Abstract
Predicting how small molecules affect diverse cell types phenotypically is central to drug discovery, yet it remains a challenging task. Modelling cell-type-specific transcriptional responses provides a scalable alternative for early candidate identification, enabling broader exploration and lower costs than exhaustive experimental exploration of the chemical space. Here we present PrePR-CT, a graph-based deep learning approach that utilizes cell-type-specific co-expression networks as an inductive bias to predict transcriptional responses to chemical perturbations. Graph attention networks learn biologically meaningful representations that capture cell-type-specific gene interactions, enabling gene-level attributions. Across five single-cell RNA sequencing datasets, including human blood and multiple cancer lines, one bulk transcriptomics dataset and a large-scale small-molecule screen, the method generalizes to unseen perturbations and previously unseen cell types under data-limited settings, achieving higher accuracy for expression variability compared to generative baselines. Attribution analyses identify high-attention genes that complement traditional differential expression analyses, highlighting pathway-specific mechanisms of small-molecule response. By combining scalability, robustness to distribution shifts and interpretability, PrePR-CT enables cell-type-resolved prediction of drug responses, providing a foundation for more precise modelling of cellular perturbations in early drug discovery.
Similar content being viewed by others
Main
Pharmaceutical research and development is constrained by rising costs and declining productivity, resulting in fewer new drugs reaching the market1,2. Artificial intelligence (AI) and generative AI hold great potential for drug development3,4. This includes de novo drug design5, repurposing6 drugs for alternative indications7 and integrating AI techniques into the drug development pipeline8. A core step in these efforts is predicting the effects of chemical compounds. This helps identify therapeutic candidates early, reduces costs and improves safety. Chemical perturbations precisely alter cellular processes, revealing the effects of drugs and their mechanisms of action. However, drug responses vary considerably between cell types and the amount of representative data is limited.
The availability of large-scale chemical perturbation datasets enables robust, interpretable predictions across diverse cell types and experimental conditions. For example, in vitro drug screens, exemplified by high-throughput screening (HTS), allow the simultaneous testing of many molecules against specific biological activities or targets. HTS typically analyses bulk RNA sequencing data to assess transcriptional responses of thousands of molecules9. However, a significant limitation of this approach is its disregard for cell-type-specific responses to individual perturbations. The transcriptional profiles generated by single-cell RNA sequencing methods provide a powerful resource for capturing the heterogeneity of cellular responses to various perturbations, including chemical compounds and CRISPR-based interventions. Nevertheless, the challenge of scaling HTS to single-cell resolution persists, primarily because of the substantial costs and technical complexities of highly multiplexed chemical experiments. Consequently, these experiments typically allow measurement of the effects of a limited number of drugs, usually fewer than a hundred10.
Recent experimental studies11,12,13,14 have developed multiplexed single-cell transcriptional profiling techniques to examine the heterogeneous responses of cancer cell lines to chemical or genetic perturbations. Such datasets motivate machine learning researchers to address the inherent experimental limitations by designing generative in silico models that can produce single-cell gene expression profiles for previously unobserved perturbations. Existing models15,16 leverage generative approaches to map the data into a lower-dimensional latent space, subsequently employing simple vector arithmetic operations within this space to extrapolate the response of a specific drug to a different cell type. Uncovering the latent space has also proved effective in drug target discovery17,18. However, a prominent challenge with generative models is their requirement for substantial data quantities to learn distribution parameters and generalize effectively to unseen conditions. One solution16 involves applying transfer learning in conjunction with bulk RNA sequencing data, such as connectivity map transcriptional data9. However, transfer learning restricts the feature space in the training set to the genes shared with the source domain dataset, which may not include the cell-type-specific marker genes.
On the other hand, recent studies have explored using prior information, such as gene–gene interaction networks, to predict cellular responses to unseen perturbations19. These models demonstrate that leveraging relational graphs yields significant improvements in predicting cellular responses. However, they ignore the cell-type-specific differences between treatment and control groups. Kana et al.20 compared these differences in the principal component analysis space and showed significant variability between cell-type centroids in magnitude and direction, suggesting that the cell-type-specific context is crucial for predicting perturbation responses. Previous methods, such as GEARS19 and chemCPA16 demonstrate strong potential for generalizing across perturbations but face challenges in generalizing to out-of-distribution cell types. Introducing a cell-type-specific context as an inductive bias into model design could overcome this limitation, supporting broader generalization across diverse biological contexts21. For example, a recent study constructed cell-type-specific protein–protein interaction networks from single-cell data to support drug discovery22, while another integrated protein–protein interactions or gene regulatory networks inferred from transcriptional data with drug target information and disease driver gene sets to identify target genes that optimally shift the cellular state towards a desired post-perturbation profile23.
To this end, we invoke an inductive prior, such as cell-type-specific co-expression networks, to predict the mean and variance of post-perturbation expression for differentially expressed genes (DEGs) in unseen cell types. We evaluate the performance of our method across five small-scale single-cell perturbation datasets and one large-scale chemical screening experiment. Additionally, we conduct a comparative analysis with two state-of-the-art generative models, focusing exclusively on chemical perturbations. Our results demonstrate the effectiveness of our approach for predicting DEG expression across diverse cellular contexts. We utilize the graph attention network (GAT) algorithm24 to extract meaningful features from the cell-type-specific networks. To enhance interpretability, we analyse the attention values learned by the GAT layers for each gene, revealing their capability to capture non-trivial patterns from the data.
Results
PrePR-CT predicts single-cell perturbation responses using cell-type-specific graphs
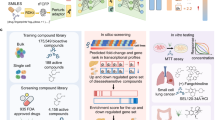
Here we introduce PrePR-CT (Predicting Perturbation Responses in Cell Types), a graph-based deep learning method integrating cell-type-specific co-expression networks with single-nucleus RNA sequencing data to predict transcriptional responses to previously unseen chemical perturbations (Fig. 1). PrePR-CT leverages cell-type-specific co-expression patterns as an inductive bias to capture the fundamental cell-type expression patterns, allowing it to generalize to unseen cell types. Such co-expression patterns are modelled as graphs, enabling the generation of cell-type feature vectors using GATs.
PrePR-CT learns cell-type-specific gene co-expression graphs from a batch of training samples using GAT layers, integrates them with unperturbed control gene expression values and pre-defined perturbation embeddings, and processes them through multilayer perceptron (MLP) layers to predict the expression response to a perturbation. Different cell types are colour-coded along the rows of the gene expression value and feature matrices as well as the cell-type graphs (Methods).
