Abstract
Programmed death 1 blockade (tislelizumab) has been approved for metastatic urothelial carcinoma but not as part of neoadjuvant therapy for muscle-invasive bladder cancer (MIBC). In this multicenter single-arm trial (ChiCTR2000037670), 65 participants with cT2-4aN0M0 MIBC received neoadjuvant gemcitabine–cisplatin plus tislelizumab; 57 of them underwent radical cystectomy (RC). The primary endpoint of pathologic complete response (pCR) rate was 50.9% (29/57, 95% confidence interval (CI) 37.3–64.4%) and the pathologic downstaging (secondary endpoint) rate was 75.4% (43/57, 95% CI 62.2–85.9%) in participants undergoing RC. Genomic and transcriptomic analyses revealed three MIBC molecular subtypes (S): S1 (immune-desert) with activated cell-cycle pathway, S2 (immune-excluded) with activated transforming growth factor-β pathway and S3 (immune-inflamed) with upregulated interferon-α and interferon-γ response. Post hoc analysis showed pCR rates of 16% (3/19, S1), 77% (10/13, S2) and 80% (12/15, S3) (P = 0.006). In conclusion, neoadjuvant gemcitabine–cisplatin plus tislelizumab for MIBC was compatible with an enhanced pCR rate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
RNA and DNA sequencing data that support the findings of this study were deposited to the Genome Sequence Archive in National Genomics Data Center, China National Center for Bioinformation and Beijing Institute of Genomics, Chinese Academy of Sciences under accession code HRA006086. The dataset derived from this resource that supports the findings of this study is available at https://bigd.big.ac.cn/gsa-human/browse/HRA006086. RNA and DNA sequencing data generated for this study are subjected to the regulations by the Ministry of Science and Technology of the People’s Republic of China. The hg19 human genome can be found at https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001405.13/. All other data supporting the findings of this study are available from T.L. (lintx@mail.sysu.edu.cn) or K.L. (likw6@mail.sysu.edu.cn) on reasonable request. Individual deidentified participant data (including data dictionaries, text and tables) that underlie the results reported in this article will be eligible for data sharing requests after January 1, 2027. The study protocol can be found in the Supplementary Information. Researchers should send methodologically sound proposals directly to T.L. (lintx@mail.sysu.edu.cn) or K.L. (likw6@mail.sysu.edu.cn) for data requests. The reuse of the individual participant data is permitted only for revalidation of the results or for meta-analysis. Source data are provided with this paper.
Code availability
All relevant package and software information is provided in the Methods. No custom code was generated in the course of this study.
References
Kamat, A. M. et al. Bladder cancer. Lancet 388, 2796–2810 (2016).
Alfred Witjes, J. et al. European association of urology guidelines on muscle-invasive and metastatic bladder cancer: summary of the 2023 guidelines. Eur. Urol. 85, 17–31 (2024).
Flaig, T. W. et al. A randomized phase II study of coexpression extrapolation (COXEN) with neoadjuvant chemotherapy for bladder cancer (SWOG S1314; NCT02177695). Clin. Cancer Res. 27, 2435–2441 (2021).
Zargar, H. et al. Multicenter assessment of neoadjuvant chemotherapy for muscle-invasive bladder cancer. Eur. Urol. 67, 241–249 (2015).
Iyer, G. et al. Neoadjuvant gemcitabine–cisplatin plus radical cystectomy–pelvic lymph node dissection for muscle-invasive bladder cancer: a 12-year experience. Clin. Genitourin. Cancer 18, 387–394 (2020).
Powles, T. et al. Clinical efficacy and biomarker analysis of neoadjuvant atezolizumab in operable urothelial carcinoma in the ABACUS trial. Nat. Med. 25, 1706–1714 (2019).
Necchi, A. et al. Pembrolizumab as neoadjuvant therapy before radical cystectomy in patients with muscle-invasive urothelial bladder carcinoma (PURE-01): an open-label, single-arm, phase II study. J. Clin. Oncol. 36, 3353–3360 (2018).
de Biasi, A. R., Villena-Vargas, J. & Adusumilli, P. S. Cisplatin-induced antitumor immunomodulation: a review of preclinical and clinical evidence. Clin. Cancer Res. 20, 5384–5391 (2014).
