Abstract
Hepatocellular carcinoma (HCC) frequently recurs from minimal residual disease (MRD), which persists after therapy. Here, we identified mechanisms of persistence of residual tumor cells using post-chemoembolization human HCC (n = 108 patients, 1.07 million cells) and a transgenic mouse model of MRD. Through single-cell high-plex cytometric imaging, we identified a spatial neighborhood within which PD-L1 + M2-like macrophages interact with stem-like tumor cells, correlating with CD8+ T cell exhaustion and poor survival. Further, through spatial transcriptomics of residual HCC, we showed that macrophage-derived TGFβ1 mediates the persistence of stem-like tumor cells. Last, we demonstrate that combined blockade of Pdl1 and Tgfβ excluded immunosuppressive macrophages, recruited activated CD8+ T cells and eliminated residual stem-like tumor cells in two mouse models: a transgenic model of MRD and a syngeneic orthotopic model of doxorubicin-resistant HCC. Thus, our spatial analyses reveal that PD-L1+ macrophages sustain MRD by activating the TGFβ pathway in stem-like cancer cells and targeting this interaction may prevent HCC recurrence from MRD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Source data for all figures have been provided as Source Data files. All other data supporting the findings of this study are available from the corresponding author on reasonable request. CODEX data (the raw data matrix), which includes cell-level information on normalized expression for all markers, centroid coordinates and the assigned cell type and cohort for each cell, have been deposited in Figshare (ref. 73). In addition, if raw images are sought, we will make them available at https://app.enablemedicine.com/portal/visualizer upon request to the corresponding author (dhanaser@stanford.edu) within 10 days of receiving the request. Nanostring data: the gene expression data matrix, which includes sample-level information on normalized expression for all genes, and the assigned AOI type and cohort for each sample, has been deposited in Figshare (ref. 74). The mouse single-cell and whole-transcriptome sequencing data generated in this study are publicly available in the National Center for Biotechnology Information Gene Expression Omnibus database under accession codes GSE243176, GSE242746, GSE242745 and GSE242743. Publicly available TCGA liver cancer data were used for analysis from https://www.cbioportal.org/study/summary?id=lihc_tcga_pan_can_atlas_2018. Source data are provided with this paper.
Code availability
No custom code or algorithms were generated in this manuscript.
References
Lee, Y.-T. et al. The mortality and overall survival trends of primary liver cancer in the United States. J. Natl. Cancer Inst. 113, 1531–1541 (2021).
Lei, J. et al. Response to transarterial chemoembolization may serve as selection criteria for hepatocellular carcinoma liver transplantation. Oncotarget 8, 91328–91342 (2017).
Kim, D. J. et al. Recurrence of hepatocellular carcinoma: importance of mRECIST response to chemoembolization and tumor size. Am. J. Transplant 14, 1383–1390 (2014).
Agopian, V. G. et al. Impact of pretransplant bridging locoregional therapy for patients with hepatocellular carcinoma within Milan criteria undergoing liver transplantation: analysis of 3601 patients from the US multicenter HCC transplant consortium. Ann. Surg. 266, 525–535 (2017).
Adeniji, N. et al. Impact of bridging locoregional therapies for hepatocellular carcinoma on post-transplant clinical outcome. Clin. Transplant. 34, e14128 (2020).
Ghajar, C. M. Metastasis prevention by targeting the dormant niche. Nat. Rev. Cancer 15, 238–247 (2015).
Lee, T. K.-W., Guan, X.-Y. & Ma, S. Cancer stem cells in hepatocellular carcinoma - from origin to clinical implications. Nat. Rev. Gastroenterol. Hepatol. 19, 26–44 (2022).
Price, T. T. et al. Dormant breast cancer micrometastases reside in specific bone marrow niches that regulate their transit to and from bone. Sci. Transl. Med. 8, 340ra73 (2016).
Shachaf, C. M. et al. MYC inactivation uncovers pluripotent differentiation and tumour dormancy in hepatocellular cancer. Nature 431, 1112–1117 (2004).
Ding, W. et al. Clinicopathologic and prognostic significance of tumor-associated macrophages in patients with hepatocellular carcinoma: a meta-analysis. PLoS ONE 14, e0223971 (2019).
Zhu, X.-D. et al. High expression of macrophage colony-stimulating factor in peritumoral liver tissue is associated with poor survival after curative resection of hepatocellular carcinoma. J. Clin. Oncol. 26, 2707–2716 (2008).
Borriello, L. et al. Primary tumor associated macrophages activate programs of invasion and dormancy in disseminating tumor cells. Nat. Commun. 13, 626 (2022).
Dhanasekaran, R. et al. MYC overexpression drives immune evasion in hepatocellular carcinoma that is reversible through restoration of proinflammatory macrophages. Cancer Res. 83, 626–640 (2023).
Dhanasekaran, R. et al. MYC and Twist1 cooperate to drive metastasis by eliciting crosstalk between cancer and innate immunity. eLife https://doi.org/10.7554/elife.50731 (2020).
Black, S. et al. CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat. Protoc. 16, 3802–3835 (2021).
Uenishi, T. et al. Cytokeratin 19 expression in hepatocellular carcinoma predicts early postoperative recurrence. Cancer Sci. 94, 851–857 (2003).
Khosla, R. et al. EpCAM+ liver cancer stem-like cells exhibiting autocrine wnt signaling potentially originate in cirrhotic patients. Stem Cells Transl. Med. 6, 807–818 (2017).