Single-cell perturbation datasets often contain sufficient cells in the control state to permit the use of metacells, here SEACells25, to derive more robust expression estimates and to mitigate the imbalance with the less frequent perturbed state cells. As matched pairs of expression measurements before and after perturbation are technically not feasible, we used SEACells to match control and perturbed cells in single-cell datasets (Methods). For bulk RNA sequencing data, we employed either random assignments19 or selected pairs with minimal Euclidean distance in gene expression between the control and perturbed samples.
Furthermore, we incorporated chemical structure information into the model by generating molecular embeddings from SMILES representations. These chemical embeddings are then combined with the basal cell-type-specific transcriptional profile embeddings. By jointly modelling these complementary representations, the deep neural network learns the relationship between chemical structures and their transcriptional effects, enabling the accurate prediction of gene expression changes following chemical perturbations.
We evaluated the performance of our method across five distinct datasets representing diverse cell types and perturbations: human peripheral blood mononuclear cells (PBMCs) from Kang et al.26, the multi-individual PBMC dataset (NeurIPS) from the Open Problems in Single-cell competition27, a diverse set of multiplexed human cancer cell lines with 18 perturbations in the McFarland dataset11, another set of human cancer cell lines with four perturbations in the Chang dataset12 and a mouse liver dataset containing 11 cell (sub)types from Nault et al.28 (see Methods for details). Leave-one-out cross-validation was used across cell types, where we removed the test cell-type graph during training and evaluated the predicted perturbation response for that cell type. When multiple perturbations were present in a dataset, compounds were stratified by E-distance10 and those with maximal and minimal transcriptional effects were selected, thereby ensuring a comprehensive assessment across varying perturbation strengths. We benchmarked our model against scGen15 and the compositional perturbation autoencoder (CPA)29 for single-perturbation settings and against chemCPA16 for multi-perturbation settings. In addition to the unseen cell type and unseen perturbation evaluations, we assessed our method under a random splitting protocol on the chemical-protein–protein interaction dataset23. This setting follows a different evaluation protocol from the out-of-distribution experiments and provides a complementary assessment of model performance (Supplementary Fig. 1).
PrePR-CT predicts perturbation responses in unseen cell types
We tested PrePR-CT’s ability to predict the transcriptional response of an unseen cell type to a known compound in five datasets, using the transcriptional response of another cell type to the testing compound as training data.
PrePR-CT accurately predicts both the mean expression levels (coefficient of determination (R2) > 0.90) and expression variability (R2 > 0.70) of PBMC cell types in response to a single dose of IFNβ (Kang dataset) (Fig. 2a,b). Prediction accuracy improves further when focusing on the top 100 DEGs that are often central to biological studies. Accordingly, predicted cell profiles colocalize with observed cells in the Uniform Manifold Approximation and Projection (UMAP) space (Fig. 2c), and PrePR-CT consistently outperforms scGen and CPA in both metrics (Fig. 2d). It also demonstrates lower R2 variability across cell types, indicating more stable predictions. To further dissect model performance, genes were stratified by transcriptional change upon perturbation in B cells (Fig. 2e). R2 scores were high for all genes (0.91), highly expressed genes (0.93), upregulated DEGs (0.99) and downregulated DEGs (0.84), and only negative for lowly expressed genes (R2 = −1.83), which likely indicates only a very weak or absent transcriptional response to the perturbation. Overall, PrePR-CT demonstrates robust modelling of transcriptional responses, especially for DEGs, which are essential for interpreting cell-specific perturbation effects.
a, UMAP visualization of cell types stimulated with IFNβ (n = 24,249 cells), as demonstrated in Kang et al. b, R2 values comparing the predicted versus observed mean (left) and standard deviation (right) of gene expression across seven cell types. Bars show the mean R2 over n = 100 bootstrap iterations with ± standard deviation error bars; black dots indicate individual bootstrap iterations. Results are shown for all genes and the top 100 DEGs of the held-out test cell type. c, UMAP visualization comparing predicted (pred) and observed samples of stimulated and control B cells (n = 3,789 cells). d, Comparison of R2 scores for the predicted mean and standard deviation of gene expression across CPA, scGen and PrePR-CT under the leave-one-cell-type-out setting (IFNβ perturbation). For each metric and gene set, R2 values are reported across the held-out cell types (n = 7). Box plots show the median (centre line) and interquartile range (25th–75th percentiles; box); whiskers extend to 1.5× the interquartile range. e, Scatter plots of predicted versus observed differential expression (mean stimulated − mean control) in B cells across gene subsets: all genes in the B cell graph (n = 3,176), highly expressed genes (n = 1,588), lowly expressed genes (n = 1,588), upregulated DEGs (n = 43) and downregulated DEGs (n = 43). Each point represents one gene. Red lines show linear regression fits and shaded areas indicate the 95% confidence intervals. R2 values are shown in each panel.
To extend the range of perturbations and cell types, we utilized the NeurIPS PBMC dataset, containing 144 small-molecule perturbations of four distinct immune cell types (Fig. 3a). This dataset structure allowed the use of cross-validation and perturbation stratification by transcriptional response strength (Fig. 3b). As illustrated in Fig. 3c, drugs with small transcriptional impact were easier to predict, allowing all models to perform well in estimating mean gene expression and variance across cell types. Nevertheless, PrePR-CT consistently achieved the highest R2 scores, particularly among the top 100 DEGs, whereas CPA and scGen showed greater variability, particularly in prediction variance.
a, UMAP visualization of cells coloured by condition (top) and cell type (bottom) across multiple drug perturbations (n = 291,793). b, E-distance between control and perturbed cells for each compound (n = 135 drugs), used to identify perturbations inducing large versus small transcriptional responses. c, R2 scores for predicted versus observed gene expression mean (left) and standard deviation (middle) across cell types for penfluridol, flutamide and SCH-58261, evaluated using all genes and the top 100 DEGs of the held-out test cell type. Each point represents one cell-type drug evaluation. The box plot (right) summarizes the performance for penfluridol across available cell types (n = 3; NK cells excluded). The centre line indicates the median and the box bounds indicate the upper and lower quartiles. Whiskers extend to the largest and smallest values within 1.5× the interquartile range. d, The same as in c, but for scriptaid, ketoconazole and dactolisib. The box plot (right) summarizes the performance for dactolisib across evaluated cell types (n = 4). e, UMAP visualization comparing predicted and observed perturbed cells for penfluridol (left; n = 4,914) and dactolisib (right; n = 4,814), shown alongside control cells in B cells.