Rose, T. L. et al. Phase II study of gemcitabine and split-dose cisplatin plus pembrolizumab as neoadjuvant therapy before radical cystectomy in patients with muscle-invasive bladder cancer. J. Clin. Oncol. 39, 3140–3148 (2021).
Funt, S. A. et al. Neoadjuvant atezolizumab with gemcitabine and cisplatin in patients with muscle-invasive bladder cancer: a multicenter, single-arm, phase II trial. J. Clin. Oncol. 40, 1312–1322 (2022).
Zhang, T. et al. The binding of an anti-PD-1 antibody to FcγRΙ has a profound impact on its biological functions. Cancer Immunol. Immunother. 67, 1079–1090 (2018).
Ye, D. et al. Tislelizumab in Asian patients with previously treated locally advanced or metastatic urothelial carcinoma. Cancer Sci. 112, 305–313 (2021).
Robertson, A. G. et al. Comprehensive molecular characterization of muscle-Invasive bladder cancer. Cell 171, 540–556 (2017).
Plimack, E. R. et al. Defects in DNA repair genes predict response to neoadjuvant cisplatin-based chemotherapy in muscle-invasive bladder cancer. Eur. Urol. 68, 959–967 (2015).
Van Allen, E. M. et al. Somatic ERCC2 mutations correlate with cisplatin sensitivity in muscle-invasive urothelial carcinoma. Cancer Discov. 4, 1140–1153 (2014).
Adib, E. et al. CDKN2A alterations and response to immunotherapy in solid tumors. Clin. Cancer Res. 27, 4025–4035 (2021).
Huang, Q. et al. Loss of TSC1/TSC2 sensitizes immune checkpoint blockade in non-small cell lung cancer. Sci. Adv. 8, eabi9533 (2022).
Lu, Q. et al. Rheb1 protects against cisplatin-induced tubular cell death and acute kidney injury via maintaining mitochondrial homeostasis. Cell Death Dis. 11, 364 (2020).
Teo, M. Y. et al. Alterations in DNA damage response and repair genes as potential marker of clinical benefit from PD-1/PD-L1 blockade in advanced urothelial cancers. J. Clin. Oncol. 36, 1685–1694 (2018).
Teo, M. Y. et al. DNA damage response and repair gene alterations are associated with improved survival in patients with platinum-treated advanced urothelial carcinoma. Clin. Cancer Res. 23, 3610–3618 (2017).
de Visser, K. E. & Joyce, J. A. The evolving tumor microenvironment: from cancer initiation to metastatic outgrowth. Cancer Cell 41, 374–403 (2023).
Tian, B. et al. Curcumin inhibits urothelial tumor development by suppressing IGF2 and IGF2-mediated PI3K/AKT/mTOR signaling pathway. J. Drug Target. 25, 626–636 (2017).
Das, R. et al. An integrated functional and clinical genomics approach reveals genes driving aggressive metastatic prostate cancer. Nat. Commun. 12, 4601 (2021).
De Vitis, C. et al. ALDOC- and ENO2-driven glucose metabolism sustains 3D tumor spheroids growth regardless of nutrient environmental conditions: a multi-omics analysis. J. Exp. Clin. Canc. Res. 42, 69 (2023).
Januchowski, R. et al. Drug transporter expression profiling in chemoresistant variants of the A2780 ovarian cancer cell line. Biomed. Pharmacother. 68, 447–453 (2014).
Maddika, S. et al. Cell survival, cell death and cell cycle pathways are interconnected: implications for cancer therapy. Drug Resist. Updat. 10, 13–29 (2007).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Kamoun, A. et al. A consensus molecular classification of muscle-invasive bladder cancer. Eur. Urol. 77, 420–433 (2020).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Liu, Z. et al. TTN/OBSCN ‘double‐hit’ predicts favourable prognosis, ‘immune‐hot’ subtype and potentially better immunotherapeutic efficacy in colorectal cancer. J. Cell. Mol. Med. 25, 3239–3251 (2021).
Zhang, L., Han, X. & Shi, Y. Association of MUC16 mutation with response to immune checkpoint inhibitors in solid tumors. JAMA Netw. Open 3, e2013201 (2020).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437–1452 (2017).
Carneiro, B. A. et al. Phase I study of elraglusib (9-ING-41), a glycogen synthase kinase-3β inhibitor, as monotherapy or combined with chemotherapy in patients with advanced malignancies. Clin. Cancer Res. 30, 522–531 (2024).
