. E., Stone, S. J., Koliwad, S., Harris, C. & Farese, R. V. DGAT enzymes and triacylglycerol biosynthesis. J. Lipid Res. 49, 2283–2301 (2008).
Miyanari, Y. et al. The lipid droplet is an important organelle for hepatitis C virus production. Nat. Cell Biol. 9, 1089–1097 (2007).
Herker, E. & Ott, M. Unique ties between hepatitis C virus replication and intracellular lipids. Trends Endocrinol. Metab. 22, 241–248 (2011).
Camus, G. et al. The hepatitis C virus core protein inhibits adipose triglyceride lipase (ATGL)-mediated lipid mobilization and enhances the ATGL interaction with comparative gene identification 58 (CGI-58) and lipid droplets. J. Biol. Chem. 289, 35770–35780 (2014).
Moriya, K. et al. Hepatitis C virus core protein induces hepatic steatosis in transgenic mice. J. Gen. Virol. 78, 1527–1531 (1997).
Nardacci, R. et al. Evidences for lipid involvement in SARS-CoV-2 cytopathogenesis. Cell Death Dis. 12, 263 (2021).
Baazim, H. et al. CD8+ T cells induce cachexia during chronic viral infection. Nat. Immunol. 6, 701–710 (2019).
Heaton, N. S. & Randall, G. Dengue virus-induced autophagy regulates lipid metabolism. Cell Host Microbe 8, 422–432 (2010).
Ayari, A. et al. Influenza infection rewires energy metabolism and induces browning features in adipose cells and tissues. Commun. Biol. 3, 237 (2020).
Wang, J. et al. Porcine reproductive and respiratory syndrome virus activates lipophagy to facilitate viral replication through downregulation of NDRG1 expression. J. Virol. 93, 1–20 (2019).
Taylor, M. P. & Karla, K. Modification of cellular autophagy protein LC3 by poliovirus. J. Virol. 81, 12543–12553 (2007).
Dreux, M., Gastaminza, P., Wieland, S. F. & Chisari, F. V. The autophagy machinery is required to initiate hepatitis C virus replication. Proc. Natl Acad. Sci. USA 106, 14046–14051 (2009).
Wong, J. et al. Autophagosome supports coxsackievirus B3 replication in host cells. J. Virol. 82, 9143–9153 (2008).
Nourbakhsh, M. et al. Arylacetamide deacetylase: a novel host factor with important roles in the lipolysis of cellular triacylglycerol stores, VLDL assembly and HCV production. J. Hepatol. 59, 336–343 (2013).
Vieyres, G. et al. ABHD5/CGI-58, the Chanarin–Dorfman syndrome protein, mobilises lipid stores for hepatitis C virus production. PLoS Pathog. 12, e1005568 (2016).
Brown, J. M. et al. CGI-58 facilitates the mobilization of cytoplasmic triglyceride for lipoprotein secretion in hepatoma cells. J. Lipid Res. 48, 2295–2305 (2007).
Shoemaker, J. P., Hoffman, R. V. & Huffman, D. G. Trypanosoma cruzi: preference for brown adipose tissue in mice by the Tulahuen strain. Exp. Parasitol. 27, 403–407 (1970).
Trindade, S. et al. Trypanosoma brucei parasites occupy and functionally adapt to the adipose tissue in mice. Cell Host Microbe 19, 837–848 (2016).
Bechah, Y. et al. Persistence of Coxiella burnetii, the agent of q fever, in murine adipose tissue. PLoS ONE 9, e97503 (2014).
Bechah, Y., Paddock, C. D., Capo, C., Mege, J.-L. & Raoult, D. Adipose tissue serves as a reservoir for recrudescent Rickettsia prowazekii infection in a mouse model. PLoS ONE 5, e8547 (2010).
Neyrolles, O. et al. Is adipose tissue a place for Mycobacterium tuberculosis persistence? PLoS ONE 1, e43 (2006).
Combs, T. P. et al. The adipocyte as an important target cell for Trypanosoma cruzi infection. J. Biol. Chem. 280, 24085–24094 (2005).
Kennedy, P. G. Clinical features, diagnosis, and treatment of human African trypanosomiasis (sleeping sickness). Lancet Neurol. 12, 186–194 (2013).
Ranjithkumar, M. et al. Hyperlipidaemia in trypanosomiasis of naturally infected horses: possible cachexia–anorexia syndrome? Trop. Anim. Health Prod. 45, 417–421 (2013).
Wang, A. et al. Opposing effects of fasting metabolism on tissue tolerance in bacterial and viral inflammation. Cell 166, 1512–1525 (2016).
Acknowledgements
R. Z. and M. S. are supported by grants from the FWF SFB Lipid Hydrolysis (grant number F7302) and SFB Immunometabolism (grant number F83), respectively. R. Z. is additionally supported by the Louis Jeantet Prize 2015 awarded by the Louis Jeantet Foundation.
Author information
Authors and Affiliations
Contributions
Conceptualization was done by M. S. and R. Z.; G. F. G., H. X., M. S. and R. Z. wrote the manuscript; figures were prepared by G. F. G. and M. S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks Dominique Langin and the other, anonymous reviewers for their contribution to the peer review of this work. Primary handling editor: Christoph Schmitt.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Grabner, G.F., Xie, H., Schweiger, M. et al. Lipolysis: cellular mechanisms for lipid mobilization from fat stores. Nat Metab 3, 1445–1465 (2021). https://doi.org/10.1038/s42255-021-00493-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-021-00493-6
This article is cited by
-
Hepatic loss of AATF attenuates MASH by suppressing AKT–mTORC1 signaling and reprogramming lipid metabolism
Molecular Medicine (2026)
-
Non-coding RNAs in lipid metabolism and their roles in atherosclerosis
Nature Reviews Cardiology (2026)
-
eIF3 musketeers: loyal in health, rogue in disease, and redeemed by therapeutic targeting
The EMBO Journal (2026)
-
Glycophagy is an ancient bilaterian pathway supporting metabolic adaptation through STBD1 structural evolution
Communications Biology (2026)
-
Canagliflozin regulates adipocyte lipolysis in vitro via a SGLT2 independent signaling pathway
International Journal of Obesity (2026)