When considering drugs that induce the largest observed transcriptional shifts, model performance becomes more variable, with a general decline in predictive accuracy (Fig. 3d). PrePR-CT maintains the highest performance, particularly in capturing gene expression variance, demonstrating its robustness under complex perturbation conditions. Again, UMAP embedding of predicted and observed B cells yields robust co-localization for two representative drugs (Fig. 3e): penfluridol, a mild perturbation (Fig. 3e, left), and dactolisib, a strong perturbation (Fig. 3e, right). For penfluridol, the predicted perturbed cells (blue) closely overlap with both the control population (grey) and the observed perturbed cells (red), indicating that the model effectively captures subtle transcriptional changes. In contrast, dactolisib-treated cells form a distinct cluster in which the predicted cells remain well-aligned with the true responses. Natural killer (NK) cells were excluded from the mild perturbation analysis (Supplementary Fig. 2) due to minimal transcriptional response and low DEG counts, resulting in low or negative R2 scores. This exclusion was justified because the absence of a strong biological signal limited the ability to make meaningful predictions. Notably, PrePR-CT did not overfit to noise in these cases, reflecting a desirable inductive bias: the model responds only when a true signal is present, rather than attempting to extract patterns from background variability. This behaviour aids its generalizability and biological relevance.
Supplementary Figs. 3 and 4 present the results for the Nault28 and McFarland11 datasets. In the Nault dataset, cells cluster distinctly by both cell type and treatment condition (Supplementary Fig. 3a), reflecting strong perturbation-driven transcriptional shifts. The obtained model performance varied substantially across cell types but remained consistent for very similar populations, such as central and portal hepatocytes (Supplementary Fig. 3b). As most DEGs in the dataset were downregulated, the model more effectively captured these changes (R2 = 0.93 for downregulated genes; Supplementary Fig. 3e). In the McFarland dataset, we found predictions were easier for drugs with minimal effects, with a decline in R2 for compounds that induced substantial transcriptional changes (Supplementary Fig. 4).
We further evaluated PrePR-CT generalization by training on the NeurIPS dataset and testing on each tissue type within the Tahoe-100M30 dataset, considering only shared compounds and highly variable common genes (Supplementary Note 1), again predicting transcriptional effects in cell types not seen during training.
To better understand the factors that influence generalization performance, we analyse the results with respect to two key variables: (1) the strength of the transcriptional response induced by each drug measured by E-distance and (2) the transcriptional similarity between training and testing cell types via Euclidean distance. At the cell-type level, a very weakly negative correlation (r = −0.26) between median perturbation E-distance to control over all considered drugs and corresponding median DEG prediction R2 indicates that cell lines that respond more strongly to perturbations are not inherently harder to predict (Supplementary Fig. 6). In contrast, at the drug level, stronger median responses for all cell types yield slightly more predictable effects (r = 0.53, and r = 0.38 when removing the outlier). Finally, when examining generalization across cell lines, we found a stronger negative correlation (r = −0.60) between the median Euclidean distance (from each Tahoe cell line to the NeurIPS training cell types) and the corresponding R2 scores, illustrating the difficulty of out-of-distribution generalization as dissimilarity from the training data increases.
We benchmarked PrePR-CT against scGen, chemCPA and a mean expression baseline in two settings: <Tahoe controls excluded, strict out-of-distribution> and <Tahoe controls included>. scGen applies only to the latter, as it requires control cells from the test cell types during training, whereas PrePR-CT and chemCPA apply to both. While the mean baseline outperformed PrePR-CT and chemCPA for most cell lines in the first setting, PrePR-CT performed better in the second setting (Supplementary Fig. 7), demonstrating better generalization with the control data.
PrePR-CT predicts the response to unseen perturbations
We also assessed the model’s ability to predict the effect of unseen compounds within a known cell type using the Chang dataset, which includes three structurally similar small molecules (Fig. 4a). We adopted a leave-one-compound-out strategy analogous to the cell-type generalization setting.
a, Molecular structures of the three evaluated compounds: GNE-104, GNE-069 and erlotinib. b, R2 scores for the predicted versus observed gene expression mean (left) and standard deviation (right) across compounds, evaluated on all genes and the top 100 DEGs. Bars show the mean R2 over n = 100 bootstrap iterations, and error bars denote ± standard deviation. c, Comparison of chemCPA and PrePR-CT performance across four metrics: mean and standard deviation of gene expression for all genes and the top 100 DEGs. Each box plot summarizes the performance across all evaluated compound cell-type test cases (n = 3 compounds). Centre lines indicate the median and boxes the interquartile range (25th–75th percentiles) and whiskers extend to 1.5× interquartile range. d, UMAP visualization of control, observed perturbed and predicted erlotinib-treated cells, showing strong alignment between predicted and observed transcriptional responses (n = 34,154 cells).
Despite the structural similarity among the compounds, PrePR-CT accurately captured the mean transcriptional response across all genes and the top 100 DEGs, achieving consistently high R2 scores (Fig. 4b, left). For gene-wise variance predictions, however, performance differed across compounds: while GNE-104 and GNE-069 showed similar accuracy, erlotinib achieved a markedly higher R2 score, especially among the top 100 DEGs (Fig. 4b, right). As scGen does not apply to unseen perturbations, we benchmarked PrePR-CT against chemCPA. PrePR-CT outperformed chemCPA in predicting gene expression variance, particularly for highly responsive genes. UMAP projections of erlotinib-treated cells showed that the predicted profiles closely match the observed perturbed cells and separate from the controls, confirming the ability of PrePR-CT to accurately capture drug-induced transcriptional shifts (Fig. 4c,d).
Next, we extended the evaluation of unseen perturbations to the LINCS dataset, which contains bulk transcriptomic profiles9. From this dataset, we constructed a subset of cell lines and perturbations based on the sample size distribution (Methods and Supplementary Fig. 8). In this setting (Supplementary Fig. 8b,c), both PrePR-CT and chemCPA achieved comparable performance in predicting mean DEGs; however, PrePR-CT outperformed chemCPA in predicting the variance of DEGs across all cell lines.