Song, Y. et al. Fibroblast growth factor receptor 3 mutation attenuates response to immune checkpoint blockade in metastatic urothelial carcinoma by driving immunosuppressive microenvironment. J. Immunother. Cancer 11, e006643 (2023).
Cathomas, R. et al. Perioperative chemoimmunotherapy with durvalumab for muscle-invasive urothelial carcinoma: primary analysis of the single-arm phase II trial SAKK 06/17. J. Clin. Oncol. 41, 5131–5139 (2023).
Brown, J. et al. HCRN GU14-188: phase Ib/II study of neoadjuvant pembrolizumab and chemotherapy for T2-4aN0M0 urothelial cancer. J. Clin. Oncol. 41, 448–448 (2023).
Gupta, S. et al. Results from BLASST-1 (Bladder Cancer Signal Seeking Trial) of nivolumab, gemcitabine, and cisplatin in muscle invasive bladder cancer (MIBC) undergoing cystectomy. J. Clin. Oncol. 38, 439–439 (2020).
Powles, T. et al. Pembrolizumab alone or combined with chemotherapy versus chemotherapy as first-line therapy for advanced urothelial carcinoma (KEYNOTE-361): a randomised, open-label, phase 3 trial. Lancet Oncol. 22, 931–945 (2021).
Bamias, A. et al. Atezolizumab monotherapy versus chemotherapy in untreated locally advanced or metastatic urothelial carcinoma (IMvigor130): final overall survival analysis from a randomised, controlled, phase 3 study. Lancet Oncol. 25, 46–61 (2024).
Van Der Heijden, M. S. et al. Nivolumab plus gemcitabine–cisplatin in advanced urothelial carcinoma. N. Engl. J. Med. 389, 1778–1789 (2023).
Bajorin, D. F. et al. Adjuvant nivolumab versus placebo in muscle-invasive urothelial carcinoma. N. Engl. J. Med. 384, 2102–2114 (2021).
Szabados, B. et al. Final results of neoadjuvant atezolizumab in cisplatin-ineligible patients with muscle-invasive urothelial cancer of the bladder. Eur. Urol. 82, 212–222 (2022).
Galsky, M. D. et al. Determinants of survival with combined HER2 and PD-1 blockade in metastatic esophagogastric cancer. Clin. Cancer Res. 29, 3633–3640 (2023).
Kitamura, H. et al. Randomised phase III study of neoadjuvant chemotherapy with methotrexate, doxorubicin, vinblastine and cisplatin followed by radical cystectomy compared with radical cystectomy alone for muscle-invasive bladder cancer: Japan Clinical Oncology Group Study JCOG0209. Ann. Oncol. 25, 1192–1198 (2014).
Grossman, H. B. et al. Neoadjuvant chemotherapy plus cystectomy compared with cystectomy alone for locally advanced bladder cancer. N. Engl. J. Med. 349, 859–866 (2003).
Strandgaard, T. et al. Field cancerization is associated with tumor development, T-cell exhaustion, and clinical outcomes in bladder cancer. Eur. Urol. 85, 82–92 (2024).
Robertson, A. G. et al. Expression-based subtypes define pathologic response to neoadjuvant immune-checkpoint inhibitors in muscle-invasive bladder cancer. Nat. Commun. 14, 2126 (2023).
Kortlever, R. M. et al. Myc cooperates with Ras by programming inflammation and immune suppression. Cell 171, 1301–1315 (2017).
Fang, X. et al. Sintilimab plus bevacizumab, oxaliplatin and capecitabine as first-line therapy in RAS-mutant, microsatellite stable, unresectable metastatic colorectal cancer: an open-label, single-arm, phase II trial. EClinicalMedicine 62, 102123 (2023).
Kent, L. N. & Leone, G. The broken cycle: E2F dysfunction in cancer. Nat. Rev. Cancer 19, 326–338 (2019).
Baldominos, P. et al. Quiescent cancer cells resist T cell attack by forming an immunosuppressive niche. Cell 185, 1694–1708 (2022).
Seiler, R. et al. Divergent biological response to neoadjuvant chemotherapy in muscle-invasive bladder cancer. Clin. Cancer Res. 25, 5082–5093 (2019).