Liu, C.-Q. et al. Expression patterns of programmed death ligand 1 correlate with different microenvironments and patient prognosis in hepatocellular carcinoma. Br. J. Cancer 119, 80–88 (2018).
Pfister, D. et al. NASH limits anti-tumour surveillance in immunotherapy-treated HCC. Nature 592, 450–456 (2021).
Cancer Genome Atlas Research Network. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341.e23 (2017).
Guo, Y. et al. The heterogeneity of immune cell infiltration landscape and its immunotherapeutic implications in hepatocellular carcinoma. Front. Immunol. 13, 861525 (2022).
Yu, S. et al. Tumor-infiltrating immune cells in hepatocellular carcinoma: Tregs is correlated with poor overall survival. PLoS ONE 15, e0231003 (2020).
Veglia, F. et al. Analysis of classical neutrophils and polymorphonuclear myeloid-derived suppressor cells in cancer patients and tumor-bearing mice. J. Exp. Med. 218, e20201803 (2021).
Salcher, S. et al. High-resolution single-cell atlas reveals diversity and plasticity of tissue-resident neutrophils in non-small cell lung cancer. Cancer Cell 40, 1503–1520.e8 (2022).
Singal, A. G. et al. AASLD practice guidance on prevention, diagnosis, and treatment of hepatocellular carcinoma. Hepatology 78, 1922–1965 (2023).
Schürch, C. M. et al. Coordinated cellular neighborhoods orchestrate antitumoral immunity at the colorectal cancer invasive front. Cell 183, 838 (2020).
Dasari, A., Grothey, A. & Kopetz, S. Circulating tumor DNA-defined minimal residual disease in solid tumors: opportunities to accelerate the development of adjuvant therapies. J. Clin. Oncol. 36, JCO2018789032 (2018).
Villanueva, A. et al. Combining clinical, pathology, and gene expression data to predict recurrence of hepatocellular carcinoma. Gastroenterology 140, 1501–12.e2 (2011).
Pello, O. M. et al. Role of c-MYC in alternative activation of human macrophages and tumor-associated macrophage biology. Blood 119, 411–421 (2012).
Li, Z. et al. Cancer-associated fibroblasts promote PD-L1 expression in mice cancer cells via secreting CXCL5. Int. J. Cancer 145, 1946–1957 (2019).
Zhang, W. et al. IL-6 promotes PD-L1 expression in monocytes and macrophages by decreasing protein tyrosine phosphatase receptor type O expression in human hepatocellular carcinoma. J. Immunother. Cancer 8, e000285 (2020).
Xi, X. et al. Interleukin-22 promotes PD-L1 expression via STAT3 in colon cancer cells. Oncol. Lett. 22, 716 (2021).
Kobori, T. et al. Interleukin-18 amplifies macrophage polarization and morphological alteration, leading to excessive angiogenesis. Front. Immunol. 9, 334 (2018).
Patel, M. & Pillai, A. Management of intermediate-stage hepatocellular carcinoma: systemic versus locoregional therapy. Surg. Oncol. Clin. N. Am. 33, 159–172 (2024).
Lu, Y. et al. A single-cell atlas of the multicellular ecosystem of primary and metastatic hepatocellular carcinoma. Nat. Commun. 13, 4594 (2022).
Ma, L. et al. Single-cell atlas of tumor cell evolution in response to therapy in hepatocellular carcinoma and intrahepatic cholangiocarcinoma. J. Hepatol. 75, 1397–1408 (2021).
Ma, L. et al. Tumor cell biodiversity drives microenvironmental reprogramming in liver cancer. Cancer Cell 36, 418–430.e6 (2019).
Ruf, B. et al. Tumor-associated macrophages trigger MAIT cell dysfunction at the HCC invasive margin. Cell 186, 3686–3705.e32 (2023).
Maestri, E. et al. Spatial proximity of tumor-immune interactions predicts patient outcome in hepatocellular carcinoma. Hepatology https://doi.org/10.1097/hep.0000000000000600 (2023).
Govaere, O. et al. Keratin 19: a key role player in the invasion of human hepatocellular carcinomas. Gut 63, 674–685 (2014).
Zhang, J. et al. Overexpression of Epcam and CD133 correlates with poor prognosis in dual-phenotype hepatocellular carcinoma. J. Cancer 11, 3400–3406 (2020).
Noh, C.-K. et al. EpCAM as a predictive marker of tumor recurrence and survival in patients who underwent surgical resection for hepatocellular carcinoma. Anticancer Res. 38, 4101–4109 (2018).
Zhou, C. et al. Disruption of SLFN11 deficiency-induced CCL2 signaling and macrophage M2 polarization potentiates anti-PD-1 therapy efficacy in hepatocellular carcinoma. Gastroenterology 164, 1261–1278 (2023).
Murai, H. et al. Multiomics identifies the link between intratumor steatosis and the exhausted tumor immune microenvironment in hepatocellular carcinoma. Hepatology 77, 77–91 (2023).
Li, X. et al. Targeting of tumour-infiltrating macrophages via CCL2/CCR2 signalling as a therapeutic strategy against hepatocellular carcinoma. Gut 66, 157–167 (2017).
Martin-Padura, I. et al. Residual dormant cancer stem-cell foci are responsible for tumor relapse after antiangiogenic metronomic therapy in hepatocellular carcinoma xenografts. Lab. Invest. 92, 952–966 (2012).
Haraguchi, N. et al. CD13 is a therapeutic target in human liver cancer stem cells. J. Clin. Invest. 120, 3326–3339 (2010).