PrePR-CT operates successfully in a small-data regime
A recurring challenge in perturbation-response modelling arises from the different ways experimental setups using limited-size single-cell datasets handle their limited size, requiring a balance between the number of conditions (usually pairs of cell type and treatment) and the resulting number of cells per condition. This typically yields (1) datasets with one cell type and few perturbations, (2) datasets with one perturbation and few cell types, both yielding many cells per perturbation, and (3) datasets containing many cell types and perturbations, yielding a dramatically lower number of cells per pair. These scenarios pose different obstacles to learning perturbation effects, particularly when the variability of the transcriptional response must be estimated from only a few perturbed cells. To systematically evaluate the behaviour of PrePR-CT under conditions similar to (1) and (2), we used subsets of varying sizes from the already small Kang PBMC dataset, consisting of 24,249 cells, for training. The model consistently converged, even when the subsampling rate was aggressively reduced to 20% of the total cells.
Mean expression and variability R2 across all genes and the top 100 DEGs show that PrePR-CT consistently outperforms CPA across all subset sizes (Fig. 5a,b). While CPA’s R2 scores for mean expression remain relatively stable across dataset sizes, PrePR-CT’s performance declines when the dataset is reduced to below 60% (~15,000 cells). For both methods, R2 for mean expression is higher for the top 100 DEGs than for all genes, likely reflecting the lower variability expected at higher gene expression levels.
a,b, R2 scores for predicting the mean (a) and standard deviation (b) of post-perturbation gene expression in B cells as a function of training set size on the Kang dataset. Results are shown for all genes in the B cell graph (n = 3,176) and the top 100 DEGs for both CPA and PrePR-CT. Each point represents the mean performance across n = 3 independent runs with different random seeds; error bars denote ± standard deviation across the three runs. PrePR-CT consistently outperforms CPA across all training set sizes, demonstrating robust performance even in the small-data regime.
In contrast, both methods exhibit a similar decline in the quality of variability predictions in datasets with fewer than ~15,000 cells. However, CPA’s variability estimates are more precise when considering all genes, while PrePR-CT’s estimates are consistently more precise for the top 100 DEGs. This difference is likely attributable to the use of SEACells to match control and treatment conditions, which enables balanced representation across cellular clusters and thereby captures the full landscape of the control population.
Graph attention network learns meaningful representations
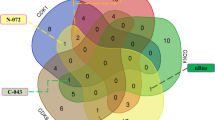
We analysed attention maps to assess the GAT network’s ability to capture meaningful patterns. High-attention genes (HAGs) were identified by summing each gene’s outgoing edge weights and ranking them accordingly. This analysis was performed on cell-type-specific graphs from the Kang dataset. GAT identifies distinct gene sets across different cell types, and the analysis of HAGs suggests that GAT extracts complementary features compared to DEG analysis. To explore the relationship between these two approaches, we compared the union of the top 50 HAGs across all PBMC cell types with that of the top 50 DEGs, yielding 333 and 130 genes for HAGs and DEGs, respectively. Of the genes, 70.5% are uniquely HAGs (Fig. 6a), indicating that a significant number of HAGs are not differentially expressed. Similarly, 24.5% are uniquely DEGs, demonstrating that not all DEGs are required for the prediction. We observed only a 5% overlap between HAGs and DEGs in PBMCs, suggesting that these sets captured complementary signals. HAGs exhibit greater cell-type specificity, while DEGs are more commonly shared, highlighting the ability of GATs to identify unique biological features and to offer complementary insights to traditional differential expression in understanding gene function. Next, we investigated the set of unique HAGs through pathway enrichment analysis and attention scores for ranking. The analysis yielded significant ontology terms, including T cell activation and immune system pathways (Fig. 6b), which align with the known mechanism of action of IFNβ within the immune system, particularly its effects on T cells. Furthermore, the existing literature supports roles for a range of HAGs in the considered perturbations. For example, CD74 is among the top 20 HAGs identified in dendritic cells, and Nguyen-Pham et al.31 demonstrated that CD74 negatively regulates dendritic cell migration, underscoring its relevance in the design of protocols aimed at enhancing dendritic cell migratory capacity in immunological research. LCK, LAT and CD3E are among the top 10 HAGs in CD4+ T cells and are members of immune system pathways, including T cell receptor signalling and T helper 17 cell differentiation. Subsequently, we examined the associations between HAGs and IFNβ-related pathways. We found that XAF1, ISG20, STAT1 and RSAD2 are involved in multiple interferon-related pathways, including the cellular response to type I interferon (GO:0071357) and the type I interferon signalling pathway (GO:0060337). These genes are among the top DEGs in response to IFNβ in the Kang dataset. This finding suggests that the GAT model effectively attends to the most critical features within the gene set.
a, Venn diagram showing the overlap between HAGs and DEGs. b, Gene set enrichment analysis of HAGs reveals a strong association with immune activation pathways, including T cell activation, cytokine signalling and antigen receptor signalling. c, R2 scores before and after random shuffling of cell-type-specific features across multiple cell types, evaluated using all genes (n = 3,176) and the top 100 DEGs. Data are shown as mean ± standard deviation over five independent random shuffling runs (different random seeds). d, R2 scores before and after masking HAGs from the input features.
To further examine the ability of GATs to attend to meaningful biological features, we investigated their capability to learn the correlation structures defining the edges in cell-type-specific graphs. To this end, we computed Spearman’s correlation between the number of neighbours and the attention rank for the genes in each cell type. The results revealed a minimum correlation of 0.68 in B cells and a maximum of 0.91 in FCGR3A+ monocytes. These correlation values across different cell types suggest that GATs effectively learn the network structure of correlations within cell-type-specific graphs. To assess the importance of features learned by the GAT model, we performed ablation studies by disrupting these features, measuring R2 changes for mean gene expression prediction under different perturbations as a proxy for predictive degradation (Fig. 6c,d). First, we randomized cell type features while preserving the overall graph structure to assess the importance of cell-type-specific information (Fig. 6c). Second, we masked the features of HAGs at test time to evaluate their direct contribution to prediction performance (Fig. 6d). These perturbations allowed us to assess the relative importance of global cell-type features and HAG-derived signals in supporting accurate predictions. Shuffling cell-type features resulted in a pronounced drop in R2 values (Fig. 6c), particularly for the top 100 DEGs, compared to the complete set of genes. While the model retained high performance across all genes before shuffling (solid lines), disrupting the alignment between cell-type features and graph structure led to a more significant reduction in prediction accuracy for DEGs (dashed pink line) across most cell types. This finding suggests that DEGs are more sensitive to the integrity of cell-type features, likely due to their stronger responsiveness to perturbations, which becomes obscured when the input feature-cell-type correspondence is disrupted. Masking HAG features at test time also led to a noticeable, albeit smaller, decrease in R2 values (Fig. 6d). Again, the top 100 DEGs (dashed pink) exhibited a greater decline in predictive performance than all genes (dashed blue), although overall model performance remained relatively high. This indicates that HAGs contribute disproportionately to the accurate prediction of perturbation-responsive genes, and their removal selectively impairs the model’s ability to reconstruct DEG expression. The model appears more robust to disruption when using only the top 100 DEGs, which likely reflects that DEGs capture more informative and biologically relevant variation, whereas using all genes introduces noise from lowly expressed or uninformative genes, making the model more susceptible to degradation when features are shuffled or masked.