Schulz, K. F., Altman, D. G., Moher, D. & CONSORT Group CONSORT 2010 statement: updated guidelines for reporting parallel group randomised trials. BMJ 340, c332–c332 (2010).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2018).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Aran, D., Hu, Z. & Butte, A. J. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 220 (2017).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Li, S. et al. Molecular signatures of antibody responses derived from a systems biology study of five human vaccines. Nat. Immunol. 15, 195–204 (2014).
Chaussabel, D. et al. A modular analysis framework for blood genomics studies: application to systemic lupus erythematosus. Immunity 29, 150–164 (2008).
Zyla, J. et al. Gene set enrichment for reproducible science: comparison of CERNO and eight other algorithms. Bioinformatics 35, 5146–5154 (2019).
Simon, R. Optimal two-stage designs for phase II clinical trials. Controlled Clin. Trials 10, 1–10 (1989).
Acknowledgements
This trial was supported by the National Key Research and Development Program of China (2018YFA0902800), the National Natural Science Foundation of China (81825016, 82341018 and U21A20383) and Sun Yat-Sen Memorial Hospital Clinical Research 5010 Program (SYS-5010Z-202401) to T.L., the National Natural Science Foundation of China (82173230 and 81961128027) to J.H., the National Natural Science Foundation of China (82173088), the Natural Science Foundation of Guangdong (2022A1515012383) and research funding of Sun Yat-sen University (23ptpy168) to K.L., the National Natural Science Foundation of China (82373254) to W.Z. and the Guangdong Provincial Clinical Research Center for Urological Diseases (2020B1111170006). BeiGene (Beijing, China) provided tislelizumab free of charge and financial support for the procurement of gemcitabine and cisplatin, as well as biomarker analysis. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. L. Ling from the Department of Medical Statistics, School of Public Health, Sun Yat-Sen University provided statistical assistance. K. Zhang from the Ivy Medical Editing (Shanghai, China) provided writing and editing assistance.
Author information
Authors and Affiliations
Contributions
Conceptualization and design, T.L., J.H., K.L. and W.Z. Administrative support: T.L. and J.H. Provision of study materials, K.L., J.F., S.Wang, D.Y., T.X., J.L., Z.L., K.W., J.W., Q.W., J.M., Z.H., F.L., Z.Z., L.Y., S.D., J.H. and T.L. Collection and assembly of data, K.L., J.F., S. Wang, D.Y., T.X., J.L., T.Q., Z.L., K.W., J.W., Q.W., J.M., Z.H., F.L., Z.Z., L.Y., S.D., J.H. and T.L. Data analysis and interpretation, K.L., W.Z., J.F., S.Wang, D.Y., T.X., J.L., S.Wu, T.Q., Z.W., L.X., Z.L., K.W., J.W., Q.W., J.M., Z.H., F.L., Z.Z., L.Y., S.D., L.H., T.Z., J.H. and T.L. Manuscript writing, K.L., W.Z., J.H. and T.L. Accountable for all aspects of the work: T.L. and J.H. All authors read and approved the final version of the manuscript. T.L. and J.H. supervised all aspects of this work.
Corresponding authors
Ethics declarations
Competing interests
L.H. and T.Z. were employees of GloriousMed at the time of the study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Joaquim Bellmunt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Trial design and participant enrollment.
a, The design of this trial; b, Flow of participants enrolled in this trial.
Extended Data Fig. 2 Comparison of response to neoadjuvant treatment by radiologic and pathologic assessment.
a, Representative magnetic resonance imaging (MRI) images showing bladder tumor status in a cT3N0M0 patient, at the time of prior-cycle 1 and cycle 3 of neoadjuvant treatment, and prior-radical cystectomy, respectively. This patient was evaluated as radiologic complete response (rCR) after neoadjuvant treatment and confirmed as pathologic complete response (pCR) after radical cystectomy. b, Comparison between rCR and pCR. ADC: apparent diffusion coefficient; DCE: dynamic contrast enhanced; DWI: diffusion weighted imaging; PD: progressive disease; T2WI: T2 weighted imaging.
Extended Data Fig. 3 Swimmer plot with treatment, response and follow-up details for 57 patients undergoing radical cystectomy.
Duration is relative to the date of first-dose neoadjuvant treatment. Clinical T/N stage is shown on the left side of the plot and displayed by bars with colors (green for T2N0M0, red for T3N0M0, and blue for T4aN0M0, respectively). Pathologic T/N stage after radical cystectomy is shown on the right side. Time points are marked by different shapes, including inverted triangle (last-dose of neoadjuvant treatment), triangle (last radiologic evaluation), diamond (radical cystectomy), circle (recurrence), cross (death), and arrow (continued follow-up).