Schrader, J. et al. Matrix stiffness modulates proliferation, chemotherapeutic response, and dormancy in hepatocellular carcinoma cells. Hepatology 53, 1192–1205 (2011).
Ravi, R. et al. Bifunctional immune checkpoint-targeted antibody-ligand traps that simultaneously disable TGFβ enhance the efficacy of cancer immunotherapy. Nat. Commun. 9, 741 (2018).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Lan, Y. et al. Enhanced preclinical antitumor activity of M7824, a bifunctional fusion protein simultaneously targeting PD-L1 and TGF-β. Sci. Transl. Med. 10, eaan5488 (2018).
Metropulos, A. E., Munshi, H. G. & Principe, D. R. The difficulty in translating the preclinical success of combined TGFβ and immune checkpoint inhibition to clinical trial. eBioMedicine 86, 104380 (2022).
Lencioni, R. & Llovet, J. M. Modified RECIST (mRECIST) assessment for hepatocellular carcinoma. Semin. Liver Dis. 30, 52–60 (2010).
Merritt, C. R. et al. Multiplex digital spatial profiling of proteins and RNA in fixed tissue. Nat. Biotechnol. 38, 586–599 (2020).
Cosi, C., Millar, M., Beltran, M., Sherry, L. & Gatti-McArthur, S. Quantitative analysis of RNAscope staining for expression in mouse brain tissue as a measure of neuronal activation. MethodsX 8, 101348 (2021).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Genin, M., Clement, F., Fattaccioli, A., Raes, M. & Michiels, C. M1 and M2 macrophages derived from THP-1 cells differentially modulate the response of cancer cells to etoposide. BMC Cancer 15, 577 (2015).
Gambari, R., Hau, D. K.-P., Wong, W.-Y. & Chui, C.-H. Sensitization of Hep3B hepatoma cells to cisplatin and doxorubicin by corilagin. Phytother. Res. 28, 781–783 (2014).
Hu, Q.-D. et al. NSC 74859 enhances doxorubicin cytotoxicity via inhibition of epithelial-mesenchymal transition in hepatocellular carcinoma cells. Cancer Lett. 325, 207–213 (2012).
Buschauer, S. et al. Hepatocellular carcinoma cells surviving doxorubicin treatment exhibit increased migratory potential and resistance to doxorubicin re-treatment in vitro. Oncol. Lett. 15 https://doi.org/10.3892/ol.2018.7887 (2018).
Namur, J. et al. Embolization of hepatocellular carcinoma with drug-eluting beads: doxorubicin tissue concentration and distribution in patient liver explants. J. Hepatol. 55, 1332–1338 (2011).
Ulbrich, J., Lopez-Salmeron, V. & Gerrard, I. BD RhapsodyTM single-cell analysis system workflow: from sample to multimodal single-cell sequencing data. Methods Mol. Biol. 2584, 29–56 (2023).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01767-y (2023).
Nielsen, M. C., Andersen, M. N. & Møller, H. J. Monocyte isolation techniques significantly impact the phenotype of both isolated monocytes and derived macrophages in vitro. Immunology 159, 63–74 (2020).
Wu, P. V. & Nusse, R. 3D culture of primary patient-derived hepatoblastoma tumoroids. Methods Mol. Biol. 2544, 259–267 (2022).
Wang, S. K., Xue, Y. & Cepko, C. L. Microglia modulation by TGF-β1 protects cones in mouse models of retinal degeneration. J. Clin. Invest. 130, 4360–4369 (2020).
Kress, S. et al. p53 mutations are absent from carcinogen-induced mouse liver tumors but occur in cell lines established from these tumors. Mol. Carcinog. 6, 148–158 (1992).
Brown, Z. J., Heinrich, B. & Greten, T. F. Establishment of orthotopic liver tumors by surgical intrahepatic tumor injection in mice with underlying non-alcoholic fatty liver disease. Methods Protoc. 1, 21 (2018).
Zabransky, D. J. et al. Profiling of syngeneic mouse HCC tumor models as a framework to understand anti-PD-1 sensitive tumor microenvironments. Hepatology 77, 1566–1579 (2023).
Lai, I. et al. Lipid nanoparticles that deliver IL-12 messenger RNA suppress tumorigenesis in MYC oncogene-driven hepatocellular carcinoma. J. Immunother. Cancer 6, 125 (2018).
Dhanasekaran, R. et al. MYC ASO impedes tumorigenesis and elicits oncogene addiction in autochthonous transgenic mouse models of HCC and RCC. Mol. Ther. Nucleic Acids 21, 850–859 (2020).
Palmer, G. M. et al. In vivo optical molecular imaging and analysis in mice using dorsal window chamber models applied to hypoxia, vasculature and fluorescent reporters. Nat. Protoc. 6, 1355–1366 (2011).
Dhanasekaran, R. Spatial analysis reveals targetable macrophage-mediated mechanisms of immune evasion in hepatocellular carcinoma minimal residual disease. Figshare https://doi.org/10.6084/m9.figshare.26539345 (2024).
Dhanasekaran, R. Nanostring_GeoMx_HCC_9_2_24.csv. Figshare https://doi.org/10.6084/m9.figshare.26898886.v1 (2024).
Acknowledgements
R.D. acknowledges NIH grant CA222676 from the National Cancer Institute, American College of Gastroenterology Junior Faculty Career Development Grant and Cancer League Award. D.F. acknowledges NIH grant CA208735 and CA253180 from the National Cancer Institute. P. Chu helped with mouse histology services.