These findings highlight the importance of cell-type features and HAG-derived representations in shaping the GAT model’s performance. DEGs are more sensitive to feature disruptions, supporting their role as key indicators of perturbation effects.
Discussion
We used a gene co-expression network-based cell type representation as an inductive bias to predict cell-type-specific responses to chemical perturbations. This strategy effectively predicted the mean and variance of post-perturbation expression for DEGs across different cell types and chemical perturbations, using leave-one-out cross-validation within each dataset to evaluate out-of-distribution scenarios. By analysing the salient features of GAT-generated cell-type-specific networks and explicitly focusing on genes with HAGs, we identified cell-type markers and perturbation-related genes. This approach offered a complementary perspective by identifying features beyond those examined in differential gene expression analysis. It is essential to note that DEG analysis tends to be more stable than HAG analysis, as HAG genes exhibit lower expression levels. Furthermore, HAG genes are more sensitive to training parameters, such as the number of attention heads in the GAT layers. These findings underscore the need to assess the stability of the model’s interpretations by evaluating their consistency across conditions. Additionally, it is essential to investigate how various model selection criteria impact the interpretability of deep learning models.
The results of our benchmarking against state-of-the-art generative models highlight the advantage of integrating cell-type-specific inductive bias to predict gene expression in response to perturbations. Importantly, our method performs well in the small-data regime, where data scarcity often limits the efficacy of alternative approaches. Moreover, the direct optimization of the Earth Mover’s Distance (EMD; Wasserstein) yields better results than reference generative models, underscoring the robustness and predictive accuracy of our approach. These results demonstrate that the method remains highly effective across diverse single-cell perturbation datasets, emphasizing its broad applicability to single-cell data analysis, particularly in scenarios with limited data availability. In addition, recent benchmarking studies32,33,34 comparing deep learning methods with linear models have investigated why simple linear baselines often outperform deep learning approaches. These studies highlight the issue of control bias and underscore the need to design deep learning methods, evaluation frameworks and metrics that can effectively distinguish between systematic variation and perturbation-specific responses.
One limitation of the GAT approach is that the graph node features model only the mean and variance of gene expression. A promising direction for future work would be to incorporate additional features, such as cell cycle states, morphological traits and tissue of origin, to improve the biological relevance of the learned representations. Expanding the method to encompass a broader range of cell types would further enhance its generalizability, enabling it to capture diverse cellular contexts more effectively and increase predictive accuracy, particularly in challenging scenarios. Our transfer learning experiment using the NeurIPS and Tahoe datasets highlights several challenges in generalizing across different experiments. We found that the transcriptional distance between training and testing cell types significantly impacts model performance, an observation aligned with current experience in machine learning35,36. We also observed that the sets of highly variable genes vary across datasets, complicating feature selection. Drugs with strong perturbation effects partly offset this mismatch by generating stronger, more predictable signals. Finally, differences in experimental covariates, such as batch effects or protocol discrepancies, can mask true effects and confound interpretation. Addressing these limitations in future work may lead to the development of more robust and generalizable models for predicting cellular responses to perturbations. The proposed input representation could offer a potential for various drug discovery applications, including predicting drug combination effects, identifying therapeutic targets and exploring interactions between different perturbations.
In summary, we have developed a method that shows strong performance in a small-data regime. This outcome resulted from incorporating an appropriate inductive, cell-specific network bias, which simultaneously enables mechanistic interpretation via the attention layers. This line of research may open the possibility of developing quantitative cell-specific systems biology models37,38, which could fuel predictive biology39.
Methods
PrePR-CT considers a perturbational single-cell dataset \(D=\{({x}_{i},{y}_{i},{g}_{i}){\}}_{i=1}^{N}=\{({x}_{i},{y}_{i},({c}_{i},{p}_{i})){\}}_{i=1}^{N}\), where \({{\bf{x}}}_{i}\) and \({{\bf{y}}}_{i}\in {R}^{m}\) are m-dimensional gene expression vectors representing, respectively, a control cell and its transcriptional response to a perturbation \({g}_{i}\). Each perturbation \({g}_{i}\) comprises a cell-type label \({c}_{i}\in C\) and a \(k\)-dimensional embedding vector \({{\bf{p}}}_{i}\). We associated each cell type with one of the cell-type-specific graphs \(C=\{{G}_{{c}_{1}},{G}_{{c}_{2}},\ldots ,{G}_{{c}_{M}}\}\), where \(M\) is the number of cell types in \(D\). We predicted the perturbation effect as a function of the cell-type graph, a set of control samples of the same cell type and a perturbation embedding.
Cell-type-specific graphs
In PrePR-CT, each cell type is represented as a graph, with genes as nodes and edges between genes that exhibit sufficiently correlated expression patterns. Smaller disconnected submodules were retained to capture cell-type-specific regulatory patterns while maximizing the overlap of shared genes across cell types. Consistent with the goal of predicting responses to unseen perturbations, we constructed cell-type graphs using expression data from unperturbed (control) cell states, as perturbation-induced measurements were not available during testing. Moreover, these cell-type graphs can contain different gene sets to reflect each cell type’s distinct expression and functions.
The co-expression graphs were constructed by aggregating single-cell expression values of unperturbed cells into metacells using SEACells25 to mitigate the inherent sparsity of single-cell expression data. We calculated the set of 5,000 highly variable genes (HVGs) before the metacell aggregation. To construct the graph for a given cell type \({G}_{{c}_{i}}\), all pairwise correlations between HVGs were computed using only metacells of the selected cell type. Edges with correlation strengths below the 99th percentile of the absolute correlation distribution were removed.