Extended Data Fig. 4 Kaplan–Meier curves of event-free, overall survival (EFS and OS) and recurrence-free survival (RFS) stratified by different pathologic responses to neoadjuvant treatment.
a, b, c, Patients with (n = 14) or without pDS (n = 43). The cutoff of landmark analysis was 6 months for EFS and OS, and 3 months for RFS. Hazard ratio (HR) was calculated by comparing patients with vs without pDS. d, e, f, Patients with pCR (n = 29), paR (n = 14) and non-pDS (n = 14). CI, confidence interval; ns: non-significant; paR, partial response, Tis/Ta/T1N0M0; pCR, pathologic complete response; pDS, pathologic downstaging. p-values were calculated by using two-sided log-rank test.
Extended Data Fig. 5 Baseline urine tumor DNA (utDNA) assessment.
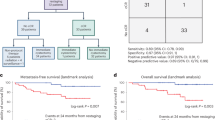
Estimated variant allele frequency (eVAF%) in baseline urine from patients with a, bladder-pCR (pT0Nx, n = 32) vs non-pT0 (n = 25); b, pCR (n = 29) vs non-pCR (n = 28), and c, pDS (n = 43) vs non-pDS (n = 14). pCR, pathologic complete response; pDS, pathologic downstaging. Two-sided Mann-Whitney U test was used.
Extended Data Fig. 6 Immune checkpoint gene expression and hallmark pathways in three subtypes.
a, Expression level of PDCD1, CD274, CTLA4, TIM3, and LAG3 gene; and b, Single-sample Gene set enrichment analysis showing pathways (based on the MSigDB v6.2 Hallmarks gene sets) with significant differences among three subtypes (subtype [S]1 [n = 19], S2 [n = 13] and S3 [n = 15]). Two-side Kruskal-Wallis test was used.
Extended Data Fig. 7 Somatic mutations and copy number variations (CNVs) in three subtypes.
a, Oncoprints for somatic mutations and CNVs in subtype 1 (S1, n = 18), S2 (n = 13) and S3 (n = 15) tumors. Horizontal bars to the right indicate the number of samples with an alteration and the types of alterations in indicated gene. Bar plots at the top show the total number of genetic alterations in the oncoprint genes. Horizontal bars and bar plots are colored by the alteration type. b, Percentage of subtypes (S1, n = 18; S2, n = 13; and S3, n = 15) tumors in patients with mutation in 13 selected genes. c, Pathways (based on the MSigDB v6.2 Hallmarks gene sets) with significant difference in patients (n = 46) with vs without mutation in 13 selected genes. Red (vs blue) discs represent for enriched (vs repressed) gene sets with disc areas for the areas-under-the-curve (AUCs) by the CERNO test.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2 and Study protocol.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Table 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed MRI images.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Table 1 (download XLSX )
Statistical source data.
Source Data Extended Data Table 2 (download XLSX )
Statistical source data.
Source Data Extended Data Table 3 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, K., Zhong, W., Fan, J. et al. Neoadjuvant gemcitabine–cisplatin plus tislelizumab in persons with resectable muscle-invasive bladder cancer: a multicenter, single-arm, phase 2 trial. Nat Cancer 5, 1465–1478 (2024). https://doi.org/10.1038/s43018-024-00822-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-024-00822-0
This article is cited by
-
Glutamine metabolism reprogramming promotes bladder cancer progression via PYCR1: a multi-omics and functional validation study
Journal of Translational Medicine (2025)
-
Regulation of drug resistance in bladder urothelial carcinoma by tumor aerobic glycolysis
Journal of Translational Medicine (2025)
-
Real-world comparison of neoadjuvant chemoimmunotherapy and chemotherapy in muscle-invasive bladder cancer
Scientific Reports (2025)
-
Disitamab vedotin (RC48-ADC) combined with immunotherapy as neoadjuvant therapy for localized muscle-invasive bladder cancer: a multicenter real-world study
npj Precision Oncology (2025)
-
Derazantinib enhances gemcitabine efficacy in PDAC by attenuating the NF-κB and MAPK pathways to suppress MUC5AC expression
Medical Oncology (2025)