Author information
Authors and Affiliations
Contributions
R.D. and D.F. conceptualized the study. L.L., N.A., A.S., R.R., J.Z., J.P., A.R., A.D., A.S.H., L.T. and V.A. carried out experiments under the supervision of R.D. and D.F. A.T., A.M., H.B.D.’A., R.P., Y.R., J.Z. and V.C. contributed to method development. L.L., N.A., A.S., A.D., A.S.H. and Y.R. analyzed the data. N. Kambham, B.C.V., M.M.D., C.A.B., N. Kothary, V.C. and R.D. who contributed through clinical data retrieval, clinical follow-up and sample acquisition. The paper was written by R.D. with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A.T.M., A.E.T., R.P. and H.B.D. are employees of Enable Medicine. All other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Bertram Bengsch, Tim Greten and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification, validation and relative proportion of major cell types using CODEX.
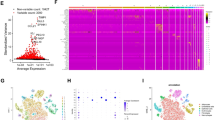
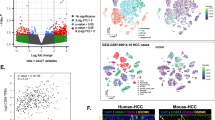
a. UMAP representation of 12 major cell subtypes identified by CODEX analysis of human HCC. b. Comparison of normalized expression patterns of stemness markers (EPCAM, CK19, CD44), survival markers (Ki67, BCL2), and mesenchymal markers (vimentin, podoplanin) between stem-like and non-stem-like tumor cell types identified by CODEX analysis of human HCCs (Tumor cell EpCAM+ n= 26622 cells, Tumor cell CK19+ n= 34436 cells, Tumor cell CK+ n= 433,918 cells). c. Comparison of normalized expression patterns of macrophage markers between three macrophage subsets identified by CODEX analysis of human HCCs (Macro 206+ n=45628 cells; Macro HLA-DR+ n=55558 cells; Macro PD-L1+ n=53713 cells). Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. d. Stacked bar chart comparing relative proportions of all 20 immune and tumor cell subtypes identified by CODEX between self-reported sex (female n=35 patients, male n=73 patients), HCC etiologies (NASH n=28 patients, ALD n=11 patients, HepC n=35 patients, and HepB n=21 patients), AJCC Stage (I n=58 vs II-III n=50 patients), and grade (1 n=40, 2 n=57, and 3 n=9 patients). e. CODEX image representations comparing exhausted CD8+ T cells, endothelial cell, and neutrophil presence in NASH HCC compared to HCV-HCC. f. Boxplot comparisons of the proportions of exhausted CD8+ T cells, endothelial cells, and neutrophils in NASH HCC (n=28) vs non-NASH HCC (n=80). Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. Two-tailed unpaired t-test was used to compare the groups. Abbreviations: UMAP- uniform manifold approximation and projection for dimension reduction, CODEX- co-detection by indexing, HCC- hepatocellular carcinoma, CD- cluster of differentiation, CK- cytokeratin, EPCAM- epithelial cell adhesion molecule, BCL2- B cell lymphoma 2, NASH-nonalcoholic steatohepatitis, ALD- alcoholic liver disease, HepC- hepatitis C, HCV- hepatitis C virus, HepB- hepatitis B, HLA-DR - major histocompatibility complex II cell surface receptor, PDL1- programmed cell death ligand 1, DAPI- 4′,6-diamidino-2-phenylindole, PanCK- pan cytokeratin.
Extended Data Fig. 2 Comparison of immune cell distribution in residual HCC.
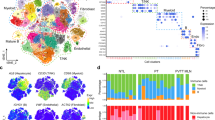
a. Stacked Bar graph containing relatively similar proportions of immune cell populations in HCC as reported by CODEX analysis in this study, the Cancer Genome Atlas Project, the ICGC project and a single cell RNA sequencing research study PMID 35355983. b. Boxplot comparisons and Voronoi plot of representative cores of NK cell proportions in primary (n=53 patients) vs residual HCCs (n=55 patients). Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. Two-tailed unpaired t-test was used to compare the groups. c. Boxplot comparisons and Voronoi plot of representative cores of neutrophil proportions in primary (n=53 patients) vs residual HCCs (n=55 patients). Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. Two-tailed unpaired t-test was used to compare the groups. d. Volcano plot showing key tumor and immune cell populations (with statistical significance) in primary vs residual HCCs in the subgroup of HCCs arising in the cirrhotic liver (n=96 patients). Two-tailed unpaired t-test was used to compare the groups. e. Boxplot comparisons of PDL1+ macrophages and exhausted CD8 T cells between peritumoral TACE-exposed cirrhotic livers (n=3 patients), primary (n=53 patients) vs residual HCCs (n=55 patients). Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. Two-tailed unpaired t-test was used to compare the groups. f. Boxplot comparisons of PDL1+ macrophages and exhausted CD8 T cells between primary (n=53 patients), residual HCC- not refractory to TACE (n=21 patients) vs residual HCC refractory to TACE (n=24 patients). Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. Two-tailed unpaired t-test was used to compare the groups. g. Stacked bar chart comparing relative proportions of all 20 immune and tumor cell subtypes identified by CODEX between primary (n=53 patients), residual HCC- not refractory to TACE (n=21 patients) vs residual HCC refractory to TACE (n=24 patients). h. H&E and Immunofluorescence for PDL1+ macrophages in hepatic tissue with complete response to TACE (n=5 patients) compared to residual HCC (n=5 patients). Bar plots compare the proportion of PDL1+ cells between the two groups. Data are presented as mean values +/− SEM. Two-tailed unpaired t-test was used to compare the groups. i. Stacked bar chart comparing relative proportions of all 20 immune and tumor cell subtypes identified by CODEX between conventional TACE (cTACE, n=6 patients) and doxorubicin-eluting beads TACE (DEB-TACE, n=49 patients). Abbreviations: HCC- hepatocellular carcinoma, CODEX- co-detection by indexing, RNA-Ribonucleic acid, NK- natural killer, CD- cluster of differentiation, PDL1- programmed cell death ligand 1, TACE-transarterial chemoembolization, H&E-hematoxylin and eosin, cTACE- conventional transarterial chemoembolization, DEB-TACE- doxorubicin-eluting beads transarterial chemoembolization.