The resulting co-expression graphs included varying numbers of genes based on the defined threshold, with some genes shared across cell types. The selected parameters depended on the processed data and constituted a trade-off between considering the most specific genes per cell type, retaining similar genes between the cell-type graphs and controlling the graph’s size, which affects the time complexity of the training. The graph’s feature matrix was initialized using the mean and standard deviation of the control state to capture the basal expression profile.
Matching control and perturbed cells in single-cell data
Because single-cell transcriptomic assays are destructive, paired measurements from the same cell before and after perturbation were not available, necessitating a strategy to match the control and perturbed cells. Within each cell line, we randomly assigned control-state cells to perturbed cells to avoid bias arising from systematic pairing. As the datasets contained substantially more control than perturbed cells, this procedure inherently subsampled the control population. To ensure that the subsampled controls remained uniformly distributed across the transcriptional manifold, we guided the selection using metacell clusters generated by SEACells. These metacells provided a coarse-grained representation of control-state variation and allowed uniform sampling independent of local differences in cell density. For each cell type and perturbation, we drew an equal number of control cells from the metacells and assigned them to the perturbed cells for training. Gene-expression samples were used as inputs and outputs; SEACell clusters served solely as a tool to align the control and perturbed groups.
Chemical structure representation of the perturbing compound
We used SMILES strings to encode the chemical structures of the perturbing compounds. SMILES provides a compact, machine-readable format for describing molecular structures and serves as input for generating molecular embeddings. We used chemical descriptors computed via the RDKit package, a widely used cheminformatics toolkit, to extract structural and physicochemical features from each SMILES string. This descriptor-based approach captured relevant molecular properties, such as atom types, bond connectivity, partial charges and topological features, which were embedded into fixed-length vectors representing each compound. These embeddings provided structured molecular features that could be integrated with transcriptional response data, allowing our model to predict drug effects across different cell types and conditions. We obtained the SMILES strings for all chemical perturbations from the DrugBank41 database or the original publications of the corresponding datasets.
Graph attention network
The GAT24 defines an attention mechanism on the graph structure by computing an attention score \({\alpha }_{i,j}\) between a given node i and each node \(j\) in its first-order neighbourhood set \({N}_{i}\). This score represents the significance of node \(j\)’s features to node i and is calculated as:
where \({{h}^{\to }}_{i}\) is a feature representation of node i transformed using the weight matrix \(W\), and \({\bf{a}}\) is a learnable weight vector. The self-attention mechanism is then extended to multi-head attention by applying \(L\) independent mechanisms, and their outputs are either concatenated or averaged:
The final output \({{h}^{\to }}_{i}^{{\prime} }\) is a high-level feature representation of node i in the graph. We employed GATs in this project because they showed successful examples of generalizing completely unobserved graphs during the training process24.
Loss function
The mean squared error (MSE) is commonly used as a loss function in regression models. However, MSE primarily focuses on the mean and does not consider the overall distribution of the actual and predicted samples. In contrast, the Wasserstein distance, or EMD, is an optimal transport measure for comparing probability distributions with disjoint supports42. The Wasserstein distance has achieved significant success in the context of generative adversarial networks, leading to more stable training and improved sample quality by providing a more meaningful and smoother gradient signal compared to traditional generative adversarial networks43. To approximate the EMD, we seek the optimal transportation cost \(\gamma\):
where \(M\) is a cost matrix corresponding to the Euclidean distances between the actual and predicted samples, and \({\mathbf{a}}\) and \({\mathbf{b}}\) are the sample weights of the source and target distributions, respectively44,45.
For the implementation, we used the mini-batch EMD function implemented by Flamary et al.46 to solve the aforementioned optimization problem. The optimal transport library in Python offers several tutorials for using this loss function46.
The final loss is defined as:
where \(T\) is the number of cell-type–perturbation pairs \({g}_{i}\) in the current batch, and \({Y}_{t}\) and \({\hat{Y}}_{t}\) denote the observed and predicted samples, respectively.
Implementation details
Utilizing a cell-type-specific gene interaction network requires mapping each gene expression sample to the graph of its respective cell type. However, this direct mapping substantially increases the training time complexity. To address this issue, we adopted an alternative formulation. First, we extracted the graphs within each mini-batch and embedded them using the graph attention layers. We then mapped the samples to the features generated by these layers according to their cell-type labels. This avoided treating each cell as an independent graph and thereby reduced computational complexity. Figure 1 demonstrates the training steps.
The model required tuning six hyperparameters during training. Supplementary Table 2 lists these parameters along with the values optimized throughout the training process using the hyperparameter optimization tool Optuna18 on a fixed random validation set. Optimizer47 was used for model training, and no activation function was applied to the output layer.
Evaluation
We evaluated model performance using the coefficient of determination (R2), which measures the agreement between the observed statistics (mean and standard deviation) and their predicted values. Our analysis focused explicitly on gene expression statistics rather than overall gene expression profiles. This distinction is crucial because other methods included in the comparative analysis map all control cells to the perturbed state, resulting in an unequal number of predicted samples compared to the actual perturbed samples in the ground truth data.
Consequently, we concentrated on capturing the distinct shifts from the control state to the perturbed state. The intentional choice of R2 as our evaluation metric offers notable advantages over alternative metrics such as Pearson correlation and MSE. Unlike Pearson correlation, which measures only linear relationships and overlooks magnitude considerations, R2 provides a more comprehensive assessment of predictive power. This deliberate selection aligned with our objective of capturing nuanced changes in gene expression.
For the gene set selected for evaluation, we assessed all the genes in the graph for the test cell type and the top 100 DEGs. The top 100 DEGs were defined using a combined ranking scheme based on the P value and log fold change statistics computed via t-tests using the scanpy Python package. First, we filtered genes with a P value of 0.05 and then applied the rank product statistic:
where \(s=2\) represents the two ranking criteria: P value and log fold change. This ranking aimed to eliminate noisy genes with extreme log fold change values and very small P-values.
Datasets
We assessed the efficacy and versatility of our method across diverse scenarios by evaluating it on multiple datasets encompassing various cell types and perturbations. Specifically, we employed five distinct datasets: human peripheral PBMCs treated with interferon-beta (IFNβ). IFNβ is known for its potent antiviral activity, growth inhibition and immune modulation. It modulates T cell behaviour by reducing activation and proliferation, limiting infiltration into sensitive tissues and promoting regulatory mechanisms, published by Kang et al.26, the McFarland dataset11, the Chang dataset12, the single-dose TCDD liver by Nault et al.28 and the cross-individual dataset (NeurIPS) from the Open Problems in Single-Cell competition27.
The McFarland dataset encompasses 13 chemical perturbations, each with a specific dose value, and comprises over 100 cell lines. We focused on the five cell lines with the largest sample counts and excluded drugs lacking SMILES identifiers in the DrugBank database41.
The Nault dataset entailed single-nucleus RNA sequencing of flash-frozen C57BL6 mouse livers. In this dataset, mice underwent subchronic administration of a specified TCDD dose via oral gavage every 4 days over 28 days. As indicated by Kana et al.20, we excluded all immune cell types, given their propensity to migrate from the lymph to the liver during TCDD administration.
The NeurIPS dataset comprises single-cell gene expression profiles obtained from human peripheral PBMCs treated with 144 compounds from the LINCS Connectivity Map database9. Measurements were conducted 24 hours after treatment, with the experiment replicated using cells from three healthy human donors. The full dataset was utilized, except for drug conditions in any cell type with fewer than 30 associated samples.
The LINCS dataset comprises 1,034,271 samples of 122,974 distinct combinations of 83 cell lines treated with 20,544 different drugs. However, the majority of cell line–drug combinations are only sparsely sampled (Supplementary Fig. 7a). We therefore excluded combinations with fewer than 300 samples to reduce potential bias due to sampling, which yielded 65,585 samples from 57 pairs (21 cell lines, 11 perturbations and DMSO as the control). Control–treatment pairs were assigned randomly between DMSO-treated control cells and perturbed cells for a given cell type, a method that has been shown to be successful previously19.
We processed all the single-cell datasets through a series of steps: removing mitochondrial and ribosomal genes; filtering for cells with at least 1,000 counts and for genes detected in fewer than 50 cells; normalizing the count matrix using scanpy.pp.normalize_total; and applying a \(\log (x+1)\) transformation.
For each dataset, we constructed a set of cell-type-specific graphs. The initial gene–gene correlation matrix contained 5,000 × 5,000 edges. After applying a threshold, we retained between 240,000 and 250,000 edges per cell type. Supplementary Table 1 provides additional details for each dataset.
For the transfer learning experiments, we used a subset of the Tahoe dataset, which initially contained 50 unique cell lines from various tissues and 380 distinct compounds. From this dataset, we selected a subset corresponding to the 18 drugs shared with the NeurIPS dataset. To ensure tissue diversity, we identified the cell line with the highest number of samples from each tissue type and subsampled these selected cell lines for testing. To define a shared gene set for transfer learning, we identified the genes common to both datasets and ranked them based on their variability scores computed using Seurat. We then averaged the ranks across the datasets and selected the top 5,000 genes as a joint feature set for downstream analysis. Supplementary Note 1 provides the pre-processing steps in more detail.
Reporting summary
Further information on the research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All datasets used in this study for reproducibility are publicly available. The Kang dataset is available from the Gene Expression Omnibus (GEO) under accession number GSE96583, the NeurIPS dataset under GSE279945 and the Nault dataset under GSE184506. The McFarland and Chang datasets are available through the scPerturb project (https://projects.sanderlab.org/scperturb/). The LINCS dataset was obtained from https://github.com/theislab/chemCPA and the Tahoe dataset from https://huggingface.co/datasets/tahoebio/Tahoe-100M. In addition, all processed datasets used in this study are available at https://github.com/reem12345/Cell-Type-Specific-Graphs. Source data are provided with this paper.
Code availability
The code used for reproducing all analyses and figures in this study is publicly available at https://github.com/reem12345/Cell-Type-Specific-Graphs. It is also available via Zenodo at https://doi.org/10.5281/zenodo.15255075 (ref. 40).
References
Paul, S. M. et al. How to improve RD productivity: the pharmaceutical industry’s grand challenge. Nat. Rev. Drug Discov. 9, 203–214 (2010).
Ringel, M. S., Scannell, J. W., Baedeker, M. & Schulze, U. Breaking Eroom’s Law. Nat. Rev. Drug Discov. 19, 833–834 (2020).
Gangwal, A. & Lavecchia, A. Unleashing the power of generative AI in drug discovery. Drug Discov. Today 29, 103992 (2024).
Paul, D. et al. Artificial intelligence in drug discovery and development. Drug Discov. Today 26, 80–93 (2021).
Tang, X. et al. A survey of generative AI for de novo drug design: new frontiers in molecule and protein generation. Brief Bioinform. 25, bbae338 (2024).
Pushpakom, S. et al. Drug repurposing: progress, challenges and recommendations. Nat. Rev. Drug Discov. 18, 41–58 (2018).
Cousins, H. C., Nayar, G. & Altman, R. B. Computational approaches to drug repurposing: methods, challenges, and opportunities. Annu. Rev. Biomed. Data Sci. 7, 15–29 (2024).
Tropsha, A., Isayev, O., Varnek, A., Schneider, G. & Cherkasov, A. Integrating QSAR modelling and deep learning in drug discovery: the emergence of deep QSAR. Nat. Rev. Drug Discov. 23, 141–155 (2023).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437–1452 (2017).
Peidli, S. et al. scPerturb: harmonized single-cell perturbation data. Nat. Methods 21, 531–540 (2024).
McFarland, J. M. et al. Multiplexed single-cell transcriptional response profiling to define cancer vulnerabilities and therapeutic mechanism of action. Nat. Commun. 11, 4296 (2020).
Chang, M. T. et al. Identifying transcriptional programs underlying cancer drug response with TraCe-seq. Nat. Biotechnol. 40, 86–93 (2022).
Srivatsan, S. R. et al. Massively multiplex chemical transcriptomics at single-cell resolution. Science 367, 45–51 (2020).
Mathur, L. et al. Combi-seq for multiplexed transcriptome-based profiling of drug combinations using deterministic barcoding in single-cell droplets. Nat. Commun. 13, 4450 (2022).
Lotfollahi, M., Wolf, F. A. & Theis, F. J. scGen predicts single-cell perturbation responses. Nat. Methods 16, 715–721 (2019).
Hetzel, L. et al. Predicting cellular responses to novel drug perturbations at a single-cell resolution. Adv. Neural Inf. Process Syst. 35, 26711–26722 (2022).
de Weerd, H. A. et al. Latent space arithmetic on data embeddings from healthy multi-tissue human RNA-seq decodes disease modules. Patterns 5, 101093 (2024).