Extended Data Fig. 3 Interactions of tumor and immune cells in residual HCC.
a. Heatmap showing different patterns of direct interactions between the four tumor cell subtypes in primary (n=53 patients) vs residual HCC (n=55 patients). Two-tailed unpaired t-test was used to compare the groups. * Indicates pAdj value<0.05. b. Dot plot showing mean frequency of interaction between tumor cell subtypes and PDL1+ macrophages in primary and residual HCC stratified by median frequency of PDL1+ macrophages in tumor. Two-tailed unpaired t-test was used to compare the groups. * Indicates pAdj value<0.05. c. Kaplan Meir curve showing survival analysis of human HCC TCGA cohort (n=372 patients) classified based on expression of genes related to M2-like macrophages and cancer stem cells. Log rank test used. d. IF analysis of human HCC cell lines Huh7 co-cultured in 3D tumoroids with THP1 macrophages polarized to M1-like (n=6 tumoroids) or M2-like (n=8 tumoroids) macrophages. Two-tailed unpaired t-test was used to compare the groups. Bar plot shows quantification of CK19 expression on the 3D tumoroids. Data are presented as mean values +/− SEM. e. Heatmap of direct interactions between immune cells identified by CODEX in primary (n=53 patients) and residual HCC (n=55 patients). f. Large neighborhood sizes demonstrate reduced, but still significant spatial autocorrelation for Ki67 and BCL2 markers. Spatial autocorrelation is calculated using Geary’s C statistic. The spatial autocorrelation for each marker is calculated independently for all tumor regions (n=1.07 million cells) and reported as C′ = 1 - C. The maximal spatial autocorrelation possible is 1. Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. g. Alluvial plot of statistically significant indirect tumor cell subtype-immune cell interactions in primary (n=53 patients) vs residual HCCs (n=55 patients). h. Heatmap showing different patterns of indirect (25-100um) interactions of tumor cell subtypes with immune cells in primary (n=53 patients) vs residual HCC (n=55 patients). Two-tailed unpaired t-test was used to compare the groups. * Indicates pAdj value<0.05. Abbreviations: HCC- hepatocellular carcinoma, PDL1- programmed cell death ligand 1, TCGA- the cancer genome atlas project, M2- Type 2 Macrophage, IF- Immunofluorescence, 3D- 3 Dimensional, THP1-, M1- Type 1 Macrophage, CK- cytokeratin, CODEX- co-detection by indexing, CD- cluster of differentiation, EPCAM- epithelial cell adhesion molecule, HLA-DR - major histocompatibility complex II cell surface receptor, NK- natural killer, Eff- Effector, Exh- Exhausted, Mem- Memory, DC- Dendritic cell, EC-, Macro- Macrophage, CSC- Cancer Stem cell, Mac- Macrophage, BCL2- B cell lymphoma 2.
Extended Data Fig. 4 Cellular neighborhood analysis of residual HCC.
a. Cross-matching of Voronoi plots showing spatial neighborhoods and the associated H&E and CODEX immunofluorescence staining images representations of the same tumor core. The canonical markers of key cells represented in each neighborhood are shown. b. Stacked bar chart comparing relative proportions of all 9 cellular neighborhoods identified by CODEX by HCC AJCC Stage (I n=58 vs II-III n=50 patients), grade (1 n=40, 2 n=57, and 3 n=9 patients), and etiologies (NASH n=28 patients, ALD n=11 patients, HepC n=35 patients, and HepB n=21 patients). c. Boxplot comparisons of CK19, EpCAM, CD3, aSMA, CD15, and CD68 expression amongst the identified cellular neighborhoods (CK19+Tumor CN=33708 cells, EpCAM+ Tumor CN=28190, Fibroinflammatory immune CN= 140,273, Innate immune CN= 53,912, M2-macrophage Immune CN= 188,407, Other= 6,130, T cell Immune CN=51,987, Pauci-immune tumor CN=310,977, Vascular inflammatory Tumor CN=251,581). Box: 25–75 percentile, whiskers: 5–95 percentile, line: median. d. Comparison of proportion of EpCAM+ tumor cells and exhausted CD8 T cells in residual HCC stratified by median frequency of M2-macrophage CN (Low n=17; High n=38 patients). e. Comparison of specific canonical marker expression on macrophage and T cells in the EpCAM+ tumor CN (macrophages n=272 cells, T cells n=13 cells), CK19+ tumor CN (macrophages n=1201 cells, T cells n=354 cells) compared to vascular inflammatory tumor CN (macrophages n=16143 cells, T cells n=1275 cells) in residual HCC. Abbreviations: H&E-hematoxylin and eosin, HCC- hepatocellular carcinoma, CODEX- co-detection by indexing, NASH-nonalcoholic steatohepatitis, ALD- alcoholic liver disease, HepC- hepatitis C, HepB- hepatitis B, CK- cytokeratin, CD- cluster of differentiation, EPCAM- epithelial cell adhesion molecule, aSMA- alpha smooth muscle actin, CN- Cellular neighborhood, PD1- programmed cell death protein 1, TIM3- T cell immunoglobulin and mucin domain-containing protein 3, Macro- macrophages, DAPI-4′,6-diamidino-2-phenylindole, PanCK- Pan-cytokeratin, ECad- E cadherin.