Dwivedi, S. K., Tjärnberg, A., Tegnér, J. & Gustafsson, M. Deriving disease modules from the compressed transcriptional space embedded in a deep autoencoder. Nat. Commun. 11, 856 (2020).
Roohani, Y., Huang, K. & Leskovec, J. Predicting transcriptional outcomes of novel multigene perturbations with GEARS. Nat. Biotechnol. 42, 927–935 (2023).
Kana, O. et al. Generative modeling of single-cell gene expression for dose-dependent chemical perturbations. Patterns 4, 100817 (2023).
Velez-Arce, A. & Zitnik, M. PyTDC: a multimodal machine learning training, evaluation, and inference platform for biomedical foundation models. In Proc. 42nd International Conference on Machine Learning 267 (PMLR, 2025).
Li, M. M. et al. Contextual AI models for single-cell protein biology. Nat. Methods 21, 1546–1557 (2024).
Gonzalez, G. et al. Combinatorial prediction of therapeutic perturbations using causally inspired neural networks. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-025-01481-x (2025).
Veličković, P. et al. Graph attention networks. In Proc. International Conference on Learning Representations (ICLR, 2018).
Persad, S. et al. SEACells infers transcriptional and epigenomic cellular states from single-cell genomics data. Nat. Biotechnol. 41, 1746–1757 (2023).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
Szałata, A. et al. A benchmark for prediction of transcriptomic responses to chemical perturbations across cell types. In Advances in Neural Information Processing Systems 37 (NeurIPS, 2024).
Nault, R. et al. Benchmarking of a Bayesian single cell RNAseq differential gene expression test for dose–response study designs. Nucleic Acids Res. 50, e48 (2022).
Lotfollahi, M. et al. Predicting cellular responses to complex perturbations in high-throughput screens. Mol. Syst. Biol. 19, e11517 (2023).
Zhang, J. et al. Tahoe-100M: a giga-scale single-cell perturbation atlas for context-dependent gene function and cellular modeling. Preprint at bioRxiv https://doi.org/10.1101/2025.02.20.639398 (2025).
Nguyen-Pham, T.-N. et al. Type I and II interferons enhance dendritic cell maturation and migration capacity by regulating CD38 and CD74 that have synergistic effects with TLR agonists. Cell Mol. Immunol. 8, 341–347 (2011).
Viñas Torné, R. et al. Systema: a framework for evaluating genetic perturbation response prediction beyond systematic variation. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02777-8 (2025).
Mejia, G. M. et al. Diversity by design: addressing mode collapse improves scRNA-seq perturbation modeling on well-calibrated metrics. Preprint at https://arxiv.org/abs/2506.22641 (2025).
Ahlmann-Eltze, C., Huber, W. & Anders, S. Deep-learning-based gene perturbation effect prediction does not yet outperform simple linear baselines. Nat. Methods https://doi.org/10.1038/s41592-025-02772-6 (2025).
Liu, J. et al. Towards out-of-distribution generalization: a survey. Preprint at https://arxiv.org/abs/2108.13624 (2021).
Ye, N. et al. OoD-Bench: quantifying and understanding two dimensions of out-of-distribution generalization. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 7947–7956 (CVPR, 2022).
Cichonska, A. et al. Computational-experimental approach to drug-target interaction mapping: a case study on kinase inhibitors. PLoS Comput. Biol. 13, e1005678 (2017).
Tegnér, J. N. et al. Computational disease modeling–fact or fiction? BMC Syst. Biol. 3, 56 (2009).
Zenil, H. et al. An algorithmic information calculus for causal discovery and reprogramming systems. iScience 19, 1160–1172 (2019).
Reem, A. Cell-type-specific-graphs: predicting and interpreting cell-type specific drug responses in the low data regime using inductive priors (v1.0.0). Zenodo https://doi.org/10.5281/zenodo.15255075 (2025).
Wishart, D. S. et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Drugbank https://www.drugbank.ca/ (2006).
Genevay, A., Peyre, G. & Cuturi, M. Learning generative models with sinkhorn divergences. In 21st International Conference on Artificial Intelligence and Statistics (eds Storkey, A. & Perez-Cruz, F.) 1608–1617 (PMLR, 2018).
Arjovsky, M., Chintala, S. & Bottou, L. Wasserstein generative adversarial networks. In Proc. 34th International Conference on Machine Learning (eds Precup, D. & Teh, Y. W.) 214–223 (PMLR, 2017).
Panaretos, V. M. & Zemel, Y. Statistical aspects of Wasserstein distances. Annu. Rev. Stat. Appl. 6, 405–431 (2019).
Bonneel, N., Van de Panne, M., Paris, S. & Heidrich, W. Displacement interpolation using Lagrangian mass transport. ACM Trans. Graph. 30, 158 (2011).
Flamary, R. et al. POT: python optimal transport. J. Mach. Learn. Res. 22, 1–8 (2021).
Kingma, D. P. & Ba, J. L. Adam: a method for stochastic optimization. In Proc. 3rd International Conference on Learning Representations (ICLR, 2015).
Acknowledgements
King Abdullah University of Science and Technology supported this work. N.A.K. was supported by the European Union’s Horizon Europe Research and Innovation Actions under grant agreement No. 101137154 (WISDOM). We thank all members of the lab for constructive critical reading.
Author information
Authors and Affiliations
Contributions
R.A., N.A.K., R.L. and J.T. conceived of the study. R.A. and R.L. designed the methodology. R.A. implemented the software, ran experiments and created visualizations with the help of S.A.K. A.M. managed the software resources and the GitHub page. The formal analyses and validation were performed by R.A. and R.L, and R.A. wrote the original draft with the supervision of R.L. and J.T. All authors contributed to reviewing and revising the original paper. R.L. and J.T. supervised the project, and J.T. secured the funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Xin Ye and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Note 1, Figs. 1–8 and Tables 1–2.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data and UMAP coordinates. UMAP of the original data in https://github.com/reem12345/Cell-Type-Specific-Graphs.
Source Data Fig. 3 (download XLSX )
Statistical source data and UMAP coordinates. UMAP of the original data in https://github.com/reem12345/Cell-Type-Specific-Graphs.
Source Data Fig. 4 (download XLSX )
Statistical source data and UMAP coordinates.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Alsulami, R., Lehmann, R., Khan, S.A. et al. Predicting and interpreting cell-type-specific drug responses in the small-data regime using inductive priors. Nat Mach Intell 8, 461–473 (2026). https://doi.org/10.1038/s42256-026-01202-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-026-01202-2