Extended Data Fig. 5 Pathway analysis of Residual tumor cells and Macrophages.
a. Characterizing the HCC cancer cell clusters C0 cluster between doxorubicin-resistant (DoxR) (n=4320 cells) and control samples (n=3627 cells). Gene set expression analysis shows enrichment of HCC progenitor signature and doxorubicin resistance signature in the cancer cells of the DoxR samples. b. Dot plot shows mean expression of key differentially expressed genes in C0 and C5 clusters between control (C0 n=3627 cells; C5 n=161 cells) and DoxR (C0 n=4320 cells; C5 n=57 cells) samples. c. Flow cytometry of monocyte-derived macrophages (MoM) from HCC peripheral blood mononuclear cells (PBMCs) were cultured for 48 hours with conditioned media from patient-derived organoid (PDO) which were either treated with doxorubicin (MoM+ DoxoR PDO CM; n=11,753 cells) or control (MoM+ ctrl PDO CM; n=2089 cells), d. Flow cytometry of monocyte-derived macrophages (MoM) from HCC peripheral blood mononuclear cells (PBMCs) were treated with doxorubicin-containing media (MoM+Doxorubicin, n=7559 cells) or doxorubicin-free control media (MoM+ctrl media, n=15,546 cells). This experiment was repeated using MoM from patients with HCC which were then treated with doxorubicin (n=3 patients) or control (n=3 patients). Two-tailed unpaired t-test used to compare the proportion of PDL1+ MoM on flow cytometry. e. Recurrence-free survival predicted by the macrophage and tumor cell signatures derived from the spatial transcriptomic analysis. Kaplan Meir analysis with log rank test was performed (n=372 patients, unique biological samples). f. TGFBR1/2 pathway displayed as an upstream regulator of the network of transcriptional changes in the tumor cell AOI and the corresponding ligand TGFB1 in the macrophage AOI of residual HCC. Upstream analysis in Ingenuity Pathway Analysis was performed using a two-tailed Fisher’s Exact Test to identify likely upstream regulators based on differential gene expression data. Abbreviations: HCC- hepatocellular carcinoma; DoxR- Doxorubicin-Resistant; PDO- Patient-derived organoid; CM- Conditioned media; Ctrl- Control; MoM- Monocyte-derived macrophage; PBMCs- Peripheral blood mononuclear cells, PDL1- programmed cell death ligand 1, TGFB1 transforming growth factor beta 1; Res- Residual; Tum- Tumor; HR- hazard ratio. AOI- Area of interest.
Extended Data Fig. 6 Translational relevance of MRD model of MYC-HCC and MT-HCC.
a. Representative H&E images and IHC for HCC-specific proteins glutamine synthetase, HNF4A and stem cell marker CK19. This was performed on multiple tumor samples (MYC-HCC=5, MT-HCC=5 mice). b. Expression of HCC-specific genes in primary (MYC-HCC=5, MT-HCC=5 mice) compared to WT FVB mouse liver (n=2 mice). Data are presented as mean values +/− SEM. c. Heatmap shows similarity between MYC-HCC=5, MT-HCC=5 and five human HCC transcriptome datasets including the TCGA cohort; the top 5 upregulated and downregulated genes across the different datasets are shown d. Representative H&E and IHC images showing phospho histone 3 (pH3) and cleaved caspase 3 (cC3) staining in primary HCC, minimal residual disease (MRD), and recurrent HCC in MYC-HCC and MYC-Twist1 HCC livers. Bar plots comparing quantification of phospho histone 3, and cleaved caspase 3, in primary tumors, MRD, and recurrent tumors in the liver of MYC-HCC. This was repeated n=5 mice in each group. Data are presented as mean values +/− SEM. e. Experimental scheme to induce NASH in MT-HCC. Gross liver images, H&E, and trichrome staining demonstrating induction of NASH, fibrosis, and HCC in this model. Experiments were performed on 3–5 animals in each group. f. Confirming induction of NASH by demonstrating hepatic inflammation, obesity, hyperlipidemia with high-fat diet (HFD) (c1 chow=5, HFD=3; c2-5 chow=4; HFD=4 mice). Data are presented as mean values +/− SEM. Two-sided unpaired t-tests were used for comparison. g. Confirming reversible residual HCC upon oncogene inactivation in MT-HCC with diet-induced NASH. Experiments were performed on 5 mice on HFD diet. h. Representative H&E and IHC images showing persistence of CD133+ stem-like cancer cells in perivascular niches of oncogene inactivated MRD in the liver, lungs and subcutaneous spaces. Experiments were performed on 5 mice in each group. i. Representative H&E and IHC images showing persistence of CK19+ stem-like cancer cells in perivascular niches of MRD in the subcutaneous spaces. Bar plots comparing quantification of CK19 and CD133 in primary tumors, MRD, and recurrent tumors in the liver of MYC-HCC. Experiments were performed on 5 mice in each group. Data are presented as mean values +/− SEM Abbreviations: H&E-hematoxylin and eosin, IHC- Immunohistochemistry, HCC- hepatocellular carcinoma, HNF4A- Hepatocyte nuclear factor-4 alpha, CK- cytokeratin, MYC-HCC- MYC-driven hepatocellular carcinoma, MT-HCC- MYC/Twist hepatocellular carcinoma, WT FVB- wild-type FVB, TCGA- the cancer genome atlas project, pH3- phospho histone 3, cC3- cleaved caspase 3, MRD- Minimal residual disease, NASH Nonalcoholic steatohepatitis, HFD- High-fat diet, ALT-Alanine aminotransferase, LDL- Low density lipoprotein, CD- cluster of differentiation.
Extended Data Fig. 7 Residual tumor cells demonstrate stemness and activation of Tgfβ pathway.
a. Experimental scheme for single cell sequencing on MYC inactivation residual tumor cells (n=4367 cells). b. Two major clusters upon MYC inactivation, stem cell-like cluster (n=1184 cells) and non-stem-like cluster (n=3183 cells). c. Heatmap shows differentially expressed genes between the stem cell-like cluster and non-stem-like clusters. d. Activation of Tgfβ1 pathway in the stem cell-like cluster of residual tumor cells. e. Top molecular pathways activated in the stem cell-like cluster. Abbreviations: TGFB1 transforming growth factor beta 1, HCC- hepatocellular carcinoma, scRNA-seq- single cell RNA sequencing.
Extended Data Fig. 8 Interaction of stem-like cancer cells and macrophages in MRD.
a. Comparison of macrophages in the spatial proximity of MRD (n=6 mice) versus areas without MRD (n=6 mice) in the lungs of MYC/Twist1 mice. Data are presented as mean values +/− SEM. b. Representative images demonstrating spatial interactions of stem-like cancer cells and macrophages in the spatial proximity of MRD in the lungs and liver of MYC/Twist1 mice. c. Comparison of immune cell subsets between livers without MRD ctrl Liver (n=4 mice) and livers with MRD (n=4 mice). Bar plots compared using two-tailed unpaired t-tests. Data are presented as mean values +/− SEM. d. Comparison of CD8 T cell subsets between livers without MRD ctrl Liver (n=4 mice) and livers with MRD (n=4 mice). Bar plots compared using two-tailed unpaired t-tests. Data are presented as mean values +/− SEM. Abbreviations: MRD- Minimal residual disease, ctrl- control, CD- cluster of differentiation, H&E-hematoxylin and eosin, DAPI-4′,6-diamidino-2-phenylindole, HCC- hepatocellular carcinoma.
Extended Data Fig. 9 Combined blockade of Tgfβr1 and Pdl1 in mouse MRD HCC.
a. Experimental scheme for treatment of oncogene-deprived subcutaneous MRD-bearing mice with control antibody or combination therapy with Tgfbr1 and Pdl1 inhibitors. Treatment is followed by oncogene reactivation to induce tumor recurrence. Kaplan–Meier analysis with log rank test was performed. b. Kaplan–Meier curves show time to recurrence in oncogene-deprived subcutaneous MRD-bearing mice with control antibody (n=4 mice) or combination therapy with Tgfbr1 and Pdl1 inhibitors (n=5 mice). c. Experimental scheme for treatment of oncogene-deprived subcutaneous MRD-bearing mice with control antibody or combination therapy with TgfbrI and anti-Pdl1 inhibitor. Residual tumor niches are then evaluated at the end of treatment. d. Macroscopic and microscopic evaluation of subcutaneous MRD sites shows elimination of residual tumor cells in mice treated with combination therapy with TgfbrI and Pdl1 inhibitors (n=4 mice) than control (n=4 mice). e. Experimental scheme for treatment of oncogene-activated primary MT-HCC with control antibody or combination therapy with TgfbrI and anti-Pdl1 inhibitor. Liver tumor burden is assessed at the end of treatment. f. Quantification of liver tumor burden, gross images, and H&E images of primary MT-HCC treated with control antibody (n=4 mice) or combination therapy with TgfbrI and anti-Pdl1 inhibitor (n=4 mice). Data are presented as mean values +/− SEM. Two-tailed unpaired t-test was used to compare the groups. g. Apoptosis measured by cleaved caspase 3+ cells in liver MRD of MYC-HCC mice treated with control (n=5 mice) versus Tgfbr1 (n=5 mice) or Pdl1 inhibitors (n=5 mice) or combination therapy with Tgfbr1 and Pdl1 inhibitors (n=5 mice). Data are presented as mean values +/− SEM. Two-tailed unpaired t-test was used to compare the groups. h. Flow cytometry quantification of activated CD4 and CD8 T cells which are CD69+/CD44high in MRD of mice treated with control antibody (n=3 mice) or combination therapy with Tgfbr1 and Pdl1 inhibitors (n=3 mice). Bar plots compare the mean between the groups with unpaired t-tests. Data are presented as mean values +/− SEM. Two-tailed unpaired t-test was used to compare the groups. Abbreviations: HCC- hepatocellular carcinoma, PDL1- programmed cell death ligand 1, TGFBr1 transforming growth factor receptor beta 1, H&E-hematoxylin and eosin, Ctrl-control, MRD- Minimal residual disease, MT-HCC- MYC/Twist hepatocellular carcinoma, CD- cluster of differentiation.
Extended Data Fig. 10 Combined blockade of Tgfβr1 and Pdl1 eliminates doxorubicin-resistant mouse HCC.
a. Schematic showing the establishment of mouse model of control (n=5 mice) or doxorubicin-resistant syngeneic orthotopic allografts (n=11 mice) which were then treated with control antibody (n=6 mice) or combination therapy with TgfbrI and anti-Pdl1 inhibitor (n=10 mice). b.Representative gross images, H&E and IHC images of control or doxorubicin-resistant orthotopic HCC-bearing mice treated with control antibody (n=6 mice) or combination therapy with TgfbrI and anti-Pdl1 inhibitor (n=10 mice). IHC for CD8 T cells shows in the bottom panel. c. Quantification of tumor burden and CD8T cells in the liver of control or doxorubicin-resistant orthotopic HCC-bearing mice treated with control antibody (n=6 mice) or combination therapy with TgfbrI and anti-Pdl1 inhibitor (n=10 mice). Bar plots compare the mean between the groups with two-sided unpaired t-tests. Data are presented as mean values +/− SEM. d. Flow cytometry-based quantification of tumor-infiltrating leukocytes, T cells, M2-like (PDL1+ or CD206+) and M1-like macrophage (CD86+ or MHCII+) subsets in doxorubicin-resistant orthotopic HCC-bearing mice treated with control antibody (n=4 mice) or combination therapy with TgfbrI and anti-Pdl1 inhibitor (n=4 mice). Bar plots compare the mean between the groups two-sided with unpaired t-tests. Data are presented as mean values +/− SEM. e. Representative flow cytometry images and quantification of M1-like and M2-like macrophages within the tumor in the doxorubicin-resistant orthotopic HCC-bearing mice treated with control antibody or combination therapy with TgfbrI and anti-Pdl1 inhibitor. f. Hep 53.4 HCC cell lines are treated with control or doxorubicin for 96 hours and viable cells are selected for orthotopic implantation into mouse liver, confirmed by MRI. g. Flow cytometry analysis shows no difference in leukocyte or T cell or macrophage infiltration in WT mice bearing control HCC treated with either control antibody (n=2 mice) of combined inhibition of Tgfbr1 and anti-Pdl1 antibody (n=3 mice). Data are presented as mean values +/− SEM. h. Schematic summarizing the main findings- i. The spatial organization of post-TACE residual HCC is unveiled through integrated analysis of single-cell spatial profiling employing CODEX and GeoMx spatial transcriptomics of human HCC. ii. Mouse model of minimal residual disease (MRD) reveals that TGFβ1 derived from PDL1+ macrophages enable the persistence of residual stem-like tumor cells and induces exhaustion in CD8T cells. iii. In two mouse models of MRD, we target the TGFβ pathway and PDL1, eliminating residual tumor cells and preventing HCC recurrence. Abbreviations: HCC- hepatocellular carcinoma, Tgfb -transforming growth factor beta, H&E-hematoxylin and eosin, Ctrl-control, Res-residual, MRD- Minimal residual disease, DoxR- doxorubicin-resistant.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–5.
Supplementary Tables 1–9 (download XLSX )
Supplementary Table 1. List of markers used to characterize the HCC tumor cell microenvironment for CODEX analysis. Supplementary Table 2. List of the 20 cell types identified by CODEX analysis. Supplementary Table 3. Demographic and clinicopathological characteristics of patients included in this study, subdivided into those with primary versus residual HCC between these groups. Supplementary Table 4. Demographic and clinicopathological characteristics of patients included in NanoString spatial transcriptomics study, subdivided into those with primary versus residual HCC. Supplementary Table 5. Differentially expressed genes in the epithelial compartment of the residual HCC versus matched liver samples. Supplementary Table 6. Differentially expressed genes in the macrophage compartment of the residual HCC versus matched liver samples. Supplementary Table 7. Differentially expressed genes in the residual tumor cell cluster versus the two hepatocyte clusters of scRNA-seq in vivo mouse MRD. Supplementary Table 8. Differentially expressed genes in the M2-like macrophage cluster versus the two other macrophage clusters of scRNA-seq in vivo mouse MRD. Supplementary Table 9. Antibodies used in multiple assays in the study.
Source data
Source Data Figs. (download XLSX )
Source data for all main figs., where each panel is presented in a single tab with the tab name corresponding to the panel label.
Source Data Extended Data Figs. (download XLSX )
Source data for all extended data figs., where each is presented in a single tab with the tab name corresponding to the figure number.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lemaitre, L., Adeniji, N., Suresh, A. et al. Spatial analysis reveals targetable macrophage-mediated mechanisms of immune evasion in hepatocellular carcinoma minimal residual disease. Nat Cancer 5, 1534–1556 (2024). https://doi.org/10.1038/s43018-024-00828-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-024-00828-8
This article is cited by
-
Cellular neighborhoods in cancer
Nature Cancer (2026)
-
Adjuvant cytokine-induced killer cell immunotherapy in hepatocellular carcinoma: real-world data and 9-year extended follow-up of a randomized controlled trial
Cancer Immunology, Immunotherapy (2026)
-
Single-cell multi-omics in cancer immunotherapy: from tumor heterogeneity to personalized precision treatment
Molecular Cancer (2025)
-
Cancer stem cells and niches: challenges in immunotherapy resistance
Molecular Cancer (2025)
-
Targeting tumor-associated macrophages to overcome immune checkpoint inhibitor resistance in hepatocellular carcinoma
Journal of Experimental & Clinical Cancer Research (2025)